Abstract
Objective
Breast cancer is a severe risk to public health and has adequately convoluted pathogenesis. Therefore, the description of key molecular markers and pathways is of much importance for clarifying the molecular mechanism of breast cancer-associated fibroblasts initiation and progression. Breast cancer-associated fibroblasts gene expression dataset was downloaded from Gene Expression Omnibus database.
Methods
A total of nine samples, including three normal fibroblasts, three granulin-stimulated fibroblasts and three cancer-associated fibroblasts samples, were used to identify differentially expressed genes (DEGs) between normal fibroblasts, granulin-stimulated fibroblasts and cancer-associated fibroblasts samples. The gene ontology (GO) and pathway enrichment analysis was performed, and protein-protein interaction (PPI) network of the DEGs was constructed by NetworkAnalyst software.
Results
Totally, 190 DEGs were identified, including 66 up-regulated and 124 down-regulated genes. GO analysis results showed that up-regulated DEGs were significantly enriched in biological processes (BP), including cell-cell signalling and negative regulation of cell proliferation; molecular function (MF), including insulin-like growth factor II binding and insulin-like growth factor I binding; cellular component (CC), including insulin-like growth factor binding protein complex and integral component of plasma membrane; the down-regulated DEGs were significantly enriched in BP, including cell adhesion and extracellular matrix organization; MF, including N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase activity and calcium ion binding; CC, including extracellular space and extracellular matrix. WIKIPATHWAYS analysis showed the up-regulated DEGs were enriched in myometrial relaxation and contraction pathways. WIKIPATHWAYS, REACTOME, PID_NCI and KEGG pathway analysis showed the down-regulated DEGs were enriched endochondral ossification, TGF beta signalling pathway, integrin cell surface interactions, beta1 integrin cell surface interactions, malaria and glycosaminoglycan biosynthesis—chondroitin sulfate/dermatan sulphate. The top 5 up-regulated hub genes, CDKN2A, MME, PBX1, IGFBP3, and TFAP2C and top 5 down-regulated hub genes VCAM1, KRT18, TGM2, ACTA2, and STAMBP were identified from the PPI network, and subnetworks revealed these genes were involved in significant pathways, including myometrial relaxation and contraction pathways, integrin cell surface interactions, beta1 integrin cell surface interaction. Besides, the target hsa-mirs for DEGs were identified. hsa-mir-759, hsa-mir-4446-5p, hsa-mir-219a-1-3p and hsa-mir-26a-5p were important miRNAs in this study.
Conclusions
We pinpoint important key genes and pathways closely related with breast cancer-associated fibroblasts initiation and progression by a series of bioinformatics analysis on DEGs. These screened genes and pathways provided for a more detailed molecular mechanism underlying breast cancer-associated fibroblasts occurrence and progression, holding promise for acting as molecular markers and probable therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer is associated with fibroblasts at entire phases of disease progression, including metastasis, and they are a substantial component of the general host response to tissue damage induced by tumour cells [1]. Cancer-associated with fibroblasts become synthetic machines that produce diverse extracellular matrix (ECM) components, growth factors, cytokines, proteases, and hormones. Cancer-associated fibroblasts have generally been characterize as α-smooth muscle actin (α-SMA) [2]. Cancer-associated fibroblast expresses biomarkers such as such as vimentin, fibroblast specific protein, fibroblast activation protein (FAP), osteonectin, and desmin [3]. However, distinct biomarkers have been diagnosed in different tumour subtypes [4], and unique biomarkers that can perceive all of the cancer-associated fibroblasts in breast cancer remain inadequate to date.
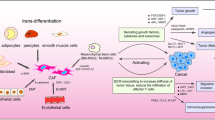
Stromal cancer-associated fibroblasts of breast cancer have four specific cellular origins [5]. The first one is activated normal stromal fibroblasts, which is primary one and 80% of breast cancer-associated fibroblasts [6]. The second one is epithelial–mesenchymal transition (EMT) or endothelial cells, which undergo endothelial–mesenchymal transition [7]. The third one is bone marrow-derived mesenchymal stem cells (MSCs), which share to the regeneration of mesenchymal tissues, and are crucial in furnishing backing for the growth and differentiation of primitive hemopoietic cells within the bone marrow microenvironment [8]. The fourth one comprise trans differentiated cells in breast tissue, such as pericytes, adipocytes, or smooth muscle cells [9]. The significances of cancer-associated fibroblast in the clinical diagnosis and prognosis of breast cancer have been studied broadly, and the possible usage of some cancer-associated fibroblast biomarkers has been diagnosed. PDGF receptor β and FAP are the biomarkers significantly associated with breast cancer [10]. Alteration and/or loss of tumor suppressor genes are also critical features of breast cancer-associated fibroblast. Significantly alteration in tumor suppressor gene Cav-1 in breast cancer-associated fibroblast [11]. So triple negative breast cancer (TNBC) is comes under stromal cancer-associated fibroblasts.
Triple receptor-negative breast carcinoma (TNBC) is the most frequent and aggressive mammary gland malignancy in female adults, and is identified by absence of the estrogen receptor (ER) and progesterone receptor (PR) and lack of over expression of human epidermal growth factor receptor 2 (HER2) that are involved with advancement of the disease [12]. It is a rapidly fatal malignancy and the majority of patients with TNBC suffer from a poor quality of life [13]. Presently, the accepted clinical treatment is surgical resection of the malignant tissues, followed by radiotherapy and chemotherapy [14]. However, patients that receive these treatments may expeditiously advance resistance to chemotherapy [15]. Recent studies have focused on the recognition of candidate biomarkers of TNBC advancement, in order to produce a more efficient therapeutic strategy [16].
With the aim to explore the mechanisms of tumor initiation, progression and metastasis and develop new targeted therapies for TNBC, studies have focused on the signalling pathways deregulation and genes alternation related to TNBC in the past few years. Gene expression profiling is a beneficial tool to find differentially expressed genes (DEGs) in human TNBC so as to find possible critical genes or transcription factors that play crucial roles in the regulation of TNBC development and progression [13]. Numerous previous studies have identified some genes which may be used as diagnostic markers or therapy targets for TNBC. For example, FOXA1, KRT18, and XBP1 are found to be over-expressed in TNBC and involved in many important cell functions contribute to tumorigenesis [17]. FOXA1, a widely cytokeratin protein participated in many physiological and pathological processes such as lymphocyte homing and activation, cell survival and migration, and tumour growth and metastasis [18]. It is reported FOXA1, KRT18, and XBP1 are expressed at both high transcriptional and translational levels in TNBC and its expression is related to the degree of malignancy [19]. Furthermore, the up-regulation of FOXA1, KRT18, and XBP1 in malignant TNBC may be an indication of tumour cell growth and migration. ASNA1, NDUFS8, NDUFV1 and NDUFB7 [20] have all been studied and their possibilities to be used as targets for diagnosis and therapy of TNBC have also been evaluated. However these studies just noted a few DEGs and the interaction among these genes were still unknown.
In 2016, Marsh et al. [21] performed a microarray data analysis based on GSE75333 to identify gene expression changes responding to TNBC between normal fibroblasts, granulin-stimulated fibroblasts and cancer-associated fibroblasts. They obtained numerous differentially expressed genes (DEGs) between normal fibroblasts, granulin-stimulated fibroblasts and cancer-associated fibroblasts and found that the DEGs and their related function. This demonstrated there still existed distance between clinical application and the laboratory test, and more tests should be done to identify more candidates for therapy for TNBC.
In order to identify more genetic candidates for therapy for TNBC, expression microarray data GSE75333 deposited in Gene Expression Omnibus (GEO) by Marsh et al. [21]. DEGs in this dataset were analyze and their interaction relation were investigated by protein-protein interaction (PPI) network. Moreover, possible functions of DEGs involved in the PPI network were evaluated by gene and pathway enrichment analyses. The functions of the significant DEGs involved in significant function terms would be argued and some of them might be treated as the candidates for TNBC therapy.
Materials and methods
Affymetrix microarray data
The gene expression profile data of GSE75333 based on theplatform of GPL570 (Affymetrix Human Genome U133_Plus_2 Array) (Affymetrix Inc., Santa Clara, CA, USA) were downloaded from Gene Expression Omnibus (GEO) database in National Center for Biotechnology Information (NCBI, http://www.ncbi.nlm.nih.gov/geo/), which was deposited by Marsh et al. [21]. The datasets available in this analysis contained nine samples, including three normal fibroblasts cultures, three granulin-stimulated fibroblasts cultures and three cancer-associated fibroblasts cultures.
Identification of DEGs
The raw data of the mRNA expression profiles were downloaded and investigated by R language software [22]. Background correction, quartile data normalization, and probe summarization were tested for the authentic data. The limma [23] technique in Bioconductor (http://www.bioconductor.org/) was used to analyze genes which were differentially expressed between normal fibroblasts, granulin-stimulated fibroblasts and cancer-associated fibroblasts; the significance of DEGs was determined by t test and was characterized by p value. To cut down the risk of false positives, p values were adjusted for multiple testing using the Benjamini–Hochberg False Discovery Rate (FDR) technique. The revised p value was characterized by FDR [24]. FDR < 0.05 were treated as the cut off values for DEG screening.
Gene ontology (GO) analysis of DEGs
Gene ontology is an effective tool for compiling an enormous number of gene annotation terms [25]. The Lynx tool for a database and knowledge extraction engine for integrative medicine [26], is bioinformatics resources consisting of an integrated biological knowledgebase and analytic tools aimed at consistently elicit biological functional annotation from enormous gene/protein lists, such as being derived from high-throughput genomic experiments. To hike the extensive sympathetic of the biological functions of DEGs, Lynx tool was used to obtain the enriched GO terms of DEGs based on the p < 0.05 was set as the threshold value.
Bio-pathway analysis of DEGs
WIKIPATHWAYS, REACTOME, NCI and KEGG are a database resource for understanding functions of genes list from molecular level [27,28,29,30]. Lynx is a valuable tool to functionally interpret results from experimental techniques in genomics [26]. This web-based application consolidate different sources of information for discovering groups of genes with similar biological meaning. The enrichment analysis of Lynx is crucial in the analysis of high-throughput experiments. In the study, Lynx software was used to test the statistical enrichment of DEGs in pathways. p < 0.05 was set as the threshold value.
