Abstract
Plant growth retardants can enhance the lodging resistance of crops and prevent yield losses. In this study, we used the crystal structure of the gibberellin receptor GIBBERELLIN INSENSITIVE DWARF1 (GID1) to aid the discovery of novel growth retardants. The Maybridge database was filtered based on multiple-screening analysis, including pharmacophore modeling, target structures, Lipinski’s rules of 5, and physicochemical properties. Finally, the 1,3-benzodioxole compounds were screened and their potential interaction mechanisms, synthesis, and biological activity on the root lengths of wild-type and mutant-type Arabidopsis thaliana were investigated. The results showed that the compounds inhibited root growth in WT A. thaliana at a range of concentrations, with optimal inhibition observed at 100 μmol/L. The inhibition by compound HTS05309 was comparable to that of uniconazole (100%). Quantitative experiments confirmed that the compounds inhibited A. thaliana growth. These results suggested that novel plant growth retardants have been identified based on computational-aided discovery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
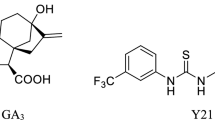
Gibberellins are a class of plant growth regulators that promote the longitudinal growth of stems and induce hydrolytic enzymes in germinating seeds (Matsuoka 2003; Petracek et al. 2013; Rademacher 2015). To date, 136 gibberellins have been identified in plants, fungi and bacteria (Garcia-Martinez and Gil 2002). Gibberellin acid (GA3) has been widely used in agricultural production as a known promoter of plant growth (Ozga and Reinecke 2003) (Fig. S1). Phthalimide compounds, including AC94377 (Suttle and Hultstr 1987), AC006, AC007, and AC015 (Yalpani et al. 1989; Suttle et al. 1992), or GA3 derivatives including 10d and 10f (Tian et al. 2017) have been identified as functional analogs of gibberellins (Fig. S1). N-Substituted phthalimides (NSPs) compete for binding to GA4 with gibberellin-binding proteins (Yalpani et al. 1989; Jiang et al. 2017). GIBBERELLIN INSENSITIVE DWARF1 (GID1) was discovered to be a soluble gibberellin receptor (Ueguchi-Tanaka et al. 2005; Nakajima et al. 2006; Hauvermale et al. 2014) which provides the foundation for target-based signaling transduction, metabolism, and molecular design.
Although gibberellin increases plant yields, it also causes conflicts between “high yield and lodging”. In the 1960s, the large-scale promotion of semi-dwarf rice and wheat varieties greatly improved crop yields and solved the food crisis (Peng et al. 1999). Furthermore, dwarf plants with gibberellin synthesis and signal transduction defects are largely resistant to lodging. Uniconazole and paclobutrazol, trinexapac-ethyl and prohexadione calcium, inhibitors of kauriene oxidase, and gibberellin 3-oxidase (GA3ox) inhibit gibberellin biosynthesis and are broad-spectrum, high-efficiency, low-residue plant growth retardants that can dwarf plants and shorten root lengths (Adriansen and Odgaard 1997). As such, plant growth retardants can increase plant yields through lodging resistance.
In this study, we combined plant growth regulators, computer-aided drug design and organic synthesis, based on multiple screening, gibberellin targets, pharmacodynamics, Lipinski’s roles of five, physicochemical properties, toxicity and biological activity determination, to identify new and highly active plant growth retardants. Moreover, molecular binding modes and binding free energy studies were also performed.
Materials and Methods
Molecular Conformation and Energy Optimization
Ligands were constructed using the Build/Edit module. The lowest-energy conformations were optimized by SYBYL 7.3 software using the steepest gradient descent method (Hirano et al. 2010). The Tripos force-field was selected and the Gasteiger–Hückel charge was loaded with 10,000 iterations and an energy convergence of 0.005 kJ/mol. Other parameters were defined as default values unless otherwise indicated.
Molecular Docking
Molecular docking was performed using SYBYL 7.3 software using the Surflex-dock algorithm (Jain 2003; Clark et al. 2002) in which the crystal structure of GA3-GID1A-DELLA from the RCSB protein date bank (PDB ID: 2ZSH) was used as the docking receptor. The receptor model was prepared with charges, atom types, protonation, and the addition of hydrogen atoms. The best protomol in the active domain of the receptor was obtained using a ligand docking mode.
