Abstract
Main conclusion
Fluorescence in situ hybridization and molecular markers have confirmed that several chromosomes from Thinopyrum intermedium ssp. trichophorum have been added to a wheat background, which originated from a cross between a wheat– Thinopyrum partial amphiploid and triticale. The lines displayed blue grains and resistance to wheat stripe rust.
Thinopyrum intermedium has been used as a valuable resource for improving the disease resistance and yield potential of wheat. With the aim to transfer novel genetic variation from Th. intermedium species for sustainable wheat breeding, a new trigeneric hybrid was produced by crossing an octoploid wheat–Th. intermedium ssp. trichophorum partial amphiploid with hexaploid triticale. Fluorescence in situ hybridization (FISH) revealed that Thinopyrum chromosomes were transmitted preferably and the number of rye chromosomes tended to decrease gradually in the selfed derivatives of the trigeneric hybrids. Four stable wheat–Th. intermedium chromosome substitution, addition and translocation lines were selected, and a 2JS addition line, two substitution lines of 4JS(4B) and 4J(4B), and a small 4J.4B translocation line were identified by FISH and molecular markers. It was revealed that the gene(s) responsible for blue grains may located on the FL0.60–1.00 of long arm of Th. intermedium-derived 4J chromosome. Disease resistance screenings indicated that chromosomes 4JS and 2JS appear to enhance the resistance to stripe rust in the adult plant stage. The new germplasm with Th. intermedium introgression shows promise for utilization of Thinopyrum chromosome segments in future wheat improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The success of current and future wheat improvement programs is dependent on a continuous supply of genetic variability, which will principally be sourced from the related species. The perennial species, hexaploid wheatgrass, Thinopyrum intermedium (Host) Barkworth & D.R. Dewey, is widely distributed in Europe (particularly in the Mediterranean region), western Asia, and northern Africa (Dewey 1984). The relative ease of crossability between wheat and Th. intermedium has enabled the development of wheat–Th. intermedium partial amphiploids (Cauderon et al. 1973; Chen et al. 1999; Fedak et al. 2000; Bao et al. 2014). Derived from these amphiploids, a number of wheat–Thinopyrum chromosome addition, substitution and translocation lines have proved to be a valuable resource for enhancing the disease resistance and yield potential of wheat (Han et al. 2003; Chen 2005; Li and Wang 2009; Wang and Wang 2016). The disease resistance genes, such as Pm40, Pm43, Yr50, Lr38 and Sr44, were transferred from Th. intermedium into common wheat (Friebe et al. 1993, 1996; Luo et al. 2009; He et al. 2009; Liu et al. 2013). Genomic in situ hybridization has revealed the complex genomic composition of Th. intermedium which includes three genomes named J, JS, and St, of which the St genome shows a high degree of similarity to that of Pseudoroegneria, while the J and JS genomes appear to be related to the modified Th. elongatum/Th. bessarabicum genomes (Chen et al. 1998; Chen 2005; Mahelka et al. 2011, 2013). However, the complete set of chromosomes across the seven linkage groups and three St, JS, J genomes still have not been transferred to a wheat background. The great geographical and climatic diversity of their native distribution, combined with open pollination, has resulted in high variability and numerous chromosomal rearrangements among Th. intermedium sub-species, which has attracted the studies into further wheat–Th. intermedium hybridization (Wang 2011; Kantarski et al. 2017).
Thinopyrum intermedium ssp. trichophorum displayed a distinctly enriched chromosomal heterochromatin constitution which differs from that of Th. intermedium ssp. intermedium (Xu and Conner 1994). A partial amphiploid between wheat–Th. intermedium ssp. trichophorum contained unique seed storage proteins and also displayed novel disease resistance in wheat (Yang et al. 2006). Recently, we isolated a disomic chromosome substitution involving chromosome 1St#2 carrying stripe rust resistance (Hu et al. 2011) and specific gliadin and high-molecular weight glutenin subunits (HMW-GS) genes (Li et al. 2013), as well as 7St addition lines (Song et al. 2013). In addition, two double disomic substitutions were developed (Li et al. 2015). Therefore, continuous developing novel wheat–Th. intermedium ssp. trichophorum introgression lines may provide effective resistance against new disease pathotypes (Xu et al. 2009).
Trigeneric hybridizations involving the genera Triticum, Aegilops, Secale, Dasypyrum, Psathyrostachys, Thinopyrum, Agropyron, and Hordeum are commonly used as bridges to transfer genes from some wild species to wheat (Fedak and Armstrong 1981; Jauhar 1992; Li and Dong 1993; Li et al. 2006; Kang et al. 2012, 2016). The objective of the present study was to transfer and characterize Th. intermedium ssp. trichophorum chromosomes to wheat using trigeneric hybridization involving a cross between an octoploid wheat–Thinopyrum partial amphiploid and hexaploid triticale. New wheat–Th. intermedium derivative lines displaying the blue grain character and resistance to stripe rust were identified by sequential FISH and molecular markers.
Materials and methods
Plant materials
The cross between an octoploid (2n = 8x = 56) wheat–Th. intermedium ssp. trichophorum partial amphiploid TE1508 (Song et al. 2013) as the female with hexaploid (2n = 6x = 42) triticale Currency as the male parent (Shu et al. 2000) was used to transfer Thinopyrum chromosomes to wheat. Lines X24C-14, X24C10, X24C5 and X24C1-1 were developed from the F6 progeny. Th. intermedium PI440028 (StJSJ genomes, 2n = 6x = 42) was obtained from the National Small Grains Collection at Aberdeen, Idaho, USA. Wheat–Th. intermedium addition lines L series from TAF46 and Z series from Zhong5 were obtained from Drs Bernd Friebe, Kansas State University, USA, and Zhijian Chang, Shanxi Academy of Agricultural Sciences, China, respectively.
