Abstract
miRNAs are important factors for post-transcriptional process that controls gene expression at mRNA level. Various biological processes, including growth and differentiation, are regulated by miRNAs. miRNAs have been demonstrated to play an essential role in development and progression of hearing loss. Nowadays, miRNAs are known as critical factors involved in different physiological, biological, and pathological processes, such as gene expression, progressive sensorineural hearing loss, age-related hearing loss, noise-induced hearing loss, cholesteatoma, schwannomas, and inner ear inflammation. The miR-183 family (miR-183, miR-96 and miR-182) is expressed abundantly in some types of sensory cells in inner ear specially mechanosensory hair cells that exhibit a great expression level of this family. The plasma levels of miR-24-3p, miR-16-5p, miR-185-5p, and miR-451a were upregulated during noise exposures, and increased levels of miR-21 have been found in vestibular schwannomas and human cholesteatoma. In addition, upregulation of pro-apoptotic miRNAs and downregulation of miRNAs which promote differentiation and proliferation in age-related degeneration of the organ of Corti may potentially serve as a helpful biomarker for the early detection of age-related hearing loss. This knowledge represents miRNAs as promising diagnostic and therapeutic tools in the near future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hearing loss also known as silent disability affects over 275 million people worldwide most of which live in low- and moderate-income countries. The number of people with debilitating from mild to severe hearing impairment has been estimated to be more than 360 million [1]. Social problems due to hearing impairment include heavy socioeconomic costs imposed on families, delayed acquisition of cognitive and verbal skills, and education achievement-related problems in children, as well as difficulties with finding or keeping job in adults. Meanwhile, costs of education of children with special needs can cause a great national economic burden. The main causes of hearing loss include infectious diseases, such as meningitis, measles, mumps, and chronic ear infections, exposure to noise, damage to ear and head, aging, and taking certain drugs with toxic effects. Hearing loss and hearing impairment can be prevented or, through the early diagnosis and appropriate management, treated (by surgery, use of certain instruments, such as hearing aid, and cochlear implant) in half of the cases [2].
The genome of eukaryotes consists of both coding and non-coding DNA. Coding DNA consists of all open reading frames (ORFs) and exons that are transcribed to RNA sequences and even translated to protein sequences. However, the function of non-coding regions, including introns has not fully understood, although some regulatory roles have been observed for introns [3].
Another important factors are miRNAs which were discovered and found to be involved in regulating expression and function of protein-coding RNAs by Lee and co-workers in 1993 [4]. These small non-coding RNAs [4] are usually 22- to 42-nucleotide long and are highly conserved in plants and animals. These regulatory elements are evolutionarily ancient component for the process of gene regulation [5]. These molecules regulate the expression of over 30% of genes in different biological processes, such as proliferation, differentiation, survival, migration, apoptosis, and death [6].
Cellular reprogramming occurs during the development of eukaryotic organisms via both extracellular [7] and intracellular agents. Among intracellular factors, histone modifications, DNA methylation, and gene expression regulation via microRNAs have been described. microRNAs are found in animals, plants, and some viruses, and they function in silencing and post-transcriptional regulation of gene expression [8]. Most of miRNAs are located within the cell, but some miRNAs are found in extracellular environment, such as serum, biological fluids, and the media of cell culture [9]. microRNAs can silence the gene expression via cleavage of the mRNA strand, shortening of the poly(A) tail, and affecting the mRNA translation into proteins by ribosomes; phenomena that can be predicted using computational biology [10].
To date, over 2500 human miRNAs have been identified [11]. Many pathological processes, such as cardiovascular and neurological diseases, diabetes, and cancers are associated with abnormal expression of miRNAs [12, 13]. Interestingly a number of studies have pointed out the importance of miRNAs in hearing-related diseases. For instance, there are evidences that miRNAs can be used as biomarker to diagnose noise-induced and age-related hearing loss [20, 21]. In this work, we have reviewed the role of microRNAs in ear-related diseases and hearing loss. We have studied the current evidences on the role of miRNAs in inner ear evolution and recent research accomplishments and developments on the role of miRNAs in inner ear biology and pathogenesis.
