Abstract
Main conclusion
Transcriptomic analysis indicates that the bacterial signalling molecule lumichrome enhances plant growth through a combination of enhanced cell division and cell enlargement, and possibly enhances photosynthesis.
Lumichrome (7,8 dimethylalloxazine), a novel multitrophic signal molecule produced by Sinorhizobium meliloti bacteria, has previously been shown to elicit growth promotion in different plant species (Phillips et al. in Proc Natl Acad Sci USA 96:12275–12280, https://doi.org/10.1073/pnas.96.22.12275, 1999). However, the molecular mechanisms that underlie this plant growth promotion remain obscure. Global transcript profiling using RNA-seq suggests that lumichrome enhances growth by inducing genes impacting on turgor driven growth and mitotic cell cycle that ensures the integration of cell division and expansion of developing leaves. The abundance of XTH9 and XPA4 transcripts was attributed to improved mediation of cell-wall loosening to allow turgor-driven cell enlargement. Mitotic CYCD3.3, CYCA1.1, SP1L3, RSW7 and PDF1 transcripts were increased in lumichrome-treated Arabidopsis thaliana plants, suggesting enhanced growth was underpinned by increased cell differentiation and expansion with a consequential increase in biomass. Synergistic ethylene–auxin cross-talk was also observed through reciprocal over-expression of ACO1 and SAUR54, in which ethylene activates the auxin signalling pathway and regulates Arabidopsis growth by both stimulating auxin biosynthesis and modulating the auxin transport machinery to the leaves. Decreased transcription of jasmonate biosynthesis and responsive-related transcripts (LOX2; LOX3; LOX6; JAL34; JR1) might contribute towards suppression of the negative effects of methyl jasmonate (MeJa) such as chlorophyll loss and decreases in RuBisCO and photosynthesis. This work contributes towards a deeper understanding of how lumichrome enhances plant growth and development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant growth progresses through different developmental phases and the transitions are controlled by distinct genetic cues that integrate endogenous and environmental signals. A variety of bacterial genera are present in soils, some of which produce substances that enhance plant growth. Such plant growth-promoting rhizobacteria (PGPR) stimulate plant growth through mobilizing nutrients in soils, producing numerous plant growth regulators, protecting plants by controlling or inhibiting phytopathogens, improving soil structure and bioremediating polluted soils by sequestering toxic heavy metal species and degrading xenobiotic compounds (Bhattacharyya and Jha 2012). Diverse symbiotic (Rhizobium, Bradyrhizobium, Mesorhizobium) and non-symbiotic (Pseudomonas, Bacillus, Klebsiella, Azotobacter, Azospirillum, Azomonas) rhizobacteria are used worldwide as bio-inoculants to promote plant growth and development, including under various stresses. Other than nitrogen fixing symbioses, many PGPR release phytohormones that promote cell growth and massively increase root hair production (Yanni et al. 2001). PGPR also facilitate plant growth directly by assisting in nutrient resource acquisition. Root-nodule bacteria synthesize signal molecules in soil that indirectly promote plant growth via an increase in nutrient availability and uptake through enhanced root absorptive capacity in the transpiration stream (Zhang and Smith 2002; Matiru and Dakora 2004, 2005a).
The discovery of lumichrome (7,8 dimethylalloxazine) as a novel plant growth promoting multitrophic signal molecule produced by the bacterium Sinorhizobium meliloti (Phillips et al. 1999), has driven research to elucidate the molecular mechanisms through which this compound induces plant growth. Previous studies reported that its application leads to enhanced root respiration which resulted in increasing concentrations of rhizospheric CO2 needed for growth of N2-fixing rhizobia and mycorrhizal fungi (Phillips et al. 1999). Lumichrome’s effect on improving plant growth and development has been attributed to enhanced leaf stomatal conductance, transpiration and enhanced photosynthetic rates in soybean and corn (Zhang and Smith 2002; Dakora 2003; Matiru and Dakora 2005b). Conversely, its addition led to significantly decreased root respiration in lupin, while in cowpea it decreased stomatal conductance, which subsequently affected CO2 intake and reduction by RuBisCo (Matiru and Dakora 2005b).
At the molecular level, a microarray gene expression study indicated changes in carbon and ethylene metabolism in roots of both Lotus japonicus and tomato treated with lumichrome. Enhanced starch accumulation was attributed to an increase in plastidial GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE (GAPDH) transcripts and NAD-dependent enzyme activity (Gouws et al. 2012), which has previously been shown to generate this phenotype (Muñoz-Bertomeu et al. 2009). Additionally, lumichrome treatment resulted in a reduction of transcript levels of genes involved in ethylene metabolism, including ethylene response factor/elements, ACC OXIDASE 1 (ACO1) and transcriptional factors AP2/EREBP and a C2H2 ZINC FINGER PROTEIN (Gouws et al. 2012). It was speculated that the interaction of lumichrome with ethylene metabolism was the result of a transient redox mimicry and independent from the biomass accumulation. Despite these physiological experimental advances in exploring the growth stimulatory mechanisms of lumichrome, a comprehensive molecular analysis remains elusive. Here, we use next generation sequencing (RNA-seq) in the model plant Arabidopsis thaliana to examine this. Although we previously worked on Lotus japonicus and tomato, the enormous variety of genetic resources offered by A. thaliana make it an obvious choice for further investigations. We, therefore, aimed to identify specific changes in gene expression relating to the enhanced growth observed in A. thaliana plants treated with lumichrome, utilising an RNA-seq approach.
Materials and methods
Plant material, treatment and growth
Arabidopsis thaliana (ecotype Columbia-0) seeds were stratified (4 °C, 48 h) prior to seed sowing. The experiment was conducted in a controlled environment growth chamber (16/8 h day/night, 22 ± 2 °C, 75% relative humidity). Pots were arranged in a factorial randomised complete block design consisting of 6 replications and blocks. Lumichrome stock solutions (100 μM) were freshly prepared for each treatment in methanol/1 M HCl (49:1, v/v) with constant stirring. Intact plants were treated with 5 nM lumichrome by a combination of root drenching and foliar application (50 ml) at intervals of 2 days throughout the entire growth period (Phillips et al. 1999). Control plants were treated using the same dilution of 49:1 methanol/HCl, without added lumichrome, in dH2O. Above-ground plant material (rosette leaves) was harvested from 5-week-old plants for fresh biomass determination. The same plant materials were then oven dried at 70 °C for 3 d and subjected to dry biomass determination.