Fluorescence in situ hybridization (FISH)
Seedling root tips of trigeneric hybrids derived lines and their parents were collected and treated with nitrous oxide followed by enzyme digestion, using the procedure of Han et al. (2006). The synthesized oligonucleotide probes Oligo-pSc119.2, Oligo-pTa535, Oligo-pTa71 and Oligo-(GAA)7 were used for identifying the wheat chromosomes according to the description of Tang et al. (2014). Probe Oligo-pSt122 is specific for terminal regions of Th. intermedium chromosomes (Li et al. 2016). Oligonucleotide probes were synthesized by Shanghai Invitrogen Biotechnology Co. Ltd. (Shanghai, China). The synthetic oligonucleotides were either 5′ end-labeled with 6-carboxyfluorescein (6-FAM) for green or 6-carboxytetramethylrhodamine (Tamra) for red signals. The protocol of non-denaturing FISH (ND-FISH) by the synthesized probes was described by Fu et al. (2015). After the oligo-based FISH, the sequential FISH with the long terminal repeat (LTR) pDb12H sequence can clearly distinguish the JS chromosomes in Th. intermedium (Liu et al. 2009). This LTR sequence was labeled with Alexa Fluor 488-5-dUTP (Invitrogen) according to Han et al. (2006). Photomicrographs of FISH chromosomes were taken with an Olympus BX-51 microscope equipped with a DP-70 CCD camera.
Molecular marker analysis
DNA was extracted from young leaves of Chinese Spring (CS), Th. intermedium, TE1508-1, lines X24C10, X2C14, X24C5, and X24C1-1. PCR-based Landmark Unique Gene (PLUG) primers were designed from rice genomic DNA sequences specific for the syntenic regions of the corresponding linkage group(s) of wheat genomes, which may amplify Triticeae genome fragments (Ishikawa et al. 2009). Polymerase chain reaction (PCR) was performed in an Icycler Thermal Cycler (Bio-Rad Laboratories, Emeryville, CA) in a 25 μl reaction, containing 10 mmol Tris–HCl (pH 8.3), 2.5 mmol MgCl2, 200 μmol of each dNTP, 100 ng template DNA, 0.2 U Taq polymerase (Takara, Japan) and 400 nmol of each primer. The cycling parameters were 94 °C for 3 min for denaturation, followed by 35 cycles at 94 °C for 1 min, 55 °C for 1 min, 72 °C for 2 min, and a final extension at 72 °C for 10 min. The amplified products were restriction enzyme digested and electrophoresis was conducted as described by Hu et al. (2012). The PLUG markers without chromosomal physical location in Ishikawa et al. (2009) were searched using the wheat genome database IWGSC WGA v0.4 from https://urgi.versailles.inra.fr/blast/, and confirmed by PCR.
Rust resistance tests
Agronomic traits under field conditions were studied at the Xindu Experimental Station, Chengdu, China during the 2013–2015 wheat growing seasons. Stripe rust (P. striiformis f. sp. Tritici, Pst) races, CYR32, CYR33 and v26 (Liu et al. 2010), were provided by the Crop Protection Institute, Sichuan Academy of Agricultural Sciences, China. All materials were tested for resistance to mixed isolates in the field. Infection type (IT) was scored 14–15 days after inoculation when rust was fully developed on the susceptible check line SY95-71 (Hu et al. 2011). The IT was recorded based on the 0–4 scale according to Bariana and McIntosh (1993).
Results
FISH karyotype analysis
Cytological analysis of the wheat–Th. intermedium ssp. trichophorum partial amphiploid TE1508 revealed two distinct variants of chromosome numbers, 56 and 54, the genotypes of which were subsequently named TE1508-1 and TE1508-2, respectively. The mitotic metaphase chromosomes of TE508-1 and TE1508-2 were hybridized with probes Oligo-pSc119.2, Oligo-pTa535, pDb12H, Oligo-(GAA)7 and Oligo-pSt122 by sequential multi-color-FISH (Fig. 1). The FISH hybridization signals of the probes Oligo-pSc119.2 and Oligo-pTa535 could easily identify the wheat chromosomes based on the standard FISH karyotype of wheat chromosomes described by Tang et al. (2014). Both TE1508-1 and TE1508-2 contained only 40 wheat chromosomes with the absence of chromosome 4B (Fig. 1a, c). The difference between TE1508-1 and TE1508-2 was in the number of Th. intermedium chromosomes. Line TE1508-1 contained eight pairs of Thinopyrum chromosomes and TE1508-2 had seven pairs of Th. intermedium chromosomes. The sequential FISH with LTR probe pDb12H on TE1508-1 revealed that three pairs of chromosomes belonged to the JS group, and were named JS-1 to JS-3. In comparison, TE1508-2 had two pairs of JS genome chromosomes but lacked JS-3 chromosomes (Fig. 1c, d). We have previously developed Th. intermedium chromosome 1St in line AS1677 (Hu et al. 2011), and identified the 6St and 7St chromosomes from the lines X482 (Li et al. 2015) and X484 (Song et al. 2013), respectively. The remaining two pairs of J-chromosomes were named J-1 and J-2 (Fig. 1). The FISH karyotype of eight pairs of Th. intermedium chromosomes of TE1508 is shown in Fig. 1e.
