Abstract
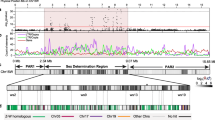
B chromosomes are found in numerous plants and animals. These nonessential, supernumerary chromosomes are often composed primarily of noncoding DNA repeats similar to those found within transcriptionally “silenced” heterochromatin. In order to persist within their resident genomes, many B chromosomes exhibit exceptional cellular behaviors, including asymmetric segregation into gametes and induction of genome elimination during early development. An important goal in understanding these behaviors is to identify unique B chromosome sequences and characterize their transcriptional contributions. We investigated these properties by examining a paternally transmitted B chromosome known as paternal sex ratio (PSR), which is present in natural populations of the jewel wasp Nasonia vitripennis. To facilitate its own transmission, PSR severely biases the sex ratio by disrupting early chromatin remodeling processes. Through cytological mapping and other approaches, we identified multiple DNA repeats unique to PSR, as well as those found on the A chromosomes, suggesting that PSR arose through a merger of sequences from both within and outside the N. vitripennis genome. The majority of PSR-specific repeats are interspersed among each other across PSR’s long arm, in contrast with the distinct “blocks” observed in other organisms’ heterochromatin. Through transcriptional profiling, we identified a subset of repeat-associated, small RNAs expressed by PSR, most of which map to a single PSR-specific repeat. These RNAs are expressed at much higher levels than those arising from A chromosome-linked repeats, suggesting that in addition to its sequence organization, PSR’s transcriptional properties differ substantially from the pericentromeric regions of the normal chromosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thousands of plant and animal genomes harbor B chromosomes, which are supernumerary to the normal, or A, chromosomes (Palestis et al. 2004). In most cases, B chromosomes are nonessential for the organism (Camacho et al. 2000; Jones 1991); they must, therefore, segregate with near-perfect fidelity or “drive” through unique cellular behaviors that allow them to be transmitted at higher-than-Mendelian frequencies, or both, in order to persist in their resident genomes. Types of drive range from asymmetric B segregation during the mitotic or meiotic divisions of gametogenesis to more extreme effects such as B-induced conversion of the organism to the sex that is more conducive to B chromosome transmission (Hewitt 1976; Jones 1991; Werren 2011). While these unusual chromosome behaviors underscore the fact that B chromosomes can behave very differently than the normal chromosomes, little is known about their underlying mechanisms.
A key to understanding the unique behaviors of B chromosomes is to discern their sequence compositions and contributions to gene expression. Generally, B chromosomes are thought to derive from the pericentromeric regions of A chromosomes (Camacho et al. 2000; Perrot-Minnot and Werren 2001; Werren and Stouthamer 2003), which are typically gene-poor and consist almost entirely of highly repetitive DNA sequences including transposable elements and noncoding satellite repeats. Indeed, previous studies have shown that many B chromosomes carry simple and complex satellite repeats, transposable elements, and in some cases, ribosomal DNA (rDNA) repeats (Huang et al. 2016; Jones et al. 2008; Klemme et al. 2013; Munoz-Pajares et al. 2011; Valente et al. 2014; Van Vugt et al. 2009). However, certain B chromosomes in rye, maize, cichlids, and grasshoppers were found to also contain partial and full copies of protein-coding genes located on the A chromosomes that are involved in mitochondrial metabolism and cell cycle/division (Martis et al. 2012; Navarro-Dominguez et al. 2017; Valente et al. 2014). A number of these B-linked protein-coding genes and also transposable elements and repeats are transcribed, although in most cases it is currently not known whether their expression is functional (Carchilan et al. 2007, 2009; Huang et al. 2016; Navarro-Dominguez et al. 2017). Whether expressed or not, most sequences identified on these B chromosomes are similar to those present on the A chromosomes, a general pattern that is consistent with the idea that these and other B chromosomes originate from A chromosomes within their respective genomes (Huang et al. 2016; Klemme et al. 2013; Martis et al. 2012; Munoz-Pajares et al. 2011; Page et al. 2001; Silva et al. 2014; Valente et al. 2014).
An important goal in B chromosome biology is to better understand the identities, subchromosome-level organization, and expression of abundant sequences present on B chromosomes. Of particular interest are highly repetitive satellite DNAs, which make up a large part of pericentromeric heterochromatin on both A and B chromosomes. Traditionally, these sequences were considered to be transcriptionally inert. However, studies in yeast, flies, and plants have shown that certain complex satellite repeats (≥ 100 bp in DNA monomer length) can generate RNA products ranging from thousands of nucleotides to as few as tens of nucleotides in length (Menon and Meller 2012; Ugarkovic 2005; Usakin et al. 2007). In some cases, longer RNAs can serve as precursors for small noncoding RNAs (21–30 nucleotides), which have been shown to participate in transcriptional and post-transcriptional gene regulation (Ugarkovic 2005). Broadly, both long and small RNAs corresponding to satellite repeats and transposable elements have been implicated in a range of functions including chromatin structure, genome stability, and regulation of gene expression (Moazed 2009; Pal-Bhadra et al. 2004; Rosic et al. 2014; Volpe et al. 2002). Understanding which B-linked DNA repeats are expressed and the function(s) of their products may provide important insights into the unique segregation and drive behaviors of B chromosomes, especially given that these noncoding sequences make up a large part of pericentromeric heterochromatin.
Previous studies identified several complex satellite repeats carried on a nonessential, supernumerary B chromosome known as paternal sex ratio (PSR), which is present in the genome of the jewel wasp Nasonia vitripennis (Eickbush et al. 1992; Nur et al. 1988). The origin of PSR is currently not known, although previous studies suggested that it either derives from a normal N. vitripennis chromosome through an intraspecific genome fragmentation or chromosome duplication event, or it instead originates from a chromosome of another wasp species that entered into the N. vitripennis genome through interspecific hybridization (Eickbush et al. 1992; Perfectti and Werren 2001). Transmitted paternally (i.e., solely via the sperm) to new progeny, PSR causes the complete elimination of the paternal half of the wasp’s genome, but not itself, during anaphase of the first embryonic mitotic division (Nur et al. 1988; Swim et al. 2012). This action converts fertilized eggs that should become diploid females into PSR-transmitting haploid males, an essential component of PSR inheritance. Screening of genomic DNA libraries made from PSR-containing males led to the identification of three complex satellite repeats, PSR2 (171 bp), PSR22 (183 bp), and PSR105 (214 bp), which are highly enriched or located exclusively on PSR (Eickbush et al. 1992; Nur et al. 1988). Three additional complex repeats, NV85 (175 bp), NV104 (162 bp), and NV126 (110 bp), were found only in the PSR(−) genome, while a fourth, NV79 (94 bp), is present on both PSR and one or more of the A chromosomes (Eickbush et al. 1992). Through Northern blotting, it was determined that none of these repeats produce transcripts at detectable levels (Eickbush et al. 1992). Interestingly, two highly conserved, AT-rich palindromic regions resembling sequences that bind certain proteins like transcription factors were discovered within the PSR2, PSR22, and PSR105 repeats (Eickbush et al. 1992). Additionally, subsequent studies showed that large deletions of the PSR chromosome that removed high copy numbers of the PSR22 repeat caused increased appearance of F1 female progeny produced from PSR crosses, indicating either reduced strength of genome elimination or instead an unstable segregation and loss of PSR resulting from deletion of PSR22 repeats (Beukeboom and Werren 1993). Based on the lack of evidence that PSR22 is transcribed and due to the presence of palindromic regions within this satellite repeat, it was proposed that PSR could induce genome elimination by titrating away essential chromatin-associated factors from the paternal genome through their binding to PSR22’s conserved palindromic regions (Eickbush et al. 1992).
More recent work, however, has demonstrated that other repetitive sequences located on PSR are indeed expressed. Transcriptional profiling of PSR(−) and PSR(+) wasp testes identified nine polyadenylated transcripts ranging between ~ 500 and 1500 bp in length that were present in PSR(+) but not PSR(−) testes (Akbari et al. 2013). Four of these sequences had low to moderate protein-coding potential, while the other five were strongly predicted to be noncoding. The corresponding chromosomal sequences for two of the three most highly expressed of these transcripts were mapped by DNA fluorescence in situ hybridization (FISH) to the PSR chromosome (Akbari et al. 2013), suggesting that these transcripts are produced by PSR. Moreover, none of these transcripts matched in sequence to the PSR2, PSR22, or PSR105 repeats.
