Abstract
Blastocystis is a unicellular, anaerobic protist inhabiting the intestinal tract of diverse animal hosts, including human. Information regarding Blastocystis in small ruminants, namely goats and sheep, is limited globally; thus, this study was carried out to investigate the distribution and determinants of Blastocystis in ruminant livestock animals from Penang, Malaysia. Fecal samples from 127 cattle, 149 goats, and 100 sheep were examined for Blastocystis by in vitro cultivation using modified Jones’ medium, while DNA barcoding was used for subtyping. Overall, 23.1% (87/376) of animals screened were positive for Blastocystis sp. The prevalence of infection was significantly higher in goats than in cattle and sheep, while the female gender, semi-intensive farming system, and the Northeast Penang Island district were identified as potential risk factors for Blastocystis infection. Blastocystis sp. ST5, ST14, and ST25 were identified in cattle; ST5, ST10, ST13, and ST14 in goats; and ST4, ST5, ST14, and ST15 in sheep. ST5 and ST14 were found to be the most abundant and widespread subtypes in the study area. To the best of our knowledge, this is the first report of ST4 from sheep and ST13 from goats, thus serving as an update to the host range of Blastocystis sp. ST4 and ST13. The isolation of ST4 and ST5 in this study suggests that ruminant livestock animals could serve as reservoirs of human infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Blastocystis sp. is a unicellular protist that belongs to the Stramenopiles, a complex assemblage of “botanical Protists” (Silberman et al. 1996; Ahmed and Karanis 2018). It has been detected worldwide in diverse animal hosts including humans and is referred to as the most frequently observed enteric eukaryotic symbiont in mammals and birds (Tan 2008; Adao and Rivera 2018). The transmission of this organism is by the fecal–oral route and mainly via the ingestion of feces-contaminated food and water (Clark et al. 2013); animal-to-human and human-to-animal transmission can occur (Hublin et al. 2020). Blastocystis is a fascinating organism; it displays morphological polymorphism and extensive genetic diversity, and its role as either a commensal or parasite in the host gut is controversial (Lepczyńska et al. 2017). Based on the phylogeny of the small subunit ribosomal RNA (SSU rRNA) gene, at least 32 genetic lineages (subtypes) of Blastocystis have been proposed in a wide range of hosts including humans, other mammals, and birds, and at least 28 subtypes (ST1–ST17, ST21, and ST23–ST32) are generally accepted as valid subtypes (Liu et al. 2022). The presence of Blastocystis sp. ST1-ST10, ST12, ST14, and ST16 reported in humans (Khaled et al. 2021; Osorio-Pulgarin et al. 2021) have also been reported in various animal hosts indicating the possible occurrence of zoonotic transmission (Higuera et al. 2021; Rauff-Adedotun et al. 2021).
Livestock-associated infectious agents have become a major threat to human health with veterinarians, slaughterhouse workers, and farmers at higher risk of infection (Klous et al. 2016). Irrespective of farming system employed in livestock production, livestock animals worldwide have been reported as hosts to Blastocystis (Hublin et al. 2020). Nevertheless, information regarding Blastocystis in small ruminants, namely goats and sheep, is still scarce globally with most reports involving a small number of samples (Hublin et al. 2020). In Malaysia, the livestock industry is continually expanding especially as ruminants livestock production in Malaysia is inadequate to meet consumer demands (International Trade Administration 2021). However, a few studies have illustrated the presence of Blastocystis sp. in ruminant livestock animals (Tan et al. 2013; Noradilah et al. 2017; Mohammad et al. 2018; Abd Razak et al. 2019; Kamaruddin et al. 2020), with even fewer reporting on the subtypes present. From available studies from Malaysia, Blastocystis sp. ST1, ST3, ST4, ST5, ST10, and ST14 were reported in cattle (Mohammad et al. 2018; Kamaruddin et al. 2020), while ST1, ST3, ST4, ST6, ST7, ST8, and ST10 were reported in goats (Tan et al. 2013; Noradilah et al. 2017). There are no reports on Blastocystis sp. subtypes in sheep in Malaysia. Subtype analysis of Blastocystis infection is essential to understand the distribution, and transmission dynamics of this organism. Notably, it is common to find surveys reporting on risk factors or predictors of Blastocystis sp. infection in humans (Lee et al. 2012; Osman et al. 2016; Hidalgo et al. 2019; Deng et al. 2020); this is rarely the case in animal surveys. Thus, the aim of this study is to describe the prevalence, potential risk factors, and subtypes of Blastocystis sp. in cattle, goats, and sheep from Penang, Malaysia.
Materials and methods
Sampling
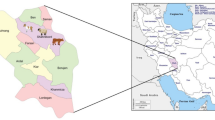
This survey was conducted in Penang, Malaysia (Fig. 1). It occupies a total area of 1048 km2 and situated at 5°24′52.2″N 100°19′45.12″E. Sampling was carried out in 40 ruminant livestock farms across four districts of Penang (Northeast Penang Island, Southwest Penang Island, North Seberang Perai, South Seberang Perai). These farms rely on the Department of Veterinary Services (DVS) Penang for health and extension services. Ruminant livestock animal groups included in this research were cattle (Bos taurus), goat (Capra hircus), and sheep (Ovis aries). Data related to the study animals (gender, age, production purpose, and fecal type) and the farms (the livestock farm management system—intensive, semi-intensive, or extensive) were recorded. The intensive farm management system rears its livestock in zero-grazing units, with feed and water adequately provided. Animals reared under the semi-intensive system are allowed to graze pasture during the day and kept in the shelter at night with supplementary feeding. The extensive system allows animals to graze during the day and night.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. Animal ethical approval was obtained from USM Institutional Animal Care and Use Committee (USM IACUC), Universiti Sains Malaysia. Meanwhile, permission for sampling activities was obtained from the DVS Penang and the Department of Veterinary Services, Ministry of Agriculture and Agro-based Industry Malaysia.
Sample collection
Animals involved in this study were chosen randomly, according to convenience, from farms scheduled for visitation by the DVS, from January to February 2021. All animals appeared healthy. Fresh fecal samples of ruminant livestock animals were collected directly from the rectum of selected animals; and when this method of collection was not achievable, freshly voided fecal samples were carefully collected from the ground. Animals from which samples had been obtained were marked to avoid multiple examination of same animal. The fecal samples were placed into labeled screw-cap stool collection containers and transported immediately to the USM Veterinary Parasitology Laboratory, School of Biological Sciences, Universiti Sains Malaysia, Penang, Malaysia, for Blastocystis examination. Information on study animals and farms involved in this study is given in Table 1.
