Abstract
The sensation of pungency generated by capsaicinoids is a characteristic trait of chili peppers (Capsicum spp.), and the presence or absence of pungency is central in determining its usage as a spice or a vegetable. In the present study, we aimed to clarify the heredity and genetic factors involved in the deficiency of pungency (quite low pungency) that is uniquely observed in the Japanese chili pepper ‘Shishito’ (Capsicum annuum). First, the F2 population (‘Shishito’ × pungent variety ‘Takanotsume’) was used for segregation analysis, and pungency level was investigated using capsaicinoid quantification with high-performance liquid chromatography. Also, restriction site associated DNA sequencing of the F2 population was performed, and genetic map construction and quantitative trait locus (QTL) mapping were implemented. The results indicated that the F2 population showed varying capsaicinoid content and two major QTLs were detected, Shql3 and Shql7, which explained 39.8 and 19.7% of the genetic variance, respectively. According to these results, the quite low pungency of ‘Shishito’ was a quantitative trait that involved at least the two loci. Further, this trait was completely separate from general non-pungent traits controlled by individual recessive genes, as described in previous studies. The present study is the first report to investigate the genetic mechanism of pungency deficiency in Japanese chili peppers, and our results provide new insights into the genetic regulation of pungency in chili pepper.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chili peppers (Capsicum spp.) are members of the Solanaceae family and are characterized by their pungent taste. The sensation of pungency is derived from chemical constituents known as capsaicinoids (Suzuki and Iwai, 1984), and their amounts determine the pungency levels of chili pepper fruits and whether they are utilized as a spice or vegetable. Capsaicinoids are indispensable ingredients in food cultures around the world, e.g., as a spice, and recently their functional health effects, e.g., anti-inflammatory, antioxidant, antitumor, and weight-loss properties, have been a subject of focus (Naves et al. 2019). In chili pepper fruits, capsaicinoids are mostly biosynthesized in the epidermal cells of the placental septum (Fujiwake et al. 1982) and their biochemical and genetic mechanisms have been investigated. First, capsaicinoid biosynthesis has been shown to consist of two intermediate pathways using radiotracer studies (Bennett and Kirby 1968). One is the phenylpropanoid pathway that synthesizes vanillylamine from phenylalanine, and the other is the branched-chain fatty acid pathway that synthesizes a series of branched-chain fatty acid moieties from valine or leucine. Capsaicinoids are eventually generated by the condensation of vanillylamine and branched-chain fatty acids. In investigating the genetics of capsaicinoids, several genome and transcriptome analyses have profiled a host of genes involved in capsaicinoid biosynthesis (Kim et al. 2014; Martínez-López et al. 2014; Qin et al. 2014; Zhang et al. 2016). In particular, several genes have been identified as necessary in generating capsaicinoids. First, Pun1, which is located on chromosome 2, is the most critical gene for synthesis. This gene was once known as the C locus controlling the presence of pungency (Deshpande 1935), and the responsible gene was identified by Stewart et al. (2005). Pun1 encodes an acyltransferase that is responsible for the final reaction of capsaicinoid biosynthesis, and mutation of the Pun1 gene results in complete deficiency of pungency (Kirii et al. 2017; Stellari et al. 2010; Stewart et al. 2007). Moreover, putative aminotransferase (pAMT) is also critical as it is responsible for synthesizing vanillylamine from vanillin in the phenylpropanoid pathway (Lang et al. 2009). pAMT is located on chromosome 3 and its various gene mutations are known to cause a drastic reduction of pungency (Tanaka et al. 2019). In addition, Koeda et al. (2019) recently discovered a novel gene called ketoacyl-ACP reductase (CaKR1) located on chromosome 6. CaKR1 is involved in branched-chain fatty acid elongation, and also is necessary for generating capsaicinoids in chili peppers. Besides these biosynthesis genes, Arce-Rodríguez and Ochoa-Alejo (2017) initially proposed that the R2R3-MYB transcription factor encoded by CaMYB31 significantly affected the expression of capsaicinoid biosynthesis genes. Around the same time, Han et al. (2019) identified the locus (chromosome 7) and dysfunctional allele of CaMYB31 (Pun3) that resulted in deficiency of pungency.