While the discovery of PSR-encoded, polyadenylated transcripts makes it possible that PSR induces genome elimination by expressing trans-acting molecules, much remains to be understood about the extent of PSR’s transcriptional contributions or the subchromosomal organization of its transcribed repetitive sequences. Indeed, these characteristics remain largely enigmatic for B chromosomes in general. Here, we have cytologically mapped the previously identified PSR-specific repeats, many of which are known to be transcribed, to the subchromosomal level. We also performed transcriptional profiling to determine if small RNAs are produced by PSR’s sequences. Our studies uncovered a subset of small RNAs deriving primarily from three PSR-specific repeats unique to PSR. These small RNAs represent a very small number of the total individual sequences expressed in the small RNA transcriptome, but they are expressed at unusually high levels compared to small RNAs from comparable repeats located on the A chromosomes. Most PSR-specific repeats that produce small or long RNAs are concentrated on PSR’s long arm and interspersed among one another. Finally, we found that PSR contains both “foreign” sequences (i.e., those not found on the A chromosomes) and those present in the normal wasp genome, suggesting that its origin is more complex than originally thought.
Results
PSR contains both unique repetitive sequences and those also present in the PSR(−) genome
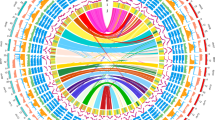
We began by focusing on three complex satellite repeats that were identified from screening of wasp genomic DNA libraries and determined to be located almost entirely or exclusively on the PSR chromosome (Eickbush et al. 1992; Nur et al. 1988). Subsequent analyses of PSR-deletion lines provided a means for generating a map approximating the distribution of these repeats; specifically, this map depicted PSR2, PSR22, and PSR105 at small, distinct regions on the long arm of PSR (McAllister et al. 2004). In order to complement these earlier studies, we performed DNA FISH to visually map the general positions and distributions of these repeats at the subchromosomal level (see “Methods”). Consistent with previous studies, all three of these repeats mapped exclusively to the PSR chromosome, although their individual locations varied (Fig. 1). Specifically, PSR22 signal spanned continuously across the long arm of PSR, except for a small region on its distal tip, and it was also located in a small region on the short arm near the constriction that is likely to be the centromere (Fig. 1). In contrast, PSR2 signal was present across PSR’s short arm and also in a small region on the extreme distal tip of the long arm, outside of the PSR22 signal (Fig. 1). PSR105 signal was confined to a single region on the proximal left arm of PSR that overlapped with the very small region of PSR22 there (Fig. 1). Together these three repeats collectively spanned the entirety of PSR, and none were detected on any of the A chromosomes (Fig. 1).
Enrichment of unique repeats on the long arm of the PSR chromosome. DNA fluorescence in situ hybridization (FISH) shows that PSR carries 11 of 12 previously identified DNA repeats in N. vitripennis. Three of these twelve repeats are present on PSR and the normal chromosomes, while eight are exclusively located on PSR. Ten of the repeats co-hybridize with one another in an overlapping manner across PSR’s long arm. White arrows in the top panels depict PSR at low magnification for each repeat; PSR and its hybridization patterns are shown at higher magnification in the lower panels for each repeat. The blue and red arrows indicate single foci of NV104 and NV85 on single A chromosomes, respectively. Bottom right schematic: the general positions of each mapped repeat are depicted across PSR’s short and long arms. PSR22 is co-stained with each of the other repeats as a point of reference. PSR104, which does not hybridize to PSR, is not shown
We similarly visualized the positions of four additional complex repeats, NV79, NV85, NV104, and NV126, which were previously identified from the analysis of PSR(−) wasp genomic DNA (Eickbush et al. 1992). NV79 signal appeared in varying intensities at distinct locations within the pericentromeric regions of four of the five A chromosomes, reflective of differing copy numbers at these locations (Fig. 1). A region of NV79 was also present on the distal half of PSR’s long arm, overlapping with PSR22; this region was approximately the same size as the largest region in the PSR(−) genome (Fig. 1). NV85 signal was present in a very small region on a single A chromosome, but, interestingly, it spanned broadly across the long arm of PSR, thus also overlapping completely with PSR22 (Fig. 1). NV126 signal showed a similar pattern to NV85, existing across the long arm of PSR as well as in a small region on each of two different A chromosomes (Fig. 1). NV104 signal was present in one region on a single A chromosome but not on PSR, based on our level of resolution (Fig. 1). Thus, three of the four repeats previously identified from the PSR(−) genome are also located on the PSR chromosome. In addition, hybridization with mixed combinations of these probes to the A chromosomes revealed that the single A chromosome containing NV104 is the same one that harbors the single region of NV85 repeat as well as a region of the NV79 repeat (Fig. S1). The presence of these three repeats on a single A chromosome, two of which are also on PSR, supports the possibility that at least part of the PSR chromosome is derived from this particular N. vitripennis chromosome.
We next visually mapped the subchromosomal locations of the repeats corresponding to the top 5 most highly expressed PSR-specific, polyadenylated transcripts in the testis (Akbari et al. 2013). All five of these sequences completely overlapped with PSR22 across PSR’s long arm and in a very small region on the proximal left arm (Fig. 1 and Fig. S2). This pattern is in contrast to many other simple and complex repeats, which are organized into tandem-repeated arrays that form distinct regions or “blocks” on the chromosomes (Bonaccorsi and Lohe 1991; Ferree et al. 2014; Ferree and Prasad 2012; Lohe et al. 1993). The majority of the PSR-specific repeat signals in our cytological experiments overlap, suggesting that these sequences are interspersed among each other across a given region, either homogeneously or as separate small tandem arrays that collectively appear at the cytological level as homogeneous.
Overall, the repetitive sequences examined here can be classified as either exclusive to PSR (PSR2, PSR22, PSR105, PSR4317, PSR1539, PSR5885, and PSR4656), exclusive to the A chromosomes (NV104), or present on both PSR and one or more A chromosomes (NV79, NV85, and NV126). PCR amplification using either PSR(−) or PSR(+) genomic DNA (gDNA) as a template largely confirmed these findings with two exceptions (Fig. 2). First, NV79 was only detected in PSR(+) gDNA, in contrast to our DNA FISH analyses (Fig. 1) and previous Southern blot analyses (Eickbush et al. 1992) which both indicated that NV79 is present on PSR and the A chromosomes. By further analyzing the NV79 repeat, we found that it shares some similarity with a common repetitive element class found in the PSR(−) genome (satellite BRP2) and is mostly likely cross-hybridizing with those repeats. Based on PCR, the canonical NV79 repeat seems to be exclusive to PSR. The second exception is PSR4656, which can be detected with PCR in both PSR(−) and PSR(+) genomic DNA but is only seen cytologically on PSR. A likely explanation for this inconsistency is that this sequence is present in the PSR(−) genome, but at too low of copy number to be detected with DNA FISH.
Certain PSR repeats are organized into tandem arrays, while others are distantly spaced. Each repeat was amplified from genomic DNA from PSR(+) males and PSR(−) males. Amplification of PSR2, PSR22, PSR105, NV79, NV85, NV104, and NV126 generates laddered products from PSR(+) gDNA (right column in each condition). The NV repeats are also amplified from PSR(−) gDNA due to the presence of these repeats on the normal N. vitripennis chromosomes. In contrast, amplification of PSR4317, PSR1539, PSR5885, PSR5643, and PSR4656, each of which produces long transcripts (Akbari et al. 2013), yielded single products (expected sizes of 241, 410, 282, 232, and 272 bp, respectively), despite that these sequences appear to be in high copy number across PSR’s long arm based on DNA FISH. These patterns suggest that the copies of these sequences are located at distances of 10 kb or greater from one another, thus preventing amplification across multiple copies. PSR4656 was amplified from PSR(−) gDNA, despite our failure to detect this sequence by DNA FISH, suggesting that this sequence or a similar one exists in very low copy number in the PSR(−) genome such that it cannot be detected with this method. The nearest size markers (ladders not shown) are indicated by arrowheads in each panel. Each individual product appears as the predicted size
Amplification of the previously identified satellite repeats produced a laddered pattern of products (Fig. 2, top row), suggesting that the individual copies of a given repeat are in close association with one another. This finding is consistent with previous results demonstrating that cloned blocks of these repeats were both tandem-repeated and homogeneous in nature (Eickbush et al. 1992). In contrast, amplification of the top 4 PSR-specific transcript sequences produced single products despite the use of long PCR extension times (Fig. 2, bottom row). This pattern suggests that these sequences are not in close proximity to one another and are most likely either embedded in larger blocks of unknown sequence or are dispersed among other repeats at a distance such that amplification with PCR is difficult or impossible.
Identification of a new PSR-specific transcript
All nine previously identified PSR transcripts were discovered through recovery of polyadenylated RNA from the PSR(+) testis (Akbari et al. 2013). In order to more exhaustively explore the transcriptional potential of PSR, we performed transcriptional profiling of long RNAs from testis that included both polyadenylated and nonadenylated sequences (see “Methods” for details). In addition to finding all nine previously identified PSR-specific polyadenylated transcripts, we identified one new, long transcript, labeled PSR8495 (Table S1). This sequence is only 250 nucleotides in length and thus strongly predicted to be noncoding (Table S1). Like the DNA sequences corresponding to the top 5 highest expressed PSR-specific polyadenylated transcripts, PSR8495 spanned across PSR’s long arm, overlapping completely with PSR22 (Fig. S2). Thus, we have been able to portray the subchromosomal distributions and relative abundances of 12 repetitive sequences located on the PSR chromosome or the A chromosomes or both. Moreover, 11 of these 12 sequences are located on the PSR chromosome, and ten span across PSR’s long arm in a visually overlapping pattern.
