Abstract
Polychaetes are famous for their outstanding ability to regenerate lost body parts. Moreover, these worms possess a number of ancestral features in anatomy, development, and genetics, making them particularly suitable for comparative studies. Thus, fundamental as well as new undisclosed so far features of regenerative processes may be revealed, using polychaetes as a model. In the present work, we aimed to analyze the molecular basis of caudal regeneration in the nereid polychaete Alitta virens (formerly Nereis virens). We focused on homologues genes of RNA helicases Vasa and PL10 and ncRNA-binding proteins Piwi. These markers are suggested to play a significant role in maintenance of undifferentiated state of primordial germ cells and multipotent stem cells across invertebrates. In normal conditions, A. virens homologues of Vasa, PL10, and Piwi were differentially expressed in the subterminal growth zone and germline cells. Caudal amputation induced expression of studied genes de novo, which further accompanies all steps of regeneration. An early appearance of the transcripts in wound epithelium and internal blastemal cells suggests involvement of these genes in the well-known cell dedifferentiation events that assure polychaete regeneration. Provided interpretation of the gene expression dynamics implies the primary restoration of the pygidium and growth zone, which promotes following segment formation. Obtained results are valuable as a molecular fingerprint of the alterations occurring in regulatory state of locally regenerating tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polychaete annelids are well-known to possess widely distributed regeneration ability after injury and during asexual reproduction. Unlike classical invertebrate models of regenerative biology (hydra and planarians), polychaete worms display a compound segmented body plan similar to hypothesized Urbilateria (Balavoine and Adoutte 2003; De Robertis 2008). Recently, an ancestral-type genome organization has been shown for the nereid polychaete Platynereis dumerilii (Raible et al. 2005), rising this species as a new valuable model organism for evolutionary and developmental biology studies. Assuming an ancestral state of both the developmental programs and the body molecular architecture in nereid polychaetes (Kulakova et al. 2007; Denes et al. 2007; Christodoulou et al. 2010; Simakov et al. 2013), these animals may be particularly useful for disclosure of ancient and fundamental features of regenerative processes.
Nereids and most of other errant (free-moving) polychaetes, regarded as more primitive forms, are incapable of the head regeneration, but can easily regrow lost tail region. Unexpectedly, the more derived sedentary (tube-dwelling) polychaetes as well as oligochaetes in addition to caudal regeneration possess a great ability of anterior regeneration, whereas leeches lack epimorphic regeneration at all (Hyman 1940). Owing to well-resolved phylogenetic relationships among annelids (Struck et al. 2011), a comparative analysis of molecular and cellular morphogenetic mechanisms can elucidate evolutionary trends of the regenerative ability in the whole phylum. In that way, it would benefit our understanding of the causality of long-distance interspecies differences. In turn, this knowledge is a key prerequisite of the progress in regenerative biomedicine.
Nereid worms (Nereis, Perinereis, Platynereis) have long been studied as models for regeneration, so comprehensive data describing specific physiological and cellular features of the process are available. Thus, more in-depth, i.e., molecular, analysis can be performed. Classical microsurgical studies on annelids have first indicated an exceptional importance of the nervous system in regeneration (Hyman 1940; Herlant-Meewis 1964). Damaged ventral nerve cord is believed to induce and promote regenerative outgrowth in the stump or even ectopically. Nereids also display a fascinating and still enigmatic hormonal control of mutually exclusive normal/regenerative growth versus sexual maturation (Herlant-Meewis 1964; Golding 1967).
From an anatomical perspective, after amputation of posterior segments, caudal regeneration in nereids starts by the constriction of the body wall musculature that brings together epidermis and intestinal epithelium in the wound surface. During the wound healing, these two epithelial layers join each other circumferentially, so that the new anus is reconstituted at the first few days post amputation (dpa) (Herlant-Meewis and Nokin 1962; Hofmann 1966; Boilly 1969; Combaz and Boilly 1974). This morphogenesis employs local tissue remodeling (histolysis, phagocytosis, wound plug formation, epithelialization) and externally results in construction of the so-called wound (cicatricial) epithelium (Herlant-Meewis 1964; Bely 2014). Typically, wounding (cicatrization) in annelids is free of cell divisions in the stump (Hill 1970; Bely 2014).
The next step of regeneration is accumulation of the internal cell mass referred to as the blastema. It consists of mesenchymal elements of diverse morphology including highly basophilic undifferentiated cells. The wound epithelium is also reported to have activated appearance (enlarged basophilic cells with hypertrophic nuclei) that indicates undifferentiated state of the cells (Herlant-Meewis 1964; Hill 1970; Korotkova 1997). Proliferation of epithelial and blastemal cells promotes further growth of the regenerative bud. Finally, the formation of a new pygidium, growth zone, and segment primordia manifests the differentiation of the regenerative bud.
