Abstract
Given that combination with multiple biomarkers may well raise the predictive value of wound age, it appears critically essential to identify new features under the limited cost. For this purpose, the present study explored whether the gene expression ratios provide unique time information as an additional indicator for wound age estimation not requiring the detection of new biomarkers and allowing full use of the available data. The expression levels of four wound-healing genes (Arid5a, Ier3, Stom, and Lcp1) were detected by real-time polymerase chain reaction, and a total of six expression ratios were calculated among these four genes. The results showed that the expression levels of four genes and six ratios of expression changed time-dependent during wound repair. The six expression ratios provided additional temporal information, distinct from the four genes analyzed separately by principal component analysis. The overall performance metrics for cross-validation and external validation of four typical prediction models were improved when six ratios of expression were added as additional input variables. Overall, expression ratios among genes provide temporal information and have excellent potential as predictive markers for wound age estimation. Combining the expression levels of genes with ratio-expression of genes may allow for more accurate estimates of the time of injury.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Assessing the interval between injury and death is a central issue in forensic practice [1]. For many decades, numerous forensic specialists have attempted to determine wound age association with murders, cases of manslaughter, and accidents [2]. Cellular and molecular responses to injury are complicated and organized [3, 4]. Studies have shown that orchestrated biological phenomena (e.g., various inflammatory cells, cytokines, and growth factors), which occur predictably with time during wound healing processes, could be used to determine the age of mechanically induced wounds [5]. Temporal expression profiles of genes involved in healing processes (e.g., matrix metalloproteases, chemokines, and growth factors) have shown great potential for wound aging assessment [6,7,8].
With the increasing exploration of time-dependent markers, combinations of biomarkers have significantly achieved more robust outcomes than single biomarkers. Birincioglu et al. [9] measured nine cytokines using a multiplex bead-based immunoassay, while Kubo et al. [10] examined the temporal expression levels of 13 genes (e.g., cytokines, chemokines, and growth factors). Although those studies indicated that combinations of biomarkers enable accurate wound age estimation, the multiple markers are usually applied as simple combinations, and the information they can provide without fully exploited.
Machine learning, which can discover new knowledge and learn from databases, extracts latent details and uses them to make predictions and provide an effective analysis tool for using multi-index to estimate wound age [11, 12]. Generally, the availability of more data enhances the accuracy of machine learning. Therefore, how identifying more diagnostic features for wound age estimation is critically essential. The ratio of biomarkers can magnify the slight difference between two molecule expressions and be characterized in a biologically meaningful manner. Concentration changes in metabolite ratios between different pathological states are measured to construct differential metabolic networks because metabolite ratios could represent metabolic pathway reactions [13, 14]. Top-scoring pair (TSP) methods were proposed in microarray data analysis based on pairwise mRNA comparisons to identify crucial feature pairs [15]. Based on the predecessor’s research foundation, we attempted to calculate the ratio of gene expressions as a potential biomarker to improve prediction accuracy.
This study aimed to explore whether ratios of gene expression (ratio-expressions) provide distinct temporal information so that they can be used as indicators for estimating wound age to achieve cost reduction and better practical application. First, the expression levels of four wound-healing genes (AT-rich interactive domain-containing protein 5a [Arid5a], immediate early response-3 [Ier3], stomatin [Stom], and lymphocyte cytosolic protein 1 [Lcp1]) were detected by reverse transcription-quantitative polymerase chain reaction (RT-qPCR). Second, six expression ratios among four genes were calculated by 2−△△Ct method. Finally, the supervised learning algorithms were employed to investigate whether prediction accuracy could be improved when ratios of expression levels among genes were regarded as input parameters.
Materials and methods
Animal model of skeletal muscle contusion
All procedures were performed following the “Guiding Principles in the Use and Care of Animals” (NIH Publication No. 85–23, Revised 1996); they were approved by the Institutional Animal Care and Use Committee of Shanxi Medical University of China [batch number of rats: SCXK (Jin) (2009–0001)]. Animals received humane care following the principles of the Guide for the Care and Use of Laboratory Animals protocol, published by the Ministry of the People’s Republic of China (issued on June 4, 2004).