PSR expresses a unique set of repeat-associated small RNAs
In order to more thoroughly investigate the transcriptional potential of PSR, we profiled RNAs that were size-selected below ~ 50 bp in both PSR(−) and PSR(+) testes and carcasses (somatic tissues without testes). In particular, we aimed to identify small noncoding RNAs (21–33 bp in length) produced uniquely by PSR, with special attention to small RNAs generated from known and perhaps new repetitive elements located on PSR. To this end, we sequenced the small RNA transcriptomes of each tissue condition and then subtracted away all sequences present in both genotypes, leaving only those expressed exclusively in the PSR genotype. Here, we report several basic characteristics of small RNAs expressed from the A chromosomes for context before discussing PSR-specific sequences.
Out of > 20 million high-quality reads per condition, a total of ~ 3.7 million distinct small RNA sequences were commonly identified in both PSR(−) and PSR(+) testes; these sequences represent the small RNA population derived from the A chromosomes (Table 1). Using specific criteria (see “Methods”), we identified from these sequences a total of 168 putative microRNAs (miRNAs), 53 of which are found in miRBase while the remaining sequences are novel, 49 putative endogenous small interfering RNAs (siRNAs), and 697 repeat-associated small interfering RNAs (or putative PIWI-associated small RNAs, or piRNAs), all of which were expressed at levels higher than 10 traces per million (TPM) (Table S2). Many of these sequences were map-able to loci in the sequenced (PSR(−)) genome (Nvit2.2) or to transposable elements or noncoding repeats in the N. vitripennis repeat library (Tables S2 and S3) (Werren et al. 2010). Interestingly, we found a relatively small set of 64 sequences expressed at ≥ 10 TPM and found exclusively in the PSR(+) genotype (Table S4). Seventeen of these 64 PSR-specific small RNAs do not match any known sequences in the annotated PSR(−) genome, the N. vitripennis repeat library, or the long RNA testis transcriptomes (Table S4) (Akbari et al. 2013). The average length of these particular sequences is 32.5 nucleotides. Additionally, many of them begin with a uracil residue and are predicted to overlap in a reverse complementary manner by 10 nucleotides at their 5′ ends with other small RNAs found in trace amounts in the PSR(+) transcriptome (Table S4), indicative of “ping pong” amplification of piRNAs (Czech and Hannon 2016). Together, these characteristics suggest that these particular sequences do not function as miRNAs or siRNAs but are likely instead piRNAs.
The remaining 47 PSR-specific RNAs vary in length, with an average value of 22 nucleotides. Each of these sequences matches at > 98% identity to one of the three previously described PSR repeats, PSR2, PSR22, and PSR105 (Table S4). Eighty-five percent of all PSR-specific small RNA reads match to these three repeats, with 64% matching specifically to the PSR22 repeat alone. The contribution of PSR-specific small RNAs to the total small RNA testis transcriptome is very small, representing roughly 0.1% of all small RNA reads and less than 0.002% of all individual small RNA sequences present in the testis, assuming that these PSR-specific small RNAs are representative of most small RNAs produced by PSR. These contributions of PSR to the small RNA transcriptome stand in strong contrast to the fact that PSR represents approximately 5.7% of the nuclear content [41]. Despite this small overall contribution by PSR, the levels of small RNAs corresponding to each of the three PSR-specific repeats are much higher than the levels of small RNAs derived from individual repeats found on the A chromosomes (Fig. S3). The fact that several of these PSR(−) DNA repeats are also present on PSR demonstrates that some but not all DNA repeats carried by PSR are expressed at unusually high levels as small RNAs.
In order to better understand the sequence characteristics of the 47 small RNAs that match the PSR-specific repeats, we performed alignments of these small RNAs to each other and also to their corresponding DNA repeats. These small RNAs formed eight clusters based on high similarity to one another (Fig. 3, Fig. S4, and Fig. S5). The sequences within each cluster differ by no more than two nucleotide polymorphisms or by five or fewer nucleotides in length. In one case, three clusters (2–4) align with one another by overlapping 5–10 nucleotides (Fig. 3). Small RNAs belonging to clusters 1, 2, 4, and 5 match to discrete regions within the PSR22 repeat (Fig. 3). Sequences belonging to cluster 6 match to a region within PSR105 (Fig. S4), and sequences in clusters 7 and 8 match to regions within PSR2 (Fig. S5). Interestingly, the most highly abundant set of small RNAs, which comprise cluster 1, overlaps the entire 12 bases of the highly conserved palindrome located toward the 5′ end of the canonical PSR22 repeat (Fig. 3). Additionally, sequences making up cluster 4 correspond to eight of the 10 bases of the second palindrome of PSR22 (Fig. 3). All small RNAs matching to the PSR2 and PSR105 DNA repeats overlapped with their 5′ palindrome regions (Fig. S4 and Fig. S5). The correspondence of small RNAs to the palindromes within the PSR2, PSR22, and PSR105 repeats may explain the high conservation of these regions (see “Discussion”).
PSR-specific small RNAs derive from a subset of noncoding repeats. a The most abundant PSR-specific small RNAs (shown in colored text) form five different “clusters” based on alignment to one another and derive from the PSR22 repeat. Trace values are shown in parentheses for each small RNA sequence. b The canonical PSR22 DNA repeat is shown, with sequences highlighted in colored text depicting the corresponding small RNAs shown in a. Many of these small RNAs overlap two highly conserved palindromic regions within the repeat (underlined). c A longer RNA precursor containing the most abundant PSR22-specific small RNAs (above arrow) is predicted to form a perfect hairpin (below arrow), a structure that is likely to be important in processing of the small RNAs from this molecule. The region corresponding to the small RNAs is shown in red (and underlined in the hairpin), and the antisense sequence is underlined in black
The biogenesis of small RNAs begins with their processing from longer RNA precursors (Kim et al. 2009). In order to identify the precursors of the PSR-specific small RNAs, we searched for identity to sequences present in the previously published transcriptome of polyadenylated transcripts in the testis (Akbari et al. 2013). These searches yielded 10 matching transcripts, five of which corresponded specifically to PSR22-derived small RNAs, three to PSR105-derived small RNAs, and two to PSR2-derived small RNAs (Table S5). These transcripts appeared to be rearranged versions of their corresponding canonical repeats, and each transcript contains between one and five of the small RNAs from their respective clusters (Fig. 3, Fig. S4, and Fig. S5). Additionally, four of the 10 precursors are perfect palindromes and are thus predicted to form perfect hairpins as secondary structure, whereas the remaining six precursors are imperfect palindromes, likely forming imperfect hairpins (Fig. 3, Fig. S4, and Fig. S5). No long transcripts were found that reflect the exact sequence composition of the canonical PSR repeats. Thus, it is likely that the corresponding small RNAs are produced from a subset of variants of these repeats, and not from the canonical repeats, themselves. Additionally, we note that these small RNAs are unique because their characteristics match those of multiple conventional small RNA classes. In particular, these small RNAs appear to be processed from hairpin precursors like miRNAs, yet they do not match complementarily to any detectible mRNAs but instead to noncoding repeats, similar to repeat-associated siRNAs (rasiRNAs). We acknowledge that the term “rasiRNA” is not commonly used because many of these repeat-associated small RNAs formerly classified as rasiRNAs have been subsequently reclassified as piRNAs due to their association with PIWI and other related proteins (Ku and Lin 2014). Similar to the 17 PSR-specific small RNAs that do not match to any known wasp sequences (see above), most of these 47 small RNAs matching to PSR2, PSR22, and PSR105 also show signatures of participation in ping pong amplification (Table S4), although their average length is shorter than piRNAs. Therefore, we cannot say with certainty if these particular small RNAs are indeed piRNAs without testing for their physical association with PIWI-family proteins, and so, we generally refer here to these small RNAs as rasiRNAs.
Finally, we tested whether the nine previously identified, long polyadenylated transcripts produced by PSR or the new one reported here serve as precursors for small RNAs. We found no traces of small RNAs matching to the nine previously identified polyadenylated transcripts, strongly suggesting that they are not precursors for small RNAs. However, we did find that the newly identified PSR transcript (PSR8495) matches to a subset of the PSR-specific rasiRNAs. However, PSR8495 is not predicted to form a stable hairpin structure, as are the 10 long precursors identified above.
