Abstract
Purpose
This study aimed to investigate the efficacy of digital mammography (DM), digital breast tomosynthesis (DBT), diffusion-weighted (DW) and dynamic contrast-enhanced (DCE) MRI separately and combined in the prediction of molecular subtypes of breast cancer.
Methods
A total of 241 patients were enrolled and underwent breast MD, DBT, DW and DCE scans. Radiomics features were calculated from intra- and peritumoral regions, and selected with least absolute shrinkage and selection operator (LASSO) regression to develop radiomics signatures (RSs). Prediction performance of intra- and peritumoral regions in the four modalities were evaluated and compared with area under the receiver-operating characteristic (ROC) curve (AUC), specificity and sensitivity as comparison metrics.
Results
The RSs derived from combined intra- and peritumoral regions improved prediction AUCs compared with those from intra- or peritumoral regions alone. DM plus DBT generated better AUCs than the DW plus DCE on predicting Luminal A and Luminal B in the training (Luminal A: 0.859 and 0.805; Luminal B: 0.773 and 0.747) and validation (Luminal A: 0.906 and 0.853; Luminal B: 0.807 and 0.784) cohort. For the prediction of HER2-enriched and TN, the DW plus DCE yielded better AUCs than the DM plus DBT in the training (HER2-enriched: 0.954 and 0.857; TN: 0.877 and 0.802) and validation (HER2-enriched: 0.974 and 0.907; TN: 0.938 and 0.874) cohort.
Conclusions
Peritumoral regions can provide complementary information to intratumoral regions for the prediction of molecular subtypes. Compared with MRI, the mammography showed higher AUCs for the prediction of Luminal A and B, but lower AUCs for the prediction of HER2-enriched and TN.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Breast cancer, as a heterogeneous disease at clinical levels, can be classified into four main intrinsic molecular subtypes: Luminal A, Luminal B, human epidermal growth factor receptor 2 (HER2) -enriched, and triple negative (TN) based on hormone status by immunohistochemical (IHC) analyses (Koboldt et al. 2012; Perou et al. 2000). Histologically similar tumors with different molecular subtypes may have substantial differences in phenotype and prognoses, and responds differently to chemotherapy and radiation therapy (Goldstein et al. 2007; Rouzier et al. 2005; Wang et al. 2011). Generally, luminal tumors are considered as good candidates for endocrine therapy (Ma et al. 2019). While, the HER2-enriched patients are often responders to targeted antibody therapy (Lam et al. 2014). The triple-negative tumors, generally considered as having the worst prognosis, may respond to chemotherapy (Huber et al. 2009). Therefore, preoperative assessments of molecular subtypes of breast cancer have been involved in individual treatment planning (Ye et al. 2020).
Routine clinical tests on subtypes of breast cancer rely on invasive tissue sampling and genetic profiling, which have inherent limitations since the biopsy performed at a single time point cannot reflect genetic heterogeneity within the breast tumor (Zardavas et al. 2015; Orlando et al. 2016). As an alternate, clinical imaging is a non-invasive and efficient approach for clinical evaluations on breast cancer (Waks et al. 2019). Previous studies have revealed associations between molecular subtypes with some imaging characteristics visually assessed with mammography and MRI (Luck et al. 2008; Uematsu et al. 2009). Recent radiomics-based evaluations of the four main intrinsic molecular subtypes in breast cancer have also been proposed by extraction and analysis of quantitative features from medical images. However, the published findings are limited due to the use of single modality and lack of comparison among different modalities (Fan et al. 2017; Monti et al. 2018). Although MRI has advantages in breast tissue characterization, it is not suitable for every patient, and has disadvantages such as high examination fees and relatively low specificity (Orel and Schnall 2001). Mammography, on the other hand, is widely used for breast screening and highly accessible for all patients. The digital breast tomosynthesis (DBT) generates 3D tomographic images reconstructed from multiple scanned images by rotating the X-ray tubes, and can decrease recall rates and increase cancer detection rates compared with full-field digital mammography (FFDM) (Fischer et al. 2002). While, to our knowledge, direct and quantified comparisons between mammography and MRI on the prediction of molecular subtypes of breast cancer have not been investigated.
