Abstract
Competitiveness for nodulation is one of the major restrictive factors in symbiotic nitrogen fixation between rhizobia and their host legumes. Soybean root exudates that include a variety of compounds are thought to act as signals to trigger the early symbiotic events between Bradyrhizobium diazoefficiens and soybeans, and thus they act as a key determinant of the competitiveness for nodulation. To gain a better understanding of the molecular mechanism of competitiveness at the level of protein expression, we compared the proteomic responses of two B. diazoefficiens strains that demonstrated completely different nodulation abilities, strain 4534 being the most competitive and strain 4222 being the least competitive in nodulation. In the proteomic analysis, 40 of the 65 and 22 of the 29 differential proteins were identified in response to soybean root exudates in strain 4534 and strain 4222, respectively. Compared to strain 4222, a higher amount and a number of differential proteins were detected in strain 4534, including S-adenosylmethionine synthetase (SAMS), PhyR-σEcfG regulon, ABC-type transporters, flagellar proteins, molecular chaperones, and proteins involved in redox state maintenance as well as several unknown proteins. Noteworthy was the induction of the PhyR-σEcfG regulon and flagellar proteins, recently demonstrated to be involved in the competitiveness for nodulation in Bradyrhizobium japonicum. Our results indicate that the role of root exudates can go far beyond inducing the expression of nodulation genes in B. diazoefficiens. Many other proteins/enzymes involved in the metabolism and environmental fitness were also upregulated when exposed to root exudates. More proteins were upregulated by the high nodulation competitive strain than that by the low, and the reasons for this need further investigation. The outcome of such study may contribute to our understanding of molecular mechanisms of different competitiveness in B. diazoefficiens as well as specific adaptation in the legume host.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biological nitrogen fixation is a fundamental process in the global N cycles and agricultural production (Xiao et al. 2010). Legume–rhizobium symbiosis accounts for about 60 % of total inputs of biologically fixed N in world agriculture (David et al. 2008). To increase crop production and reduce N fertilizer input, farmers usually treat leguminous seeds with highly efficient rhizobial inoculants (McInnes et al. 2004; Argaw 2012; Juge et al. 2012). Among rhizobia, some strains of Bradyrhizobium japonicum, now renamed Bradyrhizobium diazoefficiens, draw attention because they fix N with soybean, the most important oil and protein crop in the world (Hungria et al. 1998; Delamuta et al. 2013). This symbiosis involves a molecular cross talk of multistep dialogue based on the exchange of diffusible complex signals between the symbionts (Oldroyd et al. 2011; Cesco et al. 2012; Palacios et al. 2014).
Secondary metabolites in the rhizosphere released by plants, serving as nutritional and signaling substances, are capable of positively affecting the microbial colonization of the root and microbial survival in the rhizosphere (Kennedy and Powell 1985; Mark et al. 2005; Palacios et al. 2014). The ability to percept and interpret these signals allows bacteria to sense their environment, leading to the expression of genes, which are important for establishing symbiosis between rhizobia and their host plants, especially for a successful competition for nodule occupancy (Cesco et al. 2012). Despite the advance in the knowledge of molecular cross talks, researchers have long been puzzled as to why the so-called superior strains failed to occupy a significant portion of nodules compared with indigenous rhizobia (McInnes et al. 2004). For this reason, studies on mechanism of competitiveness are critical for the development of commercial inoculants with high-efficient N fixation and to be used to improve crop productivity. Successful nodule formation is determined by factors such as the rhizobial strain’s growth rate in the rhizosphere, communication via signal recognition, and physical attachment (Yates et al. 2011). Specific microbial communities can be selected in the rhizosphere by different plants (Cooper 2004; Mark et al. 2005; Jones et al. 2007; Kidaj et al. 2012).
In response to legume-derived flavonoids, rhizobia synthesize host-specific lipooligosaccharides (LCOs), known as nodulation (Nod) factors, which allow rhizobia to enter the host plant via root hairs (D’Haeze and Holsters 2002; Cesco et al. 2012). The backbone of the Nod factors is produced by enzymes encoded by nodABC, the common nod genes found in most rhizobia studied so far. The specificity of each Nod factor is achieved by the addition of different groups to the backbone, a process facilitated via species-specific nod gene-encoded proteins (Takakazu et al. 2002; Takeshima et al. 2013). In B. japonicum, NodD (a LysR family of prokaryotic transcriptional regulators), NodVW (the two-component regulators), and FrrA (a TetR family transcriptional regulators) positively activate the expression of the nodulation genes in response to host-specific flavonoids (Göttfert et al. 1990, 1992; Loh et al. 1997; Wenzel et al. 2012). So far, more than 4000 types of flavonoids have been found in legumes (Ferrer et al. 2008), and many of them have been studied for their effects on nod gene-inducing activity in different legume–rhizobia interactions (Barbour et al. 1991; Kape et al. 1991, 1992; Lang et al. 2008; Cesco et al. 2012). Daidzein and genistein are the two major isoflavones presented in soybean exudates that induce nod gene expression of B. japonicum (Lang et al. 2008). Recently, it was found that many non-flavonoid components also participate in the induction of nodulation genes in rhizobia such as B. japonicum (Mabood et al. 2006, 2008; Lee et al. 2012). In addition, it has also been reported that flavonoids released as root exudates by lupine roots can inhibit microbial proliferation and mineralization of root exudates (Tomasi et al. 2008). Therefore, the regulation of nod gene is complex. Despite the widely accepted fact that extracellular signals produced by plants can influence the performance of bacteria in the rhizosphere, the effects of naturally occurred soybean root exudates on the interaction of legumes with rhizobia, especially regarding the physiology and nodulation competitiveness of rhizobia, are poorly known.