RNA extraction, library preparation, and Illumina sequencing
Total RNA was isolated from Arabidopsis rosette leaf tissue material (250 mg) according to a CTAB protocol (Hu et al. 2002). RNA was further purified using the Qiagen RNase-free DNase kit (Cat #79254), and eluted in RNase-free water according to the manufacturer’s instructions. Library preparation and sequencing were performed at the Agricultural Research Council Biotechnology Platform (South Africa). For RNA-seq, total RNA was subjected to removal of ribosomal RNA using the Plant Leaf Ribo-Zero™ Magnetic Kit (Illumina), according to the manufacturer’s instructions. Ribosomal RNA-depleted RNA samples were fragmented, and first-strand cDNA synthesis performed using random hexamers and reverse transcriptase. The cDNA was converted to double-stranded cDNA, subjected to end repair and 3′ adenylation and ligated to Illumina TrueSeq’s paired-end index adaptor before the DNA fragments were PCR enriched (15 cycles). The purity and size of resulting libraries were verified on an Agilent Technologies 2100 Bioanalyzer with an expected fragment of approximately 260 bp. The prepared libraries were sequenced on the Illumina Platform, generating paired-end reads of length 125 nt.
Differential gene expression analysis
Adaptor sequences were removed from the raw sequencing reads and low-quality bases at read ends were trimmed (minimum quality 20 Phred score over a 3 nt window, minimum read length 20 nt) using Trimmomatic v. 0.33 (Bolger et al. 2014). The Tuxedo software suite v.2.2 (Bowtie, TopHat, Cufflinks, Cuffdiff; Trapnell et al. 2012) was used to compare samples and calculate differential expression. Trimmed sequencing reads were aligned against the wild type Arabidopsis (Columbia-0; TAIR10) genome and gene expression was quantified as Fragments Per Kilobase of transcript per Million mapped reads (FPKM). Differential expression was calculated based on Cuffdiff statistical tests of three replicates of treated relative to untreated samples, using a statistical significance of q (adjusted P value) < 0.05.
Validation of RNA sequencing data by reverse transcription quantitative PCR (RT-qPCR)
The same DNase-treated RNA pool as was used for library construction was used for validation of the RNA-seq data. The complementary DNA (cDNA) template was obtained via reverse transcription of 1 µg of total RNA, using an oligo (dT18) primer and M-MLV (H-) reverse transcriptase (Promega) following the manufacturer’s instruction. The integrity of cDNA template was checked with RT-qPCR using MONENSIN SENSITIVITY1 (At2g28390; MON1) as a constitutively expressed gene (Supplementary Fig. S1). To validate the reliability of the RNA-seq, we performed RT-qPCR using SYBR-Green dye in reactions containing KAPA SYBR® FAST qPCR Kit Master Mix (2X) ROX Low (KAPA Biosystems, Cape Town, South Africa), forward and reverse primers (200 nM each) and a 5X diluted cDNA template (10 ng) according to the manufacturer’s instructions. The primer pairs (Supplementary Table S1) were designed with Quant Primer tool (Arvidsson et al. 2008), and PCR reactions were run in a 7500 Real-Time PCR System (Applied Biosystems) under conditions that included initial denaturing/enzyme activation at 95 °C for 3 min and 40 cycles at 95 and 60 °C for 3 and 30 s, respectively. Data for at least three technical replicates were analysed using the Applied Biosystems SDS software (version 1.4), while the cycle threshold (Ct) was used to determine the relative expression level of a given gene using the E−ΔΔCt method [PCR efficiency (E) was calculated using LinRegPCR (version 2014.5)] (Ramakers et al. 2003). MON1, a member of the SAND family, shown by RNA-seq to be invariantly expressed in the samples, was used as a housekeeping reference control gene (Czechowski et al. 2005).
Results
Growth response of Arabidopsis upon lumichrome treatment
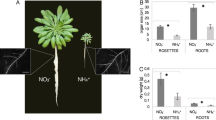
To test for the effect of lumichrome, Arabidopsis thaliana Columbia-0 plants were watered with a combination of root drenching and foliar application every 2 days with 5 nM lumichrome. Fresh and dry weights of 5-week-old rosette leaves showed significantly increased biomass production in treated plants (Fig. 1a, b). Rosettes from treated plants were visually distinguishable as larger than those of untreated plants after 5 weeks of growth (Fig. 1c).
Biomass production analysis of lumichrome-treated Arabidopsis thaliana plants. Lumichrome treatment (5 nM) enhanced fresh (a) and dry biomass production (b) in Arabidopsis compared to the untreated control. Treated plants had a visibly larger rosette than untreated plants after 5 weeks of treatment (c). Error bars represent standard error (mean ± SE, n = 6) of 6 individual plants, while asterisks represent significant differences between the treatments (P ≤ 0.05)
Functional classification and gene enrichment of differentially expressed responsive genes
A microarray study on Lotus japonicus and tomato (Gouws et al. 2012) has previously assisted in our understanding the complexity of the transcriptional regulation and its impact on phenotype after application of lumichrome. However, next-generation sequencing is a more sensitive methodology to examine differential gene expression because of its high throughput and accuracy (Szittya et al. 2008). The current study, therefore, adopted a high-throughput paired-end sequencing of ribosomal-depleted RNA on the Illumina HiSeq 2500 platform, and used the Tuxedo analysis protocol to further investigate lumichrome-associated transcriptional modulation. Statistics of clean reads in RNA sequencing are shown in Supplementary Table S1. Out of the total 32 845 different transcripts analysed, only 198 showed significant differential expression, with 66 and 132 transcripts being significantly up or down-regulated, respectively.
To gain insight into which transcripts were most highly over-represented, transcriptome data were loaded into PageMan and a Wilcoxon test was applied to each category. Based on this statistical overview, enrichment analysis of differentially expressed genes that were up-regulated in lumichrome-treated Arabidopsis rosette leaves relative to their untreated control were associated with lipid metabolism, RNA regulation of transcription was over-represented. In contrast, down-regulated genes resulted in a shift of the gene enrichment functional categories associated to secondary metabolism, hormone metabolism (jasmonates), biotic stress (pathogenesis-related and plant defensins), miscellaneous myrosinases (jacalin-lectin) and development storage proteins. Besides the over-represented biological functional categories revealed by Wilcoxon, several transcripts unique to the lumichrome treatment were associated with phytohormone-related pathways including auxin, gibberellin, ethylene, and jasmonate. In addition, a significant up-regulation of mitotic transcripts was also observed in the category cell cycle, division and development (Table 1).
