Abstract
Background
Non-small cell lung cancer is the most common subtype of lung cancer in the world. However, the survival rate of non-small cell lung cancer patients remains low currently. Immune checkpoint and long non-coding RNAs are emerging as critical roles in prognostic significance and the immunotherapeutic response of non-small cell lung cancer. It is critical to discern LncRNAs related with immune checkpoints in patients with Non-small cell lung cancer.
Methods
In this study, immune checkpoint-linked LncRNAs were determined and achieved by the co-expression analysis. Immune checkpoint-linked LncRNAs with noteworthy prognostic value (P < 0.05) gained were next utilized to separate into two cluster by non-negative matrix factorization (NMF). Univariate and a least absolute shrinkage and selection operator were applied to construct an immune checkpoint-linked LncRNAs model. Kaplan–Meier analysis, Gene Set Enrichment Analysis, and the nomogram were utilized to investigate the LncRNAs model. Lastly, the capability immunotherapy and chemotherapy prediction value of this risk model were also estimated.
Results
The model consisting of ten immune checkpoint-related LncRNAs was acknowledged to be a self-determining predictor of prognosis. Through regrouping the NSCLC patients by this model, difference between them more efficiently on immunotherapeutic response, tumor microenvironment and chemotherapy response could be discovered. This risk model related to the immune checkpoint-based LncRNAs may have an excellent clinical prediction for prognosis and the immunotherapeutic response in patients with NSCLC.
Conclusions
We performed an integrative analysis of LncRNAs linked with immune checkpoints and emphasized the significance of NSCLC subtypes classification, immune checkpoints related LncRNAs in estimating the tumor microenvironment score, immune cell infiltration of the tumor, immunotherapy, and chemotherapy response.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Non-small cell lung cancer (NSCLC) is the most common subtype of lung cancer globally, accounting for 25% of all cancer deaths (Bray et al. 2018). With the development of the diagnosis, surgery, radiotherapy, chemotherapy and immunotherapy, the clinical consequence of NSCLC patients has meaningfully improved (Ramalingam et al. 2011). Nevertheless, the five-year overall survival possibility of patients with NSCLC is hitherto at a low level (Herbst et al. 2018). At present, some evidence shows that the finding and usage of molecular markers can improve prognostic value. The growing researches have indicated that Immune Checkpoints play a vital role in cancer development and aggressiveness (Fritz and Lenardo 2019). The programmed cell death 1 (PDCD1, also known as PD-1)/ CD274 molecule (CD274, also known as PD-L1) and cytotoxic T-lymphocyte associated protein 4 (CTLA-4) pathways are implicated in tumor immune evasion, and therefore, immune checkpoint inhibitors targeting PD-1 and CTLA-4 thereby enhance anti-tumor immunity. PD-1 was first discovered as a member of the immunoglobulin gene superfamily thought to be involved in programmed cell death (Ishida et al. 1992). It is a transmembrane protein expressed by select thymocyte subsets and T and B lymphocytes, especially after antigen receptor stimulation (Shinohara et al. 1994). PD-1 is triggered by the B7 homologues, PD-L1 (B7-H1) and PD-L2 (B7-H2), constitutively reside on nonlymphoid tissues, which can be up-regulated in immune cells by proinflammatory cytokines such as IFN-γ and TNF. Immune regulation targeting PD-1 plays a vital role in anti-tumor, anti-infection, anti-autoimmune diseases and organ transplant survival (Yi et al. 2009; Keir et al. 2008). Its ligand PD-L1 can also be used as a target, and the corresponding antibody can also play the same role. The combination of PD-1 and PD-L1 initiates the programmed death of T cells and makes the tumor cells get immune escape. The effectiveness of PD-1/PD-L1 inhibition in tumor regression encouraged clinical tests in humans with advanced cancer (Topalian et al. 2012). The fully human PD-1 (nivolumab) and PD-L1 (MDX-1105 and atezolizumab) monoclonal blocking antibodies showed exciting regression in patients with diverse incurable cancers, including melanoma (Takeuchi et al. 2017) and renal cell carcinoma (McDermott et al. 2018). Especially, inhibition of immune checkpoint molecules through PD-1 or PD-L1 blockade has demonstrated significant clinical advancements for patients with lung cancer (Yang et al. 2018; Garon et al. 2015; Brahmer et al. 2015). CTLA-4 is a member of the CD28 family of receptors that are induced on the cell surface on conventional T cells by antigen activation and constitutively expressed on regulatory T (T reg) cells, a specialized subset of CD4 + T cells that can arrest T cell responses (Sansom 2000). Anti-CTLA-4 inhibitors has shown certain efficacy in the treatment of melanoma (Afzal et al. 2018; Meerveld-Eggink et al. 2017), non-small cell lung cancer (Perets et al. 2021) and other tumors (Monjazeb et al. 2021; Zhang et al. 2021). Long non-coding RNAs (LncRNAs), have been revealed to be involved in a varied of biological functions (Yao et al. 2019; Moran et al. 2012). Recent studies have proved that LncRNAs are rising as crucial monitoring elements in human immune system and act vital roles in the growth and differentiation of diverse immune cell lineages (Turner et al. 2014; Atianand et al. 2017; Chen et al. 2017). Immune checkpoints such as PD1/PD-L1, CTLA4 inhibitors become aggressively important for cancer therapy, the specific role of Immune checkpoints related LncRNA is still indistinct, thus, comprehending the mechanism of immune checkpoints related LncRNA in the progress of NSCLC may be helpful for uncovering new prognostic and therapeutic targets.
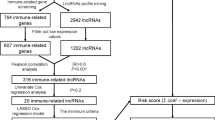
In our research, we distracted the expression matrix of 14,086 LncRNAs and 3 Immune checkpoint recognized genes (PD1, PD-L1, CTLA4) based on NSCLC patients in The Cancer Genome Atlas (TCGA) online database. Next, we identified the Immune checkpoints related LncRNAs through Pearson correlation analysis. A novel prognostic model of immune checkpoints related LncRNAs, was constructed to predict the overall survival (OS) in NSCLC patients. Furthermore, we investigate the relationship associated with chemotherapy and immunotherapy responses. Lastly, we launched a nomogram to foretell the patients’ OS of NSCLC.
Materials and methods
Acquiring data of patients with NSCLC
We attained RNA sequence transcriptome data of FPKM, relevant clinical info of patients with NSCLC from the online database TCGA (https://cancergenome.nih.gov/) (Tomczak et al. 2015). For reducing statistical bias of this analysis, we excluded NSCLC patients with OS days < 30, and this study did not require ethical approval.