Construction of protein-protein interaction (PPI) network
The online database resource InnateDB keeps individually broad coverage, and access to predicted and experimental interaction information [31]. Interactions in the InnateDB are administered with a confidence score [31]. In the current study, application of the InnateDB (http://innatedb.com/) was used to predict PPIs based on a confidence score > 0.4 and other default parameters. The PPI network was visualized using network analyst (http://www.networkanalyst.ca/) [32]. Subsequently, a PPI network was build up based on the analyzed DEGs. A hypergeometric algorithm was used, and p < 0.05 was treated to express statistically significant differences.
Prediction of target miRNAs for DEGs
DIANA-TarBasev7.0 (http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=tarbase/index) [33] is an integrated system for exploring large sets of gene. DIANA-TarBase was used to identify hsa-mir s corresponding to DEGs. Hsa-mirs with DEGs number > 2 and raw p < 0.05 were identified as target hsa-mirs. The gene hsa-mir network was visualized with network analyst (http://www.networkanalyst.ca/).
Results
Preprocessing of data
After preprocessing and data integration, we obtained 9 samples, including expression data matrix of 52,238 gene probes. Through comparing the quality of expression values of microarray before and after normalization, we found that the medians of expression value of nine samples were in a straight line (Fig. 1).
Identification of DEGs
Among the 52,238 genes identified from the gene expression profile microarray analysis, 190 differentially expressed genes were screened out (logFC| ≥ 1 and p < 0.05) (Table 1). Among these, 66 genes were up-regulated, and 124 genes were down-regulated. A “volcano plot” was constructed by plotting the differentially expressed genes between the p values and fold changes (Fig. 2). The corresponding Venn diagrams are shown in Fig. 3. The overlapping part represents the number of overlapping DEGs between groups, whereas the non-overlapping part represents the number of unique DEGs between groups.
Venn diagram of differentially expressed genes (DEGs) in three groups. A vs. B represents DEGs between A and B; A vs. C represents DEGs between A and C; B vs. C represents DEGs between B and C. A normal fibroblasts group, B granulin-stimulated fibroblasts group, C cancer-associated fibroblasts group
Hierarchical clustering analysis of DEGs
After extracting the expression values of the DEGs, hierarchical clustering analysis was conducted for the DEGs. As shown in Fig. 4, the DEGs could clearly distinguish the normal fibroblasts samples from the granulin-stimulated fibroblasts and normal fibroblasts samples from the cancer-associated fibroblasts samples. In granulin-stimulated fibroblasts and cancer-associated fibroblasts samples, there were more down-regulated genes than up-regulated genes (Fig. 4).
Heat map of differentially expressed genes. The red colour represents down-regulated gene and the green represents up-regulated gene in TNBC samples. Legend on the top left indicate log fold change of genes (A1, A2, A3: normal fibroblasts group; B1, B2, B3: granulin-stimulated fibroblasts group; C1, C2, C3: cancer-associated fibroblasts group)
GO enrichment analysis
After the GO enrichment analysis, the DEGs in the PPI network of up-regulated genes were mainly associated with the GO BP categories of cell-cell signalling(including ADM, EFNB2, NOV, SSTR1, TFAP2C and TRHDE, p = 0.0000562) and negative regulation of cell proliferation (including ADM, BCHE, CDKN2A, CDKN2B, IGFBP3, SSTR1, and TWIST2, p = 0.0000733), GO MF categories of insulin-like growth factor II binding (including IGFBP3 and IGFBP5, p = 0.000201) and insulin-like growth factor I binding (including IGFBP3 and IGFBP5, p = 0.000258) and GO CC categories of insulin-like growth factor binding protein complex (including IGFBP3and IGFBP5, p = 0.0000217) and integral component of plasma membrane (including EFNB2, ENPP2, EPHA5, FLRT3, KCND2, KCNJ8, MME, NLGN1, PTGER3, SLC38A5, SSTR1 and TRHDE, p = 0.000258) (Table 2).
Furthermore, the down-regulated genes were essentially involved in the GO BP categories of cell adhesion (including CCL2, CDH6, COL18A1, COL8A1, COMP, CXCL12, ENG, ITGA6, MCAM, NCAM1, NTM, PCDHB15, PODXL, SORBS2, SUSD5, and VCAM1, p = 9.03e−10) and extracellular matrix organization (including ADAMTS5, BMP4, COL10A1, COL11A1, COL18A1, COL8A1, COMP, EFEMP1, ITGA6, NCAM1, TGFB2 and VCAM1, p = 4.65e−8), GO MP categories of N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase activity (including CHST11and CHST15, p = 0.0000243) and calcium ion binding (including ANXA3, CDH6, CELSR1, COMP, EFEMP1, FLG, GALNT3, JAG1, MGP, PCDHB14, PCDHB15, SCUBE3 and SYTL2, p = 0.0000335) and GO CC categories of extracellular space (including ACTA2, BMP4, CCL2, COL18A1, COMP, CXCL12, EFEMP1, ENG, GPX3, IL26, KRT81, LRRC17, MCAM, PLAU, PODXL, TGFB2, TIMP3 and VCAM1, p = 0.000258) and extracellular matrix (including COL18A1, COL8A1, COMP, EFEMP1, MGP, TGFB2 and TIMP3, p = 0.000258) (Table 2).
Pathway enrichment analysis
After the pathway enrichment analysis, up-regulated genes were mainly associated with the WIKIPATHWAYS categories of myometrial relaxation and contraction pathways (including ADM, ATF5, IGFBP3 and IGFBP5, p = 0.001) (Table 3).
Furthermore, the down-regulated genes were essentially involved in the WIKIPATHWAYS categories of endochondral ossification (ADAMTS5, CHST11, COL10A1, MGP, PLAU, RUNX3, SOX6, TGFB2, TIMP3, p = 4.27e−11) and TGF beta signalling pathway (including BMP4, ENG, LEF1 and RUNX3, p = 0.00016), REACTOME categories of extracellular matrix organization (including ADAMTS5, CHST11, COL10A1, MGP, PLAU, RUNX3, SOX6, TGFB2 and TIMP3, p = 9.35e−9) and integrin cell surface interactions (including COL10A1, COL18A1, COL8A1, COMP, ITGA6 and VCAM1, p = 0.00000408) and PID_NCI categories of beta1 integrin cell surface interactions (including COL11A1, COL18A1, ITGA6, PLAU, TGM2 and VCAM1, p = 8.35e−7), and KEGG categories of malaria (including CCL2, COMP, TGFB2 and VCAM1, p = 0.000102) and glycosaminoglycan biosynthesis—chondroitin sulfate/dermatansulfate (including CHST11, CHST15 and CHSY3, p = 0.000127) (Table 3).
PPI network analysis
The PPI networks of up- and down-regulated genes are shown in Figs. 5a and 6a, respectively. The up-regulated network was constructed with 472 nodes and 529 edges (A). The proteins cyclin dependent kinase inhibitor 2A (CDKN2A, degree = 148), membrane matalloendopeptidase (MME, degree = 78), PBX homebox 1 (PBX1, degree = 39), insulin like growth factor binding protein 3 (IGFBP3, degree = 25) and transcription factor AP-2 gamma (TFAP2C, degree = 23) were hub nodes in this network. The distribution of node degrees complied with exponential distribution. R squared and correlation coefficient are 0.618 and 0.898, respectively (Fig. 7a). The down-regulated PPI network was constructed with 1143 nodes and 1374 edges (B). The proteins vascular cell adhesion molecule (VCAM1, degree = 426), keratin 18 (KRT18, degree = 89), transglutaminase 2 (TGM2, degree = 69), actin, alpha 22, smooth muscle, aorta (ACTA2, degree = 55) and STAM binding protein (STAMBP, degree = 44) were hub proteins in this network. The distribution of node degrees complied with exponential distribution. R squared and correlation coefficient are 0.684 and 0.914, respectively (Fig. 7b). After the connectivity degree analysis, the top 20 nodes with high degrees for the up- and down-regulated PPI network were screened (Table 4). The connectivity degrees of the top 20 nodes in the up-regulated and down-regulated PPI were all higher than 5.
Protein-protein interaction network of the DEGs (a). Top four PPI subnetworks in co-expression module (b–e) drawn from the protein-protein interaction network of the differentially expressed genes. The green circles represent for the up-regulated genes. The PPI pairs were identified with the required confidence (combined score) > 0.9 as a threshold, and the PPI network of these connections were constructed using NetworkAnalyst software
Protein-protein interaction network of the DEGs (a). Top four PPI subnetworks in co-expression module (b–e) drawn from the protein-protein interaction network of the differentially expressed genes. The red circles represent for the up-regulated genes. The PPI pairs were identified with the required confidence (combined score) > 0.9 as a threshold, and the PPI network of these connections were constructed using NetworkAnalyst software
Moreover, in the four subnetworks screened from up-regulated genes, several hub genes were also identified, such as MMP3 (Fig. 5b), CDH10 (Fig. 5c), LSAMP (Fig. 5d), CSGALNACT1 (Fig. 5e) and four subnetworks for down-regulated genes, such as CARD16 (Fig. 6b), COL10A1 (Fig. 6c), COL18A1 (Fig. 6d), and CELSR1 (Fig. 6e).
Prediction of target hsa-mir for DEGs
The target hsa-mir for up- and down-regulated genes are presented in Figs. 8 and 9, respectively. In the up-regulated gene hsa-mir network, hsa-mir-759 (n = 3) and hsa-mir-4446-5p (n = 4) regulated the most up-regulated genes. Insulin like growth factor binding protein 5 (IGFBP5) was regulated by 143 hsa-mirs, such as hsa-mir-219a-1-3p and hsa-mir-4446-5p. Besides, hsa-mir-26a-5p (n = 5) and hsa-mir- 301a-3p (n = 5) families regulated the most down-regulated gene hsa-mir network. Gene myosin regulatory light chain interacting proteins (MYLIP) was the hub node and was regulated by 113 hsa-mirs, including hsa-mir-26a-5p and hsa-mir- 301a-3p.