Cell cycle, division and development related metabolism
Lumichrome treatment of Arabidopsis prompted an up-regulated expression of cell wall and cell cycle transcripts such as RADIALLY SWOLLEN 7 (RSW7, At2g28620) and XYLOGLUCAN ENDOTRANSGLUCOSYLASE/HYDROLASE 9 (XTH9; At4g03210) and EXPANSIN A4 (EXPA4; At2g39700). Lumichrome treatment additionally resulted in an increase in transcription of cell mitosis CYCLIN A1;1 (CYCA1;1; At1g44110), and CYCLIN D3;3 (CYCD3;3; At3g50070). Similarly plant development-associated transcripts, namely, PROTODERMAL FACTOR 1 (PDF1; At2g42840), SPIRAL 1-LIKE3 (SP1L3; At3g02180) and UNIVERSAL STRESS PROTEIN (USP) FAMILY PROTEIN/EARLY NODULIN ENOD18 family protein (At3g03270) were also induced upon lumichrome treatment (Table 1).
Hormone metabolism and signalling
Transcripts for ethylene biosynthesis (Table 1) were altered through increased expression of the 1-AMINOCYCLOPROPANE-1-CARBOXLATE OXIDASE (ACO1; At2g19590) gene. This was accompanied by expression of several transcription factors belonging to proteins predicted to contain an ethylene-responsive element, including an AP2/EREBP/RAP2.4 transcription factor (At1g22190). Interestingly, transcripts for proteins from other hormone signalling pathways that have been implicated in cross-talk with ethylene signalling during Arabidopsis growth, such as the auxin responsive protein SMALL AUXIN RNA51 (SAUR54; At1g19830), and the gibberellin-induced transcript, GAST1 PROTEIN HOMOLOG 1 (GASA1; At1g75750), were also induced. A distinct suppression of jasmonate metabolism-related transcripts was demonstrated through the down-regulated expression of Arabidopsis lipoxygenases LOX2 (At3g45140), LOX3 (At1g17420) and LOX6 (At1g67560). In addition, we observed down-regulated transcripts for jasmonate responsive transcripts, including JACALIN-RELATED LECTIN 34 (JAL34; At3g16460) and JASMONATE RESPONSIVE 1 (JR1; At3g16470) in lumichrome-treated Arabidopsis plants.
Stress and defence response
Treating Arabidopsis rosette leaves with lumichrome resulted in a distinct decrease in transcript levels for genes involved in systemic acquired resistance against pathogens (Table 1). This includes ten plant immunity-related transcripts such as PATHOGENESIS-RELATED GENE 1 (PR1; At2g14610), PATHOGENESIS-RELATED 4 (PR4; At3g04720), DISEASE RESISTANCE FAMILY PROTEIN (At2g34930), DEFENSIN-LIKE PROTEIN 194 (At2g43530), DEFENSIN-LIKE PROTEIN 287 (At1g13609), PLANT DEFENSIN 1.2C (PDF1.2C; At5g44430), PLANT DEFENSIN 1.2 (PDF1.2A; At5g44420), DEFENSIN-LIKE (At2g43510), PLANT DEFENSIN 1.3 (PDF1.3; At2g26010) and PLANT DEFENSIN 1.2B (PDF1.2B; At2g26020). Other than reduced plant immunity-related transcripts, lumichrome-treated plants showed increased levels of transcripts encoding PLASMA MEMBRANE INTRINSIC PROTEIN 3 (PIP3A) and PLASMA MEMBRANE INTRINSIC PROTEIN 1C (PIP1C), both aquaporins involved in the salt stress response, and DEFECTIVE IN INDUCED RESISTANCE1 (DIR1), which is a lipid transfer protein involved in systemic acquired resistance.
Validation of RNA-seq gene expression by RT-qPCR
To validate RNA-seq results, a number of differentially expressed transcripts were randomly selected for expression analysis by RT-qPCR (Supplementary Table S2). The up-regulated transcripts (XTH9, RSW7, CYCA1;1, GASA1, ACO1) and down-regulated transcripts (LOX3, JAL34) were analysed by RT-qPCR using MON1 (AT2G28390) as reference gene. MON1, a member of the SAND family, has previously been identified as a useful reference gene in Arabidopsis expression studies (Czechowski et al. 2005) and was also invariant in the RNA-seq expression data. Comparisons between the RT-qPCR and RNA-seq analysis showed a positive correlation between the two approaches, indicating that the RNA-seq expression analysis performed is highly reliable (Fig. 2).
RT-qPCR validation of differentially expressed genes from lumichrome-treated Arabidopsis RNA-seq data. Histograms represent relative transcript expression levels of RNAseq and RT-qPCR of lumichrome-treated against untreated rosette leaves. Log2 fold change of transcript levels was determined from replicates (n = 3) of each sample while for qPCR, the Ct values were averaged and normalized to MON1 according to E−ΔΔCt method (Ramakers et al. 2003). Error bars represent standard error (mean ± SE, n = 3). Relative expression of all transcripts was significantly different at P ≤ 0.05
Discussion
There is ongoing rigorous research worldwide exploring a range of plant growth promoting compounds for use in improving crop production. Lumichrome is one such plant growth promoting rhizobacterial signal molecule that stimulates growth in a variety of plant taxa (Phillips et al. 1999; Zhang and Smith 2002; Dakora 2003; Matiru and Dakora 2005a, b; Khan et al. 2008; Gouws et al. 2012). Our results confirm its growth promoting effect, as demonstrated by significantly increased rosette leaf fresh and dry biomass. This prompted RNA-seq and RT-qPCR validation analysis to elucidate the molecular mechanisms allied to growth promotion on the gene transcription level.