Biogenesis of MicroRNAs
Biogenesis of miRNAs occurs in nucleus and cytoplasm. In nucleus, first, RNA polymerase II transcribes pri-miRNAs from either coding or non-coding areas in the genome. pri-miRNAs have a loop-stem structure with poly(A) tail at 3′-end and CAP at 5′-end. pri-miRNAs are converted to pre-miRNAs by an RNAse III-containing complex which is specialized to digest a double-stranded RNA (Drosha), and a double-stranded RNA-binding protein, DGCR8. pre-miRNAs are then transported to cytoplasm by exportin-5. In cytoplasm, final processing is conducted on pre-miRNA by TRBP/Dicer complex. Dicer function leads to formation of a 31 to 42-nucleotide, double-stranded RNA with a guide strand and a passenger strand. Passenger strand is destroyed and guide strand forms miRNA via binding to RNA-induced silencing complex (mi-RISC) containing members of Argonaute protein family. This complex is complementary to 3′-UTR of target mRNA. If miRNA sequence is completely complementary to target mRNA, target mRNA is destroyed after binding. However, translation is inhibited, in case the miRNA binding to mRNA is incomplete. Each miRNA can target several mRNAs [14–18] (Fig. 1).
Biogenesis of miRNA: miRNA biogenesis is a multistep process. First, miRNA genes are transcribed by RNA polymerase II in the nucleus (step 1). The resulting primary transcript is cleaved by Drosha and DGCR8 to produce pre-miRNA (step 2). After exportin-5- and RanGTP-mediated transport to the cytoplasm (step 3), the pre-miRNA undergoes its final processing step, which consists of Dicer-dependent cleavage just below the stem loop to produce a duplex molecule (step 4). The duplex is then separated and usually one strand is selected as the mature miRNA and directed to target-specific mRNAs (step 5)
MicroRNAs in hearing loss
Little is known about molecular basis of progressive hearing loss. Recently, the potential role of miRNA as a therapeutic agent in regeneration has been demonstrated in research on regulatory factors involved in ear diseases [19, 20]. miRNAs play a very important role in gene expression, physiological and pathological processes, and progressive sensorineural hearing loss (SNHL) which is mainly due to cochlear hair cells defect or loss. Role of miRNAs in pathogenesis of different ear diseases has been studied via detecting genetic and somatic mutations in miRNAs and their binding sites in target genes [21, 22]. Recently, mutations in miR-96 have been demonstrated to be associated with progressive hearing loss in humans and mice. Similar to histone modifications and DNA methylation, miRNAs have been found to play roles in survival and development of organs and cells, such as mechanosensory hair cells [23]. Moreover, miRNAs have been suggested to be essential and new regulatory factors for the formation of induced pluripotent stem cells (iPSCs) and differentiation of these cells into hair cell [24, 25]
MicroRNAs are involved in formation of the inner ear
miRNAs are considered as important factors that widely affect the development of inner ear. Microarray analysis, quantitative real-time PCR, and in situ hybridization have demonstrated that miRNA-182, miRNA-140, miRNA-200c expressions have distinct temporal and spatial patterns at E13.5 (mouse embryonic day 13.5 of development) and E16.5 [26]. mir-96, mir-182, and mir-183 are the well-investigated miRNAs that were first detected in the otic vesicle, cochlea-vestibule ganglion, and cochlear hair cells at E9.5, E11.5, and E17.5, respectively. According to some studies, the expression continued to at least P30 (Postnatal Day 30). The expression of other miRNAs detected by in situ hybridization is shown in Fig. 2 [27].