The fine-tuning of transcriptomics and mass spectrometry technology has greatly facilitated the investigation of the transcriptome-level changes of B. diazoefficiens global gene expression in response to a given compound (Donati et al. 2013). However, due to the presence of posttranslational regulation, quantitative mRNA data are insufficient to predict the function of proteins in the cells (Kawamura and Uemura 2003). Proteomics is the study of the protein expression profiles of organisms and can be used for monitoring the microbial responses to external stimuli. Proteomic profiles of rhizobia during the establishment of the symbiotic relation with leguminous plants have been reported (Nomura et al. 2010; Da Silva Batista and Hungria 2012). However, the proteomic responses of rhizobial strains with different nodulation efficiencies to root exudates as well as the role of specific compounds of root exudates in the initiation of the symbiotic relation between rhizobia and leguminous plants are still poorly understood (Giagnoni et al. 2011).
In this study, we investigated and compared the effects of soybean root exudates on the protein expression of two B. diazoefficiens strains, which are characterized by different nodulation competitiveness: one occupies 90 % of nodules and the other only 10 % when an equal amount of cells was applied to soybean roots (Xiao et al. 2010). Thus it is interesting to know the differences in the proteomic level of these two strains when they are treated by root exudates. The optimal conditions for nod genes induction by soybean root exudates were determined by quantitative reverse transcription PCR (qRT-PCR). Then we analyzed the total proteins produced by the two strains after exposure to soybean root exudates. The results of the study can contribute to a better understanding of molecular mechanisms of competitiveness in B. diazoefficiens as well as the specific adaptation to legume host.
Materials and methods
Bacterial strains, medium, and culture conditions
The bacterial strains used in this work were B. diazoefficiens 4534 and B. diazoefficiens 4222. The soybean cultivar was Zhonghuang 13, which is widely cultivated in Huang-Huai-Hai region on the North China Plain. All rhizobial strains were cultured in TY broth (Beringer 1974) shaking at 180 rpm and 30 °C. Bacterial growth was evaluated by measuring its optical absorbance at 600 nm.
Preparation of root exudates
Root exudates were collected from the soybean variety Zhonghuang 13. Soybean seeds were surface disinfected and transferred into a nitrogen-free medium (Albareda et al. 2006). Twenty-five pre-germination seeds were aseptically transferred into a polypropylene lattice placed in a glass cylinder containing 300 mL of sterilized modified N-free solution as reported by Rigaud and Puppo (1975). No seeds were sown in the control. Microcosms were arranged in a replicate-randomized block design and incubated at 30 °C for 16 h and 15 °C for 8 h. The root exudates and control eluant (liquid from blank control) were tested on TY medium to monitor the microbial contamination, no growth indicating no microbial contamination present. Soybean root exudates (SREs) were sterilized by filtration through a 0.22-μm filter (Millipore Company, USA) and stored at −20 °C until use (Mark et al. 2005). Each treatment was replicated three times.
Assays for induction of nod genes
Before proteomic analysis, the influence of root exudates on nodulation of B. diazoefficiens was evaluated. Expression levels of nod genes were chosen as criteria on judging the effective induction of root exudates. Though it is generally accepted that root exudates have profound influence on bacterial physiology, the gene expression pattern in response to root exudates is seldom studied. The optimal conditions of incubation should be established for the highest upregulation of nod genes. At the start of this study, three parameters were compared: (i) concentrations of nutrient solution, i.e., soybean plants were grown in different concentrations of nutrient solution; (ii) time intervals of root exudate collection, i.e., root exudates were collected from 0, 6, 12, and 16-day-old soybean plants; and (iii) optimal period exposure of rhizobia to root exudates for maximal expression of nod genes, i.e., the bacterial cells were harvested after exposure to root exudates for 0.25, 1, 3, and 6 h. We measured the expression of nodD1, nodD2, and nodC genes. Both B. diazoefficiens strains 4534 and 4222 were grown up to the exponential phase and then incubated with 200 mL root exudates.
RT-PCR and quantitative real-time RT-PCR
Total RNA was extracted using the SV Total RNA Isolation System (Promega, Madison, WI, USA) purified with RNase-free DNase I (Promega). The first strand of cDNA was synthesized with 1.0 μg of RNA using the ProtoScript First-Strand cDNA Synthesis Kit (New England Biolabs, Ipswich, MA, USA).
Specific primers for nodC, nodD1, and nodD2 genes to be used in the quantitative real-time PCR (Q-PCR) were designed using Primer 5 Server based on the published genomic sequence of B. diazoefficiens USDA110 targeting an amplicon size of 150–200 bp. The reaction specificity was assayed by agarose gel electrophoresis and product dissociation curves. The primers used are listed in Table 1. Quantitative RT-PCR experiments following the manufacturer’s instruction were performed as described by Yan et al. (2008). The equipment used included the 7500 Sequence Detection system (Applied Biosystems) and the SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA).