Discussion
Firstly, total 66 up-regulated and 124 down-regulated genes were identified between three normal fibroblasts cultures, three granulin-stimulated fibroblasts cultures and three cancer-associated fibroblasts cultures. The result indicates that the common DEGs might play important roles during TNBC development and progression. In order to explore the potential roles of these DEGs, we performed functional analyses to them. The GO term analysis showed that up-regulated DEGs were mainly involved in cell-cell signalling and negative regulation of cell proliferation with high significant p value. The previous study shows that cell-cell signaling is mainly involved in TNBC development [34]. ADM, EFNB2, NOV, SSTR1, TFAP2C and TRHDE are biomarkers enriched in cell-cell signalling. Smirnov et al., 2006 [35] showed that ADM (adrenomedullin) was a potent vasodilator and a hypotensive agent and mainly associated with metastatic carcinomas. Higher expression of EFNB2 (ephrinB2) controls the arterial/venous specialization and vessel branching in TNBC [36]. NOV (nephroblastoma overexpressed) is a connective tissue growth factor is highly expressed in TNBC [37]. Hormone receptor gene SSTR1 (somatostatin receptor 1) play an important role in breast cancer [38]. Most of the breast cancers expressing estrogen receptor-α (ERα). TFAP2C (transcription factor AP-2 gamma) control the expression of ERα directly by binding to the ERα promoter region [39]. TRHDE (thyrotropin releasing hormone degrading enzyme) gene might increase the risk of TNBC development [40]. The key role of negative regulation of cell proliferation is associated with cancer metastasis and angiogenesis [41]. ADM, BCHE, CDKN2A, CDKN2B, IGFBP3, SSTR1 and TWIST2 are the biomarkers enriched in negative regulation of cell proliferation. Increased expression of BCHE (butyrylcholinesterase) was found in TNBC [42]. Increased risk of cancer with germline CDKN2A (cyclin-dependent kinase inhibitor 2 A) and CDKN2B (cyclin-dependent kinase inhibitor 2 B) mutation [43, 44]. High expression of mutated version of CDKN2A and CDKN2B gene in TNBC [42, 45]. Activation of altered IGFBP3(insulin like growth factor binding protein 3) stimulates mitosis and inhibits apoptosis [46]. And Key et al. [47] found the increased expression of IGFBP3 was correlated with prognosis in patients with breast cancer. Expression of TWIST2 (twist family bHLH transcription factor 2) is a highly conserved basic helix-loop-helix transcription factor that play in embryogenesis and promotes cancer invasion [48]. Accumulating evidence showed that over-expression of TWIST2 was involved in breast cancer development [49]. Similarly, GO term analysis showed that down-regulated DEGs were mainly involved in cell adhesion and extracellular matrix organization with high significant p value. Cell-to-cell and cell-to-extracellular matrix adhesion controls the social behavior of cells in tumour development [50, 51]. Carey et al., 2010 [52] reported that cell adhesion is primarily involved in TNBC development. Molecular markers such as CCL2, CDH6, COL18A1, COL8A1, COMP, CXCL12, ENG, ITGA6, MCAM, NCAM1, NTM, PCDHB15, PODXL, SORBS2, SUSD5 and VCAM1 enriched in cell adhesion. CCL2 (C-C motif chemokine ligand 2) negotiate between cancer cells and stromal fibroblasts that control cancer progression [53]. Expression of CCL2 is involved TNBC development [54]. Expression of CDH6 (cadherin 6) in the metastatic progression of many cancer types [55]. Polymorphism of the COL18A1 (collagen type XVIII alpha 1 chain) gene in breast cancer [56]. High molecular weight keratin COL8A1 (collagen type VIII alpha 1 chain) and variable elaboration that is diagnose in ~ 10–15% of breast cancers [57]. COMP (cartilage oligomeric matrix protein) is one of the most auspicious serologic markers with regard to an ability to prognose development of cancer and also play key roles in chondrogenesis and cartilage development in cancer [58]. CXCL12 (C-X-C motif chemokine ligand 12) and its specialized receptor, CXCR4, have newly been shown to be involved in tumorigenesis, proliferation and angiogenesis [59]. CXCL12 play a critical role in the progression in breast cancer through participating in cell adhesion [60]. ENG (Endoglin) is highly expressed in the tumor-associated vascular endothelium and is an component receptor for TGF-β that has been implicated in cancer cell detachment, migration, and invasiveness [61]. Molecular marker ENG play critical role in breast cancer progression [62]. ITGA6 (integrin subunit alpha 6) is tumour suppresser genes and mutation of this gene results in cancer progression [63]. Biomarker ITGA6 is found to be strongly associated with the TNBC [64]. MCAM (melanoma cell adhesion molecule) is a cell-surface glycoprotein molecule that is actively expressed on leading human carcinoma [65]. MCAM protein is confirmed to induce cell adhesion and induces TNBC [66]. NCAM1 (neuronal cellular adhesion molecule) expression has been corresponded with the existence of perineural invasion in specimens from a variety of tumours [67]. NCAM1 highly expressed in advanced breast cancer [68]. NTM (neurotrimin) is novel cell adhesion biomarkers expressed in many cancer types [69]. It has been reported in TNBC development [70]. Mutated PCDHB15 (protocadherin beta 15) expression was associated TNBC [71]. Molecular markers PODXL (podocalyxin like) is involved in the arrangement of both adhesion and cell morphology and cancer development [72]. Castro et al. [73] showed PODXL involvement in TNBC progression. SORBS2 (sorbin and SH3 domain containing 2) is a adapter protein that plays a key role in the assembling of signalling complexes, being a link between ABL kinases and actin cytoskeleton and also express in most of cancer development [74]. SUSD5 (sushi domain containing 5) is involved in metastatic colonization of most of the cancers [75]. SUSD5 is a one of the prognostic marker in breast cancer [76]. VCAM1 (vascular cell adhesion molecule-1) is an endothelial cell membrane glycoprotein that has been implicated in leukocyte/endothelial cell interactions in cancer cell metastasis [77]. VCAM1 was a pivotal contributor to TNBC progression [78]. ADAMTS5, BMP4, COL10A1, COL11A1, COL18A1, COL8A1, COMP, EFEMP1, ITGA6, NCAM1, TGFB2 and VCAM1 are the gene highly enriched in extracellular matrix organization. Extracellular matrix enzyme ADAMTS5 (ADAM metallopeptidase with thrombospondin type 1 motif) has emerged as key players in angiogenesis and cancer development [79]. BMP4 (bone morphogenetic protein 4) controls distinct cellular processes, such as proliferation, differentiation, and apoptosis [80]. Expression of BMP4 plays a positive role in progression of breast cancer [81]. COL10A1 (collagen type X alpha 1 chain) promotes cell proliferation in cancer development [82]. Accumulating evidence has demonstrated that expression of COL10A1 is involved in TNBC development [83]. COL11A1 (collagen type XI alpha 1 chain) important in cell invasiveness and tumour formation [84]. Molecular marker COL11A1 involves in the TNBC progression [85]. The polymorphisms of EFEMP1 (EGF containing fibulin like extracellular matrix protein 1) gene were identified with breast cancer and might share to the susceptibility of the progression of TNBC [86]. A polymorphism in the promoter of TGFB2 (transforming growth factor beta 2) that intensify expression of the protein was related with lymph node metastasis in TNBC patients, pointing to a role of TGFB2 in the process of invasion [87].
The enriched Wikipathways pathways of up-regulated DEGs is myometrial relaxation and contraction pathways. Previous studies have shown that myometrial Relaxation and contraction pathways up-regulated genes in TNBC development can predict the overall survival of TNBC patients [88]. Several common genes enriched in these above pathways (ADM, ATF5, IGFBP3 and IGFBP5) showed up regulation. Biomarker ATF5 (activating transcription factor 5) encourage invasion by activating the expression of integrin-alpha2 and integrin-beta1 in several human cancer cell [89]. Accumulating evidence showed that over-expression of ATF5 was involved in TNBC [90]. Reticence of IGFBP3 (insulin like growth factor binding protein 3) and IGFBP5 (insulin like growth factor binding protein 5) imbalance their proliferative action and programmed cell death in breast cancer [91, 92]. The Wikipathways results showed that the down-regulated genes were significantly enriched in two pathways, which included endochondral ossification and TGF beta signalling pathway. Endochondral ossification play key role in vascularization particularly in cancer development [93]. TGF-beta signalling reinforce tumour recurrence through IL-8–dependent expansion of cancer stem-like cells (CSCs) [94]. Genes such as ADAMTS5, CHST11, COL10A1, MGP, PLAU, RUNX3, SOX6, TGFB2 and TIMP3, were significantly enriched in endochondral ossification. Enzyme CHST11 (carbohydrate sulfotransferase 11) catalyzes the transfer of sulfate to position 4 of the N-acetylgalactosamine (GalNAc) residue of chondroitin [95]. CHST11 may play a explicit role in advancement of breast cancer and that its expression is restrained by DNA methylation [96]. Among the proteins involved in vascular calcium metabolism, the vitamin K-dependent MGP (matrix Gla-protein) plays a dominant role in breast cancer development [97]. PLAU (plasminogen activator, urokinase) play essential roles in tumour invasion and metastasis [98]. PLAU genes have been reported participate in the breast cancer development [99]. RUNX3 (Runt-related transcription factor 3) is a contender tumour suppressor gene and that is down-regulated in diverse cancers [100] and also activation of Wnt/β-catenin signalling in TNBC [101]. SOX6 (SRY-box 6) is tumor-suppressive function and its inactivation results in cancer progression [102]. Pinto et al. [103] found that low expression of SOX6 results in breast cancer development. Clinical exercise found that TIMP3(TIMP metallopeptidase inhibitor 3) was silenced in a number of cancer types [104]. Yuan et al. [105] revealed that TIMP3 silencing results in TNBC development. BMP4, ENG, LEF1 and RUNX3 are the down-regulated genes enriched in TGF beta signalling pathway. LEF1 (lymphoid enhancer binding factor 1) aberrantly controlled signalling pathways in cancer the WNT/β-catenin pathway plays an dominant role, since it was shown to be perilously involved in a wide range of cancer developments [106]. Delaunay and colleagues [107] demonstrated that LEF1 gene play important role breast cancer development. And for reactome pathway results showed that the down-regulated genes were most significantly enriched in extracellular matrix organization and integrin cell surface interactions. ADAMTS5, CHST11, COL10A1, MGP, PLAU, RUNX3, SOX6, TGFB2 and TIMP3 are the gene enriched in included extracellular matrix organization. Modification of extracellular matrix organization results in TNBC progression [108]. Integrin cell surface interactions including 23 alterations in integrin, laminin and collagen genes results in TNBC progression [109]. COL10A1, COL18A1, COL8A1, COMP, ITGA6 and VCAM1 are the gene enriched in integrin cell surface interactions. PID_NCI pathway results showed that the down-regulated genes were most significantly enriched in beta1 integrin cell surface interactions. Beta1 integrin cell surface interactions induces cellular proliferation results in breast cancer development [110]. COL11A1, COL18A1, ITGA6, PLAU, TGM2 and VCAM1 are gene enriched in beta1 integrin cell surface interactions. Finally KEGG pathway results showed that the down-regulated genes were significantly enriched in malaria and glycosaminoglycan biosynthesis—chondroitin sulfate/dermatansulfate. Glycosaminoglycan biosynthesis is crucial in TNBC development [111]. CHST11, CHST15 and CHSY3 are the gene enriched in glycosaminoglycan biosynthesis. Therefore, these results were consistent with previous studies and we identified critical genes involved in the PPI network.