The miRNA-183 family in hair cell development and deafness
miRNA-183 family (96, 182, and 183) is expressed abundantly in specific sensory cell types in the inner ear. They play either distinct or common roles in inner ear cell-fate determination [28]. In vertebrate hair cells, such as zebrafish, the miR-183 family is highly expressed and induces extra and ectopic hair cells. Contrariwise, knockdown causes reduction in numbers of hair cells in the inner ear, smaller spiral ganglion neuron (SGNs), and defects in semicircular canals [29].
Among the members of this family, miR-96 is expressed as a sensory organ-specific miRNA in the mammalian cochlea during development. Indeed, in humans and mice, some types of non-syndromic progressive hearing loss are due to miR-96 mutations [30]. One example is the results of the study conducted by Mencia and co-workers who found that point mutations in the seed region of miR-96 result in progressive hearing loss with autosomal dominant inheritance pattern [31]. Further investigations on the role of a SNP (Single-nucleotide polymorphism) in the seed region of hsa-miR-96 revealed two key biological processes involved in progressive hearing loss. This SNP leads to a significant enrichment of two sets of allele-specific target genes of hsa-miR-96, including neurotrophin in TRK receptor signaling pathway and epidermal growth factor receptor signaling pathway [21]. miRNA profiling, target analysis, and validation suggest that miRNAs (e.g. miR-183/Taok1 target pair) are involved in regulating the degenerative process of the cochlea after acoustic overstimulation [32]. In inner ear hair cells, chloride intracellular channel 5 (CLIC5) expression is regulated by miR-183 family. In other words, CLIC5 3′-UTR contains a highly conserved sequence which is a single predicted miR-96/-182-binding site. This binding site which is located between nucleotides 760–766 is the target where miR-96/-182 can regulated the CLIC5 [33].
MicroRNAs are involved in age-related hearing loss
The main cause of age-related hearing loss (ARHL) is degeneration of the organ of Corti (OC) caused by alteration of miR-29 and miR-34 families that regulate pro-apoptotic pathways. The expression levels of miR-34a exhibit a significant stimulator to increase the cochlea, auditory cortex, and plasma in mice during aging. These increases are accompanied by elevated hearing thresholds and greater hair cells loss. Levels of silent information regulator-1 (SIRT1), B-cell lymphoma-2, and E2F transcription factor 3 as targets of miR-34a may potentially serve as useful biomarkers to detect ARHL early [34]. Moreover, several studies have shown that acetylation of p53 and apoptosis is increased via miR-34a/Sirtuin1 (SIRT1)/p53 signaling pathway in cochlear hair cells during aging. The findings, therefore, suggest a link between age-related apoptosis of cochlear hair cell and miR-34a/SIRT1/p53 signaling, which may provide a potential target for ARHL [35]. Unlike upregulation of this pro-apoptotic miRNA, many miRNA families are important for proliferation and differentiation. Two examples are miR-181 and miR-183 whose downregulation has been reported to be associated with cell death resulting in ARHL [36].
MicroRNA expression profiles in noise-induced hearing loss
There are evidences that noise stimulation or occurrence of noise-induced hearing loss (NIHL) affects miRNA expression. One of the major causes in pathogenesis of NIHL is oxidative stress, because the cochlea is metabolically active [27]. Cellular response to noise exposure can be detected as plasma miRNAs biomarker that may provide new insights in pathogenesis of NIHL. In fact, miRNA target analysis may reveal the pathways involved in NHIL and help to develop new drugs through identification of the cellular stress response components [37]. After noise exposure in animal model with noise-induced deafness, the expression of members of miRNA-183 family exhibits significant increase, which shows that this family members may play significant roles in the pathogenesis and development of NHIL [38]. Moreover, the plasma levels of miR-16-5p, miR-24-3p, miR-185-5p, and miR-451a are also known to be upregulated in noise exposures [39].