To explain the cellular mechanisms of annelid regeneration, a great attention was given to determine the cellular origin of the blastema. The first reports on this topic in oligochaetes described totipotent stem cells (called neoblasts) as the main source of blastema (Randolph 1892). Injury activates neoblasts, which become larger, begin to proliferate, and migrate over segments towards the wound. However, some oligochaete species apparently lack population of neoblasts, but all examined oligochaetes do employ cellular dedifferentiation in regenerative process (Herlant-Meewis 1964; Yoshida-Noro and Tochinai 2010; Myohara 2012).
Descriptive histological and EM studies as well as experiments utilizing autoradiography, X-ray irradiation, and transplantations undoubtedly indicated that polychaetes have no neoblast-like totipotent stem cells (Potswald 1969; Hill 1970; Korotkova 1997). Indeed, polychaete regeneration involves numerous cell dedifferentiation events in the stump along with retention of the germ layer identity in new tissues (Boilly 1968, 1969; Marilley and Thouveny 1978; Fontés et al. 1983; Paulus and Müller 2006; Bely 2014). Diverse mesodermal derivatives (muscles, cells of the coelomic (peritoneal) lining, and free coelomocytes) of the last old segment were found to dedifferentiate and accumulate under the wound epithelium, forming blastema. These blastemal cells and dedifferentiated wound epithelium begin to proliferate in the regenerative bud and redifferentiate afterwards according to their germ layer origin. The molecular basis underlying regeneration in polychaetes is still completely undiscovered and should be revealed.
The so-called germline genes encoding DEAD box RNA helicase Vasa and ncRNA-binding protein Piwi are the most famous markers of undifferentiated cells in invertebrates to date (Gustafson and Wessel 2010; Juliano et al. 2011). Moreover, these genes are believed to have a broad role in establishing and maintaining of multipotency (Juliano et al. 2010; Alié et al. 2011). Germline, progenitor, and stem cells employ a conserved set of genes referred to as the germline multipotency program (GMP) (Juliano and Wessel 2010; Solana 2013). To test probable involvement of the GMP in polychaete caudal regeneration, we have isolated and characterized gene expression patterns of two RNA helicases (Vasa and PL10) and two paralogues of Piwi in the nereid worm Alitta virens (formerly Nereis virens).
Material and methods
Animals
Mature A. virens worms were collected in natural habitat at the White Sea nearby the Marine Biological Station of St. Petersburg State University. An artificial fertilization and embryos’ culturing were performed as described earlier (Dondua 1975). Juvenile 15-segmented individuals were relaxed in 7.5 % MgCl2 mixed with an artificial sea water 1:1, following amputation of posterior third of the body carried out under dissecting binocular microscope. Regenerating worms were kept in the small Petri dishes in artificial sea water at 18 °C.
Gene cloning and phylogenetic analysis
Total RNA was isolated from regenerating A. virens of various ages using TRI Reagent (Sigma) according to the manufacturer’s instructions. Mixed RNA from different stages was subjected for the reverse transcription using RevertAid First Strand cDNA Synthesis Kits (Thermo Scientific) and SMARTer RACE cDNA Amplification Kit (Clontech Laboratories). Degenerate, 5′, and 3′ rapid amplification of cDNA ends (RACE) polymerase chain reaction (PCR) resulted in amplicons, which were cloned in the pCRII-TOPO vector (Invitrogen) and sequenced. Primer sequences and PCR conditions are available upon request. NCBI database accession numbers for Avi-vasa, Avi-pl10, Avi-piwi1, and Avi-piwi2 are KM406469–KM406472, correspondingly.
Newly cloned A. virens nucleotide sequences of candidate genes were translated into proteins, on which we mapped conserved domains using the Pfam database (http://pfam.xfam.org/). Genes’ orthologies were assigned to the sequences based on BLASTX searches in the GenBank database of the NCBI. Multiple alignments followed by maximum likelihood analyses were performed with algorithms MUSCLE 3.7 and PhyML 3.0 coupled with approximate likelihood-ratio test (aLRT) using the Phylogeny.fr online tools (www.phylogeny.fr).
In situ hybridization and imaging
Intact and amputated A. virens were fixed in 4 % formaldehyde on 1.75× PBS overnight at 4 °C. At least 20 individuals of every stage were analyzed in whole mount in situ hybridization for each gene. All treatments of the single-color in situ hybridization using NBT/BCIP staining were performed as described earlier (Tessmar-Raible et al. 2005). Dig-labeled RNA probes were prepared according to the manufacturer’s protocol (Roche). The results were imaged under DIC optics on an Axio Imager D1 microscope (Carl Zeiss) equipped with AxioCam ICc5 and MRm (Carl Zeiss) digital cameras. Artwork was done using Adobe Photoshop, ImageJ, and AxioVision 4.8 software.