Sequential FISH patterns of wheat–Th. intermedium partial amphiploid TE1508-1 (a, b), TE1508-2 (c, d) and karyotype of Th. intermedium chromosomes in TE1508 (e). The probes for FISH (a, c) were Oligo-pSc119.2 (green) + Oligo-pTa535 (red), while the probes for sequential FISH (b) were pDb12H (green), and FISH (d) were Oligo-(GAA)7 (green) + Oligo-pSt122 (red), respectively. The karyotype (e) of Th. intermedium chromosomes was shown by the probes at left with Oligo-pSt122 (green) + Oligo-pTa71 (red) and right with Oligo-pSc119.2 (green) + Oligo-pTa535 (red), while in the middle with pDb12H (green) added for JS chromosomes, respectively
The FISH probes Oligo-pSc119.2, Oligo-pTa535 and Oligo-1162 were used to identify the wheat and rye chromosomes in this triticale (Fu et al. 2015). As shown in Fig. 2a, the clear FISH patterns of individual rye chromosomes 1R to 7R can be distinguished from the 28 wheat chromosomes in Currency. The FISH patterns of rye chromosomes were different from the Thinopyrum chromosomes in TE1508. Therefore, the FISH probes enable the identification of the eight pairs of individual Th. intermedium chromosomes and seven pairs of rye chromosomes in a wheat background.
Transmission of Thinopyrum and rye chromosomes
The Oligo-based ND-FISH was able to identify the Thinopyrum and rye chromosomes in the trigeneric hybrids and their derived progenies. The chromosome compositions of the F4 to F6 progenies by FISH are shown in Table 1. Over 80% of F4 plants had five to 10 Th. intermedium chromosomes and one to six rye chromosomes including the telocentrics (Fig. 2b). In the F5, 50 plants contained two to six Th. intermedium chromosomes and zero to three rye chromosomes (Fig. 2c), and 42 plants had no rye chromosomes. In the F6, there were one to seven Th. intermedium chromosomes, but no rye chromosomes were detected (Fig. 2d), except two plants with a 1BL.1RS translocation. Thus, it seems that Th. intermedium chromosomes are more preferentially transmitted than rye chromosomes in the latter generations of selfed progenies. Among the different groups of Thinopyrum chromosomes, the JS chromosomes exhibited higher transmission rates than the J and St chromosomes. A total 36.8 and 34.3% of F4 to F6 plants contained at least one JS-2 chromosome and a 4BS.JS-3L translocation chromosome, respectively. Only 7.8 and 6.5% plants have the J-1 and J-2 chromosomes, respectively. Other wheat–Th. intermedium translocation chromosomes were also observed in about 1.2% of the plants.
Chromosome identification of stable wheat-trigeneric derived lines
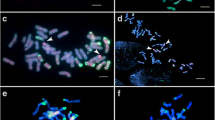
Sequential multi-color ND-FISH with probes Oligo-pSc119.2, Oligo-pTa535, and Oligo-(GAA)7 was conducted to characterize the cytologically stable lines X24C5, X24C10, X24C14 and X24C1-1 (Fig. 3). The chromosome number of X24C14 was 44, while X24C5, X24C10, and X24C1-1 all had 42 chromosomes. As shown in Fig. 3a, it was found that X24C14 includes all the 42 wheat chromosomes and two alien chromosomes. Probes Oligo-pSc119.2 and Oligo-pTa535 showed a pair of chromosomes with faint Oligo-pTa535 hybridization signals at the telomeric region of long arm, and faint hybridization signals of Oligo-pSc119.2 in the interstitial region of the short arm in X24C14 (Fig. 3a). The FISH hybridization pattern by pDb12H revealed that the added chromosomes belonged to the JS genome, and were identical to Th. intermedium chromosome JS-1 (Fig. 1e). Therefore, it was identified that the line X24C14 was a chromosome JS-1 addition line.
FISH patterns of wheat–Th. intermedium-derived lines X24C14 (a, b), X24C5 (c), X24C10 (d), and X24C1-1(e, f). The probes Oligo-pSc119.2 (green) + Oligo-pTa535 (red) were used in FISH (a, c, d, e), and the probes Oligo-(GAA)7 and pDb12H were used in sequential FISH (b, f). The arrows indicated the Thinopyrum chromosomes
Similarly, FISH with probes Oligo-pSc119.2 and Oligo-pTa535 showed that both X24C5 and X24C10 lack the 4B chromosomes of wheat (Fig. 3c, d). The comparison of the introduced alien chromosomes in X24C10 and X24C5 with TE1508 (Fig. 1e) revealed that lines X24C10 and X24C5 were JS-2(4B) and J-1(4B) substitution lines, respectively.
As shown in Fig. 3e, FISH with probes Oligo-pSc119.2 and Oligo-pTa535 revealed that line X24C1-1 contained a pair of translocated chromosomes. Based on sequential FISH with Oligo-(GAA)7 (Fig. 3f), it was found that the translocated chromosome involved the entire wheat arm 4BS and the distal portion of the long arm of the J-2 chromosome from Fraction Length (FL) 0.60 to the long arm telomere.
Molecular markers analysis
The PLUG markers were useful for developing alien chromosome-specific markers of Secale, Dasypyrum and Thinopyrum chromatin and to assign the alien chromosome to specific homoeologous linkage groups (Hu et al. 2011; Lei et al. 2012; Li et al. 2013). The chromosomal locations of most of the PLUG markers were based on the PCR amplifications as described in Ishikawa et al. (2009), and blasted to the wheat genome database “The IWGSC WGA v0.4” (Tables 2, 3). A total of 500 PLUG markers were used to test the CS, Th. intermedium and TE1508. A total of 285 markers gave rise to Th. intermedium-specific amplification identical to those in TE1508 after TaqI, HaeIII or HapII digestion of PCR products. A total of 31 pairs of TNAC primers from wheat homoeologous group 2 (Supplementary Table 1) generated the identical bands from Th. intermedium to those from the disomic addition line X24C14. These results suggested that the Th. intermedium chromosome in X24C14 belonged to homoeologous group 2. After combining these PCR results with the FISH patterns, we concluded that the X24C14 was a JS-1 addition line, and the JS-1 was chromosome 2JS.