Whole-cell protein extraction and two-dimensional electrophoresis
Bacterial cells were harvested by centrifugation at 8000×g for 5 min at 4 °C and resuspended in the mixture of 1.5 mL W1 buffer, 50 μL DTT, and 1.5 mL W2 buffer (Wang et al. 2008). Bacterial cells were lysed by 3 cycles of sonication at 35 kHz for 30 s on ice and incubated on ice for 30 s. Clear lysates obtained after centrifugation at 12,000×g for 10 min were subjected to protein precipitation by adding 1.5 mL 50 % TCA solution to 6 mL of clear lysates; then the mixture was incubated at 4 °C and then centrifuged at 12,000×g for 10 min at 4 °C; the precipitate was washed with W3 buffer for three times, dried in a vacuum dryer, and stored in an ultrafreezer (−80 °C) until analysis, and total protein concentration was determined as reported by Bradford (1976).
Immobilized pH gradient (IPG) strips (11 cm, pH 4–7; GE Healthcare Bio-Sciences) were used for isoelectric focusing (IEF). The IPG gels were rehydrated with the protein solution (about 600 μg) and covered with cover fluid (GE Healthcare) for 12 h. Sodium dodecyl sulfate (SDS)-PAGE was carried out with the following gradient: 4 and 15 % polyacrylamide gel; it was run with 0.3 % Tris, 1.44 % glycine, and 0.1 % SDS buffer. The electrophoresis was performed as described previously with minor modifications (Liu et al. 2013). The preparative gels were stained with Coomassie brilliant blue (CBB) R-250. At least three independent replicates were performed for each sample.
Gel image analysis
The two-dimensional electrophoresis (2-DE) profiles of different samples were acquired by scanning the 2-DE gels with Image Scanner III (GE Healthcare Bio-Sciences) and analyzed by ImageMatser 2D-platinum v5.0 software (GE Healthcare Bio-Sciences). The spot volumes were normalized as the proportion of the sum of total spots per gel. All selected spots were automatically detected and matched before the manual confirmation. Three well-separated gels of biological replicates were used to create “replicate groups.” After image analysis, normalized spot volumes were obtained for each gel and statistical differences were calculated between the control group and each treated group by the Student’s t test considering a significance level of 95 %. The spot were selected as differential proteins if their volume showed more than 1.5-fold of differences and a statistically significant difference (p < 0.05) between exudate-induced and non-induced treatments.
MALDI-TOF/TOF-TOF analysis and protein identification
MS and MS/MS spectra were acquired using the ultrafleXtreme™ MALDI-TOF/TOF mass spectrometer (Bruker Daltonics, Bremen, Germany) and ABI 4800 Proteomics Analyzer MALDI-TOF/TOF (Applied Biosystems, Foster City, CA, USA) and analyzed by the software GPS Explorer, 3.6 software (Applied Biosystems) and MASCOT 2.1 software with the parameter settings as described for ultrafleXtreme™. The instrument was operated in the positive reflection mode and externally calibrated using the peptide calibration kit (Bruker Daltonics). The mass accuracy and mass resolution were set as the default. The samples (1 μL) were spotted onto the AnchorChip™ MALDI target plate (Bruker Daltonics, Billerica, MA, USA) and allowed to dry at room temperature. One milliliter of matrix solution (1 mg/mL, a-cyano-4-hydroxycinnamic acid in 70 % acetonitrile containing 0.1 % trifluoroacetic acid) was then manually applied onto samples to allow cocrystallization.
MS and MS/MS data analysis was performed using the BioTools software (V3.2, Bruker Daltonics) and GPS Explorer Software (Applied Biosystems), which uploads the peptide mass fingerprinting and MS/MS ion to Mascot for database searching on the Matrix Science (London, UK) public web site (http://www.matrixscience.com) and searched against NCBInr protein databases. The parameters for searching were set as follows: an enzyme of trypsin; one missed cleavage; a fragment tolerance of ±0.5 Da; a peptide mass tolerance of 100 ppm; carbamidomethylation (Cys) set as fixed modification; and oxidation of methionines (Met) set as variable modification. Only significant hits, as defined by the MASCOT probability analysis, were accepted (Liu et al. 2013).
Results
Transcriptional analysis of two strains after exposure to root exudates
Transcriptional analysis indicated that the nod genes of both strains were upregulated to varying degrees in response to root exudates, and the patterns of both strains were quite similar. However, under the same conditions, the upregulation of nod genes was greater in B. diazoefficiens 4534 than in B. diazoefficiens 4222. By considering these results, the optimal conditions for nod gene expression were determined: the root exudates were collected from soybean grown in one-third strength of modified N-free Rigaud–Puppo solution for 7 days and a 1-h incubation period for gene induction; this gave the highest upregulation of nodC, nodD1, and nodD2 genes.
Proteomic analysis of two strains treated with soybean root exudates
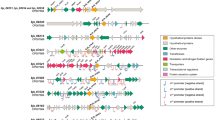
Sixty-five differential proteins were detected in B. diazoefficiens 4534 (Fig. 1(b)) in response to root exudate induction. Only 40 of the 65 differentially expressed proteins were excised for tryptic digestion and identified by MALDI-TOF (Table 2), because the remaining ones showed a low abundance. Although 29 differential protein spots were detected, only 22 differential protein spots were selected in B. diazoefficiens 4222 in response to the root exudates induction, as shown by comparing Fig. 1(c) with Fig. 1(d) (Table 3).