5-Bromo-2′-deoxyuridine incorporation assay
Alive 2 dpa worms were incubated for 4 h in pasteurized sea water containing 0.1 % of 5-bromo-2′-deoxyuridine (BrdU). BrdU can be incorporated into the newly synthesized DNA during the S phase of the cell cycle. As cells pulsed with BrdU may be photosensitive, the incubation was performed in darkness. Afterwards, the animals were briefly washed in several changes of fresh sea water and fixed in Bouin solution (picric acid, saturated; formaldehyde, 37–40 %; glacial acetic acid; 15:5:1) for 2 h. Specimens were washed in 70 % ethanol and stored at −20 °C for a short period. Following BrdU detection steps were performed as described earlier (de Rosa et al. 2005).
Results
Using degenerate, 5′, and 3′ RACE PCR, we cloned and reconstructed the full CDS containing fragments of the four A. virens genes named Avi-vasa (2.7 kb), Avi-pl10 (2.4 kb), Avi-piwi1 (3.6 kb), and Avi-piwi2 (3.6 kb). Sequence analysis revealed typical domains for each of the gene families and apparent phylogenetic relations of these genes (Figs. 1 and 2). Considering branch lengths and topography of the phylogenetic trees, one can see that, A. virens sequences show relatively low divergence speed and constantly cluster with other lophotrochozoan orthologues.
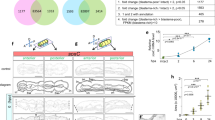
Sequence analysis of Avi-vasa and Avi-pl10 genes. a Domain structure of the inferred protein products. Conserved amino acid motifs are depicted: zf Zn-fingers, DEAD DEAD box helicase domain, Hel C helicase C-terminal domain. b Maximum-likelihood phylogenetic tree. DEAD box helicases form distinct DDX4/Vasa, DDX3/PL10, and p68 subgroups. A. virens sequences cluster with their lophotrochozoan orthologues. Node labels on a branch show approximate likelihood-ratio test (aLRT) results. NCBI accession numbers are indicated
Sequence analysis of Avi-piwi1 and Avi-piwi2 genes. a Domain structure of the inferred protein products. Conserved amino acid motifs are depicted: PAZ domain and Piwi domain. b Maximum-likelihood phylogenetic tree. The two A. virens Piwi paralogues fall apart within Piwi-1-like and Piwi-2-like subgroups. Node labels on a branch show aLRT results. NCBI accession numbers are indicated
Avi-vasa and Avi-pl10 are predicted to encode proteins of 808 and 769 amino acid residues, correspondingly. The sequences share two conserved domains: the DEAD/DEAH box helicase and the helicase C-terminal domain, which are responsible for ATP binding, nucleic acid binding, and unwinding of nucleic acid duplexes. In addition, six CCHC-type zinc finger motifs are detected at the N-terminus of Avi-vasa (Fig. 1a). Phylogenetic tree confirms Avi-vasa and Avi-pl10 as corresponding representatives of DDX4/Vasa and DDX3/PL10 subfamilies (Fig. 1b).
Two assembled A. virens Piwi homologues were translated into proteins of 869 and 951 amino acid residues. Both sequences contain characteristic PAZ and Piwi domains (Fig. 2a). Maximum likelihood analysis revealed two clear clades separating metazoan Piwi-1-like from Piwi-2-like genes (Fig. 2b). Thus, two A. virens paralogues were named Avi-piwi1 and Avi-piwi2. A single known P. dumerilii Piwi gene clustered with Avi-piwi1.
Next, we have addressed the question what regions of postlarval A. virens body are distinguished by molecular signs of multipotency and stemness. Differential expression of PL10, Vasa, and Piwi homologues were found in a posterior region of the 10–15-segmented juvenile worms (Fig. 3). These genes had evident expression in the subterminal growth zone (GZ). Avi-pl10 was expressed more broadly both in the posteriormost nascent segments and the GZ (Fig. 3a–c). The latter domain includes superficial (ectodermal) and internal (mesodermal) cells arranged into a narrow ring adjacent to the pygidium. Relatively higher expression level in the mesodermal part of the GZ was detected for Avi-vasa and Avi-piwi1 (Fig. 3d–g). These two genes also marked germline cells in the primary gonad and gonial clusters in parapodia (Fig. 4) similar to described localization of Vasa-positive cells in the polychaete P. dumerilii (Rebscher et al. 2007). Expression of Avi-piwi2 was detected only in the mesodermal part of the GZ region (Fig. 3h). Immediately after posterior amputation, there is no any PL10-, Vasa-, or Piwi-positive cells at the wound (Fig. 3i–l).