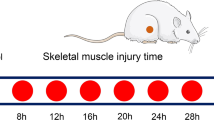
In total, 70 male Sprague–Dawley rats (6–8 weeks old, weighing 250–300 g) were obtained from the Animal Center of Shanxi Medical University. All rats were housed in cages with access to liberal amounts of food and water under a 12-h light–dark cycle at 22–24 °C and 40–60% relative humidity. In total, 56 rats were randomly sorted into a control group (n = 8) and 4-, 8-, 12-, 16-, 20-, and 24-h contusion groups (n = 8/group). Additionally, 14 rats were randomly allocated to control and contusion groups (n = 2/group) to validate the models. In brief, each rat was anesthetized with pentobarbital and placed on an experimental table in the supine position. Then, a 500-g counterpoise (1.1 cm in diameter) was dropped from a height of 30 cm through a clear Lucite guide tube onto the thigh muscles of the right posterior limb to simulate a mechanical injury that resulted in edema and hemorrhage, usually followed by muscle regeneration. This paradigm closely mimicked the inflammatory response and muscle healing process associated with violent muscle injury.
The rats were sacrificed at 4, 8, 12, 16, 20, and 24 h post-contusion (n = 8 per time point) using a lethal dose of pentobarbital (350 mg/kg body weight, intraperitoneal injection). Approximately 100 mg of muscle was sampled from the wound site and equally divided into two parts in each rat. In the control group, specimens were harvested from the same site after anesthetization with an overdose of pentobarbital. All muscle samples were immediately frozen with liquid nitrogen and then stored at − 80 °C until real-time PCR analysis.
Hematoxylin and eosin staining
The injured muscles were fixed in 4% paraformaldehyde in 0.1 M phosphate buffer (pH 7.4), processed in an automated tissue processor, and embedded in paraffin. Four-micron-thick coronal sections were cut on a microtome and stained with hematoxylin and eosin (H&E). Following staining, the slides were imaged using a Tissue Fax Plus 2000 slide scanner (Tissuen Gnostics) at a magnification of 200 × , and two different pathologies described the image of H&E sections.
Total RNA preparation
Total RNA was extracted from derived muscle specimens (approximately 50 mg/sample) using RNAiso Plus 9108 (Takara, Shiga, Japan). The quality and quantity of total RNA were determined using the Agilent 2100 bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) using an Agilent RNA 6000 Nanokit and the Infinite M200 Pro microplate reader (Tecan, Männedorf, Switzerland). Only RNAs with OD260/OD280 ratios of 1.8–2.2 and RNA integrity numbers > 7.0 were used for cDNA synthesis.
Real-time PCR
cDNA conversion was conducted using the Prime Script RT-PCR Kit (Takara) following the standard protocol. Reverse transcription was performed using 0.4 μg of total RNA per reaction in accordance with the manufacturer’s protocol. Subsequently, the reaction mixture was incubated for 5 min at 37 °C and 15 s at 85 °C. Primers and TaqMan fluorescent probes were designed using AlleleID 6 software (Premier Biosoft International, Palo Alto, CA, USA), verified with BLAST, and synthesized by Sangon Biotech (Shanghai, China). Sense and anti-sense primers were designed to span genomic DNA introns, thus avoiding the amplification of genomic DNA. Ribosomal protein L13 (RPL13) mRNA and ribosomal protein L32 (RPL32) mRNA, which are stably expressed in contused skeletal muscle [16], were selected as reference genes. The primers, probes, and fluorescent labeling sequences are listed in Table 1.
PCR reaction mixtures were prepared with 12.5 μL Premix Ex Taq, 2.0 μL 10% dimethyl sulfoxide, 3 μL RNase-free H2O, 1.5 μL cDNA, and 0.5 μL of primers and probes for each gene. Each mixture consisted of primers and probes for four genes (two reference genes and two target genes), allowing simultaneous measurement of those four genes in a single well. The amplification process was performed using a Bio-Rad real-time PCR system (CFX384; Bio-Rad, Hercules, CA, USA) and a protocol of pre-denaturation at 95 °C for 10 s, followed by 40 cycles of denaturation at 95 °C for 5 s and annealing/extension at 60 °C for 40 s.