Subcellular localization of PSR-expressed RNAs
In order to investigate the subcellular localization of PSR-specific RNAs, which could provide additional clues regarding their functions, we used RNA FISH to visualize the highest expressed small RNA and long transcript produced by PSR—specifically a PSR22-derived small RNA and the PSR4317 transcript, respectively—in whole-mount testes (see “Methods”). We focused on testes from male pupae in the yellow body-red eye stage; at this time, the testis contains cysts of mitotically dividing germ cells, or cystocytes, at the anterior end, as well as more advanced germ cells that are undergoing spermiogenesis at the posterior end (Fig. 4a). To visualize PSR4317, we used a pool of fluorescently labeled DNA probes that tile in a complementary manner across the PSR4317 transcript. In PSR(+) testes, we observed a single bright focus of PSR4317 in the nucleus of each cystocyte (Fig. 4b). Additionally, PSR4317 signal was concentrated in a granular appearance within the cytoplasm of these cells (Fig. 4b). Treatment with RNase A caused all PSR4317 signal to disappear, demonstrating that the patterns reflect hybridization to RNA and not cognate DNA sequences (Fig. S6). Additionally, no signal above background was observed in PSR(−) testes (Fig. S6), further arguing that the detected signal was PSR4317 RNA.
Subcellular localization of PSR-expressed RNAs in the testis. a A whole testis from a yellow body-red eye pupa stained for DNA is shown. The cysts of mitotically dividing cystocytes are circumscribed by white lines, and red arrows point to large nuclei intermixed among the cysts. b Top row: the PSR4317 transcript (red) appears within large polyploidy nuclei (red arrows) that are intermixed among the cysts of mitotically dividing germ cells (circumscribed by white line). PSR4317 RNA also localizes within the germ cell cytoplasm but not in their nuclei. This pattern suggests that 4317 is transcribed within the polyploidy cells and transported into the germ cells. Middle row: a different polyadenylated transcript (NV11286, shown in green), which encodes an uncharacterized chromatin protein, similarly localizes within the cytoplasm of the germ cells but is not present at all in the polyploidy nuclei. Additionally, one or two small foci of this transcript can be seen in some germ cell nuclei; these foci are likely the sites of transcription. Bottom row: RNA polymerase II phosphorylated at serine residue 2 (blue), a transcriptionally active form of this enzyme, localizes within both the large polyploid nuclei as well as the germ cell nuclei. Scale bar is 10 μM. c A region within the testis of a PSR(+) male showing localization of the highest expressed small RNA (red, also indicated by red arrows in the middle row), which matches to the PSR22 DNA repeat. This small RNA is seen in small foci within the large polyploid nuclei, but not within the germ cell nuclei. Additionally, the PSR22 small RNAs (red arrowheads in the bottom row) localize adjacent to the large number of PSR22 DNA repeats green) within the polyploid nuclei. These small RNAs only overlap with a very small amount of the PSR22 repeats. Scale bar is 5 μM
We speculated that the single focus of PSR4317 RNA in each nucleus is the site of nascent transcription, as has been shown for other transcripts (Femino et al. 1998), and that the perinuclear signal reflects PSR4317 transcripts that have been exported to the cytoplasm. To indirectly test this hypothesis, we visualized a different testis-specific transcript (NV11286) encoding a putative chromatin-associated protein that was uncovered in a previous study (Ferree et al. 2015). Localization of this transcript mirrored that of PSR4317; that is, this transcript localized to a single bright region in the nucleus, likely the site of transcription, and also within the cytoplasm (Fig. 4b). Thus, the cytoplasmic localization of PSR4317 is consistent with the fact that this transcript is polyadenylated and the notion that this transcript, which contains a predicted open reading frame, may be translated. To more broadly confirm the transcriptional activity of the cystocytes, we stained testes with an antibody that recognizes active RNA polymerase II (Fig. 4b). As expected, the nuclei of these cells showed bright staining of the active form of this enzyme, thereby adding further credence to the conclusion that the signals of PSR4317 and NV11286 are transcripts produced from the cystocytes themselves.
We also employed a locked nucleic acid (LNA) probe to visualize the top-most expressed PSR22 rasiRNA in PSR(+) testes (see “Methods”). Interestingly, this small RNA was cytologically absent from the cystocytes but appeared instead in multiple bright foci within large nuclei that are interspersed among the cystocytes (Fig. 4c). These nuclei are ~ 5-fold larger in size than the nuclei of the cystocytes, and they have a heterogeneous appearance that resembles polyploid nuclei present in the ovarian nurse cells of Drosophila melanogaster (Dej and Spradling 1999). In support of the likelihood that these large nuclei are polyploid, hybridization with a DNA probe that recognizes PSR22 DNA repeats showed multiple, distinct foci within each large nucleus (Fig. 4c). The same pattern was also found by using a probe that hybridizes to rDNA, which is located in a single locus on an A chromosome (Fig. S7). As expected, neither the PSR22 LNA probe nor the PSR22 DNA probe provided any signal in PSR(−) testes (data not shown). Interestingly, in PSR(+) testes, the PSR22 small RNA foci overlapped with very minor regions of PSR22 DNA but not with the largest regions of this repeat (Fig. 4). These PSR22 small RNA foci may reflect either the site of precursor transcription from a subpopulation of PSR22 repeats, or instead, they may be sites of mature small RNA accumulation.
Expression of PSR repeats across development
In order to better understand the expression of PSR-specific RNAs, we used reverse transcription PCR (RT-PCR) to amplify a handful of these sequences from different tissues and developmental stages. Previous transcriptional profiling experiments (Akbari et al. 2013) and those presented here detected substantial levels of long transcripts produced by PSR in the testis. However, it was not known whether these transcripts are also expressed in somatic tissues. To test this possibility, we examined for the presence of the top 4 highest expressed long transcripts, PSR4317, PSR1539, PSR5885, and PSR4656, in complementary DNA (cDNA) made from somatic and testis tissues. We were able to successfully amplify each of these transcripts from both tissue types (Fig. 5a). Our results suggest that these four PSR-specific transcripts are expressed in easily detectable amounts in the soma, demonstrating they are not preferentially transcribed in the testis.
PSR-specific RNAs are expressed in both somatic and germ line tissues and during most developmental stages. a Nonquantitative RT-PCR amplification shows that PSR-expressed RNAs are present in both the testis and carcass (somatic tissues). RNA expressed from the NV126 repeat, which is also present in the PSR(−) N. vitripennis genome, is also present in both tissue types. Two RNA products were amplified for PSR1539 and PSR4656; in these cases, the larger product (asterisks) correspond in size to the monomer product amplified from gDNA. Thus, the smaller product(s) may reflect spliced products. b All examined PSR-expressed RNAs and NV126 are present in mid-stage (15 h) PSR(+) embryos and larvae, and all but PSR4317 were absent in 0–2-h embryos, when the embryo’s genome has become transcriptionally active. In the case of PSR4317, very slight amplification was detected. c PSR4317 RNA was clearly present in very young embryos immediately following fertilization, suggesting that this transcript may be paternally transmitted via the sperm into the egg cytoplasm
We also tested for expression of these particular transcripts across several key developmental stages: 0–2-h embryos, in which there should be no transcripts produced from the embryo’s genome but instead only from maternal loading; ~ 15-h embryos, which have undergone activation of zygotic genome expression; and larvae in the third (final) instar stage. All four transcripts were present in 15-h embryos and in third instar larvae (Fig. 5b). We detected no expression of PSR1539, PSR5885, or PSR4656 in 0–2-h embryos (Fig. 5b). However, we observed light amplification of PSR4317 at this early developmental time (Fig. 5). In order to confirm this pattern, we tested for expression of PSR4317 in 0–45-min embryos. Interestingly, we obtained stronger amplification of PSR4317 at this earliest time point (Fig. 5c). Amplification using primers for the PSR(−) repeat, NV126, and the PSR-exclusive repeat, PSR22, produced many multiple products—likely reflecting transcriptional read-through of multiple adjacent repeats—and revealed transcription in a similar pattern to the PSR-specific repeats (Fig. 5a, b). Thus, it appears that PSR indiscriminately expresses these RNAs in both somatic and germ line tissues. Additionally, these results also suggest that, while all examined sequences are transcribed by mid-embryogenesis, it is possible that PSR4317 and perhaps other RNAs are transmitted at low but detectable levels into the egg cytoplasm by way of the sperm.
Finally, we point out that amplification of several transcripts, including PSR1539 and PSR4656, produced two major products of differing lengths (Fig. 5a, b). In each case, the longer products matched in length to the single amplified products of the repeat observed when using genomic DNA as a template and the smaller products matched in length to the RNA-seq-identified transcripts (Akbari et al. 2013). While the two amplified products for PSR1539 are present during mid-embryogenesis, only its shorter product is present at the third instar larval stage and in the testis (Fig. 5a, b). It is therefore possible that these product size differences reflect unspliced and spliced products that, at least in the case of PSR1539, are dependent on the particular developmental stage and tissue.