Expression patterns in the growing intact juvenile A. virens (a–h) and just after amputation (i–l): a–c l Avi-pl10, b is enlargement of the bordered square in (a); d–f, k Avi-piwi1, e is enlargement of the bordered square in (d); g, j Avi-vasa; h–i Avi-piwi2. Transcriptional expression is indicated in subterminal ectodermal (black arrows) and mesodermal (black arrowheads) cells comprising the GZ, in segmental mesoderm at the worm posterior end (white arrows). Dotted line marks an amputation plane. Dark color of the spinning glands (asterisks) and gut content (double asterisks) is a non-specific staining in some specimens. a, b, d, e, i–l are ventral views; с and f are dorsal views; g and h are deep focal planes from dorsal orientation. Anterior is to the left
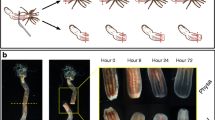
During caudal regeneration of the amputated A. virens juveniles in the used experimental conditions, wound healing is completed by 1 dpa. Wound epithelium combines with the intestinal one, but no outgrowth can be observed in the stump. The next 1–2 days, epimorphic blastema grows at the site of wound. Subsequent differentiation of the regenerative bud shows up for the first time by 3 dpa, when pygidial cirrus anlagen become apparent. Following regenerative bud segmentation occurs the next days (4–5 dpa). By 6–7 dpa, new segment primordia start to appear sequentially, indicating resumption of the normal posterior growth.
Among all examined stages, we found no scattered expressing cells in the whole body (which otherwise could be interpreted as migrating stem cells). At the time point 1 dpa, all four studied genes demonstrate nearly identical expression pattern: it appears de novo at the site of injury. We observed expression in the wound epithelium and in the internal cells underneath (Fig. 5a, f, k, p). Proposed dedifferentiated internal cells are situated on the ventral side of the stump on the level of longitudinal muscles or slightly deeper. In particular, the expression of Avi-pl10 is more extensive around the wound (Fig. 5a). Avi-vasa- and Avi-piwi2-positive cells show up as two lateral patches on each side of the anus (Fig. 5f, p). Avi-piwi1 is expressed in the wound epithelium wider and stronger than in the deep mesodermal cells (Fig. 5k). A day later, Avi-pl10, Avi-vasa, and Avi-piwi1 transcripts are broadly distributed across the whole regenerative bud (Fig. 5b, g, l), while Avi-piwi2 signal is found exclusively in the blastema comprised of accumulating internal cells (Fig. 5q). In the case of each gene, the internal expressing cells at this stage form a bilateral protrusion covered by the wound epithelium. At the 3 dpa stage, the expression of Avi-pl10, Avi-vasa, and Avi-piwi1 disappears from the most terminal region of the regenerative bud corresponding to pygidium anlage (Fig. 5c, h, m). Ectodermal cells adjacent to this border express Avi-vasa and Avi-piwi1 on relatively higher level than the rest regenerated epidermis does (Fig. 5h, m). The mesodermal expression domains of all four genes have a dumbbell-like shape on the ventral side. The dumbbell-shaped region comprises restricted to three–four cells in width middle part and wider lateral parts. At 4–5 dpa, the mesodermal domain extends in lateral parts anteriorly, thereby shifting the ventral middle region towards subterminal position (Fig. 5d, i, s). At the similar subterminal position, Avi-pl10, Avi-vasa, and Avi-piwi1 are prominently expressed in the ectodermal epithelium on the dorsal and lateral sides, forming a narrow one- to two-cell-wide stripe (Fig. 6). Later on, segment primordia become apparent in the regenerative bud and the expression of Avi-vasa and Avi-piwi2 gradually fades in the nascent segments (Fig. 5j, t). At the same time (6 dpa), Avi-pl10 and Avi-piwi1 demonstrate rather vast expression despite inferred differentiation that have begun in the anteriormost segmented part of the regenerative bud (Fig. 5e, o). These differences between gene expression patterns resemble the situation observed in unamputated growing worms.
Expression patterns in the regenerating A. virens: a–e Avi-pl10; f–j Avi-vasa; k–o Avi-piwi1; p–t Avi-piwi2. Stages of 1, 2, 3, 4–5, and 6 dpa are shown sequentially in each row. Amputation site (dotted line); wound epithelium (double arrowheads); mesodermal blastemal cells (white arrows); and subterminal ectodermal (black arrows) and mesodermal (black arrowheads) cells comprising the GZ are indicated. Ventral views, anterior is to the left
To address the role of cell divisions during regenerative growth, we performed BrdU (an analog of thymidine) incorporation experiment. Cells in the S phase are abundant throughout regenerative bud at about 2.5 dpa (Fig. 7). BrdU-labeled nuclei are randomly scattered across the epidermis (Fig. 7a). A different intensity of the signal in neighboring nuclei indicates asynchronous proliferation of the ectodermal cells. The strongest labeling throughout the ectodermal layer was observed in primordia of the pygidial cirri (Fig. 7b). Internal BrdU-incorporated nuclei in the blastema are restricted to ventrolateral positions (Fig. 7b). Thus, labeled mesodermal cells underlay actively proliferating ectodermal ones.