Selection of immune checkpoint genes and immune checkpoint-related LncRNAs
We obtain the expression matrix of LncRNAs and 3 immune checkpoint genes (PD1(PDCD1), PD-L1(CD274) and CTLA4) from the online TCGA database. We screened immune checkpoint-related LncRNAs by Pearson correlation analysis. In this study, the cutoff criteria were |Pearson R|> 0.3, and p value < 0.001.
NSCLC subtypes identification
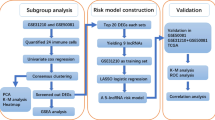
The R package called “survival” was applied to Univariate Cox regression analysis for estimating all candidate genes’ association with overall survival. Immune checkpoint-linked LncRNAs with important prognostic value (P < 0.05) obtained were later utilized to non-negative matrix factorization (NMF) clustering (Gaujoux and Seoighe 2010). The unsupervised NMF clustering methods were conducted by using the “ConsensusClusterPlus” and “limma” R package on the metadata set, then the best cluster number was selected as the coexistence correlation coefficient K value = 2.
Constructing an immune checkpoint-related LncRNAs signature
TCGA’s entire cohort was randomized divided into training cohort and the testing cohort (Table 1). The training cohort was used to fabricate the immune checkpoint-related LncRNAs risk model. The entire cohort and the testing cohort were applied to corroborate this established risk model. In conjunction with NSCLC survival data in TCGA, we marked off the prognosis of immune checkpoint-related LncRNAs from above LncRNAs in the NSCLC data of TCGA dataset (P < 0.05) and univariate Cox regression analyses was utilized in this process. R package “glmnet” was used to perform LASSO Cox regression (The penalty parameter assessed by tenfold cross-validation), then, we discovered that 22 immune checkpoint-related LncRNAs were definitely correlated to the OS of NSCLC patients, and an immune checkpoint-related LncRNAs risk model was established ultimately. The following formula was utilized to compute the risk score: risk score = coefficient1 × expression (LncRNA1) + coefficient2 × expression (LncRNA2) + … + coefficientn × expression (LncRNAn), where the coefficient of LncRNAs associated with overall survival. Based on the median risk score, we recognized the two subcategories (including high- and low-risk groups).
Assessment and validation of immune checkpoint-linked LncRNAs signature
After fabricating the immune checkpoint-linked LncRNAs risk model, we utilized it to the testing and entire cohort to confirm the model’s accuracy. The receiver operating characteristic curve (ROC) was also applied to measure the models’ prediction capability in the cohort of training, testing, and entire, respectively.
Independence of the immune checkpoint-related LncRNA model and establishing and proving a predictive nomogram
To confirm whether the prognostic signature could be self-determining among clinical parameters, including age, gender, stage, risk score, univariate and multivariate Cox regression model method was performed with a stepwise forward procedure. Statistical significance was defined as a P value < 0.05. All data were processed using the R package “survival ROC”. Furthermore, a nomogram was formed based on the self-determining prognostic factors acknowledged by multivariate cox analysis to explore the 1, 3, and 5 year OS of NSCLC patients. The nomogram and calibration plots were generated using the RMS R package (Version: 5.1–3).The prediction ability of the nomogram and other predictors (age, gender, stage, risk score) for 1, 3, and 5 year OS were established.
Assessment of tumor microenvironment and tumor-infiltrating immune cells
Stromal, Immune and Estimate scores of tumor microenvironment (TME) were evaluated by the R package (“ESTIMATE”) (Yoshihara et al. 2013). CIBERSORT is an analytical tool developed by Newman et al. to provide an estimation of 22 immune cell types in a mixed cell population using gene expression data, and it is highly consistent with ground-truth estimations in many cancers (Newman et al. 2015). Hence, we used a combination of the normalized NSCLC gene expression matrix and the LM22 signature matrix to estimate the scores of 22 human immune cell phenotypes between high- and low-risk patients. For all sample, the amount of estimated 22 immune cell scores is equal to 1. P < 0.05 is considered to be statistically significant.
Gene set enrichment analysis
Gene set enrichment analysis (GSEA) (Kuleshov et al. 2019; Reimand et al. 2019) was used to investigate the possible active immune-related Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways in the cluster 1 and cluster 2, high-risk and low-risk groups. P < 0.05 is considered to be statistically significant.
Investigation of the risk model in the immunotherapeutic and drug therapeutic response
We utilized the online Tumor Immune Dysfunction and Exclusion (TIDE) algorithm to foresee the possibility of the immunotherapy response (Fu et al. 2020). The R package (“pRRophetic”) was utilized to assess the drug therapy response determined by the half-maximal inhibitory concentration (IC50) of each patients with NSCLC based on the Genomics of Drug Sensitivity in Cancer (GDSC) website (Garnett et al. 2012; Yang et al. 2012; Iorio et al. 2016).
Statistical analyses
In this study, statistical analyses were executed by R software (Version 4.0.2). Cox regression analyses (univariate, lasso and multivariate) were utilized to assess prognostic value. Kaplan–Meier (KM) survival analysis, was utilized to explore the OS time of the two different risk groups. Then, we verified the accurateness of this immune checkpoint-related LncRNAs model in the testing cohort and the entire cohort. A P value < 0.05 was concerned as a statistically noteworthy variance.
Results
Identification of immune checkpoint-linked LncRNAs in patients of NSCLC
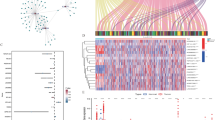
The expression matrixes of three immune checkpoint genes (PD1, PD-L1, CTLA4) and 14,086 LncRNAs were separated from the TCGA database. We characterized immune checkpoint-related LncRNA as the expression level of LncRNA, which was significantly relevant to greater than or equal to one of the three immune checkpoint genes (cutoff value: Pearson R|> 0.3, P < 0.001). At last, the immune checkpoint–LncRNA co-expression network were visualized by using the network diagram and Sankey diagram in Fig. 1A and B, respectively. 164 LncRNAs were determined as immune checkpoint-related LncRNAs.
Classification of NSCLC based on immune checkpoint-related LncRNAs
From the TGGA RNA-seq data, gene expression matrix of 1037 samples and three immune checkpoint-related genes were obtained. A total of 22 immune checkpoint-related LncRNAs were selected by Univariate cox regression analysis, based on the significant prognostic value, P < 0.05(Fig. 2A). There was a significant difference in the expression level of these LncRNAs between NSCLC and normal tissues (Fig. 2B, C). Based on the 22 immune checkpoint-related LncRNAs expression matrix and overall survival time of NSCLC patients, the non-negative matrix factorization was utilized to divide NSCLC samples into two obviously different clusters (cluster 1 and cluster 2). The determination of non-negative matrix factorization is to detect potential characteristics in gene expression landscapes by separating the original matrix into two obviously different non-negative matrices. We utilized a full correlation coefficient to ascertain the k value. The best total cluster number was set to k = 2 (the two subtypes were separated as cluster 1 and cluster 2). For example, when the cluster number k = 2, the heat map of consensus matrix showed a clear and sharp boundary, which indicated that these patients had steady and robust clusters (Fig. 3A, B).
