Abstract
Purpose
MiR-361-5p has been reported to act as tumor suppressor in several types of cancers. Retinoblastoma (RB) is the most common ocular tumor in childhood. The current study aimed to investigate the expression pattern and biological function of miR-361-5p in RB.
Methods
Quantitative real time was utilized to determine and compare the expression of miR-361-5p in RB cells and normal retinal pigment epithelial cell line ARPE-19. CCK-8 and Edu assay were performed to assess cell proliferation. Cell apoptosis was evaluated using flow cytometry assay. Bioinformatics databases and luciferase reporter assay were applied to predict and confirm the target gene of miR-361-5p in RB cells.
Results
Here, we found miR-361-5p was significantly downregulated in RB cells compared with normal retinal pigment epithelial cell line ARPE-19. MiR-361-5p overexpression significantly inhibited or silencing promoted cell proliferation in Y79 and SO-RB50 cells, respectively. Flow cytometry assay showed a significantly decreased cell apoptosis in miR-361-5p silencing Y79 cells and increased cell apoptosis in miR-361-5p overexpressing SO-RB50 cells. Moreover, miR-361-5p directly bound to the 3′ untranslated region of claudin 8 (CLDN8) and inhibited the expression of CLDN8. Furthermore, we found knockdown of CLDN8 photocopied the effect of miR-361-5p on cell proliferation and apoptosis in RB cells.
Conclusion
These results indicated that overexpression of miR-361-5p might act as a suppressor in RB by targeting CLDN8 to inhibit the cellular function.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Retinoblastoma (RB) is the most common ocular tumor in childhood arising from the retinas of early childhood [1]. The incidence rate of RB should be around one in 15,000 to 20,000 live births, approximately 9000 new cases are added annually worldwide [2]. Mutations of both the RB-susceptibility gene (RB1) is considered the triggering event in the development of RB [3]. Although intraarterial and intravitreal chemotherapies have a major role in enhancing treatment outcomes, the patients’ survival still remains low than 30% globally [4]. Therefore, it is urgently needed to explore the key molecular mechanisms underlying in the initiation and development of RB to improve its clinical outcomes. The human genome sequence era and the discovery of microRNAs (miRNAs) have brought conceptual breakthrough in the investigation of human cancers [5]. To date, more than 2588 human mature miRNAs (small noncoding RNAs, 19–22 bases in length) have been registered in miRNA database (miRBase) [6]. It has been recognized that large numbers of miRNAs are transcribed in the genome and function as new players in cancer carcinogenesis for their post-transcriptional and translational modifications of their target genes [7, 8]. A growing body of evidence suggests that miRNAs are frequently dysregulated in human cancers and implicated in fundamental processes including proliferation, apoptosis, differentiation, and etc. [9]. MiR-361-5p as an evolutionarily conserved miRNA has been reported to be downregulated in several types of human malignancies, including colorectal cancer [10], gastric cancer [10], prostate cancer [11], and lung cancer [12]. Previous studies have also shown that miR-361-5p regulates tumor cell biological behavior, including cell proliferation, survival, and cell cycle progression by targeting staphylococcal nuclease domain containing-1, signal transducer, activator of transcription-6, FOXM1, and ABCA1 [10,11,12,13]. However, the expression and function of miR-361-5p still remain unclear in the pathogenesis of RB.
In claudins (CLDNs) with 18 to 27 kDa, the critical structural and functional components of tight junctions are recognized by their C-terminal cytoplasmic domain and two extracellular loops [14]. The family of CLDNs consists of at least 24 members, encoding tetraspan membrane proteins, and plays key roles in controlling paracellular ion flux and maintaining cell polarity in epithelial and endothelial cellular sheets [15, 16]. The expression of CLDNs has been extensively reported to exhibit epithelial differentiation in a variety of cancers [16]. Specially, disturbance of tight junction function is an important factor in the development of inflammatory bowel disease, cancer metastasis, celiac disease, and autoimmune disease [17,18,19]. It is noteworthy that claudins are often deregulated in several cancers [20]. For example, claudin 8 (CLDN8) has been found to be reduced in human osteosarcoma [21] and prostate cancer [22]. What is more, depletion of CLDN8 in osteosarcoma U2OS and SW1353 cells results in decreased cell viability and proliferation [21]. In prostate cancer, CLDN8 is regulated by androgen and seems to trigger cancer cell proliferation and migration [22]. Notably, the role of CLDN8 in RB was rarely reported.