Cell division pattern revealed by BrdU labeling. a Superficial focal plane; b deep focal plane. BrdU-positive nuclei of ectodermal cells (black arrows) and mesodermal blastemal cells (white arrows) in 2.5 dpa regenerative bud are indicated. Asterisks mark primordia of the pygidial cirri. Ventral views, anterior is to the left
Discussion
In the present study, we have attempted to analyze a molecular profile of regenerating and normally growing tail of the polychaete A. virens. The used here so-called germline genes were expressed in the GZ of juvenile worms, corresponding to the results on other polychaete and oligochaete species (Rebscher et al. 2007; Dill and Seaver 2008; Oyama et al. 2008; Sugio et al. 2008; Giani et al. 2011; Kostyuchenko et al. 2012; Gazave et al. 2013). Thus, PL10, Vasa, and Piwi homologues can be considered as conserved markers of the annelid GZ. We suppose this particular localization reflects an undifferentiated state of the GZ cells. At the same time, subtle differences in the observed expression patterns indicate a complexity of this region.
After tail amputation, an expression of Avi-pl10, Avi-vasa, Avi-piwi1, and Avi-piwi2 accompanies all steps of caudal regeneration. Signal of all tested genes emerges de novo in the both wound epithelium and mesodermal blastemal cells. It corresponds well with the known cellular dedifferentiation events initiated at the beginning of polychaete regeneration (see “Introduction”). Thus, the proposed dedifferentiating ectodermal and mesodermal cells seem to utilize the GMP components since the initial step of regeneration (0–1 dpa). No signs of scattered or migrating stem-like cells (i. e., expressing the GMP genes) were found across all segments of A. virens body. It further corroborates an absence of multipotent stem cells in polychaetes. A recent proliferation survey on regenerating polychaete Dorvillea bermudensis inferred similar conclusions (Paulus and Müller 2006). It is opposite to results obtained on the oligochaete Enchytraeus japonensis, which possesses a system of Vasa-positive stem cells (neoblasts). In E. japonensis, neoblasts apparently contribute to regeneration blastema and are distinct from the germline and the GZ cells for their molecular fingerprint and proliferative activity (Yoshida-Noro and Tochinai 2010). The neoblast lineage also appears much later in the development, suggesting an idea of its evolutionary origin as oligochaete-specific deviation. Altogether, it implies an essential variability of the cellular origin of regeneration blastema across annelids.
During the A. virens blastema growth and development (1–3 dpa), expression patterns demonstrate subtle distinctions (more general distribution of Avi-pl10 transcript, restricted mesodermal signal of Avi-piwi2), indicating diversification of these four genes’ specificity as well as compound molecular architecture of the regenerative bud. According to BrdU assay results, the regenerative bud increases in size at this time on account of extensive cellular proliferation. In regenerated epidermis, the area of active BrdU incorporation is rather wider than expression domains, whereas only a ventrolateral subset of expressing cells in blastema is proliferative. Hence, the function of the GMP genes here is not restricted to promoting cell divisions, while the latter can occur (e.g., in primordia of the pygidial cirri) in cells lacking the expression of tested genes.
Since differentiation of regenerative bud shows up (3 dpa), a strong expression of Avi-vasa and Avi-piwi1 delineates the newly formed subterminal GZ. At 4–6 dpa, the corresponding subterminal region of expression becomes more apparent, comprising dorsal ectodermal one- to two-cell-wide arc and ventral mesodermal three- to four-cell-wide arc. The former part of the GZ on late stages of regeneration of nereids P. dumerilii and Perinereis nuntia has the same outward and a synchronized long-term cell cycle (de Rosa et al. 2005; Gazave et al. 2013; Niwa et al. 2013). The rest regenerative bud epidermis excluding the pygidium proliferates actively till the full segment size has been achieved (de Rosa et al. 2005).
Therefore, we speculate that the initial cell dedifferentiation and following proliferation aim to restore the pygidium and the GZ, utilizing a molecular state of the latter. Accordingly, further regenerative bud growth (3 dpa onward) is based on the activity of the GZ, which gives rise to new segments and shifts posteriorly meanwhile.
A few works were published on the molecular basis of regeneration in polychaetes to date. The results on Piwi gene expression in regenerating Capitella teleta (Giani et al. 2011) confirm the use of GMP in somatic undifferentiated cells across polychaetes. However, the expression was not detected prior to obvious blastema outgrowth at 3 dpa. Vasa homologue expression but not Piwi was shown for GZ in the clitellate worm E. japonensis (Sugio et al. 2008). In contrast, in another oligochaete Tubifex tubifex Vasa is not expressed in the GZ (Oyama and Shimizu 2007). Probably its function has been assigned to p68, another RNA helicase of the DEAD box family (Oyama et al. 2008). In our own studies, we have found both Vasa and Piwi homologues to be expressed posteriorly during caudal regeneration and normal growth in naidids Pristina longiseta and Nais communis (Kostyuchenko et al. 2012). Thus, although some important cellular events of regeneration in clitellate and non-clitellate annelids might be different, restoration of the GZ and de novo segmentation occurs by involving GMP in these processes. Moreover, Piwi homologues are required for regeneration across as diverse models as flatworm, ascidian, and amphibian species (Reddien et al. 2005; Rinkevich et al. 2010; Zhu et al. 2012). In the latter case, the expression of two Piwi paralogues was activated upon injury in dedifferentiating ectodermal and mesodermal cells (Zhu et al. 2012). Depletion of these genes by morpholino oligonucleotides interfered with forelimb regeneration by decreasing cell proliferation and increasing cell death in the blastema. On the contrary, planarian regeneration is based on a population of pluripotent stem cells, whose self-renewal and differentiation are assured by Piwi and Vasa genes (Reddien et al. 2005; Gustafson and Wessel 2010; Rink 2013). Considering the profound difference in cellular mechanisms of tissue restoration in animals, it is extremely curious how GMP have evolved. Whether an ancestral state was a reprogramming of the cell fate upon inductive cues or specification of distinct lineage of germline/stem cells, the answer might lie at the root of the metazoan tree, in studies of the very first and oldest multicellular organisms, such as sponges, which possess both unique totipotent stem cell populations (Funayama 2013) and highly conserved developmental processes (Korotkova 1997; Ereskovsky et al. 2013).