Identification of immune checkpoints linked LncRNAs of prognostic value in patients with NSCLC. A Univariate Cox regression analysis reveals that 22 obtained immune checkpoints linked LncRNAs significantly correlate with clinical prognosis. B Heatmap for the expression difference of immune checkpoints related LncRNAs between tumor and normal tissue. C Boxplot for the expression difference of immune checkpoints related LncRNAs between tumor and normal tissue (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001)
Identification of NSCLC subtypes using NMF consensus clustering in NSCLC patients. A, B NMF clustering based on 22 immune checkpoints linked LncRNAs. Patients were separated into two clusters (cluster 1 and cluster 2). C Survival analysis of patients in Clusters 1 and 2 in the NSCLC cohort. D Heatmap of two clusters well-defined by the immune checkpoints linked LncRNAs expression. E–G The difference of PD1, PD-L1, CTLA4 expression between lung cancer tissue and normal tissue. H–J The difference of PD1, PD-L1, CTLA4 expression between cluster 1 and cluster 2
Clinical correlation analysis of cluster 1 and cluster 2
Kaplan–Meier (KM) survival analysis showed Patients in cluster 1 had an obviously worse OS than their corresponding counterparts in cluster 2 (P < 0.004, Fig. 3C). The survival rate of 50% for cluster 1 was four years; in contrast, that for cluster 2 was six years. Based on the features (Stage, Gender, Age, T, M, N, cluster), the heatmap Fig. 3D showed that 22 immune checkpoint-related LncRNAs of differences in expression levels. Compared with normal tissues, PD1 and CTLA4 were highly expressed in cancer tissues, while PD-L1 expression was low in cancer tissues (Fig. 3E–G). Compared to cluster 1, we discovered that PD1, PD-L1, and CTLA4 were highly expressed in cluster 2 (Fig. 3H–J).
TME, TIDE, immune cell infiltration landscape, and GSEA between cluster 1 and cluster 2
We found that there were noteworthy differences in ImmuneScore, StromalScore and ESTIMATEScore between the cluster 1 and cluster 2 (Fig. 4A–C). However, there is no significant difference in TIDE_score between cluster 1 and cluster 2 (Fig. 4D). Furthermore, we investigated the difference of tumor immune cell infiltration between cluster 1 and cluster 1 patients via the CIBERSORT algorithm in 22 immune cell infiltrates of NSCLC.
Identification of immune cell infiltration landscape, tumor microenvironment score, Kegg-pathway, and TIDE_Score in the two clusters. A–C ImmuneScore, StromalScore and ESTIMATEScore between cluster 1 and cluster 2. D The difference of TIDE_score between the two clusters. E The difference of immune cell infiltration between cluster 1 and cluster 2. F Kegg Pathway analysis between cluster 1 and cluster
The Wilcoxon rank-sum test was also accurately applied to explore this difference. We found that several immune cells conferred a significantly high infiltration density in the cluster 2, including T cells CD4 memory resting (P < 0.001), Dendritic cells resting (P < 0.001), Mast cells resting(P < 0.002); On other hand, B cells naïve (P = 0.038), B cells memory(P < 0.001), Plasma cells (P = 0.007), NK cells resting (P < 0.004), Macrophages M0 (P < 0.001), Mast cells activated (P = 0.003) have a high infiltration density in the cluster 1 (Fig. 4E). In light of the above analysis, we hypothesized that these two clusters were linked to immune cell infiltrates and have implications for survival outcomes. Based on Gene Set Enrichment Analysis, we found that the main active pathways of cluster 2 are KEGG_T_CELL_RECEPTOR_SIGNALING_PATHWAY, KEGG_B_CELL_RECEPTOR_SIGNALING_PATHWAY and KEGG_NATURAL_KILLER_CELL_MEDIATED_CYTOTOXICITY; On other hands, the main active pathways of cluster 1 are KEGG_CELL_CYCLE, KEGG_DNA_REPLICATION and P53_SIGNALING_PATHWAY (Fig. 4F).
Construction and validation of an immune checkpoint-related LncRNAs risk in NSCLC patients
We investigated prognostic LncRNAs from 164 immune checkpoint-associated LncRNAs based on the training cohort through univariate Cox regression analysis. The result indicated that 22 immune checkpoint-linked LncRNAs in the training cohort of TCGA dataset were meaningfully associated with OS (Fig. 2A). These 22 immune checkpoint-linked LncRNAs were used for LASSO-penalized Cox analysis and model construction. Hence, 10 immune checkpoint-related LncRNAs(including MMP2-AS1, HLA-DQB1-AS1, AC099343.3, AC006369.1, AC104971.3, LINC01281, LINC02084, AL035530.2, LINC00944, AC243960.3) are picked out from the LASSO-penalized Cox analysis (Fig. 5A, B). As a result, 10 immune checkpoint-linked LncRNAs were the prognostic features associated with OS in the TCGA training cohort and constructed to be a risk model to calculate patients’ prognosis risk with NSCLC (Table 2). NSCLC samples were categorized into low-risk and high-risk groups on the bias of the median value of the prognostic risk grade.
A risk model for patients with NSCLC from immune checkpoints linked LncRNAs. A The LASSO coefficient profile of 22 OS-correlated immune checkpoints linked LncRNAs and the perpendicular imaginary line was drawn at the value selected by tenfold cross-validation. B The tuning parameters (log λ) of patients' OS related immune checkpoints linked LncRNAs were chosen to cross validate the error curve. Along with the minimal criterion and 1-se criterion, the perpendicular imaginary line is dragged at the ideal value. C Kaplan–Meier survival curves of patients’ OS in the low- and high-risk groups. D The ROC analyses of this model in the training cohort. E The distribution of risk scores based on the immune checkpoints related LncRNAs, vital statuses of patients sorted by risk score and the ten-LncRNAs expression heatmap in the training cohort
The survival analysis proved that the patients’ OS in the high risk group was obviously shorter than that of the low risk (P < 0.001) (Fig. 5C). The ROC analyses found that this immune checkpoint-related LncRNAs model was a valuable tool to predict NSCLC patients’ OS (AUC = 0.665, Fig. 5D). The distribution of risk scores based on the ten LncRNAs, vital status of patients sorted by risk grade, and the ten LncRNAs expression were also according to these findings (Fig. 5E). To validate the prognostic competence of this recognized model, we computed risk scores for every patients in the testing cohort and the entire cohort by using the above formula. Moreover, Kaplan–Meier survival analyses carried on patients of the testing cohort and the entire cohort have no differences in the outcomes from the training cohort: the OS time of NSCLC patients with higher risk grades was worse than that of patients with lower risk grades (Fig. 6A, D). The ROC analyses uncovered that this model was an effectual instrument to predict NSCLC patient OS in the testing cohort (AUC = 0.631, Fig. 6B) and the entire cohort, respectively (AUC = 0.647, Fig. 6E). The distribution of risk scores, array of survival status and time, the expression of the immune checkpoint-related LncRNAs were showed in the testing cohort (Fig. 6C) and the entire cohort (Fig. 6F).