Proteins responsible for changes in the proteome profile of the two B. diazoefficiens strains, identified in this study, fell in the following six functional categories with numbers of the different proteins associated with different functions in strain 4534 and strain 4222: (i) signal transduction (2, 0); (ii) transport and substrate binding (4, 1); (iii) chemotaxis and motility (1, 0); (iv) metabolic fitness and energy metabolism (12, 9); (v) transcription, translation, and protein folding (5, 2); and (vi) unclassified conserved and hypothetical proteins (16, 11).
Discussion
Proteins related to signal transduction
We observed that the S-adenosylmethionine (SAM) synthetase was increased in B. diazoefficiens 4534 by incubation with root exudates, which is consistent with what was previously reported for the proteome of B. japonicum induced by genistein (Da Silva Batista and Hungria 2012). SAM is the methyl donor for the synthesis of N-acyl-homoserine lactones (acyl-HSLs), which positively activates the expression of quorum sensing (QS)-dependent genes. Previous studies confirmed that QS was involved in several processes in rhizobia, including biofilm formation, exopolysaccharide synthesis, swimming motility, nodulation efficiencies, and nitrogen fixation efficiencies (Sanchez-Contreras et al. 2007; Mueller and González 2011). Biofilm formation of rhizobia is important for nodule infection (Williams et al. 2008) and is influenced by environmental conditions as well as legume-produced compounds such as lectins (Perez-Giménez et al. 2009) and flavonoids (Fujishige et al. 2008). In addition, EPS biosynthesis is also regulated by QS and is important for rhizobium infection, attachment, and competitiveness (Davies and Walker 2007; Edwards et al. 2009).
Another important protein for the establishment of the symbiosis identified in this study was the two-component regulator PhyR, a special type of response regulator consisting of a receiver domain and an extracytoplasmic function sigma factor-like domain; the latter linked in the signaling cascade following the interaction with the anti-sigma factor PhyR-σEcfG regulon, was involved in the Bradyrhizobium–legume interaction, and was a general stress response whose mechanism is still unknown (Gourion et al. 2008). The PhyR-σEcfG-lacking mutants of B. japonicum were deficient in nodulation (Cytryn et al. 2007; Gourion et al. 2009).
We observed a clear difference in the upregulation of the signal transduction proteins between the high and the low competitiveness strains: two differential proteins (e.g., SAM synthetase and PhyR) associated with signal transduction were only detected in B. diazoefficiens 4534, whereas none was detected in B. diazoefficiens 4222.
Our results suggest that the root exudate-induced signal transduction plays an important role in nodulation competitiveness of B. diazoefficiens, which is important for the nodulation efficiency (Gourion et al. 2009).
Proteins related to transport and substrate binding
ABC-type transporter substrate-binding protein (ATP-binding cassette), one of the largest superfamilies of membrane transport proteins, was differentially expressed in the presence of soybean root exudates. ABC-type transporters can participate in nutrient uptake, polysaccharide secretion, signal transduction, and drug resistance through energy produced by ATP hydrolysis (Nicolás et al. 2007). It is clear that efficient transport systems are an essential requisite for nutrient competition, which leads to the difference in competitiveness (Sarma and Emerich 2006).
An abundance of ABC-type transporters was confirmed in B. japonicum, and the relative gene expression of proteins was induced by genistein (Da Silva Batista and Hungria 2012). Here, four different proteins related to transport and substrate-binding were upregulated in B. diazoefficiens 4534, but only one transporter protein was upregulated in B. diazoefficiens 4222. Therefore, root exudates led to the increased expression of ABC-type transporters and that expression differed in the two strains.
Proteins related to chemotaxis and motility
Here, flagellar proteins were significantly induced by root exudates. Recent evidence suggests that flagellum biosynthesis is critical for bacterial adaptation to numerous environmental conditions and enhanced competitiveness for nodule occupancy under laboratory conditions (Mongiardini et al. 2009; Covelli et al. 2013). Microarray experiments indicated that all genes within the flagellar cluster are slightly upregulated in response to genistein (Lang et al. 2008), and highly abundant flagellar proteins were detected in the secretome analysis of B. japonicum (Süß et al. 2006; Hempel et al. 2009). The mutants of B. japonicum with increased motility had the advantage when competing with indigenous strains for nodulation (López-García et al. 2002, 2009; Bogino et al. 2008).
Here, the significantly upregulated proteins related to chemotaxis and motility only occurred in B. diazoefficiens 4534 (Table 2). Since B. diazoefficiens 4534 is highly efficient in nodulation, the upregulation of chemotaxis and motility-related proteins by root exudates indicates that chemotaxis and motility are probably important for nodulation competitiveness. In addition, compared with the results by Lang et al. (2008), root exudates are more efficient in inducing the upregulation of chemotaxis and motility-related proteins than genistein (Lang et al. 2008).