The data regarding Hox genes’ profiles during regeneration of the nereid worms P. dumerilii and A. virens (Pfeifer et al. 2012; Novikova et al. 2013) strengthen our hypothesis of the primary GZ restoration. Since the expression of the GZ-specific markers Hox-2 (a mesodermal one) and Hox-3 (an ectodermal one) is observed quite early (1 dpa onward) and their following domains refine position of the GZ within the regenerative bud, one may propose the first labeled (dedifferentiated) cells to be determined for the GZ fate. Even further, there are a substantial data proving a sequential homeogenetic induction during the posterior segmentation of the nereid polychaete P. nuntia (Niwa et al. 2013). In P. nuntia, superficial ectodermal cells at the segment/pygidium boundary are periodically induced to synchronously enter the cell cycle and remodel the chromatin state. In a similar way, de Rosa et al. (2005) interpreted the GZ in P. dumerilii. Hence, we favor dedifferentiation-based explanation of the polychaete GZ in normal condition and after amputation rather the existence of posterior teloblast-like stem cells.
In future studies, an attempt to reconstruct the whole gene regulatory network of the polychaete regeneration should be given, emphasizing on the developmentally relevant transcription factors and signaling cascades. Taking into account recent elaboration of new functional techniques for P. dumerilii (Backfisch et al. 2014; Bannister et al. 2014; Zantke et al. 2014), dissecting the exact role of GMP genes in polychaetes would be of great significance. The presented data can serve as a reference of cellular dedifferentiation and GZ restoration during caudal regeneration in nereids.
References
Alié A, Leclère L, Jager M et al (2011) Somatic stem cells express Piwi and Vasa genes in an adult ctenophore: ancient association of “germline genes” with stemness. Dev Biol 350:183–197. doi:10.1016/j.ydbio.2010.10.019
Backfisch B, Kozin VV, Kirchmaier S et al (2014) Tools for gene-regulatory analyses in the marine annelid Platynereis dumerilii. PLoS ONE 9:e93076. doi:10.1371/journal.pone.0093076
Balavoine G, Adoutte A (2003) The segmented urbilateria: a testable scenario. Integr Comp Biol 43:137–147. doi:10.1093/icb/43.1.137
Bannister S, Antonova O, Polo A et al (2014) TALENs mediate efficient and heritable mutation of endogenous genes in the marine annelid Platynereis dumerilii. Genetics 197:77–89. doi:10.1534/genetics.113.161091
Bely AE (2014) Early events in annelid regeneration: a cellular perspective. Integr Comp Biol icu109. doi:10.1093/icb/icu109
Boilly B (1968) Origine des cellules de régénération chez Aricia foetida Clap. (Annélide Polychète). Arch d’Anat Microscop Morphol Experiment 57:297–308
Boilly B (1969) Origine des cellules régénératrices chez Nereis diversicolor O. F. Müller (Annélide Polychète). Roux Arch Entwm 162:286–305. doi:10.1007/BF00576934
Christodoulou F, Raible F, Tomer R et al (2010) Ancient animal microRNAs and the evolution of tissue identity. Nature 463:1084–1088. doi:10.1038/nature08744
Combaz A, Boilly B (1974) Étude expérimentale et histologique de la régénération caudale en l’absence de chaine nerveuse chez les Nereidae (Annélides Polychètes). Ann Embryol Morphog 7:171–197
De Robertis EM (2008) Evo-devo: variations on ancestral themes. Cell 132:185–195. doi:10.1016/j.cell.2008.01.003
De Rosa R, Prud’homme B, Balavoine G (2005) Caudal and even-skipped in the annelid Platynereis dumerilii and the ancestry of posterior growth. Evol Dev 7:574–587. doi:10.1111/j.1525-142X.2005.05061.x
Denes AS, Jékely G, Steinmetz PRH et al (2007) Molecular architecture of annelid nerve cord supports common origin of nervous system centralization in bilateria. Cell 129:277–288. doi:10.1016/j.cell.2007.02.040
Dill KK, Seaver EC (2008) Vasa and nanos are coexpressed in somatic and germ line tissue from early embryonic cleavage stages through adulthood in the polychaete Capitella sp. I. Dev Genes Evol 218:453–463. doi:10.1007/s00427-008-0236-x
