Abstract
The domestication of cultivated crops from their wild relatives narrowed down their genetic diversity in a bottleneck effect. Subsequently, the cultivation areas of crops have expanded all over the world into various environmental conditions from the original area along with human migration after domestication. Here, we demonstrated the genetic changes in the adaptation of rice to Hokkaido (41°2–45°3N latitude), Japan, from the tropics of their origin in Asian cultivated rice, Oryza sativa L. Although cultivated rice originated from the tropics, Hokkaido is one of the northern-limits of rice cultivation worldwide. Population genomics focusing on the local populations showed the varieties had genetically distinct classes with limited genetic diversity. In addition, some varieties in the class carried unique genotypes for flowering time, exhibiting extremely early flowering time. Certain mutations in unique genotypes can split off the varieties that are able to grow in Hokkaido. Furthermore, the changes in the genotype for flowering time during rice cultivation in Hokkaido demonstrated novel combinations of genes for flowering time owing to the intensive artificial selection on natural variation and rice breeding programs to achieve stable rice production in Hokkaido.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cultivated crop species have been grown all over the world, whereas the wild species of cultivated crops grow in their original areas. The genetic diversity of local gene pools has been narrowed down through various genetic bottlenecks, such as domestication from wild species, adaptability to local environmental conditions including country and latitudinal levels, and human demands in local culture (Tenaillon et al. 2004; Hyten et al. 2006; Haudry et al. 2007). Crop domestication has produced dramatic changes in morphological traits of plant architecture to meet human demands (Doebley et al. 2006; Ross-Ibarra et al. 2007; Purugganan and Fuller 2009). Therefore, it is unclear which genetic mechanisms drive the evolution of the wide adaptability of crops in agriculture.
Asian cultivated rice, Oryza sativa L., was domesticated from O. rufipogon around 10,000 years ago (Fuller 2011; Huang et al. 2012; Yang et al. 2012; Choi et al. 2017) (Fig. 1a). Rice is the principal staple food responsible for feeding people worldwide. The geographic distribution of O. rufipogon is limited to tropical and subtropical Asia, around 10–20°N, (Khush 1997), whereas cultivated rice is distributed across an extremely wide range of climatic conditions, from 53°N to 40°S latitude, due to the extensive efforts of rice breeding programs and through human migration all over the world (Lu and Chang 1980; Agrama et al. 2010). Hokkaido (41°2–45°3N latitude) is the northern-most region of Japan and one of the northern-limits of rice cultivation (Fig. 1a). The climatic conditions of Hokkaido, including a natural daylength of more than 15 h, are severe in terms of rice growth. During the last century, genetic improvements in varieties and the modernization of cultivation systems have enabled the stable production of rice in Hokkaido (Shinada et al. 2014; Fujino et al. 2015, 2017).
Extremely early flowering time of rice varieties in Hokkaido. a Locations of Ganges and Yangtze, where domestication of Asian cultivated rice (closed circle) started, and one of the northern-limits of rice cultivation in the world, Hokkaido (open circle). b Days to heading (DTH) of Nipponbare (NP), Koshihikari (KSH), Akitakomachi (AKT), and Hoshinoyume (HS) in natural field conditions in Hokkaido, 2012. ND; not determined due to lateness, > 130 days. Different letters indicate significant differences (P < 0.05; Tukey’s significant difference test). c Rice varieties HS (left) and NP (right) grown for 130 days in a natural paddy field in Hokkaido
Flowering time control may play a role in adaptive responses to changing environmental conditions. Photoperiodic sensitivity is the most important factor in determining flowering time (expressed as the heading date in crops), which ensures crop adaptation to specific ecological conditions and environmental variations. Mutations generated during the expansion of crop cultivation, including maize, wheat, and barley, may have accelerated the variation in flowering time and driven the spread of such crops all over the world (Turner et al. 2005; Beales et al. 2007; Faure et al. 2012; Hung et al. 2012; Zakhrabekova et al. 2012; Yang et al. 2014; Huang et al. 2018).