The PLUG markers of wheat homoeologous group 4 (Table 2) also amplified the specific bands with DNA from both X24C5 and X24C10 (Fig. 4), indicating that they both contained Th. intermedium chromosomes which belong to group 4. A total of 38 of 42 markers from X24C5 produced Th. intermedium-specific polymorphic bands, while 26 markers amplified Th. intermedium-specific polymorphic bands from X24C10. Only six markers give identical Th. intermedium-specific bands for lines X24C5 and X24C10, indicating that these two group 4 Th. intermedium chromosomes were genetically different. In combination with the FISH results, it was indicated that line X24C5 was a 4JS(4B) substitution, while line X24C10 was a 4J(4B) substitution. The PLUG markers indicated that chromosome 4JS was more closely related to wheat group 4 chromosomes than was chromosome 4J.
The 4J polymorphic TNAC markers from line X24C10 were further used to amplify in line X24C1-1 (Table 2). Of the 13 4J polymorphic markers on arms 4BL and 4DL, seven markers mapped to the segment distal to FL0.6 and had identical amplification for X24C10 and X24C1-1, while six proximal markers showed no amplification in X24C1-1. Therefore, TNAC markers also confirmed the breakpoints in chromosome 4JL of X24C1-1, which is consistent with the FISH results (Fig. 5).
FISH patterns of chromosomes 4B, 4J and 4J.4B translocation for blue grain. The chromosomes 4B from Currency, 4J from X24C10, and 4J.4B location from X24C1-1. The probes for the chromosomes to the left were Oligo-pSc119.2 (green) + Oligo-pTa535 (red), and probe for the chromosomes to the right was Oligo-(GAA)7 (green). Arrow shows the putative position for the translocation breakpoint
Plant and grain characters
A set of agronomic traits were measured on each of the ten plants of Currency triticale, TE1508, and the stable wheat–Th. intermedium lines grown under field conditions (Table 3). Line X24C10 had significantly reduced plant height and an increase in the numbers of spikes, which possibly implies that the 4J chromosome carries strong dwarfing and tillering gene(s) expressed in the wheat background. The 1000-kernel weight was increased in X24C1-1 and thus probably contributed to a favorable effect on grain yield relative to controls in the 4JL lines.
Grain color of TE1508 displayed a light green shade, those of the triticale line Currency were red, while the wheat–Th. intermedium lines X24C10 and X24C1-1 were blue grained (Fig. 6). Based on the karyotypic comparison among the blue-grained lines, it was found that TE1508 and X24C10 contained chromosome 4J. Line X24C1-1 carrying the 4B-4J translocation also showed blue grain. Thus, we conclude that the blue grain is encoded by gene(s) located on the FL0.6–1.00 long arm of chromosome 4J (Fig. 5).
When challenged with CYR32, CYR33 and v26 races of stripe rust, the parents Currency and TE1508 showed resistance to stripe rust at adult plant stages. Lines X24C14, X24C10 and X24C5 were resistant, while X24C1-1 was susceptible. This indicates that the Th. intermedium chromosomes 2JS, 4J, and 4JS carry a gene(s) for stripe rust resistance. The wheat–Th. intermedium derivatives might be useful resource in wheat breeding programs.
Discussion
Crossing different amphiploids is an effective and rapid method for producing trigeneric hybrids, which offers the possibility of transferring several alien characters concurrently to cultivated wheat (Orellana et al. 1989).In the present study, we produced trigeneric hybrids between TE1508 and the triticale cultivar Currency. The results indicated that Th. intermedium chromosomes remained in the progenies, whereas rye chromosomes were lost quickly during selfing of the populations. Hence, we can conclude that the Th. intermedium genomes are probably more closely related to wheat genomes than is the rye genome to wheat. Moreover, the different transmission rates among JS, J and St-chromosomes in the offspring of trigeneric hybrids also possibly indicate that the JS-chromosomes may be more closely related to wheat than the other two genomes, which is also supported by the molecular-based evidence (Table 2). Due to distinctly different genetic divergences among the alien genomes and the wheat A, B, D genomes, the transmission rates of the different alien chromosomes led to variations in chromosome numbers in the progenies of trigeneric hybrids (Kosina and Heslop-Harrison 1996). The chromosome rearrangements and intergenomic translocations of wheat and those of alien origin have been previously observed in different trigeneric amphiploid hybrids (Kang et al. 2012, 2016). We also produced translocation lines between JS and J chromosome and wheat chromosomes in the trigeneric hybrids progenies described here. The lines carry only a small number of alien genes into wheat, which may be of some use to breeders for wheat improvement.
