Abstract
Main conclusion
We projected meta-QTL (MQTL) for drought, salinity, and waterlogging tolerance to the physical map of barley through meta-analysis. The positions of these MQTL were refined and candidate genes were identified.
Drought, salinity and waterlogging are three major abiotic stresses limiting barley yield worldwide. Breeding for abiotic stress-tolerant crops has drawn increased attention, and a large number of quantitative trait loci (QTL) for drought, salinity, and waterlogging tolerance in barley have been detected. However, very few QTL have been successfully used in marker-assisted selection (MAS) in breeding. In this study, we summarized 632 QTL for drought, salinity and waterlogging tolerance in barley. Among all these QTL, only 195 major QTL were used to conduct meta-analysis to refine QTL positions for MAS. Meta-analysis was used to map the summarized major QTL for drought, salinity, and waterlogging tolerance from different mapping populations on the barley physical map. The positions of identified meta-QTL (MQTL) were used to search for candidate genes for drought, salinity, and waterlogging tolerance in barley. Both MQTL3H.4 and MQTL6H.2 control drought tolerance in barley. Fine-mapped QTL for salinity tolerance, HvNax4 and HvNax3, were validated on MQTL1H.4 and MQTL7H.2, respectively. MQTL2H.1 and MQTL5H.3 were also the target regions for improving salinity tolerance in barley. MQTL4H.4 is the main region controlling waterlogging tolerance in barley with fine-mapped QTL for aerenchyma formation under waterlogging conditions. Detected and refined MQTL and candidate genes are crucial for future successful MAS in barley breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Continued crop improvement is of paramount importance for feeding an increasing human population. Global breeding efforts over the past century have made significant contributions to increased yield potential and stability, as well as cultivars with more durable levels of tolerance to a diverse array of abiotic (drought, freezing, salinity and waterlogging) stresses (Khush 2001). Breeding crops that are tolerant to abiotic stresses is still the best approach to increase crop production (Gill and Tuteja 2010; Tester and Langridge 2010).
Quantitative trait loci (QTL) analysis is a powerful tool in agriculture and other fields. It provides knowledge of the chromosomal location of the target loci and can be applied in breeding programs using marker-assisted selection (MAS). Molecular markers linked to specific QTL have provided plant breeders with a method to improve selecting desirable recombinants from superior varieties and accelerating breeding programs (Khush 2001). MAS, combined with conventional breeding, has been utilized in many parts of the world and on many crops (Singh et al. 2009).
The number of publications reporting the identification of new QTL has been increasing tremendously during the past two decades, involving many crop plants and all types of agronomic traits (Xu and Crouch 2008). However, reports of QTL mapping, to date, are mostly based on a relatively low amount of markers, providing limited marker–trait association; and few of the QTL reported have been efficiently used for MAS in plant breeding (William et al. 2007). Many QTL could be identified for one trait, but most of them explain a small proportion of phenotypic variances of the traits (Tuberosa 2012). Therefore, plant breeding programs have not been able to take full advantage of these QTL (Eagles et al. 2001; Xu and Crouch 2008). Positional cloning (DNA sequence identification) of the QTL that explains more than 15 % phenotypic variance can greatly increase the effectiveness of using MAS in breeding programs (Salvi and Tuberosa 2005).
Drought, salinity and waterlogging are three major abiotic stresses limiting the yield of crops, causing extensive losses worldwide (Mittler 2006; Qin et al. 2011). Numerous QTL for drought, salinity and waterlogging tolerance in barley have been described. A meta-analysis can be used to combine different experimental results in one single study. At the QTL level, meta-analysis is able to map the QTL on the same linkage group from different mapping populations of different traits and lower the confidence of interval of QTL to identify more effective candidate genes (Goffinet and Gerber 2000). So far, meta-analysis has been successfully used in studying QTL for flowering time in maize (Chardon et al. 2004; Wang et al. 2016b), drought tolerance in rice (Khowaja et al. 2009), agronomic traits in cotton (Said et al. 2015), leaf senescence in Arabidopsis (Chardon et al. 2014) and yield-related traits in wheat (Zhang et al. 2010).
In this study, we summarized 632 QTL for drought, salinity and waterlogging tolerance in barley. Among all these QTL, only 195 major QTL were used to perform meta-analysis to refine QTL positions for MAS. We also identified candidate genes for each of the meta-QTL. Identified meta-QTL from meta-analysis provide resources for further MAS and various omics studies.
Materials and methods
Development of databases
Overall, 632 QTL identified from 1994 to 2015 for drought, salinity and waterlogging tolerance from 32 peer-reviewed publications were summarized in barley (Supplementary Table S1). Each QTL represents QTL for different traits from different studies with some of them being located in similar positions. Major QTL with the LOD value above 3 and the value of phenotypic variance exceeding 10 % were selected for the meta-analysis, as only QTL with these qualities can potentially be used in MAS and positional cloning (Collard et al. 2005). Parameters under control conditions are able to provide the tolerance coefficients caused by stresses with the relative changes of parameters (stressed/control). Therefore, many QTL for the tolerance coefficients and QTL for traits under control conditions are reported. We included QTL for the tolerance coefficients, but excluded QTL for the traits under control conditions. Although plant architecture traits, such as reduced height, increased number of tillers and erect leaves were also reported to be effective in breeding under control conditions (Khush 2001), these QTL are not relevant to the present investigation. Therefore, we reduced the number of QTL to 195 (Supplementary Table S2). All of these 195 major QTL were used for meta-analysis.
Consensus map and QTL projection
The physical map of barley was used as the consensus map in this study (http://barleygenomeapplications.com/default_2.aspx). BioMercator V4.2 (Arcade et al. 2004) (https://urgi.versailles.inra.fr/Tools/BioMercator-V4) was used to project QTL and refine QTL positions from different populations and studies onto one consensus map. The projection of QTL on barley physical map was based on LOD scores, phenotypic variation explained by each QTL, confidence intervals and QTL positions. The positions of the 195 major QTL were based on the positions of flanking markers on the consensus map. In terms of markers without physical positions, the closest markers of the QTL flanking markers from the reference were used to project QTL on the physical map. For those QTL lacking flanking markers and confidence intervals, positions of the closest markers to these QTL were selected as the positions of QTL on the reference map. A 95 % confidence interval was calculated based on the approach: confidence interval = 530/N × R 2 (Darvasi and Soller 1997), where N was the population size and R 2 was the proportion of phenotypic variance of QTL.
Meta-analysis of QTL
A meta-QTL is an integrated QTL resulting from several experiments. It is the “actual” QTL location underlying the distribution of the observed QTL on the genome (Goffinet and Gerber 2000). Meta-analysis was conducted with BioMercator V4.2, including algorithms from the MetaQTL software (Arcade et al. 2004; Veyrieras et al. 2007; Goffinet and Gerber 2000; Sosnowski et al. 2012) (https://urgi.versailles.inra.fr/Tools/BioMercator-V4). Meta-analysis first determined the number of meta-QTL (MQTL) in the physical map on each chromosome from different experiments based on AIC (Akaike information content), AICc (AIC correction), AIC3 (AIC 3 candidate models), BIC (Bayesian information criterion) and AWE (average weight of evidence). The number was considered the best fit to carry out meta-analysis when the values of the model selection criteria were the lowest in at least three of the five models (Chardon et al. 2014). Calculated QTL from the optimum model are regarded as the meta-QTL (MQTL) (Goffinet and Gerber 2000). The positions and 95 % confidence intervals of each MQTL were calculated.
