Abstract
Verticilllium wilt of cotton is a devastating soil-borne disease, which is caused by Verticillium dahliae Kleb. Bacillus velezensis strain AL7 was isolated from cotton soil. This strain efficiently inhibited the growth of V. dahliae. But the mechanism of the biocontrol strain AL7 remains poorly understood. To understand the possible genetic determinants for biocontrol traits of this strain, we conducted phenotypic, phylogenetic and comparative genomics analysis. Phenotypic analysis showed that strain AL7 exhibited broad-spectrum antifungal activities. We determined that the whole genome sequence of B. velezensis AL7 is a single circular chromosome that is 3.89 Mb in size. The distribution of putative gene clusters that could benefit to biocontrol activities was found in the genome. Phylogenetic analysis of Bacillus strains by using single core-genome clearly placed strain AL7 into the B. velezensis. Meantime, we performed comparative analyses on four Bacillus strains and observed subtle differences in their genome sequences. In addition, comparative genomics analysis showed that the core genomes of B. velezensis are more abundant in genes relevant to secondary metabolism compared with B. subtilis strains. Single mutant in the biosynthetic genes of fengycin demonstrated the function of fengycin in the antagonistic activity of B. velezensis AL7. Here, we report a new biocontrol bacterium B. velezensis AL7 and fengycin contribute to the biocontrol efficacy of the strain. The results showed in the research further sustain the potential of B. velezensis AL7 for application in agriculture production and may be a worthy biocontrol strain for further studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton (Gossypium hirsutum L.) is a great economic importance crop in some developing and developed countries (Zhang et al. 2018). Among worldwide cotton-producing regions, Verticillium wilt of cotton caused by Verticillium dahliae is one of the most serious diseases (Zhang et al. 2019). It has been estimated that almost 26–30% of the yield losses for sugar beet, cotton, and wheat are caused by fungal pathogens alone (Oerke and Dehne 2004). Therefore, controlling plant disease plays an important role in protecting the capacity and quality of crop production. The management of Verticillium wilts includes the use of resistant cultivars, application of chemical fungicides, and crop rotation (Li et al. 2013). These methods have specific concerns and limitations. The use of biological agents is a promising approach against pathogens (Eljounaidi et al. 2016). The biological control of plant disease by using microorganisms can be a safe, cost-effective, and efficient method for controlling plant diseases.
Several biocontrol bacteria, such as Bacillus, Pseudomonas, and Streptomyces were able to reduce Verticillium disease (Erdogan and Benlioglu 2010; Li et al. 2013; Xue et al. 2016). However, spore-forming Bacillus spp. which used for biological preparations are preferred because of their long-term viability facilitates the development of commercial products (Wu et al. 2015). In general, the genus Bacillus is considered as a group of soil inhabitants. However, Bacillus spp. can be isolated from varied environment, including air, water, human and animal gut, vegetables, and food (Kotb 2015; Tidjiani Alou et al. 2015). A wide range of plant disease have been controlled using Bacillus-based biocontrol agents. For example, B. velezensis FZB42 is commercially used as an efficient biocontrol bacterium against fungal and bacterial pathogens (Wu et al. 2015). The biocontrol strains B. subtilis GLB191 and B. velezensis PG12 have the most robust preventive effects against grape downy mildew and apple ring rot (Chen et al. 2016; Zhang et al. 2017). The biocontrol strain B. subtilis 9407 displayed a strong antibacterial activity against bacterial fruit blotch (Fan et al. 2017a, b). The mechanisms can be divided into direct and indirect forms, including facilitation of resource acquisition, competition for nutrients, niche exclusion, production of antibiotic metabolites (Paterson et al. 2017).
Antibiotic production plays a significant role in biocontrol activities (Wu et al. 2015). From the biotechnological point of view, the most noteworthy feature of Bacillus species is their diverse lipopeptides (LPs), which are synthesized non-ribosomally and are structurally diverse. These LPs are the main contributor to the biocontrol activity of Bacillus (Shafi et al. 2017). For example, B. amyloliquefaciens FZB42 produces bacillomycin D and fengycin, which display synergistic antagonistic activity against the pathogen Fusarium oxysporum (Koumoutsi et al. 2004). Bacillus strains UMAF6614 and UMAF6639 have direct antagonism toward pathogens by mediating the production of iturin and fengycin lipopeptides (Romero et al. 2007; Zeriouh et al. 2011). Apart from antimicrobial properties, lipopeptide-associated biocontrol induces systemic resistance in host plants and facilitates biofilm formation and colonization of the plant roots. The bacillomycin D contributes to biofilm formation of strain B. amyloliquefaciens SQR9 (Xu et al. 2013). Surfactin and bacillomycin L in B. subtilis 916 conduce differently but synergistically to prevent growth of rice sheath blight through antifungal activity, colonization, and biofilm formation (Luo et al. 2015). Furthermore, fengycin and surfactin in B. subtilis GLB191 stimulate the plant’s defenses to plant pathogen (Li et al. 2019).
The strain B. velezensis AL7 was isolated from the soil of a cotton plantation in Xinjiang Province, China and exhibited biocontrol effects against cotton Verticillium wilt under greenhouse conditions. However, limited information is known about the mechanism of the strain AL7 on biocontrol. The study aimed to elucidate the biocontrol performance of strain AL7. The main information is currently assisted by implementing genomic approaches, which help to determine the presence of potentially beneficial features in strains. The present study provides a foundation for further studies of related genes and functions and facilitate genetic engineering of B. velezensis AL7 to promote agricultural and industrial applications.