In the present study, we determined the involvement of miR-361-5p and CLDN8 in RB. We investigated the expression level of miR-361-5p in RB cell lines and analyzed their biological function on cell proliferation and apoptosis. These preliminary results might help provide “entry point” for therapeutic intervention for RB treatment.
Materials and methods
Cell lines and culture
The human RB cell lines, Y79 and SO-RB50, and a normal retinal pigment epithelial cell line ARPE-19 were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). Y79 and SO-RB50 cells were cultured in RPMI 1640 medium with 10% fetal bovine serum (FBS, Gibco, Grand Island, NY, USA). ARPE-19 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS. All the above cell lines were incubated at 37 °C in a humidified atmosphere containing 5% CO2.
Cell transfection
The oligonucleotides of the miR-361-5p mimics, scrambled negative control miRNA (miR-NC), miR-361-5p inhibitor, and scrambled antago-miRNA (anti-miR) were synthesized from Genepharma Co., Ltd. (Shanghai, China). The small interfering RNA targeting human CLDN8 (siCLDN8) and scrambled siRNA (siNC) were purchased from Guangzhou RiboBio Co., Ltd. For cell transfection, RB cells were seeded into six-well plates at a density of 1.5 × 105 cells per well. MiR-361-5p overexpression in SO-RB50 cells and knockdown in Y79 cells were accomplished by transfecting with miR-361-5p mimics and miR-361-5p inhibitor, respectively, at a concentration of 100 nM. For CLDN8 knockdown, SO-RB50 cells were transfected with 0.6 μg siCLDN8 or siNC. Transfection was performed using Lipofectamine 2000 reagent (Invitrogen, Waltham, MA, USA) according to the manufacturer’s protocol. After 48 h following transfection, cells were harvested for the RNA analysis and protein analysis.
Prediction of miRNA targeting CLDN8
Three Bioinformatics databases, including TargetScan (http://www.targetscan.org/vert_71/), PicTar (http://www.pictar.org/), and miRanda (http://www.microrna.org/microrna/home.do), were used to predict miR-361-5p targeting CLDN8 and their binding regions.
Dual-luciferase reporter assay
For luciferase reporter assay, the wild-type 3′-untranslated regions (UTRs) of CLDN8 mRNA that had a putative miR-361-5p binding site were cloned into the pmiRGLO luciferase vector (Promega, Madison, WI, USA) to generate Wt CLDN8 plasmid. Using Wt CLDN8 3′-UTR as a template, a site-directed mutagenesis kit (Enzynomic, Daejeon, Korea) was used to amplified point mutations in the putative miR-361-5p-binding seed regions to obtain Mut CLDN8 plasmid. Subsequently, 293T cells were co-transfected with miR-361-5p mimics or miR-NC and Wt CLDN8 plasmid or Mut CLDN8 plasmid for 48 h using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instruction. Luciferase activity was determined using the Dual Luciferase Reporter Assay system (Promega) following the manufacturer’s protocols.