Searching for candidate genes
The confidence intervals (cM) of identified MQTL on the barley physical map were used to search for the candidate genes in barley on the website (http://barleygenomeapplications.com/default_2.aspx) with ‘annotated gene’ tool.
Results
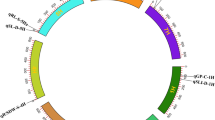
The 195 major QTL for abiotic stress tolerance were projected on different chromosomes (Fig. 1; Table 1). Chromosome 2H had the largest number of major QTL (55) and chromosome 6H had the least number of major QTL (15) for abiotic stress tolerance. There were 72 major QTL for drought tolerance, 70 major QTL for salinity tolerance, 48 major QTL for waterlogging tolerance, and 5 major QTL for combined salinity and waterlogging tolerance in barley. Each chromosome had at least seven major QTL for drought tolerance. Most major QTL for salinity tolerance were on chromosomes 2H (23) and 5H (21). In terms of waterlogging tolerance, chromosome 2H had the most number of major QTL (15).
Summarized major QTL for abiotic stress tolerance [drought (red), salinity (green), waterlogging (blue), combined salinity and waterlogging (yellow), and calculated MQTL (black)] on the barley physical map. Common used markers and genetic distance (cM) are shown on the right of chromosomes. The dotted line on the left of chromosome is the density curve of QTL on each chromosome (Chardon et al. 2004)
A total of 37 MQTL (~19 %) of the initial 195 major QTL for abiotic stress tolerance were detected based on meta-analysis (Fig. 1; Table 2). Apart from chromosome 5H, all the other chromosomes showed the peaks of density curve (Fig. 1), suggesting the target regions to improve abiotic stress tolerance in barley. There were six MQTL on chromosome 2H, with 53 initial major QTL. Each MQTL on chromosome 2H was formed with at least three initial QTL. Only three MQTL were detected on chromosome 6H. Among all the 37 identified MQTL, two MQTL were formed with QTL from six different populations and four MQTL were formed with QTL from five different populations. The QTL from different populations appeared to be unique. Meta-analysis also reduces the confidence intervals of MQTL from original 18.7 cM on average to 5.5 cM on average of each MQTL. Each MQTL had an average of 112 candidate genes (Supplementary Table S3) based on the physical positions of MQTL (Table 2). MQTL6H.1 had the lowest confidence interval of 0.1 cM (38.1–38.2 cM on chromosome 6H), resulting in no candidate genes on MQTL6H.1. No candidate genes were found on MQTL2H.6 and MQTL7H.5 due to confidence intervals of less than 1.5 cM. There were more than 600 candidate genes on MQTL3H.2 and MQTL4H.2.
Discussion
Drought tolerance in barley
Among all the abiotic stresses limiting crop yield, drought is one of the most important in agriculture, and breeders have made great efforts trying to improve drought tolerance in crops (Cattivelli et al. 2008; Tuberosa and Salvi 2006). Drought is a complex quantitative trait, controlled by many genes and numerous physiological mechanisms, such as early flowering time, plant height, higher K+ contents and osmotic adjustment (Cattivelli et al. 2008; Shabala and Pottosin 2014). Accurate phenotyping of drought tolerance remains the challenge for plant breeders to select drought-tolerant genotypes (Hu and Xiong 2014; Tuberosa 2012). Different traits have been used to identify drought-tolerance QTL (Supplementary Table S2). These traits include late leaf senescence (Guo et al. 2008; Sayed et al. 2012), root system (Chen et al. 2010), osmotic adjustment (Diab et al. 2004), relative water content (Teulat et al. 2003) and yield-related traits (Korff et al. 2008).
MQTL1H.4 were formed with five initial QTL for drought tolerance as shown in two studies (Korff et al. 2008; Sayed et al. 2012). The traits used as tolerance criteria include wilting score (Sayed et al. 2012), heading date, early vigour, days of maturity and days of grain-filling period. All of these traits were positively correlated with yield (Korff et al. 2008). Early flowering has been regarded as an effective trait to improve drought tolerance (Blum 2005; Salvi and Tuberosa 2005), escaping drought stress during flowering stage (Tuberosa 2012). Meta-analysis of flowering traits also refined positions of QTL in maize (Chardon et al. 2004).
On MQTL1H.4, totally 30 candidate genes were identified. Putative ATP-dependent Clp protease ATP-binding subunit ClpX1 (CLPX) and cytocrome P450 family protein were both expressed in drought-susceptible rice (Rabello et al. 2008). Overexpression of lipid transfer protein 3 enhanced drought tolerance in Arabidopsis (Guo et al. 2013).
MQTL3H.3 included four QTL for drought tolerance from two different studies (Fan et al. 2015; Korff et al. 2008) with three being based on agronomic traits, i.e., plant height, peduncle length and peduncle extrusion (Korff et al. 2008). MQTL3H.3 included 51 candidate genes. Different zinc-finger protein genes were found to improve drought tolerance in different plant species. Expression of CCCH-type zinc-finger gene OsTZF1 is induced by drought stress in rice (Jan et al. 2013). A C2H2-type zinc-finger protein gene GmZFP3 in soybean showed negative impact on drought tolerance in transgenic Arabidopsis (Zhang et al. 2016a). IBZFP1 is encoding a C2/H2 zinc-finger protein gene from sweet potato, improving drought tolerance in transgenic Arabidopsis (Wang et al. 2016a). Overexpression of another C2H2-type zinc-finger protein gene GsZFP1 in transgenic Arabidopsis also enhanced drought tolerance (Luo et al. 2012).
MQTL3H.4 was formed with eight droughttolerant QTL from two populations based on different physiological traits, wilting score, peduncle length, water-soluble carbohydrate contents, and plant height (Sayed et al. 2012; Diab et al. 2004; Korff et al. 2008). MQTL6H.2 was formed with seven QTL for drought tolerance based on grain yield, kernel weight, and plant height (Korff et al. 2008) and chlorophyll content (This et al. 2000). MQTL3H.4 and MQTL6H.2 had relatively large confidence intervals (7.0 and 6.8 cM), resulting in the large amount of candidate genes (more than 100).
Salinity tolerance in barley
Salinity tolerance is also a complex trait, controlled by many minor QTL (Flowers and Flowers 2005). Slow progress was made to improve salinity tolerance with MAS in crops during the past few years although many QTL for salinity tolerance were identified (Ashraf and Foolad 2013). There are two phases of growth reactions in crops to salinity stress (Munns and Tester 2008). The first phase is the osmotic effect to crops, reducing water uptake by crops, that is similar to drought effects. The second phase is the ion toxicity caused by Na+ and/or Cl− that inhibit crop growth.
Many physiological traits are regarded as salinity-tolerant mechanisms (Colmer et al. 2005; Munns 2005). This includes: osmotic adjustment; Na+ exclusion from uptake; control of xylem ion loading; efficient vacuolar Na+ sequestration; reactive oxygen species (ROS) detoxification; and cytosolic K+ homeostasis (Flowers and Colmer 2008; Munns and Tester 2008).
The fine-mapped QTL for salinity tolerance were on chromosome 1H and 7H. HvNax4 is the locus lowering the shoot Na+ contents in barley on MQTL1H.4 (Rivandi et al. 2011). This locus was fine-mapped and 34 candidate genes were identified (Rivandi et al. 2011). Possibly, the detected QTL for salinity tolerance at seedling stage on MQTL1H.4 had the same genes with HvNax4 (Mano and Takeda 1997). MQTL1H.4 was also the hot spot to improve drought tolerance, including five drought-tolerant QTL (discussed above), showing the possibility of improving drought tolerance and salinity tolerance simultaneously.