We constructed the PPI network with up-regulated DEGs and list the top degree hub genes: CDKN2A, MME, PBX1, IGFBP3 and TFAP2C. CDKN2A was identified as one of the hub genes exhibiting the highest degree of connectivity. Over expression of MME (membrane metalloendopeptidase) plays a key role in the pathogenesis of TNBC [112]. Another hub gene PBX1 (PBX homeobox 1) is a TALE homeodomain protein and a proto-oncogene involved in the development of different types of cancers [113]. Moreover, proto oncogene PBX1 promotes the breast cancer development [114]. TFAP2C (transcription factor AP-2 gamma) is a transcription factor, which plays a very important role in the control of both estrogen receptor-alpha (ERα) and c-ErbB2/HER2 (Her2) and also promotes breast cancer development [115]. We also constructed PPI network for down-regulated genes and VCAM1, KRT18, TGM2, ACTA2 and STAMBP are the top five hub genes. VCAM1 was identified as one of the hub genes exhibiting the highest degree of connectivity. KRT18 (keratin 81) is a epithelial differentiation marker and it encodes the type I intermediate filament chain keratin 18 in breast cancer [116]. TGM2 (Transglutaminase 2) plays a crucial role in cancer cell growth and endurance through the antiapoptosis signalling pathway [117]. Down regulation of TGM2 in cancer cells is an important pathogenic factor in breast cancer [118]. Down regulation of epithelial–mesenchymal transition-associated (EMT-associated) gene ACTA2 (actin, alpha 2, smooth muscle, aorta) was correlated to invasion in TNBC [119]. Mutation of STAMBP (STAM binding protein) gene results in cancer development [120].
Subnetwork analysis of the PPI network for up-regulated genes revealed that the development of TNBC was associated with MMP3, CDH10, LSAMP and CSGALNACT1 genes. MMP3 (matrix metallopeptidase 3) is class of matrix metalloproteinase enzyme and this gene is associated with tumour cell invasion and metastasis with their promoter polymorphisms regulating the level of transcription [121]. MMP3 was a pivotal contributor to TNBC progression and could function as a potential therapeutic target [122]. CDH10 (cadherin 10) gene encodes a member of the cadherin family of calcium-dependent glycoproteins that mediate cell adhesion and controls many cellular events during cancer development [123]. CDH10 is important biomarker in maintenance of cell adhesion and polarity, alterations of which contribute to TNBC development [124]. LSAMP (limbic system-associated membrane protein) is a tumour suppressor gene and mutation of this gene results in cancer development [125]. Evidence reflected that inactivation of LSAMP gene may result in TNBC progression [126]. CSGALNACT1 (chondroitin sulfate N-acetylgalactosaminyltransferase 1) gene for enzymes generating chondroitin sulfate glycosaminoglycans and altered expression of this gene results in breast cancer development [127]. We also extracted subnetwork form PPI network of down-regulated genes and CARD16, COL10A1, COL18A1 and CELSR1 hub genes in this subnetwork with highest degree of connectivity. Mutation of CARD16 (caspase recruitment domain family member 16) gene and harbour this specific genes relevant to breast cancer development [128]. The CELSR1 (cadherin EGF LAG seven-pass G-type receptor 1) gene express a biomarker that is a member of the flamingo subfamily, which is factor of the cadherin superfamily and is main factor for breast cancer development [129].
Micro RNA play essential role in cancer progression [130]. Aberrant microRNA expression profiles have been identified in breast cancer [131]. Apart from DEGs and their functions, hsa-mirs such as hsa-mir-759 and hsa-mir-4446-5p for up-regulated genes and hsa-mir-26a-5p and hsa-mir-301a-3p for down-regulated genes may be important for the progression of TNBC. Recently, it has been reported that the expression of the fibrinogen alpha gene regulated by hsa-mir-759 was associated with susceptibility to TNBC [132]. Micro RNA (hsa-mir-4446-5p) is binding site for SHANK2(SH3 and multiple ankyrin repeat domains protein 2) and is responsible for breast cancer progression [133]. Micro RNA (hsa-miR-26a-5p) act as tumour suppressors in several types of cancers targeting oncogenic genes, such as breast cancer [134], nasopharyngeal carcinoma [135], and hepatocellular carcinoma [136]. Previuos report said that hsa-miR-301a-3p acts as an oncogene in TNBC [137].
In conclusion, our data provide a comprehensive bioinformatics analysis of DEGs, which may be involved in the progress of TNBC. As a result of this preliminary study, we confirm that these DEGs, including CDKN2A, MME, PBX1, IGFBP3, TFAP2C, VCAM1, KRT18, TGM2, ACTA2, and STAMBP, may play a role in the TNBC development and could be candidate molecular targets for the treatment of TNBC. In addition, extracellular matrix organization and cell adhesion may play important roles in promoting development of TNBC. The study provides a set of useful targets for future investigation into the molecular mechanisms and biomarkers. However, further molecular biological experiments are required to confirm the function of the identified genes in TNBC.
References
Jung YY, Kim HM, Koo JS (2016) The role of cancer-associated fibroblasts in breast cancer pathobiology. Histol Histopathol 31(4):371–378. doi:10.14670/HH-11-700
Luo H, Tu G, Liu Z, Liu M (2015) Cancer-associated fibroblasts: a multifaceted driver of breast cancer progression. Cancer Lett 361(2):155–163. doi:10.1016/j.canlet.2015.02.018
Qiao A, Gu F, Guo X, Zhang X, Fu L (2016) Breast cancer-associated fibroblasts: their roles in tumor initiation, progression and clinical applications. Front Med 10(1):33–40. doi:10.1007/s11684-016-0431-5
Sugimoto H, Mundel TM, Kieran MW, Kalluri R (2006) Identification of fibroblast heterogeneity in the tumor microenvironment. Cancer Biol Ther 5(12):1640–1646
Pasanen I, Lehtonen S, Sormunen R, Skarp S, Lehtilahti E, Pietilä M, Sequeiros RB, Lehenkari P, Kuvaja P (2016) Breast cancer carcinoma-associated fibroblasts differ from breast fibroblasts in immunological and extracellular matrix regulating pathways. Exp Cell Res 344(1):53–66. doi:10.1016/j.yexcr.2016.04.016
Sappino AP, Skalli O, Jackson B, Schürch W, Gabbiani G (1998) Smooth-muscle differentiation in stromal cells of malignant and non-malignant breast tissues. Int J Cancer 41(5):707–712
Polyak K, Weinberg RA (2009) Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer 9(4):265–273. doi:10.1038/nrc2620
Weber CE, Kothari AN, Wai PY, Li NY, Driver J, Zapf MA, Franzen CA, Gupta GN, Osipo C, Zlobin A, Syn WK, Zhang J, Kuo PC, Mi Z (2015) Osteopontin mediates an MZF1-TGF-β1-dependent transformation of mesenchymal stem cells into cancer associated fibroblasts in breast cancer. Oncogene 34(37):4821–4833. doi:10.1038/onc.2014.410
Rønnov-Jessen L, Petersen OW, Koteliansky VE, Bissell MJ (1995) The origin of the myofibroblasts in breast cancer. Recapitulation of tumor environment in culture unravels diversity and implicates converted fibroblasts and recruited smooth muscle cells. J Clin Investig 95(2):859–873
Paulsson J, Sjöblom T, Micke P, Pontén F, Landberg G, Heldin CH, Bergh J, Brennan DJ, Jirström K, Ostman A (2009) Prognostic significance of stromal platelet-derived growth factor beta-receptor expression in human breast cancer. Am J Pathol 175(1):334–341. doi:10.2353/ajpath.2009.081030
El-Gendi SM, Mostafa MF, El-Gendi AM (2012) Stromal caveolin-1 expression in breast carcinoma. Correlation with early tumor recurrence and clinical outcome. Pathol Oncol Res 18(2):459–469. doi:10.1007/s12253-011-9469-5
Bauer KR, Brown M, Cress RD, Parise CA, Caggiano V (2007) Descriptive analysis of estrogen receptor (ER)-negative, progesterone receptor (PR)-negative, and HER2-negative invasive breast cancer, the so-called triple-negative phenotype: a population-based study from the California cancer Registry. Cancer 109(9):1721–1728. doi:10.1002/cncr.22618