In addition, previous works related to the evaluation of four intrinsic molecular subtypes all focused on intratumoral regions, which was inadequate to consider areas surrounding the breast tumor. While, increasing evidences have highlighted that peripheral breast tissues may also hold great biological information related to lymphatics and blood vessels and immune responses (Braman et al. 2019). Some recent attempts have suggested associations between features hidden in the peritumoral regions with breast tumor characteristics and gene status (Braman et al. 2017). While, to our knowledge, power of peritumoral regions on the prediction of the four intrinsic molecular subtypes in breast cancer has yet to be studied. Therefore, this study aims to widen the understanding of assessment for molecular subtypes by analyzing intratumoral and peritumoral radiomics, and directly and quantitatively compare the prediction efficacy of breast mammography and MRI individually and in combination.
Materials and methods
Patients
The retrospective research was approved by the ethics committee of our hospital, and the informed requirement was waived. A total of 583 patients were enrolled between January 2016 and Match 2021. The inclusion criteria were: (1) patients pathology-proven breast cancer; and (2) underwent breast mammography and MRI examinations before treatment. Exclusion criteria were: (1) patients with incomplete pathological data; (2) combined with other tumor diseases; (3) received radiotherapy or chemotherapy before mammography or MRI examinations; and (4) during pregnancy or lactation periods. According to the criteria, 241 patients were finally included, 162 were used as a training group and 79 as a validation group. Among the patients, 54 were Luminal A, 96 were Luminal B, 47 were HER2-enriched, and 44 were triple negative (TN). Clinical factors including age, menopausal status and family history were retrieved from the electronic medical recodes system (EMRS) of our hospital. Pathological data include estrogen receptor (ER), progesterone receptor (PR), antigen identified by monoclonal antibody (Ki-67), human epidermal growth factor receptor 2 (HER2), and lymph node metastatic (LNM) status. Figure 1 shows the process for recruitment patients.
Pathological assessment
Pathology reports for each patient that included the expression levels of ER, PR, and antigen identified by monoclonal antibody Ki-67/HER2 were based on postoperative tissue samples. Molecular subtypes that included Luminal A, Luminal B, HER2-enriched and TN were determined according to a previous report (Goldhirsch et al. 2011). ER and/or PR positive, HER2 negative and low Ki-67 cases were classified as the Luminal A subtype. ER and/or PR positive, HER2 negative and high expression of Ki-67 or PR negative cases were classified as the Luminal B- (HER2 negative) subtype. ER and/or PR positive, and HER2 positive cases were classified as the Luminal B + (HER2 positive) subtype. ER negative, PR negative and HER2 positive cases were classified as the HER2-enriched subtype. ER negative, PR negative and HER2 negative cases were classified as the TN subtype. The fluorescence in situ hybridization (FISH) analysis (Fehrenbacher et al. 2020) was performed to further determine the HER2 status for HER2 (2+) with the result of 2.0 or higher indicated positivity.
Data acquisition
Breast mammography screenings were performed using a Hologic scanner (Hologic Selenia Dimensions, HOLGIC, USA). The mammography images were reconstructed with 1-mm intersection spacing to generate a three-dimensional view of the breast tissues and inserted on a Hologic breast computer-aided diagnosis (CAD) workstation (SecureViewDx; Holo gic) equipped with two 5-megapixel monitors. The voltage on the X-ray tubes ranges from 20.0 to 49.0 kV (step: 1.0 kV), reconstruction time: 2.0–5.0 s, nominal power: 3.0 kW, current time range: 300–400 m As, scanning time < 4.0 s, and pixel size: 70 μm.