Among the identified 30 candidate genes on MQTL1H.4, overexpression of heavy metal transport/detoxification superfamily protein was detected in transgenic Arabidopsis under salinity conditions (Yokotani et al. 2013). Based on meta-analysis in rice, the pentatricopeptide repeat (PPR)-containing protein-like gene was identified as the candidate gene for improving rice yield on different chromosomes (Swamy et al. 2011).
MQTL7H.2 formed a fine-mapped major locus for salinity tolerance HvNax3, explaining 51 % phenotypic variance with an LOD value of 9.9 (Shavrukov et al. 2010). Neither HvNax3 nor HvNax4 was able to influence K+ contents in barley (Rivandi et al. 2011; Shavrukov et al. 2010), while HvNax3 was shown to lower the sodium accumulation in leaves. The physiological mechanisms of this reduction remain a matter of conjecture. Several candidate genes were identified in the HvNax3 locus with colinearity in rice and Brachypodium. From meta-analysis, 67 candidate genes were also identified in the locus HvNax3. Which of these candidate genes play a role in controlling Na+ content in the shoot remain to be investigated in future experiments. It was suggested earlier that other Nax loci, Nax 1 and Nax 2, enhance the retrieval of Na+ back into the root stele via HKT1;4 or HKT1;5 (Munns et al. 2012). However, more recent studies have shown that Nax loci also reduce the rate of Na+ loading into the xylem via SOS1 Na+/H+ exchanger in wheat (Zhu et al. 2016). It remains to be determined which of these mechanisms is conferred by Nax3 loci. QTL for germination speed under salinity stress (Mano and Takeda 1997) and salinity tolerance score (Fan et al. 2015) were also located in MQTL7H.2. It is probable that these two QTL are also controlled by the locus HvNax3.
The TaMyb1 gene was suggested to be involved in the signalling pathways of waterlogging and salinity stresses (Lee et al. 2007). Overexpression of another Myb transcription factor gene, JAmyb, contributed to salinity tolerance by stimulating abiotic stress-tolerant genes, such as osmotic adjustment and ROS scavenging, in rice and Arabidopsis (Yokotani et al. 2013). Another Myb transcription factor, SRM1, is able to regulate the ABA biosynthesis and signalling-related genes in Arabidopsis under salinity stress (Wang et al. 2015). A calmodulin-like protein OsMSR2 identified in rice was found to improve drought and salinity tolerance by regulating stress-related genes in ABA-mediated pathways. Expression of OsMSR2 showed improved drought and salinity tolerance in Arabidopsis (Xu et al. 2011).
Generally, root K+ retention ability is strongly associated with salinity tolerance in barley (Chen et al. 2005, 2007). One QTL for root K+ under salinity stress was identified on chromosome 1H, located on MQTL1H.3 (Nguyen et al. 2013). MQTL1H.3 was formed with four initial QTL for salinity tolerance from three different experiments from the same mapping population (Steptoe/Morex). Different traits were used among these three initial QTL: leaf injury, root K+, and chlorophyll content. All of these three salinity-tolerant QTL on MQTL1H.3 were from the seedling growth stages.
MQTL2H.1 was formed with 21 major QTL, including two for waterlogging tolerance, one for combined salinity and waterlogging tolerance, three for drought tolerance, and 15 for salinity tolerance. This region is the main area contributing to salinity tolerance in barley, at both seedling and vegetative growth stages. MQTL2H.1 included QTL for salinity tolerance based on leaf yellowing, number of yellow leaves, leaf dry matter, and proline, Na+, K+, and Cl− contents in leaves.
There were 25 candidate genes on MQTL2H.1, including two candidate genes of particular interest. In soybean seedlings, the protein flavonol 4′-sulfotransferase was downregulated when placed under combined salinity and waterlogging conditions (Alam et al. 2011). The cytochrome P450-like gene was upregulated in waterlogged rape seedlings (Lee et al. 2014). These two candidate genes on MQTL2H.1 can be further explored to improve abiotic stress tolerance in barley.
MQTL5H.3 was formed with six QTL for salinity tolerance from three different studies. Glutamate receptor was the candidate gene on MQTL5H.3. Glutamate receptor is one of the factors inducing K+ efflux under abiotic stresses (Demidchik et al. 2014). Maintaining high cytosolic K+ level with lower K+ efflux is crucial for abiotic stress tolerance in barley (Shabala and Pottosin 2014), and a causal link exists between cytosolic K+ concentration and the ability of a cell to undergo programmed cell death (e.g., senescence) (Demidchik et al. 2014; Shabala et al. 2010).
Waterlogging tolerance in barley
The factor that has impeded the progress of improving waterlogging tolerance in barley is the low heritability of plant yield under waterlogging conditions (Collaku and Harrison 2005; Zhou 2010). Despite the advanced genotyping technology, accurate phenotyping remains to be a challenge in plant breeding for waterlogging tolerance (Zhou 2011). Agronomic traits were widely used to screen waterlogging tolerance in barley, rice and maize (Qiu et al. 2007; Xu and Mackill 1996; Zhou 2010). Visual symptom of leaf yellowing is the main indicator of waterlogging tolerance in barley breeding programs (Supplementary Table S1). Utilizing physiological traits associated with waterlogging tolerance, such as higher K+ contents, is required in waterlogging breeding programs (Shabala 2011; Shabala et al. 2014). Even in breeding, only a few physiological traits have been utilized and none of the genes encoding these traits have been cloned (Collins et al. 2008). More convenient and reliable physiological traits should be further explored to screen waterlogging tolerance.
Aerenchyma formation in roots is a reliable and faster method to detect waterlogging tolerance, compared with leaf chlorosis (Zhang et al. 2015, 2016b). Root porosity is the percentage of gas volume per root volume, widely used as an indicator of aerenchyma formation (Colmer 2003). Aerenchyma provides an internal system of gas-filled spaces to improve oxygen supply in waterlogged roots (Evans 2004). MQTL4H.4 was formed with nine QTL for waterlogging tolerance, including one fine-mapped QTL for aerenchyma formation and two QTL for root porosity under waterlogging conditions (Zhang et al. 2016b). The seven QTL were from the population of Yerong/Franklin and two QTL from YYXT/Franklin (Broughton et al. 2015; Li et al. 2008; Zhou 2011; Zhou et al. 2012). MQTL4H.4 was positioned at 98.6 cM with confidence interval of 1.2. MQTL4H.4 and can be used in MAS in breeding to improve waterlogging tolerance in barley.
There were 58 candidate genes on MQTL4H.4 contributing to waterlogging tolerance in barley. Members of the family of NAC domain proteins were increased during leaf senescence in Arabidopsis (Buchanan-Wollaston et al. 2005). Also, the NAC domain-containing gene ANAC102 was induced as an important regulator of seed germination under waterlogging conditions (Christianson et al. 2009). In waterlogging-tolerant maize, calcium-dependent lipid-binding (CaLB domain) protein showed increasing abundance (Yu et al. 2015). Catalase was one of the antioxidant enzymes reducing the oxidative stress under waterlogging conditions (Zhang et al. 2015). Cytochrome P450-like gene and glutathione-S-transferase on MQTL4H.4 were upregulated in waterlogged rape seedlings (Lee et al. 2014). However, glutathione-S-transferase gene was downregulated in waterlogged cucumber (Qi et al. 2012). Glycosyltransferase genes, which are involved in cytokinin inactivation, showed decreased expression under waterlogging conditions (Christianson et al. 2010; Qi et al. 2012). LOB DOMAIN-CONTAINING PROTEIN 41 (LBD41) is likely a repressing factor in submerged Arabidopsis (Voesenek et al. 2016). The mitochondrial serine acetyltransferase gene was upregulated in waterlogged rape seedlings (Christianson et al. 2010).