Proteins related to metabolic fitness and energy metabolism
In rhizosphere, many substrates coexist at relatively low concentrations; therefore, the ability to efficiently utilize different nutrients by a strain should be an ecological advantage (Prell and Poole 2006). In this experiment, an abundance of proteins related to metabolic fitness was induced in both strains when incubated with root exudates. Proteins involved in purine metabolism were only found upregulated in B. diazoefficiens 4534, which confirmed previous studies. The purine pathway in Rhizobium is important during the nodulation processes, since it is significantly and positively correlated with competitive nodulation abilities of the bacteria (Xie et al. 2009).
Proteins associated with C and N cycling were generally more upregulated in B. diazoefficiens 4534 than in B. diazoefficiens 4222 by root exudates. Phosphopyruvate hydratase is one enzyme of the glycolysis pathway, which is a catabolic pathway. Acetyl-CoA acetyltransferase, an enzyme involved in generating poly-β-hydroxybutyrate (PHB), was also found to be upregulated. The ability to build up PHB is a key factor in the survival of bacteria, which can use the accumulated PHB under nutrient-limited conditions (Aneja et al. 2005; Marroquí et al. 2001).
Furthermore, intermediate metabolites and energy are indispensable for the proliferation of rhizobia during rhizobium–legume symbiotic nodulation (Li et al. 2011). Our proteome analysis showed that under the induction by soybean root exudates, more proteins related to metabolic fitness were found upregulated in B. diazoefficiens 4534 than in B. diazoefficiens 4222, suggesting that these metabolic properties may also be essential traits in determining the competitiveness of rhizobia.
Proteins related to transcription, translation, and protein folding
The molecular chaperone GroEL in B. diazoefficiens 4534 was significantly induced by root exudates. Previous studies indicated that GroEL chaperones are essential for the formation of a functional nitrogenase during the initial bacteroid developmental stage, and the expression of the relative gene seems to be regulated during the symbiotic development through a stringent regulatory circuit (Sarma and Emerich 2005). Molecular chaperones are a ubiquitous family of abundant proteins in cellular regulation (Wang et al. 2004) and play a pivotal role in preventing both new synthesis of polypeptide chains and assembly of subunits, preventing newly synthesized polypeptide chains from folding into nonfunctional protein; in addition, they facilitate protein refolding (Yan et al. 2006).
Root exudates also upregulated the trigger factor (TF) that is the only ribosome-associated chaperone known in bacteria. This protein ensures efficient protein folding and translocation of newly synthesized polypeptide chains (Hoffmann et al. 2010). This observation is in agreement with the downregulation of chaperone SecB in B. diazoefficiens 4534 because its access to nascent chains is normally restrained by TF (Ullers et al. 2004). Our data support the conclusion that protein folding is secured by the action of several chaperones which, despite their different mechanisms of action, are able to substitute each other, thus endowing B. diazoefficiens with maximal flexibility in response to varying environmental conditions (Hoffmann et al. 2010; Nomura et al. 2010).
Five proteins associated with transcription and translation were detected in B. japonicum 4534, but only two were found in B. diazoefficiens 4222. These data might reflect the activity of cellular metabolism during critical stages of symbiotic infection with alterations in the proteome pattern.
Conclusions
A comparison of proteomic analyses of the two strains induced by soybean root exudates reveals that proteins related to transport and substrate binding, metabolic fitness and energy metabolism, transcription, translation, and protein folding in B. diazoefficiens 4534 were more abundant than those in B. diazoefficiens 4222. The expression of proteins involved in signal transduction, chemotaxis, and motility was only detected in strain 4534, not in strain 4222. This may be due to variations in the response of individual B. diazoefficiens genotypes to soybean root exudates. Although the different regulation mechanisms of different metabolic processes are not fully characterized, it may act in the early establishment of rhizobium–legume symbiosis, demonstrating greater complexity of responses to soybean root exudates. The striking difference between high nodulation competitive strain 4534 and low competitive strain 4222 is that more proteins were upregulated when exposed to soybean root exudates. A possible explanation for this phenomenon is that high competitive strain might be more sensitive to the induction of special compounds in the root exudates.
We are unaware that other plant signals may change the protein expression of B. diazoefficiens during competition for nodulation. Flavonoids (genistein, coumestrol, daidzein, etc.) and non-flavonoids (jasmonates) can only partially substitute soybean root exudates (Mabood et al. 2006; Lee et al. 2012). Our data not only improve our understanding of the genetic and functional responses of B. diazoefficiens in competitive nodulation but also support further studies on functions of the 20 hypothetical proteins only induced by root exudates, not by pure flavonoids.