Breast MR screenings were performed using a 1.5-T MR scanner (HDx, GE Healthcare), using an 8-channel breast dedicated coil in prone position. Parameters for the MR scans were as follows: DWI MRI, b-value: 800 s/mm2, TR: 5000 ms, TE: 64 ms, slice thickness: 6 mm. DCE MRI, TR: 6.2 ms, TE: 3.0 ms; slice thickness: 3.2 mm; 48 slices per volume. The contrast agent was injected intravenously (0.1 mmoL/kg of Gd-DTPA-MBA, Omniscan, GE Healthcare), followed by a 20 mL saline flush, both at rates of 3 mL/s. After intravenous injection, continuous non-interval scans were performed in eight phases, with a scan time of 43 s for each phase. All scanned images were stored in the Picture Archiving and Communication System (PACS) in our hospital in Digital Imaging and Communications in Medicine (DICOM) format.
Tumor segmentation and mask dilation
The region of interests (ROIs) were drawn by a senior radiologists with 10 years of work experience in the DM, DBT and MRI image using the ITK-SNAP software (version 3.6.0) to generate masks of intratumoral ROIs, and stored in a .NII format. The radiologist was blinded to the pathologic results. To explore predictive values of peritumoral regions, the original mask of the intratumoral ROI was radially dilated with a distance of 4 mm outside the tumor. Then, the intratumoral ROI was subtracted from the dilated ROI to generate the peritumoral ROI. The masks of intratumoral and peritumoral ROIs were used to extract radiomics features from intra- and peritumoral regions, respectively. Figure 2 shows an example of the ROI segmentation and mask dilation process.
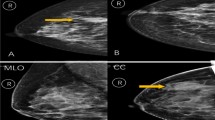
Examples of the breast cancer images and ROIs used in this study. DM (a and e), DBT (b and f), DWI (c and g) and DCE (d and h) images of a 41-year-old female TN breast cancer patient. The above row represents original images, whereas the bottom row represents intra- and peritumoral ROIs. Blue regions represent the manually delineated intratumoral ROIs covering the whole tumor, whereas red regions indicate peritumoral ROIs
Radiomics feature extraction and selection
Three categories of radiomics features were extracted from DM, DBT, DW and DCE MRI: first-order, shape-based and texture features. All features were extracted using the “Pyradiomics” package in Python v.3.6. The texture features consist of gray level cooccurrence matrix (GLCM), gray level run length matrix (GLRLM), gray level size zone matrix (GLSZM), neighboring gray tone difference matrix (NGTDM), and gray level-dependence matrix (GLDM). The original images were transformed by eight filters (Wavelet, LoG, Square, SquareRoot, Logarithm, Exponential, Gradient and LocalBinaryPattern2D) to generate eight filtered image types, from which the first-order and texture features were also derived. The feature extraction methodology, feature types, filters and parameter settings were described in the Pyradiomics documentation (https://pyradiomics.readthedocs.io/).
The intraclass correlation coefficient (ICC) analysis was performed with 40 randomly selected patients to evaluate the reproducibility of the features. Feature with ICC > 0.80 was considered to have good reproducibility and remained. Then, the Mann–Whitney U test was used to select the features, with P < 0.05 were considered significantly different. Afterwards, the least absolute shrinkage and selection operator (LASSO) was applied to select the most predictive features with tenfold cross-validation for selecting the parameter lambda using the 1 standard error of the minimum criteria (1-SE criteria). Finally, the features were selected with the logistic regression model using Akaike information criterion (AIC) as the stopping rule (Sauerbrei et al. 2007).
Construction of the radiomics signatures
Radiomics signatures (RSs) were constructed separately to predict the four molecular subtypes (Luminal A vs. non-Luminal A, Luminal B vs. non-Luminal B, HER2-enriched vs. non-HER2-enriched and TN vs. non-TN) based on the selected features and their corresponding LASSO coefficients. RSs derived from the intra- and peritumoral regions and their combination were developed and named as intra-RS, peri-RS and com-RS, respectively.