Lysigenous aerenchyma formation candidate genes have been identified in maize (Rajhi et al. 2011). The identified candidate genes, NAC domain transcription factor gene and glutathione-S-transferase gene, were both located to MQTL4H.4 (Rajhi et al. 2011). MQTL4H.4 also included one fine mapped QTL for aerenchyma formation under waterlogging conditions in barley (Zhang et al. 2016b). Further studies are needed to identify genes controlling aerenchyma formation under waterlogging conditions in barley.
Based on chlorophyll fluorescence, there were two major QTL under hypoxia conditions identified on chromosome 6H, explaining 39.8 % of the phenotypic variance (Bertholdsson et al. 2015). This suggests that QTL can also be fine mapped and used for MAS. Meta-analysis projected these two QTL on MQTL6H.4 on the physical map.
MQTL7H.2 also included two QTL for waterlogging tolerance. Ethylene response factors gene is also located in the region of MQTL7H.2 and can be the candidate gene for waterlogging-tolerant QTL on MQTL7H.2 (Xu et al. 2006). Increased transcripts of a Myb transcription factor TaMyb1 gene were identified in wheat under waterlogging conditions and combined salinity and waterlogging stress (Lee et al. 2007).
Combined drought and salinity stresses
Plants are usually subjected to combined drought and salinity in both natural and agricultural systems (Roy et al. 2011). The direct effect from drought and salinity stresses is the reduction of photosynthesis and cell growth (Chaves et al. 2009). Osmotic adjustment is one of the crucial mechanisms of drought tolerance in crops, enhancing photosynthetic rates through water uptake and cell turgor (Cattivelli et al. 2008). Osmotic adjustment is also the key trait for salinity tolerance in the first phase of salinity stress (Munns and Tester 2008). MQTL2H.4, MQTL5H.1 and MQTL6H.2 were all from one QTL for leaf osmotic potential under drought stress (Teulat et al. 2001, 1998) and QTL for salinity tolerance. These findings suggested the possibility of using osmotic adjustment to improve drought and salinity tolerance simultaneously, as well as improving combined drought and salinity tolerance.
MQTL1H.4 was formed with five QTL for drought tolerance and two QTL for salinity tolerance. One QTL for salinity tolerance, HvNax4, which lowers the shoot Na+ content in barley, is mapped to a 200-kb interval within this region (Rivandi et al. 2011). MQTL1H.4 can be a possible region controlling combined drought and salinity tolerance in barley. Until now, experiments regarding plant response to combined drought and salinity stresses are limited (Ahmed et al. 2013). QTL for combined drought and salinity tolerance have not been identified yet.
Drought would aggravate the ion toxicity caused by Na+ and Cl−, thereby impeding plant growth (Ahmed et al. 2013). Under combined drought and salinity stresses, relatively more tolerant wild barley genotypes were shown to have higher K+ contents and K+/Na+ ratio than the relatively intolerant genotypes (Ahmed et al. 2013). MQTL2H.1 was formed with 16 QTL for salinity tolerance and three QTL for drought tolerance. One QTL for salinity tolerance was based on shoot K+ contents. MQTL3H.1 was formed with two drought-tolerant QTL from two different populations (Diab et al. 2004; Zhang et al. 2005), and two salinity-tolerant QTL based on the plant Na+/K+ ratio and shoot Na+/K+ ratio (Nguyen et al. 2013). MQTL2H.1 and MQTL3H.1 both illustrated that maintaining higher K+ contents helped plants to adapt better to the drought, salinity, and combined drought and salinity stresses.
Combined salinity and waterlogging stresses
Waterlogged soils can be also affected by salinity. Under combined salinity and waterlogging stresses, severe damage occurs in barley (Colmer et al. 2005). Oxygen deprivation in waterlogged soils inhibited the ATP production in plants (Bailey-Serres and Voesenek 2008). Reduced ATP in plants leads to increased Na+ and decreased K+ levels in leaves under combined stress (Barrett-Lennard and Shabala 2013; Zeng et al. 2013). Aerenchyma provides an internal system of gas-filled spaces to improve oxygen supply to waterlogged roots, leading to increased energy in plants (Evans 2004). Therefore, it is proposed that aerenchyma formation can be an effective mechanism in plants under combined salinity and waterlogging stresses (Colmer and Flowers 2008). Maintaining lower Na+ and higher K+ content in leaves is another key mechanism for improving combined salinity and waterlogging tolerance (Zeng et al. 2013). So far, there is only one experimental study detecting QTL for combined salinity and waterlogging tolerance in barley (Ma et al. 2015).
In our study, MQTL4H.4 was formed with nine QTL for waterlogging tolerance, including one fine-mapped QTL for aerenchyma formation and two QTL for root porosity under waterlogging conditions (Zhang et al. 2016b). However, no salinity-tolerant QTL were located on MQTL4H.4. Another QTL for root porosity under waterlogging conditions was on MQTL6H.2, and without any QTL for salinity tolerance. Also, MQTL1H.4 with fine-mapped salinity-tolerant QTL HvNax4, was not formed with QTL for waterlogging tolerance. MQTL7H.2 was formed with fine-mapped salinity-tolerant QTL HvNax3 and two QTL for waterlogging tolerance.
QTL for combined salinity and waterlogging tolerance were projected on MQTL1H.1, MQTL1H.3, MQTL2H.1, MQTL5H.6, and MQTL7H.3 (Ma et al. 2015). MQTL2H.1 and MQTL7H.3 both included QTL for salinity tolerance and waterlogging tolerance. MQTL2H.1 and MQTL7H.3 indicated the possibility to improve combined salinity and waterlogging tolerance.
Importance of marker validation and limitations of meta-analysis
A major objective of QTL studies is to find QTL that can be implemented into breeding programs via MAS. The major objective of barley breeding is high yield, combined with greater malting quality and insensitivity to biotic and abiotic stresses. QTL has been successful for introgressing and pyramiding major-effect genes. However, there are still many traits of interest facing great challenges since traits are controlled by many QTL with small effects.
A meta-analysis of QTL associated with abiotic stresses has been performed in barley (Li et al. 2013). Overall, 35 experiments under both control and stress conditions, with 337 major or minor QTL on drought, salinity, waterlogging, low temperature, mineral toxicity or deficiency were included in their study (Li et al. 2013). In our study, a larger number and the latest QTL (632 overall) for drought, salinity, and waterlogging tolerance were investigated. Before meta-analysis, we excluded the parameters under control conditions and the QTL with minor effects. The QTL controlling the yield-related traits under stresses might be the QTL for yield-related traits, rather than the stress-tolerant QTL (Jones 2007). The change of traits under stress conditions should be compared with the traits under control conditions. We only used major QTL for stress tolerance in barley to perform meta-analysis since MAS was successful in crop breeding with one or two major genes controlling stress tolerance. To our knowledge, this is the first meta-analysis that projected all the QTL on the barley physical map, with previous meta-analysis generating the consensus map from the markers common to the different population maps (Khowaja et al. 2009; Li et al. 2013; Zhang et al. 2010). A limited number of common markers from different populations resulted in the inaccurate QTL positions on the consensus map. In our study, we used the position of the QTL flanking markers on the barley physical map to refine the positions of abiotic stress-tolerant QTL from different studies. The positions of MQTL on the barley physical map were also used to search the candidate genes. Identified candidate genes on the physical map provide meaningful information for further MAS and positional cloning.