MicroRNA in inner ear inflammation
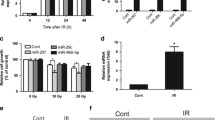
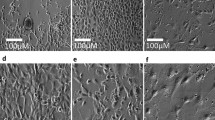
Inflammation of the inner ear can cause several forms of immune-mediated hearing loss. Therefore, understanding the inflammatory pathways and its molecular components is very helpful to develop drugs and treatments for these types of hearing loss [40, 41]. A number of studies have discussed the mechanisms of neuroprotective agents against cytotoxicity [42, 43]. In addition to identification of genetic factors involved in inflammatory hearing loss, miRNAs may contribute to gene expression regulation and affect the outcome of inflammation in ear. The biological effects of miRNAs have been found by ionizing radiation (IR)-induced cell death in auditory cells using miRNA mimics or inhibitors. Studies conducted by Tan and co-workers on house ear institute OC 1 (HEI-OC1) cells revealed that miR-207 is upregulated following IR resulting in IR-induced apoptosis and enhanced DNA damage. Therefore, inhibition or downregulation of miR-207 could be a potential strategy for protecting cochlear hair cells against IR [44]. Another miRNA involved in inner ear inflammation is miR-224 which is a transcriptional target of the key mediator of innate immunity, nuclear factor kappa B pathway. It is well documented that miR-224 diminishes the innate immune response through downregulating Ptx3 expression that stimulates the innate immune response [40].
MicroRNAs and epigenetics in inner ear
DNA methylation and histone modifications are epigenetic mechanisms that are implicated in human deafness, suggesting that different levels of non-coding genes (such as miRNAs) are required for normal hearing [23]. Sensory hair cells and supporting cells in the auditory organ of the mammalian inner ear arise from a common sensory progenitor. Cell fate in developing of organ of Corti is controlled by miR-124 which is an epigenetic protective agent for two inhibitors of the Wnt pathway, Sfrp4 (secreted frizzled-related protein4), and Sfrp5. These proteins play essential roles in fine-tuning of the expression of genes critical for cell patterning (e.g., Hes1 and Hes5) during cochlear differentiation [23].
microRNA expression in cholesteatoma and schwannoma
The role of miR-21 has been described in some of ear-related diseases, such as human cholesteatoma growth and proliferation, and vestibular schwannomas [45, 46]. Upregulation of miR-21 is implicated in potential harmful growth in the middle ear or in the mastoid bone [45]. Increased levels of miR-21 have been also observed in vestibular schwannomas [46].
Other miRNAs in hearing loss
Experimental analyses of vestibular hair cells have shown that expression of PSIP1-P75 which is a transcriptional co-activator in the inner ear may be regulated by miR-135b. Indeed, some differences between the cochlear and vestibular hair cells are determined by miR-135b which acts as a cellular effector [22]. Furthermore, since protection of SGNs from ongoing degeneration is an essential step to prevent progressive hearing loss, miRNA-base strategies can be designed to prevent or reverse SGN damages. Downregulation of TMPRSS3 (transmembrane protease, serine3) has been reported to be necessary for SGNs development through overexpression of miR-204. Therefore, alteration of miR-204 may serve as a potential therapeutic target in SNHL [47]. Another key regulator in development of hearing organs is miR-6716-3p. This miRNA is involved in actin reorganization, sensory hair cell bundle development, cell adhesion, and inner ear morphogenesis [48]. Since miRNAs can regulate important signaling pathways, it is not surprising that they are also involved in cytotoxicity. Indeed, recent findings have suggested a role of miR34a and miR34c in antibiotic-induced ototoxicity in a dose-dependent manner in cochlear cells [42].