Because of its small genome size and importance as a model crop, the molecular and genetic networks of flowering time control in short-day rice have been proposed (Tsuji et al. 2011; Hori et al. 2016). However, it was unclear which genes contribute to the adaptability to local environmental conditions. Varieties cultivated in Hokkaido have unique characteristics, including extremely low photoperiod sensitivity to avoid low-temperature stress at the maturing stage in autumn in Hokkaido (Tanisaka et al. 1992; Okumoto et al. 1996). Previously, we demonstrated the genetic basis of extremely early flowering time involving the loss of two photoperiod sensitivity genes, qDTH7-1/Ghd7 (Xue et al. 2008) and qDTH7-2/Hd2/OsPRR37 (Koo et al. 2013; Gao et al. 2014) was significant for the adaptability of rice to cultivation in Hokkaido (Fujino and Sekiguchi 2005a, b; Nonoue et al. 2008; Shibaya et al. 2011; Fujino et al. 2019).
Here, we demonstrated the evolutionary process of the establishment of genotypes for the extremely early flowering time with good fitness for agriculture. Four flowering time genes have played to the extremely early flowering time. First is Heading date 1 (Hd1), which is the rice ortholog of the Arabidopsis floral activator CONSTANS (Yano et al. 2000). Second is Grain number, plant height and heading date 7 (Ghd7) encoding a CCT (CO, CO-LIKE, and TIMING OF CAB1) domain protein (Xue et al. 2008). Third is Oryza sativa Pseudo-Response Regulator37 (OsPRR37), which is an ortholog of the circadian clock genes PRR3/7 in Arabidopsis (Nakamichi et al. 2005; Murakami et al. 2007; Koo et al. 2013; Gao et al. 2014). Finally, fourth is Days to heading 8 (DTH8) encoding a putative HAP3 subunit of the CCAAT-box-binding transcription factor (Wei et al. 2010). The extremely early flowering time has a pleiotropic effect on increased panicle number, thereby limiting grain yield. The fitness of the negative correlation between flowering time and panicle number could make rice cultivation possible in Hokkaido. Finally, we discuss the potential of current rice breeding programs for generating the novel genotype for flowering time, and for expanding rice cultivation areas.
Materials and methods
Plant material and growth conditions
Three rice populations were used in this study. First, 50 varieties from the Japanese rice core collection (JRC) were used (Table S1), which represented the genetic diversity among the ancestral gene pool of varieties bred in rice breeding programs in Japan (Ebana et al. 2008). Second, 43 varieties were collected from the Genebank, Japan (http://www.gene.affrc.go.jp/), as the Hokkaido Landrace Rice Panel (HLP) (Table S2). HLP involved varieties with adaptability to Hokkaido (aHLP) and without adaptability to Hokkaido (nHLP). Because these varieties were stored as landraces in Hokkaido, it was unclear which varieties exhibit the adaptability. The third population was 62 varieties of the Hokkaido Rice Core Panel (HRCP) representing the genetic diversity of the 100-year span of rice breeding programs in Hokkaido (Shinada et al. 2014; Fujino et al. 2015a, b, 2017) (Table S3). In addition, six near isogenic lines (NILs) for genes controlling flowering time were used as a series with Hoshinoyume (HS) as a genetic background. HSGhd7, HShd1, and HShd5 have already been described (Fujino et al. 2013, 2019). Hd5 is an allelic to DTH8 (Wei et al. 2010; Fujino et al. 2013). HSOsPRR37 was derived from repeated backcrossing between HS and Arroz Da Terra with marker-assisted selection (MAS) using RM1306 linked to OsPRR37 (Fujino and Sekiguchi 2005). Three rice varieties, Akitakomachi (AKT) from Akita, Koshihikari (KSH) from Niigata, and Nipponbare (NP) from Aichi, which are popular in Japan, were used as controls (Fig. S1).
All rice materials were cultivated in experimental paddy fields at Hokkaido Agricultural Research Center, Sapporo, Japan (43°00′N latitude), in accordance with Fujino et al. (2015a, b). Sowing and transplanting were performed in late April and late May, respectively. All materials were planted with a 15.0-cm spacing between plants within each row and 30.0-cm spacing between rows. Cultivation management followed the standard procedures used at Hokkaido Agricultural Research Center.
Phenotype evaluation
Seven agronomic traits (Table S4), days to heading (DTH), culm length (CL), panicle length (PL), panicle number (PN), 1000-grain weight (TGW), low-temperature germination (LTG), and seeds per panicle (SP), were measured in accordance with Fujino et al. (2017).
DNA analysis
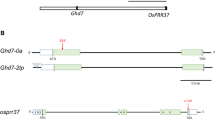
Total DNA was isolated from young leaves using the CTAB method (Murray and Thompson 1980). A total of 36 Indel markers (Fujino et al. 2015) were used for phylogenomic analysis with the JRC and HLP cultivars (Table S5). The genotype of three genes, Hd1, OsPRR37, and Ghd7 controlling flowering time, were determined by PCR, STS, and CAPS, targeted for functional nucleotide polymorphism (Tables 1, S6). PCR and sequencing were performed as described by Fujino et al. (2004, 2005).