Statistical analysis
Categorical variables were compared with the Chi-Square test and Fisher’s exact test. Continuous values were evaluated with the Student’s t-test and Mann–Whitney U test. The normality test for continuous variables was verified by the Shapiro–Wilk test. All hypothesis tests were two-sided, with P < 0.05 considered to indicate statistically significant differences. Prediction performance of the RSs were evaluated based on receiver-operating characteristic (ROC) analysis with the maximum Youden index (Ruopp et al. 2008) for obtaining the optimal cutoff values using the “pROC” package in R v.3.6.
Results
Clinical characteristics
Clinical and pathologic characteristics of each patient are shown in Table 1. Median ages of the patients were 48, 54, 53 and 50 years for the Luminal A, Luminal B, HER2-enriched and TN, respectively. There was no significant difference in the mean age, menopausal status, family history and LNM status among the patients (P > 0.05) in training and validation cohorts, except for the LMN status with P = 0.043 on Luminal B in the validation cohort.
Prediction performance of intratumoral, peritumoral regions, and their combination
Prediction performance of Intra-RS, Peri-RS and Com-RS in DM, DBT, DWI and DCE are shown in Table 2. The Intra-RS and Peri-RS derived from the intratumoral and peritumoral regions, respectively, generated similar prediction capabilities in terms of AUC, sensitivity and specificity. While, by combing information from intratumoral and peritumoral regions, the Com-RS significantly improved AUCs compared with the Intra-RS on predicting the four molecular subtypes.
Comparison of prediction performance of mammography and MRI
Predictive performance of Com-RSs derived from mammography and MRI techniques is evaluated and listed in Table 3. Figure 3 shows ROC curves of the Com-RSs. For the prediction of Luminal A and Luminal B, the DM plus DBT yielded better AUCs than DM or DBT alone. The DM plus DBT generated better AUCs and sensitivities than the DW plus DCE. For the prediction of HER2-enriched or TN, the DCE plus DW generated better AUCs than DCE or DW used alone (compare Table 3 with Table 2). The DW plus DCE outperformed DM plus DBT yielded in terms of AUCs, sensitivities and specificities.
Prediction performance of the most important features selected from mammography and MRI are listed in Tables S1, S2, respectively. A total of 5, 6, 4 and 5 features were selected from DM and DBT for predicting Luminal A, Luminal B, HER2-enriched and TN, respectively. While, a total of 4, 6, 6 and 7 features were selected from DW and DCE for predicting Luminal A, Luminal B, HER2-enriched and TN, respectively. Most of the features represent acceptable AUCs and P values.
Discussion
Our study performed comprehensive radiomics analyses to quantitatively evaluate predictive efficiencies of four commonly used breast cancer-screening modalities on molecular subtypes. Previous studies related to our work focused solely within the extent of the tumor itself (Fan et al. 2017; Sutton et al. 2016), ignoring information from areas surrounding the tumor. We found that the intra- and peritumoral regions can provide complementary information since combinations of the two regions efficiently improved prediction capabilities in terms of AUC, for predicting the four molecular subtypes. This was partially consistent with previous reports which also demonstrated that peritumoral areas of breast cancer may hold diagnostic and predictive information (Braman et al. 2019, 2017; Penn et al. 2020; Zhou et al. 2020).