Huo D, Ikpatt F, Khramtsov A, Dangou JM, Nanda R, Dignam J, Zhang B, Grushko T, Zhang C, Oluwasola O, Malaka D, Malami S, Odetunde A, Adeoye AO, Iyare F, Falusi A, Perou CM, Olopade OI (2009) Population differences in breast cancer: survey in indigenous African women reveals over-representation of triple-negative breast cancer. J Clin Oncol 27(27):4515–4521. doi:10.1200/JCO.2008.19.6873
Liedtke C, Mazouni C, Hess KR, André F, Tordai A, Mejia JA, Symmans WF, Gonzalez-Angulo AM, Hennessy B, Green M, Cristofanilli M, Hortobagyi GN, Pusztai L (2008) Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol 26(8):1275–1281. doi:10.1200/JCO.2007.14.4147
O’Shaughnessy J, Osborne C, Pippen JE, Yoffe M, Patt D, Rocha C, Koo IC, Sherman BM, Bradley C (2011) Iniparib plus chemotherapy in metastatic triple-negative breast cancer. N Engl J Med 364(3):205–214. doi:10.1056/NEJMoa1011418
Rakha EA, El-Sayed ME, Green AR, Lee AH, Robertson JF, Ellis IO (2007) Prognostic markers in triple-negative breast cancer. Cancer 109(1):25–32. doi:10.1002/cncr.22381
Farmer P, Bonnefoi H, Becette V, Tubiana-Hulin M, Fumoleau P, Larsimont D, Macgrogan G, Bergh J, Cameron D, Goldstein D, Duss S, Nicoulaz AL, Brisken C, Fiche M, Delorenzi M, Iggo R (2005) Identification of molecular apocrine breast tumours by microarray analysis. Oncogene 24(29):4660–4671. doi:10.1038/sj.onc.1208561
Hu Z, Fan C, Oh DS, Marron JS, He X, Qaqish BF, Livasy C, Carey LA, Reynolds E, Dressler L, Nobel A, Parker J, Ewend MG, Sawyer LR, Wu J, Liu Y, Nanda R, Tretiakova M, Ruiz Orrico A, Dreher D, Palazzo JP, Perreard L, Nelson E, Mone M, Hansen H, Mullins M, Quackenbush JF, Ellis MJ, Olopade OI, Bernard PS, Perou CM (2006) The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genom 7:96. doi:10.1186/1471-2164-7-96
Lal S, McCart Reed AE, de Luca XM, Simpson PT (2017) Molecular signatures in breast cancer. Methods S1046–2023(17):30058. doi:10.1016/j.ymeth.2017.06.032
Kreike B, van Kouwenhove M, Horlings H, Weigelt B, Peterse H, Bartelink H, van de Vijver MJ (2007) Gene expression profiling and histopathological characterization of triple-negative/basal-like breast carcinomas. Breast Cancer Res 9(5):R65. doi:10.1186/bcr1771
Marsh T, Wong I, Sceneay J, Barakat A, Qin Y, Sjödin A, Alspach E, Nilsson B, Stewart SA, McAllister SS (2016) Hematopoietic age at onset of triple-negative breast cancer dictates disease aggressiveness and progression. Cancer Res 76(10):2932–2943. doi:10.1158/0008-5472.CAN-15-3332
Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) Affy analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20(3):307–315. doi:10.1093/bioinformatics/btg405
Smyth GK (2005) LIMMA: linear models for microarray data. Bioinformatics and computational biology solutions using R and Bioconductor. Springer, New York, pp 397–420
Efron B, Tibshirani R (2002) Empirical Bayes methods and false discovery rates for microarrays. Genet Epidemiol 23(1):70–86. doi:10.1002/gepi.1124
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25(1):25–29. doi:10.1038/75556
Sulakhe D, Balasubramanian S, Xie B, Feng B, Taylor A, Wang S, Berrocal E, Dave U, Xu J, Börnigen D, Gilliam TC, Maltsev N (2014) Lynx: a database and knowledge extraction engine for integrative medicine. Nucleic Acids Res 42(Database issue):D1007–D1112. doi:10.1093/nar/gkt1166
Kelder T, van Iersel MP, Hanspers K, Kutmon M, Conklin BR, Evelo CT, Pico AR (2012) WikiPathways: building research communities on biological pathways. Nucleic Acids Res 40(Database issue):D1301–D1307. doi:10.1093/nar/gkr1074
D’Eustachio P (2011) Reactome knowledgebase of human biological pathways and processes. Methods Mol Biol 694:49–61. doi:10.1007/978-1-60761-977-2_4
Schaefer CF, Anthony K, Krupa S, Buchoff J, Day M, Hannay T, Buetow KH (2009) PID: the pathway interaction database. Nucleic Acids Res 37(Database issue):D674–D679. doi:10.1093/nar/gkn653
Minoru K, Susumu G (2000) KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 28(1):27–30. doi:10.1093/nar/28.1.27
Xia J, Gill EE, Hancock RE (2015) NetworkAnalyst for statistical, visual and network-based meta-analysis of gene expression data. Nat Protoc 10(6):823–844. doi:10.1038/nprot.2015.052
Breuer K, Foroushani AK, Laird MR, Chen C, Sribnaia A, Lo R, Winsor GL, Hancock RE, Brinkman FS, Lynn DJ (2013) InnateDB: systems biology of innate immunity and beyond—recent updates and continuing curation. Nucleic Acids Res 41(Database issue):D1228–D1233. doi:10.1093/nar/gks1147
Vlachos IS, Paraskevopoulou MD, Karagkouni D, Georgakilas G, Vergoulis T, Kanellos I, Anastasopoulos IL, Maniou S, Karathanou K, Kalfakakou D, Fevgas A, Dalamagas T, Hatzigeorgiou AG (2015) DIANA-TarBase v7.0: indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res 43(Database issue):D153–D159. doi:10.1093/nar/gku1215
Finn RS, Dering J, Ginther C, Wilson CA, Glaspy P, Tchekmedyian N, Slamon DJ (2007) Dasatinib, an orally active small molecule inhibitor of both the src and abl kinases, selectively inhibits growth of basal-type/”triple-negative” breast cancer cell lines growing in vitro. Breast Cancer Res Treat 105(3):319–326. doi:10.1007/s10549-006-9463-x
Smirnov DA, Foulk BW, Doyle GV, Connelly MC, Terstappen LW, O’Hara SM (2006) Global gene expression profiling of circulating endothelial cells in patients with metastatic carcinomas. Cancer Res 66(6):2918–2922. doi:10.1158/0008-5472.CAN-05-4003
da Motta LL, Ledaki I, Purshouse K, Haider S, De Bastiani MA, Baban D, Morotti M, Steers G, Wigfield S, Bridges E, Li JL, Knapp S, Ebner D, Klamt F, Harris AL, McIntyre A (2017) The BET inhibitor JQ1 selectively impairs tumour response to hypoxia and downregulates CA9 and angiogenesis in triple negative breast cancer. Oncogene 36(1):122–132. doi:10.1038/onc.2016.184
Haque I, Banerjee S, Mehta S, De A, Majumder M, Mayo MS, Kambhampati S, Campbell DR, Banerjee SK (2011) Cysteine-rich 61-connective tissue growth factor-nephroblastoma-overexpressed 5 (CCN5)/Wnt-1-induced signaling protein-2 (WISP-2) regulates microRNA-10b via hypoxia-inducible factor-1α-TWIST signaling networks in human breast cancer cells. J Biol Chem 286(50):43475–43485. doi:10.1074/jbc.M111.284158
Santagata S, Thakkar A, Ergonul A, Wang B, Woo T, Hu R, Harrell JC, McNamara G, Schwede M, Culhane AC, Kindelberger D, Rodig S, Richardson A, Schnitt SJ, Tamimi RM, Ince TA (2014) Taxonomy of breast cancer based on normal cell phenotype predicts outcome. J Clin Investig 124(2):859–870. doi:10.1172/JCI70941
Woodfield GW, Horan AD, Chen Y, Weigel RJ (2007) TFAP2C controls hormone response in breast cancer cells through multiple pathways of estrogen signaling. Cancer Res 67(18):8439–8443. doi:10.1158/0008-5472.CAN-07-2293
He DX, Gu F, Gao F, Hao JJ, Gong D, Gu XT, Mao AQ, Jin J, Fu L, Ma X (2016) Genome-wide profiles of methylation, microRNAs, and gene expression in chemoresistant breast cancer. Sci Rep 6:24706. doi:10.1038/srep24706
Liotta LA, Steeg PS, Stetler-Stevenson WG (1991) Cancer metastasis and angiogenesis: an imbalance of positive and negative regulation. Cell 64(2):327–336
Burstein MD, Tsimelzon A, Poage GM, Covington KR, Contreras A, Fuqua SA, Savage MI, Osborne CK, Hilsenbeck SG, Chang JC, Mills GB, Lau CC, Brown PH (2015) Comprehensive genomic analysis identifies novel subtypes and targets of triple-negative breast cancer. Clin Cancer Res 21(7):1688–1698. doi:10.1158/1078-0432.CCR-14-0432
Sargen MR, Merrill SL, Chu EY, Nathanson KL (2016) CDKN2A mutations with p14 loss predisposing to multiple nerve sheath tumours, melanoma, dysplastic naevi and internal malignancies: a case series and review of the literature. Br J Dermatol 175(4):785–789. doi:10.1111/bjd.14485
Suzuki H, Zhou X, Yin J, Lei J, Jiang HY, Suzuki Y, Chan T, Hannon GJ, Mergner WJ, Abraham JM (1995) Intragenic mutations of CDKN2B and CDKN2A in primary human esophageal cancers. Hum Mol Genet 4(10):1883–1887
Branham MT, Marzese DM, Laurito SR, Gago FE, Orozco JI, Tello OM, Vargas-Roig LM, Roqué M (2012) Methylation profile of triple-negative breast carcinomas. Oncogenesis 1:e17. doi:10.1038/oncsis.2012.17