Dondua AK (1975) Effect of actinomycin D and sibiromycin on the embryonic and larval development of Nereis virens (Sars.). Ontogenez 6:475–484
Ereskovsky AV, Renard E, Borchiellini C (2013) Cellular and molecular processes leading to embryo formation in sponges: evidences for high conservation of processes throughout animal evolution. Dev Genes Evol 223:5–22. doi:10.1007/s00427-012-0399-3
Fontés M, Coulon J, Delgross M-H, Thouveny Y (1983) Muscle dedifferentiation and contractile protein synthesis during post-traumatic regeneration by Owenia fusiformis (polychaete annelid). Cell Different 13:267–282. doi:10.1016/0045-6039(83)90037-4
Funayama N (2013) The stem cell system in demosponges: suggested involvement of two types of cells: archeocytes (active stem cells) and choanocytes (food-entrapping flagellated cells). Dev Genes Evol 223:23–38. doi:10.1007/s00427-012-0417-5
Gazave E, Béhague J, Laplane L et al (2013) Posterior elongation in the annelid Platynereis dumerilii involves stem cells molecularly related to primordial germ cells. Dev Biol 382:246–267. doi:10.1016/j.ydbio.2013.07.013
Giani VC, Yamaguchi E, Boyle MJ, Seaver EC (2011) Somatic and germline expression of piwi during development and regeneration in the marine polychaete annelid Capitella teleta. EvoDevo 2:10. doi:10.1186/2041-9139-2-10
Golding DW (1967) Endocrinology, regeneration and maturation in Nereis. Biol Bull 133:567–577
Gustafson EA, Wessel GM (2010) Vasa genes: emerging roles in the germ line and in multipotent cells. Bioessays 32:626–637. doi:10.1002/bies.201000001
Herlant-Meewis H (1964) Regeneration in annelids. Adv Morphog 4:155–215
Herlant-Meewis H, Nokin A (1962) Cicatrisation et premiers stades de régénération pygidiale chez Nereis diversicolor. Ann Soc R Zool Belg 93:137–154
Hill SD (1970) Origin of the regeneration blastema in polychaete annelids. Am Zool 10:101–112
Hofmann DK (1966) Untersuchungen zur Regeneration des Hinterendes bei Platynereis dumerilii (Audouin et Milne-Edwards) (Annelida, Polychaeta). Zool Jahrb allg Zool 72:374–430
Hyman LH (1940) Aspects of regeneration in annelids. Am Nat 74:513–527
Juliano C, Wessel G (2010) Versatile germline genes. Science 329:640–641. doi:10.1126/science.1194037
Juliano CE, Swartz SZ, Wessel GM (2010) A conserved germline multipotency program. Development 137:4113–4126. doi:10.1242/dev.047969
Juliano C, Wang J, Lin H (2011) Uniting germline and stem cells: the function of Piwi proteins and the piRNA pathway in diverse organisms. Annu Rev Genet 45:447–469. doi:10.1146/annurev-genet-110410-132541
Korotkova GP (1997) Regeneration in animals. Saint-Petersburg University Press, Saint-Petersburg
Kostyuchenko RP, Kozin VV, Smirnova NP, Babakhanova RA (2012) Formation of fission zone and growth zone as multipotent cell populations in development and regeneration of annelids. Euro Evo Devo Lisbon 2012 July 10-13 Abstract Book. Lisbon, p 193
Kulakova M, Bakalenko N, Novikova E, et al (2007) Hox gene expression in larval development of the polychaetes Nereis virens and Platynereis dumerilii (Annelida, Lophotrochozoa). Dev Genes Evol 217:39–54. doi: 10.1007/s00427-006-0119-y
Marilley M, Thouveny Y (1978) DNA synthesis during the first stages of anterior regeneration in the polychaete annelid Owenia fusiformis (dedifferentiation and early phases of differentiation). J Embryol Exp Morphol 44:81–92
Myohara M (2012) What role do annelid neoblasts play? A comparison of the regeneration patterns in a neoblast-bearing and a neoblast-lacking enchytraeid oligochaete. PLoS ONE 7:e37319. doi:10.1371/journal.pone.0037319
Niwa N, Akimoto-Kato A, Sakuma M et al (2013) Homeogenetic inductive mechanism of segmentation in polychaete tail regeneration. Dev Biol 381:460–470. doi:10.1016/j.ydbio.2013.04.010
Novikova EL, Bakalenko NI, Nesterenko AY, Kulakova MA (2013) Expression of Hox genes during regeneration of nereid polychaete Alitta (Nereis) virens (Annelida, Lophotrochozoa). EvoDevo 4:14. doi:10.1186/2041-9139-4-14
Oyama A, Shimizu T (2007) Transient occurrence of vasa-expressing cells in nongenital segments during embryonic development in the oligochaete annelid Tubifex tubifex. Dev Genes Evol 217:675–690. doi:10.1007/s00427-007-0180-1