qRT-PCR
Total RNA was extracted from cell lines by the TRIzol reagent (Invitrogen) in accordance with the manufacturer’s protocol. For qRT-PCR analysis, cDNA was synthesized from 1 μg of total RNA using the microRNA reverse transcription kit (TaKaRa, Dalian, China) for miR-361-5p and Prime-Script RT reagent kit (TaKaRa) for CLDN8 and then were quantified with SYBR Green PCR Master Mix kit (Applied Biosystems; Thermo Fisher Scientific, Inc.) in triplicate on ABI 7500 HT fast real-time PCR system (Applied Biosystems). The primers used in this study were described as follows: miR-361-5p forward, 5′-AGCCATTGACTAGCCATTCC-3′, and reverse, 5′-GCCTTAGCATGACGCCATGT-3′; U6 forward, 5′-TGTGGGCATCAATGATTTGG-3′ and reverse, 5′-ACACCATGTATTCCGGGTCAAT-3′; CLDN8 forward, 5′-CCGTGATGTCCTTCTTGGCTTTC-3′ and reverse, 5′-CTCTGATGATGGCATTGGCAACC-3′; and GAPDH forward 5′-CCATGTTCGTCATGGTGTG-3′ and reverse, 5′-GGTGCTAAGCAGTTGGTGGTG-3′. U6 and GAPDH were used as internal control to normalize the miR-361-5p and CLDN8 mRNA, respectively. Relative fold-change expression levels were calculated using the 2−ΔΔCT method.
Cell proliferation assay
Cell proliferation was determined using Cell Counting Kit-8 (CCK-8, Beyotime Institute of Biotechnology). Briefly, Y79 and SO-RB50 cells were trypsinized, resuspended, and were seeded into a 96-well plate at a density of 5 × 103 per well. At the indicated culturing time (24, 48, 72 and 96 h), 10 μl CCK-8 solution was added to each well and additionally incubated for 2 h at 37 °C. The absorbance was detected at the wavelength of 450 nm using spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The cell growth curves were plotted by the absorbance of each time point.
EdU incorporation assay
The RB cell proliferation potential was further evaluated using the Cell-Light TM EdU DNA cell kit (RiboBio, Guangzhou, China). Briefly, transfected RB cells were seeded in a 96-well plate at a density of 1 × 105 per well and cultured with 10 mM EdU for 2 h. After fixed with 4% paraformaldehyde, cells were stained with 1 × Apollo567 solution for 30 min in the dark and the cell nuclei were stained with DAPI solution. Finally, stained cells were observed and the percentage of EdU-positive nucleus (red) relatively to blue fluorescent nucleus was calculated under a fluorescent microscope (Olympus Corporation, Tokyo, Japan).
Cell apoptosis assay
Cell apoptosis was determined using Annexin V-FITC Apoptosis Detection kit (KeyGEN, Franklin Lake, NJ, USA) according to the instructions of the manufacturer. In brief, transfected RB cells were harvested, washed with PBS, and resuspended in 500 μl of binding buffer containing 5 μl of Annexin V-fluorescein isothiocyanate (FITC) and 5 μl of propidium iodide (PI). The apoptotic cells, including AnnexinV+/PI- and AnnexinV+/PI+, were measured by a BD Accuri™ C6 flow cytometer (BD Biosciences).
Western blotting
RB cells were harvested and lysed with Cell Lysis buffer (Beyotime Institute of Biotechnology, Shanghai, China), following by quantification a bicinchoninic acid assay kit (Beyotime Institute of Biotechnology). A total of 30 μg protein were subjected to 10% SDS-polyacrylamide gel electrophoresis and transferred to polyvinylidene difluoride (PVDF) membranes (Bio-Rad Laboratories, Inc.). Then, the membranes were incubated with primary antibodies against CLDN8 (1:1000, ab211439, Abcam, Cambridge, MA, USA) and GAPDH (1:5000, ab14247, Abcam) overnight at 4 °C, followed by incubated with horseradish peroxidase (HRP)-conjugated corresponding secondary antibody (1:5000, ab151318, Abcam) for 2 h at room temperature. The protein bands were detected using the enhanced chemiluminescence (ECL) kit (Amersham) and visualized by exposure to X-ray film.
Statistical analysis
Statistical analysis was performed using SPSS 17.0 software (SPSS, Chicago, IL, USA). All data are expressed as the mean ± SD (standard deviation) from at least three independent experiments. Differences were analyzed with two-tailed Student’s t test between the groups or one-way analysis of variance with Dunnett’s post hoc for more than two groups. All differences were considered to be statistically significant when p values of less than 0.05.