References
Albareda M, Dardanell MS, Sousa C, Megías M, Temprano F, Rodríguez-Navarro DN (2006) Factors affecting the attachment of rhizospheric bacteria to bean and soybean roots. FEMS Microbiol Lett 259:67–73
Aneja P, Zachertowska A, Charles TC (2005) Comparison of the symbiotic and competition phenotypes of Sinorhizobium meliloti PHB synthesis and degradation pathway mutants. Can J Microbiol 51:599–604
Argaw A (2012) Evaluation of co-inoculation of Bradyrhizobium japonicum and phosphate solubilizing Pseudomonas spp. effect on soybean (Glycine max L. (Merr.)) in Assossa area. J Agric Sci Tech-Iran 14:213–224
Barbour WM, Hattermann DR, Stacey G (1991) Chemotaxis of Bradyrhizobium japonicum to soybean exudates. Appl Environ Microbiol 57:2635–2639
Beringer JE (1974) R factor transfer in Rhizobium leguminosarum. J Gen Microbiol 120:421–429
Bogino P, Banchio E, Bonfiglio C, Giordano W (2008) Competitiveness of a Bradyrhizobium sp. strain in soils containing indigenous rhizobia. Curr Microbiol 56:66–72
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein dye binding. Anal Biochem 72:248–254
Cesco S, Mimmo T, Tonon G, Tomasi R, Pinton R, Terzano R, Neumann G, Weisskopf L, Renella G, Landi L, Nannipieri P (2012) Plant-borne flavonoids released into the rhizosphere: impact on soil bio-activities related to plant nutrition. A review. Biol Fertil Soils 48:123–150
Cooper JE (2004) Multiple responses of rhizobia to flavonoids during legume root infection. Adv Bot Res 41:1–62
Covelli JM, Althabegoiti MJ, López MF, Lodeiro AR (2013) Swarming motility in Bradyrhizobium japonicum. Res Microbiol 164:136–144
Cytryn EJ, Sangurdekar DP, Streeter JG, Franck WL, Chang WS, Gar S, Emerich DW, Joshi T, Xu D, Sadowsky MJ (2007) Transcriptional and physiological responses of Bradyrhizobium japonicum to desiccation-induced stress. J Bacteriol 189:6751–6762
D’Haeze W, Holsters M (2002) Nod factor structures, responses, and perception during initiation of nodule development. Glycobiology 12:79–105
Da Silva Batista JS, Hungria M (2012) Proteomics reveals differential expression of proteins related to a variety of metabolic pathways by genistein-induced Bradyrhizobium japonicum strains. J Proteomics 75:1211–1219
David F, Peoples MB, Boddey RM (2008) Global inputs of biological nitrogen fixation in agricultural systems. Plant Soil 311:1–18
Davies BW, Walker GC (2007) Identification of novel Sinorhizobium meliloti mutants compromised for oxidative stress protection and symbiosis. J Bacteriol 189:2110–2113
Delamuta JR, Ribeiro RA, Ormeño-Orrillo E, Melo IS, Martínez-Romero E, Hungria M (2013) Polyphasic evidence supporting the reclassification of Bradyrhizobium japonicum group Ia strains as Bradyrhizobium diazoefficiens sp. Nov Evol Microbiol 63:3342–3351
Donati AJ, Lee HI, Leveau JH, Chang WS (2013) Effects of indole-3-acetic acid on the transcriptional activities and stress tolerance of Bradyrhizobium japonicum. PLoS One 8:76559
Edwards A, Frederix M, Wisniewski F, Jones J, Zorreguieta A, Downie JA (2009) The cin and rai quorum-sensing regulatory systems in Rhizobium leguminosarum are coordinated by ExpR and CinS, a small regulatory protein coexpressed with CinI. J Bacteriol 191:3059–3067
Ferrer JL, Austin MB, Stewart CJ, Noel JP (2008) Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol Biochem 46:356–370
Fujishige NA, Lum MR, De Hoff PL, Whitelegge JP, Faull KF, Hirsch AM (2008) Rhizobium common nod genes are required for biofilm formation. Mol Microbiol 67:504–515
Giagnoni L, Magherini F, Landi L, Taghavi S, Modesti A, Bini L, Nannipieri P, Vanderlelie D, Renella G (2011) Extraction of microbial proteome from soil: potential and limitations assessed through a model study. Eur J Soil Sci 62:74–81
Göttfert MP, Grob H, Hennecke (1990) Proposed regulatory pathway encoded by the nodV and nodW genes, determinants of host specificity in Bradyrhizobium japonicum. Proc Natl Acad Sci U S A 87:2680–2684
Göttfert MP, Holzhäuser D, Bäni D, Hennecke H (1992) Structural and functional analysis of two different nodD genes in Bradyrhizobium japonicum USDA110. Mol Plant Microbe Interact 5:257–265
Gourion B, Francez- Charlot A, Vorholt JA (2008) PhyR is involved in the general stress response of Methylobacterium extorquens AM1. J Bacteriol 190:1027–1035
Gourion B, Sulser S, Frunzke J, Francez-Charlot A, Stiefel P, Pessi G, Vorholt JA, Fischer HM (2009) The PhyR-σEcfG signalling cascade is involved in stress response and symbiotic efficiency in Bradyrhizobium japonicum. Mol Microbiol 73:291–305
Hempel J, Zehner S, Göttfert M, Patschkowski T (2009) Analysis of the secretome of the soybean symbiont Bradyrhizobium japonicum. J Bacteriol 140:51–58