Data analysis
Phylogenetic analysis with the PHYLIP software suite (Felsenstein 1989) and STRUCTURE program ver. 2.3.4 (Pitchard et al. 2000; Evanno et al. 2005) was performed as described by Shinada et al. (2014). The polymorphism information content (PIC) was calculated for each marker in accordance with Nei (1973): PIC = 1 − Σhk2, where hk is the frequency of kth allele.
Results
Extremely early flowering time in rice varieties adapted to Hokkaido
To characterize the flowering time of varieties from different local breeding programs, four rice varieties, HS, AKT, NP, and KSH, were grown in natural environmental conditions in Hokkaido. In the paddy field in Tsukuba, Japan (36.3°N), the flowering time of HS and NP were approx. 80 and 110 days, respectively (Nonoue et al. 2008). Whereas only HS could exhibit flowering time to mature rice seeds in agriculture, 95.0 of DTH (Fig. 1b, c). Flowering time of AKT was 108.6 days, and the flowering times of KSH and NP were more than 130 days (Fig. 1b, c).
Genetic population structure of rice varieties adapted to Hokkaido
Phylogenetic analysis was conducted to understand the genetic relationship of two different populations, the JRC and HLP collections. A UPGM dendrogram using Nei’s distance showed that there were three groups, I, II, and III (Fig. 2). The JRC was divided into two major groups, groups II and III, which corresponded to upland and lowland habitats, respectively (Kojima et al. 2005). HLP varieties were clearly distinguished from those in the JRC. Most varieties in the HLP, 30 of 43, were classified into group I. Three and nine varieties were classified into groups II and III, respectively. The three populations with K = 3 in the STRACTURE analysis (Fig. S2) corresponded well with the three groups obtained by phylogenetic analysis. Degree on admixtures of genetic structures between each group, I/II and II/III, were observed, showing genetic relationships among each group. Genetic diversity represented as PIC varied (Table S7). The PIC value of group I, 0.16, was lower than those of group II, 0.25, and group III, 0.32, suggesting that there may have been genetic bottlenecks caused by intensive artificial selection on natural variation adapted to Hokkaido.
UPGMA dendrogram and population structures of varieties in the JRC and the HLP with controls, Nipponbare (NP) and Hoshinoyume (HS). The varieties were divided into three groups, I–III. Triangles and squares indicate the growth habits, lowland and upland, of varieties in the JRC, respectively. Circles indicate varieties in the HLP. Colors indicated the genotype of the two genes, Ghd7 and OsPRR37; ghd7osprr37 (red), Ghd7osprr37 (orange), ghd7OsPRR37 (green) and Ghd7OsPRR37 (blue). Numbers following the figure indicate the serial number of each population. (Color figure online)
Phenotypic variation among HLP varieties
Genetic differences between ancestral and selected individuals (Fig. 2) may contribute to phenotypic yield differences in agriculture. To elucidate which traits were targeted by artificial selection, seven traits were evaluated among the HLP varieties (Table S8, Fig. 3). A wide range of phenotypic variation was observed, with continuous distributions in five traits, whereas relatively bimodal distributions were observed in DTH and CL, which are pleiotropic and interact in each other.
Next, we determined the genotype for flowering time, Ghd7 and OsPRR37, which are known to be involved in the expression of extremely early flowering time (Fujino and Sekiguchi 2005a, b; Fujino et al. 2019). The genotype was clearly correlated with phenotypic variation in DTH among HLP varieties and phylogenetic classification (Fig. 2, Table S9). Most varieties in group I, 21/30, had ghd7osprr37. Whereas ghd7OsPRR37 and Ghd7osprr37 were only present in four and five varieties in group I. Ghd7OsPRR37 was not found in group I.
To investigate whether variation has the potential to confer a selectable fitness advantage, all seven agronomic traits were compared between varieties with/without ghd7osprr37. We found that significant differences were detected in only DTH and CL (Table S10). Agronomic traits are results of interactions between QTLs or between QTLs and the particular genetic and environmental background (Li et al. 2003; Ando et al. 2008). In addition, genes for flowering time control the traits for yield (Xue et al. 2008; Endo-Higashi and Izawa 2011; Koo et al. 2013; Gao et al. 2014; Tsuji et al. 2015). We found relatively low coefficient values between DTH/CL and other traits, − 0.37 < r < 0.47 (Table S11) among aHLP. We could not find a significant correlation of DTH with agronomic traits among varieties with ghd7osprr37, indicating that extremely early flowering time had no drastic changes in phenotype.