In vitro cultivation
A pea-sized amount of each freshly collected fecal sample or caecal content was inoculated into a sterile screw-top tube containing 3 ml of Jones’ medium supplemented with 10% heat-activated horse serum as described by Suresh and Smith (2004). Each sample was incubated vertically at 37 °C and examined after 48 h for the presence of various forms of Blastocystis sp. by placing a drop of the sediment onto a microscope slide and viewing it using a light microscope at 400 × magnification. Samples from which the various forms of Blastocystis sp. were observed were recorded as positive. Sediments from Blastocystis sp. positive cultures were stored at − 20 °C until molecular studies. However, samples from which no form of Blastocystis sp. was observed from first culture were sub-cultured and re-observed on the third day. Samples were then considered negative if no forms of Blastocystis sp. were observed on day 3 of subculture.
DNA barcoding
Genomic DNA was extracted from each Blastocystis sp.-positive culture using the Nucleospin DNA Stool Kit (Macherey–Nagel, Germany) according to the manufacturer’s protocol and stored at − 20 °C until polymerase chain reaction (PCR) analyses. DNA extracts were submitted to a single-step PCR for amplification of the barcode region of the SSU rRNA gene of Blastocystis using primers RD5 (5′-ATCTGGTTGATCCTGCCAGT-3′) and BhRDr (5′-GAGCTTTTTAACTGCAACAACG-3′) (Clark 1997; Scicluna et al. 2006).
The PCR was performed in a 50-µl reaction volume containing 25 µl of Vivantis 2 × Taq Master Mix, 2.5 mM MgCl2, 0.5 µl of each primer (at 10 µM primer concentration), and 2 µl of DNA. PCR conditions consisted of an initial denaturing step of 95 °C for 5 min, followed by 30 cycles of 95 °C for 1 min, 56.3 °C for 1 min 30 s, and 72 °C for 1 min, then followed by a final elongation step of 72 °C at 10 min. All PCR amplifications were completed with a Bio-Rad Thermo Cycler (USA). Electrophoresis was carried out using 1.5% agarose gel in Tris–Acetate-EDTA buffer. PCR products with visible bands of about 600 bp were sent to Apical Scientific, Malaysia, for purification and sequencing.
Blastocystis subtyping and phylogenetic analysis
The SSU rDNA sequences obtained were compared with published Blastocystis sp. homologous sequences resulting from Basic Local Alignment Search Tool (BLAST) calls in the National Center for Biotechnology Information (NCBI) database (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The subtypes were determined by exact matches or above 95% similarity with published reference sequences from the GenBank database. The SSU rDNA sequences obtained were also queried against the Blastocystis (18S) and Sequence Typing (MLST) databases (https://pubmlst.org/Blastocystis/) sited at the University of Oxford (Jolley et al. 2018) for identification to allele levels.
Nucleotide sequences obtained in this study and full-length reference nucleotide sequences for all accepted Blastocystis sp. subtypes obtained from the reference database (http://entamoeba.lshtm.ac.uk/blastorefseqs.htm) on 12 December 2021 were included in phylogenetic analysis. Complete SSU rRNA gene sequence of Proteromonas lacerate (U37108), an organism of close phylogenetic relation to Blastocystis sp., was also obtained to serve as the outgroup. Multiple sequence alignment was carried out with the Muscle algorithm in Mega 11 software (http://www.megasoftware.net/) (Tamura et al. 2021) and the alignment was edited to include only the barcode region. Evolutionary trees were reconstructed using the Neighbor-Joining (NJ) and Maximum-Likelihood (ML) methods by the Tamura-3-parameter substitution model using Mega 11. The reliability of the clades generated by the trees was assessed by bootstrap analysis with 1000 replicates.
Newly generated nucleotide sequences of the barcoding region of Blastocystis sp. SSU rRNA gene obtained in this survey were deposited in GenBank under the accession numbers ON738374-ON738382, ON738385-ON738404, and ON738409-ON738418.
Statistical analysis
Analysis of data was performed using the SPSS version 26.0 (SPSS Inc., Chicago, IL, USA). Frequency and percentages were used to present the prevalence of infections. Chi-square test (χ2) was used to compare the prevalence of Blastocystis sp. within host groups, age groups, genders, and other variables. Differences were considered statistically significant when p-values < 0.05. Bivariate logistic regression analysis was used to determine the associated risk factors for Blastocystis sp. infection. The p-values less than 0.05 was considered statistically significant.
Results
Prevalence and potential risk factors of Blastocystis sp. infection
A total of 376 ruminant livestock animals from 40 livestock farms were examined for Blastocystis sp. infection by in vitro cultivation. Farm animals involved were 127 cattle, 149 goats, and 100 sheep. Overall, 23.1% (87/376) of animals were examined positive for Blastocystis sp. The prevalence of infection was significantly higher in goats (35.6%) than in cattle and sheep (x2 = 21.545, p < 0.001) (Table 2). In general, 65% (26/40) of the livestock farms involved had one or more infected animals. Infection was the most common in sheep-rearing farms with 80% (8/10) of them having with one or more Blastocystis-infected; however, the frequency of livestock farms with infection based on type of animal reared did not vary significantly (x2 = 1.905, p = 0.386).
Within ruminant livestock animal groups, the prevalence of infection differed based on the different demographic characteristics (Table 3). Mostly, these variations were not of statistical significance. A significantly higher frequency of Blastocystis sp. infection was observed in cattle reared in a semi-intensive system than in the intensive husbandry system (x2 = 5.480, p = 0.019), and in goats raised within South Seberang Perai than those of other districts of Penang x2 = 10.111, p = 0.018). Likewise, sheep aged less than 1 year had a significantly lower prevalence of infection than those aged from 1 to 2 years old (x2 = 4.233, p = 0.040).
Results from bivariate logistic regression analysis of potential risk factors associated with Blastocystis sp. infection among ruminant livestock animals in Penang are presented in Table 4. Demographic characteristics were used for this analysis. Three factors were identified namely, gender, farming system, and district from which the animals were sampled. Male animals were less likely to have infections compared to females (OR = 0.531, 95% CI, 0.290–0.973, p = 0.041), while animals raised under semi-intensive system were more at risk of infection when compared to animals raised under intensive system (OR = 2.266 95% CI, 1.315–3.904, p = 0.003). Furthermore, ruminant livestock animals from Northeast Penang Island District were the most at risk of Blastocystis sp. infection.