As mentioned above, Pun1, pAMT, CaKR1, and CaMYB31 are qualitative genes responsible for the presence of pungency in chili peppers. Therefore, it is suggested that the mechanism of pungency deficiency in most chili peppers could involve the dysfunction of these four genes; however, several Japanese chili peppers could be exceptions. In Japan, there are several quite low pungency varieties of C. annuum, such as ‘Shishito’, ‘Manganji’ and ‘Fushimi amanaga’, which are utilized as vegetables. Although these varieties appear to be the same as typical non-pungent sweet chili peppers, they are distinguished by the occurrence of pungency. Specifically, capsaicinoids are never or hardly synthesized in general non-pungent varieties due to the dysfunctional genes, and the occurrence of pungency is rarely observed. Conversely, Japanese non-pungent varieties are known to become pungent depending on environmental factors, and the presence of these pungent fruits are known to cause problems in cultivation and distribution in Japan (Ishikawa et al. 2004; Minamiyama et al. 2012; Murakami et al., 2006). Our research group investigated this phenomenon using the variety ‘Shishito’ and that revealed parthenocarpy, lack of seeds fruits, was related to the occurrence of pungency (Kondo et al. 2021a, b). We also observed that several capsaicinoid biosynthesis genes, including Pun1, pAMT, CaKR1, and CaMYB31, are hardly expressed in non-pungent ‘Shishito’ fruits but highly expressed in pungent fruits. This implied that the loss of pungency in ‘Shishito’ was not dependent on dysfunctional gene mutation in the capsaicinoid biosynthesis pathways described above. Therefore, these Japanese varieties presumably repress their pungency according to unique mechanisms. In an effort to clarify these mechanisms, we focused on the quite low pungency trait observed in ‘Shishito’ and investigated the heredity and genetic factors associated with pungency. In the present study, we prepared a F2 population (‘Shishito’ × pungent variety ‘Takanotsume’), and the pungency level was investigated by quantifying the capsaicinoid content. Next, we implemented quantitative trait locus (QTL) mapping involving pungency level using the F2 population, and investigated the candidate genetic loci responsible for pungency deficiency in ‘Shishito’.
Materials and methods
Plant materials
For the segregation analysis of a low pungency in ‘Shishito’ (C. annuum), we constructed F1 progeny by crossing ‘Shishito’ and the pungent C. annuum variety ‘Takanotsume’, with the F2 progenies being obtained by self-pollination. Plants were field-cultivated in 2020 and 2021 at a farm at Shinshu University (Nagano, Japan). In each year, ten F1 individuals and 160 F2 individuals were cultivated. Further, two kinds of F1 progeny were generated by crossing ‘Shishito’ and two non-pungent cultivars: ‘Shishito’ × ‘California Wonder’ (C. annuum), which is a completely non-pungent cultivar owing to the dysfunctional Pun1 allele (pun11), and ‘Shishito’ × ‘Himo’ (C. annuum), which exhibits quite low pungency attributable to a dysfunctional pAMT allele (pamt2). Ten F1 plants were cultivated for each cross in 2020 only.
Capsaicinoid extraction and quantification
For phenotyping of pungency traits in several progenies, we performed high-performance liquid chromatography (HPLC) to quantify capsaicinoid content in the placental septum. To normalize the stage of fruit development, we referenced the color of the fruit surface and only collected fruits in which the color began to change from deep green to brown. Capsaicinoids were extracted from grouped placental septa of ten fruits per plant. In the capsaicinoid extraction, the placental septa were initially lyophilized using a freeze-drier (FDU-200, EYELA, Tokyo, Japan). The capsaicinoids were then extracted from 200 mg of the powdered dried tissue using acetone, and the contents were quantified by HPLC as described by Kondo et al. (2021a). In the present study, capsaicinoids were defined as the total contents of capsaicin and dihydrocapsaicin, then the capsaicinoid content per unit dry weight of placental septa (µg • gDW−1) was calculated.
RAD-seq and genotyping of the F2 population
Genetic analysis was conducted by restriction site associated DNA sequencing (RAD-seq) using 104 F2 individuals that were randomly selected from a total of 160 F2 progenies in 2020. As for their two parents, three plants were used for RAD-seq respectively. gDNA was extracted from young leaves of individual plants using a DNeasy Plant Pro Kit (Qiagen, Hilden, Germany). Library construction for Next Generation Sequencing (NGS) was performed by digesting gDNA with EcoRIII and EcoRI and subsequently ligating adaptors with different barcodes. On average, 400 bp DNA fragments were obtained and pair-end sequencing was performed using a HiseqX (Illumina, San Diego, CA, USA). After sequencing, adaptor trimming and quality filtering were conducted using Fastp v0.20.0 (Chen et al. 2018), and sequence reads with quality values less than Q15 were trimmed. Burrows-Wheeler Alignment (BWA v0.7.8; Li and Durbin 2009) was used to map the trimmed sequence reads to the C. annuum genome of ‘Zunla 1’ (Ref_v1.0), and variant call was subsequently performed using the Genome Analysis Toolkit (GATK v4.1.2.0; McKenna et al. 2010). The raw genotyped data were then filtered according to the following criteria: QD < 2.0, QUAL < 30.0, SOR > 4.0, FS > 60.0, MG < 40.0, MQ < 40.0, MQRankSum < − 12.5, ReadPosRankSum < − 8.0. Finally, we obtained Single Nucleotide Polymorphism (SNP) data in a Variant Calling Format (VCF) file. To determine the genotype of the F2 progenies, we screened parents-endemic SNPs from the VCF files of the two parents using TASSEL 5 (Bradbury et al. 2007), then the SNPs genotyped in less than 70% of F2 progenies (73 individuals) were removed. Consequently, a total of 17,085 SNPs were obtained that distinguished the genotype of the parents.