The prognostic value of the risk model of the ten immune checkpoints-related LncRNAs in TCGA testing and entire cohorts. A Kaplan–Meier survival curves of patients’ OS in the low- and high-risk groups for the testing cohort. B The ROC analyses of this model in the testing cohort. C The distribution of risk scores based on the immune checkpoints related LncRNAs, survival status of NSCLC patients sorted by risk score and the ten-LncRNAs expression heatmap in the testing cohort. D Kaplan–Meier survival curves of patients OS in the low- and high-risk groups for the entire cohort. E The ROC analyses of this model in the entire cohort. F The distribution of risk scores based on the immune checkpoints related LncRNAs, survival status of NSCLC patients sorted by risk score and the ten-LncRNAs expression heatmap in the entire cohort
Estimate the risk model of the immune checkpoints-linked LncRNAs and clinical characters of NSCLC patients
Univariate and multivariate Cox regression analyses was utilized to estimate whether this risk model of immune checkpoint-related LncRNAs was an independent prognostic character for NSCLC. The univariate and multivariate cox analyses revealed that Stage and risk score are independent prognostic issues of patients with NSCLC (Table 3).
Formation and assessment of the nomogram
A nomogram including the risk score and clinical features was manufactured to foresee 1, 3 and 5 year OS rates. Through comparing with clinical features, the risk score of the risk model showed powerful predictive capability in the nomogram (Fig. 7A). Correction charts presented that the detected versus predicted values of 1, 3 and 5 year OS revealed ideal consistence (Fig. 7C–E). The conformance index of the nomogram was calculated. This processes were handled to better assess the particularity and susceptibleness of risk grades in predicting OS in patients with NSCLC. With the increase of time, we found that the nomogram's concordance index was continuously greater than other clinical features, signifying that the nomogram could better predict the prognosis of patients with NSCLC (Fig. 7B).
Clinical correlation analysis and tumor immune microenvironment based on the risk model
Just as depicted in Fig. 8A–F, according to the subcategories sorted by gender, age, or tumor stage, the patients' OS of the low-risk group continuous are better to that in high-risk group. We found that there were significant differences in risk score in clinical features, cluster, age, gender, stage, T, N, and TME. The risk score of cluster 1 was significantly higher than that of cluster 2 (Fig. 8G). There was no significant difference in risk score between groups older than 65 and less than 65 (Fig. 8H). The risk score of male patients was higher than that of female patients (Fig. 8I). Patients with advanced clinical stage had higher risk scores (Fig. 8J–L). There was no significant difference in risk scores between patients with metastasis and those without metastasis due to the small number of patients considering metastasis (Fig. 8M). There was a significant difference in tumor microenvironment scores (ImmuneScore, StromalScore, and ESTIMATEScore) of the risk score (Fig. 8N–P). Based on Gene Set Enrichment Analysis, we found that the main active immune-related pathways of the low_risk group are KEGG_B_CELL_RECEPTOR_SIGNALING_PATHWAY, KEGG_CHEMOKINE_SIGNALING_PATHWAY, KEGG_NATURAL_KILLER_CELL_MEDIATED_CYTOTOXICITY, EGG_T_CELL_RECEPTOR_SIGNALING_PATHWAY, and KEGG_TGF_BETA_SIGNALING_PATHWAY; On other hand, the main active pathways of the high_risk group are KEGG_CELL_CYCLE, and KEGG_DNA_REPLICATION (Fig. 8Q).
Estimation of tumor immune microenvironment and clinical characteristics with the immune checkpoints linked LncRNAs model in the entire cohort. A–F Kaplan–Meier curves of differences in overall survival classified by gender, age, or stage between the high-risk and low-risk groups in the entire cohort. G–M The relationship between risk score and clinical characteristics. N–P Differences of risk scores in high and low tumor microenvironment score groups (ImmuneScore, StromalScore, and ESTIMATEScore). Q Kegg Pathway analysis between the high- and low-risk groups
Therapeutic responses for NSCLC
We next explored the correlations between the immune checkpoint-linked LncRNA model and immunotherapeutic markers. Compared with the high-risk group, the expression of PD1, PD-L1 and CTLA4 were dramatically higher in the low-risk group (Fig. 9A–C).
Estimation of immune checkpoint gene expression, cancer immunotherapy response with the model in the entire cohort. A–C Boxplot showed the difference of PD1, PD-L1, CTLA4 expression between high and low-risk groups. D Immune cells infiltration density between high- and low-risk patients. E TIDE prediction score between high- and low-risk patients. F–H Kaplan–Meier survival curves of OS among four patient groups stratified by the risk score and PD-1 (F), PD-L1 (G), CTLA-4 (H)
We also discovered that some immune cells conferred a expressively high infiltration density in the low-risk group, including T cells CD8 (P < 0.002), T cells CD4 memory resting (P < 0.007, T cells regulatory (Tregs) (P < 0.001), Dendritic cells resting (P < 0.001); on the other hand, macrophages M0 (P < 0.001), have a high infiltration density in the high-risk group (Fig. 9D). The correlation between risk score and immune cell infiltration is shown in supplement Fig. 1.
Unsurprisingly, we discovered that the low-risk groups could be further possible to reply to immunotherapy than high-risk groups, indicating that this immune checkpoint-related LncRNAs classifier index might be a marker for predicting TIDE (Fig. 9E).
At the same time, we compared the survival of four groups of patients with high/low-risk scores and stratified gene expression levels at immune checkpoints. As shown in Fig. 9F, the survival rate of patients with low-risk scores and low PD-1 was significantly better than that of patients with other stratification (P < 0.001). Patients with low-risk scores and high PD1 expression had more prolonged survival than patients with high-risk scores and low PD1 expression (P < 0.001) (Fig. 9F). Patients with high-risk scores and high PD1 expression had the shortest survival time. When we repeated a similar analysis using risk score and PD-L1 expression or CTLA-4 expression, we found that patients with a risk score and PD-L1 expression or CTLA-4 expression stratification showed a similar survival pattern to PD-1 (Fig. 9G, H).