Subtype and allele distribution of Blastocystis sp.
All 87 samples that were examined positive using in vitro cultivation were subjected to DNA barcoding. From these, 39 isolates were successfully corresponded to a single subtype Blastocystis infection indicated by single peaks visible from the chromatogram of each sequence. Other isolates revealed poor-quality sequences, which were associated with faint bands on agarose gel and were likely caused by inadequate amounts of DNA. Based on the NCBI nucleotide BLAST results, Blastocystis sp. subtypes identified from the livestock animals screened were ST4, ST5, ST10, ST13, ST14, ST15, and ST25. As shown in Table 2, three Blastocystis sp. subtypes, namely ST5, ST14, and ST25, were identified in cattle, with ST14 being the most frequent (66.7%) and ST25 the least detected (11.1%). In goats and sheep, four subtypes were observed; ST5 was the most common subtype in both animal groups (45% in goats, 50% in sheep), while ST10 and ST15 were the least observed in goats (5%) and sheep (10%), respectively. Blastocystis sp. ST5 and ST14 were found to be widespread across all three ruminant livestock animals groups examined. Blastocystis 18S allele calling revealed exact matches for thirteen isolates belonging to Blastocystis sp. ST5—alleles 16 and 115. Allele 16 was identified across all three ruminant livestock animal groups, while allele 115 was identified in goats alone (Fig. 2).
Phylogenetic analyses of Blastocystis sp. isolates
Generally, the evolutionary trees inferred by the NJ (Fig. 3) and ML (Fig. 4) methods, from the barcoding region of the SSU rRNA gene sequences analyzed, showed similar topologies. BLAST calls in the NCBI database were confirmed by phylogenetic analysis, for the isolates obtained, by both NJ and ML methods. Blastocystis sp. sequences from ruminant livestock animals of Penang, Malaysia, nested within clades of reference sequences belonging to their corresponding subtypes with moderate to high bootstrap support for both trees. From both NJ and ML trees, the reference isolates for ST14 formed two separate clades (“the mouflon” and “the others”), so did the ST14 isolates obtained from ruminant livestock animals.
Phylogenetic relationships of nucleotide sequences of Blastocystis SSU rRNA genes using the NJ method. The number on the branches are percent bootstrapping values from 1000 replicates, with values of more than 50% shown in the tree. Reference sequences are identified by accession number, subtype, and host. Newly generated sequences are identified by accession number, host, and location. Circle icons represent sequences from cattle, square icons represent sequences from goats, and triangle icons represent sequences from sheep
Phylogenetic relationships of nucleotide sequences of Blastocystis SSU rRNA genes using the ML method. The number on the branches are percent bootstrapping values from 1000 replicates, with values of more than 50% shown in the tree. Reference sequences are identified by accession number, subtype, and host. Newly generated sequences are identified by accession number, host, and location. Circle icons represent sequences from cattle, square icons represent sequences from goats, and triangle icons represent sequences from sheep
Discussion
The study of Blastocystis and its genetic variants is important across different hosts and geographical regions for a robust epidemiological picture of this interesting organism. This is the first report on the presence of Blastocystis in ruminant livestock animals in Penang, Malaysia. Bovid infection with Blastocystis is widespread despite the varying rates revealed from different geographical locations, and prevalence of Blastocystis in cattle documented all around the world has ranged widely from 1.8% in Spain (Quílez et al. 1995) to 100% in Indonesia and Colombia (Suwanti et al. 2020; Higuera et al. 2021). When compared with studies from Malaysia, the 15.7% Blastocystis infection rate observed in cattle from Penang in this study is lower than 34.5% prevalence reported from Perak (Hemalatha et al. 2014). However, several prevalence rates (33.3%, 43.8%, and 29.3%) have been reported from different regions of Pahang (Mohammad et al. 2018; Kamaruddin et al. 2020; Abd Razak and Mohammad 2022). The dwelling conditions of animals screened by Hemalatha et al. (2014) were not stated; however, animals screened by Kamaruddin et al. (2020) and Mohammad et al. (2018) were from rural environments, and were either allowed to graze freely and drank mainly from rivers or were raised in cages with poor sanitation practices and facilities. Animals screened by Abd Razak and Mohammad (2022) were reared under a semi-intensive system that allowed animals to graze for 4 to 6 h a day and housed them in the shelter at night. A majority (71.7%; 91/127) of the cattle involved in the present study were reared under the intensive husbandry system, while none was raised by the extensive husbandry system. These animals may be less exposed to contamination; consequently, the low prevalence of Blastocystis infection observed in cattle in this study.
Universally, descriptions of Blastocystis in small ruminants are limited (Hublin et al. 2020). In sheep from Penang, the prevalence of Blastocystis infection observed (14%) is higher than the report on sheep from China (5.5%) (Wang et al. 2018b) but lower than findings from Perak, Malaysia (57.9%) (Hemalatha et al. 2014); Pahang, Malaysia (43.07%) (Abd Razak and Mohammad 2022); and Brazil (33.3%) (Moura et al. 2018). The low prevalence of Blastocystis infection detected in sheep in this study could be attributable to the intensive husbandry system under which all of them were reared and their healthy status as none of them showed any diarrhea symptoms. A moderate infection rate of Blastocystis (35.6%) was detected in goats in Penang in this study. This corresponds to observations from goats in Selangor and Perak in Malaysia (Tan et al. 2013), but is higher than the infection rates described in goats from Pahang, Malaysia (29.16%) (Abd Razak and Mohammad 2022), and Nepal (0.75%) (Ghimire and Bhattarai 2019). The observed prevalence of Blastocystis in goats in this study is lower than findings from Perak (65%) (Hemalatha et al. 2014), France (50%) (Cian et al. 2017), and Thailand (94.7%) (Udonsom et al. 2018). The observation of a significantly higher prevalence of infection in goats compared to other livestock animals is in concordance with studies of Blastocystis sp. infection in livestock animals carried out by Hemalatha et al. (2014) in Perak, Malaysia. Surprisingly, a similar research design by Wang et al. (2018b) recorded an absence of Blastocystis in goats in Heilongjiang Province of China; this could be because of the small sample size (13) involved. Goats are believed to be less selective and scrupulous than cattle and sheep in feeding (Dias-Silva and Abdalla Filho 2021) which might increase the chances of their exposure to contamination and gastrointestinal infections, and thus, result in the higher Blastocystis prevalence recorded for goats than sheep and cattle recorded.