Genetic map construction and QTL mapping involved in pungency level
QTL mapping of pungency level factors was performed by constructing a linkage map using the R package ‘qtl’ (Broman et al. 2003). Initially, the segregation suitability of each SNP in the F2 population was investigated using the chi-square test, and SNPs in which segregations corresponded to the expected ratio (1:2:1, p > 0.01) were identified. A genetic map was then constructed using the Kosambi function according to the following criteria: maximum recombination fraction of 0.35, and threshold of LOD score of 3.0. Subsequently, QTL mapping of capsaicinoid content (µg • gDW−1) was also conducted with the R package ‘qtl’. Mapping was performed using the logarithm of the odds (LOD) score calculated by composite interval mapping (CIM), and the threshold was determined by 1000 permutation tests at a significance level of p < 0.05. We detected QTLs based on the threshold and estimated the additive effect, dominant effect, and genotypic variance explained (PVG) for each QTL. Furthermore, the physical positions of detected QTLs were compared with those of known QTLs and capsaicinoid biosynthesis genes. The physical positions of the detected QTLs in the ‘Zunla 1’ genome were converted to those in the ‘CM334’ genome (Pepper.v.1.55) using the genome browser of the Solanaceae Genomics Network (https://solgenomics.net) and the physical position data of capsaicinoid biosynthesis genes and known QTLs in the ‘CM334’ genome which was integrated by Han et al. (2018), was referenced. Finally, we merged the physical position data of the known QTLs and the QTLs detected herein; the merged data are listed in Tables S1, S2.
Verification of genotypic effects in the vicinity of candidate QTLs
In the present study, two high LOD peaks were observed on chromosomes 3 and 7, and we designed a Cleaved Amplified Polymorphic Sequences (CAPS) marker and Amplification Refractory Mutation System (ARMS) markers adjacent to the two loci, respectively. Details of the marker design method are described in Figs. S1, S2, and the primer sequences employed are listed in Table S3. Using these two markers, the genotypes of the F2 progenies were investigated. In 2020, 56 F2 individuals that were not used for RAD-seq were investigated, and 160 plants cultivated in 2021 were subsequently investigated.
Genomic variant analysis in several capsaicinoid biosynthesis genes
We targeted seven capsaicinoid biosynthesis genes (ketoacyl-acyl carrier protein synthase III (KASIII), 4-coumaroyl coenzyme A ligase (4CL), caffeoyl shikimate esterase (CSE), hydroxycinnamoyl transferase (HCT), CaMYB31, Pun1, and pAMT) located mainly around the two detected QTLs, and explored the gene mutations in their isoforms reported in previous studies (Table S2). First, the gDNAs of ‘Shishito’ and ‘Takanotsume’ were subjected to whole-genome resequencing by a sequencing service (Macrogen Japan, Tokyo, Japan); on average, 350 bp DNA fragments were applied for pair-end sequencing using a NovaSeq 6000 (Illumina, San Diego, CA, USA). Sequence reads were subsequently mapped to the ‘Zunla 1’ genome as described in the RAD-seq analysis, and genomic variants were detected by two steps. First, we broadly explored large Indels within a 40 kbp genomic region including the gene isoform at the center using Pindel v0.2.5 (Ye et al. 2009), which can detect structure variants (SVs) based on the mapping files (BAM format files). In the analysis, we detected variants that were supported by at least 4 reads (read coverages). Then, we specified Indels and SNPs in the narrow region ranging from the gene upstream region (2000 bp from the initial point of transcription) to the 3' untranslated region (UTR), using not only Pindel v0.2.5 but also GATK v4.1.2.0 as described in the RAD-seq analysis. Among the detected variants, those harboring polymorphism between ‘Shishito’ and ‘Takanotsume’ were selected. Then, we trimmed the variants that were non-allelic between ‘Shishito’ and ‘Zunla 1’ (pungent variety), as these variants rarely seemed to induce gene mutations that resulted in dysfunction.
Results
Pungency traits in ‘Shishito’ and the various F1 progenies
Capsaicinoid contents in ‘Shishito’ fruits were 218 and 424 µg • gDW−1, while those in ‘Takanotsume’ were 23,108 and 35,005 µg • gDW−1 in 2020 and 2021, respectively (Table 1). While the contents varied among individuals in both cultivars, pungency levels of ‘Shishito’ were at least 30 times lower than ‘Takanotsume’ in both years. The F1 progenies ‘Shishito’ × ‘Takanotsume’ had 10,820 and 13,602 µg • gDW−1 of capsaicinoid contents in 2020 and 2021, respectively, which were in between those of the two parents. In comparison, the F1 progenies ‘Shishito’ × ‘California Wonder’ [pun11/pun11] showed 118 µg • gDW−1 of capsaicinoid content, which was intermediate pungency level relative to the two parents. Conversely, the F1 progenies ‘Shishito’ × ‘Himo’ [pamt2/pamt2] contained 12,541 µg • gDW−1, which largely exceeded their parents.