Phenotype of NILs for flowering time
To elucidate the role of genes for flowering time on agronomic traits, we compared six agronomic traits among six NILs (Tables 2, S12). The DTH of each line was significantly different depending on the genotype for flowering time. The NILs did not exhibit drastic changes in plant architecture along with genotype or DTH. In addition, each NIL exhibited significant differences in the agronomic traits compared with HS of the genetic background (Table 2).
The selection of the best suitable lines for local environmental conditions could have pleiotropic effects on not only flowering time but also grain yield in agriculture. Compared with HS, functional Ghd7 elongated PL and reduced PN and functional OsPRR37 and loss-of-function hd1 reduced PN significantly (Table 2). Whereas loss-of-function hd5 increased PN and SP and reduced PL and CL, HShd5 exhibited the earliest flowering time among the NILs (Table 2). There were trends for alterations in agronomic traits along with DTH (Table S13). Depending on DTH, three traits and PN were positively and negatively correlated with higher coefficient values (Table S13). Rice panicle phenotypes including PN limit yield and grain quality and are under selection in rice breeding programs. NILs for earlier flowering time tend to have greater PN.
Genes for flowering time may coordinately regulate DTH and PN as yield potential (Wang and Li 2008; Liang et al. 2014; Wang et al. 2015a, b). Variations in flowering time and yield may be intensively selected to maximize grain production for local environmental conditions. Pleiotropic effects of genes for flowering time on yield impacted on adaptation to new local environmental conditions.
Dramatic changes in genotype for flowering time during rice cultivation in Hokkaido
Only two varieties from the JRC representing genetic diversity across Japan, 48/50, carried ghd7osprr37, which were collected from Hokkaido. Whereas among 43 varieties in the HLP, 27 (62.8%) carried ghd7osprr37 (Fig. 4a, Table S14). In addition, most varieties from the HRCP, 49/62 (80.3%) carried ghd7osprr37. Given the need for meeting the demands of society, extremely early flowering time expressed by ghd7osprr37 was selected for adaptability to Hokkaido.
Dramatic changes in allele frequencies in the genotypes of genes for flowering time among the populations and between generations. a Frequency (%) of the combination of Ghd7 and OsPRR37, among the three populations. Boxes indicate the four genotypes; Ghd7OsPRR37 (white), Ghd7osprr37 (hatched), ghd7OsPRR37 (gray), and ghd7osprr37 (black). b Frequency (%) of alleles in Hd1 among the three populations. c Change in allele frequency (%) in Hd1 among HRCP between four generations. White, black, and gray boxes indicate Akage (AK), Ginbouzu (G), and other alleles of Hd1, respectively
Loss-of-function allele hd1 of the AK allele was infrequent in JRC varieties, 4 of 37, whereas most the HLP population carried the AK allele, 32 of 43 varieties (Fig. 4b, Table S15). In addition, a high frequency of hd1, five among seven varieties, was observed in the early phase of rice breeding programs in Hokkaido during 1900–1940 (Fig. 4c, Table S16). Whereas all varieties, 11/11, released during 1991–2010 carried functional Hd1. Previously, we showed that functional Hd1 promotes flowering time in the genotype of ghd7osprr37 in long-day conditions (Fujino et al. 2019). Once functional Hd1, the G allele, was introduced into the gene pool in to Hokkaido varieties from Honshu varieties carrying the Hd1, it was immediately dispersed within the population.
The increase in the genotype of ghd7osprr37 during rice cultivation in Hokkaido and the functional allele Hd1 during rice breeding programs in Hokkaido suggested an effect of direct selection for flowering earliness on the genes, which are the main drivers of adaptation to northern regions with naturally long daylength.
Discussion
The middle area of the Pearl River, China, was the first place where rice cultivation was developed (Huang et al. 2012). Cultivated rice has since spread over diverse geographic regions. Here, we demonstrated the genetic and phenotypic bases of the establishment of local rice varieties from Hokkaido, a northern-limit of rice cultivation. Artificial selection on natural variation and reconstruction of genotypes for flowering time by plant breeding programs may optimize flowering time for better fitness for agriculture. Genetic variations among local populations are limited due to the lack of recombination from the inbreeding of cultivated rice, which can generate adaptability rapidly. Once desirable traits/genes are introduced into populations in new environments, they immediately spread within the population.