Meta-analysis is able to integrate the different QTL from different populations into one consensus map. Meta-analysis has also successfully validated the major QTL for abiotic stress tolerance in barley reducing the confidence interval of MQTL. After primary QTL mapping, the mapped QTL was located within a chromosome region so that the confidence interval was up to 50 cM (Supplementary Table S2). Chromosome regions within 10 cM include several hundreds of genes (de Dorlodot et al. 2007; Salvi and Tuberosa 2005). Fine mapping is widely used to refine the QTL less than 1 cM between flanking markers to search for candidate genes and positional cloning of QTL for abiotic stress tolerance (de Dorlodot et al. 2007; Rivandi et al. 2011; Semagn et al. 2013; Shavrukov et al. 2010). QTL cloning has enhanced the exploitation of functions of tolerant genes and the allelic variation in germplasm (Ashraf and Foolad 2013). Meta-analysis provided another method to refine the locations of QTL by lowering the confidence interval (de Dorlodot et al. 2007). The calculated meta-QTL provides breeders with target regions on consensus map for further MAS. However, how effective and accurate is the reduction of confidence interval for searching candidate genes by meta-analysis remains unknown unless that the number of observed QTL is more than five (Veyrieras et al. 2007). Also, recombination might break the linkage between markers and target QTL. Further experiments are, therefore, required to explore the MQTL from meta-analysis before it can become an effective tool in crop breeding.
Conclusions
Both MQTL3H.4 and MQTL6H.2 were target regions controlling drought tolerance in barley. Further experiments are required to fine-map these regions for the effective use of MAS in drought tolerance breeding in barley. Fine-mapped QTL for salinity tolerance, HvNax4 and HvNax3, were validated on MQTL1H.4 and MQTL7H.2, respectively. MQTL1H.4 was formed with fine-mapped salinity-tolerant HvNax4 and five initial major QTL for drought tolerance. MQTL1H.4 provides breeders with the possibility of improving drought tolerance and salinity tolerance simultaneously and thereby improving barley performance under combined drought and salinity stresses. MQTL7H.2 was formed with a fine-mapped major locus for salinity tolerance, HvNax3, two other QTL for salinity tolerance and two QTL for waterlogging tolerance. Genes for ethylene response factors and Myb transcription factor are possible candidate genes for salinity-tolerant locus HvNax3. Improved salinity tolerance, waterlogging tolerance, and combined salinity and waterlogging tolerance can be achieved by selecting MQTL7H.2. MQTL2H.1 and MQTL5H.3 were also target regions improving salinity tolerance. MQTL4H.4 is the main region controlling waterlogging tolerance in barley, including fine-mapped QTL for aerenchyma formation under waterlogging conditions. The genes for NAC domain transcription factor and glutathione-S-transferase are candidate genes for aerenchyma formation under waterlogging conditions in barley. Identified MQTL and candidate genes provide breeders with target regions to improve drought, salinity, and waterlogging tolerance in barley.
Author contribution statement
XZ conducted the data analysis and wrote the paper; SS, AK, and LS contributed to writing; MZ designed and wrote the paper.
References
Ahmed IM, Cao F, Zhang M, Chen X, Zhang G, Wu F (2013) Difference in yield and physiological features in response to drought and salinity combined stress during anthesis in Tibetan wild and cultivated barleys. PLoS One 8(10):e77869. doi:10.1371/journal.pone.0077869
Alam I, Sharmin S, Kim K-H, Kim Y-G, Lee J, Bahk J, Lee B-H (2011) Comparative proteomic approach to identify proteins involved in flooding combined with salinity stress in soybean. Plant Soil 346(1–2):45–62. doi:10.1007/s11104-011-0792-0
Arcade A, Labourdette A, Falque M, Mangin B, Chardon F, Charcosset A, Joets J (2004) BioMercator: integrating genetic maps and QTL towards discovery of candidate genes. Bioinformatics 20(14):2324–2326
Ashraf M, Foolad MR (2013) Crop breeding for salt tolerance in the era of molecular markers and marker-assisted selection. Plant Breed 132(1):10–20. doi:10.1111/pbr.12000
Bailey-Serres J, Voesenek LA (2008) Flooding stress: acclimations and genetic diversity. Annu Rev Plant Biol 59:313–339. doi:10.1146/annurev.arplant.59.032607.092752
Barrett-Lennard EG, Shabala SN (2013) The waterlogging/salinity interaction in higher plants revisited—focusing on the hypoxia-induced disturbance to K+ homeostasis. Funct Plant Biol 40(9):872–882. doi:10.1071/FP12235
Bertholdsson NO, Holefors A, Macaulay M, Crespo-Herrera LA (2015) QTL for chlorophyll fluorescence of barley plants grown at low oxygen concentration in hydroponics to simulate waterlogging. Euphytica 201(3):357–365. doi:10.1007/s10681-014-1215-0
Blum A (2005) Drought resistance, water-use efficiency, and yield potential—are they compatible, dissonant, or mutually exclusive? Aust J Agric Res 56(11):1159–1168. doi:10.1071/AR05069
Broughton S, Zhou G, Teakle N, Matsuda R, Zhou M, O’Leary R, Colmer T, Li C (2015) Waterlogging tolerance is associated with root porosity in barley (Hordeum vulgare L.). Mol Breed 35(1):1–15. doi:10.1007/s11032-015-0243-3
Buchanan-Wollaston V, Page T, Harrison E, Breeze E, Lim PO, Nam HG, Lin J-F, Wu S-H, Swidzinski J, Ishizaki K, Leaver CJ (2005) Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J 42(4):567–585. doi:10.1111/j.1365-313X.2005.02399.x
Cattivelli L, Rizza F, Badeck F-W, Mazzucotelli E, Mastrangelo AM, Francia E, Mare C, Tondelli A, Stanca AM (2008) Drought tolerance improvement in crop plants: an integrated view from breeding to genomics. Field Crops Res 105(1–2):1–14. doi:10.1016/j.fcr.2007.07.004
Chardon F, Virlon B, Moreau L, Falque M, Joets J, Decousset L, Murigneux A, Charcosset A (2004) Genetic architecture of flowering time in maize as inferred from quantitative trait loci meta-analysis and synteny conservation with the rice genome. Genetics 168(4):2169–2185. doi:10.1534/genetics.104.032375
Chardon F, Jasinski S, Durandet M, Lécureuil A, Soulay F, Bedu M, Guerche P, Masclaux-Daubresse C (2014) QTL meta-analysis in Arabidopsis reveals an interaction between leaf senescence and resource allocation to seeds. J Exp Bot 65(14):3949–3962. doi:10.1093/jxb/eru125