We also uncovered that 44 drugs had significant variances in assessed IC50 between low-risk and high-groups; especially, patients in the low-risk group had lower IC50 values (P < 0.05; Fig. 10 and supplement Fig. 2). Together, these results supported that the immune checkpoints related LncRNAs risk model may play a critical role in predicting therapeutic drug response.
Discussion
NSCLC, at present, as the most ordinary subtype of lung cancer globally, has been paid more attention by many medical researchers to study the occurrence, development and treatment of NSCLC during the past years (Ramalingam et al. 2011; Herbst et al. 2018). Accumulating studies show that various lung tumor subtypes had diverse clinical features and outcomes, thus, more and more effective biomarkers have been found to predict survival in NSCLC nowadays (Rodríguez et al. 2021; Pradhan et al. 2021; Tang et al. 2021). PD1/PD-L1and CTLA4 were the most intensely studied immune checkpoint at present, and related immune checkpoint inhibitors have been widely used in the treatment of advanced cancer, especially in melanoma (Liu et al. 2019), non-small cell lung cancer (Yang et al. 2018; Chen et al. 2020). Traditional immune checkpoint inhibitors, such as pembrolizumab and nivolumab have been used in all kinds of solid tumors, which have produced an exciting results (Llovet et al. 2021; Morad et al. 2021; Carlino et al. 2021). While new drugs such as M7824 and YM101 (Grote et al. 2020; Cheng et al. 2022; Paz-Ares et al. 2020) in NSCLC have showed good results. M7824 (Lind et al. 2020) and YM101 (Yi et al. 2021a) can simultaneously targeting TGF-β and PD-L1, provides a new choice for immunotherapy. The researchers found that M7824 and YM101 (Yi et al. 2021b) can overcome immunotherapy resistance, It provides a new choice for clinical decision-making. Immunotherapy combined with other targeted therapy will become a choice for more patients in the future. At the same time, drugs with immune checkpoint inhibition and a variety of targeted functions will also become a new direction for researchers.
However, the expression level of these three immune checkpoint genes in cancer is not directly related to the effect of immunotherapy and overall survival (Goodman et al. 2018).LncRNAs, a recently recognized vital player of the genome regulatory network, have also been discovered to act key roles in the growth and motivation of immune cells, may serve as peculiar molecular markers for infiltrating immune cells in TME (Hong et al. 2020; Zhang et al. 2020; Xu et al. 2019; Huang et al. 2018; Jiang et al. 2018). However, genome-wide screening of definite immune checkpoint-related LncRNA and their value in evaluating immune cell infiltrate of tumor, tumor microenvironment, therapeutic responses and clinical outcome has barely been explored. In this study, we first reannotated immune checkpoint-related LncRNA of NSCLC from the TCGA dataset. 164 LncRNAs were discerned as immune checkpoint-linked LncRNAs. Moreover the TCGA NSCLC dataset approved the prognostic value of 22 immune checkpoint-related LncRNAs, which were applied to NMF to divide NSCLC samples into two different clusters (cluster 1 and cluster 2). There was a significant difference in overall survival between cluster 1 and cluster 2. There were significant differences in gene expression of immune checkpoint genes, immune cell infiltration, tumor microenvironment score (ImmuneScore, StromalScore, and ESTIMATEScore) and active Kegg pathway between the two clusters. However, no significant difference in the TIDE_score used to evaluate immunotherapy response between the two clusters. This may be related to the small sample size of cluster2. Ten immune checkpoint-linked LncRNAs (including MMP2-AS1, HLA-DQB1-AS1, AC099343.3, AC006369.1, AC104971.3, LINC01281, LINC02084, AL035530.2, LINC00944, AC243960.3) were applied to construct an immune checkpoint-linked LncRNA model to foresee the OS of NSCLC patients. Except for LINC00944, others are protective factors to the patient with NSCLC. Generally speaking, there are few studies on non-coding RNA related to immune checkpoints.
Some researchers reported that HLA-DQB1-AS1 is an immune-related non-coding RNA associated with the prognosis of patients with lung cancer (Jin et al. 2020). The researchers found that HLA-DQB1-AS1 is related to the occurrence and development of chronic obstructive pulmonary disease (Qian et al. 2018). Recent research has found LINC01281 is related to the prognosis of patients with laryngeal carcinoma (Zhang et al. 2019). Through bioinformatics analysis, the researchers found that LINC02084 is associated with the prognosis of liver cancer (Lv et al. 2018) and renal clear cell carcinoma (Sun et al. 2020). LINC00944 is associated with breast cancer prognosis (Santiago et al. 2021). The research on these non-coding RNAs (AC099343.3, MMP2-AS1, AC006369.1, AC104971.3, AL035530.2, AC243960.3) has not been reported. The mechanism of the role of these non-coding RNAs in cancer is worthy of further study.
According to the above ten immune checkpoint-related LncRNAs mode, patients with NSCLC were separated into high-risk and low-risk groups. We found that the high-risk set had an outwardly poor clinical result. Multivariate Cox regression analyses displayed that the immune checkpoints associated LncRNAs model was an independent risk factors of OS. We also constructed a nomogram displaying wonderful consistency between the surveyed and forecast rates for 1, 3 and 5 year OS of NSCLC patients. Lastly, the observed 1, 3 and 5 year OS predicting rates presented excellent consistency. Based on the model, we found that there were some differences and correlations in risk score between the two cluster, clinical stages, tumor microenvironment score, gene expression of the immune checkpoint, TIDE_score and immune cell infiltration. Above all, the model based on ten immune checkpoint-related LncRNAs, which are self-reliantly associated with OS is equally accurate. Calculation of the TIDE score suggested that patients with low-risk subgroup have a better response to immunotherapy. Giving the above consequences, we suppose that this forecast model may have the capacity to provide dependable immune biomarkers for cancer therapy. In the meantime, it could give us a new mind in exploring the molecular biological mechanism of the immune checkpoint-related LncRNA in NSCLC. We know that the stage is the critical feature of the prognosis of NSCLC. But, patients with NSCLC at the similar stage constantly have diverse clinical outcomes, signifying that the existing periodization classifications in providing dependable forecasts and revealing the heterogeneity of NSCLC are imprecise. Thus, the underlying predictive and therapeutic markers should be discovered. The established immune checkpoint-related LncRNA model delivers us a new technique for prognostic expectation in NSCLC patients. The outcomes also afford thinking for future investigations on the procedure and mechanism of immune checkpoints modification of LncRNAs.