The genetic structure and phenotypic shifts (Figs. 2, 3, Table S10) in the population from Hokkaido clearly provide insights into the significant contributions of genetically induced mutations for phenotypic variation. Our results clearly proposed a hypothetical model on the establishment of early flowering time (Figs. 5, S3, Table S17). In the initial step in the direction to early flowering time following intensive artificial selection along with human migration all over the world, loss-of-function hd1 generated varieties with early flowering time. Then, the Southern–Northern distribution among Japanese rice was regulated by Hd1 versus hd1 (Yokoo and Kikuchi 1977; Yokoo et al. 1980). Subsequently, the Northern distribution with hd1, which showed extremely early flowering time, was generated by loss-of-function ghd7 and osprr37 selected together.
Model of the establishment process in rice varieties with early flowering time. a Relationships with the mutations for population structure in Japan. Both functional and loss-of-function alleles in Hd1 were distributed across Japan. Two mutations for loss-of-function in Ghd7 and OsPRR37 generated group I for early flowering time. The mutation for loss-of-function in Hd5 happened during the initial phase of rice breeding programs in Hokkaido (Fujino et al. 2013). Red, green, orange, and blue stars indicate mutation events of flowering time genes. Red arrows indicated the introgression of functional Hd1 into rice breeding programs in Hokkaido using exotic germplasm. b Changes of the genotype for early flowering time in three different populations. 3M; triple mutant, 4M; quadruple mutant, EARLY-t; EARLY TRIO, EARLY-q; EARLY QUARTETTE. Red and blue bold figures indicated the loss-of-function mutation allele and the functional allele transferred by rice breeding programs, respectively. (Color figure online)
Adaptability can be defied by various kinds of phenotype. Optimization of flowering time to maximize yield has been shown to adaptively differentiate plant populations from different environments. Flowering time variation correlated with fitness played as indicators of adaptation to different environments. Earlier flowering time, short vegetative phase, fewer seeds, and small reproductive phase. The degree of adaptation differed across measures of fitness components. The early flowering time with greater PN might be a break-through in fitness for agriculture.
During the process for earlier flowering time (Fig. 4), functional Hd1, which was lost in the initial phase of selection for earlier flowering (Ebana et al. 2011; Hori et al. 2015), was introgressed into the local gene pool with hd1. Furthermore, intensive selection focusing on Hd1 for earlier flowering time exhibited not only early flowering but also greater tillering, tuning the fitness of yield (Table 2). Hd1, a ortholog of CONSTANS in Arabidopsis, plays a central role in flowering time control (Yano et al. 2000). Previously, we demonstrated that the introgression around Hd1 of japonica into the proto-aus group might generate aus subspecies, which is an ecotype with early flowering time (Fujino et al. 2010). This is generally consistent with differentiation among cultivated rice O. sativa, L. by global population genomics (Huang et al. 2012). The aus subspecies has a small geographic distribution derived from a cross between japonica and local wild rice, proto-aus (Huang et al. 2012). In addition, loss-of-function hd1 alleles played a role in the adaptation to northern areas (Ebana et al. 2011; Hori et al. 2015). Furthermore, a functional Hd1 allele might achieve stable rice production at the northern-limit of rice cultivation. Taken together, Hd1 might have been an evolutionary force for the further expansion of cultivated rice worldwide.
Understanding of the genetic mechanisms found in the adaptability to a northern-limit to cultivation can facilitate adaptive divergence within genetically improved local populations, thereby expanding the species range. The loss-of-function alleles in Ghd7 and OsPRR37 could split off a variety group from the JRC. Intensive selection on natural variation in flowering time might drive this adaptability. The introduction of functional Hd1, which was lost during the process for earlier flowering time, could achieve a stable rice production with ghd7osprr37. The genotype of the combination of Hd1 with ghd7osprr37 may be rare or impossible in natural variation among cultivated rice with inbreeding, achieving stable rice production. Plant breeding programs could generate novel genotypes other than those in natural populations, exploring phenotypic variation.