In general, Blastocystis could be referred to as moderately prevalent in livestock in Penang since it was widespread among the studied host types with an overall prevalence of 23.1%. Findings from this study support findings from other studies on livestock animals that these livestock animals are very commonly infected by Blastocystis, but at different levels. Factors such as study design, geographic location, group of animals included in the study (age, gender, or dairy/beef cattle), rearing condition, or the detection method used serve as reasons for variations in values from several reports (Hublin et al. 2020; Higuera et al. 2021). The immune system of young ruminant livestock animals is considered weaker compared to adults, consequently making them more susceptible to infectious agents (Gunathilaka et al. 2018; Ngongolo et al. 2019). On the contrary, lower prevalence of infection was recorded in the age range of < 1 year than in older age groups, although differences in Blastocystis infection across age groups were of statistical significance in sheep only (p = 0.040). Findings from previous studies also revealed a lower infection rate in younger animals (Fayer et al. 2012; Tan et al. 2013; Zhu et al. 2017; Lee et al. 2018; Maloney et al. 2019a). The reason for this observation may be that the risk of exposure to contamination is reduced in young animals due to the special system of care they receive (Maloney et al. 2019a). Significant difference in prevalence of Blastocystis infection was observed between the two farming systems employed in the livestock farms. Infection was found significantly higher in cattle reared in the semi-intensive husbandry system than in the intensive system (p = 0.019). This is expected considering the semi-intensive husbandry system exposes animals to risk of gastrointestinal infections by exposure to contaminated food and water. The presence of Blastocystis cysts has been detected in various water sources (Eroglu and Koltas 2010; Lee et al. 2012; Karaman et al. 2017; Koloren et al. 2018) and vegetables (Soares and Cantos 2006; Al-Megrin 2010) suggesting that these are sources of infection to humans and animals. Results obtained also revealed that female livestock animals, animals reared under semi-intensive livestock management system, and animals reared within the Northeast Penang Island district were more at risk of Blastocystis sp. infection in Penang. Female livestock animals have been reported to be significantly associated with increased odds of gastrointestinal parasite infection among small ruminants, since stress of pregnancy and lactation are capable of altering the immunity of female animals and predisposing them to infectious diseases (Paul et al. 2020). About 85% of farms involved in this study from Northeast Penang Island district employed the semi-intensive farm management system, while other districts of Penang had less than 50% of farms practicing the semi-intensive farm management system; hence, the likely reason for the Northeast Penang Island district being more at risk of Blastocystis sp. infection in Penang.
Unfortunately, 48 out of 87 positive isolates examined using in vitro cultivation were not corresponded to any Blastocystis subtype. Perhaps the amounts of DNA extracted were inadequate in these samples, causing the yield of PCR amplicon to be insufficient to obtain good quality sequences and seen as faint bands on the agarose gel.
Out of at least 28 genetic lineages of Blastocystis sp. that have been acknowledged, seven were identified in the livestock animals examined, namely Blastocystis sp. ST4, ST5, ST10, ST13, ST14, ST15, and ST25. The observation of ST5 and ST14 in cattle is similar to findings of Kamaruddin et al. (2020) from Pahang, Malaysia; Maloney et al. (2019a) from the USA; Masuda et al. (2018) from Japan; Zhu et al. (2017) from China; and Badparva et al. (2015) from Iran. Blastocystis sp. ST25 was also identified in cattle from Penang; this subtype was previously isolated from cattle and sheep in Belgium and the United States of America (USA) (Maloney et al. 2019a; b). Although Blastocystis ST10 was not identified in cattle in this study, ST10 and ST14 are considered to be the most widespread subtypes in cattle generally (Hublin et al. 2020; Shams et al. 2021). Nevertheless, studies on Blastocystis in small ruminant animals are limited. Blastocystis ST1, ST3-ST7, ST10, ST12, and ST14 have been described in goats, while ST1, ST3, ST5, ST10, ST14, and ST15 have been reported in sheep from different parts of the world (Hublin et al. 2020; Chang et al. 2021). The isolation of ST4 from sheep and ST13 from goats has not been previously reported in the literature, to the best of our knowledge. Rodents are perceived to be the original hosts to ST4, a subtype also commonly observed in humans (Stensvold et al. 2012), however, this subtype has equally been detected in artiodactyls such as cattle (Zhu et al. 2017; Maloney et al. 2019a; b; Kamaruddin et al. 2020), goats (Lee et al. 2012; Song et al. 2017), and buffalo (Lee et al. 2012), and in rabbits, pigs, cats, dogs, and non-human primates (NHPs) (Hublin et al. 2020). And previous descriptions of ST13 have been in quokkas from Australia (Parkar et al. 2010), birds from Iran (Asghari et al. 2019), NHP from China and Bangladesh (Zhao et al. 2017; Li et al. 2019), reindeers from China (Wang et al. 2018a), and flying squirrels from China (Xiao et al. 2019). Thus, the observation of ST4 in sheep and ST13 in goats in this study provides an update to the host range of Blastocystis sp. ST4 and ST13.
Just as enzootic subtypes (ST10, ST14, ST25) adapted to ruminant livestock animals were detected in the present study, potentially zoonotic subtypes (ST4, ST5) were equally identified. Blastocystis sp. ST4 and ST5 have been observed in humans from around the world (Deng et al. 2021; Popruk et al. 2021), and from different parts of Malaysia (Nithyamathi et al. 2016; Nemati et al. 2021). The observation of potentially zoonotic Blastocystis sp. subtypes (ST4, ST5) suggests that livestock animals are possible sources of human infection. Interestingly, ST10 and ST14 were recently observed in children in Senegal (Khaled et al. 2020) and a Syrian population in Lebanon (Khaled et al. 2021), thus adding to the list of subtypes potentially transmissible between humans and animals. Khaled et al. (2021) suggest that the transmission of ST10 may be through the consumption of water contaminated by ST10 parasite cysts, noting that ST10 had been isolated from river water samples in Malaysia by Noradilah et al. (2016).