Data availability
The data in this study are available from the author for correspondence upon reasonable request.
Results
The expression of miR-361-5p in RB cell lines
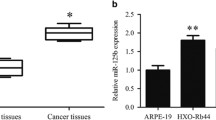
To investigate the biological function of miR-361-5p in RB in vitro, we first compared the expression of miR-361-5p in RB cell lines and a normal retinal pigment epithelial cell line ARPE-19. As a result, miR-361-5p was significantly lower in Y79 (p < 0.01) and SO-RB50 cells (p < 0.001) compared with ARPE-19 cells (Fig. 1a). Since miR-361-5p expression was higher in Y79 cells than SO-RB50 cells, we thus selected Y79 cells for stable transfection with miR-361-5p inhibitor, while SO-RB50 cells for transfection with miR-361-5p mimics. Obviously, miR-361-5p expression was markedly decreased by miR-361-5p inhibitor and significantly increased by miR-361-5p mimic transfection in Y79 cells than SO-RB50 cells, respectively, using qRT-PCR analysis (Fig. 1b, p < 0.001).
MiR-361-5p expression level in RB cells. a MiR-361-5p expression was significantly decreased in the RB cell lines (Y79 and SO-RB50) compared with a normal retinal pigment epithelial cell line (ARPE-19). b The miR-361-5p expression levels in Y79 cells transfected with miR-361-5p inhibitor and SO-RB50 cells transfected with miR-361-5p mimics were determined by qRT-PCR. Data are expressed as means ± SD of three independent experiments. **p < 0.01; ***p < 0.001
MiR-361-5p suppressed RB cell proliferation in vitro
Subsequently, we examined the effect of miR-361-5p expression on cell proliferation in RB cells. Using the CCK-8 assay (Fig. 2a), the growth curve indicated that miR-361-5p inhibitor-transfected Y79 cells exhibited a significant increased proliferation compared with the anti-miR transfection (p < 0.01), while miR-361-5p mimics transfection exerted the opposite results in SO-RB50 cells (p < 0.001). Similarly, EdU incorporation assay (Fig. 2b) revealed that miR-361-5p knockdown markedly promoted cell proliferation in Y79 cells (p < 0.001), but overexpression suppressed cell proliferation in SO-RB50 cells (p < 0.01), as described by the percentage of EdU-positive cells. Taken together, these results indicated that miR-361-5p could exert its inhibitory effect on RB cell proliferation.
Effects of miR-361-5p silencing or overexpression on RB cell proliferation in vitro. The effects of miR-361-5p silencing in Y79 cells or overexpression in SO-RB50 cells on the proliferation ability were measured by a CCK-8 assay and b EdU assay. Data are expressed as means ± SD of three independent experiments. **p < 0.01; ***p < 0.001
MiR-361-5p promoted RB cell apoptosis in vitro
Furthermore, flow cytometry analysis was performed to analyze cell apoptosis in RB cells. As shown in Fig. 3a, the percentage of cells in early apoptosis (8.32% ± 0.08% vs. 6.61% ± 0.02%, p < 0.05) and late apoptosis (12.41% ± 0.08% vs. 7.17% ± 0.11%, p < 0.01) was significantly reduced from anti-miR to miR-361-5p inhibitor group in Y79 cells. On the contrary, miR-361-5p overexpression remarkably elevated the percentage of cells from 10.75 ± 0.58 to 20.17% ± 0.61% in early apoptosis and from 5.90 ± 0.22 to 10.08% ± 0.13% in late apoptosis in SO-RB50 cells (Fig. 3b, p < 0.01, p < 0.001). These results indicated that the enforced overexpression of miR-361-5p decelerated cell apoptosis, which might be associated with impaired cell proliferation in RB cells.