Most of previous studies on the prediction of molecular subtypes in breast cancer focused on the MRI data (Fan et al. 2017; Sutton et al. 2016). However, the MRI is an expensive modality, and not suitable for all patients (Orel and Schnall 2001). While, the mammography is more widely used for breast screening and more accessible for patients. We performed direct and quantitative comparisons of the prediction effects between mammography and MRI, and found that for the prediction of Luminal A and B, the mammography (DM plus DBT) always outperformed MRI (DCE plus DW) in terms of AUC and sensitivity. This may be explainable since the X-ray was more sensitive to structural distortion and calcification, which were common in the Luminal A and B breast cancers (Cen et al. 2017; Zhang et al. 2019). The DBT showed higher AUCs for the prediction of HER2 compared with the DCE or DW alone, which may be because the HER2-enriched breast cancer also exist calcified lesions that can be detected by X-ray (Zhang et al. 2019). While, the DM plus DBT generated lower AUC, sensitivity and specificity than the DCE plus DW. This was partially in line with a previous study that also demonstrated great power of MRI on the prediction of HER2-enriched breast cancer because of the quantitative enhancement parameters of DCE can reflect the blood flow and vascular permeability, which had the correlation with HER2 breast cancers (Mazurowski et al. 2014). For the prediction of TN, the DM generated similar prediction performance compared with DBT. While, the DM plus DBT showed lower AUC, sensitivity and specificity than the DW plus DCE. This may be explainable considering the TN breast cancer are more likely to present as a non-calcified masses with a relatively circumscribed margin and this type breast cancers are rich in neovascularization, and negatively correlated with structural distortion (Son et al. 2020; Clauser et al. 2015). While, the DW can reflect tissue microenvironment and membrane integrity through depicting the diffusivity of the tissues meanwhile, the quantitative enhancement parameters of DCE can reflect the blood flow and vascular permeability, which had the correlation with TN breast cancers (Koo et al. 2012), and thus, compared with X-ray, MRI can better predict TN breast cancers. Prior study also pointed out the fact that the predominance of the presentation of the mass type on MRI is observed even more than with mammogram (Dogan et al. 2010), which was in line with the results of our research.
We identified the most important features for the prediction of each molecular subtype from the four modalities. For predicting Luminal A, the selected strength feature that was calculated from mammogram measures the primitives and quantifies the complicate textures of the tumor and unclear edges. Our results showed that values of the strength feature were higher in the Luminal A group than those in the non-Luminal A group, which indicated that the tumor with Luminal A may be associated with unclear margins (Boisserie-Lacroix et al. 2013). For predicting Luminal B, we identified the kurtosis feature from mammogram, which measures whether the data are heavy-tailed or light-tailed relative to a normal distribution. This feature has been considered as a biomarker reflecting tumor heterogeneity (Hempel et al. 2018). Our results showed that values of this feature were greater in the Luminal B group than those in the non-Luminal B group, which indicates that the Luminal B tumor may represent more heterogeneous tissue architectures. This was in line with a previous report that also found the Luminal B tumor was associated with architectural distortions (Taneja et al. 2008). For predicting HER2, the selected glszm_ZoneVariance feature was computed from MRI. This feature describes the roughness of the tumoral edge, with a higher value indicates the rougher edge. We found that the values of this feature were higher in the HER2-enriched group than the non-HER2-enriched group, which suggests that the margins of HER2-enriched tumor tends to be unclear. Our finding was accordant with a previous research that showed the HER2-enriched tumor represents indistinct margins (Taneja et al. 2008). For predicting TN, the selected Elongation and MinorAxisLength features were calculated from MRI. These features describe shapes of the ROIs, with higher values indicate rounder shapes. Our findings revealed that the values of these features were higher in the TN group than the non-TN group, which indicated that the TN tumor may tend to be rounder. The gldm_DependenceNonUniformityNormalized feature from MRI measures the similarity of the tumoral margin, with a lower value indicates a clearer margin. The value of this feature was lower in the TN group, which suggests the TN tumor may tend to be clearer margin. This was accordant with a recent effort that showed the TN tumor was associated with the round or oval mass and circumscribed margin (Kim and Choi 2013).
Limitations
There are certain limitations. First, this is a retrospective study with all samples enrolled from a single institution. We planned to enlarge the sample size and adopt a multicenter study design to overcome the limitation of the current study. Second, all radiomics features were extracted based on the manually drawn ROIs that may be subjective. To address this limitation, features with low ICCs were excluded to ensure the robustness of the features. While, our future work will employ deep learning-based automatic algorithms to improve the segmentation accuracy. Third, we did not evaluate other commonly used breast scanning technologies, such as ultrasonography or computed tomography (CT). We believe these would be warranted attentions and may be assessed in our future work.