Analysis of subtype alleles revealed intra-subtype variations within Blastocystis ST5 identified in ruminant livestock animals. Blastocystis ST5 allele 115 identified in goats was also reported in sheep from China (Li et al. 2018) and farm pigs from Italy (Gabrielli et al. 2021). Allele 16 was the mostly identified allele of Blastocystis ST5, and it was found in each ruminant animal group in this study. This allele has hardly ever been reported in literature. However, when Blastocystis sp. homologous sequences (accession numbers MG831436, MF541116, AB070999, MK801414, etc.) resulting from NCBI BLAST calls of sequences obtained were subjected to the pubmlst.org/Blastocystis/sequence query, they were found to belong to allele 16. These were sequences of Blastocystis sp. isolates from cattle, goats, and most commonly, pigs from different regions of the world including Malaysia. It could, thus, be implied that intra-subtype variation of the barcode region of Blastocystis sp. is under-reported.
The phylogenetic trees inferred by the NJ and ML methods showed clustering and branching orders identical to those described previously (Maloney and Santin 2021; Song et al. 2021; Liu et al. 2022). The phylogenetic trees inferred by the NJ and ML methods supported the assignment of isolates to Blastocystis sp. ST4, ST5, ST10, ST13, ST14, ST15, and ST25. The close sequence similarity between ST14 and ST25 (Maloney and Santin 2021) was observed by phylogenetic analyses in this study. Blastocystis sp. STs 5, 13, 14, and 25 have been mostly isolated from the artiodactyls and other hoofed animals; these subtypes also share common ancestry on the phylogenetic tree. This observation could suggest some form of association between the phylogeny of these Blastocystis subtypes and their preferred host.
In conclusion, Blastocystis infection has been revealed as moderate and widespread in cattle, goats, and sheep in Penang, Malaysia. Gender, husbandry management system, and district were identified as potential predictors of Blastocystis infection in the ruminant livestock animals; and for the first time, Blastocystis ST4 and ST13 were reported in sheep and goats respectively. Furthermore, the isolation of Blastocystis ST4 and ST5 which have been commonly observed in humans, and ST10 which has been sporadically observed in humans, suggests the likely role of ruminant livestock animals in transmission of Blastocystis to humans. Therefore, adequate hygiene practices on animal farms such as the use of appropriate personal protective covering by animal handlers and proper disposal of wastes are recommended as prevention and control measures against the spread of Blastocystis to humans and within animals. Future studies designed to involve animals and their in-contact humans will aid in revealing the transmission dynamics of Blastocystis.
Data availability
All data generated or analyzed during this study are included in this article. The newly generated sequences were deposited in the GenBank database under the accession numbers ON738374-ON738382, ON738385-ON738404, and ON738409-ON738418.
References
AbdRazak NA, Mohammad M (2022) Prevalence of Blastocystis sp. in cattle, goat and sheeep reared by different farm management systems in Pahang. Malaysia. Malaysian Appl Biol 51:47–55. https://doi.org/10.55230/mabjournal.v51i3.2165
AbdRazak NA, Yusof AM, Mohammad M (2019) Identification of Blastocystis sp. infection from cattle, goat and sheep isolated from farms in Pahang Malaysia. Int J Allied Heal Sci 3:810
Ahmed SA, Karanis P (2018) Blastocystis spp., ubiquitous parasite of human, animals and environment. Ref Modul Earth Syst Environ Sci 1–6. https://doi.org/10.1016/b978-0-12-409548-9.10947-9
Al-Megrin WAI (2010) Prevalence of intestinal parasites in leafy vegetables in Riyadh, Saudi Arabia. Int J Zool Res 6:190–195. https://doi.org/10.3923/ijzr.2010.190.195
Asghari A, Sadraei J, Pirestani M, Mohammadpour I (2019) First molecular identification and subtype distribution of Blastocystis sp. isolated from hooded crows (Corvus cornix) and pigeons (Columba livia) in Tehran Province. Iran Comp Immunol Microbiol Infect Dis 62:25–30. https://doi.org/10.1016/j.cimid.2018.11.013
Badparva E, Sadraee J, Kheirandish F (2015) Genetic diversity of Blastocystis isolated from cattle in Khorramabad. Iran Jundishapur J Microbiol 8:4–7. https://doi.org/10.5812/jjm.14810
Chang Y, Yan Y, Han H, Wu Y, Li J, Ning C (2021) Prevalence of Blastocystis infection in free-range Tibetan sheep and Tibetan goats in the Qinghai-Tibetan Plateau in China. One Heal 13:100347
Cian A, El Safadi D, Osman M, Moriniere R, Gantois N, Benamrouz-Vanneste S, Delgado-Viscogliosi P, Guyot K, Li LL, Monchy S, Noël C, Poirier P, Nourrisson C, Wawrzyniak I, Delbac F, Bosc S, Chabé M, Petit T, Certad G, Viscogliosi E (2017) Molecular epidemiology of Blastocystis sp. in various animal groups from two French zoos and evaluation of potential zoonotic risk. PLoS ONE 12:1–29. https://doi.org/10.1371/journal.pone.0169659
Clark CG (1997) Extensive genetic diversity in Blastocystis hominis. Mol Biochem Parasitol 87:79–83. https://doi.org/10.1016/S0166-6851(97)00046-7
Clark CG, van der Giezen M, Alfellani MA, Stensvold CR (2013) Recent developments in Blastocystis research. Adv Parasitol 82:1–32. https://doi.org/10.1016/B978-0-12-407706-5.00001-0
DE Adao V, Rivera WL (2018) Recent advances in Blastocystis sp. research. Philipp Sci Lett 11:39–60. https://doi.org/10.1136/bmj.320.7246.1385