Effects of miR-361-5p silencing or overexpression on RB cell apoptosis in vitro. Flow cytometry analysis was performed to analyze cell apoptosis in a Y79 cells after transfected with miR-361-5p inhibitor and b SO-RB50 cells after transfected with miR-361-5p mimics. Data are expressed as means ± SD of three independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001
CLDN8 is a direct target of miR-361-5p in RB cells
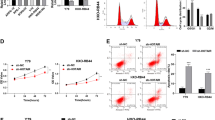
Using algorithm prediction, it was observed that miR-361-5p targets CLDN8 through a conserved domain within the 3′-UTR of CLDN8 (Fig. 4a). Next, a luciferase activity assay was performed to confirm the targeted regulation between miR-361-5p and CLDN8. As shown in Fig. 4b, restoration of miR-361-5p expression markedly decreased the luciferase activity of the Wt CLDN8, but not the Mut CLDN8 in 293T cells (p < 0.01). We further detected whether the mRNA and protein expression levels of CLDN8 in RB cells are regulated by miR-361-5p. As expected, miR-361-5p silencing upregulated, but overexpression downregulated the expression of CLDN8 in Y79 (Fig. 4c, p < 0.001) and SO-RB50 (Fig. 4d, p < 0.001) cells, respectively, at both the mRNA and protein levels. These results revealed that CLDN8 may be a direct target of miR-361-5p in RB cells.
CLDN8 was a direct target of miR-361-5p in RB cells. a The predicted conserved domain within the 3′-UTR of CLDN8 with a potential miR-361-5p binding site was shown. b The luciferase activity of the Wt-CLDN8 3′-UTR and Mut-CLDN8 3′-UTR co-transfected with miR-361-5p mimics or miR-NC was detected in 293T cells. The expression of CLDN8 was detected by qRT-PCR and western blot analysis in c Y79 and d SO-RB50 cells following transfection with miR-361-5p inhibitor and miR-361-5p mimics, respectively. Data are expressed as means ± SD of three independent experiments. **p < 0.01; ***p < 0.001
Knockdown of CLDN8 inhibited cell proliferation and induced apoptosis in RB cells
Since CLDN8 was a potential target gene of miR-361-5p in RB cells, we then speculated it might be a positive regulator for RB cell proliferation. To confirm our hypothesis, SO-RB50 cells were selected for loss-of-function assays using siCLDN8 transfection. The knockdown efficiency of siCLDN8 was evaluated using qRT-PCR and western blotting. The results showed that the expression of CLDN8 was significantly downregulated at mRNA (Fig. 5a, p< 0.001) and protein (Fig. 5b) levels in SO-RB50 cells. Similar to miR-361-5p overexpression, we found CLDN8 knockdown significantly suppressed cell proliferation, as well as promoted cell early and late apoptosis in SO-RB50 cells, as determined by CCK-8 (Fig. 5c, p < 0.001) and flow cytometry analysis (Fig. 5d, p < 0.001).
Effects of CLDN8 knockdown on RB cell proliferation and apoptosis. SO-RB50 cells were transfected with siCLDN8 or siNC, respectively. The expression of CLDN8 was detected using a qRT-PCR and b western blot analysis. c CCK-8 assay was performed to determine cell proliferation. d Flow cytometry analysis was performed to analyze cell apoptosis. Data are expressed as means ± SD of three independent experiments. ***p < 0.001
Discussion
Investigation on the precise molecular mechanisms underlying RB development and progression are pivotal in developing more potent therapeutic strategy for RB patients. In recent decades, miRNAs as a new class of non-coding RNAs has attracted widespread attention for they are responsible for 30–60% protein-coding genes in the human genome [23]. Among these miRNAs, miR-361-5p has been reported to be associated with the progression of cancer [10,11,12,13]. For example, a significant inverse relationship between miR-361-5p and VEGFA levels was observed in human cutaneous squamous cell carcinoma [24]. Subsequently, post-transcriptional regulation by miR-361-5p was involved in the progression of malignancies, including prostate cancer, gastric cancer, and cervical cancer. Similarly, we observed absent/low-expression of miR-361-5p in RB cells. The anti-tumor effect of miR-361-5p validated by both gain and loss-of function assays further supports its role as a tumor suppressor in RB cell proliferation. Moreover, miR-361-5p binds to the 3′-UTR of CLDN8 to direct its posttranscriptional depression. Silencing of CLDN8 could partially imitate the suppressive effects of miR-361-5p overexpression on RB cell proliferation, which suggests that miR-361-5p might inhibit the growth of RB cells through targeting CLDN8.