Pungency traits in the F2 population of ‘Shishito’ × ‘Takanotsume’
In each cultivation year, the F2 population ‘Shishito’ × ‘Takanotsume’ showed varying capsaicinoid contents; contents were in the range of 113–35,211 µg • gDW−1 in 2020 and 476–37,074 µg • gDW−1 in 2021. This indicated that the low pungency trait of ‘Shishito’ was a quantitative trait. Then, multiple peaks were observed in the frequency distribution (Fig. 1). In 2020, three clear peaks were observed at 800, 12,000, and 20,000 µg • gDW−1, whereas the frequency distribution in 2021 showed two peaks at 5950 and 17,850 µg • gDW−1.
QTL mapping related to pungency level
QTL mapping was conducted using 104 F2 individuals (‘Shishito’ × ‘Takanotsume’) cultivated in 2020. A genetic map was constructed using 221 SNPs obtained by RAD-seq, which consisted of 12 linkage groups with a total length of 2,750 cM and the mean distance of the marker intervals was 13.2 cM. QTL mapping detected two significant LOD peaks (p < 0.05) (Fig. 2). One peak showed an LOD of 6.96 within 50.0–71.0 cM on chromosome 3, and the other peak had an LOD of 6.69 within 202.0–225.0 cM on chromosome 7; these major QTLs were named Shql3 and Shql7, respectively (Table 2). The physical position of Shql3 was in the marker interval between ST3.4 and ST3.5 (5,167,786–20,615,780 bp on chromosome 3), while Shql7 was located in the marker interval between ST7.9–ST7.13 (218,675,433–221,842,745 bp on chromosome 7). Assessment of the genetic effects indicated that the additive effect and dominant effect of Shql3 were − 6294 and − 646, while those of Shql7 were − 4155 and 2220, respectively. The genotypic variance explained (GVE) for Shql3 and Shql7 was 39.8 and 19.7%, respectively. This implied that the ‘Shishito’ alleles of Shql3 and Shql7 were two of all genetic factors related to the low pungency trait of ‘Shishito’.
Logarithm of the odds (LOD) scores calculated by the quantitative trait locus mapping related to capsaicinoid content in the F2 population in 2020 (n = 104). The black line indicates LOD scores in the composite interval mapping, and the black dotted line is the threshold determined by 1000 permutation tests at a significance level of p < 0.05
In proximity to the Shql3 and Shql7 loci, we investigated known QTLs and capsaicinoid biosynthesis genes involved in the amount of capsaicin, dihydrocapsaicin, nordihydrocapsaicin or total capsaicinoid content. Figure 3a, b shows the physical positions of known QTLs on chromosomes 3 and 7 in the ‘CM334’ genome. Nine QTLs exist along with four capsaicinoid biosynthesis genes in proximity to Shql3 (237–252 Mbp, chromosome 3) (Fig. 3a). Notably, TH-total3.3 (Han et al. 2018), cap3.1 • total3.1 (Ben-Chaim et al. 2006), 4CL, and HCT were within the Shql3 region. Additionally, in proximity to Shql7 (219–231 Mbp, chromosome 7), three QTLs were located along with CaMYB31, which were more than 15 Mbp from that of Shql7 (Fig. 3b).
Physical position of known quantitative trait loci (QTLs) and capsaicinoid biosynthesis genes around Shql3 and Shql7. The physical positions in chromosomes 3 a and 7 b of the ‘CM334’ genome (Pepper.v.1.55). Horizontal lines indicate the positional range of QTLs, and diamonds indicate those of capsaicinoid biosynthesis genes. The complete data are also shown in Tables S1, S2
Effect of genotypes linking Shql3 and Shql7 to pungency level
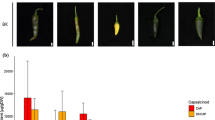
To verify the genotypic effects of Shql3 and Shql7, we determined the genotypes in the vicinity of the two loci and investigated the relationship between these genotypes and capsaicinoid contents in the F2 population. Figure 4a illustrates the capsaicinoid contents in the three genotypes for each locus. Over two cultivation years, the capsaicinoid content in the F2 population changed depending on the genotypes of both loci. Regarding Shql3, the capsaicinoid contents were respectively 17,817 and 18,245 µg • DW−1 in the homozygous allele of ‘Takanotsume’ (TK/TK), while those in the homozygous allele of ‘Shishito’(SH/SH) were 7958 and 10,658 µg • DW−1 in 2020 and 2021. As for Shql7, TK/TK individuals showed 16,003 and 18,556 µg • DW−1, while SH/SH individuals showed 6444 and 11,741 µg • DW−1 in 2020 and 2021, respectively. Integrating the above, the genotypic change from TK/TK to SH/SH in Shql3 resulted in 55.3 and 41.6% reduction of capsaicinoid in the F2 population. Meanwhile, 59.7 and 36.7% reductions were observed in the case of Shql7. Additionally, there was a significant positive correlation (p < 0.001) in the average contents of capsaicinoid between the two years (Fig. 4b); the contents became gradually lower along with the increase in the number of ‘Shishito’ alleles in each locus. This demonstrates that the genotype of Shql3 and Shql7 certainly affected pungency levels regardless of differences in the cultivation environment.