In addition, the function of LncRNAs and possible interaction with the immune checkpoints gene need to be demonstrated by in vivo and in vitro experiments.
In conclusion, we performed an integrative analysis of LncRNAs related to immune checkpoints and emphasized the significance and value of NSCLC subtypes identification, immune checkpoints related LncRNAs in estimating the Tumor microenvironment score, immune cell infiltration of the tumor, immunotherapy, and chemotherapy response. Additionally, for the first time, our research identified and validated an immune checkpoints-related LncRNA signature based on ten LncRNAs, which has self-governing prognostic implication for NSCLC patients. Finally, we suggested the potential of immune checkpoints-related LncRNA signature as a valuable predictive markers of immunotherapy response to allow a more precise selection of patients with NSCLC who will take advantage of immune checkpoint inhibitor immunotherapy, which need more immunotherapy data sets to further validating.
Data availability
Data are available in a public, open access repository. The data sets analyzed during the current study are available in The Cancer Genome Atlas (TCGA) (https://cancergenome.nih.gov/).
References
Afzal M, Mercado R, Shirai K (2018) Efficacy of metformin in combination with immune checkpoint inhibitors (anti-PD-1/anti-CTLA-4) in metastatic malignant melanoma. J Immunother Cancer 6:64
Atianand MK, Caffrey DR, Fitzgerald KA (2017) Immunobiology of long noncoding RNAs. Annu Rev Immunol 35:177–198
Brahmer J, Reckamp KL, Baas P, Crinò L, Eberhardt WE, Poddubskaya E, Antonia S, Pluzanski A, Vokes EE, Holgado E, Waterhouse D, Ready N, Gainor J, Arén Frontera O, Havel L, Steins M, Garassino MC, Aerts JG, Domine M, Paz-Ares L, Reck M, Baudelet C, Harbison CT, Lestini B, Spigel DR (2015) Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med 373:123–135
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(2018):394–424
Carlino M, Larkin J, Long G (2021) Immune checkpoint inhibitors in melanoma. Lancet (lond, England) 398:1002–1014
Chen YG, Satpathy AT, Chang HY (2017) Gene regulation in the immune system by long noncoding RNAs. Nat Immunol 18:962–972
Chen D, Menon H, Verma V, Guo C, Ramapriyan R, Barsoumian H, Younes A, Hu Y, Wasley M, Cortez MA, Welsh J (2020) Response and outcomes after anti-CTLA4 versus anti-PD1 combined with stereotactic body radiation therapy for metastatic non-small cell lung cancer: retrospective analysis of two single-institution prospective trials. J Immunother Cancer 8:e000492
Cheng B, Ding K, Chen P, Ji J, Luo T, Guo X, Qiu W, Ma C, Meng X, Wang J, Yu J, Liu Y (2022) Anti-PD-L1/TGF-βR fusion protein (SHR-1701) overcomes disrupted lymphocyte recovery-induced resistance to PD-1/PD-L1 inhibitors in lung cancer. Cancer Commun (lond, England) 42:17–36
de Santiago P, Blanco A, Morales F, Marcelain K, Harismendy O, Sjöberg Herrera M, Armisén R (2021) Immune-related lncRNA LINC00944 responds to variations in ADAR1 levels and it is associated with breast cancer prognosis. Life Sci 268:118956
Fritz JM, Lenardo MJ (2019) Development of immune checkpoint therapy for cancer. J Exp Med 216:1244–1254
Fu J, Li K, Zhang W, Wan C, Zhang J, Jiang P, Liu XS (2020) Large-scale public data reuse to model immunotherapy response and resistance. Genome Med 12:21
Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW, Greninger P, Thompson IR, Luo X, Soares J, Liu Q, Iorio F, Surdez D, Chen L, Milano RJ, Bignell GR, Tam AT, Davies H, Stevenson JA, Barthorpe S, Lutz SR, Kogera F, Lawrence K, McLaren-Douglas A, Mitropoulos X, Mironenko T, Thi H, Richardson L, Zhou W, Jewitt F, Zhang T, O’Brien P, Boisvert JL, Price S, Hur W, Yang W, Deng X, Butler A, Choi HG, Chang JW, Baselga J, Stamenkovic I, Engelman JA, Sharma SV, Delattre O, Saez-Rodriguez J, Gray NS, Settleman J, Futreal PA, Haber DA, Stratton MR, Ramaswamy S, McDermott U, Benes CH (2012) Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483:570–575
Garon EB, Rizvi NA, Hui R, Leighl N, Balmanoukian AS, Eder JP, Patnaik A, Aggarwal C, Gubens M, Horn L, Carcereny E, Ahn MJ, Felip E, Lee JS, Hellmann MD, Hamid O, Goldman JW, Soria JC, Dolled-Filhart M, Rutledge RZ, Zhang J, Lunceford JK, Rangwala R, Lubiniecki GM, Roach C, Emancipator K, Gandhi L (2015) Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 372:2018–2028
Gaujoux R, Seoighe C (2010) A flexible R package for nonnegative matrix factorization. BMC Bioinform 11:367
Goodman AM, Piccioni D, Kato S, Boichard A, Wang HY, Frampton G, Lippman SM, Connelly C, Fabrizio D, Miller V, Sicklick JK, Kurzrock R (2018) Prevalence of PDL1 amplification and preliminary response to immune checkpoint blockade in solid tumors. JAMA Oncol 4:1237–1244
Grote H, Feng Z, Schlichting M, Helwig C, Ruisi M, Jin H, Scheuenpflug J, Gann C, Su Z, Reck M, Vokes E, Kerr K (2020) Programmed death-ligand 1 immunohistochemistry assay comparison studies in NSCLC: characterization of the 73–10 assay. J Thorac Oncol: off Publ Int Assoc Study Lung Cancer 15:1306–1316
Herbst RS, Morgensztern D, Boshoff C (2018) The biology and management of non-small cell lung cancer. Nature 553:446–454
Hong W, Liang L, Gu Y, Qi Z, Qiu H, Yang X, Zeng W, Ma L, Xie J (2020) Immune-Related lncRNA to construct novel signature and predict the immune landscape of human hepatocellular carcinoma. Mol Ther Nucleic Acids 22:937–947
Huang D, Chen J, Yang L, Ouyang Q, Li J, Lao L, Zhao J, Liu J, Lu Y, Xing Y, Chen F, Su F, Yao H, Liu Q, Su S, Song E (2018) NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat Immunol 19:1112–1125