CLDNs are protein components of many epithelial tight junctions and play an indispensable role for the paracellular barrier in mammalian epithelial cell sheets [25]. Moreover, tight junctions controlling cell differentiation and proliferation through coordinate multiple signaling and trafficking have been studied in recent years [14]. The notable features of CLDNs include two extracellular loops, a N- and C-cytoplasmic domains [25]. Note that tail of the C-terminal cytoplasmic domain has an effect on tight junction barrier properties through regulation of protein stability [26]. Furthermore, C-terminus has a PDZ-binding motif, which binds to PDZ domain proteins such as ZO-1, ZO-2, and ZO-3 [27]. Interestingly, ZO-1 and ZO-2 have been shown to be involved in proliferation, apoptosis, as well gene expression [28,29,30]. Consistently with these evidences, we further found CLDN8 contributes to prostate cancer and osteosarcoma cells proliferation. Mechanically, Ashikari et al. [22] showed that phosphorylation of Akt and ERK1/2 is inactivated by deficiency of CLDN8. Both serine–threonine protein kinase Akt (also known as PKB) and ERK1/2 play key roles in mediating signal transduction processes [31, 32]. In the present study, we found that miR-361-5p negatively regulates the transcriptional and translational levels of CLDN8. Therefore, we supposed that the tight junction barrier function and multiple signaling pathways in RB cells might be disturbed, leading to a significant diminution in cell proliferation and survival, which needed to be further validated in our next experiments.
Conclusions
In conclusion, miR-361-5p emerges as a tumor suppressor in RB through inhibiting CLDN8. Despite the precise mechanism for regulating RB cells proliferation and apoptosis by miR-361-5p/CLDN8 axis remains to be investigated, the tumor suppression of miR-361-5p in RB cells may serve as a basis for the exploration of new potential strategies.
References
Fernandes AG, Pollock BD, Rabito FA (2018) Retinoblastoma in the United States: a 40-year incidence and survival analysis. J Pediatr Ophthalmol Strabismus 55:182–188. https://doi.org/10.3928/01913913-20171116-03
Theriault BL, Dimaras H, Gallie BL, Corson TW (2014) The genomic landscape of retinoblastoma: a review. Clin Exp Ophthalmol 42:33–52. https://doi.org/10.1111/ceo.12132
Li WL, Buckley J, Sanchez-Lara PA, Maglinte DT, Viduetsky L, Tatarinova TV, Aparicio JG, Kim JW, Au M, Ostrow D, Lee TC, O’Gorman M, Judkins A, Cobrinik D, Triche TJ (2016) A rapid and sensitive next-generation sequencing method to detect RB1 mutations improves care for retinoblastoma patients and their families. J Mol Diagn 18:480–493. https://doi.org/10.1016/j.jmoldx.2016.02.006
Dimaras H, Corson TW, Cobrinik D, White A, Zhao J, Munier FL, Abramson DH, Shields CL, Chantada GL, Njuguna F, Gallie BL (2015) Retinoblastoma. Nature Reviews Disease Primers 1:15021. https://doi.org/10.1038/nrdp.2015.21 https://www.nature.com/articles/nrdp201521#supplementary-information
Nishikawa R, Chiyomaru T, Enokida H, Inoguchi S, Ishihara T, Matsushita R, Goto Y, Fukumoto I, Nakagawa M, Seki N (2015) Tumour-suppressive microRNA-29s directly regulate LOXL2 expression and inhibit cancer cell migration and invasion in renal cell carcinoma. FEBS Lett 589:2136–2145. https://doi.org/10.1016/j.febslet.2015.06.005