Genotypic effects of Shql3 and Shql7 on the capsaicinoid contents in the F2 population in 2020 and 2021. a Boxplots show capsaicinoid contents in three genotypes of each locus and red circles indicate the mean values. b The scatter plot shows a comparison of the average contents between the two years. TK/TK, TK/SH, and SH/SH denote the homozygous allele of ‘Takanotsume’, heterozygous, and homozygous allele of ‘Shishito’, respectively. In the scatter plot, values indicate the mean ± standard error in each year, and their genotypes are represented by combinations of marker color and symbol. The Pearson’s correlation coefficient is shown as r. ***p < 0.001
We also investigated the relationship between capsaicinoid content and genotypic combinations of the two loci in the F2 population (Fig. 5a). In 2020, we clearly observed genotypic effects of Shql3, i.e., the SH allele resulted in reductions of capsaicinoid contents for each Shql7 genotype. Similarly, possessing the SH/SH genotype of Shql7 also resulted in reductions in all of the Shql3 genotypes. Accordingly, both genotypes seemed to independently change the pungency levels, and extraordinary epistasis was not observed in 2020. A similar tendency was observed in 2021, but the phenotypic changes were sometimes unclear (Fig. 5a). For example, in the TK/TK individuals of Shql7, capsaicinoid contents hardly differed among the three Shql3 genotypes. Moreover, compared to 2020, large differences in the capsaicinoid contents between SH/SH and other Shql7 genotypes were not observed in the TK/TK genotype of Shql3. However, a significant positive correlation (p < 0.001) in the average contents between the two years was observed (Fig. 5b), indicating similarity of their genetic effects in both years. Notably, the F2 individuals possessing both TK/TK genotypes in the two loci showed 21,153 and 20,996 µg • gDW−1 of capsaicinoid in each year, which decreased to 5290 and 8998 µg • gDW−1, respectively, when both loci were replaced with SH/SH genotypes. This corresponded to 75.0 and 57.1% reductions in capsaicinoid contents, respectively, which were large reductions compared to the case of the independent locus alone, as described above.
Effects of the genotypic combination of Shql3 and Shql7 on capsaicinoid contents in the F2 population in 2020 and 2021. a Boxplots show capsaicinoid contents in the nine genotypic combinations, and red circles indicate the mean values. b The scatter plot shows a comparison of the average contents between two years. TK/TK, TK/SH, and SH/SH denote the homozygous allele of ‘Takanotsume’, heterozygous, and homozygous allele of ‘Shishito’ respectively. In the scatter plot, values indicate the mean ± standard error in each year, and their genotypes are represented by combinations of marker color and symbol. r shows Pearson’s correlation coefficients. ***p < 0.001
Genomic variants in several capsaicinoid biosynthesis genes
As another attempt to explore the genetic factors involved in the quite low pungency of ‘Shishito’, we implemented whole-genome resequencing of ‘Shishito’ and ‘Takanotsume’, and explored genomic mutations in known capsaicinoid biosynthesis genes (KASIII, 4CL, CSE, HCT, CaMYB31, Pun1, and pAMT), which, except for Pun1 and pAMT, were located mainly around Shql3 and Shql7. In the whole-genome resequencing, a total of 436,167,062 and 603,425,568 reads were obtained for ‘Shishito’ and ‘Takanotsume’ after filtering, respectively, which were equivalent to approximately 65.8 and 91.0 Gb. Then, genomic variants in ‘Shishito’ were detected within the 40 kbp region, including the gene isoform at the center, which were allelic with both ‘Takanotsume’ and ‘Zunla 1’ (Table 3). At this scale, several large Indels with variant sizes that exceeded 100 bp were observed in ‘Shishito’; however, such Indels were never observed in the narrow genomic region upstream of the gene (− 2000 bp from the transcription start site) to the 3'UTR region (Table 3). When we focused on this region, most of the variants existed upstream of the gene or in introns at modifier levels (Tables S4, S5), which rarely caused mutations in the splice acceptor or donor sites. Then, only nine variants were detected in exons, including seven SNPs and two Indels (Table 4). Among the nine variants, most were in the 3'UTR and 5'UTR or synonymous substitution in exons, which could never affect the coding sequences of amino acids after translation. On the other hand, two significant variants were observed in CSE and pAMT. First, ‘Shishito’ had a 5 bp deletion in the 5'UTR region in one of the CSE isoforms (XM_016710287), which could induce a frameshift mutation. However, another CSE isoform (XM_016710288) never showed such variants. Meanwhile, one SNP at exon 15 of pAMT could cause a non-synonymous substitution in ‘Shishito’.