Deng Y, Zhang S, Ning C, Zhou Y, Teng X, Wu X, Chu Y, Yu Y, Chen J, Tian L, Tian L, Wang W (2020) Molecular epidemiology and risk factors of Blastocystis sp. Infections among general populations in Yunnan Province, Southwestern China. Risk Manag Healthc Policy 13:1791–1801. https://doi.org/10.2147/RMHP.S269664
Deng L, Wojciech L, Gascoigne NRJ, Peng G, Tan KSW (2021) New insights into the interactions between Blastocystis, the gut microbiota, and host immunity. PLoS Pathog 17:1–15. https://doi.org/10.1371/JOURNAL.PPAT.1009253
Dias-Silva TP, Abdalla Filho AL (2021) Sheep and goat feeding behavior profile in grazing systems. Acta Sci. Anim Sci 43:e51265. https://doi.org/10.4025/actascianimsci.v43i1.51265
Eroglu F, Koltas IS (2010) Evaluation of the transmission mode of B. hominis by using PCR method. Parasitol Res 107:841–845. https://doi.org/10.1007/s00436-010-1937-4
Fayer R, Santin M, MacArisin D (2012) Detection of concurrent infection of dairy cattle with Blastocystis, Cryptosporidium, Giardia, and Enterocytozoon by molecular and microscopic methods. Parasitol Res 111:1349–1355. https://doi.org/10.1007/s00436-012-2971-1
Gabrielli S, Palomba M, Furzi F, Brianti E, Gaglio G, Napoli E, Rinaldi L, Alburqueque RA, Mattiucci S (2021) Molecular subtyping of Blastocystis sp. Isolated from farmed animals in southern Italy. Microorganisms 9:1–10. https://doi.org/10.3390/microorganisms9081656
Ghimire TR, Bhattarai N (2019) A survey of gastrointestinal parasites of goats in a goat market in Kathmandu. Nepal J Parasit Dis 43:686–695. https://doi.org/10.1007/s12639-019-01148-w
Gunathilaka N, Niroshana D, Amarasinghe D, Udayanga L (2018) Prevalence of gastrointestinal parasitic infections and assessment of deworming program among cattle and buffaloes in Gampaha District. Sri Lanka Biomed Res Int 2018:30483. https://doi.org/10.1155/2018/3048373
Hemalatha C, Chandrawathani P, Suresh Kumar G, Premaalatha B, Geethamalar S, Lily Rozita M, Farah Haziqah MT, Sabapathy D, Ramlan M (2014) The diagnosis of Blastocystis Sp. from animals — an emerging zoonosis. Malaysian J Vet Res 5:15–22
Hidalgo L, Salvador F, Sulleiro E, López I, Balladares M, García E, Paz C, Sánchez-Montalvá A, Bosch-Nicolau P, Sao-Avilés A, Molina I (2019) Evaluation of risk factors associated to detection of Blastocystis sp. in fecal samples in population from Barcelona, Spain: a case-control study. Eur J Clin Microbiol Infect Dis 38:1241–1247. https://doi.org/10.1007/s10096-019-03532-z
Higuera A, Herrera G, Jimenez P, García-Corredor D, Pulido-Medellín M, Bulla-Castañeda DM, Pinilla JC, Moreno-Pérez DA, Maloney JG, Santín M, Ramírez JD (2021) Identification of multiple Blastocystis subtypes in domestic animals from colombia using amplicon-based next generation sequencing. Front Vet Sci 8. https://doi.org/10.3389/fvets.2021.732129
Hublin JSY, Maloney JG, Santin M (2020) Blastocystis in domesticated and wild mammals and birds. Res Vet Sci 135:260–282. https://doi.org/10.1016/j.rvsc.2020.09.031
International Trade Administration (2021) Malaysia - agricultural sector. In: Ctry. Commer. Guid. https://www.trade.gov/country-commercial-guides/malaysia-agricultural-sector. Accessed 11 Feb 2022
Jolley KA, Bray JE, Maiden MCJ (2018) Open-access bacterial population genomics: BIGSdb software, the PubMLST. org website and their applications [version 1; referees: 2 approved]. Wellcome Open Res 3:1–20. https://doi.org/10.12688/wellcomeopenres.14826.1
Kamaruddin SK, Mat Yusof A, Mohammad M (2020) Prevalence and subtype distribution of Blastocystis sp. in cattle from Pahang. Malaysia Trop Biomed 37:127–141
Karaman Ü, Kolören Z, Seferoğlu O, Ayaz E, Demirel E (2017) Presence of parasites in environmental waters in Samsun and its districts. Turkiye Parazitolojii Derg 41:19–21. https://doi.org/10.5152/tpd.2017.3574
Khaled S, Gantois N, Ly AT, Senghor S, Even G, Dautel E, Dejager R, Sawant M, Baydoun M, Benamrouz-Vanneste S, Chabé M, Ndiaye S, Schacht AM, Certad G, Riveau G, Viscogliosi E (2020) Prevalence and subtype distribution of blastocystis sp. In senegalese school children. Microorganisms 8:1–17. https://doi.org/10.3390/microorganisms8091408
Khaled S, Gantois N, Ayoubi A, Even G, Sawant M, El Houmayraa J, Nabot M, Benamrouz-Vanneste S, Chabé M, Certad G, Hamze M, Viscogliosi E (2021) Blastocystis sp. Prevalence and subtypes distribution amongst Syrian refugee communities living in North Lebanon. Microorganisms 9:1–13. https://doi.org/10.3390/microorganisms9010184
Klous G, Huss A, Heederik DJJ, Coutinho RA (2016) Human-livestock contacts and their relationship to transmission of zoonotic pathogens, a systematic review of literature. One Heal 2:65–76. https://doi.org/10.1016/j.onehlt.2016.03.001
Koloren Z, Gulabi BB, Karanis P (2018) Molecular identification of Blastocystis sp. subtypes in water samples collected from Black sea. Turkey Acta Trop 180:58–68. https://doi.org/10.1016/j.actatropica.2017.12.029
Lee H, Lee SH, Seo MG, Kim HY, Kim JW, Lee YR, Kim JH, Kwon OD, Kwak D (2018) Occurrence and genetic diversity of Blastocystis in Korean cattle. Vet Parasitol 258:70–73. https://doi.org/10.1016/j.vetpar.2018.06.010
Lee L, Chye T, Karmacharya M, Govind S (2012) Blastocystis sp.: waterborne zoonotic organism, a possibility? Parasit Vectors 5:130
Lepczyńska M, Białkowska J, Dzika E, Piskorz-Ogórek K, Korycińska J (2017) Blastocystis: how do specific diets and human gut microbiota affect its development and pathogenicity? Eur J Clin Microbiol Infect Dis 36:1531–1540. https://doi.org/10.1007/s10096-017-2965-0