Lumichrome induces genes involved in plant growth reconfiguration through the control of turgor-driven cell elongation and mitotic cell cycle
In plants, the balance between growth and cell cycle progression requires coordinated regulation of four different processes: macromolecular synthesis (cytoplasmic growth), turgor-driven cell wall extension, mitotic cycle expansion and endocycle. The up-regulated expression of XTH9 and EXPA4 transcripts following lumichrome treatment suggested the disruption of elaborate microtubule arrays, cellulose deposition and cell-wall thickening, thereby allowing cell wall loosening and turgor-driven cell enlargement (Wolf et al. 2012). In Arabidopsis, XTHs tend to be expressed strongly in rapidly dividing and expanding tissues (Hyodo et al. 2003; Jan et al. 2004), whilst expression of EXPANSINs improved cell wall loosening and promoted above-ground biomass in transgenic rice (Choi et al. 2003). EXPANSINs initiate the development of the leaf primordium, which later recapitulates the entire process of leaf formation (Pien et al. 2001), enhancing leaf cell size and results in larger leaves (Kuluev et al. 2012).
Besides this turgor-driven growth, increased expression of the core mitotic CYCD3.3 and CYCA1.1 transcripts suggested concurrent cell proliferation and cell differentiation during leaf development (Braybrook and Kuhlemeier 2010), which in turn possibly contributed to an increase in biomass. Cell cycle control and plant development are mainly integrated at the cell cycle checkpoints (G1/S and G2/M) and the molecular machineries involved (De Veylder et al. 2003; Inzé 2005). Expression of CYCD3 is promoted by cytokinin (CK), hence loss of CYCD3;1-3 activity reduces the capacity for exogenous CK to initiate shoot formation (Riou-Khamlichi 1999; Dewitte et al. 2007). Conversely, over-expression of CYCD3;1 is sufficient to confer CK-independent shoot formation from callus tissue (Riou-Khamlichi 1999). In that regard, a positive correlation of increased biomass and increased expression of CYCD3.3 following lumichrome treatment in our study could be due to a CK-independent activity that directly enhanced G2/M phase transition (Sorrell et al. 1999). In addition, the promoted G1/S transition phase was suggested through an increase in mitotic CYCA1;1 transcripts. Increased core mitotic cycle transcripts were coupled with an increased CASEIN KINASE ALPHA 1 (CKA1) expression, suggesting their reciprocal function in regulating mitotic cell cycle in plants (Reichheld et al. 1996). This is in agreement with reports that tissues with high mitotic activity, such as meristems, show a higher level of CK2 transcripts, indicating a role for CK2 in cell proliferation in these tissues. This has previously been shown in dominant negative mutant of CK2 in Arabidopsis, which demonstrated an up-regulated expression of the core cell cycle-related genes at the G2/M transition (Moreno-Romero et al. 2011). Therefore, an increase in another kinase such as CKA1 in lumichrome-treated plants suggested signalling of increased mitotic cell cycle transcripts, which further suggests increased cell division and expansion with a consequential increase in biomass.
Other than the increased core mitotic cyclins, transcripts that are involved in spindle assembly and cell cycle progression from G2 phase to metaphase, namely SPIRAL1-LIKE3 (SP1L3; At3g02180) and RADIALLY SWOLLEN 7 (RSW7; At2g28620) were also increased. SP1L3 is required for cortical microtubule directional control of rapidly expanding Arabidopsis cells through directional deposition of cellulose microfibrils (Nakajima et al. 2006; Foteinopoulos and Mulder 2014). Defects in SP1L3 phosphorylation impair events that participate in the spatiotemporal regulation of acentrosomal spindles, leading to mitotic defects which in turn result in enhanced ploidy, development arrest of apical meristems, ectopic meristem formation and defects in tissue patterning (Petrovská et al. 2012). RSW7 plays a role in the formation of unique cellular structures such as the phragmoplast and the cell plate, both of which are required to divide the cell after nuclear division (Gillmor et al. 2016). Mutations in RSW7 retard growth by disrupting the normal pattern of wall placement (Wiedemeier et al. 2002).
Still on the theme of cell expansion and differentiation, our results demonstrated an induced expression of PROTODERMAL FACTOR 1 (PDF1), which encodes a proline-rich cell-wall protein that is expressed exclusively in the protodermal tunica layer (L1) of shoot meristems (Abe et al. 2001). Together with Arabidopsis thaliana MERISTEM LAYER1 (AtML1) and PROTODERMAL FACTOR2 (PDF2), PDF1 encodes an L1 box-binding homeodomain protein with high homeobox sequence similarity and shows expression exclusively in the L1 of vegetative meristem and epidermis of the Arabidopsis shoot apical meristem (SAM) and throughout the shoot development (Lu et al. 1996; Abe et al. 2003). Failure to differentiate epidermal cells in a PDF2-1 and AtML1-1 double mutant explains the role of these genes in the differentiation of epidermal cells from the L1 of shoot meristems (Abe et al. 2003). In that respect, due to their high homeobox sequence similarities, an increase in PDF1 expression in the current study suggested its role in the differentiation of epidermal cells from the L1 of shoot meristems giving rise to cell division and differentiation of the SAM, resulting into an increase in above-ground biomass (Fletcher and Meyerowitz 2000).
Reciprocal hormonal cross-talk and signalling may play a role in enhanced Arabidopsis growth
Plant hormones play essential roles in coordinating external and internal signals to elicit the appropriate growth and developmental responses to precisely regulate responses to both temporal and spatial stimuli. A pronounced reduction in transcripts of genes involved in ethylene metabolism, such as ACO1, AP2/EREBP and a C2H2 zinc finger protein, was previously reported in Lotus japonicus and tomato (Gouws et al. 2012). We observed, however, increased expression of the ethylene biosynthesis-related ACO1 transcript and the AP2.4/EREBP transcription factor (At1g22190), hence suggesting that the response might be species-specific. In addition, we observed increased transcript levels of a primary auxin response transcript, SAUR54, which positively regulates cell expansion to promote hypocotyl growth and leaf cell expansion (Spartz et al. 2012; Stamm and Kumar 2013). This response also suggested an auxin and ethylene cross-talk mechanism. Ethylene is produced from methionine (Met) via S-adenosyl-l-methionine (AdoMet) and 1-aminocyclopropane-1-carboxylate (ACC) in which ACC synthase and ACC oxidase (ACO1), respectively, catalyze the last two steps in this biosynthetic pathway (Kende and Zeevaart 1997). Auxin is known to greatly stimulate ethylene production in vegetative tissues by inducing the synthesis or activation of ACC SYNTHASE, CS1 and ACO1 (Abel et al. 1995). As such, the increased SAUR54 expression could have resulted in activation of ACO1 and AP2.4/EREBP transcription in lumichrome-treated plants. Consistent with a previous study (Li et al. 2015), an increase in SAUR54 expression might confer reduced sensitivity to ethylene, resulting in enhanced rosette growth in lumichrome-treated plants. Simultaneous increases in expression of SAUR54 and endoplasmic reticulum (ER) localized RING MEMBRANE-ANCHOR 2 (At4g28270; RMA2) transcripts might have positively regulated auxin transport across the ER membrane, thereby regulating cellular auxin homoeostasis and hormonal control of Arabidopsis vegetative growth (Peret et al. 2012; Bou-Torrent et al. 2014). A reciprocal increase in SAUR54 and GASA1 expression may be linked to increases in auxin and gibberellin signalling, respectively, which may in turn enhance growth via synergistic hormonal cross-talk between these two hormones. Classically, gibberellins are a growth-promoting class of phytohormones, regulating a wide range of growth and developmental processes throughout the life cycle of a plant, including leaf expansion, induction of flowering, as well as flower and seed development (Davière and Achard 2013). Auxin has previously been shown to regulate the expression of a number of gibberellic acid (GA) metabolic genes involved in the synthesis of active GAs in pea stem and Arabidopsis seedlings (Frigerio et al. 2006; Chapman et al. 2012). Our results suggest that an increase in SAUR54 transcripts may have increased levels of auxin, which in turn up-regulated the expression of GASA1 for GA metabolism. The transcriptional regulation between auxins and GA hormone pathways and their signalling role is, therefore, likely to be important for the synergistic cross-talk mediated cell division and cell expansion (Ross et al. 2000).