Renehan AG, Zwahlen M, Minder C, O’Dwyer ST, Shalet SM, Egger M (2004) Insulin-like growth factor (IGF)-I, IGF binding protein-3, and cancer risk: systematic review and meta-regression analysis. Lancet 363(9418):1346–1353
Key TJ, Appleby PN, Reeves GK, Roddam AW (2010) Insulin-like growth factor 1 (IGF1), IGF binding protein 3 (IGFBP3), and breast cancer risk: pooled individual data analysis of 17 prospective studies. Lancet Oncol 11(6):530–542. doi:10.1016/S1470-2045(10)70095-4
Liu AY, Cai Y, Mao Y, Lin Y, Zheng H, Wu T, Huang Y, Fang X, Lin S, Feng Q, Huang Z, Yang T, Luo Q, Ouyang G (2014) Twist2 promotes self-renewal of liver cancer stem-like cells by regulating CD24. Carcinogenesis 35(3):537–545. doi:10.1093/carcin/bgt364
Fang X, Cai Y, Liu J, Wang Z, Wu Q, Zhang Z, Yang CJ, Yuan L, Ouyang G (2011) Twist2 contributes to breast cancer progression by promoting an epithelial–mesenchymal transition and cancer stem-like cell self-renewal. Oncogene 30(47):4707–4720. doi:10.1038/onc.2011.181
Willrodt AH, Beffinger M, Vranova M, Protsyuk D, Schuler K, Jadhav M, Heikenwalder M, van den Broek M, Borsig L, Winter CH (2017) Stromal expression of activated leukocyte cell adhesion molecule (ALCAM) promotes lung tumor growth and metastasis. Am J Pathol S0002–9440(17):30330–30339. doi:10.1016/j.ajpath.2017.07.008
Kass L, Erler JT, Dembo M, Weaver VM (2017) Mammary epithelial cell: influence of extracellular matrix composition and organization during development and tumorigenesis. Int J Biochem Cell Biol 39(11):1987–1994. doi:10.1016/j.biocel.2007.06.025
Carey L, Winer E, Viale G, Cameron D, Gianni L (2010) Triple-negative breast cancer: disease entity or title of convenience? Nat Rev Clin Oncol 7(12):683–692. doi:10.1038/nrclinonc.2010.154
Tsuyada A, Chow A, Wu J, Somlo G, Chu P, Loera S, Luu T, Li AX, Wu X, Ye W, Chen S, Zhou W, Yu Y, Wang YZ, Ren X, Li H, Scherle P, Kuroki Y, Wang SE (2012) CCL2 mediates cross-talk between cancer cells and stromal fibroblasts that regulates breast cancer stem cells. Cancer Res 72(11):2768–2779. doi:10.1158/0008-5472.CAN-11-3567
Bonapace L, Coissieux MM, Wyckoff J, Mertz KD, Varga Z, Junt T, Bentires-Alj M (2014) Cessation of CCL2 inhibition accelerates breast cancer metastasis by promoting angiogenesis. Nature 515(7525):130–133. doi:10.1038/nature13862
Gugnoni M, Sancisi V, Gandolfi G, Manzotti G, Ragazzi M, Giordano D, Tamagnini I, Tigano M, Frasoldati A, Piana S, Ciarrocchi A (2017) Cadherin-6 promotes EMT and cancer metastasis by restraining autophagy. Oncogene 36(5):667–677. doi:10.1038/onc.2016.237
Lourenço GJ, Cardoso-Filho C, Gonçales NS, Shinzato JY, Zeferino LC, Nascimento H, Costa FF, Gurgel MS, Lima CS (2006) A high risk of occurrence of sporadic breast cancer in individuals with the 104NN polymorphism of the COL18A1 gene. Breast Cancer Res Treat 100(3):335–338. doi:10.1007/s10549-006-9259-z
Vargas AC, McCart Reed AE, Waddell N, Lane A, Reid LE, Smart CE, Cocciardi S, da Silva L, Song S, Chenevix-Trench G, Simpson PT, Lakhani SR (2012) Gene expression profiling of tumour epithelial and stromal compartments during breast cancer progression. Breast Cancer Res Treat 135(1):153–165. doi:10.1007/s10549-012-2123-4
Neidhart M, Müller-Ladner U, Frey W, Bosserhoff AK, Colombani PC, Frey-Rindova P, Hummel KM, Gay RE, Häuselmann H, Gay S (2000) Increased serum levels of non-collagenous matrix proteins (cartilage oligomeric matrix protein and melanoma inhibitory activity) in marathon runners. Osteoarthritis Cartil 8(3):222–229. doi:10.1053/joca.1999.0293
Schrader AJ, Lechner O, Templin M, Dittmar KE, Machtens S, Mengel M, Probst-Kepper M, Franzke A, Wollensak T, Gatzlaff P, Atzpodien J, Buer J, Lauber J (2002) CXCR4/CXCL12 expression and signalling in kidney cancer. Br J Cancer 86(8):1250–1256. doi:10.1038/sj.bjc.6600221
Gil M, Seshadri M, Komorowski MP, Abrams SI, Kozbor D (2013) Targeting CXCL12/CXCR4 signaling with oncolytic virotherapy disrupts tumor vasculature and inhibits breast cancer metastases. Proc Natl Acad Sci USA 110(14):E1291–E1300. doi:10.1073/pnas.1220580110
Pérez-Gómez E, Del Castillo G, Juan Francisco S, López-Novoa JM, Bernabéu C, Quintanilla M (2010) The role of the TGF-β coreceptor endoglin in cancer. Sci World J 10:2367–2384. doi:10.1100/tsw.2010.230
Adhemar LF, José Manuel L, Fernando CS (2010) Angiogenesis and breast cancer. J Oncol 10:576384–576391. doi:10.1155/2010/576384
Kinoshita T, Nohata N, Hanazawa T, Kikkawa N, Yamamoto N, Yoshino H, Itesako T, Enokida H, Nakagawa M, Okamoto Y, Seki N (2013) Tumour-suppressive microRNA-29s inhibit cancer cell migration and invasion by targeting laminin-integrin signalling in head and neck squamous cell carcinoma. Br J Cancer 109(10):2636–2645. doi:10.1038/bjc.2013.607
Klahan S, Wu MS, Hsi E, Huang CC, Hou MF, Chang WC (2014) Computational analysis of mRNA expression profiles identifies the ITG family and PIK3R3 as crucial genes for regulating triple negative breast cancer cell migration. Biomed Res Int 2014:536591. doi:10.1155/2014/536591
Johnson JP, Bar-Eli M, Jansen B, Markhof E (1997) Melanoma progression-associated glycoprotein MUC18/MCAM mediates homotypic cell adhesion through interaction with a heterophilic ligand. Int J Cancer 73(5):769–774
Zeng Q, Li W, Lu D, Wu Z, Duan H, Luo Y, Feng J, Yang D, Fu L, Yan X (2012) CD146, an epithelial–mesenchymal transition inducer, is associated with triple-negative breast cancer. Proc Natl Acad Sci USA 109(4):1127–1132. doi:10.1073/pnas.1111053108
Takahashi S, Kato K, Nakamura K, Nakano R, Kubota K, Hamada H (2011) Neural cell adhesion molecule 2 as a target molecule for prostate and breast cancer gene therapy. Cancer Sci 102(4):808–814. doi:10.1111/j.1349-7006.2011.01855.x
Cavallaro U, Christofori G (2004) Multitasking in tumor progression: signaling functions of cell adhesion molecules. Ann N Y Acad Sci 1014:58–66
Struyk AF, Canoll PD, Wolfgang MJ, Rosen CL, D’Eustachio P, Salzer JL (1995) Cloning of neurotrimin defines a new subfamily of differentially expressed neural cell adhesion molecules. J Neurosci 15(3 Pt 2):2141–2156
Xie X, Ma L, Xi K, Zhang W, Fan D (2017) MicroRNA-183 suppresses neuropathic pain and expression of AMPA receptors by targeting mTOR/VEGF signaling pathway. Cell Physiol Biochem 41(1):181–192. doi:10.1159/000455987
Newman S, Howarth KD, Greenman CD, Bignell GR, Tavaré S, Edwards PA (2013) The relative timing of mutations in a breast cancer genome. PLoS One 8(6):e64991. doi:10.1371/journal.pone.0064991
Dallas MR, Chen SH, Streppel MM, Sharma S, Maitra A, Konstantopoulos K (2012) Sialofucosylated podocalyxin is a functional E- and L-selectin ligand expressed by metastaticpancreatic cancer cells. Am J Physiol Cell Physiol 303(6):C616–C624. doi:10.1152/ajpcell.00149.2012
Castro NP, Fedorova-Abrams ND, Merchant AS, Rangel MC, Nagaoka T, Karasawa H, Klauzinska M, Hewitt SM, Biswas K, Sharan SK, Salomon DS (2015) Cripto-1 as a novel therapeutic target for triple negative breast cancer. Oncotarget 6(14):11910–11929. doi:10.18632/oncotarget.4182
Liesenfeld M, Mosig S, Funke H, Jansen L, Runnebaum IB, Dürst M, Backsch C (2013) SORBS2 and TLR3 induce premature senescence in primary human fibroblasts and keratinocytes. BMC Cancer 13:507. doi:10.1186/1471-2407-13-507
Luis-Ravelo D, Antón I, Zandueta C, Valencia K, Ormazábal C, Martínez-Canarias S, Guruceaga E, Perurena N, Vicent S, De Las Rivas J, Lecanda F (2014) A gene signature of bone metastatic colonization sensitizes for tumor-induced osteolysis and predicts survival in lung cancer. Oncogene 33(43):5090–5099. doi:10.1038/onc.2013.440
Drake PM, Schilling B, Niles RK, Prakobphol A, Li B, Jung K, Cho W, Braten M, Inerowicz HD, Williams K, Albertolle M, Held JM, Iacovides D, Sorensen DJ, Griffith OL, Johansen E, Zawadzka AM, Cusack MP, Allen S, Gormley M, Hall SC, Witkowska HE, Gray JW, Regnier F, Gibson BW, Fisher SJ (2012) Lectin chromatography/mass spectrometry discovery workflow identifies putative biomarkers of aggressive breast cancers. J Proteome Res 11(4):2508–2520. doi:10.1021/pr201206w
Chen Q, Zhang XH, Massagué J (2011) Macrophage binding to receptor VCAM-1 transmits survival signals in breast cancer cells that invade the lungs. Cancer Cell 20(4):538–549. doi:10.1016/j.ccr.2011.08.025