Discussion
Chili pepper varieties can be broadly defined as either a spice or a vegetable depending on the presence of pungency. Thus, the genetic regulation of pungency has been the focus of many researchers. According to previous studies, the presence of pungency is known as a qualitative trait that is controlled by individual recessive genes; four genes: Pun1, pAMT, CaKR1, and CaMYB31 were discovered as the related genes (Stewart et al. 2005; Lang et al. 2009; Koeda et al. 2019; Han et al. 2019). The recessive alleles could have been used for breeding non-pungent cultivars, i.e., the genetic mechanism of pungency deficiency might be explained by the effect of any of the genes in most sweet cultivars. However, our research groups previously proposed the feasibility of the Japanese variety ‘Shishito’ as an exception, due to the occurrence of highly pungent fruits that were rarely observed in non-pungent cultivars (Kondo et al. 2021a, b). Therefore, we attempted to clarify the genetic regulation of pungency deficiency in ‘Shishito’ by investigating the inheritance and genetic factors of pungency.
In the present study, segregation analysis of pungency traits using F1 and F2 progenies (‘Shishito’ × ‘Takanotsume’) revealed that pungency deficiency in ‘Shishito’ was a quantitative trait. Thus, this trait is completely separate from qualitative traits controlled by recessive single genes such as those described above. This was also assisted by phenotypic analysis in other F1 progenies. The results showed that the ‘Shishito’ × ‘Himo’ F1 progenies exhibited obviously higher capsaicinoid contents than their parents due to genetic complementation, demonstrating that ‘Shishito’ possessed a functional pAMT, although ‘Shishito’ had several genomic variants compared to ‘Takanotsume’ (Tables 3, 4, S4, S5). Further, capsaicinoids were also detected in ‘Shishito’ × ‘California Wonder’ F1 progenies, at a level not exceeding that in ‘Shishito’. Although it could not be determined that ‘Shishito’ possessed a functional Pun1 allele according to this result alone, it is possible that ‘Shishito’ has a normal allele. This is because pun11 of ‘California Wonder’ is known as an allele that results in a complete deficiency of pungency (Stewart et al. 2005). Then, we were unable to detect a significant QTL in chromosome 2 where Pun1 is located (Fig. 2), and significant genomic mutations were never observed in Pun1 and the proximal region in ‘Shishito’ (Table 3). Accordingly, it was considered that ‘Shishito’ possessed either a functional and semi-functional Pun1 allele, except for pun11 at least, and Pun1 or pAMT might not be a major genetic factor in the low pungency of ‘Shishito’.
To elucidate the genetic factors of low pungency in ‘Shishito’, we implemented QTL mapping of capsaicinoid content in the F2 population. As the result, we identified two loci involved in the reduction of capsaicinoid content, designated Shql3 and Shql7 (Fig. 2, Table 2). The total GVE of the two loci was approximately 60%, suggesting that Shql3 and Shql7 were major genetic factors responsible for the pungency level of the F2 population. Although several QTLs related to pungency level have been elucidated in previous studies, most were identified from QTL analysis using interspecific crossed progenies, such as C. annuum × C. frutescens and C. annuum × C. chinense combinations (Blum et al. 2003; Ben-Chaim et al. 2006; Yarnes et al. 2013; Lee et al. 2016; Han et al. 2018). Therefore, it should be mentioned that the present QTLs derived from intraspecific crossed progenies (C. annuum × C. annuum) seem to be uncommon. Also, when we considered their high GVEs, there is a high possibility that the responsible genes will be identified in a future study. Moreover, the genetic effects of these loci were verified regardless of differences in the cultivation environment via genotyping analysis in the vicinity of Shql3 and Shql7 (Fig. 4a, b). Then, it was revealed that the ‘Shishito’ allele of the two loci additively decreased capsaicinoid contents in the absence of obvious epistasis (Fig. 5a, b). Therefore, it was suggested that changes in the pungency level of the F2 population resulted mainly from the independent accumulation of Shql3 and Shql7 alleles. However, some exceptions were also observed in 2021 with several genotypic combinations (Fig. 5a); therefore, the presence or absence of epistasis among the two loci remains controversial.