Discussion
B chromosomes have long been viewed as enigmatic elements of eukaryotic genomes regarding their origins and how they interact with their cellular environments in order to facilitate their own transmission. As a means for addressing these aspects, we have more comprehensively investigated the organization of repetitive elements on the paternal sex ratio B chromosome in N. vitripennis. Complementing previous transcriptomic efforts, we also performed transcriptional profiling of small RNAs in order to identify new RNAs produced by PSR and gain insights into the potential functions of such factors.
Unique sequence organization of the PSR chromosome
Previous studies identified a collection of complex repeats located on the PSR chromosome, in addition to several repeats thought to be located only on the wasp’s A chromosomes (Akbari et al. 2013; Eickbush et al. 1992; Nur et al. 1988). Through cytological mapping, we found that most of the repeats located on PSR span across its long arm in an overlapping manner. Our PCR analyses suggest that each of the PSR2, PSR22, and PSR105 repeats is organized into tandem arrays, while the repeats matching the long PSR-specific transcripts are not organized into tandem arrays but, instead, they are distributed apart from one another by more than 10 kb. This interspersion of multiple repeats stands in contrast to other complex repeats in the genomes of different organisms including Drosophila species; in these cases, the repeats are organized into distinct, separated blocks (Bonaccorsi and Lohe 1991; Ferree and Prasad 2012; Lohe et al. 1993). Additionally, several N. vitripennis repeats are organized into separate blocks, including PSR2, PSR105, and several repeats located on the A chromosomes.
Our cytological mapping efforts presented here provide insights into a couple of facets about PSR’s sequence composition and its origin. First, whereas previous mapping of PSR repeats by Southern blotting and chromosome deletion analysis portrayed certain repeats, such as PSR22, as being located in isolated regions (McAllister et al. 2004), our studies reveal that PSR22 spans a much larger and more contiguous distance across the PSR chromosome. Moreover, PSR derivatives deleted for PSR22 also would have concomitantly lost many other overlapping repeats, thus confounding any interpretation of PSR22 as being solely involved in genome elimination based on the deletion analyses alone. Second, PSR carries more repeats that are also present on the A chromosomes than previously thought. In particular, PSR contains large regions of the A chromosome repeats, NV79, NV85, and NV126, or at least very similar variants of these repeats. Interestingly, two of these three repeats, NV79 and NV126, are located together on a single A chromosome. These findings leave open the possibility that PSR originated from the pericentromeric region of this particular chromosome containing these two repeats.
A caveat to this idea, however, is that PSR’s organization with regard to these specific repeats is not identical to their organization on the A chromosomes. Moreover, the existence of PSR-linked repeats that are not present on the A chromosomes must be taken into account. To explain the occurrence of such PSR-specific repeats, it was previously proposed that PSR may have originated from a chromosome of another wasp species through an interspecies hybridization. This possibility was supported by the finding of several of these DNA repeats in the genomes of other wasp species outside of the Nasonia genus (McAllister and Werren 1997). In light of our findings, we propose a modified scenario: that PSR may have arisen through a combination of interspecies hybridization, in which a chromosome or chromosomal fragment containing repeats unique to another species entered into the genome of N. vitripennis through hybridization and underwent subsequent chromosomal rearrangement(s) with one or more N. vitripennis chromosomal regions, thus producing the unique combination of repeats currently present on PSR.
PSR expresses both long transcripts and small noncoding RNAs
Our experiments here extend on previous transcriptomic work in N. vitripennis (Akbari et al. 2013), showing that many of the repeats carried by PSR are transcribed either into long (~ 500–1500 bp in length) polyadenylated RNAs or hairpin-forming precursor RNAs that are processed into rasiRNAs. The initial discovery of polyadenylated transcripts produced by PSR led to speculation that they may function as noncoding RNAs involved in chromatin structure or remodeling (Akbari et al. 2013). In this case, these transcripts would be expected to localize within the nucleus. However, it was also proposed that some of these transcripts might instead encode small proteins. To begin to address these possibilities, we visualized PSR4317, which is the highest PSR-expressed transcript, present in the testis at levels three times higher than the next highest PSR-expressed transcript, and has the largest predicted ORF for any of the PSR-expressed transcripts. Our RNA FISH experiments showed that PSR4317 localizes primarily to the cytoplasm of the cystocytes during interphase between their mitotic divisions, with a single bright region within each nucleus, which is likely the site of their transcription. These patterns closely mirror those of a different, male-specific transcript that is expressed from the PSR(−) genome and predicted to encode an uncharacterized chromatin protein (Ferree et al. 2015). The lack of broad nuclear localization of PSR4317 and its presence in the cytoplasm is consistent with it expressing a protein. This finding underscores the importance of future studies aimed at testing this hypothesis, as well as further analyses of the remaining PSR-expressed transcripts.
The small RNA repertoire expressed by PSR appears to contain many fewer distinct sequences than expected based on the size of this B chromosome. At 5.7% of the total nuclear content (Reed 1993), PSR would be expected to contribute ~ 185,000 unique small RNA sequences. Instead, we detected only 64 unique small RNAs produced by PSR. However, interestingly, the expression levels of small RNAs corresponding to each of the three repeats PSR2, PSR22, and PSR105 are at least 10-fold higher than those from small RNAs matching other repeats produced from the PSR(−) genome. In addition, a large fraction of small RNAs corresponding to PSR22 overlap the two highly conserved palindromes within this repeat. Although the function(s) of these small RNAs are currently unknown, their matching to the palindromes and their high expression levels indicate that the small RNAs may indeed be functionally relevant.
Perhaps, the most interesting hypothesis is that RNAs produced by PSR may function in paternal genome elimination. For example, small RNAs could act as guides for recruiting chromatin proteins to the cognate chromosomal repeats, in a manner similar to what is believed to occur with small RNA-mediated facilitation of heterochromatin formation and maintenance (Holoch and Moazed 2015). One idea is that such an effect may alter chromatin so that PSR itself is somehow immune to the genome elimination effect. Given that much of PSR’s length contains PSR22 repeats, this “immunity” would be expected to protect much of the chromosome. Indeed, a recent study showed that the chromatin state of PSR is distinct from the rest of the nuclear chromatin. In particular, PSR was largely devoid of two important chromatin marks, H3K27me1 and H4K20me1 (Aldrich et al. 2017). Another idea is that PSR-expressed RNAs themselves, or perhaps proteins encoded by PSR4317 or other long transcripts, may somehow specifically target the paternally inherited A chromosomes.
PSR is distinct from other B chromosomes regarding sequence composition and expression
Previous groups have explored the sequence composition and expression of other B chromosomes in a wide range of distantly related organisms. In light of these studies, our work provides insights into ways in which PSR is both similar to and distinct from these other B chromosomes. For example, certain B chromosomes in cichlid fish, rye, grasshoppers, and other organisms have been found to carry a range of protein-coding genes, pseudogenes, transposable elements, and tandem arrayed noncoding repeats (Huang et al. 2016; Jones et al. 2008; Klemme et al. 2013; Munoz-Pajares et al. 2011; Valente et al. 2014; van Vugt et al. 2005). It is known that PSR contains at least one retrotransposon known as NATE, as well as an abundance of complex satellite repeats (Akbari et al. 2013; Eickbush et al. 1992; McAllister 1995). However, it is currently not known if PSR also carries protein-coding genes, especially those that are homologous to genes located on the wasp’s A chromosomes; addressing this specific issue will benefit from the application of long-read DNA sequencing technologies. Nevertheless, a clear difference is that whereas most other examined B chromosomes contain DNA sequences, including both protein-coding and noncoding sequences, which are largely similar to the ones located on the A chromosomes of their respective genomes (Carchilan et al. 2009; Klemme et al. 2013; Martis et al. 2012; Munoz-Pajares et al. 2011; Navarro-Dominguez et al. 2017; Valente et al. 2014), the majority of PSR’s sequences appear to be unique, either nearly or entirely absent from the wasp’s A chromosomes. Additionally, consistent with these patterns, PSR produces RNAs that are distinct from those present in the PSR(−) transcriptome.
The fact that PSR’s DNA sequence composition and expressed RNAs are largely distinct from those of the PSR(−) genome and transcriptome may be explained by (i) a partial, interspecific origin of this B chromosome and (ii) PSR’s mode of transmission being solely through the male germ line, in which there is no recombination (Whiting 1968), thereby preventing large-scale movement of sequences from the A chromosomes onto PSR. It also may be the case that movement of DNA sequences between the A chromosomes and PSR through transposable element mobility, as has been proposed to occur in other B chromosome systems (Valente et al. 2014), may be somehow hindered in the male germ line of N. vitripennis. Indeed, we found only trace reads corresponding to the PSR-enriched retrotransposon, NATE, in our current small RNA transcriptome analyses, and also in the long testis transcriptome (Akbari et al. 2013). Any of these features, or a combination thereof, may have facilitated the distinctness of PSR from the A chromosomes, a characteristic that is likely to be important for the ability of PSR to impose such an extreme form of drive—complete elimination of the paternally inherited half of the wasp’s genome during each generation.