Li J, Karim MR, Li D, RahamanSumon SMM, Siddiki SHMF, Rume FI, Sun R, Jia Y, Zhang L (2019) Molecular characterization of Blastocystis sp. in captive wildlife in Bangladesh National Zoo: non-human primates with high prevalence and zoonotic significance. Int J Parasitol Parasites Wildl 10:314–320. https://doi.org/10.1016/j.ijppaw.2019.11.003
Li WC, Wang K, Gu Y (2018) Occurrence of Blastocystis sp. and Pentatrichomonas hominis in sheep and goats in China. Parasit Vectors 11:1–7. https://doi.org/10.1186/s13071-018-2671-5
Liu X, Ni F, Wang R, Li J, Ge Y, Yang X, Qi M (2022) Occurrence and subtyping of Blastocystis in coypus (Myocastor coypus) in China. Parasit Vectors 15:1–7. https://doi.org/10.1186/s13071-021-05126-1
Maloney JG, Lombard JE, Urie NJ, Shivley CB, Santin M (2019a) Zoonotic and genetically diverse subtypes of Blastocystis in US pre-weaned dairy heifer calves. Parasitol Res 118:575–582. https://doi.org/10.1007/s00436-018-6149-3
Maloney JG, Molokin A, Santin M (2019b) Next generation amplicon sequencing improves detection of Blastocystis mixed subtype infections. Infect Genet Evol 73:119–125. https://doi.org/10.1016/j.meegid.2019.04.013
Maloney JG, Santin M (2021) Mind the gap: New full-length sequences of Blastocystis subtypes generated via oxford nanopore minion sequencing allow for comparisons between full-length and partial sequences of the small subunit of the ribosomal rna gene. Microorganisms 9. https://doi.org/10.3390/microorganisms9050997
Masuda A, Sumiyoshi T, Ohtaki T, Matsumoto J (2018) Prevalence and molecular subtyping of Blastocystis from dairy cattle in Kanagawa, Japan. Parasitol Int 67:702–705. https://doi.org/10.1016/j.parint.2018.07.005
Mohammad NA, Al-Mekhlafi HM, Anuar TS (2018) Subtype distribution of Blastocystis isolated from humans and associated animals in an indigenous community with poor hygiene in Peninsular Malaysia. Trop Biomed 35:849–860
Moura RGF, de Oliveira-Silva MB, Pedrosa AL, Nascentes GAN, Cabrine-Santos M (2018) Occurrence of Blastocystis spp. in domestic animals in Triângulo Mineiro area of Brazil. Rev Soc Bras Med Trop 51:240–243. https://doi.org/10.1590/0037-8682-0484-2016
Nemati S, Zali MR, Johnson P, Mirjalali H, Karanis P (2021) Molecular prevalence and subtype distribution of Blastocystis sp. in Asia and in Australia. J Water Health 19:687–704. https://doi.org/10.2166/wh.2021.011
Ngongolo K, Estes AB, Hudson PJ, Gwakisa PS (2019) Assessing risk factors for trypanosome infections in cattle in wildlife interface areas in Northern Tanzania. J Infect Dis Epidemiol 5:078
Nithyamathi K, Chandramathi S, Kumar S (2016) Predominance of Blastocystis sp. infection among school children in Peninsular Malaysia. PLoS ONE 11:1–14. https://doi.org/10.1371/journal.pone.0136709
Noradilah SA, Lee IL, Anuar TS, Salleh FM, Abdul Manap SNA, Mohd Mohtar NSH, Azrul SM, Abdullah WO, Moktar N (2016) Occurrence of Blastocystis sp. in water catchments at Malay villages and Aboriginal settlement during wet and dry seasons in Peninsular Malaysia. PeerJ 4:e2541. https://doi.org/10.7717/peerj.2541
Noradilah SA, Anuar TS, Moktar N, Lee IL, Salleh FM, Azreen SNAM, Husnie NSMM, Azrul SM, Abdullah WO, Nordin A, Abdullah SR (2017) Molecular epidemiology of Blastocystis sp. in animals reared by the aborigines during wet and dry seasons in rural communities, pahang, Malaysia. Southeast Asian J Trop Med Public Health 48:1151–1160
Osorio-Pulgarin MI, Higuera A, Beltran-álzate JC, Sánchez-Jiménez M, Ramírez JD (2021) Epidemiological and molecular characterization of Blastocystis infection in children attending daycare centers in medellín, Colombia. Biology 10:1–14. https://doi.org/10.3390/biology10070669
Osman M, El Safadi D, Cian A, Benamrouz S, Nourrisson C, Poirier P, Pereira B, Razakandrainibe R, Pinon A, Lambert C, Wawrzyniak I, Dabboussi F, Delbac F, Favennec L, Hamze M, Viscogliosi E, Certad G (2016) Prevalence and risk factors for intestinal protozoan infections with Cryptosporidium, Giardia, Blastocystis and Dientamoeba among Schoolchildren in Tripoli, Lebanon. PLoS Negl Trop Dis 10(3):e0004496. https://doi.org/10.1371/journal.pntd.0004496
Parkar U, Traub RJ, Vitali S, Elliot A, Levecke B, Robertson I, Geurden T, Steele J, Drake B, Thompson RCA (2010) Molecular characterization of Blastocystis isolates from zoo animals and their animal-keepers. Vet Parasitol 169:8–17. https://doi.org/10.1016/j.vetpar.2009.12.032
Paul BT, Jesse FFA, Chung ELT, Che’amat A, Lila MAM (2020) Risk factors and severity of gastrointestinal parasites in selected small ruminants from Malaysia. Vet Sci 7:1–14. https://doi.org/10.3390/vetsci7040208
Popruk S, DE Adao V, Rivera WL (2021) Epidemiology and subtype distribution of Blastocystis in humans: a review. Infect Genet Evol 95:105085. https://doi.org/10.1016/j.meegid.2021.105085
Quílez J, Sanchez-Acedo C, Clavel A, Causape AC (1995) Occurrence of Blastocystis sp. in cattle in Aragon, northeastern Spain. Parasitol Res 81:703–705
Rauff-Adedotun AA, Meor Termizi FH, Shaari N, Lee IL (2021) The coexistence of Blastocystis spp. in humans, animals and environmental sources from 2010–2021 in Asia. Biology 10:990. https://doi.org/10.3390/biology10100990
Scicluna SM, Tawari B, Clark CG (2006) DNA barcoding of Blastocystis. Protist 157:77–85. https://doi.org/10.1016/j.protis.2005.12.001