Chromosome painting has been used to identify alien chromosomes, including chromosome number and characterizing structural aberrations in wheat-alien hybrids (Jiang and Gill 1993). Considering that the inter- and intra-genomic chromosomal rearrangements also occur commonly in Th. intermedium and the wheat–Th. intermedium derivatives (Chen et al. 1999; Yang et al. 2006; Deng et al. 2013; Li et al. 2015), identification of individual chromosomes of the complex Th. intermedium was considered also crucial. The genomic in situ hybridization (GISH) technique has been employed to characterize genomes and chromosomes in polyploid Thinopyrum species (Chen et al. 1998; Tang et al. 2000; Mahelka et al. 2011). The multi-color GISH (mcGISH) approach has also been sequentially performed to identify the Thinopyrum chromosomes in wheat background (Han et al. 2003; Kruppa and Molnár-Láng 2016). The complete hybridization signals of the St genome chromosomes were recently widely accepted by researchers, since they are stable in different Th. intermedium accessions. We recently isolated a genome-specific long terminal repeat (LTR) sequence and were able to discriminate between the JS and J genomes by FISH (Liu et al. 2009; Tang et al. 2011). We further carried out analysis of 11 globally distributed Th. intermedium accessions by FISH using the probe pDb12H, and found that the number of JS-chromosomes was constant 14 chromosomes. We thus concluded that the FISH hybridization signals of pDb12H were specific to the JS-chromosome of Th. intermedium. ND-FISH using the probes Oligo-pSc119.2 and Oligo-pTa535 can enable construction of the karyotype of several Th. intermedium chromosomes in the wheat background (Fig. 1e). We recently generated a terminal repetitive probe Oligo-pSt122, which can also assist in distinguishing individual Th. intermedium chromosomes in the wheat background (Li et al. 2016). In the present study, the wheat–Th. intermedium partial amphiploid TE1508 was identified as having unique Th. intermedium ssp. trichophorum JS- and J-chromosomes, which can be used to compare the different origins of Th. intermedium chromosomes transferred to wheat.
The chromosomes from Thinopyrum species have been successfully introduced into wheat to develop blue-grained germplasm (Li et al. 1986; Zeven 1991; Zhang et al. 2002). The disomic substitution lines consisting of a pair of wheat 4D chromosomes replaced by a pair of Th. ponticum 4Ag chromosomes revealed that a gene conferring blue grain is located on this Thinopyrum chromosome (Li et al. 1986). Zheng et al. (2006) physically mapped the blue grain gene to the region of FL0.71–0.80 on the long arm of chromosome 4Ag. A blue-grained gene in Th. bessarabicum was located on chromosome 4J (Zhang et al. 2002), and was recently located to the region between the centromere and FL0.52 on long arm of chromosome 4J (Shen et al. 2013). Whelan (1989) first reported the grain of Th. intermedium ssp. trichophorum (formerly named Agropyron trichophorum) displayed the blue seeds color. In the present study, we found that the wheat–Th. intermedium partial amphiploid TE1508 exhibited blue grains (Fig. 6). Line X24C10 contained a Th. intermedium J-chromosome substituted for a pair of 4B chromosomes and displayed the blue grain phenotype (Fig. 3c). Molecular marker analysis in the present study also revealed that the J-chromosome responsible for blue grain belonged to linkage group 4. Moreover, we located the blue grain on the long arm of 4J based on the translocation of 4BS.4JL, and further located on the terminal region of 4JL at the FL0.60–1.00 based on the small segment translocation in X24C1-1 by FISH (Fig. 5).
Thinopyrum intermedium homoeologous group 4 chromosomes have been known to possess novel sources of resistance to both eyespot (Tapesia yallundae) and WSMV (Wheat Streak Mosaic Virus) for a long time. Chromosome 4Ai#2 carrying the Wsm1 gene was further designated as 4JS-chromosome by St-based GISH (Li et al. 2004). Other gene(s) for resistance to WSMV were also identified on the long arm of chromosome 4Ai#3 (J-chromosome) from Th. intermedium (Friebe et al. 1996; Li et al. 2005). We observed that the addition line L4 with a 4Ai#1 chromosome (St-genome) was susceptible to stripe rust. In the present study, we determined that the substitution line X24C5 contained the 4JS chromosomes originally from Th. intermedium ssp. trichophorum. We observed that the terminal region of the original short arm of 4JS of X24C5 contained the strong signals with Oligo-pSc119.2 and Oligo-pSt122, which were absent in 4JS-St chromosomes of line X482 previously described by Li et al. (2016) (Supplementary Fig. 1). The distribution of pDb12H hybridization signals confirmed that the 4JS-St chromosome had been involved in a translocation event resulting in loss of about 40% of 4JS short arm, which was substituted by a 4St-chromosome, as described by Li et al. (2015). In the field-based stripe rust resistance study, line X482 showed high susceptibility to stripe rust, while X24C5 showed high levels of resistance at the adult plant stage. We, therefore, assume that it is possible that the stripe rust resistance gene in 4JS may be located in the distal region of its short arm at about FL0.60–1.00. Furthermore, we found that the number of polymorphic bands associated with chromosome 4JS when compared to the wheat 4A and 4D wheat chromosomes was fewer than lines carrying the 4J chromosome. Therefore, chromosome 4JS may represent a promising candidate for future homoeologous recombination studies with 4A or 4D.
Three disomic chromosome addition lines, Z1, Z2, and Z6 derived from Zhong 5 and containing chromosome 2Ai#2 from Th. intermedium, also consistently showed high levels of resistance to barley yellow dwarf virus (BYDV) (Xin et al. 1988; Larkin et al. 1995). Chromosome 2Ai#2 is an St-JS translocation in which majority contains St chromatin and the partial long arm of a JS chromosome (Tang et al. 2000; Chen et al. 2003). The BYDV resistance gene may locate on the St-segment of 2Ai#2 (Wang et al. 2010). The breakpoint of St-JS of chromosome 2Ai#2 of Z2 was possibly at the FL0.40 of long arm based on the hybridization with probe pDb12H (Supplementary Fig. 2). The present study identified the Th. intermedium 2JS chromosome in X24C14 and characterized that it was a complete JS chromosome according to the FISH and molecular marker analyses. The FISH patterns with the terminal repetitive probe Oligo-pSt122 showed that the 2Ai#2 chromosome has two distinct hybridization signals in both terminal regions (Supplementary Fig. 2); however, the Oligo-pSt122 did not have any hybridization signal in 2JS of X24C14. Therefore, we have convincingly shown that the chromosomes of the different Th. intermedium-derived germplasm can be distinguished by FISH. We found that line X24C14 was resistant to stripe rust, and line Z2 was also responsible for novel rust resistance (Tang et al. 2000). This material is, therefore, considered to be essential for the future studies for transferring novel rust genes from linkage group 2 of Th. intermedium into wheat (Hou et al. 2016).