Hoffmann A, Bukau B, Kramer G (2010) Structure and function of the molecular chaperone trigger factor. Biochim Biophys Acta 1803:650–661
Hungria M, Boddey LH, Santos MA, Vargas MA (1998) Nitrogen fixation capacity and nodule occupancy by Bradyrhizobium japonicum and B. elkanii strains. Biol Fertil Soils 27:393–399
Jones KM, Kobayashi H, Davies BW, Taga ME, Walker GC (2007) How symbionts invade plants: the Sinorhizobium-Medicago model. Nat Rev Microbiol 5:619–633
Juge C, Prévost D, Bertrand A, Bipfubusa M, Chalifour FP (2012) Growth and biochemical responses of soybean to double and triple microbial associations with Bradyrhizobium, Azospirillum and arbuscular mycorrhizae. Appl Soil Ecol 61:147–157
Kape R, Parniske M, Werner D (1991) Chemotaxis and nod gene activity of Bradyrhizobium japonicum in response to hydroxycinnamic acids and isoflavonoids. Appl Environ Microbiol 57:316–319
Kape R, Parniske M, Brandt S, Werner D (1992) Isoliquiritigenin, a strong nod gene- and glyceollin resistance-inducing flavonoid from soy-bean root exudate. Appl Environ Microbiol 58:1705–1710
Kawamura Y, Uemura M (2003) Mass spectrometric approach for identifying putative plasma membrane proteins of Arabidopsis leaves associated with cold acclimation. Plant J 36:141–154
Kennedy JA, Powell HKJ (1985) Polyphenol interactions with aluminum(III) and iron(III)—their possible involvement in podzolization process. Aust J Chem 38:879–888
Kidaj D, Wielbo J, Skorupska A (2012) Nod factors stimulate seed germination and promote growth and nodulation of pea and vetch under competitive conditions. Microbiol Res 167:44–50
Lang K, Lindemann A, Hauser F, Göttfert M (2008) The genistein stimulon of Bradyrhizobium japonicum. Mol Genet Genomics 279:203–211
Lee HI, Lee JH, Park H, Sangurdekar D, Chang WS (2012) Effect of soybean coumestrol on Bradyrhizobium japonicum nodulation ability, biofilm formation, and transcriptional profile. Appl Environ Microbiol 78:2896–2903
Li J, Xiao WL, Ma MC, Guan DW, Jiang X, Cao FM, Shen DL, Chen HJ, Li L (2011) Proteomic study on two Bradyrhizobium japonicum strains with different competitivenesses for nodulation. Agric Sci China 10:10721079
Liu H, Wang C, Komatsu S, He M, Liu G, Shen S (2013) Proteomic analysis of the seed development in Jatropha curcas: from carbon flux to the lipid accumulation. J Proteomics 91:23–40
Loh J, Garcia M, Stacey G (1997) NodV and NodW, a second flavonoid recognition system regulating nod gene expression in Bradyrhizobium japonicum. J Bacteriol 179:3013–3020
López-García SL, Vázquez TE, Favelukes G, Lodeiro AR (2002) Rhizobial position as a main determinant in the problem of competition for nodulation in soybean. Environ Microbiol 4:216–224
López-García SL, Perticari A, Piccinetti C (2009) In-furrow inoculation and selection for higher motility enhances the efficacy of Bradyrhizobium japonicum nodulation. Agric J 101:357–363
Mabood F, Souleimanov A, Khan W, Smith DL (2006) Jasmonates induce Nod factor production by Bradyrhizobium japonicum. Plant Physiol Biochem 44:759–765
Mabood F, Jung WJ, Smith DL (2008) Signals in the underground: microbial signaling and plant productivity. In: Nautiyal CS, Dion PE, Chopra VL (eds) Molecular mechanisms of plant and microbe coexistence. Springer, Berlin, pp 291–318
Mark GL, Dow JM, Kiely PD, Higgins H, Haynes J, Baysse C, Abbas A, Foley T, Franks A, Morrissey J, O’Gara F (2005) Transcriptome profiling of bacterial responses to root exudates identifies genes involved in microbe-plant interactions. Proc Natl Acad Sci U S A 102:17454–17459
Marroquí S, Zorreguieta A, Santamaría C, Temprano F, Soberón M, Megías M, Downie JA (2001) Enhanced symbiotic performance by Rhizobium tropici glycogen synthase mutants. J Bacteriol 183:854–864
McInnes A, Thies JE, Abbott LK, Howieson JG (2004) Structure and diversity among rhizobial strains, populations and communities—a review. Soil Biol Biochem 36:1295–1308
Mongiardini EJ, Pérez-Gimenez J, Althabegoiti MJ, Covelli J, Ignacio Quelas J, López-García SL, Lodeiro AR (2009) Overproduction of the rhizobial adhesin RapA1 increases competitiveness for nodulation. Soil Biol Biochem 41:2017–2020
Mueller K, González JE (2011) Complex regulation of symbiotic functions is coordinated by MucR and quorum sensing in Sinorhizobium meliloti. J Bacteriol 193:485–496
Nicolás MF, Barcellos FG, Hess PN, Hungria M (2007) ABC transporters in Mycoplasma hyopneumoniae and Mycoplasma synoviae: insights into evolution and pathogenicity. Genet Mol Biol 30:2002–2011
Nomura M, Arunothayanan H, VanDao T (2010) Differential protein profiles of Bradyrhizobium japonicum USDA110 bacteroid during soybean nodule development. Soil Sci Plant Nutr 56:579–590
Oldroyd GE, Murray JD, Poole PS, Downie JA (2011) The rules of engagement in the legume-rhizobial symbiosis. Annu Rev Genet 45:119–144