Jasmonates are oxylipin signalling molecules, including 12-oxo-phytodienoic acid (OPDA), jasmonic acid (JA), and derivatives such as the methyl ester and amino acid conjugates of JA. JA is synthesized from linolenic acid, which is first oxygenated by lipoxygenase, to yield 13(S)-hydroperoxy linolenic acid (13-HPOT) (Vick and Zimmerman 1984). The Arabidopsis genome contains six lipoxygenase genes, of which LOX2 (At3g45140), LOX3 (At1g17420), and LOX6 (At1g72520) contain chloroplast signalling peptides and show 13S-lipoxygenase activity, both features that are required for JA biosynthesis upon wounding and during senescence in leaves (Bell et al. 1995; He et al. 2002; Chung et al. 2008; Seltmann et al. 2010). The leaf senescence-promoting effect of methyl jasmonate (MeJa) is accompanied by chlorophyll loss and decreases in RuBisCO and photosynthesis (Weidhase et al. 1987), which is detrimental for plant growth and productivity. However, LOX2, LOX3, LOX6, JAL34 and JR1 were all down-regulated in response to lumichrome treatment. These genes are essential for JA biosynthesis in wounded or senescing leaves (Bell et al. 1995; Seltmann et al. 2010) except for LOX3, which is transcribed in the roots and the transcript is transported to leaves and activated during leaf senescence (He et al. 2002; Chung et al. 2008). In that regard, decreased levels of these transcripts in the current study suggested a substantive role of lumichrome in delayed leaf senescence through the suppression of the effects of MeJa, namely chlorophyll loss and decreases in RuBisCO and photosynthesis. In addition, this response suggested a decrease in plant immunity against pathogen-induced injury through the expression of LOX6, which is essential for stress-induced jasmonate accumulation in Arabidopsis leaves (Chauvin et al. 2013). Reduced transcripts encoding PATHOGENESIS-RELATED (PR1; PR4), PLANT DEFENSINS, (PDF1.2C; PDF1.2A; PDF1.3; PDF1.2B) and PLANT DEFENSIN-LIKE (At2g43530, At2g43530 At1g13609) also suggested reduced plant immunity against pathogens. Arabidopsis thaliana mutants that are impaired in JA production or perception exhibit enhanced susceptibility to a variety of pathogens, including fungal pathogens (Thomma et al. 1998; Norman-Setterblad et al. 2000). Similarly, growth promotion effects in lumichrome-treated plants were accompanied by down-regulation of a large suite of plant defence genes, including the jasmonic acid biosynthetic pathway, PLANT PATHOGENESIS and DEFENSINS. However, this study did not expose Arabidopsis to pathogenic attack or fungal infection, highlighting the need to challenge lumichrome-treated Arabidopsis with pests and pathogens to test the validity of the idea that their defence responses may be impaired.
Despite the possibility of reduced plant immunity against pathogens including fungi, lumichrome-treated plants exhibited increased expression of transcripts encoding CYSTEINE-RICH RECEPTOR-LIKE PROTEIN KINASE 41 (CRK41, also designated DUF26 26 [Wrzaczek et al. 2010]; Table 1). The extracellular domain of CRKs encompasses two copies of the DUF26 (Domain of Unknown Function 26) domain, which contains three cysteine residues in a conserved configuration (C-X8-C-X2-C). The presence and spacing of the conserved cysteines in the DUF26 domain suggest that CRKs might be connected to ROS and redox signalling (Chen et al. 2004; Wrzaczek et al. 2010, 2013). Elevated transcript levels of several CRK genes trigger intracellular signalling cascades, allowing cells to respond and adapt to internal and external stimuli. For instance, CRKs play an important role as mediators of signalling specificity during regulation of stomatal aperture (Bourdais et al. 2015). Therefore, increased levels of transcripts encoding CRK41, coupled with up-regulated PLASMA MEMBRANE INTRINSIC PROTEIN 3 (PIP3A) and PLASMA MEMBRANE INTRINSIC PROTEIN 1C (PIP1C), suggest a mutual functional relationship of redox modifications and plasma membrane permeabilities for better water and nutrient uptake and movement of sugars for metabolism in the plant (Viger et al. 2014). These responses are also known to improve photosynthetic efficiency and to increase biomass production (Farquhar and Sharkey 1982; Foyer and Shigeoka 2011).