Iorio F, Knijnenburg TA, Vis DJ, Bignell GR, Menden MP, Schubert M, Aben N, Gonçalves E, Barthorpe S, Lightfoot H, Cokelaer T, Greninger P, van Dyk E, Chang H, de Silva H, Heyn H, Deng X, Egan RK, Liu Q, Mironenko T, Mitropoulos X, Richardson L, Wang J, Zhang T, Moran S, Sayols S, Soleimani M, Tamborero D, Lopez-Bigas N, Ross-Macdonald P, Esteller M, Gray NS, Haber DA, Stratton MR, Benes CH, Wessels LFA, Saez-Rodriguez J, McDermott U, Garnett MJ (2016) A landscape of pharmacogenomic interactions in cancer. Cell 166:740–754
Ishida Y, Agata Y, Shibahara K, Honjo T (1992) Induced expression of PD-1, a novel member of the immunoglobulin gene superfamily, upon programmed cell death. EMBO J 11:3887–3895
Jiang M, Zhang S, Yang Z, Lin H, Zhu J, Liu L, Wang W, Liu S, Liu W, Ma Y, Zhang L, Cao X (2018) Self-recognition of an inducible Host lncRNA by RIG-I feedback restricts innate immune response. Cell 173:906-919.e913
Jin D, Song Y, Chen Y, Zhang P (2020) Identification of a seven-lncRNA immune risk signature and construction of a predictive nomogram for lung adenocarcinoma. Biomed Res Int 2020:7929132
Keir ME, Butte MJ, Freeman GJ, Sharpe AH (2008) PD-1 and its ligands in tolerance and immunity. Annu Rev Immunol 26:677–704
Kuleshov MV, Diaz JEL, Flamholz ZN, Keenan AB, Lachmann A, Wojciechowicz ML, Cagan RL, Ma’ayan A (2019) modEnrichr: a suite of gene set enrichment analysis tools for model organisms. Nucleic Acids Res 47:W183-w190
Lind H, Gameiro S, Jochems C, Donahue R, Strauss J, Gulley J, Palena C, Schlom J (2020) Dual targeting of TGF-β and PD-L1 via a bifunctional anti-PD-L1/TGF-βRII agent: status of preclinical and clinical advances. J Immunother Cancer 8:e000433
Liu D, Schilling B, Liu D, Sucker A, Livingstone E, Jerby-Arnon L, Zimmer L, Gutzmer R, Satzger I, Loquai C, Grabbe S, Vokes N, Margolis C, Conway J, He M, Elmarakeby H, Dietlein F, Miao D, Tracy A, Gogas H, Goldinger S, Utikal J, Blank C, Rauschenberg R, von Bubnoff D, Krackhardt A, Weide B, Haferkamp S, Kiecker F, Izar B, Garraway L, Regev A, Flaherty K, Paschen A, Van Allen E, Schadendorf D (2019) Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat Med 25:1916–1927
Llovet J, Castet F, Heikenwalder M, Maini M, Mazzaferro V, Pinato D, Pikarsky E, Zhu A, Finn R (2021) Immunotherapies for hepatocellular carcinoma. Nat Rev Clin Oncol 19:151
Lv Y, Wei W, Huang Z, Chen Z, Fang Y, Pan L, Han X, Xu Z (2018) Long non-coding RNA expression profile can predict early recurrence in hepatocellular carcinoma after curative resection. Hepatol Res: off J Japan Soc Hepatol 48:1140–1148
McDermott DF, Huseni MA, Atkins MB, Motzer RJ, Rini BI, Escudier B, Fong L, Joseph RW, Pal SK, Reeves JA, Sznol M, Hainsworth J, Rathmell WK, Stadler WM, Hutson T, Gore ME, Ravaud A, Bracarda S, Suárez C, Danielli R, Gruenwald V, Choueiri TK, Nickles D, Jhunjhunwala S, Piault-Louis E, Thobhani A, Qiu J, Chen DS, Hegde PS, Schiff C, Fine GD, Powles T (2018) Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat Med 24:749–757
Meerveld-Eggink A, Rozeman E, Lalezari F, van Thienen J, Haanen J, Blank C (2017) Short-term CTLA-4 blockade directly followed by PD-1 blockade in advanced melanoma patients: a single-center experience. Ann Oncol: off J Eur Soc Med Oncol 28:862–867
Monjazeb A, Giobbie-Hurder A, Lako A, Thrash E, Brennick R, Kao K, Manuszak C, Gentzler R, Tesfaye A, Jabbour S, Alese O, Rahma O, Cleary J, Sharon E, Mamon H, Cho M, Streicher H, Chen H, Ahmed M, Mariño-Enríquez A, Kim-Schulze S, Gnjatic S, Maverakis E, Marusina A, Merleev A, Severgnini M, Pfaff K, Lindsay J, Weirather J, Ranasinghe S, Spektor A, Rodig S, Hodi S, Schoenfeld J (2021) A randomized trial of combined PD-L1 and CTLA-4 inhibition with targeted low-dose or hypofractionated radiation for patients with metastatic colorectal cancer. Clin Cancer Res: off J Am Assoc Cancer Res 27:4940
Morad G, Helmink B, Sharma P, Wargo J (2021) Hallmarks of response, resistance, and toxicity to immune checkpoint blockade. Cell 184:5309
Moran VA, Perera RJ, Khalil AM (2012) Emerging functional and mechanistic paradigms of mammalian long non-coding RNAs. Nucleic Acids Res 40:6391–6400
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, Hoang CD, Diehn M, Alizadeh AA (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12:453–457
Paz-Ares L, Kim T, Vicente D, Felip E, Lee D, Lee K, Lin C, Flor M, Di Nicola M, Alvarez R, Dussault I, Helwig C, Ojalvo L, Gulley J, Cho B (2020) Bintrafusp alfa, a bifunctional fusion protein targeting TGF-β and PD-L1, in second-line treatment of patients with NSCLC: results from an expansion cohort of a phase 1 trial. J Thorac Oncol: off Publ Int Assoc Study Lung Cancer 15:1210–1222
Perets R, Bar J, Rasco D, Ahn M, Yoh K, Kim D, Nagrial A, Satouchi M, Lee D, Spigel D, Kotasek D, Gutierrez M, Niu J, Siddiqi S, Li X, Cyrus J, Chackerian A, Chain A, Altura R, Cho B (2021) Safety and efficacy of quavonlimab, a novel anti-CTLA-4 antibody (MK-1308), in combination with pembrolizumab in first-line advanced non-small-cell lung cancer. Ann Oncol: off J Eur Soc Med Oncol 32:395–403
Pradhan M, Chocry M, Gibbons D, Sepesi B, Cascone T (2021) Emerging biomarkers for neoadjuvant immune checkpoint inhibitors in operable non-small cell lung cancer. Transl Lung Cancer Res 10:590–606
Qian Y, Mao Z, Shi Y, Liu Z, Cao Q, Zhang Q (2018) Comprehensive analysis of miRNA-mRNA-lncRNA networks in non-smoking and smoking patients with chronic obstructive pulmonary disease. Cell Physiol Biochem : Int J Expe Cell Physiol Biochem Pharmacol 50:1140–1153