MicroRNA editing and transfer
In addition to all above-mentioned roles for various miRNAs, an important factor affecting the function of miRNAs is their proper trafficking and transfer. An example is a membrane protein Cx26. This protein that is required for cochlear development is responsible for intercellular communication, including miRNA intercellular transfer between native cochlear supporting cells. Deficiency of Cx26 disrupts miRNA intercellular transfer in the cochlea and leads to cochlear developmental disorders and congenital deafness [49]. In addition to proper transit, all miRNAs require a fully functional editing system in the cell; therefore, any failure in editing may result in vigorous outcome in organ development. For instance, a spectrum of non-syndromic to syndromic hearing loss is associated with mutations in phosphoribosyl pyrophosphate synthetase 1 (PRPS1), due to the failure of unedited pri-miR-376 cluster (miR-376a-3p, b-3p, c-3p) in regulating the activity of PRPS1 in the inner ear [50] (Table 1).
Discussion
To develop effective therapeutic strategies to treat hair cell damage, a fundamental and significant step is to identify regulatory mechanisms involved [32]. Recently, a common mechanism has been demonstrated to be involved in evolution and reprogramming of inner ear cells in a number of different species of vertebrate. The regulatory role of miRNAs, particularly miRNA-183 family, and certain genes, such as GFI1, POU4F3, and ATOH1, has been confirmed in differentiation of stem cells into hair cells and protection of hair cells [24]. Moreover, since hundreds of transcripts may be regulated simultaneously by the same miRNA, miRNAs can certainly be considered as potential therapeutic agents to repair or regenerate hair cells at least in animal models [41]. Similarly, since a large proportion of human transcriptome is regulated by miRNAs, they can be used in diagnosis and prognosis as well as development of drugs. Despite advances in the use of cochlear implant to treat SNHL, the rate of hearing improvement is not fully satisfying in people undergoing this therapy. The combination of molecular-based methods and stem cell therapies can be a promising alternative to cochlear implant. Supporting cells differentiation into hair cells that does not occur under normal conditions may theoretically be induced in adult mammals by alteration and modification of miRNAs level in the cells as well as administration of their transit and cellular trafficking. Molecule-based therapies using anti-miRNA LNA (miravirsen) and mimic miRNA (MRX34) are being investigated in clinical trials, but most of miRNA-based therapies are being studied in preclinical studies [17]. Recently, circulating miRNAs have attracted attention, because they are considered to be frequent and stable biomarkers in serum and plasma. Further studies are required to shed light onto the role of miRNA profile alteration during the hearing loss process before miRNAs can be used as a biomarker to diagnose hearing loss [34, 53–57]. Briefly, in the near future, the patient’s serum miRNAs levels might be used, as molecular markers, for diagnosis and prognosis of hearing loss. Moreover, they are potential therapeutic agents for repair or regenerate hair cells, cell reprogramming, and regenerative medicine.
Conclusion
Efforts are being made to offer an appropriate molecular marker with capability to predict hearing loss or even being used as a diagnostic and prognostic agent alongside other pathological or clinical approaches. Among them, miRNAs have been demonstrated to become dysfunctional during hearing loss and play essential roles in the development and progression of this disorder. miRNAs have the potential to be used in treating hearing loss. Considering the nature of miRNAs and the fact that most common approaches to screen hearing loss at the early stages fail to diagnose this disorder, it is a highly promising strategy to study and identify circulating miRNAs as molecular markers for progression of hearing loss due to any causes, such as ARHL, NIHL, inflammation, Schwannomas, and cholesteatoma.