Materials and methods
Strain used in this study
The newly isolated strain B. velezensis AL7 (from the cotton soil) was used in this study. The strain AL7 with outstanding biocontrol performance was selected for whole genome sequencing to obtain the biocontrol informations (Liu et al. 2020).
Microorganisms and growth conditions
The AL7 was routinely cultured at 37 °C on Lurina-Bertani broth (LB) or on solid LB medium supplemented with 1.5% agar (Fan et al. 2017a, b). MSgg medium was used for assays of biofilm formation. The formula for MSgg medium is as follows: 5 mM potassium phosphate (pH 7.0), 100 mM morpholine propane sulfonic acid (pH 7.0), 2 mM MgCl2, 700 mM CaCl2, 50 mM MnCl2, 50 mM FeCl3, 1 mM ZnCl2, 2 mM thiamine, 0.5% glycerol, 0.5% glutamic acid, 50 mg/mL tryptophan, 50 mg/mL threonine, and 50 mg/mL phenylalanine (Branda et al. 2001; Fan et al. 2017a, b). Tetracycline (10 µg/mL) as an antibiotic was added when necessary.
In vitro antifungal activity of AL7
In the plate confrontation assay, the plant pathogen was grown on potato-dextrose-agar (PDA) plates at 28 °C for 3–5 days. A 5 mm-diameter block of mycelium culture was cut and transferred into the center of fresh PDA plates (90 mm). After 24 h of cultivation, 2 µL of the overnight cultures of strain AL7 grown in LB medium was added on the PDA plate 2.5 cm away from the center, where the mycelium agar block was placed (Cao et al. 2018). Antifungal activity was checked by measuring the diameter of mycelium after 5–7 days of incubation at 28 °C.
Cotton was used to evaluate the biocontrol activity of strain AL7. Ten cotton plants were grown in each of the 18 pots under natural lighting at 28 °C. Inoculation was performed by adding 50 mL of strain AL7 (108 cfu/mL) and 50 mL of V. dahliae spores (107 conidia/mL). Non-bacterialed cotton (control) were just drenched with 50 mL of LB, whereas 50 mL of V. dahliae spores (107 conidia/mL) were added to the cotton plant as the positive control. Six pots were employed for each replicate, and values were recorded as the means of six replicates for each treatment. Cotton plants were checked for severity based on the disease index as described previously (Gomez-Lama Cabanas et al. 2018).
Phylogenetic analysis
A maximum-likelihood (ML) phylogenetic tree of Bacillus species was established based on the single-copy core proteins sequence shared by Bacillus genomes and the genome of Paenibacillus polymyxa M1 according to the following methods: (1) multiple alignment of amino acid sequences were carried out via mafft (version 7.310) (Katoh and Standley, 2013); (2) conserved blocks from multiple alignment of protein were selected through Gblocks (Castresana, 2000); and (3) ML tree were established using the software of RAxML (version 8.2.10) using the PROTGAMMALGX model with 100 bootstrap replicates (Zeng et al. 2018). The tree was displayed using a web-based software (http://itol.embl.de/). Then, the average nucleotide identity (ANI) values between two genome sequences were calculated using OrthoANI (Yoon et al. 2017).
Pan-genome analysis
Pan-genome analysis of all strains was performed with a pan-genome analysis pipeline (PGAP) software after genome annotation in Prokka (Zhao et al. 2012). The requirements of the pan-genome analysis is as follows. Core orthologs with at least 50% protein sequence identity to each other were clustered and 50% overlap with the longest sequence with an e-value 1e-5. Core genomes and strain-unique genomes were selected from the pan-genome table by using own Perl script (Zeng et al. 2018). Functional annotation of the core genomes of B. subtilis and B. velezensis was performed using the COG database.
Core genome, pan-genomes, and their estimated respective sizes and trajectories were made using models and regression algorithms proposed by Tettelin and colleagues (Tettelin et al. 2008). Core genome and pan-genomes were visualized through PanGP v1.0.1 software. PanGP was run using default parameters generating distribution plots of (i) total genes and (ii) conserved genes found upon progressive sampling of “n” genomes (Zhao et al. 2014).
Whole-genome sequence comparisons
Genome comparisons were conducted by using program of BLAST ring image generator (BRIG) version 0.95. The blastn option was selected for analysis. The reference sequence is the genome sequence of AL7, whereas the genome sequences of other Bacillus strains were used as query sequences. To investigate genome rearrangements in strain AL7 and related bacteria, we adopted the Mauve v2.3.1 software of progressive Mauve program and to visualize multiple genome alignments of the four Bacillus strains (Darling et al. 2004).