Conclusion
The addition of 5 nM lumichrome elicited a growth promotion effect in Arabidopsis (Fig. 1). Based on our RNAseq data, we propose a model for growth enhancement via hormonal cross talk, intracellular signalling cascades and mitotic cell differentiation and expansion in lumichrome-treated plants. Firstly, increased abundance of XTH9 and AtEXP4 transcripts suggested loosening and rearrangement of the cell wall and subsequent cell expansion. Likewise, an increase in specific mitotic cell cycle genes (CYCA1;1; CYCD3;3; SP1L3; RSW7) suggested that the integration of cell division and expansion to developing leaves was promoted through the regulation of mitotic cell cycle phase and microtubule cellular organization and proliferation. Secondly, the reciprocal over-expression of ACO1 and SAUR54 suggested the synergistic ethylene–auxin cross-talk effect in which ethylene activated the auxin-signalling pathway and regulated Arabidopsis growth by both stimulating the auxin biosynthesis and by modulating the auxin transport machinery to the leaves (SAUR54; RMA2). Thirdly, simultaneous reduced expression levels of lipoxygenases (LOX2; LOX3; LOX6) and jasmonate-related transcripts (JAL34; JR1) suggested delayed jasmonate-associated leaf senescence, which might further contribute to improved chlorophyll biosynthesis and photosynthetic productivity. Although our study did not experimentally expose plants to stress conditions, we also observed an upregulated expression of stress induced genes associated to an ABA-independent dehydration and salinity stress signalling. While these findings gave us new insights and enhanced our knowledge of how lumichrome induces growth promotion in Arabidopsis, reverse genetic analysis with mutants and over-expressor lines, using some of these genes, will help show that these genes are actually involved.
Author contribution statement
MP, JK, JL and PH designed the research. The research presented in this manuscript forms a part of the PhD dissertation for MP at Stellenbosch University and which may be accessed online at http://scholar.sun.ac.za/handle/10019.1/101430. MP conducted all the experiments and analysed the data. BC and HM conducted the RNA-seq bioinformatics analysis and PY the RT-qPCR experiments. MP, JK, JL and PH prepared the manuscript. All authors read and approved the manuscript.
Abbreviations
- CK:
-
Cytokinin
- GA:
-
Gibberellic acid
- JA:
-
Jasmonic acid
- PGPR:
-
Plant growth-promoting rhizobacteria
References
Abe M, Takahashi T, Komeda Y (2001) Identification of a cis-regulatory element for L1 layer-specific gene expression, which is targeted by an L1-specific homeodomain protein. Plant J 26:487–494. https://doi.org/10.1046/j.1365-313X.2001.01047.x
Abe M, Katsumata H, Komeda Y, Takahashi T (2003) Regulation of shoot epidermal cell differentiation by a pair of homeodomain proteins in Arabidopsis. Development 130:635–643. https://doi.org/10.1242/dev.00292
Abel S, Nguyen MD, Chow W, Theologis A (1995) ACS4, a primary indoleacetic acid-responsive gene encoding 1-aminocyclopropane-1-carboxylate synthase in Arabidopsis thaliana. Structural characterization, expression in Escherichia coli, and expression characteristics in response to auxin. J Biol Chem 270:19093–19099
Arvidsson S, Kwasniewski M, Riaño-Pachón DM, Mueller-Roeber B (2008) QuantPrime–a flexible tool for reliable high-throughput primer design for quantitative PCR. BMC Bioinform 9:465. https://doi.org/10.1186/1471-2105-9-465
Bell E, Creelman RA, Mullet JE (1995) A chloroplast lipoxygenase is required for wound-induced jasmonic acid accumulation in Arabidopsis. Proc Natl Acad Sci USA 92:8675–8679. https://doi.org/10.1073/pnas.92.19.8675
Bhattacharyya PN, Jha DK (2012) Plant growth-promoting rhizobacteria (PGPR): Emergence in agriculture. World J Microbiol Biotechnol 28:1327–1350. https://doi.org/10.1007/s11274-011-0979-9
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bourdais G, Burdiak PP, Gauthier A et al (2015) Large-scale phenomics identifies primary and fine-tuning roles for CRKs in responses related to oxidative stress. PLoS Genet 11:e1005373. https://doi.org/10.1371/journal.pgen.1005373
Bou-Torrent J, Galstyan A, Gallemí M et al (2014) Plant proximity perception dynamically modulates hormone levels and sensitivity in Arabidopsis. J Exp Bot 65:2937–2947. https://doi.org/10.1093/jxb/eru083
Braybrook SA, Kuhlemeier C (2010) How a plant builds leaves. Plant Cell 22:1006–1018. https://doi.org/10.1105/tpc.110.073924
Chapman EJ, Greenham K, Castillejo C et al (2012) Hypocotyl transcriptome reveals auxin regulation of growth-promoting genes through GA-dependent and -independent pathways. PLoS One 7:e36210. https://doi.org/10.1371/journal.pone.0036210
Chauvin A, Caldelari D, Wolfender JL, Farmer EE (2013) Four 13-lipoxygenases contribute to rapid jasmonate synthesis in wounded Arabidopsis thaliana leaves: a role for LIPOXYGENASE 6 in responses to long-distance wound signals. New Phytol 197:566–575. https://doi.org/10.1111/nph.12029
Chen K, Fan B, Du L, Chen Z (2004) Activation of hypersensitive cell death by pathogen-induced receptor-like protein kinases from Arabidopsis. Plant Mol Biol 56:271–283. https://doi.org/10.1007/s11103-004-3381-2
Choi D, Lee Y, Cho HT, Kende H (2003) Regulation of expansin gene expression affects growth and development in transgenic rice plants. Plant Cell 15:1386–1398. https://doi.org/10.1105/tpc.011965
Chung HS, Koo AJK, Gao X et al (2008) Regulation and function of Arabidopsis JASMONATE ZIM-domain genes in response to wounding and herbivory. Plant Physiol 146:952–964. https://doi.org/10.1104/pp.107.115691
Czechowski T, Stitt M, Altmann T, Udvardi MK (2005) Genome-wide identification and testing of superior reference genes for transcript normalization. Plant Physiol 139:5–17. https://doi.org/10.1104/pp.105.063743.1