Chaves MM, Flexas J, Pinheiro C (2009) Photosynthesis under drought and salt stress: regulation mechanisms from whole plant to cell. Ann Bot 103(4):551–560. doi:10.1093/aob/mcn125
Chen Z, Newman I, Zhou M, Mendham N, Zhang G, Shabala S (2005) Screening plants for salt tolerance by measuring K+ flux: a case study for barley. Plant, Cell Environ 28(10):1230–1246. doi:10.1111/j.1365-3040.2005.01364.x
Chen Z, Pottosin II, Cuin TA, Fuglsang AT, Tester M, Jha D, Zepeda-Jazo I, Zhou M, Palmgren MG, Newman IA, Shabala S (2007) Root plasma membrane transporters controlling K+/Na+ homeostasis in salt-stressed barley. Plant Physiol 145(4):1714–1725
Chen G, Krugman T, Fahima T, Chen K, Hu Y, Roder M, Nevo E, Korol A (2010) Chromosomal regions controlling seedling drought resistance in Israeli wild barley, Hordeum spontaneum C. Koch. Genet Resour Crop Evol 57(1):85–99. doi:10.1007/s10722-009-9453-z
Christianson JA, Wilson IW, Llewellyn DJ, Dennis ES (2009) The low-oxygen-induced NAC domain transcription factor ANAC102 affects viability of Arabidopsis seeds following low-oxygen treatment. Plant Physiol 149(4):1724–1738
Christianson JA, Llewellyn DJ, Dennis ES, Wilson IW (2010) Global gene expression responses to waterlogging in roots and leaves of cotton (Gossypium hirsutum L.). Plant Cell Physiol 51(1):21–37. doi:10.1093/pcp/pcp163
Collaku A, Harrison SA (2005) Heritability of waterlogging tolerance in wheat. Crop Sci 45:722–727
Collard BCY, Jahufer MZZ, Brouwer JB, Pang ECK (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142(1–2):169–196. doi:10.1007/s10681-005-1681-5
Collins NC, Fo Tardieu, Tuberosa R (2008) Quantitative trait loci and crop performance under abiotic stress: where do we stand? Plant Physiol 147(2):469–486
Colmer TD (2003) Aerenchyma and an inducible barrier to radial oxygen loss facilitate root aeration in upland, paddy and deep-water rice (Oryza sativa L.). Ann Bot 91(2):301–309
Colmer TD, Flowers TJ (2008) Flooding tolerance in halophytes. New Phytol 179(4):964–974. doi:10.1111/j.1469-8137.2008.02483.x
Colmer TD, Munns R, Flowers TJ (2005) Improving salt tolerance of wheat and barley: future prospects. Aust J Exp Agr 45:1425–1443
Darvasi A, Soller M (1997) A simple method to calculate resolving power and confidence interval of QTL map location. Behav Genet 27(2):125–132. doi:10.1023/a:1025685324830
de Dorlodot S, Forster B, Pagès L, Price A, Tuberosa R, Draye X (2007) Root system architecture: opportunities and constraints for genetic improvement of crops. Trends Plant Sci 12(10):474–481. doi:10.1016/j.tplants.2007.08.012
Demidchik V, Straltsova D, Medvedev SS, Pozhvanov GA, Sokolik A, Yurin V (2014) Stress-induced electrolyte leakage: the role of K+-permeable channels and involvement in programmed cell death and metabolic adjustment. J Exp Bot 65(5):1259–1270. doi:10.1093/jxb/eru004
Diab A, Teulat-Merah B, This D, Ozturk N, Benscher D, Sorrells M (2004) Identification of drought-inducible genes and differentially expressed sequence tags in barley. Theor Appl Genet 109(7):1417–1425. doi:10.1007/s00122-004-1755-0
Eagles HA, Bariana HS, Ogbonnaya FC, Rebetzke GJ, Hollamby GJ, Henry RJ, Henschke PH, Carter M (2001) Implementation of markers in Australian wheat breeding. Aust J Agric Res 52:1349–1356
Evans DE (2004) Aerenchyma formation. New Phytol 161:35–49
Fan Y, Shabala S, Ma Y, Xu R, Zhou M (2015) Using QTL mapping to investigate the relationships between abiotic stress tolerance (drought and salinity) and agronomic and physiological traits. BMC Genom 16(1):43
Flowers TJ, Colmer TD (2008) Salinity tolerance in halophytes. New Phytol 179(4):945–963. doi:10.1111/j.1469-8137.2008.02531.x
Flowers TJ, Flowers SA (2005) Why does salinity pose such a difficult problem for plant breeders? Agric Water Manage 78(1–2):15–24. doi:10.1016/j.agwat.2005.04.015
Gill SS, Tuteja N (2010) Polyamines and abiotic stress tolerance in plants. Plant Signal Behav 5(1):26–33
Goffinet B, Gerber S (2000) Quantitative trait loci: a meta-analysis. Genetics 155:463–473
Guo P, Baum M, Varshney R, Graner A, Grando S, Ceccarelli S (2008) QTLs for chlorophyll and chlorophyll fluorescence parameters in barley under post-flowering drought. Euphytica 163(2):203–214. doi:10.1007/s10681-007-9629-6
Guo L, Yang H, Zhang X, Yang S (2013) Lipid transfer protein 3 as a target of MYB96 mediates freezing and drought stress in Arabidopsis. J Exp Bot 64(6):1755–1767. doi:10.1093/jxb/ert040
Hu H, Xiong L (2014) Genetic engineering and breeding of drought-resistant crops. Annu Rev Plant Biol 65(1):715–741. doi:10.1146/annurev-arplant-050213-040000
Jan A, Maruyama K, Todaka D, Kidokoro S, Abo M, Yoshimura E, Shinozaki K, Nakashima K, Yamaguchi-Shinozaki K (2013) OsTZF1, a CCCH-tandem zinc finger protein, confers delayed senescence and stress tolerance in rice by regulating stress-related genes. Plant Physiol 161(3):1202–1216
Jones HG (2007) Monitoring plant and soil water status: established and novel methods revisited and their relevance to studies of drought tolerance. J Exp Bot 58(2):119–130. doi:10.1093/jxb/erl118
Khowaja F, Norton G, Courtois B, Price A (2009) Improved resolution in the position of drought-related QTLs in a single mapping population of rice by meta-analysis. BMC Genom 10(1):276
Khush GS (2001) Green revolution: the way forward. Nat Rev Genet 2(10):815–822
Korff M, Grando S, Del Greco A, This D, Baum M, Ceccarelli S (2008) Quantitative trait loci associated with adaptation to Mediterranean dryland conditions in barley. Theor Appl Genet 117(5):653–669. doi:10.1007/s00122-008-0787-2
Lee TG, Jang CS, Kim JY, Kim DS, Park JH, Kim DY, Seo YW (2007) A Myb transcription factor (TaMyb1) from wheat roots is expressed during hypoxia: roles in response to the oxygen concentration in root environment and abiotic stresses. Physiol Plant 129(2):375–385. doi:10.1111/j.1399-3054.2006.00828.x
Lee Y-H, Kim K-S, Jang Y-S, Hwang J-H, Lee D-H, Choi I-H (2014) Global gene expression responses to waterlogging in leaves of rape seedlings. Plant Cell Rep 33(2):289–299. doi:10.1007/s00299-013-1529-8
Li H, Vaillancourt R, Mendham N, Zhou M (2008) Comparative mapping of quantitative trait loci associated with waterlogging tolerance in barley (Hordeum vulgare L.). BMC Genom 9:401. doi:10.1186/1471-2164-9-401