Construction of ΔfenA mutant and tagging strains with green fluorescence protein (GFP)
A fenA-deletion mutant was generated through homologous recombination. A tetracycline resistance gene was amplified from plasmid pGFP78 using primer pair tet-F and tet-R. Primers fenA-U-F and fenA-U-R were used to amplify the upstream region (1329 bp) of fenA and fenA-D-F and fenA-D-R were used to amplify the downstream region (1270 bp). Then, all of them combined through fusion polymerase chain reaction (PCR) and ligated into plasmid PMD19T via gateway cloning technology. For the transformations process, competent cells were prepared as describe by published papers with slight modifications (Zhang et al. 2017). Brief, 1 mL of the overnight cultures was added to 40 mL growth medium (LB medium with 0.5 M sorbitol) and cultured at 37 °C, 200 rpm to OD600 = 0.5. Then, the pellets were harvested by centrifugation at 8000 rpm and 4 °C for 10 min and washed four times with an equal volume of electroporation buffer (0.5 M sorbitol, 0.5 M mannitol, 10% glycerol, pH = 7). The plasmid was added to the resulting competent cells and immediately given a 1.8 kV electric shock and cell suspensions immediately were diluted with 1 mL LB medium containing 0.5 M sorbitol and 0.38 M mannitol. Finally, it incubated at 37 °C with shaking at 160 rpm for 3 h. The cells were spread onto LB plates amended with tetracycline. Then, the mutant ΔfenA was selected, identified via PCR with primer pair fenA-out-F and fenA-out-R, and subsequently confirmed by sequencing (Table S1 and S2). The pGFP78 plasmid containing a GFP tag was transformed into B. velezensis AL7. The transformants AL7-gfp with positive green fluorescence emission were selected.
In vitro antifungal activity of ΔfenA
Activities of ΔfenA against spore germination of V. dahliae were evaluated as follows. Spores of V. dahliae were diluted to 105 or 106 mL−1 and 300 µL were inoculated to PDA plates. Then, the plates were permitted to dry under a laminar flow hood. Portions (2 µL) of culture of strain AL7 were added in the center of plate. The plates were cultured at 28 °C, and inhibition zones were measured after 3 to 5 days (Luo et al. 2015).
Biofilm formation and swarming motility assay
The cultures of AL7 was shaken at 37 °C until optical density (OD600) was 0.8 using LB liquid medium (5 mL). 1 mL of cells were collected through centrifugation at 6000 × g for 5 min, washed with phosphate-buffered saline (PBS), and resuspended in 100 µL PBS (Fan et al. 2017a, b). The swarming motility of AL7 was tested using standard protocols with minor modification. LB plates containing 0.7% agar were dried in a laminar flow hood for 20 min, and then 3 µL of the cell suspension of strain AL7 was added on the center of each plate. The plates were cultivated at room temperature for 7 h to allow cell growth to obviously visualize the zone of swarming. Then, the swarming agar plates were dried for another 2 h in a laminar flow hood, and the diameter of the swarming zone was measured (Fan et al. 2017a, b). The experiment was conducted with three independent assays, each with at least three technical replicates.
Biofilm formation was analyzed in MSgg medium to monitor pellicle formation. B. velezensis AL7 was cultured in LB medium at 37 °C overnight. Then, 4 µL of culture was added to the MSgg medium (a 1000-fold dilution) in a 12-well plate and cultured statically at 28 °C for 72 h (Fan et al. 2017a, b). To quantify biofilm formation, we used the reported methods (Fan et al. 2017a, b; Weng et al. 2013). The assay was performed with three independent experiments. For colony architecture, 2 µL of the culture was added onto the surface of an MSgg plate containing 1.5% agar (Fan et al. 2017a, b).
Nucleotide sequence accession numbers
The complete genome sequence of B. velezensis AL7 has been deposited in NCBI under the GenBank accession number CP045926.
Results
Strain AL7 is an efficient biocontrol agent against Verticillium wilt of cotton
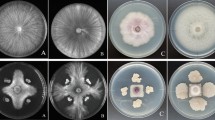
Strain AL7 inhibited spore germination with a clear inhibition zone and the diameter of the inhibition zone is 23.3 ± 0.58 mm (Fig. 1a and b). Meantime, strain AL7 remarkably prevented the hyphae growth of other fungal pathogens with an obvious inhibition zone (Fig. 1c, d). Then, the potential biological control abilities of strain AL7 on Verticillium wilt of cotton was assessed on greenhouse condition. The disease index of the AL7-treated cotton plant was lower than the LB-treated control plant for a period of 70 days after inoculation (17.9 vs 78.2). The results suggested that strain AL7 could effectively control the Verticillium wilt of cotton on greenhouse condition. As expected, strain AL7 formed robust, wrinkled pellicles on MSgg liquid medium (Fig. 1e–g). Strain AL7 also exhibited remarkable swarming motility and it could colonize more than half the plate after 7 h of culture (Fig. 1i–h). Furthermore, cotton plants were treated with gfp-tagged strains and were examined via confocal scanning laser microscopy (CSLM) 7 days later to detect the colonization of strain AL7. The results displayed that the gfp-tagged strains were easily observed in cotton plant tissue and confirmed that the strain AL7 could recolonize in plant tissue.
Biocontrol efficacy of strain AL7. a Effect of AL7 treatments on the Verticillium wilt of cotton. b Diameter of inhibition zone of strain AL7. Error bars indicate ± SD of three replicates. c Antagonistic activity of AL7 against nine pathogenic fungi on PDA plate. d Quantification of antagonistic activity of AL7. *Shows obvious differences at P < 0.01 compared with the control. e Pellicle formation by strain AL7 on Msgg liquid media. f OD readings from 12-well-plate experiments of biofilm formation by AL7. Data are expressed as the mean ± SD (n = 6). g The colony morphology of strain AL7 was monitored on MSgg plates. h Data presented are the swarming ability of AL7 on LB plates 8 h later. Data are expressed as mean ± SD (n = 5). CK represents the LB medium, and AL7 represents the strain. i Swarming ability of AL7 was assessed
In summary, we have isolated the strain AL7, which showed broad-spectrum antagonistic activities and displayed stronger biofilm formation and swarming motility. Above all, strain AL7 is a beneficial microbe with potential for biotechnological application.