For amplification efficiency, EASY Dilution solutions (Takara) were used for serial dilution of the synthesized cDNAs (fivefold 1, 1:51, 1:52, 1:53, and 1:54 or threefold 1, 1:31, 1:32, 1:33, and 1:34). Negative controls were also performed with deionized water during each run; the real-time PCR procedure was repeated at least three times for each sample. The amplification efficiencies of the genes were within 90–110%, and all R2 values were > 0.99.
Data analysis
The expression levels of four target mRNAs and six ratios of expression among the four genes were computed using the 2−△△Ct method. The means and standard errors of the means were calculated for all parameters investigated in the study. One-way analysis of variance was used for the analysis, and P < 0.05 was considered statistically significant.
Formula for calculating the relative expression of the four target genes:
Formula for calculating the six ratios of expression among the four genes:
Principal component analysis (PCA) was performed using SIMCA-P software (version 14.1; Umetrics, Malmo, Sweden) to extract temporal information from the expression data. Logistic regression, random forest, support vector machine, and multilayer perceptron classification models were established in the Python environment (version 3.8). For model evaluation, eightfold cross-validation and external validation were applied; the accuracy, recall rate, precision rate, and F1 score (i.e., comprehensive evaluation index) were calculated.
Results
Histological examination of contused skeletal muscle
To examine the morphological changes in skeletal muscle tissue at various injury time points, H&E staining was employed on the skeletal muscle tissue. The findings revealed that the muscle cells in the control group displayed clarity, with well-organized muscle bundles (Fig. 1a). Conversely, the contused skeletal muscles exhibited a gradual occurrence of hemorrhage, edema, myocyte degeneration, and inflammatory response following injury (Fig. 1b–g). Specifically, at 4 h post-injury, there was conspicuous infiltration of red blood cells, disarrayed arrangement of skeletal muscle cells, and a minor presence of infiltrating neutrophils. As time progressed, the number of neutrophils within the injured tissue steadily increased, culminating in a significant influx of neutrophils at 24 h post-injury.
Hematoxylin–eosin staining (H&E) of skeletal muscle samples in rats. a The morphology of normal skeletal muscle without contusion (control) showed that the muscle cells in the control group displayed clarity, with well-organized muscle bundles. b–g The morphology of contused skeletal muscle at 4, 8, 12, 16, 20, and 24 h after injury. The contused skeletal muscles exhibited a gradual occurrence of hemorrhage, edema, myocyte degeneration, and inflammatory response following injury. Specifically, at 4 h post-injury, there was conspicuous infiltration of red blood cells, disarrayed arrangement of skeletal muscle cells, and a minor presence of infiltrating neutrophils. As time progressed, the number of neutrophils within the injured tissue steadily increased, culminating in a significant influx of neutrophils at 24 h post-injury
Expression levels of Arid5a, Ier3, Stom, and Lcp1 during wound healing
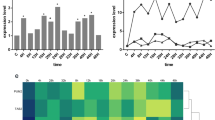
The relative expression levels of Arid5a, Ier3, Stom, and Lcp1 were detected by RT-qPCR (Table 2 and Fig. 2b). The expression profiles of all four genes exhibited time-dependent patterns after muscle contusion; all expression levels were significantly increased at 4 h after injury compared with the control group. The expression levels were consistently elevated within 24 h after injury, except Ier3 at 20 h and Stom at 8 h. Although these genes were upregulated after injury, their expression levels and profiles were distinct over time, implying specific temporal information that could aid wound age estimation.
Expression levels of Arid5a, Ier3, Stom, and Lcp1 were analyzed by RT-qPCR throughout the injury period. a Animal groups in the study. In total, 56 rats were randomly sorted into control (n = 8) and 4-, 8-, 12-, 16-, 20-, and 24-h contusion groups (n = 8/group). b Relative expression levels of Arid5a, Ier3, Stom, and Lcp1 after muscle contusion; comparisons among control and contusion groups were performed by one-way analysis of variance. Data are shown as means ± standard deviations; *P < 0.05
Changes in the ratio expressions among the four genes after muscle injury
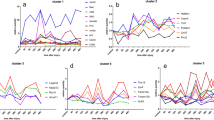
Because the four genes exhibited different expression profiles, we calculated six expression ratios among them to explore whether they varied in a time-dependent manner. As shown in Fig. 3, the six expression ratios among the four genes significantly changed after muscle contusion; they exhibited distinct patterns over time after injury. The level of Arid5a/Ier3 decreased from 4 to 16 h, then significantly increased at 20 h. Within 24 h after injury, Arid5a/Lcp1 and Stom/Lcp1 expression ratios were lower than in the control group. The ratios of Arid5a/Stom, Ier3/Lcp1, and Lcp1/Stom expression rapidly increased to a peak at 8 h, and then gradually decreased. These findings indicated that the ratio expressions among these four genes changed distinctly over time, implying the presence of temporal information for wound aging assessment.