Palacios OA, Bashan Y, de-Bashan LE (2014) Proven and potential involvement of vitamins in interactions of plants with plant growth-promoting bacteria—an overview. Biol Fertil Soils 50:415–432
Perez-Giménez J, Mongiardini EJ, Althabegoiti MJ, Covelli J, Quelas JI, López-García SL, Lodeiro AR (2009) Soybean lectin enhances biofilm formation by Bradyrhizobium japonicum in the absence of plants. Int J Microbiol 1. doi: 10.1155/2009/719367
Prell J, Poole P (2006) Metabolic changes of rhizobia in legume nodules. Trends Microbiol 14:161–168
Rigaud J, Puppo A (1975) Indole-3-acetic acid catabolism by soybean bacteroids. J Gen Microbiol 88:223–228
Sanchez-Contreras M, Bauer WD, Gao M, Robinson JB, Allan Downie J (2007) Quorum-sensing regulation in rhizobia and its role in symbiotic interactions with legumes. Philos Trans R Soc Lond B Biol Sci 362:1149–1163
Sarma AD, Emerich DW (2005) Global protein expression pattern of Bradyrhizobium japonicum bacteroids: a prelude to functional proteomics. Proteomics 5:4170–4784
Sarma AD, Emerich DW (2006) A comparative proteomic evaluation of culture grown vs nodule isolated Bradyrhizobium japonicum. Proteomics 6:3008–3028
Süß C, Hempel J, Zehner S, Krause A, Patschkowski T, Göttfert M (2006) Identification of genistein-inducible and type III-secreted proteins of Bradyrhizobium japonicum. J Biotechnol 126:69–77
Takakazu K, Yasukazu N, Shusei S (2002) Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res 9:189–197
Takeshima K, Hidaka T, Wei M, Yokoyama T, Minamisawa K, Mitsui H, Itakura M, Kaneko T, Tabata S, Saeki K, Oomori H, Tajima S, Uchiumi T, Abe M, Tokuji Y, Ohwada T (2013) Involvement of a novel genistein-inducible multidrug efflux pump of Bradyrhizobium japonicum early in the Interaction with Glycine max (L.) Merr. Microbes Environ 28:414–421
Tomasi N, Weisskopf L, Renella G, La L, Pinton R, Varanini Z, Nannipieri P, Torrent J, Martinoia E (2008) Flavonoids of white lupin roots participate in phosphorus mobilization from soil. Soil Biol Biochem 40:1971–1974
Ullers RS, Luirink J, Harms N, Schwager F, Georgopoulos C, Genevaux P (2004) SecB is a bona fide generalized chaperone in Escherichia coli. Proc Natl Acad Sci U S A 101:7583–7588
Wang W, Vinocur B, Shoseyov O, Arie A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252
Wang XQ, Yang PF, Gao Q, Liu X, Kuang T, Shen S, He Y (2008) Proteomic analysis of the response to high-salinity stress in Physcomitrella patens. Planta 228:167–177
Wenzel M, Lang K, Günther T, Bhandari A, Weiss A, Lulchev P, Szentgyörgyi E, Kranzusch B, Göttfert M (2012) Characterization of the flavonoid-responsive regulator FrrA and its binding sites. J Bacteriol 194:2363–2370
Williams A, Wilkinson A, Krehenbrink M, Russo DM, Zorreguieta A, Downie JA (2008) Glucomannan-mediated attachment of Rhizobium leguminosarum to pea root hairs is required for competitive nodule infection. J Bacteriol 190:4706–4715
Xiao WL, Guan DW, Li J, Cao FM, Chen HJ (2010) Evaluation on the competitiveness of strains of soybean rhizobia marking with gfp and rfp genes. Soybean Sci 29:336–369
Xie B, Chen D, Cheng G, Ying Z, Xie F, Li Y, Zhou J (2009) Effects of the purL gene expression level on the competitive nodulation ability of Sinorhizobium fredii. Curr Microbiol 59:193–198
Yan SP, Zhang QY, Tang ZC, Su WA, Sun WN (2006) Comparative proteomic analysis provides new insights into chilling stress responses in rice. Mol Cell Proteomics 5:484–496
Yan Y, Yang J, Dou Y, Chen M, Ping S, Peng J, Lu W, Zhang W, Yao Z, Li H (2008) Nitrogen fixation island and rhizosphere competence traits in the genome of root-associated Pseudomonas stutzeri A1501. Proc Natl Acad Sci U S A 21:7564–7569
Yates RJ, Howieson JG, Reeve WG, Graham WO (2011) A reappraisal of the biology and terminology describing rhizobial strain success in nodule occupancy of legumes in agriculture. Plant Soil 348:255–267
Acknowledgments
This work was supported by the Special Fund for Establishment of Modern Agricultural R&D System; Ministry of Finance and Ministry of Agriculture, China (CARS-04); the National Basic Research Program (973 Program) (2015CB150506); the National Natural Science Foundation of China For Young Scholars (31200388); the National High-tech R&D Program of China (863 Program) (2013AA102802-04); and National Nonprofit Institute Research Grant of CAAS (IARRP-2014-4).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, Y., Guan, D., Jiang, X. et al. Proteins involved in nodulation competitiveness of two Bradyrhizobium diazoefficiens strains induced by soybean root exudates. Biol Fertil Soils 51, 251–260 (2015). https://doi.org/10.1007/s00374-014-0969-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00374-014-0969-9