Genome characteristics of biocontrol strain AL7
The genome sequence of AL7 is 3,894,709 bp in length with a GC content of 46.64%. The number of predicted protein-coding genes of AL7 was 3706. Among the predicted protein-coding genes, 2872 of them could be allocated a putative function and 1,060 genes were predicted to encode hypothetical proteins (Figure S1). The average length of protein-coding genes is 930 bp. The protein-coding genes account for 88.50% of the genome sequence. In addition, a total of 86 tRNA-coding genes and 27 rRNA genes were annotated in the genome sequence of strain AL7 (Table 1).
The functional classification assigned the protein-coding genes in different COG functional groups (Figure S2). This analysis found that the highest number of genes (278) are participated in amino acid transport and metabolism. A group of 209 genes is involved in transcription roles, and a group of 64 genes is involved in secondary metabolites. As expected, surfactin, iturin, and fengycin gene clusters were annotated in the genome sequence of AL7 (Fig. 2). All of the secondary metabolism was displayed in Table S3.
Strain AL7 was identified as B. velezensis
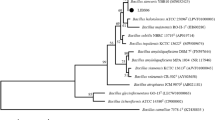
The phylogenetic analysis based on 16S rDNA sequence indicated that strain AL7 has a close relationship with B. velezensis (Figure S3). Meantime, the phylogenetic tree which constructed through the ML method based on the 903 single-copy core genes showed that B. velezensis strains were clustered together and formed a single cluster (Fig. 3A). In this cluster, AL7 grouped together with other sequenced B. velezensis strains. We also found that B. velezensis strains were more closely related to B. amyloliquefaciens strains and formed a single large clade make up of two subclades (Fig. 3A). Moreover, genome relatedness of strain AL7 was analyzed based on ANI. The ANI values of strain AL7 against all B. velezensis strains were > 98%, which is above the threshold of 96% (Fig. 3B) (Richter and Rossello-Mora 2009). Strain AL7 has a great nucleotide identity (98.67%) with strain FZB42, which is a type strain for B. velezensis species. We also performed clustering analysis based on ANI values among each strain to find out evolutionary relationship of Bacillus strains and found that B. velezensis strains clustered together. However, some discrepancies were found when comparing with the phylogenetic tree constructed based on core-genome sequence (Fig. 3B).
Pan-genome analysis of B. velezensis strains
We carried out pan-genome analysis to gain clue into core and variable gene pool among B. velezensis isolated from diverse ecological origin. The 30 examined B. velezensis strains produced a pan-genome size of 7167 genes. Based on the result, the core genome includes 3108 genes, the accessory genome is comprised of 1991 genes, and the unique genome contains 2068 genes (Fig. 4A). B. velezensis strain 9912D has the highest number of unique genes, whereas B. velezensis strains LS69 and S3-1 have the least number of unique genes.
Pan-genomes of B. velezensis strains. A Number of specific genes for each strain of the B. velezensis. The inner circle displays the core genomes shared between all strains. The unique genes for each strain are displayed in each of the outer circles. B Curves for B. velezensis pan-genomes and core genomes. The blue dots represent the pan-genome size of B. velezensis for each genome comparison while the green dots represent the core genome size of B. velezensis for each genome comparison. The median values were linked to display the relationship between number of genomes and gene families
The core-genome and pan-genome sizes of B. velezensis were calculated by extrapolation of the selected genome data. The generated pan-genome curves of B. velezensis are cross-sectional by the Heaps law mathematical functions: \(Y\; = \;77.46X^{0.49\;} + \;2973.52\) , where Y represents the size of pan-genome and X indicates to the number of selected strains. According to above results, the pan-genome size appeared to reach infinity with the number of strains increasing (Fig. 4B). Therefore, our data suggest that B. velezensis hold open pan-genomes, which illustrates that the species have infinite genomes. The curve of the core genome was also asymptotic, with 3108 core genes after addition of the 30th genome. The infinite pan-genome indicates the bacteria will keep gaining other new genes as they evolve independently over evolutionary time.
Comparative genomics analysis of four biocontrol Bacillus strains
To obtain an extensive overview of the occurrence of biocontrol activity in Bacillus strains, we selected three biocontrol Bacillus strains (BSD-2, HJ5 and CAU B946) for study, which are well-known for their inhibition ability of plant disease, especially for Verticillium wilt of cotton (Blom et al. 2012; Li et al. 2013; Liu et al. 2016). Firstly, a global alignment of the four Bacillus strains was performed using progressive Mauve and BRIG software. The result showed a close genetic relatedness of these strains and displayed that most sections within their genomes were conserved (Figure S4). In addition, the comparison of strains using progressive Mauve software illustrated that some collinear blocks were present with several regions of rearrangement (Figure S5). Meanwhile, the genome features for each of the biocontrol Bacillus strains analyzed are summarized in Table S4. The studied genomes had the same genome sizes (4 M) and number of protein-coding genes (3873 in average). However, their GC contents were different. The GC content in B. velezensis strain is higher than that in B. subtilis strains (Table S4). Furthermore, we performed phylogenetic analysis based on the core-genomes of Bacillus was estimated from 903 proteins sequence. The resulting phylogenetic tree grouped the B. velezensis and B. subtilis strains into two species clades (Fig. 5A).