Finally, we arranged the existence of QTLs around Shql3 and Shql7. Surprisingly, the edge of chromosome 3 was the QTLs-enriched region which was contained not only Shql3 but also various QTLs related to the capsaicinoid content, regardless of differences in the mapping population used in individual QTL analyses (Fig. 3a, Table S1). This suggested that the edge of chromosome 3 was one of the critical regions involved in the regulation of pungency levels in chili peppers, which is indirectly supported by the high density of capsaicinoid biosynthesis genes: KASIII, 4CL, CSE, HCT (Fig. 3a, Table S2). Relatively fewer QTLs seemed to be in proximity to Shql7, with only CaMYB31, which is responsible for the presence of pungency, being in the vicinity (Fig. 3b, Tables S1, S2). These five proximal capsaicinoid biosynthesis genes (KASIII, 4CL, CSE, HCT, and CaMYB31) were considered to be the responsible genes for the two loci; thus, we conducted whole-genome resequencing and explored their genomic variants in ‘Shishito’. As the result, significant variants that apparently induced gene dysfunction were not observed in most genes (KASIII, 4CL, HCT, and CaMYB31), although they had various Indels and SNPs in the upstream region and in introns at modifier levels (Tables 3, S4, S5). This implied the normality of their gene functions, which was supported by our previous studies. Our research group previously revealed that 4CL and HCT, located in the physical range of Shql3, were normally expressed in non-pungent ‘Shishito’ fruit; moreover, their expression levels had no significant correlation with capsaicinoid content when ‘Shishito’ experienced pungency fluctuation (Kondo et al. 2021a, b). In contrast, KASIII and CaMYB31 (linked to Shql3 and Shql7, respectively) showed low expression in non-pungent ‘Shishito’ fruits and high expression in pungent fruits, thereby implying they were functional (Rathnayaka et al. 2021; Kondo et al. 2021b). Therefore, it was difficult to definitively determine that these genes were responsible for Shql3 and Shql7. However, transcriptional levels do not always correspond with the phenotype in the case of nonsense-mediated mRNA decay (Shaul 2015), or detected Indels and SNPs in the upstream region and in introns of these genes could furtively affect their function; thus, our hypothesis should be thoroughly investigated in future studies. On the other hand, we remarkably found one isoform of CSE (XM_016710287) in ‘Shishito’ with a probable frameshift mutation due to the 5 bp deletion in the 5'UTR region (Table 4). Thus, this variant was considered as a candidate gene of Shql3, while another isoform (XM_016710288) did not show such mutation, and it was also considered that this isoform retained its normal function. To demonstrate validity, transcriptional and gene function analyses via reverse genetic approaches will be necessary. Regardless, since it was difficult to determine the responsible genes for Shql3 and Shql7 in the present study, further narrowing of their candidate genomic regions is required. Actually, more than 800 and 200 genes were located on Shql3 and Shql7, respectively, based on the ‘Zunla 1’ genome database (data not shown), indicating that further precise and elaborate QTL mapping will be necessary for screening candidate genes.
In conclusion, the present study was the first to investigate the inheritance and genetic factors of pungency deficiency in Japanese chili peppers. The present results provide novel insights regarding the genetics of pungency traits in Capsicum spp. The absence of pungency in chili peppers has been generally acknowledged as a qualitative trait controlled by individual recessive genes; however, the present study has newly identified an exception. The results of our study will provide helpful information for further research in clarifying the genetic regulation of pungency in chili peppers.
References
Arce-Rodríguez ML, Ochoa-Alejo N (2017) An R2R3-MYB transcription factor regulates capsaicinoid biosynthesis. Plant Physiol 174(3):1359–1370
Ben-Chaim A, Borovsky Y, Falise M et al (2006) QTL analysis for capsaicinoid content in Capsicum. Theor Appl Genet 113:1481–1490
Bennett DJ, Kirby GW (1968) Constitution and biosynthesis of capsaicin. J Chem Soc C: Organic 442–446
Blum E, Mazourek M, O’Connell M et al (2003) Molecular mapping of capsaicinoid biosynthesis genes and quantitative trait loci analysis for capsaicinoid content in Capsicum. Theor Appl Genet 108:79–86
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 23(19):2633–2635
Broman KW, Wu H, Sen Ś, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19(7):889–890
Chen S, Zhou Y, Chen Y, Gu J (2018) Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):884–890
Deshpande RB (1935) Studies in indian chillies: 4. Inheritance of pungency in Capsicum annuum L. Indian J Agric Sci 5:514–516
Fujiwake H, Suzuki T, Iwai K (1982) Capsaicinoid formation in the protoplast from the placenta of Capsicum fruits. Agric Biol Chem 46(10):2951–2952
Han K, Lee HY, Ro NY et al (2018) QTL mapping and GWAS reveal candidate genes controlling capsaicinoid content in Capsicum. Plant Biotechnol J 16:1546–1558