Li W-T, Liu C, Liu Y-X, Pu Z-E, Dai S-F, Wang J-R, Lan X-J, Zheng Y-L, Wei Y-M (2013) Meta-analysis of QTL associated with tolerance to abiotic stresses in barley. Euphytica 189(1):31–49. doi:10.1007/s10681-012-0683-3
Luo X, Bai X, Zhu D, Li Y, Ji W, Cai H, Wu J, Liu B, Zhu Y (2012) GsZFP1, a new Cys2/His2-type zinc-finger protein, is a positive regulator of plant tolerance to cold and drought stress. Planta 235(6):1141–1155. doi:10.1007/s00425-011-1563-0
Ma Y, Shabala S, Li C, Liu C, Zhang W, Zhou M (2015) Quantitative trait loci for salinity tolerance identified under drained and waterlogged conditions and their association with flowering time in barley (Hordeum vulgare. L). PLoS One 10(8):e0134822. doi:10.1371/journal.pone.0134822
Mano Y, Takeda K (1997) Mapping quantitative trait loci for salt tolerance at germination and the seedling stage in barley (Hordeum vulgare L.). Euphytica 94(3):263–272. doi:10.1023/a:1002968207362
Mittler R (2006) Abiotic stress, the field environment and stress combination. Trends Plant Sci 11(1):15–19. doi:10.1016/j.tplants.2005.11.002
Munns R (2005) Genes and salt tolerance: bringing them together. New Phytol 167(3):645–663. doi:10.1111/j.1469-8137.2005.01487.x
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59(1):651–681. doi:10.1146/annurev.arplant.59.032607.092911
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D, Gilliham M (2012) Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat Biotech 30(4):360–364. doi:10.1038/nbt.2120
Nguyen V, Ribot S, Dolstra O, Niks R, Visser RF, van der Linden CG (2013) Identification of quantitative trait loci for ion homeostasis and salt tolerance in barley (Hordeum vulgare L.). Mol Breed 31(1):137–152. doi:10.1007/s11032-012-9777-9
Qi X-H, Xu X-W, Lin X-J, Zhang W-J, Chen X-H (2012) Identification of differentially expressed genes in cucumber (Cucumis sativus L.) root under waterlogging stress by digital gene expression profile. Genomics 99(3):160–168. doi:10.1016/j.ygeno.2011.12.008
Qin F, Shinozaki K, Yamaguchi-Shinozaki K (2011) Achievements and challenges in understanding plant abiotic stress responses and tolerance. Plant Cell Physiol 52(9):1569–1582. doi:10.1093/pcp/pcr106
Qiu F, Zheng Y, Zhang Z, Xu S (2007) Mapping of QTL associated with waterlogging tolerance during the seedling stage in maize. Ann Bot 99(6):1067–1081. doi:10.1093/aob/mcm055
Rabello AR, Guimarães CM, Rangel PH, da Silva FR, Seixas D, de Souza E, Brasileiro AC, Spehar CR, Ferreira ME, Mehta  (2008) Identification of drought-responsive genes in roots of upland rice (Oryza sativa L.). BMC Genom 9(1):1–13. doi:10.1186/1471-2164-9-485
Rajhi I, Yamauchi T, Takahashi H, Nishiuchi S, Shiono K, Watanabe R, Mliki A, Nagamura Y, Tsutsumi N, Nishizawa NK, Nakazono M (2011) Identification of genes expressed in maize root cortical cells during lysigenous aerenchyma formation using laser microdissection and microarray analyses. New Phytol 190(2):351–368. doi:10.1111/j.1469-8137.2010.03535.x
Rivandi J, Miyazaki J, Hrmova M, Pallotta M, Tester M, Collins NC (2011) A SOS3 homologue maps to HvNax4, a barley locus controlling an environmentally sensitive Na+ exclusion trait. J Exp Bot 62(3):1201–1216. doi:10.1093/jxb/erq346
Roy SJ, Tucker EJ, Tester M (2011) Genetic analysis of abiotic stress tolerance in crops. Curr Opin Plant Biol 14(3):232–239. doi:10.1016/j.pbi.2011.03.002
Said J, Song M, Wang H, Lin Z, Zhang X, Fang D, Zhang J (2015) A comparative meta-analysis of QTL between intraspecific Gossypium hirsutum and interspecific G. hirsutum × G. barbadense populations. Mol Genet Genomics 290(3):1003–1025. doi:10.1007/s00438-014-0963-9
Salvi S, Tuberosa R (2005) To clone or not to clone plant QTLs: present and future challenges. Trends Plant Sci 10(6):297–304. doi:10.1016/j.tplants.2005.04.008
Sayed M, Schumann H, Pillen K, Naz A, Leon J (2012) AB-QTL analysis reveals new alleles associated to proline accumulation and leaf wilting under drought stress conditions in barley (Hordeum vulgare L.). BMC Genet 13(1):61
Semagn K, Beyene Y, Warburton M, Tarekegne A, Mugo S, Meisel B, Sehabiague P, Prasanna B (2013) Meta-analyses of QTL for grain yield and anthesis silking interval in 18 maize populations evaluated under water-stressed and well-watered environments. BMC Genom 14(1):313
Shabala S (2011) Physiological and cellular aspects of phytotoxicity tolerance in plants: the role of membrane transporters and implications for crop breeding for waterlogging tolerance. New Phytol 190(2):289–298. doi:10.1111/j.1469-8137.2010.03575.x
Shabala S, Pottosin I (2014) Regulation of potassium transport in plants under hostile conditions: implications for abiotic and biotic stress tolerance. Physiol Plant 151(3):257–279. doi:10.1111/ppl.12165
Shabala S, Shabala S, Cuin TA, Pang J, Percey W, Chen Z, Conn S, Eing C, Wegner LH (2010) Xylem ionic relations and salinity tolerance in barley. Plant J 61(5):839–853. doi:10.1111/j.1365-313X.2009.04110.x
Shabala S, Shabala L, Barcelo J, Poschenrieder C (2014) Membrane transporters mediating root signalling and adaptive responses to oxygen deprivation and soil flooding. Plant, Cell Environ 37(10):2216–2233. doi:10.1111/pce.12339
Shavrukov Y, Gupta N, Miyazaki J, Baho M, Chalmers K, Tester M, Langridge P, Collins N (2010) HvNax3—a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp. spontaneum). Funct Integr Genomics 10(2):277–291. doi:10.1007/s10142-009-0153-8
Singh S, Mackill DJ, Ismail AM (2009) Responses of SUB1 rice introgression lines to submergence in the field: yield and grain quality. Field Crops Res 113(1):12–23. doi:10.1016/j.fcr.2009.04.003
Sosnowski O, Charcosset A, Joets J (2012) BioMercator V3: an upgrade of genetic map compilation and quantitative trait loci meta-analysis algorithms. Bioinformatics 28(15):2082–2083. doi:10.1093/bioinformatics/bts313
Swamy BPM, Vikram P, Dixit S, Ahmed HU, Kumar A (2011) Meta-analysis of grain yield QTL identified during agricultural drought in grasses showed consensus. BMC Genom 12(1):319
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327(5967):818–822. doi:10.1126/science.1183700
Teulat B, This D, Khairallah M, Borries C, Ragot C, Sourdille P, Leroy P, Monneveux P, Charrier A (1998) Several QTLs involved in osmotic-adjustment trait variation in barley (Hordeum vulgare L.). Theor Appl Genet 96(5):688–698. doi:10.1007/s001220050790