References
Ware SL (2014) Human hearing loss. PeerJ PrePrints 2:e378v1
Tucci D, Merson MH, Wilson BS (2010) A summary of the literature on global hearing impairment: current status and priorities for action. Otol Neurotol 31(1):31–41
Jami MS, Hemati S, Salehi Z, Tavassoli M (2008) Association between the length of a CA dinucleotide repeat in the EGFR and risk of breast cancer. Cancer Investig 26(4):434–437
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854
Garzon R, Calin GA, Croce CM (2009) MicroRNAs in Cancer. Annu Rev Med 60:167–179
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19(1):92–105
García-Estrada C, Barreiro C, Jami M-S, Martín-González J, Martín J-F (2013) The inducers 1, 3-diaminopropane and spermidine cause the reprogramming of metabolism in Penicillium chrysogenum, leading to multiple vesicles and penicillin overproduction. J Proteom 85:129–159
Mahmoodian sani MR, Hashemzadeh-Chaleshtori M, Mehri-Ghahfarrokhi A, Ghasemi-Dehkordi P, Saidijam M, Jami M-S (2016) MicroRNA-183 family in inner ear: hair cell development and deafness. J Audiol Otol 20(3):131–138
Ahmadinejad F, Honardoost MA, Mowla SJ, Teimori H, Ghaedi K (2016) miR-218 as a multifunctional regulator of oncogenic processes in different solid tumors. Genet 3rd Millenn 14(2):4276–4295
Honardoost MA, Naghavian R, Ahmadinejad F, Hosseini A, Ghaedi K (2015) Integrative computational mRNA-miRNA interaction analyses of the autoimmune-deregulated miRNAs and well-known Th17 differentiation regulators: An attempt to discover new potential miRNAs involved in Th17 differentiation. Gene 572(2):153–162
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucl Acids Res 39(Database issue):D152–D157
Miller BH, Wahlestedt C (2010) MicroRNA dysregulation in psychiatric disease. Brain Res 1338:89–99
Guay C, Roggli E, Nesca V, Jacovetti C, Regazzi R (2011) Diabetes mellitus, a microRNA-related disease? Transl Res J Lab Clin Med 157(4):253–264
Tolia NH, Joshua-Tor L (2007) Slicer and the argonautes. Nat Chem Biol 3(1):36–43
Lee I, Ajay SS, Yook JI, Kim HS, Hong SH, Kim NH et al (2009) New class of microRNA targets containing simultaneous 5′-UTR and 3′-UTR interaction sites. Genome Res 19(7):1175–1183
Farazi TA, Hoell JI, Morozov P, Tuschl T (2013) MicroRNAs in human cancer. Adv Exp Med Biol 774:1–20
Ling H, Fabbri M, Calin GA (2013) MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov 2(11):847–865
Macfarlane LA, Murphy PR (2010) MicroRNA: biogenesis, function and role in cancer. Curr Genom 11(7):537–561
Lewis MA, Quint E, Glazier AM, Fuchs H, De Angelis MH, Langford C et al (2009) An ENU-induced mutation of miR-96 associated with progressive hearing loss in mice. Nat Genet 41(5):614–618
Ushakov K, Rudnicki A, Avraham KB (2013) MicroRNAs in sensorineural diseases of the ear. Front Mol Neurosci 6:52
Bhattacharya A, Cui Y (2015) Knowledge-based analysis of functional impacts of mutations in microRNA seed regions. J Biosci 40(4):791–798
Elkan-Miller T, Ulitsky I, Hertzano R, Rudnicki A, Dror AA, Lenz DR et al (2011) Integration of transcriptomics, proteomics, and microRNA analyses reveals novel microRNA regulation of targets in the mammalian inner ear. PloS One 6(4):e18195
Friedman LM, Avraham KB (2009) MicroRNAs and epigenetic regulation in the mammalian inner ear: implications for deafness. Mamm Genome 20(9–10):581–603
Beisel K, Hansen L, Soukup G, Fritzsch B (2008) Regenerating cochlear hair cells: quo vadis stem cell. Cell Tissue Res 333(3):373–379