Comparative genomics analysis of the four biocontrol Bacillus strains. A Phylogenetic tree showing the relationship of B. velezensis and B. subtilis strains. B Genomic diversity of the four biocontrol Bacillus strains. C Organization of lipopeptides genes in the four Bacillus strains. Surfactin, iturin, and fengycin are marked with different colors
In total, 3706, 3947, 3943 and 3896 protein-coding genes (including hypothetical proteins) of the Bacillus strain AL7, BSD-2, HJ5, and CAU B946 were annotated, respectively. The analysis of the four biocontrol Bacillus genomes identified 2965 predicted protein-coding genes which formed the core genome (Fig. 5B). The predicted proteins were functionally classified using the COG database, and COGs classifications were compared among the genomes of the four biocontrol Bacillus strains. COGs displayed analogous distributions among the studied strains (Table S5). The results verified that those acquired from the pan-genome analysis, where the studied biocontrol Bacillus strains are very similar, with several genes participated in different functions. However, we found that B. velezensis strains had more specific genes than B. subtilis strains (Fig. 5B). Thus, it stimulates us to compare the functions of core genomes of B. velezensis and B. subtilis strains. We compared the core genome of B. velezensis and B. subtilis strains by using COG distributions to decide whether there were distinctions in the proportion of the core genome attributable to a specific cellular process. As excepted, we found that the core genome of B. velezensis was disproportionately enriched in secondary metabolism genes (Fisher’s exact test; P value < 0.01, Table 2). Furthermore, we found that B. velezensis strains harbored the three types of lipopeptides (surfactin, iturin and fengycin), whereas B. subtilis strains only included surfactin and fengycin (Fig. 5C). However, all of the Bacillus strains inhibited cotton Verticillium wilt. Fengycin may play an important role in the inhibition cotton Verticillium wilt.
A ΔfenA mutant exhibited decreased biocontrol efficacy
To determine the potential role of the fengycin in the biocontrol ability of AL7, a mutant was constructed in the biosynthetic genes of fengycin. We investigated whether the production of the fengycin compounds would influence the biocontrol efficacy in strain AL7 by utilizing the ΔfenA mutant. The prevention abilities of strain AL7 against plant pathogens were attenuated in the ΔfenA mutant (Fig. 6). The result indicated that the production of fengycin affect the biocontrol ability of strain AL7.
Antagonistic activities of B. velezensis AL7 and its mutant against cotton Verticillium wilt. A 300 µL of V. dahliae spores (106 conidial/mL) was added on the plate and 2 µL of AL7 and ΔfenA also was added in the center of PDA plate. Obvious inhibition zones were observed after cultivation at 28 °C for 3 days. B Investigation of the inhibition zones of AL7 and mutant against V. dahliae. Data are the average inhibition zone region of three triplicates ± the standard deviation of three independent assays. *Represented that there are significantly different (P value < 0.05) according to t test
Discussion
In the present study, biocontrol strain AL7 displayed significant inhibitory effect on V. dahliae in plate and greenhouse condition. However, the control efficiency of biocontrol strains is influenced by several factors, such as environmental conditions, colonization, inoculation methods, and antifungal compound production (Tan et al. 2019). For example, environmental conditions could affect the efficacy of endophytes under field conditions, despite biocontrol strain have displayed strong in vitro suppression against the plant disease (Martin et al. 2015). Therefore, we need to check the biocontrol efficacy of B. velezensis AL7 in different field. In addition, plants and communities of microorganisms that play important roles for the host’s development and health have co-evolved. (Berg et al. 2017). It is reported that phage treatment did not affect the existing rhizosphere microbiome and enriched some bacterial species that were antagonistic toward Ralstonia solanacearum (Wang et al. 2019). Moreover, how does Bacillus control plant diseases through regulating the microbial community and we could learn the relationship between Bacillus and community structure to understand the mechanism of Bacillus and to apply it in the agricultural production.
Previously, the definitive relationship of these strains is very difficult to obtain. Because of many Bacillus strains show very similar phenotypic and physiological properties and 16S rRNAs gene sequences. With the development of sequencing technique, more and more genome sequence have been published. The constructed species tree based on genome sequence will help us to correct and improve the taxonomy of strains. For example, strains CECT 8237 and CECT 8238 are classified as B. subtilis based on 16S rDNA and metabolic profiles. While, the two biocontrol strain were identified as B. amyloliquefaciens group using 11 conserved protein sequence (Magno-Perez-Bryan et al. 2015). In the study, we identified that strain AL7 belongs to B. velezensis by using 16S rDNA and core-genome sequence (Fig. 3 and Figure S3). Meantime, B. amyloliquefaciens group strains were divided into three taxonomic units that could be regarded as ‘subspecies’, they were re-classified into other species through the phylogenomic analysis. A phylogenetic tree was constructed using core-genome sequence and divided into three tightly linked branches, including three species (Wu et al. 2015; Fan et al. 2017a, b). Confirming the taxonomic status of Bacillus using whole-genome sequence would likely contribute to further insights into their phylogeny and adaptation.