PCA of temporal information contained in the four genes and six ratios of expression
PCA was applied to the expression data to more fully elucidate the temporal information provided by these indicators. The expression levels of Arid5a, Ier3, Stom, and Lcp1 changed during wound repair; the groups were distinct from each other except at 4, 12, and 16 h (Fig. 4a). Notably, the 4, 12, and 16 h samples were distinct in the scatter plots of the six expression ratios among these four genes, suggesting the presence of distinct temporal information (Fig. 4b). Moreover, after combining the expression data of the four genes with the six ratios of expression among them, we found that the samples were separated according to wound age; samples were more closely clustered within each group (Fig. 4c). Overall, the expression data of the four genes combined with the six ratios of expression among them may provide novel information for wound age estimation.
Performance comparison of models built without or with the six expression ratios among the four genes
In order to understand the performance of the expression ratios among those genes that contribute to estimating wound age, the classification models were developed using logistic regression, random forest, support vector machine, and multilayer perceptron methods. The accuracy, precision, recall, and F1 scores were measured to determine whether the predictive performance could be improved when the six expression ratios were additional input variables.
As a result, Table 3 and Fig. 5a show the performance evaluation metrics of the proposed classifiers during eight-fold cross-validation of the training data. Compared with the models built using four genes (Arid5a, Ier3, Stom, and Lcp1), the overall performance metrics were better for all models that included the four genes plus six expression ratios. The accuracy of the logistic regression, random forest, support vector machine, and multilayer perceptron models increased by 16.1%, 5.4%, 5.4%, and 8.9%, respectively.
Performance comparisons among models trained using the expression data of four genes, alone and in combination with six ratios of expression. a Accuracies and F1 scores of proposed predictors during eightfold cross-validation. b Accuracies, precision rates, recall rates, and F1 scores of external validation. Models without the six ratios of expression are shown in blue; models with the six ratios of expression are shown in red
Moreover, the total evaluation index F1 scores were improved by 17.5%, 6.8%, 5.8%, and 8.9%, respectively. These metrics demonstrated minor variances during eightfold cross-validation. Furthermore, models that included the four genes plus six ratios of expression achieved better classification performance during external validation; their accuracies, recall rates, precision rates, and F1 scores were improved by 14.3–21.5% (Fig. 5b). These findings confirmed that the six expression ratios provided additional temporal information, implying potential use as biomarkers.
Discussion
Skin injuries and muscle contusions are common injuries encountered in forensic practice due to the distribution of skeletal muscles and skin tissues throughout the body. Compared to skin tissues, skeletal muscles are less susceptible to the external environment as they are not in direct contact with the outside world. In forensic medicine, the study of skeletal muscle injury time inference has gradually increased [17,18,19]. Previous studies have shown that standard histological examination may not be able to determine the time of the wound in the first few minutes or hours after injury [20]. Indeed, the delay before polymorphonuclear neutrophil (PMN) infiltration ranges from 10 min to 4 h, making applying it widely in practice difficult [21]. Therefore, this study established a skeletal muscle contusion model with a wound age of 24 h and explored the expression changes of four mRNAs more suitable for the early wound age at an initial stage after 4 h of survival, providing a new idea for forensic injury time inference.
Wound vitality and progression have been widely investigated since the first reports by Raekallio using scientific experiments in the 1960s [22,23,24]. Nowadays, it has come from single biomarkers that were initially used; currently, combinations of multiple biomarkers are under consideration for diagnostic applications. For such combinations, machine learning can be used to identify complex patterns. Recently, using machine learning in forensic investigations has shown great potential for wound age estimation. Barington et al. [25] established a partial least square prediction model based on gene expression profiles, which could determine porcine bruise age with a precision of ± 2 h. Peyron et al. [26] performed multivariate logistic regression to distinguish between vital and early postmortem wounds using a combination of cytokines.