Oyama A, Yoshida H, Shimizu T (2008) Embryonic expression of p68, a DEAD-box RNA helicase, in the oligochaete annelid Tubifex tubifex. Gene Expr Patterns 8:464–470. doi:10.1016/j.gep.2008.02.003
Paulus T, Müller MCM (2006) Cell proliferation dynamics and morphological differentiation during regeneration in Dorvillea bermudensis (Polychaeta, Dorvilleidae). J Morphol 267:393–403. doi:10.1002/jmor.10280
Pfeifer K, Dorresteijn AWC, Fröbius AC (2012) Activation of Hox genes during caudal regeneration of the polychaete annelid Platynereis dumerilii. Dev Genes Evol 222:165–179. doi:10.1007/s00427-012-0402-z
Potswald HE (1969) Cytological observations on the so-called neoblasts in the serpulid Spirorbis. J Morphol 128:241–259. doi:10.1002/jmor.1051280207
Raible F, Tessmar-Raible K, Osoegawa K et al (2005) Vertebrate-type intron-rich genes in the marine annelid Platynereis dumerilii. Science 310:1325–1326. doi:10.1126/science.1119089
Randolph H (1892) The regeneration of the tail in Lumbriculus. J Morphol 7:317–344. doi:10.1002/jmor.1050070304
Rebscher N, Zelada-González F, Banisch TU et al (2007) Vasa unveils a common origin of germ cells and of somatic stem cells from the posterior growth zone in the polychaete Platynereis dumerilii. Dev Biol 306:599–611. doi:10.1016/j.ydbio.2007.03.521
Reddien PW, Oviedo NJ, Jennings JR et al (2005) SMEDWI-2 is a PIWI-like protein that regulates planarian stem cells. Science 310:1327–1330. doi:10.1126/science.1116110
Rink JC (2013) Stem cell systems and regeneration in planaria. Dev Genes Evol 223:67–84. doi:10.1007/s00427-012-0426-4
Rinkevich Y, Rosner A, Rabinowitz C et al (2010) Piwi positive cells that line the vasculature epithelium, underlie whole body regeneration in a basal chordate. Dev Biol 345:94–104. doi:10.1016/j.ydbio.2010.05.500
Simakov O, Larsson TA, Arendt D (2013) Linking micro- and macro-evolution at the cell type level: a view from the lophotrochozoan Platynereis dumerilii. Brief Funct Genom els049. doi:10.1093/bfgp/els049
Solana J (2013) Closing the circle of germline and stem cells: the primordial stem cell hypothesis. EvoDevo 4:2. doi:10.1186/2041-9139-4-2
Struck TH, Paul C, Hill N et al (2011) Phylogenomic analyses unravel annelid evolution. Nature 471:95–98. doi:10.1038/nature09864
Sugio M, Takeuchi K, Kutsuna J et al (2008) Exploration of embryonic origins of germline stem cells and neoblasts in Enchytraeus japonensis (Oligochaeta, Annelida). Gene Expr Patterns 8:227–236. doi:10.1016/j.gep.2007.12.008
Tessmar-Raible K, Steinmetz PRH, Snyman H et al (2005) Fluorescent two-color whole mount in situ hybridization in Platynereis dumerilii (Polychaeta, Annelida), an emerging marine molecular model for evolution and development. BioTechniques 39:460–462
Yoshida-Noro C, Tochinai S (2010) Stem cell system in asexual and sexual reproduction of Enchytraeus japonensis (Oligochaeta, Annelida). Dev Growth Differ 52:43–55. doi:10.1111/j.1440-169X.2009.01149.x
Zantke J, Bannister S, Rajan VBV et al (2014) Genetic and genomic tools for the marine annelid Platynereis dumerilii. Genetics 197:19–31. doi:10.1534/genetics.112.148254
Zhu W, Pao GM, Satoh A et al (2012) Activation of germline-specific genes is required for limb regeneration in the Mexican axolotl. Dev Biol 370:42–51. doi:10.1016/j.ydbio.2012.07.021
Acknowledgments
This work was supported by the RFBR grant 12-04-00523 and the Saint Petersburg State University grant 1.38.209.2014. We are grateful to the research resource center “Molecular and Cell Technologies” of St. Petersburg State University for a technical support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Volker G. Hartenstein
Rights and permissions
About this article
Cite this article
Kozin, V.V., Kostyuchenko, R.P. Vasa, PL10, and Piwi gene expression during caudal regeneration of the polychaete annelid Alitta virens . Dev Genes Evol 225, 129–138 (2015). https://doi.org/10.1007/s00427-015-0496-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-015-0496-1