References
Agrama H, Yan W, Jia M, Fjellstrom R, McClung A (2010) Genetic structure associated with diversity and geographic distribution in the USDA rice world collection. Nat Sci 2:247–291
Ando T, Yamamoto T, Shimizu T, Ma XF, Shomura A, Takeuchi Y, Lin SY, Yano M (2008) Genetic dissection and pyramiding of quantitative traits for panicle architecture by using chromosomal segment substitution lines in rice. Theor Appl Genet 116:881–890
Beales J, Turner A, Griffiths S, Snape J, Laurie D (2007) A pseudo-response regulator is misexpressed in the photoperiod insensitive Ppd-D1a mutant of wheat (Triticum aestivum L.). Theor Appl Genet 115:721–733
Choi JY, Platts AE, Fuller DQ, Hsing YI, Wing RA, Purugganan MD (2017) The rice paradox: multiple origins but single domestication in Asian rice. Mol Biol Evol 34:969–979
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321
Ebana K, Kojima Y, Fukuoka S, Nagamine T, Kawase M (2008) Development of mini core collection of Japanese rice landrace. Breed Sci 58:281–291
Ebana K, Shibaya T, Wu J, Matsubara K, Kanamori H et al (2011) Uncovering of major genetic factors generating naturally occurring variation in heading date among Asian rice cultivars. Theor Appl Genet 122:1199–1210
Endo-Higashi N, Izawa T (2011) Flowering time genes Heading date 1 and Early heading date 1 together control panicle development in rice. Plant Cell Physiol 52:1083–1094
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Faure S, Turner AS, Gruszka D, Christodoulou V, Davis SJ, von Korff M, Laurie DA (2012) Mutation at the circadian clock gene EARLY MATURITY 8 adapts domesticated barley (Hordeum vulgare) to short growing seasons. Proc Natl Acad Sci USA 109:8328–8333
Felsenstein J (1989) PHYlIP-phylogeny inference package (version 3.2). Cladistics 5:164–166
Fujino K, Sekiguchi H (2005a) Mapping of QTLs conferring extremely early heading in rice (Oryza sativa L.). Theor Appl Genet 111:393–398
Fujino K, Sekiguchi H (2005b) Identification of QTLs conferring genetic variation for heading date among rice varieties at the northern-limit of rice cultivation. Breed Sci 55:141–146
Fujino K, Sekiguchi H, Sato T, Kiuchi H, Nonoue Y, Takeuchi Y, Ando T, Lin SY, Yano M (2004) Mapping of quantitative trait loci controlling low-temperature germinability in rice (Oryza sativa L.). Theor Appl Genet 108:794–799
Fujino K, Sekiguchi H, Kiguchi T (2005) Identification of an active transposon in intact rice plants. Mol Gen Genet 273:150–157
Fujino K, Wu J, Sekiguchi H, Ito T, Izawa T, Matsumoto T (2010) Multiple introgression events surrounding the Hd1 flowering-time gene in cultivated rice, Oryza sativa L. Mol Genet Genom 284:137–146
Fujino K, Yamanouchi U, Yano M (2013) Roles of the Hd5 gene controlling heading date for adaptation to the northern limits of rice cultivation. Theor Appl Genet 126:611–618
Fujino K, Obara M, Shimizu T, Koyanagi KO, Ikegaya T (2015a) Genome-wide association mapping focusing on a rice population derived from rice breeding programs in a region. Breed Sci 65:403–410
Fujino K, Obara M, Ikegaya T, Tamura K (2015b) Genetic shift in local rice populations during rice breeding programs in the northern limit of rice cultivation in the world. Theor Appl Genet 128:1739–1746
Fujino K, Nishimura T, Kiuchi H, Hirayama Y, Sato T (2017) Phenotypic changes during 100-year rice breeding programs in Hokkaido. Breed Sci 67:528–534
Fujino K, Yamanouchi U, Nonoue Y, Obara M, Yano M (2019) Switching genetic effects of the flowering time gene Hd1 under LD conditions by Ghd7 and OsPRR37 in rice. Breed Sci. https://doi.org/10.1270/jsbbs.18060
Fuller DQ (2011) Pathways to Asian civilizations: tracing the origins and spread of rice and rice cultures. Rice 4:78–92
Gao H, Jin M, Zheng XM, Chen J, Yuan D, Xin Y, Wang M, Huang D, Zhang Z, Zhou K et al (2014) Days to heading 7, a major quantitative locus determining photoperiod sensitivity and regional adaptation in rice. Proc Natl Acad Sci USA 111:16337–16342