Teulat B, Borries C, This D (2001) New QTLs identified for plant water status, water-soluble carbohydrate and osmotic adjustment in a barley population grown in a growth-chamber under two water regimes. Theor Appl Genet 103(1):161–170. doi:10.1007/s001220000503
Teulat B, Zoumarou-Wallis N, Rotter B, Ben Salem M, Bahri H, This D (2003) QTL for relative water content in field-grown barley and their stability across Mediterranean environments. Theor Appl Genet 108(1):181–188. doi:10.1007/s00122-003-1417-7
This D, Borries C, Souyris I, Teulat B (2000) QTL study of chlorophyll content as a genetic parameter of drought tolerance in barley. Barley Genet Newslett 30:20–23
Tuberosa R (2012) Phenotyping for drought tolerance of crops in the genomics era. Front Physiol 3:347. doi:10.3389/fphys.2012.00347
Tuberosa R, Salvi S (2006) Genomics-based approaches to improve drought tolerance of crops. Trends Plant Sci 11(8):405–412. doi:10.1016/j.tplants.2006.06.003
Veyrieras J-B, Goffinet B, Charcosset A (2007) MetaQTL: a package of new computational methods for the meta-analysis of QTL mapping experiments. BMC Bioinform 8(1):49
Voesenek LACJ, Sasidharan R, Visser EJW, Bailey-Serres J (2016) Flooding stress signaling through perturbations in oxygen, ethylene, nitric oxide and light. New Phytol 209(1):39–43. doi:10.1111/nph.13775
Wang T, Tohge T, Ivakov AA, Mueller-Roeber B, Fernie AR, Mutwil M, Schippers JH, Persson S (2015) Salt-Related MYB1 (SRM1) coordinates abscisic acid biosynthesis and signaling during salt stress in Arabidopsis. Plant Physiol 169:1027–1041
Wang F, Tong W, Zhu H, Kong W, Peng R, Liu Q, Yao Q (2016a) A novel Cys2/His2 zinc finger protein gene from sweetpotato, IbZFP1, is involved in salt and drought tolerance in transgenic Arabidopsis. Planta 243(3):783–797. doi:10.1007/s00425-015-2443-9
Wang Y, Xu J, Deng D, Ding H, Bian Y, Yin Z, Wu Y, Zhou B, Zhao Y (2016b) A comprehensive meta-analysis of plant morphology, yield, stay-green, and virus disease resistance QTL in maize (Zea mays L.). Planta 243(2):459–471. doi:10.1007/s00425-015-2419-9
William HM, Trethowan R, Crosby-Galvan EM (2007) Wheat breeding assisted by markers: CIMMYT’s experience. Euphytica 157(3):307–319. doi:10.1007/s10681-007-9405-7
Xu Y, Crouch JH (2008) Marker-Assisted selection in plant breeding: from publications to practice. Crop Sci 48(2):391–407. doi:10.2135/cropsci2007.04.0191
Xu K, Mackill D (1996) A major locus for submergence tolerance mapped on rice chromosome 9. Mol Breed 2(3):219–224. doi:10.1007/bf00564199
Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail AM, Bailey-Serres J, Ronald PC, Mackill DJ (2006) Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442:705–708
Xu G-Y, Rocha PCF, Wang M-L, Xu M-L, Cui Y-C, Li L-Y, Zhu Y-X, Xia X (2011) A novel rice calmodulin-like gene, OsMSR2, enhances drought and salt tolerance and increases ABA sensitivity in Arabidopsis. Planta 234(1):47–59. doi:10.1007/s00425-011-1386-z
Yokotani N, Ichikawa T, Kondou Y, Iwabuchi M, Matsui M, Hirochika H, Oda K (2013) Role of the rice transcription factor JAmyb in abiotic stress response. J Plant Res 126(1):131–139. doi:10.1007/s10265-012-0501-y
Yu F, Han X, Geng C, Zhao Y, Zhang Z, Qiu F (2015) Comparative proteomic analysis revealing the complex network associated with waterlogging stress in maize (Zea mays L.) seedling root cells. Proteomics 15(1):135–147. doi:10.1002/pmic.201400156
Zeng F, Shabala L, Zhou M, Zhang G, Shabala S (2013) Barley responses to combined waterlogging and salinity stress: separating effects of oxygen deprivation and elemental toxicity. Front Plant Sci 4:1–13
Zhang F, Chen G, Huang Q, Orion O, Krugman T, Fahima T, Korol AB, Nevo E, Gutterman Y (2005) Genetic basis of barley caryopsis dormancy and seedling desiccation tolerance at the germination stage. Theor Appl Genet 110(3):445–453. doi:10.1007/s00122-004-1851-1
Zhang L-Y, Liu D-C, Guo X-L, Yang W-L, Sun J-Z, Wang D-W, Zhang A (2010) Genomic distribution of quantitative trait loci for yield and yield-related traits in common wheat. J Integr Plant Biol 52(11):996–1007. doi:10.1111/j.1744-7909.2010.00967.x
Zhang X, Shabala S, Koutoulis A, Shabala L, Johnson P, Hayes D, Nichols D, Zhou M (2015) Waterlogging tolerance in barley is associated with faster aerenchyma formation in adventitious roots. Plant Soil 394(1–2):355–372. doi:10.1007/s11104-015-2536-z
Zhang D, Tong J, Xu Z, Wei P, Xu L, Wan Q, Huang Y, He X, Yang J, Shao H, Ma H (2016a) Soybean C2H2-Type zinc finger protein GmZFP3 with conserved QALGGH motif negatively regulates drought responses in transgenic Arabidopsis. Front Plant Sci 7:325. doi:10.3389/fpls.2016.00325
Zhang X, Zhou G, Shabala S, Koutoulis A, Shabala L, Johnson P, Li C, Zhou M (2016b) Identification of aerenchyma formation-related QTL in barley that can be effective in breeding for waterlogging tolerance. Theor Appl Genet 129(6):1167–1177. doi:10.1007/s00122-016-2693-3
Zhou M (2010) Improvement of plant waterlogging tolerance. In: Mancuso S, Shabala S (eds) Waterlogging signalling and tolerance in plants. Springer-Verlag, Heidelberg, pp 267–285. doi:10.1007/978-3-642-10305-6_13,#Springer-Verlag
Zhou MX (2011) Accurate phenotyping reveals better QTL for waterlogging tolerance in barley. Plant Breed 130(2):203–208. doi:10.1111/j.1439-0523.2010.01792.x
Zhou M, Johnson P, Zhou G, Li C, Lance R (2012) Quantitative trait loci for waterlogging tolerance in a barley cross of Franklin × YuYaoXiangTian Erleng and the relationship between waterlogging and salinity tolerance. Crop Sci 52(5):2082–2088. doi:10.2135/cropsci2012.01.0008
Zhu M, Zhou M, Shabala L, Shabala S (2016) Physiological and molecular mechanisms mediating xylem Na+ loading in barley in the context of salinity stress tolerance. Plant, Cell Environ:n/a-n/a. doi:10.1111/pce.12727
Acknowledgments
This work was supported by the Australian Research Council Linkage grant (LP120200516) and Grains Research and Development Corporation (GRDC) of Australia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared that no conflict of interest exists.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, X., Shabala, S., Koutoulis, A. et al. Meta-analysis of major QTL for abiotic stress tolerance in barley and implications for barley breeding. Planta 245, 283–295 (2017). https://doi.org/10.1007/s00425-016-2605-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-016-2605-4