Conclusions
Our results here suggest that PSR’s DNA sequence composition and transcriptional contributions are remarkably distinct from those of the N. vitripennis A chromosomes. These differences likely stem from PSR’s origin, its uniparental mode of inheritance, and unique reproductive developmental characteristics of the wasp. We propose that these characteristics have ultimately shaped the ability of PSR to cause complete elimination of the wasp’s paternal genome. Additionally, these studies have provided new factors that can be experimentally tested as potential effectors of this extreme manifestation of genome conflict.
Methods
Wasp lines and propagation
Our experiments employed two different wasp lines: a PSR(−) symbiont-free line, AsympC, and another of the same genetic background that contains the PSR chromosome. Because the presence of PSR leads to all-male broods and, therefore, no females, we maintained PSR by crossing PSR(+) males from this line to AsympC virgin females pairwise, and selecting for all-male broods.
Testis dissection and fixation
Testes were dissected from male pupae in the yellow body-red eye stage. Tissues were removed from male pupae in 1× PBT (phosphate-buffered saline and 0.5% Triton-X 100). For whole-mount work, testes were fixed in 800 μl of heptane and 200 μl of 4% paraformaldehyde for 20 min on a platform rocker, and subsequently washed three times in 1 ml of 1× PBT. For preparation of chromosome spreads, testes were fixed for 4 min in a drop of 50% acetic acid/2.5% paraformaldehyde on a coverslip, squashed onto a clean slide, snap frozen in liquid nitrogen, dehydrated for 10–20 min in 100% ethanol, and air-dried. A complete demonstration of this procedure can be viewed in Larracuente and Ferree (2015).
Fluorescence in situ hybridization and antibody staining
For DNA FISH to testis chromosome spreads, we followed a previously described procedure (Larracuente and Ferree 2015) with no deviations. The probes for DNA FISH were chemically synthesized DNA oligonucleotides (25–30mers) that were conjugated with either Alexa488, 555, or 633 (Integrated DNA Technologies). DNA FISH probe sequences are shown in Table S6.
To visualize the PSR4317 transcript, we employed a probe cocktail of 33 different DNA oligonucleotides each ranging between 19 and 21 nucleotides in length and conjugated with the Quasar 570 fluorophore (Stellaris), which tiled complementarily across the length of each of these transcripts. Similarly, to visualize a control testis-specific protein-coding transcript (NV11286), we used a cocktail of 48 different DNA oligonucleotides of similar length as described above and conjugated with Quasar 670 (Stellaris). To visualize the highest expressed PSR-specific small RNA, we utilized an LNA/DNA-based probe (Exiqon) with the following sequence: 5′-AATATCCAATCATAAGTCGAGACTTT-3′. This probe was DIG-labeled at both 5′ and 3′ ends. The identity of the specific LNA nucleotides is proprietary, but the batch number (623899) can be referenced in order to request synthesis of the same probe. For both RNA probe types, we performed RNA FISH on whole-mount fixed testes according to a procedure provided by Exiqon. This procedure involved the use of the company’s hybridization buffer. The hybridization temperature for the LNA/DNA probe was 37 °C, whereas the temperature for the DNA probe cocktails was 50 °C.
Immunostaining
A primary mouse antibody recognizing active RNA polymerase II phosphorylated at serine residue 2 (Abcam) was used at a dilution of 1:200 in 1× PBT. Tissues were hybridized to diluted antibodies overnight at 4 °C on a platform rocker, washed three times with 1× PBT, stained with an Alexa633-conjugated, antimouse secondary antibody (Thermo Fisher Scientific) for 1 h, and washed again three times in 1× PBT. Following removal of all wash buffer, stained tissues were slide mounted in Vectashield containing DAPI (Vector Laboratories).
Fluorescence microscopy
Fluorescent signals on chromosome spreads were collected by using a Leica epifluorescence microscope. For stained, whole-mount tissues, images were collected with a Leica TCS SPE confocal microscope. Images collected in separate channels were merged by using Adobe Photoshop CS5 v. 12.
Nucleic acid purification, reverse transcription, and polymerase chain reaction
RNA was purified using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. Prior to precipitation with ethanol, RNA was further extracted with phenol:choloroform (5:1 pH 4.5) and chloroform. RNA was resuspended in nuclease-free water and genomic DNA was digested using the TURBO DNA-free kit (Ambion) according to the manufacturer’s instruction. Third instar larvae and pupae samples were prepared from 20 individuals, while each embryo sample contained approximately 100 carefully aged eggs collected from ~ 20 PSR(−) females mated with ~ 20 PSR(+) males. For the 0–2-h embryo sample, embryos were collected immediately after 2 h of laying, while embryos for the 15-h sample were allowed to age 13 h before collection.
DNase-treated RNA (1.6 μg) was used for each 20 μl reverse transcription reaction along with 200 U SuperScript II reverse transcriptase (Invitrogen), 1× first strand buffer, 1 μM dNTPs, 5 mM DTT, 25 μM random hexamer primers, and 5 μM oligo dT primers. Nucleic acids were denatured alone at 65 °C for 5 min before the addition of other reagents. Reactions were then incubated at 25 °C for 10 min followed by 50 °C for 50 min; 2 μl cDNA was used for each 25 μl PCR reaction.
PCR reactions were carried out using GoTaq reagents (Promega). Each 25 μl reaction contained 1× reaction buffer, 2 mM MgCl2, 1 mM dNTPs, 0.4 μM of each primer (Table S7), and 1.25 U GoTaq DNA polymerase. Thermocycling conditions were as follows: 95 °C for 2 min followed by 35 cycles of 95 °C for 30 s, 54–60 °C for 30 s, and 72 °C for 30 s. Reactions were separated on a 1% agarose gel containing 0.5 μg/ml ethidium bromide and visualized using a BioDoc-It imaging system (UVP).
Genomic DNA for PCR was prepared by homogenizing 20 Nasonia adult males in 400 μl squish buffer (10 mM Tris-Cl pH 8.0, 1 mM EDTA, 25 mM NaCl, 1% (w/v) SDS, and 50 μg/ml proteinase K). Samples were incubated for 1 h at 65 °C before being extracted twice with phenol:chloroform:isoamyl alcohol (25:24:1 pH 8.0) and once with chloroform. DNA was precipitated with ethanol and resuspended in water, and 80 ng was used for each 25 μl PCR reaction. PCR reaction and thermocycler conditions were the same as those used for RT-PCR with the exception that the extension time was increased from 30 s to 3 min.
Transcriptional profiling
Identification of PSR-specific long transcripts
We used the Ambion mirVana mRNA isolation kit (Ambion, USA). Specifically, we collected total RNA from pooled sets of ~ 50 testis pairs taken from AsympC or PSR(+) yellow body-red eyed male pupae. Both polyadenylated and nonadenylated sequences were included in our preparations. Extracted RNA samples were depleted of rRNA using the thermos Ribominus kit, DNase-treated (Ambion/Applied Biosystems), and assessed for purity and quality on a Bioanalyzer 2100 (Agilent Technologies) and a Nanodrop 1000 UV-Vis spectrophotometer (NanoDrop Technologies/Thermo Scientific). Paired-end sequenced RNA libraries were prepared for sequencing with the Illumina mRNA-Seq Sample Preparation Kit (Illumina), and the libraries were multiplex-sequenced by using an Illumina HiSequation 2000 sequencer system.
Long RNA reads for both PSR(+) (31,340,421 reads) and WT (28,532,338 reads) testes samples were used to build de novo transcriptomes for each sample independently using Oases v0.2.08 and Velvet v.1.2.08 (Schulz et al. 2012). Oases runs were performed with k-mer sizes ranging from 51 to 93 generating a set of transcripts for the WT testes sample and PSR(+) testes sample. To find transcripts specific to the PSR(+) sample, the transcripts produced from the aforementioned WT sample and PSR(+) sample were blasted to each other, producing PSR(+) loci that had no hits against WT with an e-value cutoff of 0.1. To further filter down these transcripts, a bowtie database was produced from these transcripts and the poly(A)+ transcriptome reads were aligned for both samples with settings –v 0, −k 50, and –m 50, and transcript abundance was calculated as reads per million (RPM). Transcripts that had reads mapping to them from the WT sample were excluded, and we required that the PSR-specific long transcripts were abundantly expressed and had at least 50 reads mapping to them. This stringent filtering resulted in one nonpoly(A)+ PSR-specific transcript.
Identification of PSR-specific small RNAs
Testes were dissected from ~ 100 AsympC or PSR(+) male pupae in the yellow body-red eye stage, and size-selected RNAs (< 50 bp) were purified from the isolated testes as well as the carcasses without testes (somatic tissues) for each genotype. We only used carcasses in which the entire male reproductive tract was completely removed; any dissections in which the reproductive tract was damaged or incompletely extracted were discarded.