Although machine learning is a powerful approach for diagnosis and prediction, prediction outcomes depend on whether the data contain sufficient helpful information. Generally, additional time-dependent markers provide more significant temporal information for wound age estimation. However, assessment of additional biomarkers often has higher costs. Previous studies have explored changes in ratio relationships for feature pair selection defined potential ratio biomarkers [27, 28]. Thus, this study amid to explore whether ratios of gene expression contained temporal information and measure the discriminative ability of ratio features for wound age estimation, which is the difference from previous studies.
To our knowledge, this is the first description of ratio-expression among genes associated with wound aging. In this study, six expression ratios, calculated based on four wound-healing genes, exhibited time-dependent patterns after injury. PCA showed that the samples clustered distinctly according to wound age based on the six expression ratios, indicating temporal information distinct from the four genes. When the six ratios of expression were added as a new character input data of the machine learning algorithms, the performances of four models predicting wound age were improved. Overall, our results illustrated that six expression ratios provide distinct temporal information and could be used for wound age estimation.
The following reason may be the additional ratio-expressions that improve the accuracy of multi-indicator estimating injury time. First, genes exhibit distinct expression patterns depending on their function during healing. The ratios of expression levels among genes thus change over time after injury and may provide specific temporal information. Second, housekeeping genes are usually measured to normalize the mRNA level of target genes. However, the results will be biased due to the variability of the reference gene. In present study, the heterogeneity of housekeeping genes was eliminated by the ratio-expression of genes. Third, the ratios of expression levels among genes offer the full exploitation of the available data and magnify the slight difference between the two genes.
In this study, the four wound-healing genes (Arid5a, Ier3, Stom, and Lcp1) had distinct expression profiles associated with their different responses to injury. Compared with genes that exhibit similar expression patterns, genes that exhibit distinct patterns may be more suitable for wound age estimation because they provide additional temporal information. These four genes exhibit different functions after injury, associated with distinct expression patterns over time; these patterns determine the six ratios of expression among the four genes, which provide additional information for wound aging assessment. According to the literature, Arid5a, which is in the nucleus under normal conditions, translocates to the cytoplasm as a unique RNA-binding protein to stabilize various inflammatory cytokine mRNAs; it thus contributes to the inflammatory response [29, 30]. Ier3, a key regulatory factor in the immune response, is rapidly and transiently activated by numerous stimuli, such as growth factors, cytokines (such as tumor necrosis factor and IL-1b), ionizing radiation, viral infection, and other cellular stresses [31, 32]. Stom is a ubiquitously expressed membrane protein presumed to complex with ion channels, skeleton proteins, and transporters [33]. IL-6 has also induced Stom in a human amniotic cell line [34]. Lcp1, an important β-actin-bundling protein, is predominantly found in leukocytes and is required for cell migration; S-glutathionylation of Lcp1 impairs chemotaxis, polarization, and bactericidal activity in neutrophils [35].
Together, this study explored whether ratio-expressions among genes provided distinct temporal information and could be used for wound age estimation. The ratio-expression levels of four wound-healing genes (Arid5a, Ier3, Stom, and Lcp1) were calculated. Expression pattern examination and PCA showed that the six expression ratios provided distinct temporal information that could enhance prediction model performance, indicating the potential for use as diagnostic markers. Therefore, combined assessment using gene expression levels with the ratio-expressions among those genes could improve time of injury estimations.
Conclusion
The gene ratios were calculated to obtain more indicators to achieve a lower cost but more accurate estimation of wound age, making it more likely to reach a higher generalization ability. Although the ratio-expressions among genes show good performance and great potential for application on rat data, there remain translational challenges. Further studies are required according to the limitations of animal experiments. Next, we will collect human samples and apply these results to real medicolegal cases.
Data Availability
All relevant data and code are available from the corresponding author upon request.