To determine the specific traits of Bacillus strain, we selected four Bacillus strains for comparative genome analysis. We found that B. velezensis strains possess more specific genes than B. subtilis genes (Fig. 5B). Furthermore, comparison of the core genome between B. velezensis and B. subtilis strains based on COGs database displayed that the genes participated in secondary metabolites were more abundant in B. velezensis strains (Table 2). Genomic and evolutionary features of B. velezensis and B. subtilis are obtaining great attention because of their potential in agriculture. B. velezensis and B. subtilis comprise an evolutionary compact (Zhang et al. 2016), but their phenotypic difference remains unclear. Secondary metabolism possesses multiple functions, including antifungal activities, motility, and biofilm formation (Luo et al. 2015). The major adaptive colonization strategies of Bacillus strains are motility and biofilm formation in new environments (Zhao et al. 2017). We should compare the antifungal activities and colonization abilities of B. subtilis and B. velezensis strains isolated from the same environment in further research. However, interactions and competitions between organisms usually have been happening in rhizosphere which is the highly dynamic font; microbes within the environment have to compete with each other for resources, by nutrient/space competition and direct suppression (Zhang et al. 2016). Plant-associated strains hold more intermediary metabolism and secondary metabolites biosynthesis genes when compared with non-plant-associated strains (Zhang et al. 2016). Overall, different species and strains from different source display the enrichment of functional genes, especially those involved in secondary metabolism. Secondary metabolism also plays a vital role in the plant-bacteria interaction. We are also interested in the difference between endophyte and non-endophyte strains to help us to understand the evolutionary relationship of Bacillus strain.
We found that fengycin gene cluster from biocontrol strain B. velezensis AL7 plays an important role in inhibiting V. dahliae (Fig. 6). Iturins can induce cell death in V. dahliae and serve as activators to induce the expression of some defence genes in cotton plants (Han et al. 2015). In the current study, we compared the B. velezensis and B. subtilis strains and found their difference. Based on the mentioned results, B. velezensis strains may have good control effect than B. subtilis strains. We also need to compare the efficacy of different Bacillus species. Meantime, we should focus on the iturin gene cluster to study the function of the gene cluster in the inhibition effect of B. velezensis AL7 against cotton Verticillium wilt. Our results revealed that the strain AL7 exhibits broad spectrum antifungal activities and contains genes related to antibiotic production. Importantly, the fengycin gene cluster of strain AL7 has an effect against cotton Verticillium wilt. Further research on the biocontrol mechanisms of this beneficial strain will aid the development of biocontrol agents for plant diseases.
References
Berg G, Koeberl M, Rybakova D, Mueller H, Grosch R, Smalla K (2017) Plant microbial diversity is suggested as the key to future biocontrol and health trends. FEMS Microbiol Ecol 93:fix050. https://doi.org/10.1093/femsec/fix050
Blom J, Rueckert C, Niu B, Wang Q, Borriss R (2012) The complete genome of Bacillus amyloliquefaciens subsp plantarum CAU B946 contains a gene cluster for nonribosomal synthesis of Iturin A. J Bacteriol 194:1845–1846
Branda SS, Gonzalez-Pastor JE, Ben-Yehuda S, Losick R, Kolter R (2001) Fruiting body formation by Bacillus subtilis. P Natl Acad Sci USA 98:11621–11626
Cao Y, Pi H, Chandrangsu P, Li Y, Wang Y, Zhou H, Xiong H, Helmann JD, Cai Y (2018) Antagonism of two plant-growth promoting Bacillus velezensis isolates against Ralstonia solanacearum and Fusarium oxysporum. Sci Rep 8:4360
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Chen X, Zhang Y, Fu X, Li Y, Wang Q (2016) Isolation and characterization of Bacillus amyloliquefaciens PG12 for the biological control of apple ring rot. Postharvest Biol Tec 115:113–121
Darling A, Mau B, Blattner FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14:1394–1403
Eljounaidi K, Lee SK, Bae H (2016) Bacterial endophytes as potential biocontrol agents of vascular wilt diseases—review and future prospects. Biol Control 103:62–68
Erdogan O, Benlioglu K (2010) Biological control of Verticillium wilt on cotton by the use of fluorescent Pseudomonas spp. under field conditions. Biol Control 53:39–45
Fan B, Blom J, Klenk H, Borriss R (2017a) Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis form an “operational group B. amyloliquefaciens” within the B. subtilis species complex. Front Microbiol 8:22
Fan H, Zhang Z, Li Y, Zhang X, Duan Y, Wang Q (2017b) Biocontrol of bacterial fruit blotch by Bacillus subtilis 9407 via surfactin-mediated antibacterial activity and colonization. Front Microbiol 8:1973