For small RNA extractions, we used the mirVana miRNA isolation kit (AM1561, Ambion/Applied Biosystems). The quality of the extracted small RNA was evaluated using BioAnalyzer 2100 (Agilent Technologies). Small RNAs ≤ 50 nucleotides (nt) were gel-purified and ligated to the 3′ adaptor and 5′ adaptor oligonucleotides. RNA was prepared for sequencing using the Illumina TruSeq Small RNA Sample Preparation Kit (RS-200-0012, Illumina). Libraries were clustered onto a flow cell using TruSeq® Rapid SR Cluster Kit – HS (GD-402-4001, Illumina). Samples were sequenced by SBC (Shanghai Biotechnology Corporation, Shanghai, China) using TruSeq® Rapid SBS Kits – HS (FC-402-4002, Illumina) on the HiSeq 2500 Sequencing System (Illumina).
Small RNA sequences were identified by alignment of traces by using Megablast, and the sequences were mapped with the reference N. vitripennis genome version 2.1. Conserved microRNAs were identified by aligning mapped sequences to miRBase Release 21, while novel miRNAs were found by using miRDeep2. The genome locations of these miRNAs were identified by searching for regions with exact sequence matches, along with 100 bases flanking on either side. Novel siRNAs were preliminarily identified as those with > 250 traces and lengths between 20 and 25 bases. Subsequently, these sequences were aligned to themselves by using Megablast, and we identified sequences that matched to reads in a plus/minus manner. We mapped these particular sequences to the N. vitripennis transcriptome (http://trace.ncbi.nlm.nih.gov/Traces/sra/?run=SRR940321, http://trace.ncbi.nlm.nih.gov/Traces/sra/?run=SRR940323). The secondary structures of retrieved hairpin precursor transcripts were predicted by using mfold; sequences that matched perfectly to long transcripts so as to leave two nucleotide overhangs on each 3′ end were considered to be strong candidate siRNAs. Finally, to predict repeat-associated small RNAs (rasiRNAs) or putative piRNAs, we initially selected sequences that begin with a 5′ uracil residue and whose lengths are between 26 and 32 bases. We also tested for signatures of ping pong amplification by blasting all small RNA sequences against the entire PSR(+) testis transcriptome and searching for matches to small RNAs that overlap in a reverse complementary manner by exactly 10 nucleotides at the matching molecules’ 5′ ends.
PSR-specific small RNAs were identified bioinformatically by blasting all small RNAs sequenced from the testis and carcass populations against both the PSR(−) (https://www.ncbi.nlm.nih.gov/pubmed/?term=SRR940321) and PSR(+) (https://www.ncbi.nlm.nih.gov/pubmed/?term=SRR940323) transcriptomes (Akbari et al. 2013). Sequences were considered preliminarily to be PSR-specific small RNAs if they mapped to the PSR(+) transcriptome but not the PSR(−) transcriptome.
All raw sequencing data generated through this study is openly available at the National Center for Biotechnology Information (NCBI) Short Read Archive (SRA) under project number PRJNA387600.
Change history
19 August 2017
An erratum to this article has been published.
References
Akbari OS, Antoshechkin I, Hay BA, Ferree PM (2013) Transcriptome profiling of Nasonia vitripennis testis reveals novel transcripts expressed from the selfish B chromosome, paternal sex ratio. G3 (Bethesda) 3:1597–1605
Aldrich JC, Leibholz A, Cheema MS, Ausio J, Ferree PM (2017) A ‘selfish’ B chromosome induces genome elimination by disrupting the histone code in the jewel wasp Nasonia vitripennis. Sci Rep-Uk. 7
Beukeboom LW, Werren JH (1993) Deletion analysis of the selfish B-chromosome, paternal sex-ratio (Psr), in the parasitic wasp Nasonia-vitripennis. Genetics 133:637–648
Bonaccorsi S, Lohe A (1991) Fine mapping of satellite DNA sequences along the Y chromosome of Drosophila melanogaster: relationships between satellite sequences and fertility factors. Genetics 129:177–189
Camacho JPM, Sharbel TF, Beukeboom LW (2000) B-chromosome evolution. Philos T Roy Soc B 355:163–178
Carchilan M, Delgado M, Ribeiro T, Costa-Nunes P, Caperta A, Morais-Cecilio L, Jones RN, Viegas W, Houben A (2007) Transcriptionally active heterochromatin in rye B chromosomes. Plant Cell 19:1738–1749
Carchilan M, Kumke K, Mikolajewski S, Houben A (2009) Rye B chromosomes are weakly transcribed and might alter the transcriptional activity of A chromosome sequences. Chromosoma 118:607–616
Czech B, Hannon GJ (2016) One loop to rule them all: the ping-pong cycle and piRNA-guided silencing. Trends Biochem Sci 41:324–337
Dej KJ, Spradling AC (1999) The endocycle controls nurse cell polytene chromosome structure during Drosophila oogenesis. Development 126:293–303
Eickbush DG, Eickbush TH, Werren JH (1992) Molecular characterization of repetitive DNA-sequences from a B-chromosome. Chromosoma 101:575–583
Femino AM, Fay FS, Fogarty K, Singer RH (1998) Visualization of single RNA transcripts in situ. Science 280:585–590
Ferree PM, Fang C, Mastrodimos M, Hay BA, Amrhein H, Akbari OS (2015) Identification of genes uniquely expressed in the germ-line tissues of the Jewel wasp Nasonia vitripennis. G3-Genes Genom Genet 5:2647–2653
Ferree PM, Gomez K, Rominger P, Howard D, Kornfeld H, Barbash DA (2014) Heterochromatin position effects on circularized sex chromosomes cause filicidal embryonic lethality in Drosophila melanogaster. Genetics 196:1001–1005
Ferree PM, Prasad S (2012) How can satellite DNA divergence cause reproductive isolation? Let us count the chromosomal ways. Genetics Research International 2012:430136
Hewitt GM (1976) Meiotic drive for B-chromosomes in primary oocytes of Myrmeleotettix-maculatus (Orthoptera-Acrididae). Chromosoma 56:381–391
Holoch D, Moazed D (2015) RNA-mediated epigenetic regulation of gene expression. Nat Rev Genet 16:71–84
Huang, W., Y. Du, X. Zhao, and W.W. Jin. 2016. B chromosome contains active genes and impacts the transcription of A chromosomes in maize (Zea mays L.). Bmc Plant Biol. 16
Jones RN (1991) B-chromosome drive. Am Nat 137:430–442
Jones RN, Viegas W, Houben A (2008) A century of B chromosomes in plants: so what? Ann Bot-London 101:767–775
Kim VN, Han J, Siomi MC (2009) Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10:126–139
Klemme S, Banaei-Moghaddam AM, Macas J, Wicker T, Novak P, Houben A (2013) High-copy sequences reveal distinct evolution of the rye B chromosome. New Phytol 199:550–558
Ku HY, Lin HF (2014) PIWI proteins and their interactors in piRNA biogenesis, germline development and gene expression. Natl Sci Rev 1:205–218
Larracuente, A.M., and P.M. Ferree. 2015. Simple method for fluorescence DNA in situ hybridization to squashed chromosomes. Jove-J Vis Exp.