Dakora FD (2003) Defining new roles for plant and rhizobial molecules in sole and mixed plant cultures involving symbiotic legumes. New Phytol 158:39–49. https://doi.org/10.1046/j.1469-8137.2003.00725.x
Davière J-M, Achard P (2013) Gibberellin signaling in plants. Development 140:1147–1151. https://doi.org/10.1242/dev.087650
De Veylder L, Joubès J, Inzé D (2003) Plant cell cycle transitions. Curr Opin Plant Biol 6:536–543. https://doi.org/10.1016/j.pbi.2003.09.001
Dewitte W, Scofield S, Alcasabas AA et al (2007) Arabidopsis CYCD3 D-type cyclins link cell proliferation and endocycles and are rate-limiting for cytokinin responses. Proc Natl Acad Sci USA 104:14537–14542. https://doi.org/10.1073/pnas.0704166104
Farquhar GD, Sharkey TD (1982) Stomatal conductance and photosynthesis. Annu Rev Plant Physiol 33:317–345. https://doi.org/10.1146/annurev.pp.33.060182.001533
Fletcher JC, Meyerowitz EM (2000) Cell signaling within the shoot meristem. Curr Opin Plant Biol 3:23–30. https://doi.org/10.1016/S1369-5266(99)00033-3
Foteinopoulos P, Mulder BM (2014) The effect of anisotropic microtubule-bound nucleations on ordering in the plant cortical array. Bull Math Biol 76:2907–2922. https://doi.org/10.1007/s11538-014-0039-3
Foyer CH, Shigeoka S (2011) Understanding oxidative stress and antioxidant functions to enhance photosynthesis. Plant Physiol 155:93–100. https://doi.org/10.1104/pp.110.166181
Frigerio M, Alabadí D, Pérez-Gómez J et al (2006) Transcriptional regulation of gibberellin metabolism genes by auxin signaling in Arabidopsis. Plant Physiol 142:553–563. https://doi.org/10.1104/pp.106.084871
Gillmor CS, Roeder AHK, Sieber P et al (2016) A genetic screen for mutations affecting cell division in the Arabidopsis thaliana embryo identifies seven loci required for cytokinesis. PLoS One 11:e0146492. https://doi.org/10.1371/journal.pone.0146492
Gouws LM, Botes E, Wiese AJ et al (2012) The plant growth promoting substance, lumichrome, mimics starch, and ethylene-associated symbiotic responses in lotus and tomato roots. Front Plant Sci 3:1–20. https://doi.org/10.3389/fpls.2012.00120
He Y, Fukushige H, Hildebrand DF, Gan S (2002) Evidence supporting a role of jasmonic acid in Arabidopsis leaf senescence. Plant Physiol 128:876–884. https://doi.org/10.1104/pp.010843
Hu CG, Honda C, Kita M et al (2002) A simple protocol for RNA isolation from fruit trees containing high levels of polysaccharides and polyphenol compounds. Plant Mol Biol Rep 20:69. https://doi.org/10.1007/BF02801935
Hyodo H, Yamakawa S, Takeda Y et al (2003) Active gene expression of a xyloglucan endotransglucosylase/hydrolase gene, XTH9, in inflorescence apices is related to cell elongation in Arabidopsis thaliana. Plant Mol Biol 52:473–482. https://doi.org/10.1023/A:1023904217641
Inzé D (2005) Green light for the cell cycle. EMBO J 24:657–662. https://doi.org/10.1038/sj.emboj.7600561
Jan A, Yang G, Nakamura H et al (2004) Characterization of a XYLOGLUCAN ENDOTRANSGLUCOSYLASE gene that is up-regulated by gibberellin in rice. Plant Physiol 136:3670–3681. https://doi.org/10.1104/pp.104.052274
Kende H, Zeevaart J (1997) The five “classical” plant hormones. Plant Cell 9:1197–1210. https://doi.org/10.1105/tpc.9.7.1197
Khan W, Prithiviraj B, Smith DL (2008) Nod factor [Nod Bj V (C18:1, MeFuc)] and lumichrome enhance photosynthesis and growth of corn and soybean. J Plant Physiol 165:1342–1351. https://doi.org/10.1016/j.jplph.2007.11.001
Kuluev BR, Knyazev AB, Lebedev YP, Chemeris AV (2012) Morphological and physiological characteristics of transgenic tobacco plants expressing expansin genes: AtEXP10 from Arabidopsis and PnEXPA1 from poplar. Russ J Plant Physiol 59:97–104. https://doi.org/10.1134/S1021443712010128
Li Z-G, Chen H-W, Li Q-T et al (2015) Three SAUR proteins SAUR76, SAUR77 and SAUR78 promote plant growth in Arabidopsis. Sci Rep 5:12477. https://doi.org/10.1038/srep12477
Lu P, Porat R, Nadeau JA, O’Neill SD (1996) Identification of a meristem L1 layer-specific gene in Arabidopsis that is expressed during embryonic pattern formation and defines a new class of homeobox genes. Plant Cell 8:2155–2168. https://doi.org/10.1105/tpc.8.12.2155
Matiru VN, Dakora FD (2004) Potential use of rhizobial bacteria as promoters of plant growth for increased yield in landraces of African cereal crops. Afr J Biotechnol 3:1–7
Matiru VN, Dakora FD (2005a) Xylem transport and shoot accumulation of lumichrome, a newly recognized rhizobial signal, alters root respiration, stomatal conductance, leaf transpiration and photosynthetic rates in legumes and cereals. New Phytol 165:847–855. https://doi.org/10.1111/j.1469-8137.2004.01254.x
Matiru VN, Dakora FD (2005b) The rhizosphere signal molecule lumichrome alters seedling development in both legumes and cereals. New Phytol 166:439–444. https://doi.org/10.1111/j.1469-8137.2005.01344.x
Moreno-Romero J, Armengot L, Marquès-Bueno MM et al (2011) About the role of CK2 in plant signal transduction. Mol Cell Biochem 356:233–240. https://doi.org/10.1007/s11010-011-0970-7
Muñoz-Bertomeu J, Cascales-Miñana B, Mulet JM et al (2009) Plastidial glyceraldehyde-3-phosphate dehydrogenase deficiency leads to altered root development and affects the sugar and amino acid balance in Arabidopsis. Plant Physiol 151:541–558. https://doi.org/10.1104/pp.109.143701
Nakajima K, Kawamura T, Hashimoto T (2006) Role of the SPIRAL1 gene family in anisotropic growth of Arabidopsis thaliana. Plant Cell Physiol 47:513–522. https://doi.org/10.1093/pcp/pcj020
Norman-Setterblad C, Vidal S, Palva ET (2000) Interacting signal pathways control defense gene expression in Arabidopsis in response to cell wall-degrading enzymes from Erwinia carotovora. Mol Plant-Microbe Interact 13:430–438. https://doi.org/10.1094/MPMI.2000.13.4.430