Ghasemi-Dehkordi P, Allahbakhshian-Farsani M, Abdian N, Mirzaeian A, Saffari-Chaleshtori J, Heybati F et al (2015) Comparison between the cultures of human induced pluripotent stem cells (hiPSCs) on feeder-and serum-free system (Matrigel matrix), MEF and HDF feeder cell lines. J Cell Commun Signal 9(3):233–246
Wang XR, Zhang XM, Zhen J, Zhang PX, Xu G, Jiang H (2010) MicroRNA expression in the embryonic mouse inner ear. Neuroreport 21(9):611–617
Rudnicki A, Avraham KB (2012) microRNAs: the art of silencing in the ear. EMBO Mol Med 4(9):849–859
Rezaei H, Movahedi R (2010) High frequency of 35delG mutation in GJB2 associated with autosomal recessive nonsyndromic hearing loss (ARNSHL) in the province of Isfahan-Iran. Genet 3rd Millenn 8(3):2074–2078
Li H, Kloosterman W, Fekete DM (2010) MicroRNA-183 family members regulate sensorineural fates in the inner ear. J Neurosci 30(9):3254–3263
Kuhn S, Johnson SL, Furness DN, Chen J, Ingham N, Hilton JM et al (2011) miR-96 regulates the progression of differentiation in mammalian cochlear inner and outer hair cells. Proc Natl Acad Sci USA 108(6):2355–2360
Mencia A, Modamio-Hoybjor S, Redshaw N, Morin M, Mayo-Merino F, Olavarrieta L et al (2009) Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss. Nat Genet 41(5):609–613
Patel M, Cai Q, Ding D, Salvi R, Hu Z, Hu BH (2013) The miR-183/Taok1 target pair is implicated in cochlear responses to acoustic trauma. PLoS One 8(3):e58471
Gu C, Li X, Tan Q, Wang Z, Chen L, Liu Y (2013) MiR-183 family regulates chloride intracellular channel 5 expression in inner ear hair cells. Toxicol Vitro Int J Publ Assoc BIBRA 27(1):486–491
Pang J, Xiong H, Yang H, Ou Y, Xu Y, Huang Q et al (2016) Circulating miR-34a levels correlate with age-related hearing loss in mice and humans. Exp Gerontol 76:58–67
Xiong H, Pang J, Yang H, Dai M, Liu Y, Ou Y et al (2015) Activation of miR-34a/SIRT1/p53 signaling contributes to cochlear hair cell apoptosis: implications for age-related hearing loss. Neurobiol Aging 36(4):1692–1701
Zhang Q, Liu H, McGee J, Walsh EJ, Soukup GA, He DZ (2013) Identifying microRNAs involved in degeneration of the organ of corti during age-related hearing loss. PloS One 8(4):e62786
Mendell JT, Olson EN (2012) MicroRNAs in stress signaling and human disease. Cell 148(6):1172–1187
Zhang Z, Liu K, Chen Y, Li Z, Yan N, Zhang J (2014) The expression of miR-183 family in the pathogenesis and development of noise-induced deafness. Lin chuang er bi yan hou tou jing wai ke za zhi = J Clin Otorhinolaryngol Head Neck Surg 28(7):468–472
Ryan AF, Harris JP, Keithley EM (2002) Immune-mediated hearing loss: basic mechanisms and options for therapy. Acta Oto-laryngol Suppl (548):38–43
Rudnicki A, Shivatzki S, Beyer LA, Takada Y, Raphael Y, Avraham KB (2014) microRNA-224 regulates Pentraxin 3, a component of the humoral arm of innate immunity, in inner ear inflammation. Hum Mol Genet 23(12):3138–3146
Li H, Fekete DM (2010) MicroRNAs in hair cell development and deafness. Curr Opin Otolaryngol Head Neck Surg 18(5):459–465.
Jami MS, Pal R, Hoedt E, Neubert TA, Larsen JP, Moller SG (2014) Proteome analysis reveals roles of L-DOPA in response to oxidative stress in neurons. BMC Neurosci 15:93
Jami MS, Salehi-Najafabadi Z, Ahmadinejad F, Hoedt E, Chaleshtori MH, Ghatrehsamani M et al (2015) Edaravone leads to proteome changes indicative of neuronal cell protection in response to oxidative stress. Neurochem Int 90:134–141
Tan PX, Du SS, Ren C, Yao QW, Zheng R, Li R, et al. MicroRNA-207 enhances radiation-induced apoptosis by directly targeting Akt3 in cochlea hair cells. Cell Death Dis. 2014;5:e1433.