References
Li N, Du Q, Bai R et al (2018) Vitality and wound-age estimation in forensic pathology: review and future prospects. Forensic Sci Res 5(1):15–24. https://doi.org/10.1080/20961790.2018.1445441
Grellner W (2007) Madea B Demands on scientific studies: vitality of wounds and wound age estimation. Forensic Sci Int 165(2–3):150–154. https://doi.org/10.1016/j.forsciint.2006.05.029
Ceafalan L, Dobre M, Milanesi E et al (2020) Gene expression profile of adhesion and extracellular matrix molecules during early stages of skeletal muscle regeneration. J Cell Mol Med. https://doi.org/10.1111/jcmm.15624
Li N, Bai RF, Li C et al. Insight into molecular profile changes after skeletal muscle contusion using microarray and bioinformatics analyses. Bioscience reports. 2021;41(1). https://doi.org/10.1042/bsr20203699
Kondo T (2010) Ishida Y Molecular pathology of wound healing. Forensic Sci Int 203(1–3):93–98. https://doi.org/10.1016/j.forsciint.2010.07.004
Wang Y, Yamamoto Y, Kuninaka Y et al (2015) Forensic potential of MMPs and CC chemokines for wound age determination. J Forensic Sci 60(6):1511–1515. https://doi.org/10.1111/1556-4029.12831
Yu T-S, Li Z, Zhao R et al (2016) Time-dependent expression of MMP-2 and TIMP-2 after rats skeletal muscle contusion and their application to determine wound age. J Forensic Sci 61(2):527–533. https://doi.org/10.1111/1556-4029.13001
Casse JM, Martrille L, Vignaud JM et al (2016) Skin wounds vitality markers in forensic pathology: an updated review. Med Sci Law 56(2):128–137. https://doi.org/10.1177/0025802415590175
Birincioglu I, Akbaba M, Alver A et al (2016) Determination of skin wound age by using cytokines as potential markers. J Forensic Leg Med 44:14–19. https://doi.org/10.1016/j.jflm.2016.08.011
Kubo H, Hayashi T, Ago K et al (2014) Temporal expression of wound healing-related genes in skin burn injury. Leg Med (Tokyo) 16(1):8–13. https://doi.org/10.1016/j.legalmed.2013.10.002
Deo RC (2015) Machine learning in medicine. Circulation 132(20):1920–1930. https://doi.org/10.1161/circulationaha.115.001593
Bi Q, Goodman KE, Kaminsky J et al (2019) What is machine learning? A primer for the epidemiologist. Am J Epidemiol 188(12):2222–2239. https://doi.org/10.1093/aje/kwz189
Huang X, Zhou Y, Tang H et al (2021) Differential metabolic network construction for personalized medicine: study of type 2 diabetes mellitus patients’ response to gliclazide-modified-release-treated. J Biomed Inform 118:103796. https://doi.org/10.1016/j.jbi.2021.103796
Netzer M, Weinberger KM, Handler M et al (2011) Profiling the human response to physical exercise: a computational strategy for the identification and kinetic analysis of metabolic biomarkers. Journal of clinical bioinformatics 1(1):34. https://doi.org/10.1186/2043-9113-1-34
Geman D, d’Avignon C, Naiman DQ et al (2004) Classifying gene expression profiles from pairwise mRNA comparisons. Stat Appl Genet Mol Biol 3:19. https://doi.org/10.2202/1544-6115.1071
Sun JH, Nan LH, Gao CR et al (2012) Validation of reference genes for estimating wound age in contused rat skeletal muscle by quantitative real-time PCR. Int J Legal Med 126(1):113–120. https://doi.org/10.1007/s00414-011-0604-3
Hassan Gaballah M, Fukuta M, Maeno Y et al (2016) Simultaneous time course analysis of multiple markers based on DNA microarray in incised wound in skeletal muscle for wound aging. Forensic Sci Int 266:357–368. https://doi.org/10.1016/j.forsciint.2016.06.027
Tian ZL, Jiang SK, Zhang M et al (2016) Detection of satellite cells during skeletal muscle wound healing in rats: time-dependent expressions of Pax7 and MyoD in relation to wound age. Int J Legal Med 130(1):163–172. https://doi.org/10.1007/s00414-015-1251-x
Fan YY, Zhang ST, Yu LS et al (2014) The time-dependent expression of alpha7nAChR during skeletal muscle wound healing in rats. Int J Legal Med 128(5):779–786. https://doi.org/10.1007/s00414-014-1001-5