Peret B, Swarup K, Ferguson A et al (2012) AUX/LAX genes encode a family of auxin influx transporters that perform distinct functions during Arabidopsis development. Plant Cell 24:2874–2885. https://doi.org/10.1105/tpc.112.097766
Petrovská B, Cenklová V, Pochylová Ž et al (2012) Plant Aurora kinases play a role in maintenance of primary meristems and control of endoreduplication. New Phytol 193:590–604. https://doi.org/10.1111/j.1469-8137.2011.03989.x
Phillips DA, Joseph CM, Yang GP et al (1999) Identification of lumichrome as a sinorhizobium enhancer of alfalfa root respiration and shoot growth. Proc Natl Acad Sci USA 96:12275–12280. https://doi.org/10.1073/pnas.96.22.12275
Pien S, Wyrzykowska J, McQueen-Mason S et al (2001) Local expression of expansin induces the entire process of leaf development and modifies leaf shape. Proc Natl Acad Sci USA 98:11812–11817. https://doi.org/10.1073/pnas.191380498
Ramakers C, Ruijter JM, Lekanne Deprez RH, Moorman AFM (2003) Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci Lett 339:62–66. https://doi.org/10.1016/S0304-3940(02)01423-4
Reichheld JP, Chaubet N, Shen WH et al (1996) Multiple A-type cyclins express sequentially during the cell cycle in Nicotiana tabacum BY2 cells. Proc Natl Acad Sci USA 93:13819–13824. https://doi.org/10.1073/pnas.93.24.13819
Riou-Khamlichi C (1999) Cytokinin activation of Arabidopsis cell division through a D-Type cyclin. Science 283:1541–1544. https://doi.org/10.1126/science.283.5407.1541
Ross JJ, O’Neill DP, Smith JJ et al (2000) Evidence that auxin promotes gibberellin A1 biosynthesis in pea. Plant J 21:547–552. https://doi.org/10.1046/j.1365-313X.2000.00702.x
Seltmann MA, Stingl NE, Lautenschlaeger JK et al (2010) Differential impact of lipoxygenase2 and jasmonates on natural and stress-induced senescence in Arabidopsis. Plant Physiol 152:1940–1950. https://doi.org/10.1104/pp.110.153114
Sorrell DA, Combettes B, Chaubet-Gigot N et al (1999) Distinct cyclin D genes show mitotic accumulation or constant levels of transcripts in tobacco bright yellow-2 cells. Plant Physiol 119:343–352. https://doi.org/10.1104/pp.119.1.343
Spartz AK, Lee SH, Wenger JP et al (2012) The SAUR19 subfamily of SMALL AUXIN UP RNA genes promote cell expansion. Plant J 70:978–990. https://doi.org/10.1111/j.1365-313X.2012.04946.x
Stamm P, Kumar PP (2013) Auxin and gibberellin responsive Arabidopsis SMALL AUXIN UP RNA36 regulates hypocotyl elongation in the light. Plant Cell Rep 32:759–769. https://doi.org/10.1007/s00299-013-1406-5
Szittya G, Moxon S, Santos DM et al (2008) High-throughput sequencing of Medicago truncatula short RNAs identifies eight new miRNA families. BMC Genom 9:593. https://doi.org/10.1186/1471-2164-9-593
Thomma BPHJ, Eggermont K, Penninckx IAMA et al (1998) Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc Natl Acad Sci USA 95:15107–15111. https://doi.org/10.1073/pnas.95.25.15107
Trapnell C, Roberts A, Goff L et al (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578. https://doi.org/10.1038/nprot.2012.016
Vick BA, Zimmerman DC (1984) Biosynthesis of jasmonic acid by several plant species. Plant Physiol 75:458–461. https://doi.org/10.1104/pp.75.2.458
Viger M, Hancock RD, Miglietta F, Taylor G (2014) More plant growth but less plant defence? First global gene expression data for plants grown in soil amended with biochar. GCB Bioenergy 7:658–672. https://doi.org/10.1111/gcbb.12182
Weidhase RA, Kramell HM, Lehmann J et al (1987) Methyljasmonate-induced changes in the polypeptide pattern of senescing barley leaf segments. Plant Sci 51:177–186. https://doi.org/10.1016/0168-9452(87)90191-9
Wiedemeier AMD, Judy-March JE, Hocart CH et al (2002) Mutant alleles of Arabidopsis RADIALLY SWOLLEN 4 and 7 reduce growth anisotropy without altering the transverse orientation of cortical microtubules or cellulose microfibrils. Development 129:4821–4830
Wolf S, Hématy K, Höfte H (2012) Growth control and cell wall signaling in plants. Annu Rev Plant Biol 63:381–407. https://doi.org/10.1146/annurev-arplant-042811-105449
Wrzaczek M, Brosché M, Salojärvi J et al (2010) Transcriptional regulation of the CRK/DUF26 group of receptor-like protein kinases by ozone and plant hormones in Arabidopsis. BMC Plant Biol 10:95. https://doi.org/10.1186/1471-2229-10-95
Wrzaczek M, Brosché M, Kangasjärvi J (2013) ROS signaling loops—production, perception, regulation. Curr Opin Plant Biol 16:575–582. https://doi.org/10.1016/j.pbi.2013.07.002
Yanni YG, Rizk RY, El-Fattah FKA et al (2001) The beneficial plant growth-promoting association of Rhizobium leguminosarum bv. trifolii with rice roots. Aust J Plant Physiol 28:845–870. https://doi.org/10.1071/Pp01069
Zhang F, Smith DL (2002) Interorganismal signaling in suboptimum environments: the legume-rhizobia symbiosis. Adv Agron 76:125–161. https://doi.org/10.1016/S0065-2113(02)76004-5
Acknowledgements
We are very grateful to Professor Martin Kidd for aid in the statistical analysis. Computations were performed using the University of Stellenbosch Central Analytical Facilities’ HPC2: http://www.sun.ac.za/hpc.
Funding
This research was funded by the National Research Foundation (SARChi Research Chair “Genetic tailoring of biopolymers”) of South Africa.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table S1
Statistics of clean reads in RNA sequencing. Supplementary Table S2 Primer pairs for the selected genes for RT-qPCR validation (DOC 37 kb)
Suppl. Fig. S1
RNA integrity denatured in 1% (w/v) agarose gel and cDNA transcribed from RNA for RT-qPCR (PPT 217 kb)
Rights and permissions
About this article
Cite this article
Pholo, M., Coetzee, B., Maree, H.J. et al. Cell division and turgor mediate enhanced plant growth in Arabidopsis plants treated with the bacterial signalling molecule lumichrome. Planta 248, 477–488 (2018). https://doi.org/10.1007/s00425-018-2916-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-018-2916-8