Author contribution statement
ZY and GL designed the experiments. JL, BL, HW and ZY performed the experiments, TL and EY analysis the data, ZY and GL wrote the paper.
References
Bao YG, Wu X, Zhang C, Li XF, He F, Qi XL, Wang HG (2014) Chromosomal constitutions and reactions to powdery mildew and stripe rust of four novel wheat–Thinopyrum intermedium partial amphiploids. J Genet Genomics 12:663–666
Bariana HS, McIntosh RA (1993) Cytogenetic studies in wheat. XV. Location of rust resistance genes in VPM1 and their genetic linkage with other disease resistance genes in chromosome 2A. Genome 36:476–482
Cauderon Y, Saigne B, Dauge M (1973) The resistance to wheat rusts of Agropyron intermedium and its use in wheat improvement. In: Sears ER, Sears LMS (eds) Proceedings of the 4th international wheat genetics symposium, University of Missouri, Columbia, pp 401–407
Chen Q (2005) Detection of alien chromatin introgression from Thinopyrum into wheat using S genomic DNA as a probe—a landmark approach for Thinopyrum genome research. Cytogenet Genome Res 109:350–359
Chen Q, Conner R, Laroche A, Thomas J (1998) Genome analysis of Thinopyrum intermedium and Thinopyrum ponticum using genomic in situ hybridization. Genome 41:580–586
Chen Q, Conner RL, Laroche A, Ji W, Armstrong KC, Fedak G (1999) Genomic in situ hybridization analysis of Thinopyrum chromatin in a wheat–Th. intermedium partial amphiploid and six derived chromosome addition lines. Genome 42:1217–1223
Chen Q, Conner RL, Li HJ, Sun SC, Ahmad F, Laroche A, Graf RJ (2003) Molecular cytogenetic discrimination and reaction to wheat streak mosaic virus and the wheat curl mite in Zhong series of wheat–Thinopyrum intermedium partial amphiploids. Genome 46:135–145
Deng CL, Bai LL, Fu SL, Yin WB, Zhang YX, Chen YH, Wang RRC, Zhang XQ, Han FP, Hu ZM (2013) Microdissection and chromosome painting of the alien chromosome in an addition line of wheat–Thinopyrum intermedium. PLoS One 8:e72564
Dewey DR (1984) The genomic system of classification as a guide to intergeneric hybridization in the perennial Triticeae. In: Gustafson JP (ed) Gene manipulation in plant improvement. Plenum, New York, pp 209–279
Fedak G, Armstrong KC (1981) Cytogenetics of the trigeneric hybrid, (Hordeum vulgare × Triticum aestivum) × Secale cereale. Theor Appl Genet 60:215–219
Fedak G, Chen Q, Conner RL, Laroche A, Petroski R, Armstrong KW (2000) Characterization of wheat–Thinopyrum partial amphiploids by meiotic analysis and genomic in situ hybridization. Genome 43:712–719
Friebe B, Jiang J, Gill BS, Dyck PL (1993) Radiation-induced nonhomoeologous wheat–Agropyron intermedium chromosomal translocations conferring resistance to leaf rust. Theor Appl Genet 86:141–149
Friebe B, Gill KS, Tuleen NA, Gill BS (1996) Transfer of wheat streak mosaic virus resistance from Agropyron intermedium. Crop Sci 36:857–861
Fu S, Chen L, Wang Y, Li M, Yang Z, Qiu L, Yan B, Ren Z, Tang Z (2015) Oligonucleotide probes for ND-FISH analysis to identify rye and wheat chromosomes. Sci Rep 5:10552
Han FP, Fedak G, Benabdelmouna A, Armstrong K, Ouellet T (2003) Characterization of six wheat × Thinopyrum intermedium derivatives by GISH, RFLP, and multicolor GISH. Genome 46:490–495
Han FP, Lamb JC, Birchler JA (2006) High frequency of centromere inactivation resulting in stable dicentric chromosomes of maize. Proc Natl Acad Sci USA 103:3238–3243
He R, Chang Z, Yang Z, Liu Z, Zhan H, Zhang X, Liu J (2009) Inheritance and mapping of a powdery mildew resistance gene Pm43 introgressed from Thinopyrum intermedium into wheat. Theor Appl Genet 118:1173–1180
Hou L, Jia J, Zhang X, Li X, Yang Z, Ma J, Guo H, Zhan H, Qiao L, Chang Z (2016) Molecular mapping of stripe rust resistance gene Yr69 in wheat chromosome 2AS. Plant Dis 100:1717–1724
Hu LJ, Li GR, Zeng ZX, Chang ZJ, Liu C, Zhou JP, Yang ZJ (2011) Molecular cytogenetic identification of a new wheat–Thinopyrum substitution line with stripe rust resistance. Euphytica 177:169–177
Hu LJ, Zeng ZX, Liu C, Li GR, Yang ZJ (2012) Isolation and chromosomal localization of new MITE-like sequences from Secale. Biologia 67:126–131
Ishikawa G, Nakamura T, Ashida T, Saito M, Nasuda S, Endo T, Wu J, Matsumoto T (2009) Localization of anchor loci representing five hundred annotated rice genes to wheat chromosomes using PLUG markers. Theor Appl Genet 118:499–514
Jauhar PP (1992) Synthesis and cytological characterization of trigeneric hybrids involving durum wheat, Thinopyrum bessarabicum, and Lophopyrum elongatum. Theor Appl Genet 84:511–519
Jiang J, Gill BS (1993) Sequential chromosome banding and in situ hybridization analysis. Genome 36:792–795