Shams M, Shamsi L, Sadrebazzaz A, Asghari A, Badali R, Omidian M, Hassanipour S (2021) A systematic review and meta-analysis on the global prevalence and subtypes distribution of Blastocystis sp. infection in cattle: a zoonotic concern. Comp Immunol Microbiol Infect Dis 76:101650. https://doi.org/10.1016/j.cimid.2021.101650
Silberman JD, Sogin ML, Leipe DD, Clark CG (1996) Human parasite finds taxonomic home. Nature 380:398. https://doi.org/10.1038/380398a0
Soares B, Cantos GA (2006) Detecção de estruturas parasitárias em hortaliças comercializadas na cidade de Florianópolis, SC, Brasil. Rev Bras Ciencias Farm J Pharm Sci 42:455–460. https://doi.org/10.1590/S1516-93322006000300015
Song J, Yang X, Ma X, Wu X, Wang Y, Li Z, Liu G, Zhao G (2021) Molecular characterization of Blastocystis sp. in Chinese bamboo rats (Rhizomys sinensis). Parasite 28:81. https://doi.org/10.1051/parasite/2021081
Song JK, Yin YL, Yuan YJ, Tang H, Ren GJ, Zhang HJ, Li ZX, Zhang YM, Zhao GH (2017) First genotyping of Blastocystis sp. in dairy, meat, and cashmere goats in northwestern China. Acta Trop 176:277–282. https://doi.org/10.1016/j.actatropica.2017.08.028
Stensvold CR, Alfellani M, Clark CG (2012) Levels of genetic diversity vary dramatically between Blastocystis subtypes. Infect Genet Evol 12:263–273. https://doi.org/10.1016/j.meegid.2011.11.002
Suresh K, Smith H (2004) Comparison of methods for detecting Blastocystis hominis. Eur J Clin Microbiol Infect Dis 23:509–511. https://doi.org/10.1007/s10096-004-1123-7
Suwanti LT, Susana Y, Hastutiek P, Suprihati E, Lastuti NDR (2020) Blastocystis spp. Subtype 10 infected beef cattle in Kamal and Socah, Bangkalan, Madura, Indonesia. Vet World 13:231–237. https://doi.org/10.14202/vetworld.2020.231-237
Tamura K, Stecher G, Kumar S (2021) MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 38:3022–3027. https://doi.org/10.1093/molbev/msab120
Tan K (2008) New insights on classification, identification, and clinical relevance of Blastocystis spp. Clin Microbiol Rev 21:639–665. https://doi.org/10.1128/CMR.00022-08
Tan TC, Tan PC, Sharma R, Sugnaseelan S, Suresh KG (2013) Genetic diversity of caprine Blastocystis from Peninsular Malaysia. Parasitol Res 112:85–89. https://doi.org/10.1007/s00436-012-3107-3
Udonsom R, Prasertbun R, Mahittikorn A, Mori H, Changbunjong T, Komalamisra C, Pintong AR, Sukthana Y, Popruk S (2018) Blastocystis infection and subtype distribution in humans, cattle, goats, and pigs in central and western Thailand. Infect Genet Evol 65:107–111. https://doi.org/10.1016/j.meegid.2018.07.007
Wang J, Gong B, Liu X, Zhao W, Bu T, Zhang W, Liu A, Yang F (2018a) Distribution and genetic diversity of Blastocystis subtypes in various mammal and bird species in northeastern China. Parasit Vectors 11:1–7. https://doi.org/10.1186/s13071-018-3106-z
Wang J, Gong B, Yang F, Zhang W, Zheng Y, Liu A (2018b) Subtype distribution and genetic characterizations of Blastocystis in pigs, cattle, sheep and goats in northeastern China’s Heilongjiang Province. Infect Genet Evol 57:171–176. https://doi.org/10.1016/j.meegid.2017.11.026
Xiao X, Zhou SH, Jiang N, Tian DZ, Zhou ZM, Zhang M, Ke H, Jiang XC, Lv WL, Gao QH (2019) First record of Leptospira and Blastocystis infections in captive flying squirrels (Trogopterus xanthipes) from Enshi County, China. Acta Trop 197. https://doi.org/10.1016/j.actatropica.2019.105065
Zhao GH, Hu XF, Liu TL, Hu RS, Yu ZQ, Yang WB, Wu YL, Yu SK, Song JK (2017) Molecular characterization of Blastocystis sp. in captive wild animals in Qinling Mountains. Parasitol Res 116:2327–2333. https://doi.org/10.1007/s00436-017-5506-y
Zhu W, Tao W, Gong B, Yang H, Li Y, Song M, Lu Y, Li W (2017) First report of Blastocystis infections in cattle in China. Vet Parasitol 246:38–42. https://doi.org/10.1016/j.vetpar.2017.09.001
Acknowledgements
The authors would also like to thank the members of staff of the Department of Veterinary Services (DVS), Penang, for their tremendous assistance rendered during sampling.
Funding
This research was supported by Universiti Sains Malaysia (Short Term Grant: 304/PBIOLOGI/6315156 and Incentive Grant 2020: 1001.PBIOLOGI.AUPS001), and Short Term Grants of UniSHAMS (100–161(RMU)01/STG/2020 Bil. (64)).
Author information
Authors and Affiliations
Contributions
Farah Haziqah Meor Termizi, Ii Li. Lee, and Nurshafarina Shaari conceived and designed the study. Adedolapo Aminat Rauff-Adedotun and Nabilah Abd Talib conducted sampling. Adedolapo Aminat Rauff-Adedotun carried out laboratory works. Adedolapo Aminat Rauff-Adedotun analyzed and interpreted the data, and wrote the article Farah Haziqah Meor Termizi, Ii Li Lee, Nurshafarina Shaari, and Zary Shariman Yahaya revised the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. Animal ethical approval was obtained from USM Institutional Animal Care and Use Committee (USM IACUC), Universiti Sains Malaysia. Meanwhile, permission for sampling activities was obtained from the Department of Veterinary Services (DVS) Penang and the Department of Veterinary Services, Ministry of Agriculture and Agro-based Industry Malaysia.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Kevin Tan
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rauff-Adedotun, A.A., Lee, I.L., Abd Talib, N. et al. Prevalence, potential risk factors and genetic diversity of Blastocystis in ruminant livestock animals from Penang, Malaysia. Parasitol Res 122, 2193–2205 (2023). https://doi.org/10.1007/s00436-023-07920-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-023-07920-5