Wushou A, Jiang YZ, Hou J, Liu YR, Guo XM, Shao ZM (2015) Development of triple-negative breast cancer radiosensitive gene signature and validation based on transcriptome analysis. Breast Cancer Res Treat 154(1):57–62. doi:10.1007/s10549-015-3611-0
Kumar S, Rao N, Ge R (2012) Emerging roles of ADAMTSs in angiogenesis and cancer. Cancers (Basel) 4(4):1252–1299. doi:10.3390/cancers4041252
Alarmo EL, Kuukasjärvi T, Karhu R, Kallioniemi A (2007) A comprehensive expression survey of bone morphogenetic proteins in breast cancer highlights the importance of BMP4 and BMP7. Breast Cancer Res Treat 103(2):239–246. doi:10.1007/s10549-006-9362-1
Cyr-Depauw C, Northey JJ, Tabariès S, Annis MG, Dong Z, Cory S, Hallett M, Rennhack JP, Andrechek ER, Siegel PM (2016) Chordin-like 1 suppresses bone morphogenetic protein 4-induced breast cancer cell migration and invasion. Mol Cell Biol 36(10):1509–1525. doi:10.1128/MCB.00600-15
Chapman KB, Prendes MJ, Sternberg H, Kidd JL, Funk WD, Wagner J, West MD (2012) COL10A1 expression is elevated in diverse solid tumor types and is associated with tumor vasculature. Future Oncol 8(8):1031–1040. doi:10.2217/fon.12.79
Yang F, Liu YH, Dong SY, Yao ZH, Lv L, Ma RM, Dai XX, Wang J, Zhang XH, Wang OC (2016) Co-expression networks revealed potential core lncRNAs in the triple-negative breast cancer. Gene. doi:10.1016/j.gene.2016.07.002
Wu YH, Chang TH, Huang YF, Huang HD, Chou CY (2014) COL11A1 promotes tumor progression and predicts poor clinical outcome in ovarian cancer. Oncogene 33(26):3432–3440. doi:10.1038/onc.2013.307
Kim Y, Kim J, Lee HD, Jeong J, Lee W, Lee KA (2013) Spectrum of EGFR gene copy number changes and KRAS gene mutation status in Korean triple negative breast cancer patients. PLoS One 8(10):e79014. doi:10.1371/journal.pone.0079014
Mathe A, Wong-Brown M, Morten B, Forbes JF, Braye SG, Avery-Kiejda KA, Scott RJ (2015) Novel genes associated with lymph node metastasis in triple negative breast cancer. Sci Rep 5:15832. doi:10.1038/srep15832
Hou C, Yang Z, Kang Y, Zhang Z, Fu M, He A, Zhang Z, Liao W (2015) MiR-193b regulates early chondrogenesis by inhibiting the TGF-beta2 signaling pathway. FEBS Lett 589(9):1040–1047. doi:10.1016/j.febslet.2015.02.017
Liu C, Louhimo R, Laakso M, Lehtonen R, Hautaniemi S (2015) Identification of sample-specific regulations using integrative network level analysis. BMC Cancer 15:319. doi:10.1186/s12885-015-1265-2
Gho JW, Ip WK, Chan KY, Law PT, Lai PB, Wong N (2008) Re-expression of transcription factor ATF5 in hepatocellular carcinoma induces G2-M arrest. Cancer Res 68(16):6743–6751. doi:10.1158/0008-5472.CAN-07-6469
Karagoz K, Sinha R, Arga KY (2015) Triple negative breast cancer: a multi-omics network discovery strategy for candidate targets and driving pathways. OMICS 19(2):115–130. doi:10.1089/omi.2014.0135
Gill ZP, Perks CM, Newcomb PV, Holly JM (1997) Insulin-like growth factor-binding protein (IGFBP-3) predisposes breast cancer cells to programmed cell death in a non-IGF-dependent manner. J Biol Chem 272(41):25602–25607
Voudouri K, Berdiaki A, Tzardi M, Tzanakakis GN, Nikitovic D (2015) Insulin-like growth factor and epidermal growth factor signaling in breast cancer cell growth: focus on endocrine resistant disease. Anal Cell Pathol (Amst) 2015:975495. doi:10.1155/2015/975495
Han B, Bhowmick N, Qu Y, Chung S, Giuliano AE, Cui X (2017) FOXC1: an emerging marker and therapeutic target for cancer. Oncogene 36(28):3957–3963. doi:10.1038/onc.2017.48
Bhola NE, Balko JM, Dugger TC, Kuba MG, Sánchez V, Sanders M, Stanford J, Cook RS, Arteaga CL (2013) TGF-β inhibition enhances chemotherapy action against triple-negative breast cancer. J Clin Investig 123(3):1348–1358. doi:10.1172/JCI65416
Oliveira-Ferrer L, Heßling A, Trillsch F, Mahner S, Milde-Langosch K (2015) Prognostic impact of chondroitin-4-sulfotransferase CHST11 in ovarian cancer. Tumour Biol 36(11):9023–9030. doi:10.1007/s13277-015-3652-3
Herman D, Leakey TI, Behrens A, Yao-Borengasser A, Cooney CA, Jousheghany F, Phanavanh B, Siegel ER, Safar AM, Korourian S, Kieber-Emmons T, Monzavi-Karbassi B (2015) CHST11 gene expression and DNA methylation in breast cancer. Int J Oncol 46(3):1243–1251. doi:10.3892/ijo.2015.2828
Hirota S, Ito A, Nagoshi J, Takeda M, Kurata A, Takatsuka Y, Kohri K, Nomura S, Kitamura Y (1995) Expression of bone matrix protein messenger ribonucleic acids in human breast cancers. Possible involvement of osteopontin in development of calcifying foci. Lab Investig 72(1):64–69
Kenny HA, Leonhardt P, Ladanyi A, Yamada SD, Montag A, Im HK, Jagadeeswaran S, Shaw DE, Mazar AP, Lengyel E (2011) Targeting the urokinase plasminogen activator receptor inhibits ovarian cancer metastasis. Clin Cancer Res 17(3):459–471. doi:10.1158/1078-0432.CCR-10-2258
Mali AV, Joshi AA, Hegde MV, Kadam ShS (2017) Enterolactone suppresses proliferation, migration and metastasis of MDA-MB-231 breast cancer cells through inhibition of uPA induced plasmin activation and MMPs-mediated ECM remodeling. Asian Pac J Cancer Prev 18(4):905–915. doi:10.22034/APJCP.2017.18.4.905
Nishina S, Shiraha H, Nakanishi Y, Tanaka S, Matsubara M, Takaoka N, Uemura M, Horiguchi S, Kataoka J, Iwamuro M, Yagi T, Yamamoto K (2011) Restored expression of the tumor suppressor gene RUNX3 reduces cancer stem cells in hepatocellular carcinoma by suppressing Jagged1-Notch signaling. Oncol Rep 26(3):523–531. doi:10.3892/or.2011.1336
Ouyang M, Li Y, Ye S, Ma J, Lu L, Lv W, Chang G, Li X, Li Q, Wang S, Wang W (2014) MicroRNA profiling implies new markers of chemoresistance of triple-negative breast cancer. PLoS One 9(5):e96228. doi:10.1371/journal.pone.0096228
Qin YR, Tang H, Xie F, Liu H, Zhu Y, Ai J, Chen L, Li Y, Kwong DL, Fu L, Guan XY (2011) Characterization of tumor-suppressive function of SOX6 in human esophageal squamous cell carcinoma. Clin Cancer Res 17(1):46–55. doi:10.1158/1078-0432.CCR-10-1155
Pinto R, De Summa S, Pilato B, Tommasi S (2014) DNA methylation and miRNAs regulation in hereditary breast cancer: epigenetic changes, players in transcriptional and post- transcriptional regulation in hereditary breast cancer. Curr Mol Med 14(1):45–57. doi:10.2174/1566524013666131203101405
Lin H, Zhang Y, Wang H, Xu D, Meng X, Shao Y, Lin C, Ye Y, Qian H, Wang S (2012) Tissue inhibitor of metalloproteinases-3 transfer suppresses malignant behaviors of colorectal cancer cells. Cancer Gene Ther 19(12):845–851. doi:10.1038/cgt.2012.70
Yuan ZY, Dai T, Wang SS, Peng RJ, Li XH, Qin T, Song LB, Wang X (2014) Overexpression of ETV4 protein in triple-negative breast cancer is associated with a higher risk of distant metastasis. Onco Targets Ther 7:1733–1742. doi:10.2147/OTT.S66692
Gehrke I, Gandhirajan RK, Kreuzer KA (2009) Targeting the WNT/beta-catenin/TCF/LEF1 axis in solid and haematological cancers: multiplicity of therapeutic options. Eur J Cancer 45(16):2759–2767. doi:10.1016/j.ejca.2009.08.003
Delaunay S, Rapino F, Tharun L, Zhou Z, Heukamp L, Termathe M, Shostak K, Klevernic I, Florin A, Desmecht H, Desmet CJ, Nguyen L, Leidel SA, Willis AE, Büttner R, Chariot A, Close P (2016) Elp3 links tRNA modification to IRES-dependent translation of LEF1 to sustain metastasis in breast cancer. J Exp Med 213(11):2503–2523. doi:10.1084/jem.20160397
Giricz O, Calvo V, Pero SC, Krag DN, Sparano JA, Kenny PA (2012) GRB7 is required for triple-negative breast cancer cell invasion and survival. Breast Cancer Res Treat 33(2):607–615. doi:10.1007/s10549-011-1822-6
Shah SP, Roth A, Goya R, Oloumi A, Ha G, Zhao Y, Turashvili G, Ding J, Tse K, Haffari G, Bashashati A, Prentice LM, Khattra J, Burleigh A, Yap D, Bernard V, McPherson A, Shumansky K, Crisan A, Giuliany R, Heravi-Moussavi A, Rosner J, Lai D, Birol I, Varhol R, Tam A, Dhalla N, Zeng T, Ma K, Chan SK, Griffith M, Moradian A, Cheng SW, Morin GB, Watson P, Gelmon K, Chia S, Chin SF, Curtis C, Rueda OM, Pharoah PD, Damaraju S, Mackey J, Hoon K, Harkins T, Tadigotla V, Sigaroudinia M, Gascard P, Tlsty T, Costello JF, Meyer IM, Eaves CJ, Wasserman WW, Jones S, Huntsman D, Hirst M, Caldas C, Marra MA, Aparicio S (2012) The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486(7403):395–399. doi:10.1038/nature10933