Conclusion
Peritumoral regions can provide complementary information to intratumoral regions in breast mammography and MRI for the prediction of molecular subtypes. Compared with MRI, mammography performed better than MRI in the prediction of Luminal A and B, but worse than MRI in the prediction of HER2 or TN. Our findings may widen the understanding of intra- and peritumoral regions in different modalities and have the potential to predict the molecular subtypes.
Availability of data and materials
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
The code used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Au-Yong IT, Evans AJ, Taneja S et al (2009) Sonographic correlations with the new molecular classification of invasive breast cancer. Eur Radiol 19:2342–2348. https://doi.org/10.1007/s00330-009-1418-2
Boisserie-Lacroix M, Hurtevent-Labrot G, Ferron S et al (2013) Correlation between imaging and molecular classification of breast cancers. Diagn Interv Imaging 94:1069–1080. https://doi.org/10.1016/j.diii.2013.04.010
Braman NM, Etesami M, Prasanna P et al (2017) Intratumoral and peritumoral radiomics for the pretreatment prediction of pathological complete response to neoadjuvant chemotherapy based on breast DCE-MRI. Breast Cancer Res 19:57. https://doi.org/10.1186/s13058-017-0846-1
Braman N, Prasanna P, Whitney J et al (2019) Association of peritumoral radiomics with tumor biology and pathologic response to preoperative targeted therapy for HER2 (ERBB2)-positive breast cancer. JAMA Netw Open 2:e192561. https://doi.org/10.1001/jamanetworkopen.2019.2561
Cen D, Xu L, Li N et al (2017) BI-RADS 3–5 microcalcifications can preoperatively predict breast cancer HER2 and Luminal a molecular subtype. Oncotarget 8:13855–13862. https://doi.org/10.18632/oncotarget.14655
Clauser P, Carbonaro LA, Pancot M et al (2015) Additional findings at preoperative breast MRI: the value of second-look digital breast tomosynthesis. Eur Radiol 25:2830–2839. https://doi.org/10.1007/s00330-015-3720-5
Dogan BE, Gonzalez-Angulo AM, Gilcrease M et al (2010) Multimodality imaging of triple receptor-negative tumors with mammography, ultrasound, and MRI. AJR Am J Roentgenol 194:1160–1166. https://doi.org/10.2214/AJR.09.2355
Fan M, Li H, Wang S, Zheng B, Zhang J, Li L (2017) Radiomic analysis reveals DCE-MRI features for prediction of molecular subtypes of breast cancer. PLoS ONE 12:e0171683. https://doi.org/10.1371/journal.pone.0171683
Fehrenbacher L, Cecchini RS, Geyer CE Jr et al (2020) NSABP B-47/NRG oncology phase III randomized trial comparing adjuvant chemotherapy with or without trastuzumab in high-risk invasive breast cancer negative for HER2 by FISH and With IHC 1+ or 2. J Clin Oncol 38:444–453. https://doi.org/10.1200/JCO.19.01455
Fischer U, Baum F, Obenauer S et al (2002) Comparative study in patients with microcalcifications: full-field digital mammography vs screen-film mammography. Eur Radiol 12:2679–2683. https://doi.org/10.1007/s00330-002-1354-x
Goldhirsch A, Wood WC, Coates AS et al (2011) Strategies for subtypes–dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol 22:1736–1747. https://doi.org/10.1093/annonc/mdr304
Goldstein NS, Decker D, Severson D, Schell S, Vicini F, Margolis J, Dekhne NS (2007) Molecular classification system identifies invasive breast carcinoma patients who are most likely and those who are least likely to achieve a complete pathologic response after neoadjuvant chemotherapy. Cancer 110:1687–1696. https://doi.org/10.1002/cncr.22981
Hempel JM, Schittenhelm J, Bisdas S et al (2018) In vivo assessment of tumor heterogeneity in WHO 2016 glioma grades using diffusion kurtosis imaging: diagnostic performance and improvement of feasibility in routine clinical practice. J Neuroradiol 45:32–40. https://doi.org/10.1016/j.neurad.2017.07.005