Oehmichen M (2004) Vitality and time course of wounds. Forensic Sci Int 144(2–3):221–231. https://doi.org/10.1016/j.forsciint.2004.04.057
Gauchotte G, Wissler MP, Casse JM et al (2013) FVIIIra, CD15, and tryptase performance in the diagnosis of skin stab wound vitality in forensic pathology. Int J Legal Med 127(5):957–965. https://doi.org/10.1007/s00414-013-0880-1
Raekallio J (1960) Enzymes histochemically demonstrable in the earliest phase of wound healing. Nature 188:234–235. https://doi.org/10.1038/188234a0
Fatteh A (1966) Histochemical distinction between antemortem and postmortem skin wounds. J Forensic Sci 11(1):17–27
Raekallio J (1966) Applications of histochemistry to forensic medicine. Med Sci Law 6(3):142–146. https://doi.org/10.1177/002580246600600304
Barington K, Jensen HE (2017) Skovgaard K Forensic aspects of gene expression signatures for age determination in bruises as evaluated in an experimental porcine model. Forensic Sci Med Pathol 13(2):151–160. https://doi.org/10.1007/s12024-017-9869-2
Peyron PA, Colomb S, Becas D et al (2021) Cytokines as new biomarkers of skin wound vitality. Int J Legal Med 135(6):2537–2545. https://doi.org/10.1007/s00414-021-02659-z
Chen Y, Wang L, Li L et al (2016) Informative gene selection and the direct classification of tumors based on relative simplicity. BMC Bioinformatics 17:44. https://doi.org/10.1186/s12859-016-0893-0
Wang H, Zhang H, Dai Z et al (2013) TSG: a new algorithm for binary and multi-class cancer classification and informative genes selection. BMC Med Genomics 6 Suppl 1(Suppl 1):S3. https://doi.org/10.1186/1755-8794-6-s1-s3
Nyati KK, Agarwal RG, Sharma P et al (2019) Arid5a regulation and the roles of Arid5a in the inflammatory response and disease. Front Immunol 10:2790. https://doi.org/10.3389/fimmu.2019.02790
Higa M, Oka M, Fujihara Y et al (2018) Regulation of inflammatory responses by dynamic subcellular localization of RNA-binding protein Arid5a. Proc Natl Acad Sci USA 115(6):E1214-e1220. https://doi.org/10.1073/pnas.1719921115
Arlt A (2011) Schäfer H Role of the immediate early response 3 (IER3) gene in cellular stress response, inflammation and tumorigenesis. Eur J Cell Biol 90(6–7):545–552. https://doi.org/10.1016/j.ejcb.2010.10.002
Kochan J, Wawro M, Kolka A et al (2014) Transcription factor Elk-1 participates in the interleukin-1β-dependent regulation of expression of immediate early response gene 3 (IER3). Cytokine 70(2):120–125. https://doi.org/10.1016/j.cyto.2014.06.021
Lapatsina L, Brand J, Poole K et al (2012) Stomatin-domain proteins. Eur J Cell Biol 91(4):240–245. https://doi.org/10.1016/j.ejcb.2011.01.018
Snyers L (1994) Content J Induction of metallothionein and stomatin by interleukin-6 and glucocorticoids in a human amniotic cell line. Eur J Biochem 223(2):411–418. https://doi.org/10.1111/j.1432-1033.1994.tb19008.x
Dubey M, Singh AK, Awasthi D et al (2015) L-Plastin S-glutathionylation promotes reduced binding to β-actin and affects neutrophil functions. Free Radic Biol Med 86:1–15. https://doi.org/10.1016/j.freeradbiomed.2015.04.008
Funding
This work was supported by grants from the National Natural Science Foundation of China (grant number 81971795) and Natural Science Foundation for Excellent Young Scientists of Shanxi Province (grant number 20191D211351).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants performed by any of the authors. The principles of the Guide for the Care and Use of Laboratory Animals protocol, published by the Ministry of the People’s Republic of China, were followed.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, N., Liang, Xr., Bai, X. et al. Novel ratio-expressions of genes enables estimation of wound age in contused skeletal muscle. Int J Legal Med 138, 197–206 (2024). https://doi.org/10.1007/s00414-023-03095-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-023-03095-x