Lohe AR, Hilliker AJ, Roberts PA (1993) Mapping simple repeated DNA sequences in heterochromatin of Drosophila melanogaster. Genetics 134:1149–1174
Martis MM, Klemme S, Banaei-Moghaddam AM, Blattner FR, Macas J, Schmutzer T, Scholz U, Gundlach H, Wicker T, Simkova H, Novak P, Neumann P, Kubalakova M, Bauer E, Haseneyer G, Fuchs J, Dolezel J, Stein N, Mayer KFX, Houben A (2012) Selfish supernumerary chromosome reveals its origin as a mosaic of host genome and organellar sequences. Proc Natl Acad Sci U S A 109:13343–13346
McAllister BF (1995) Isolation and characterization of a retroelement from B chromosome (PSR) in the parasitic wasp Nasonia vitripennis. Insect Mol Biol 4:253–262
McAllister BF, Beukeboom LW, Werren JH (2004) Mapping of paternal-sex-ratio deletion chromosomes localizes multiple regions involved in expression and transmission. Heredity 92:5–13
McAllister BF, Werren JH (1997) Hybrid origin of a B chromosome (PSR) in the parasitic wasp Nasonia vitripennis. Chromosoma 106:243–253
Menon DU, Meller VH (2012) A role for siRNA in X-chromosome dosage compensation in Drosophila melanogaster. Genetics 191:1023–U1623
Moazed D (2009) Small RNAs in transcriptional gene silencing and genome defence. Nature 457:413–420
Munoz-Pajares A, Martinez-Rodriguez L, Teruel M, Cabrero J, Camacho JPM, Perfectti F (2011) A single, recent origin of the accessory B chromosome of the grasshopper Eyprepocnemis plorans. Genetics 187:853–863
Navarro-Dominguez B, Ruiz-Ruano FJ, Cabrero J, Corral JM, Lopez-Leon MD, Sharbel TF, Camacho JPM (2017) Protein-coding genes in B chromosomes of the grasshopper Eyprepocnemis plorans. Sci Rep-Uk 7
Nur U, Werren JH, Eickbush DG, Burke WD, Eickbush TH (1988) A “selfish” B chromosome that enhances its transmission by eliminating the paternal genome. Science 240:512–514
Page BT, Wanous MK, Birchler JA (2001) Characterization of a maize chromosome 4 centromeric sequence: evidence for an evolutionary relationship with the B chromosome centromere. Genetics 159:291–302
Pal-Bhadra M, Leibovitch BA, Gandhi SG, Rao M, Bhadra U, Birchler JA, Elgin SCR (2004) Heterochromatic silencing and HP1 localization in Drosophila are dependent on the RNAi machinery. Science 303:669–672
Palestis BG, Trivers R, Burt A, Jones RN (2004) The distribution of B chromosomes across species. Cytogenet Genome Res 106:151–158
Perfectti F, Werren JH (2001) The interspecific origin of B chromosomes: experimental evidence. Evolution 55:1069–1073
Perrot-Minnot MJ, Werren JH (2001) Meiotic and mitotic instability of two EMS-produced centric fragments in the haplodiploid wasp Nasonia vitripennis. Heredity 87:8–16
Reed KM (1993) Cytogenetic analysis of the paternal sex-ratio chromosome of Nasonia-vitripennis. Genome 36:157–161
Rosic S, Kohler F, Erhardt S (2014) Repetitive centromeric satellite RNA is essential for kinetochore formation and cell division (vol 207, pg 335, 2014). J Cell Biol 207:673–673
Schulz MH, Zerbino DR, Vingron M, Birney E (2012) Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28:1086–1092
Silva DMZD, Pansonato-Alves JC, Utsunomia R, Araya-Jaime C, Ruiz-Ruano FJ, Daniel SN, Hashimoto DT, Oliveira C, Camacho JPM, Porto-Foresti F, Foresti F (2014) Delimiting the origin of a B chromosome by FISH mapping, chromosome painting and DNA sequence analysis in Astyanax paranae (Teleostei, Characiformes). PLoS One 9
Swim MM, Kaeding KE, Ferree PM (2012) Impact of a selfish B chromosome on chromatin dynamics and nuclear organization in Nasonia. J Cell Sci 125:5241–5249
Ugarkovic D (2005) Functional elements residing within satellite DNAs. EMBO Rep 6:1035–1039
Usakin L, Abad J, Vagin VV, de Pablos B, Villasante A, Gvozdev VA (2007) Transcription of the 1.688 satellite DNA family is under the control of RNA interference machinery in Drosophila melanogaster ovaries. Genetics 176:1343–1349
Valente GT, Conte MA, Fantinatti BEA, Cabral-de-Mello DC, Carvalho RF, Vicari MR, Kocher TD, Martins C (2014) Origin and evolution of B chromosomes in the cichlid fish Astatotilapia latifasciata based on integrated genomic analyses. Mol Biol Evol 31:2061–2072
Van Vugt JJFA, de Jong H, Stouthamer R (2009) The origin of a selfish B chromosome triggering paternal sex ratio in the parasitoid wasp Trichogramma kaykai. P R Soc B 276:4149–4154
van Vugt JJFA, de Nooijer S, Stouthamer R, de Jong H (2005) NOR activity and repeat sequences of the paternal sex ratio chromosome of the parasitoid wasp Trichogramma kaykai. Chromosoma 114:410–419
Volpe TA, Kidner C, Hall IM, Teng G, Grewal SIS, Martienssen RA (2002) Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297:1833–1837
Werren JH (2011) Selfish genetic elements, genetic conflict, and evolutionary innovation. Proc Natl Acad Sci U S A 108(Suppl 2):10863–10870
Werren JH, Richards S, Desjardins CA, Niehuis O, Gadau J, Colbourne JK (2010) Functional and evolutionary insights from the genomes of three parasitoid Nasonia species (vol 327, pg 343, 2010). Science 327:1577–1577
Werren JH, Stouthamer R (2003) PSR (paternal sex ratio) chromosomes: the ultimate selfish genetic elements. Genetica 117:85–101
Whiting PW (1968) The chromosomes of Mormoniella. The Journal of Heredity 59:19–22
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was funded by a USDA National Institute of Food and Agriculture (NIFA) Hatch Project grant (1009509) to O. S. A. and a US National Science Foundation CAREER award (NSF-1451839) to P.M.F.
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not present any studies with human participants or animals performed by any of the authors.
Additional information
The original version of this article was revised: During the steps of corrections, the author name “Omar A. Akbari” went wrong. The corrected author name is shown above.
An erratum to this article is available at https://doi.org/10.1007/s00412-017-0642-9.
Electronic supplementary material
Fig. S1
The PSR chromosome contains abundant copies of repeats that are also present on the A chromosomes. DNA FISH shows that NV79 and NV126 (both in red) are present on a single A chromosome that uniquely contains PSR104 (green). (GIF 85 kb)
Fig. S2
DNA FISH shows that PSR harbors repeats (PSR4656 and PSR8495) that generate polyadenylated transcripts. These repeats co-localize across PSR’s long arm, and are not detectable on the A chromosomes using this method. (GIF 65 kb)
Fig. S3
PSR-specific repeats generate small RNAs at levels much higher than those expressed by repeats in the PSR(−) genome. Levels of small RNAs from A chromosome repeats and PSR-specific repeats are shown in grey and red, respectively. The Y axis portrays the total amount of small RNA traces per repeat in the testis. (GIF 15 kb)
Fig. S4
Depiction of small RNAs deriving from PSR105. (A) Alignment of small RNAs corresponding to PSR105 (red text). (B) The canonical PSR105 repeat is shown, with the sequence of the corresponding small RNAs (highlighted in red text). The two highly conserved palindromic regions of this repeat are underlined. (C) A long RNA precursor for the PSR105-matching small RNAs is shown, with the region generating the small RNAs highlighted in red, and the complementary region shown in black underline. This RNA is predicted to form an imperfect hairpin (below arrow). The region matching to the small RNAs is marked by a red line. (GIF 356 kb)
Fig. S5
Depiction of small RNAs deriving from PSR2. (A) Alignment of small RNAs corresponding to PSR2 (red text). These RNAs form two distinct ‘clusters.’ (B) The canonical PSR2 repeat is shown, with the sequences of the corresponding small RNAs (highlighted in green text and underlined in blue). The two highly conserved palindromic regions of this repeat are underlined). (C) Two long RNA precursors for the PSR2-matching small RNAs are shown, with the region generating each cluster of small RNAs heighted in green and blue, respectively. Each of these RNAs is predicted to form hairpin structures (shown below each arrow). The regions matching to the small RNAs are marked by either a green or a blue line. (GIF 299 kb)
Fig. S6
Control conditions for RNA FISH of the 4317 transcript. Top row depicts PSR4317 probe hybridization to PSR(−) testes, resulting in no signal. Similarly, no signal results from hybridization of this same probe to PSR(+) testes pre-treated with RNase A (bottom row). Only the PSR(+) untreated testes show punctate hybridization signals with this probe. (GIF 434 kb)
Fig. S7
Large nuclei in the N. vitripennis testis are polyploid. (A) The large nuclei show multiple, distinct foci of PSR22, in addition to rDNA located at a single locus on an A chromosome. Nuclei of cyst cells following S-phase (B) and mature sperm (C) have exactly two foci and one focus, respectively, of PSR22 and rDNA. (GIF 89 kb)
Table S1
Identification of a new PSR-specific transcript from unfiltered total RNA (XLSX 11 kb)
Table S2
miRNAs, siRNAs, and rasi−/putative piRNAs identified from N. vitripennis testis and carcass (somatic tissues) of PSR(−) and PSR(+) males. Each of these small RNA classes is shown in a separate tab. (XLSX 103 kb)
Table S3
Additional information for N. vitripennis miRNAs and piRNAs. (XLSX 159 kb)
Table S4
Small RNAs that are exclusive to the PSR(+) genotype. (XLSX 72 kb)
Table S5
Putative precursor RNAs for small RNAs derived from PSR repeats PSR2, PSR22, and PSR105. (XLSX 47 kb)
Table S6
Oligonucleotides used for DNA FISH (XLSX 89 kb)
Table S7
Oligonucleotides used for PCR (XLSX 112 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Jing, X.A., Aldrich, J.C. et al. Unique sequence organization and small RNA expression of a “selfish” B chromosome. Chromosoma 126, 753–768 (2017). https://doi.org/10.1007/s00412-017-0641-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-017-0641-x