Huber KE, Carey LA, Wazer DE (2009) Breast cancer molecular subtypes in patients with locally advanced disease: impact on prognosis, patterns of recurrence, and response to therapy. Semin Radiat Oncol 19:204–210. https://doi.org/10.1016/j.semradonc.2009.05.004
Kim MY, Choi N (2013) Mammographic and ultrasonographic features of triple-negative breast cancer: a comparison with other breast cancer subtypes. Acta Radiol 54:889–894. https://doi.org/10.1177/0284185113488580
Koboldt DC, Fulton RS, McLellan MD, Schmidt H, Kalicki-Veizer J, McMichael JF, Fulton LL, Dooling DJ, Ding L, Mardis ER (2012) Comprehensive molecular portraits of human breast tumours. Nature 490:61–70. https://doi.org/10.1038/nature11412
Koo HR, Cho N, Song IC et al (2012) Correlation of perfusion parameters on dynamic contrast-enhanced MRI with prognostic factors and subtypes of breast cancers. J Magn Reson Imaging 36:145–151. https://doi.org/10.1002/jmri.23635
Lam SW, Jimenez CR, Boven E (2014) Breast cancer classification by proteomic technologies: current state of knowledge. Cancer Treat Rev 40:129–138. https://doi.org/10.1016/j.ctrv.2013.06.006
Luck AA, Evans AJ, James JJ et al (2008) Breast carcinoma with basal phenotype: mammographic findings. AJR Am J Roentgenol 191:346–351. https://doi.org/10.2214/AJR.07.2659
Ma W, Zhao Y, Ji Y et al (2019) Breast cancer molecular subtype prediction by mammographic radiomic features. Acad Radiol 26:196–201. https://doi.org/10.1016/j.acra.2018.01.023
Mazurowski MA, Zhang J, Grimm LJ et al (2014) Radiogenomic analysis of breast cancer: luminal B molecular subtype is associates with enhancement dynamics at MR imaging. Radiology 273:365–372. https://doi.org/10.1148/radiol.14132641
Monti S, Aiello M, Incoronato M et al (2018) DCE-MRI pharmacokinetic-based phenotyping of invasive ductal carcinoma: a radiomic study for prediction of histological outcomes. Contrast Media Mol Imaging 2018:5076269. https://doi.org/10.1155/2018/5076269
Orel SG, Schnall MD (2001) MR imaging of the breast for the detection, diagnosis, and staging of breast cancer. Radiology 220:13–30. https://doi.org/10.1148/radiology.220.1.r01jl3113
Orlando L, Viale G, Bria E et al (2016) Discordance in pathology report after central pathology review: Implications for breast cancer adjuvant treatment. Breast 30:151–155. https://doi.org/10.1016/j.breast.2016.09.015
Penn AI, Medved M, Dialani V, Pisano ED, Cole EB, Brousseau D, Karczmar GS, Gao G, Reich BD, Abe H (2020) Discrimination of benign from malignant breast lesions in dense breasts with model-based analysis of regions-of-interest using directional diffusion-weighted images. BMC Med Imaging 20:61. https://doi.org/10.1186/s12880-020-00458-3
Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lønning PE, Børresen-Dale AL, Brown PO, Botstein D (2000) Molecular portraits of human breast tumours. Nature 406:747–752. https://doi.org/10.1038/35021093
Rouzier R, Perou CM, Symmans WF et al (2005) Breast cancer molecular subtypes respond differently to preoperative chemotherapy. Clin Cancer Res 11:5678–5685. https://doi.org/10.1158/1078-0432.CCR-04-2421
Ruopp MD, Perkins NJ, Whitcomb BW et al (2008) Youden index and optimal cut-point estimated from observations affected by a lower limit of detection. Biom J 50:419–430. https://doi.org/10.1002/bimj.200710415
Sauerbrei W, Royston P, Binder H (2007) Selection of important variables and determination of functional form for continuous predictors in multivariable model building. Stat Med 26:5512–5528. https://doi.org/10.1002/sim.3148