Ramalingam SS, Owonikoko TK, Khuri FR (2011) Lung cancer: new biological insights and recent therapeutic advances. CA Cancer J Clin 61:91–112
Reimand J, Isserlin R, Voisin V, Kucera M, Tannus-Lopes C, Rostamianfar A, Wadi L, Meyer M, Wong J, Xu C, Merico D, Bader GD (2019) Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat Protoc 14:482–517
Rodríguez M, Ajona D, Seijo L, Sanz J, Valencia K, Corral J, Mesa-Guzmán M, Pío R, Calvo A, Lozano M, Zulueta J, Montuenga L (2021) Molecular biomarkers in early stage lung cancer. Transl Lung Cancer Res 10:1165–1185
Sansom D (2000) CD28, CTLA-4 and their ligands: who does what and to whom? Immunology 101:169–177
Shinohara T, Taniwaki M, Ishida Y, Kawaichi M, Honjo T (1994) Structure and chromosomal localization of the human PD-1 gene (PDCD1). Genomics 23:704–706
Sun Z, Jing C, Xiao C, Li T (2020) Long non-coding RNA profile study identifies an immune-related lncRNA prognostic signature for kidney renal clear cell carcinoma. Front Oncol 10:1430
Takeuchi Y, Tanemura A, Tada Y, Katayama I, Kumanogoh A, Nishikawa H (2017) Clinical response to PD-1 blockade correlates with a sub-fraction of peripheral central memory CD4+ T cells in patients with malignant melanoma. Int Immunol 30:13–22
Tang S, Li S, Liu T, He Y, Hu H, Zhu Y, Tang S, Zhou H (2021) MicroRNAs: emerging oncogenic and tumor-suppressive regulators, biomarkers and therapeutic targets in lung cancer. Cancer Lett 502:71–83
Tomczak K, Czerwinska P, Wiznerowicz M (2015) The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (poznan, Poland) 19:A68-77
Topalian S, Hodi F, Brahmer J, Gettinger S, Smith D, McDermott D, Powderly J, Carvajal R, Sosman J, Atkins M, Leming P, Spigel D, Antonia S, Horn L, Drake C, Pardoll D, Chen L, Sharfman W, Anders R, Taube J, McMiller T, Xu H, Korman A, Jure-Kunkel M, Agrawal S, McDonald D, Kollia G, Gupta A, Wigginton J, Sznol M (2012) Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med 366:2443–2454
Turner M, Galloway A, Vigorito E (2014) Noncoding RNA and its associated proteins as regulatory elements of the immune system. Nat Immunol 15:484–491
Xu M, Xu X, Pan B, Chen X, Lin K, Zeng K, Liu X, Xu T, Sun L, Qin J, He B, Pan Y, Sun H, Wang S (2019) LncRNA SATB2-AS1 inhibits tumor metastasis and affects the tumor immune cell microenvironment in colorectal cancer by regulating SATB2. Mol Cancer 18:135
Yang W, Jorge S, Patricia G, Edelman EJ, Howard L, Simon F, Nidhi B, Dave B, Smith JA, Richard TI (2012) Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 41:D955
Yang X, Yin R, Xu L (2018) Neoadjuvant PD-1 blockade in resectable lung cancer. N Engl J Med 379:e14
Yao RW, Wang Y, Chen LL (2019) Cellular functions of long noncoding RNAs. Nat Cell Biol 21:542–551
Yi T, Chen Y, Wang L, Du G, Huang D, Zhao D, Johnston H, Young J, Todorov I, Umetsu DT, Chen L, Iwakura Y, Kandeel F, Forman S, Zeng D (2009) Reciprocal differentiation and tissue-specific pathogenesis of Th1, Th2, and Th17 cells in graft-versus-host disease. Blood 114:3101–3112
Yi M, Zhang J, Li A, Niu M, Yan Y, Jiao Y, Luo S, Zhou P, Wu K (2021a) The construction, expression, and enhanced anti-tumor activity of YM101: a bispecific antibody simultaneously targeting TGF-β and PD-L1. J Hematol Oncol 14:27
Yi M, Niu M, Zhang J, Li S, Zhu S, Yan Y, Li N, Zhou P, Chu Q, Wu K (2021b) Combine and conquer: manganese synergizing anti-TGF-β/PD-L1 bispecific antibody YM101 to overcome immunotherapy resistance in non-inflamed cancers. J Hematol Oncol 14:146
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RGW (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612
Zhang G, Fan E, Zhong Q, Feng G, Shuai Y, Wu M, Chen Q, Gou X (2019) Identification and potential mechanisms of a 4-lncRNA signature that predicts prognosis in patients with laryngeal cancer. Hum Genom 13:36
Zhang L, Xu W, Gao X, Li W, Qi S, Guo D, Ajayi O, Ding S, Wu Q (2020) lncRNA sensing of a viral suppressor of RNAi activates non-canonical innate immune signaling in drosophila. Cell Host Microbe 27:115-128.e118
Zhang L, Sun L, Zhou Y, Yu J, Lin Y, Wasan H, Shen M, Ruan S (2021) Association of survival and immune-related adverse events with anti-PD-1/PD-L1 and anti-CTLA-4 inhibitors, alone or their combination for the treatment of cancer: a systematic review and meta-analysis of 13 clinical trials. Front Oncol 11:575457
Acknowledgements
We sincerely acknowledge the contributions from the TCGA project.
Funding
There is no funding.
Author information
Authors and Affiliations
Contributions
HZ, ML and GD designed the experiments and wrote the paper, contributed equally to this work; BY, XM, YG, and LC, collected data and revised this manuscript; XL and BT conducted and Financial the experiments.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Patient consent for publication
Not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2022_3940_MOESM2_ESM.tif
Supplementary file2 Sup_Figure2: Different chemotherapy responses in low- and high- risk NSCLC patients. (P < 0.05) (TIF 11547 KB)
Rights and permissions
About this article
Cite this article
Zhang, H., Liu, M., Du, G. et al. Immune checkpoints related-LncRNAs can identify different subtypes of lung cancer and predict immunotherapy and prognosis. J Cancer Res Clin Oncol 148, 1597–1612 (2022). https://doi.org/10.1007/s00432-022-03940-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-022-03940-3