Kang H, Zeng J, Xie Q, Tao S, Zhong M, Zhang H, Fan X, Sha L, Xu L, Zhou Y (2012) Molecular cytogenetic characterization and stripe rust response of a trigeneric hybrid involving Triticum, Psathyrostachys, and Thinopyrum. Genome 55:383–390
Kang HY, Huang J, Zhu W, Li DY, Diao CD, Tang L, Wang Y, Xu LL, Zeng J, Fan X, Sha LN, Zhang HQ, Zheng YL, Zhou YH (2016) Cytogenetic behavior of trigeneric hybrid progeny involving wheat, rye and Psathyrostachys huashanica. Cytogenet Genome Res 148:74–82
Kantarski T, Larson S, Zhang X, DeHaan L, Borevitz J, Anderson J, Poland J (2017) Development of the first consensus genetic map of intermediate wheatgrass (Thinopyrum intermedium) using genotyping-by-sequencing. Theor Appl Genet 130:137–150
Kosina R, Heslop-Harrison JS (1996) Molecular cytogenetics of an amphiploid trigeneric hybrid between Triticum durum, Thinopyrum distichum and Lophopyrum elongatum. Ann Bot (Lond) 78:583–589
Kruppa K, Molnár-Láng M (2016) Simultaneous visualization of different genomes (J, JSt and St) in a Thinopyrum intermedium × Thinopyrum ponticum synthetic hybrid (Poaceae) and in its parental species by multicolour genomic in situ hybridization (mcGISH). Comp Cytogenet 10:283–293
Larkin PJ, Banks PM, Lagudah ES, Apple R, Chen X, Xin ZY, Ohm HW, McIntosh RA (1995) Disomic Thinopyrum intermedium addition lines in wheat with barley yellow dwarf virus resistance and with rust resistances. Genome 38:385–394
Lei MP, Li GR, Liu C, Yang ZJ (2012) Characterization of new wheat–Secale africanum derivatives reveals evolutionary aspects of chromosome 1R in rye. Genome 55:765–774
Li LH, Dong YS (1993) A self-fertile trigeneric hybrid, Triticum aestivum × Agropyron michnoi × Secale cereale. Theor Appl Genet 87:361–368
Li H, Wang X (2009) Thinopyrum ponticum and the promising source of resistance to fungal and viral diseases of wheat. J Genet Genomics 36:557–565
Li ZS, Mu SM, Zhou HP, Wu J (1986) The establishment and application of blue-grained monosomics in wheat chromosome engineering. Cereal Res Commun 14:133–137
Li HJ, Conner RL, Chen Q, Graf RJ, Laroche A, Ahmad F, Kuzyk AD (2004) Promising genetic resources for resistance to wheat streak mosaic virus and the wheat curl mite in wheat–Thinopyrum partial amphiploids and their derivatives. Genet Resour Crop Evol 51:827–835
Li HJ, Arterburn M, Jones SS, Murray TD (2005) Resistance to eyespot of wheat, caused by Tapesia yallundae, derived from Thinopyrum intermedium homoeologous group 4 chromosomes. Theor Appl Genet 111:932–940
Li XF, Song ZQ, Liu SB, Gao JR, Wang HG (2006) Cytogenetic study of a trigeneric (triticale × tritileymus) hybrid. Euphytica 150:117–122
Li GR, Liu C, Li CH, Zhao JM, Zhou L, Dai G, Yang EN, Yang ZJ (2013) Introgression of a novel Thinopyrum intermedium St-chromosome-specific HMW-GS gene into wheat. Mol Breed 31:843–853
Li G, Lang T, Dai G, Li D, Li C, Song X, Yang Z (2015) Precise identification of two wheat–Thinopyrum intermedium substitutions reveals the compensation and rearrangement between wheat and Thinopyrum chromosomes. Mol Breed 35:1
Li G, Wang H, Lang T, Li J, La S, Yang E, Yang Z (2016) New molecular markers and cytogenetic probes enable chromosome identification of wheat–Thinopyrum intermedium introgression lines for improving grain quality. Planta 244:865–876
Liu C, Yang ZJ, Jia JQ, Li GR, Zhou JP, Ren ZL (2009) Genomic distribution of a long terminal repeat (LTR) Sabrina-like retrotransposon in Triticeae species. Cereal Res Commun 37:363–372
Liu GT, Peng YL, Chen WQ, Zhang ZY (2010) First detection of virulence in Puccinia striiformis f. sp. tritici in China to resistance genes Yr24 (= Yr26) present in wheat cultivar Chuanmai 42. Plant Dis 94:1163
Liu J, Chang Z, Zhang X, Yang Z, Li X, Jia J, Zhan H, Guo H, Wang J (2013) Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor Appl Genet 126:265–274
Luo PG, Hu XY, Chang ZJ, Zhang M, Zhang HQ, Ren ZL (2009) A new stripe rust resistance gene transferred from Thinopyrum intermedium to hexaploid wheat (Triticum aestivum). Phytoprotection 90:57–63
Mahelka V, Kopecky D, Pastova L (2011) On the genome constitution and evolution of intermediate wheatgrass (Thinopyrum intermedium: Poaceae, Triticeae). BMC Evol Biol 11:127
Mahelka V, Kopecky D, Baum BR (2013) Contrasting patterns of evolution of 45S and 5S rDNA families uncover new aspects in the genome constitution of the agronomically important grass Thinopyrum intermedium (Triticeae). Mol Biol Evol 30:2065–2086
Orellana J, Vazquez JF, Carrillo JM (1989) Genome analysis in wheat–rye–Aegilops caudata trigeneric hybrids. Genome 32:169–172