dos Santos PB, Zanetti JS, Ribeiro-Silva A, Beltrão EI (2012) Beta 1 integrin predicts survival in breast cancer: a clinicopathological and immunohistochemical study. Diagn Pathol 7:104. doi:10.1186/1746-1596-7-104
Mathe A, Wong-Brown M, Locke WJ, Stirzaker C, Braye SG, Forbes JF, Clark SJ, Avery-Kiejda KA, Scott RJ (2016) DNA methylation profile of triple negative breast cancer-specific genes comparing lymph node positive patients to lymph node negative patients. Sci Rep 6:33435. doi:10.1038/srep33435
Muñoz M, Coveñas R (2016) Neurokinin-1 receptor antagonists as antitumor drugs in gastrointestinal cancer: a new approach. Saudi J Gastroenterol 22(4):260–268. doi:10.4103/1319-3767.187601
Magnani L, Ballantyne EB, Zhang X, Lupien M (2011) PBX1 genomic pioneer function drives ERα signaling underlying progression in breast cancer. PLoS Genet 7(11):e1002368. doi:10.1371/journal.pgen.1002368
Buas MF, Rho JH, Chai X, Zhang Y, Lampe PD, Li C (2015) Candidate early detection protein biomarkers for ER+/PR+ invasive ductal breast carcinoma identified using pre-clinical plasma from the WHI observational study. Breast Cancer Res Treat 153(2):445–454. doi:10.1007/s10549-015-3554-5
Kulak MV, Cyr AR, Woodfield GW, Bogachek M, Spanheimer PM, Li T, Price DH, Domann FE, Weigel RJ (2013) Transcriptional regulation of the GPX1 gene by TFAP2C and aberrant CpG methylation in human breast cancer. Oncogene 32(34):4043–4051. doi:10.1038/onc.2012.400
Powell AA, Talasaz AH, Zhang H, Coram MA, Reddy A, Deng G, Telli ML, Advani RH, Carlson RW, Mollick JA, Sheth S, Kurian AW, Ford JM, Stockdale FE, Quake SR, Pease RF, Mindrinos MN, Bhanot G, Dairkee SH, Davis RW, Jeffrey SS (2012) Single cell profiling of circulating tumor cells: transcriptional heterogeneity and diversity from breast cancer cell lines. PLoS One 7(5):e33788. doi:10.1371/journal.pone.0033788
Miyoshi N, Ishii H, Mimori K, Tanaka F, Hitora T, Tei M, Sekimoto M, Doki Y, Mori M (2010) TGM2 is a novel marker for prognosis and therapeutic target in colorectal cancer. Ann Surg Oncol 17(4):967–972. doi:10.1245/s10434-009-0865-y
Ai L, Kim WJ, Demircan B, Dyer LM, Bray KJ, Skehan RR, Massoll NA, Brown KD (2008) The transglutaminase 2 gene (TGM2), a potential molecular marker for chemotherapeutic drug sensitivity, is epigenetically silenced in breast cancer. Carcinogenesis 29(3):510–518. doi:10.1093/carcin/bgm280
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Investig 121(7):2750–2767. doi:10.1172/JCI45014
McCullough J, Row PE, Lorenzo O, Doherty M, Beynon R, Clague MJ, Urbé S (2006) Activation of the endosome-associated ubiquitin isopeptidase AMSH by STAM, a component of the multivesicular body-sorting machinery. Curr Biol 16(2):160–165. doi:10.1016/j.cub.2005.11.073
Hinoda Y, Okayama N, Takano N, Fujimura K, Suehiro Y, Hamanaka Y, Hazama S, Kitamura Y, Kamatani N, Oka M (2002) Association of functional polymorphisms of matrix metalloproteinase (MMP)-1 and MMP-3 genes with colorectal cancer. Int J Cancer 102(5):526–529. doi:10.1002/ijc.10750
Han J, Bae SY, Oh SJ, Lee J, Lee JH, Lee HC, Lee SK, Kil WH, Kim SW, Nam SJ, Kim S, Lee JE (2014) Zerumbone suppresses IL-1β-induced cell migration and invasion by inhibiting IL-8 and MMP-3 expression in human triple-negative breast cancer cells. Phytother Res 28(11):1654–1660. doi:10.1002/ptr.5178
An CH, Je EM, Yoo NJ, Lee SH (2015) Frameshift mutations of cadherin genes DCHS2, CDH10 and CDH24 genes in gastric and colorectal cancers with high microsatellite instability. Pathol Oncol Res 21(1):181–185. doi:10.1007/s12253-014-9804-8
Yang C, Zhao X, Cui N, Liang Y (2017) Cadherins associate with distinct stem cell-related transcription factors to coordinate the maintenance of stemness in triple-negative breast cancer. Stem Cells Int 2017:5091541. doi:10.1155/2017/5091541
Kresse SH, Ohnstad HO, Paulsen EB, Bjerkehagen B, Szuhai K, Serra M, Schaefer KL, Myklebost O, Meza-Zepeda LA (2009) LSAMP, a novel candidate tumor suppressor gene in human osteosarcomas, identified by array comparative genomic hybridization. Genes Chromosomes Cancer 48(8):679–693. doi:10.1002/gcc.20675
Subramaniam M, Hefferan TE, Tau K, Peus D, Pittelkow M, Jalal S, Riggs BL, Roche P, Spelsberg TC (1998) Tissue, cell type, and breast cancer stage-specific expression of a TGF-beta inducible early transcription factor gene. J Cell Biochem 68(2):226–236
Iida J, Dorchak J, Clancy R, Slavik J, Ellsworth R, Katagiri Y, Pugacheva EN, van Kuppevelt TH, Mural RJ, Cutler ML, Shriver CD (2015) Role for chondroitin sulfate glycosaminoglycan in NEDD9-mediated breast cancer cell growth. Exp Cell Res 330(2):358–370. doi:10.1016/j.yexcr.2014.11.002
Tuna M, Smid M, Zhu D, Martens JW, Amos C (2010) Association between acquired uniparental disomy and homozygous mutations and HER2/ER/PR status in breast cancer. PLoS One 5(11):e15094. doi:10.1371/journal.pone.0015094
Jason BN, Walter CL, Paul AB (2012) Prognosis of treatment response (pathological complete response) in breast cancer. Biomark Insights 7:59–70. doi:10.4137/BMI.S9387
Hata A, Kashima R (2016) Dysregulation of microRNA biogenesis machinery in cancer. Crit Rev Biochem Mol Biol 51(3):121–134. doi:10.3109/10409238.2015.1117054
Bottai G, Diao L, Baggerly KA, Paladini L, Győrffy B, Raschioni C, Pusztai L, Calin GA, Santarpia L (2017) Integrated microRNA-mRNA profiling identifies oncostatin M as a marker of mesenchymal-like ER-negative/HER2-negative breast cancer. Int J Mol Sci 18(1):E194. doi:10.3390/ijms18010194
Guo X, Cai Q, Bao P, Wu J, Wen W, Ye F, Zheng W, Zheng Y, Shu XO (2016) Long-term soy consumption and tumor tissue MicroRNA and gene expression in triple-negative breast cancer. Cancer 122(16):2544–2551. doi:10.1002/cncr.29981
Berillo O, Régnier M, Ivashchenko A (2013) Binding of intronic miRNAs to the mRNAs of host genes encoding intronic miRNAs and proteins that participate in tumourigenesis. Comput Biol Med 43(10):1374–1381. doi:10.1016/j.compbiomed.2013.07.011
Li J, Li X, Kong X, Luo Q, Zhang J, Fang L (2014) MiRNA-26b inhibits cellular proliferation by targeting CDK8 in breast cancer. Int J Clin Exp Med 7(3):558–565
Lu J, He ML, Wang L, Chen Y, Liu X, Dong Q, Chen YC, Peng Y, Yao KT, Kung HF, Li XP (2011) MiR-26a inhibits cell growth and tumorigenesis of nasopharyngeal carcinoma through repression of EZH2. Cancer Res 71(1):225–233. doi:10.1158/0008-5472.CAN-10-1850
Zhao N, Wang R, Zhou L, Zhu Y, Gong J, Zhuang SM (2014) MicroRNA-26b suppresses the NF-κB signaling and enhances the chemosensitivity of hepatocellular carcinoma cells by targeting TAK1 and TAB 3. Mol Cancer 13:35. doi:10.1186/1476-4598-13-35
Martignetti L, Tesson B, Almeida A, Zinovyev A, Tucker GC, Dubois T, Barillot E (2015) Detection of miRNA regulatory effect on triple negative breast cancer transcriptome. BMC Genom 16:S4. doi:10.1186/1471-2164-16-S6-S4
Acknowledgements
We thank Marsh T., very much, the author who deposited their microarray dataset, GSE75333, into the public Gene Expression Omnibus (GEO) database.
Author information
Authors and Affiliations
Contributions
BV carried out the design of this study, performed the statistical analysis and drafted the manuscript. CV collected important background information and software. AKT and SI participated in its design and coordination, visualizations and also helped to draft the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
No informed consent because this study does not contain human or animals participants.
Rights and permissions
About this article
Cite this article
Vastrad, B., Vastrad, C., Tengli, A. et al. Identification of differentially expressed genes regulated by molecular signature in breast cancer-associated fibroblasts by bioinformatics analysis. Arch Gynecol Obstet 297, 161–183 (2018). https://doi.org/10.1007/s00404-017-4562-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00404-017-4562-y