Haudry A, Cenci A, Ravel C, Bataillon T, Brunel D, Poncet C, Hochu I, Poirier S, Santoni S, Glemin S, David J (2007) Grinding up wheat: a massive loss of nucleotide diversity since domestication. Mol Biol Evol 24:1506–1517
Hori K, Nonoue Y, Ono N, Shibaya T, Ebana K, Matsubara K, Ogiso-Tanaka E, Tanabata T, Sugimoto K, Taguchi-Shiobara F et al (2015) Genetic architecture of variation in heading date among Asian rice accessions. BMC Plant Biol 15:115
Hori K, Matsubara K, Yano M (2016) Genetic control of flowering time in rice: integration of Mendelian genetics and genomics. Theor Appl Genet 129:2241–2252
Huang X, Kurata N, Wei X, Wang ZX, Wang A, Zhao Q, Zhao Y, Liu K, Lu H, Li W, Guo Y, Lu Y, Zhou C, Fan D, Weng Q, Zhu C, Huang T, Zhang L, Wang Y, Feng L, Furuumi H, Kubo T, Miyabayashi T, Yuan X, Xu Q, Dong G, Zhan Q, Li C, Fujiyama A, Toyoda A, Lu T, Feng Q, Qian Q, Li J, Han B (2012) A map of rice genome variation reveals the origin of cultivated rice. Nature 490:497–501
Huang C, Sun H, Xu D, Chen Q, Liang Y, Wang X, Xu G, Tian J, Wang C, Li D, Wu L, Yang X, Jin W, Doebley JF, Tian F (2018) ZmCCT9 enhances maize adaptation to higher latitudes. Proc Natl Acad Sci USA 115:E334–E341
Hung HY, Shannon LM, Tian F, Bradbury PJ, Chen C, Flint-Garcia SA, McMullen MD, Ware D, Buckler ES, Doebley JF, Holland JB (2012) ZmCCT and the genetic basis of day-length adaptation underlying the postdomestication spread of maize. Proc Natl Acad Sci USA109:E1913–E1921
Hyten DL, Song Q, Zhu Y, Choi IY, Nelson RL, Costa JM, Specht JE, Shoemaker RC, Cregan PB (2006) Impacts of genetic bottlenecks on soybean genome diversity. Proc Natl Acad Sci USA 103:16666–16671
Khush GS (1997) Origin, dispersal, cultivation and variation of rice. Plant Mol Biol 35:25–34
Kojima Y, Ebana K, Fukuoka S, Nagamine T, Kawase M (2005) Development of an RFLP-based rice diversity research set of germplasm. Breed Sci 55:431–440
Koo BH, Yoo SC, Park JW, Kwon CT, Lee BD, An G, Zhang Z, Li J, Li Z, Paek NC (2013) Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes. Mol Plant 6:1877–1888
Li X, Qian Q, Fu Z, Wang Y, Xiong G, Zeng D, Wang X, Liu X, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li J (2003) Control of tillering in rice. Nature 422:618–621
Liang WH, Shang F, Lin QT, Lou C, Zhang J (2014) Tillering and panicle branching genes in rice. Gene 537:1–5
Lu JJ, Chang TT (1980) Rice in its temporal and spatial perspectives. In: Luh BS (ed) Rice: production and utilization. AVI Publishing Co., Inc., Westport, pp 1–74
Murakami M, Tago Y, Yamashino T, Mizuno T (2007) Comparative overviews of clock-associated genes of Arabidopsis thaliana and Oryza sativa. Plant Cell Physiol 48:110–121
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acid Res 8:4321–4325
Nakamichi N, Kita M, Ito S, Yamashino T, Mizuno T (2005) PSEUDO-RESPONSE REGULATORS, PRR9, PRR7 and PRR5, together play essential roles close to the circadian clock of Arabidopsis thaliana. Plant Cell Physiol 46:686–698
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nonoue Y, Fujino K, Hirayama Y, Yamanouchi U, Lin SY, Yano M (2008) Detection of quantitative trait loci controlling extremely early heading in rice. Theor Appl Genet 116:715–722
Okumoto Y, Ichitani K, Inoue H, Tanisaka T (1996) Photoperiod insensitivity gene essential to the varieties grown in the northern limit region of paddy rice (Oryza sativa L.) cultivation. Euphytica 92:63–66
Pitchard JK, Stephens M, Donnely P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Purugganan MD, Fuller DQ (2009) The nature of selection during plant domestication. Nature 457:843–848
Ross-Ibarra J, Morrell PL, Gaut BS (2007) Plant domestication, a unique opportunity to identify the genetic basis of adaptation. Proc Natl Acad Sci USA 104(Suppl 1):8641–8648
Shibaya T, Nonoue Y, Ono N, Yamanouchi U, Hori K, Yano M (2011) Genetic interactions involved in the inhibition of heading by heading date QTL, Hd2 in rice under long-day conditions. Theor Appl Genet 123:1133–1143