Shen Y, Shen J, Dawadondup Zhuang L, Wang Y, Pu J, Feng Y, Chu C, Wang X, Qi Z (2013) Physical localization of a novel blue-grained gene derived from Thinopyrum bessarabicum. Mol Breed 31:195–204
Shu HL, Yang ZJ, Li GR (2000) Genetic analysis of several wheat–rye introgression lines. J Sichuan Agric Univ 18:223–227
Song XJ, Li GR, Zhan HX, Liu C, Yang ZJ (2013) Molecular identification of a new wheat–Thinopyrum intermedium ssp. trichophorum addition line for resistance to stripe rust. Cereal Res Commun 41:211–220
Tang S, Li Z, Jia X, Larkin PJ (2000) Genomic in situ hybridization (GISH) analyses of Thinopyrum intermedium, its partial amphiploid Zhong 5, and disease-resistant derivatives in wheat. Theor Appl Genet 100:344–352
Tang ZX, Yang ZJ, Fu SL, Yang MY, Li GR, Zhang HQ, Tan FQ, Ren Z (2011) A new long terminal repeat (LTR) sequence allows the identify J genome from Js and St genomes of Thinopyrum intermedium. J Appl Genet 52:31–33
Tang ZX, Yang ZJ, Fu SL (2014) Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J Appl Genet 55:313–318
Wang RRC (2011) Agropyron and Psathyrostachys. In: Kole C (ed) Wild crop relatives: genomic and breeding resources, cereals. Springer, Berlin
Wang Y, Wang H (2016) Characterization of three novel wheat–Thinopyrum intermedium addition lines with novel storage protein subunits and resistance to both powdery mildew and stripe rust. J Genet Genomics 43:45–48
Wang MJ, Zhang Y, Lin ZS, Ye XG, Yuan YP, Ma W, Xin ZY (2010) Development of EST-PCR markers for Thinopyrum intermedium chromosome 2Ai#2 and their application in characterization of novel wheat-grass recombinants. Theor Appl Genet 121:1369–1380
Whelan EDP (1989) Transmission of an alien telocentric addition chromosome in common wheat that confers blue seed color. Genome 32:30–34
Xin ZY, Brettell RIS, Cheng EM, Waterhouse PM, Appels R, Banks PM, Zhou GH, Chen X, Larkin PJ (1988) Characterization of a potential source of barley yellow dwarf virus resistance for wheat. Genome 30:250–257
Xu J, Conner RL (1994) Intravarietal variation in satellites and C-banded chromosomes of Agropyron intermedium ssp. trichophorum cv. Greenleaf. Genome 37:305–310
Xu SS, Jin Y, Klindworth DL, Wang RRC, Cai X (2009) Evaluation and characterization of seedling resistance to stem rust Ug99 races in wheat-alien species derivatives. Crop Sci 49:2167–2175
Yang ZJ, Li GR, Chang ZJ, Zhou JP, Ren ZL (2006) Characterization of a partial amphiploid between Triticum aestivum cv. Chinese Spring and Thinopyrum intermedium ssp. trichophorum. Euphytica 149:11–17
Zeven AC (1991) Wheats with purple and blue grains: a review. Euphytica 56:243–258
Zhang JY, Li XM, Wang RRC, Cortes A, Rosas V, Mujeeb-Kazi A (2002) Molecular cytogenetic characterization of Eb-genome chromosomes in Thinopyrum bessarabicum disomic addition lines of bread wheat. Plant Sci 163:167–174
Zheng Q, Li B, Mu SM, Zhou HP, Li ZS (2006) Physical mapping of the blue-grained gene(s) from Thinopyrum ponticum by GISH and FISH in a set of translocation lines with different seed colors in wheat. Genome 49:1109–1114
Acknowledgements
We gratefully thank Drs I. Dundas at the University of Adelaide, and Dr. Peng Zhang at University of Sydney, Australia for reviewing the manuscript, and to the support of The National Key Research and Development Program of China (2016YFD0102000), Basic and Applied Project from Department of Science and Technology, Sichuan (2016JY0075), and National Natural Science Foundation of China (31171542).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2017_2669_MOESM1_ESM.jpg
Supplementary material 1 Chromosome comparison of 4JS from X24C5 and 4JS-St from X482. Arrow shows the putative position for the St-JS translocation breakpoint. Left showed the signals of Oligo-pSc119.2 (green) + Oligo-pTa535 (red), middle showed the signals of pDb12H (green), and right showed the signals of Oligo-pSt122 (green) (JPEG 39 kb)
425_2017_2669_MOESM2_ESM.jpg
Supplementary material 2 FISH patterns of Th. intermedium chromosomes in Z2 and X24C14. Arrow indicates the possible position of the breakpoint. Left chromosomes showed the signals of pDb12H (green), middle showed the signals of Oligo-pSc119.2 (green) + Oligo-pTa535 (red), and right indicated the signals of Oligo-pSt122 (green) (JPEG 31 kb)
Rights and permissions
About this article
Cite this article
Li, J., Lang, T., Li, B. et al. Introduction of Thinopyrum intermedium ssp. trichophorum chromosomes to wheat by trigeneric hybridization involving Triticum, Secale and Thinopyrum genera. Planta 245, 1121–1135 (2017). https://doi.org/10.1007/s00425-017-2669-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2669-9