Gomez-Lama Cabanas C, Legarda G, Ruano-Rosa D, Pizarro-Tobias P, Valverde-Corredor A, Niqui JL, Trivino JC, Roca A, Mercado-Blanco J (2018) Indigenous Pseudomonas spp. strains from the Olive (Olea europaea L) rhizosphere as effective biocontrol agents against Verticillium dahlia: from the host roots to the bacterial genomes. Front Microbiol 9:277
Han Q, Wu F, Wang X, Qi H, Shi L, Ren A, Liu Q, Zhao M, Tang C (2015) The bacterial lipopeptide iturins induce Verticillium dahliae cell death by affecting fungal signalling pathways and mediate plant defence responses involved in pathogen-associated molecular pattern-triggered immunity. Environ Microbiol 17:1166–1188
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kotb E (2015) Purification and partial characterization of serine fibrinolytic enzyme from Bacillus megaterium KSK-07 isolated from kishk, a traditional Egyptian fermented food. Appl Biochem Micro 51:34–43
Koumoutsi A, Chen XH, Henne A, Liesegang H, Hitzeroth G, Franke P, Vater J, Borriss R (2004) Structural and functional characterization of gene clusters directing nonribosomal synthesis of bioactive cyclic lipopeptides in Bacillus amyloliquefaciens strain FZB42. J BACTERIOL 186:1084–1096
Li S, Zhang N, Zhang Z, Luo J, Shen B, Zhang R, Shen Q (2013) Antagonist Bacillus subtilis HJ5 controls Verticillium wilt of cotton by root colonization and biofilm formation. Biol Fert Soils 49:295–303
Li Y, Heloir M, Zhang X, Geissler M, Trouvelot S, Jacquens L, Henkel M, Su X, Fang X, Wang Q, Adrian M (2019) Surfactin and fengycin contribute to the protection of a Bacillus subtilis strain against grape downy mildew by both direct effect and defence stimulation. Mol Plant Pathol 20:1037–1050
Liu H, Yin S, An L, Zhang G, Cheng H, Xi Y, Cui G, Zhang F, Zhang L (2016) Complete genome sequence of Bacillus subtilis BSD-2, a microbial germicide isolated from cultivated cotton. J Biotechnol 230:26–27
Liu H, ZengQ Wang W, Zhang R, Yao J, Gill SR (2020) Complete genome sequence of Bacillus velezensis strain AL7, a biocontrol agent for suppression of cotton Verticillium wilt. Microbiol Resour Announc 9:e01595–19. https://doi.org/10.1128/MRA.01595-19
Luo C, Zhou H, Zou J, Wang X, Zhang R, Xiang Y, Chen Z (2015) Bacillomycin L and surfactin contribute synergistically to the phenotypic features of Bacillus subtilis 916 and the biocontrol of rice sheath blight induced by Rhizoctonia solani. Appl Microbiol Biot 99:1897–1910
Magno-Perez-Bryan MC, Martinez-Garcia PM, Hierrezuelo J, Rodriguez-Palenzuela P, Arrebola E, Ramos C, de Vicente A, Perez-Garcia A, Romero D (2015) Comparative genomics within the Bacillus genus reveal the singularities of two robust Bacillus amyloliquefaciens biocontrol strains. Mol Plant Microbe in 28:1102–1116
Martin JA, Macaya-Sanz D, Witzell J, Blumenstein K, Gil L (2015) Strong in vitro antagonism by elm xylem endophytes is not accompanied by temporally stable in planta protection against a vascular pathogen under field conditions. Eur J Plant Pathol 142:185–196
Oerke EC, Dehne HW (2004) Safeguarding production—losses in major crops and the role of crop protection. Crop Prot 23:275–285
Paterson J, Jahanshah G, Li Y, Wang Q, Mehnaz S, Gross H (2017) The contribution of genome mining strategies to the understanding of active principles of PGPR strains. FEMS Microbiol Ecol 93:fiw249. https://doi.org/10.1093/femsec/fiw249
Richter M, Rossello-Mora R (2009) Shifting the genomic gold standard for the prokaryotic species definition. P Natl Acad Sci USA 106:19126–19131
Romero D, de Vicente A, Rakotoaly RH, Dufour SE, Veening J, Arrebola E, Cazorla FM, Kuipers OP, Paquot M, Perez-Garcia A (2007) The iturin and fengycin families of lipopeptides are key factors in antagonism of Bacillus subtilis toward Podosphaera fusca. Mol Plant Microbe in 20:430–440
Shafi J, Tian H, Ji M (2017) Bacillus species as versatile weapons for plant pathogens: a review. Biotechnol Biotec Eq 31:446–459
Tan T, Zhu J, Shen A, Li J, Yu Y, Zhang M, Zhao M, Li Z, Chen J, Gao C, Cheng Y, Guo L, Yan L, Sun X, Zeng L, Yan Z (2019) Isolation and identification of a Bacillus subtilis HZ-72 exhibiting biocontrol activity against flax seedling blight. Eur J Plant Pathol 153:825–836
Tettelin H, Riley D, Cattuto C, Medini D (2008) Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol 11:472–477
Tidjiani Alou M, Rathored J, Khelaifia S, Michelle C, Brah S, Diallo BA, Raoult D, Lagier J (2015) Bacillus rubiinfantis sp. nov. strain mt2T, a new bacterial species isolated from human gut. New Microbes New Infect 8:51–60
Wang X, Wei Z, Yang K, Wang J, Jousset A, Xu Y, Shen Q, Friman V (2019) Phage combination therapies for bacterial wilt disease in tomato. Nat Biotechnol 37:1513–1520
Weng J, Wang Y, Li J, Shen Q, Zhang R (2013) Enhanced root colonization and biocontrol activity of Bacillus amyloliquefaciens SQR9 by abrB gene disruption. Appl Microbiol Biot 97:8823–8830
Wu L, Wu H, Qiao J, Gao X, Borriss R (2015) Novel routes for improving biocontrol activity of Bacillus based bioinoculants. Front Microbiol 6:1395
Xu Z, Shao J, Li B, Yan X, Shen Q, Zhang R (2013) Contribution of Bacillomycin D in Bacillus amyloliquefaciens SQR9 to antifungal activity and biofilm formation. Appl Environ Microb 79:808–815