Desvignes T, Batzel P, Berezikov E, Eilbeck K, Eppig JT, McAndrews MS, Singer A, Postlethwait JH (2015) miRNA nomenclature: a view incorporating genetic origins, biosynthetic pathways, and sequence variants. Trends Genet 31:613–626. https://doi.org/10.1016/j.tig.2015.09.002
Wilczynska A, Bushell M (2015) The complexity of miRNA-mediated repression. Cell Death Differ 22:22–33. https://doi.org/10.1038/cdd.2014.112
Oliveto S, Mancino M, Manfrini N, Biffo S (2017) Role of microRNAs in translation regulation and cancer. World J Biol Chem 8:45–56. https://doi.org/10.4331/wjbc.v8.i1.45
Acunzo M, Romano G, Wernicke D, Croce CM (2015) MicroRNA and cancer--a brief overview. Adv Biol Regul 57:1–9. https://doi.org/10.1016/j.jbior.2014.09.013
Ma F, Song H, Guo B, Zhang Y, Zheng Y, Lin C, Wu Y, Guan G, Sha R, Zhou Q, Wang D, Zhou X, Li J, Qiu X (2015) MiR-361-5p inhibits colorectal and gastric cancer growth and metastasis by targeting staphylococcal nuclease domain containing-1. Oncotarget 6:17404–17416. https://doi.org/10.18632/oncotarget.3744
Liu D, Tao T, Xu B, Chen S, Liu C, Zhang L, Lu K, Huang Y, Jiang L, Zhang X, Huang X, Zhang L, Han C, Chen M (2014) MiR-361-5p acts as a tumor suppressor in prostate cancer by targeting signal transducer and activator of transcription-6(STAT6). Biochem Biophys Res Commun 445:151–156. https://doi.org/10.1016/j.bbrc.2014.01.140
Hou XW, Sun X, Yu Y, Zhao HM, Yang ZJ, Wang X, Cao XC (2017) miR-361-5p suppresses lung cancer cell lines progression by targeting FOXM1. Neoplasma 64:526–534. https://doi.org/10.4149/neo_2017_406
Zhang X, Shao R, Gao W, Sun G, Liu Y, Fa X (2018) Inhibition of miR-361-5p suppressed pulmonary artery smooth muscle cell survival and migration by targeting ABCA1 and inhibiting the JAK2/STAT3 pathway. Exp Cell Res 363:255–261. https://doi.org/10.1016/j.yexcr.2018.01.015
Haseloff RF, Dithmer S, Winkler L, Wolburg H, Blasig IE (2015) Transmembrane proteins of the tight junctions at the blood-brain barrier: structural and functional aspects. Semin Cell Dev Biol 38:16–25. https://doi.org/10.1016/j.semcdb.2014.11.004
Tsukita S, Furuse M, Itoh M (2001) Multifunctional strands in tight junctions. Nat Rev Mol Cell Biol 2:285–293. https://doi.org/10.1038/35067088
Ouban A, Ahmed AA (2010) Claudins in human cancer: a review. Histol Histopathol 25:83–90. https://doi.org/10.14670/HH-25.83
Jauregi-Miguel A, Fernandez-Jimenez N, Irastorza I, Plaza-Izurieta L, Vitoria JC, Bilbao JR (2014) Alteration of tight junction gene expression in celiac disease. J Pediatr Gastroenterol Nutr 58:762–767. https://doi.org/10.1097/MPG.0000000000000338
Lerner A, Matthias T (2015) Changes in intestinal tight junction permeability associated with industrial food additives explain the rising incidence of autoimmune disease. Autoimmun Rev 14:479–489. https://doi.org/10.1016/j.autrev.2015.01.009
Landy J, Ronde E, English N, Clark SK, Hart AL, Knight SC, Ciclitira PJ, Al-Hassi HO (2016) Tight junctions in inflammatory bowel diseases and inflammatory bowel disease associated colorectal cancer. World J Gastroenterol 22:3117–3126. https://doi.org/10.3748/wjg.v22.i11.3117