Son J, Lee SE, Kim EK et al (2020) Prediction of breast cancer molecular subtypes using radiomics signatures of synthetic mammography from digital breast tomosynthesis. Sci Rep 10:21566. https://doi.org/10.1038/s41598-020-78681-9
Sutton EJ, Dashevsky BZ, Oh JH et al (2016) Breast cancer molecular subtype classifier that incorporates MRI features. J Magn Reson Imaging 44:122–129. https://doi.org/10.1002/jmri.25119
Taneja S, Evans AJ, Rakha EA et al (2008) The mammographic correlations of a new immunohistochemical classification of invasive breast cancer. Clin Radiol 63:1228–1235. https://doi.org/10.1016/j.crad.2008.06.006
Uematsu T, Kasami M, Yuen S (2009) Triple-negative breast cancer: correlation between MR imaging and pathologic findings. Radiology 250:638–647. https://doi.org/10.1148/radiol.2503081054
Waks AG, Winer EP (2019) Breast cancer treatment: a review. JAMA 321:288–300
Wang Y, Yin Q, Yu Q et al (2011) A retrospective study of breast cancer subtypes: the risk of relapse and the relations with treatments. Breast Cancer Res Treat 130:489–498. https://doi.org/10.1007/s10549-011-1709-6
Ye DM, Wang HT, Yu T (2020) The application of radiomics in breast MRI: a review. Technol Cancer Res Treat 19:1533033820916191. https://doi.org/10.1177/1533033820916191
Zardavas D, Irrthum A, Swanton C et al (2015) Clinical management of breast cancer heterogeneity. Nat Rev Clin Oncol 12:381–394. https://doi.org/10.1038/nrclinonc.2015.73
Zhang L, Huang Y, Feng Z, Wang X, Li H, Song F, Liu L, Li J, Zheng H, Wang P, Song F, Chen K (2019) Comparison of breast cancer risk factors among molecular subtypes: a case-only study. Cancer Med 8:1882–1892. https://doi.org/10.1002/cam4.2012
Zhou J, Zhang Y, Chang KT, Lee KE, Wang O, Li J, Lin Y, Pan Z, Chang P, Chow D, Wang M, Su MY (2020) Diagnosis of benign and malignant breast lesions on DCE-MRI by using radiomics and deep learning with consideration of peritumor tissue. J Magn Reson Imaging 51:798–809. https://doi.org/10.1002/jmri.26981
Acknowledgements
The study was funded by Climbing Fund of National Cancer Center (NCC201806B011), National Natural Science Foundation of China (81872363), Major Technology Plan Project of Shenyang (17-230-9-07), Special foundation for the central government guides the development of local science and technology of Liaoning Province (2018416029), Education Department Foundation of Liaoning (LQNK201744), Key Program of Ministry of Science and Technology of China (17YFC1309100) China National Natural Science Foundation (31770147) and Medical-Engineering Joint Fund for Cancer Hospital of China Medical University and Dalian University of technology (LD202029), Wu Jieping Medical Foundation (20.6750.2020-08-22)
Author information
Authors and Affiliations
Contributions
SN and WJ contributed to the conception and design of the study. NZ and TY contributed to the acquisition of the data. SN and WJ contributed to the data analysis and interpretation. Tao Jiang and Yue Dong contributed to statistical analysis of this study. XJ and SN participated in manuscript preparation and manuscript editing of this study. XJ and YL participated in the review of manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have declared that no conflict of interest exists.
Ethics approval
The Ethics Committee of the Cancer Hospital of China Medical University, Liaoning Cancer Hospital and Institute (Shenyang, China) approved the use of patient materials in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Niu, S., Jiang, W., Zhao, N. et al. Intra- and peritumoral radiomics on assessment of breast cancer molecular subtypes based on mammography and MRI. J Cancer Res Clin Oncol 148, 97–106 (2022). https://doi.org/10.1007/s00432-021-03822-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-021-03822-0