Shinada H, Yamamoto T, Yamamoto E, Hori K, Yonemaru J, Matsuba S, Fujino K (2014) Historical changes in population structure during rice breeding programs in the northern limits of rice cultivation. Theor Appl Genet 127:995–1004
Tanisaka T, Inoue H, Uozu S, Yamagata H (1992) Basic vegetative growth and photoperiod sensitivity of heading-time mutants induced in rice. Jpn J Breed 42:657–668
Tenaillon MI, U’Ren J, Tenaillon O, Gaut BS (2004) Selection versus demography: a multilocus investigation of the domestication process in maize. Mol Biol Evol 21:1214–1225
Tsuji H, Taoka K, Shimamoto K (2011) Regulation of flowering in rice: two florigen genes, a complex gene network, and natural variation. Curr Opin Plant Biol 14:45–52
Tsuji H, Tachibana C, Tamaki S, Taoka K, Kyozuka J, Shimamoto K (2015) Hd3a promotes lateral branching in rice. Plant J 82:256–266
Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034
Wang Y, Li J (2008) Molecular basis of plant architecture. Ann Rev Plant Biol 59:253–279
Wang L, Lu Q, Wen X, Lu C (2015a) Enhanced sucrose loading improves rice yield by increasing grain size. Plant Physiol 169:2848–2862
Wang L, Sun S, Jin J, Fu D, Yang X, Weng X, Xu C, Li X, Xiao J, Zhang Q (2015b) Coordinated regulation of vegetative and reproductive branching in rice. Proc Natl Acad Sci USA 112:15504–15509
Wei X, Xu J, Guo H, Jiang L, Chen S, Yu C, Zhou Z, Hu P, Zhai H, Wan J (2010) DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol 153:1747–1758
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X, Zhang Q (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yang J, Zhao X, Cheng K, Du H, Ouyang Y, Chen J, Qiu S, Huang J, Jiang Y, Jiang L, Ding J, Wang J, Xu C, Li X, Zhang Q (2012) A killer-protector system regulates both hybrid sterility and segregation distortion in rice. Science 337:1336–1340
Yang S, Murphy RL, Morishige DT, Klein PE, Rooney WL, Mullet JE (2014) Sorghum phytochrome B inhibits flowering in long days by activating expression of SbPRR37 and SbGHD, repressors of SbEHD1, SbCN8 and SbCN12. PLoS One 9:e105352
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12:2473–2484
Yokoo M, Kikuchi F (1977) Multiple allelism of the locus controlling heading time of rice, detecting using close linkage with blast-resistance. Jpn J Breed 21:123–130
Yokoo M, Kikuchi F, Nakane A, Fujimaki H (1980) Genetical analysis of heading time by aid of close linkage with blast, Pyricularia oryzae, resistance in rice. Bull Natl Inst Agric Sci Ser D 31:95–126
Zakhrabekova S, Gough SP, Braumann I, Muller AH, Lundqvist J, Ahmann K, Dockter C, Matyszczak I, Kurowska M, Druka A, Waugh R, Graner A, Stein N, Steuernagel B, Lundqvist U, Hansson M (2012) Induced mutations in circadian clock regulator Mat-a facilitated short-season adaptation and range extension in cultivated barley. Proc Natl Acad Sci USA 109:4326–4331
Acknowledgements
This work was supported in part by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries and Food Industry) (to KF) and JSPS KAKENHI Grant number 25450015 (to KF).
Funding
This work was supported in part by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Science and Technology Research Promotion Program for Agriculture, Forestry, Fisheries and Food Industry) (to KF) and JSPS KAKENHI Grant number 25450015 (to KF).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments and wrote the manuscript: KF. Performed the experiments, analyzed the data, and approved the final manuscript: TI, MO, KF.
Corresponding author
Ethics declarations
Conflict of interest
All author declares that he/she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Bing Yang.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fujino, K., Obara, M. & Ikegaya, T. Establishment of adaptability to the northern-limit of rice production. Mol Genet Genomics 294, 729–737 (2019). https://doi.org/10.1007/s00438-019-01542-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-019-01542-2