Xue L, Gu M, Xu W, Lu J, Xue Q (2016) Antagonistic Streptomyces enhances defense-related responses in cotton for biocontrol of wilt caused by phytotoxin of Verticillium dahliae. Phytoparasitica 44:225–237
Yoon S, Ha S, Lim J, Kwon S, Chun J (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Anton Leeuw Int J G 110:1281–1286
Zeng Q, Xie J, Li Y, Gao T, Xu C, Wang Q (2018) Comparative genomic and functional analyses of four sequenced Bacillus cereus genomes reveal conservation of genes relevant to plant-growth- promoting traits. Sci Rep-UK 8:17009. https://doi.org/10.1038/s41598-018-35300-y
Zeriouh H, Romero D, Garcia-Gutierrez L, Cazorla FM, de Vicente A, Perez-Garcia A (2011) The iturin-like lipopeptides are essential components in the biological control arsenal of Bacillus subtilis against bacterial diseases of cucurbits. Mol Plant Microbe in 24:1540–1552
Zhang X, Zhou Y, Li Y, Fu X, Wang Q (2017) Screening and characterization of endophytic Bacillus for biocontrol of grapevine downy mildew. Crop Prot 96:173–179
Zhang F, Li X, Zhu S, Ojaghian MR, Zhang J (2018) Biocontrol potential of Paenibacillus polymyxa against Verticillium dahliae infecting cotton plants. Biol Control 127:70–77
Zhang L, Li W, Tao Y, Zhao S, Yao L, Cai Y, Niu Q (2019) Overexpression of the key virulence factor 1,3–1,4-beta-D-glucanase in the endophytic bacterium Bacillus halotolerans Y6 to improve Verticillium resistance in cotton. J Agr Food Chem 67:6828–6836
Zhang N, Yang D, Kendall JRA, Borriss R, Druzhinina IS, Kubicek CP, Shen Q, Zhang R (2016) Comparative genomic analysis of Bacillus amyloliquefaciens and Bacillus subtilis reveals evolutional traits for adaptation to plant-associated habitats. Front Microbiol 7:2039
Zhao Y, Wu J, Yang J, Sun S, Xiao J, Yu J (2012) PGAP: pan-genomes analysis pipeline. Bioinformatics 28:416–418
Zhao Y, Jia X, Yang J, Ling Y, Zhang Z, Yu J, Wu J, Xiao J (2014) PanGP: a tool for quickly analyzing bacterial pan-genome profile. Bioinformatics 30:1297–1299
Zhao H, Shao D, Jiang C, Shi J, Li Q, Huang Q, Rajoka MSR, Yang H, Jin M (2017) Biological activity of lipopeptides from Bacillus. Appl Microbiol Biot 101:5951–5960
Acknowledgements
This study was funded by the project of renovation capacity building for the young sci-tech talents sponsored by Xinjiang academy of agricultural sciences (xjnkq-2020015), The National Key Research and Development Program of China (No. 2017YFD020030303), Science and technology support project of Xinjiang Autonomous Region (2019E0244).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Haiyang Liu declares that he has no conflict of interest. Qingchao Zeng declares that he has no conflict of interest. Nuerziya Yalimaimaiti declares that she has no conflict of interest. Wei Wang declares that he has no conflict of interest. Renfu Zhang declares that he has no conflict of interest. Ju Yao declares that he has no conflict of interest.
Ethical approval
The artical does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
438_2021_1816_MOESM1_ESM.tif
Supplementary file1 Figure S1 Genome map of B. velezensisstrainAL7. Each ring of the circle represents a different genome information: rings from the outside inward: (1) scale marks (unit: Mb), (2, 3) protein-coding genes on the forward and reverse strands, respectively (different color represent the different functional classifications), (4, 5) rRNA (blue) and tRNA (red) on the forward and reverse strands, respectively, (6) GC content (positive: red; negative: blue), and (7) GC skew (above average: aquamarine; below average: orange). (TIF 19,804 KB)
438_2021_1816_MOESM4_ESM.tif
Supplementary file4 Figure S4 Genome comparisons of other B. velezensisstrains against reference genome (strain FZB42) generating through BRIG version 0.95. The circular map explains the whole genome comparison of biocontrol strain AL7 against the other three selected B. velezensis. Theinner cycle displays the genome sequence of the reference strain AL7 and the shade of colors display the similarities between each strains with biocontrol strain AL7. (TIF 4,038 KB)
438_2021_1816_MOESM5_ESM.tif
Supplementary file5 Figure S5 Genome alignment showing synthetic blocks between Bacillusstrains. The same colored blocks represent the homologous DNA regions among the strains, and gaps display the non-homologous regions. (TIF 1,864 KB)
438_2021_1816_MOESM8_ESM.xlsx
Supplementary file8 Table S3 Genes and gene cluster encoding for secondary metabolites in Bacillus velezensis AL7. (XLSX 10 KB)
438_2021_1816_MOESM10_ESM.xlsx
Supplementary file10 Table S5 Comparative analysis of the functional classifications based on COG of the protein-coding genes of the Bacillusstrains analyzed. (XLSX 10 KB)
Rights and permissions
About this article
Cite this article
Liu, H., Zeng, Q., Yalimaimaiti, N. et al. Comprehensive genomic analysis of Bacillus velezensis AL7 reveals its biocontrol potential against Verticillium wilt of cotton. Mol Genet Genomics 296, 1287–1298 (2021). https://doi.org/10.1007/s00438-021-01816-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-021-01816-8