Friedland DR, Eernisse R, Erbe C, Gupta N, Cioffi JA (2009) Cholesteatoma growth and proliferation: post-transcriptional regulation by microRNA-21. Otol Neurotol 30(7):998
Cioffi JA, Yue WY, Mendolia-Loffredo S, Hansen KR, Wackym PA, Hansen MR (2010) MicroRNA-21 over-expression contributes to vestibular schwannoma cell proliferation and survival. Otol Neurotol 31(9):1455
Li Y, Peng A, Ge S, Wang Q, Liu J (2014) miR-204 suppresses cochlear spiral ganglion neuron survival in vitro by targeting TMPRSS3. Hear Res 314:60–64
Rudnicki A, Isakov O, Ushakov K, Shivatzki S, Weiss I, Friedman LM et al (2014) Next-generation sequencing of small RNAs from inner ear sensory epithelium identifies microRNAs and defines regulatory pathways. BMC Genom 15:484
Zhu Y, Zong L, Mei L, Zhao HB (2015) Connexin26 gap junction mediates miRNA intercellular genetic communication in the cochlea and is required for inner ear development. Sci Rep 5:15647
Yan D, Xing Y, Ouyang X, Zhu J, Chen ZY, Lang H et al (2012) Analysis of miR-376 RNA cluster members in the mouse inner ear. Int J Exp Pathol 93(6):450–457
Chen J, Johnson SL, Lewis MA, Hilton JM, Huma A, Marcotti W et al (2014) A reduction in Ptprq associated with specific features of the deafness phenotype of the miR-96 mutant mouse diminuendo. Eur J Neurosci 39(5):744–756
Chiang DY, Cuthbertson DW, Ruiz FR, Li N, Pereira FA (2013) A coregulatory network of NR2F1 and microRNA-140. PloS One 8(12):e83358
Huyghe A, Van den Ackerveken P, Sacheli R, Prevot PP, Thelen N, Renauld J et al (2015) MicroRNA-124 Regulates Cell Specification in the Cochlea through Modulation of Sfrp4/5. Cell Rep 13(1):31–42
Sivakumaran TA, Resendes BL, Robertson NG, Giersch AB, Morton CC (2006) Characterization of an abundant COL9A1 transcript in the cochlea with a novel 3′ UTR: Expression studies and detection of miRNA target sequence. J Assoc Res Otolaryngol JARO 7(2):160–172
Thoenes M, Zimmermann U, Ebermann I, Ptok M, Lewis MA, Thiele H et al (2015) OSBPL2 encodes a protein of inner and outer hair cell stereocilia and is mutated in autosomal dominant hearing loss (DFNA67). Orphanet J Rare Dis 10:15
Wang XR, Zhang XM, Du J, Jiang H (2012) MicroRNA-182 regulates otocyst-derived cell differentiation and targets T-box1 gene. Hear Res 286(1–2):55–63
Ding L, Liu J, Shen HX, Pan LP, Liu QD, Zhang HD, et al. (2016) Analysis of plasma microRNA expression profiles in male textile workers with noise-induced hearing loss. Hear Res 333:275–282
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study funded by Research and Technology Deputy of Hamadan University of Medical Sciences (Hamadan, Iran) and Shahrekord University of Medical Sciences, Shahrekord, Iran (Grant number 1724).
Conflict of interest
Authors declare no conflict of interest for the manuscript.
Rights and permissions
About this article
Cite this article
Mahmoudian-sani, MR., Mehri-Ghahfarrokhi, A., Ahmadinejad, F. et al. MicroRNAs: effective elements in ear-related diseases and hearing loss. Eur Arch Otorhinolaryngol 274, 2373–2380 (2017). https://doi.org/10.1007/s00405-017-4470-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-017-4470-6