Han K, Jang S, Lee JH et al (2019) A MYB transcription factor is a candidate to control pungency in Capsicum annuum. Theor Appl Genet 132:1235–1246
Ishikawa K, Sasaki S, Matsufuji H, Nunomura O (2004) High ß-carotene and Capsaicinoid contents in seedless fruits of ‘Shishitoh’ pepper. HortSci 39(1):153–155
Kim S, Park M, Yeom SI et al (2014) Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat Genet 46:270–278
Kirii E, Goto T, Yoshida Y et al (2017) Non-pungency in a Japanese chili pepper landrace (Capsicum annuum) is caused by a novel loss-of-function pun1 allele. Japan Soc Hort Sci 86:61–69
Koeda S, Sato K, Saito H et al (2019) Mutation in the putative ketoacyl-ACP reductase CaKR1 induces loss of pungency in Capsicum. Theor Appl Genet 132:65–80
Kondo F, Hatakeyama K, Sakai A et al (2021a) Parthenocarpy induced fluctuations in pungency and expression of capsaicinoid biosynthesis genes in a Japanese pungency-variable sweet chili pepper ‘Shishito’ (Capsicum annuum). Japan Soc Hort Sci 90:48–57
Kondo F, Hatakeyama K, Sakai A et al (2021b) The pungent-variable sweet chili pepper ‘Shishito’ (Capsicum annuum) provides insights regarding the relationship between pungency, the number of seeds, and gene expression involving capsaicinoid biosynthesis. Mol Genet Genomics 296:591–603
Lang Y, Kisaka H, Sugiyama R et al (2009) Functional loss of pAMT results in biosynthesis of capsaicinoids, capsaicinoid analogs, in Capsicum annuum cv. CH-19 sweet. Plant 59:953–961
Lee J, Park SJ, Hong SC et al (2016) QTL mapping for capsaicin and dihydrocapsaicin content in a population of Capsicum annuum “NB1” × Capsicum chinense “Bhut Jolokia.” Plant Breed 135:376–383
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Martínez-López LA, Ochoa-Alejo N, Martínez O (2014) Dynamics of the chili pepper transcriptome during fruit development. BMC Genomics 15:143
Mazourek M, Pujar A, Borovsky Y et al (2009) A Dynamic interface for capsaicinoid systems biology. Plant Physiol 150(4):1806–1821
McKenna A, Hanna M, Banks E et al (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Minamiyama Y, Furutani N, Inaba K et al (2012) A new sweet pepper variety, ‘Kyoto Manganji No. 2’, with non-pungent. Fruit Hort Res (japan) 11(3):411–416
Murakami K, Ido M, Masuda M (2006) Fruit pungency of ‘Shishito’ pepper as affected by a dark interval in continuous fluorescent illumination with temperature alteration. SHITA J 18:284–289
Naves ER, de Ávila SL, Sulpice R et al (2019) Capsaicinoids: Pungency beyond Capsicum. Trends Plant Sci 24:109–120
Qin C, Yu C, Shen Y et al (2014) Whole-genome sequencing of cultivated and wild peppers provides insights into Capsicum domestication and specialization. Proc Natl Acad Sci USA 111:5135–5140
Rathnayaka RMSMB, Kondo F, Prabandaka SS et al (2021) Drought stress induced an increase in the pungency and expression of capsaicinoid biosynthesis genes in chili pepper (Capsicum annuum L.). Japan Soc Hort Sci 90:410–419
Shaul O (2015) Unique Aspects of Plant Nonsense-Mediated mRNA Decay. Trends Plant Sci 20:767–779
Stellari GM, Mazourek M, Jahn MM (2010) Contrasting modes for loss of pungency between cultivated and wild species of Capsicum. Heredity (edinb) 104:460–471
Stewart C, Kang BC, Liu K et al (2005) The Pun1 gene for pungency in pepper encodes a putative acyltransferase. Plant J 42:675–688
Stewart C, Mazourek M, Stellari GM et al (2007) Genetic control of pungency in C. chinense via the Pun1 locus. J Exp Bot 58:979–991
Suzuki T, Iwai K (1984) Chapter 4 Constituents of Red Pepper Species: Chemistry, Biochemistry, Pharmacology, and food Science of the Pungent Principle of Capsicum Species. The Alkaloids: Chemistry and Pharmacology 23:227–299
Tanaka Y, Asano T, Kanemitsu Y et al (2019) Positional differences of intronic transposons in pAMT affect the pungency level in chili pepper through altered splicing efficiency. Plant J 100:693–705
Tsurumaki K, Sasanuma T (2019) Discovery of novel unfunctional pAMT allele pamt10 causing loss of pungency in sweet bell pepper (Capsicum annuum l.). Breed Sci 69:133–142
Yarnes SC, Ashrafi H, Reyes-Chin-Wo S et al (2013) Identification of QTLs for capsaicinoids, fruit quality, and plant architecture-related traits in an interspecific Capsicum RIL population. Genome 56:61–74
Ye K, Schulz MH, Long Q et al (2009) Pindel: A pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25:2865–2871
Zhang ZX, Zhao SN, Liu GF et al (2016) Discovery of putative capsaicin biosynthetic genes by RNA-Seq and digital gene expression analysis of pepper. Sci Rep 6:34121. https://doi.org/10.1038/srep3412
Acknowledgements
This research was financially supported by Yawataya Isogoro Inc. and the Shinshu Foundation for Promotion of Agricultural and Forest Science. Computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Bing yang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kondo, F., Umeda, K., Sudasinghe, S.P. et al. Genetic analysis of pungency deficiency in Japanese chili pepper ‘Shishito’ (Capsicum annuum) revealed its unique heredity and brought the discovery of two genetic loci involved with the reduction of pungency. Mol Genet Genomics 298, 201–212 (2023). https://doi.org/10.1007/s00438-022-01975-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-022-01975-2