Singh AB, Dhawan P (2015) Claudins and cancer: fall of the soldiers entrusted to protect the gate and keep the barrier intact. Semin Cell Dev Biol 42:58–65. https://doi.org/10.1016/j.semcdb.2015.05.001
Xu J, Yang Y, Hao P, Ding X (2015) Claudin 8 contributes to malignant proliferation in human osteosarcoma U2OS cells. Cancer Biother Radiopharm 30:400–404. https://doi.org/10.1089/cbr.2015.1815
Ashikari D, Takayama KI, Obinata D, Takahashi S, Inoue S (2017) CLDN8, an androgen-regulated gene, promotes prostate cancer cell proliferation and migration. Cancer Sci 108:1386–1393. https://doi.org/10.1111/cas.13269
Mataki H, Enokida H, Chiyomaru T, Mizuno K, Matsushita R, Goto Y, Nishikawa R, Higashimoto I, Samukawa T, Nakagawa M, Inoue H, Seki N (2015) Downregulation of the microRNA-1/133a cluster enhances cancer cell migration and invasion in lung-squamous cell carcinoma via regulation of Coronin1C. J Hum Genet 60:53–61. https://doi.org/10.1038/jhg.2014.111
Kanitz A, Imig J, Dziunycz PJ, Primorac A, Galgano A, Hofbauer GF, Gerber AP, Detmar M (2012) The expression levels of microRNA-361-5p and its target VEGFA are inversely correlated in human cutaneous squamous cell carcinoma. PLoS One 7:e49568. https://doi.org/10.1371/journal.pone.0049568
Tsukita S, Tanaka H, Tamura A (2019) The claudins: from tight junctions to biological systems. Trends Biochem Sci 2018. https://doi.org/10.1016/j.tibs.2018.09.008
Van Itallie CM, Colegio OR, Anderson JM (2004) The cytoplasmic tails of claudins can influence tight junction barrier properties through effects on protein stability. J Membr Biol 199:29–38
Van Itallie CM, Anderson JM (2014) Architecture of tight junctions and principles of molecular composition. Semin Cell Dev Biol 36:157–165. https://doi.org/10.1016/j.semcdb.2014.08.011
Gonzalez-Mariscal L, Miranda J, Raya-Sandino A, Dominguez-Calderon A, Cuellar-Perez F (2017) ZO-2, a tight junction protein involved in gene expression, proliferation, apoptosis, and cell size regulation. Ann N Y Acad Sci 1397:35–53. https://doi.org/10.1111/nyas.13334
Cordenonsi M (2015) ZO-oming on growth control by junctional proteins. Cell Cycle 14:472. https://doi.org/10.1080/15384101.2015.1006557
Chidiac R, Zhang Y, Tessier S, Faubert D, Delisle C, Gratton JP (2016) Comparative phosphoproteomics analysis of VEGF and angiopoietin-1 signaling reveals ZO-1 as a critical regulator of endothelial cell proliferation. Mol Cell Proteomics 15:1511–1525. https://doi.org/10.1074/mcp.M115.053298
Bai L, Mao R, Wang J, Ding L, Jiang S, Gao C, Kang H, Chen X, Sun X, Xu J (2015) ERK1/2 promoted proliferation and inhibited apoptosis of human cervical cancer cells and regulated the expression of c-Fos and c-Jun proteins. Med Oncol 32:57. https://doi.org/10.1007/s12032-015-0490-5
Lawlor MA, Alessi DR:2001 PKB/Akt. A key mediator of cell proliferation, survival and insulin responses? , 114:2903–2910
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that there is no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, B., Lu, B., Wang, X. et al. MiR-361-5p inhibits cell proliferation and induces cell apoptosis in retinoblastoma by negatively regulating CLDN8. Childs Nerv Syst 35, 1303–1311 (2019). https://doi.org/10.1007/s00381-019-04199-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00381-019-04199-9