Building Pharmacophore Models
Seven reported gibberellins including GA3, GA4, GA7 and functional analog NSPs including AC94377, AC006, AC007, and AC015 were used to generate a 3D pharmacophore model using the GALAHAD method. This method encompassed five features including: hydrogen bond donor and acceptor atoms; positive nitrogen; and negative hydrophobic centers. Twenty pharmacophore models were obtained and evaluated, and representative models were selected. Parameters including population size, max generations, mols required to N hit, and keep best N models were set to 45, 45, 5 and 20, respectively. Other parameters used default values, unless stated otherwise.
Virtual Screening
The Maybridge database containing a collection of 58,698 molecules was used for virtual screening. To ensure accuracy of screening protocols, the energy of the molecules was minimized prior to the screening process. Initially, the Maybridge database was filtered using Lipinski’s rules of five (M < 450) (Sander et al. 2015; Kelder et al. 1999). LogS and ADME/T for the molecules was then predicted using the open software DataWarrior in which low solubility and toxic compounds were filtered. The resultant molecules were finally screened against the pharmacophore models of gibberellins and the active pocket of GID1A using SYBYL 7.3 software. The screening molecules were ranked by score clustering and the top 15% were selected. Four of the selected were synthesized based on their solubility and available synthesis.
Molecular Dynamics Simulations
The complex crystal structure of GA3-GID1A-DELLA was used to mimic the receptor–ligand interaction between through molecular dynamics simulations. During the preparation of the receptor structure, the protonation states of histidine residues were manually inspected to ensure consistency with the local chemical environment. All systems were optimized using Amber14 software. The force-field parameters for the proteins were generated using Amber ff99SB (Hornak et al. 2006), whilst those for ligands were generated using General Amber Force Field (GAFF) (Wang et al. 2004). A rectangular water box was used to fill the gaps between the receptor and ligands, which was generated with TIP3P water molecules with an edge of 12 Å. Na+ ions were added to neutralize the system and data collection of 10 ns of the production run was performed through molecular dynamics simulations (Chen et al. 2018). Finally, calculations of the free binding energy were performed using the MM/GBSA method available in Amber14 software (Zhu et al. 2014).
Biological Activity Assays
The bioactivity of the synthesized compounds was determined in vivo by measuring the root length of A. thaliana. 1/2 MS (Murashige–Skoog) culture media were prepared by mixing distilled water (1 L) with MS (1.08 g), sucrose (5 g), and agar (4 g), and pH was adjusted to 5.9 with Sodium hydroxide. The media was sterilized in a high-pressure steam sterilization pot and A. thaliana seeds were sterilized in 1% sodium hypochlorite for 15 min. Following sterilization with sodium hypochlorite, seeds were washed five times with sterile distilled water. Seeds were planted in 1/2 MS media with four concentrations (0.1, 1, 10, and 100 μmol L−1). GA3 and uniconazole were included as controls. Each medium was planted with 12 seeds and maintained at 4 °C for two days. Then, seeds were incubated under consecutive light (400 μmol−2 s−1 of photon flux density) for 7 days. Root lengths were measured using ImageJ software (Tian et al. 2017).
Results
Molecular Docking of GAs and Their Functional Analogs
Molecular simulations can be used in drug design, but their reliability is dependent on their correlation to experimental data. To verify the accuracy of our molecular simulations, GA3, GA4, and GA7, and four functional analogs, AC94377, AC006, AC007, and AC015 (Fig. S1) were docked onto the GID1A receptor to predict the correlation between binding scores and experimental protein-binding activity.
As shown in Fig. S2, the predicted docking scores correlated with the protein-binding activity, pIC50, and high R2 values (approximately equal to 0.71). The high-predicted docking scores indicated a high affinity between the ligand and receptor. The conformations of high-scoring ligands were then superimposed (Table S1). The NSPs were found to be highly similar to gibberellins and could fit into the binding pocket of the GID1A receptor, indicating their potential as competitive inhibitors to gibberellins.
Pharmacophore Models
Seven molecules with diverse structures, including GA3, GA4, GA7, AC94377, AC006, AC007, and AC015, could fit into the binding cavity of GID1A with good binding ability. These compounds were used as training sets for the construction of pharmacophore models using the GALAHAD method (Abrahamian et al. 2003; Richmond et al. 2006). The best pharmacophore model with the highest number of features was selected amongst the 20 generated pharmacophore models using the GALAHAD method (Fig. S3). The generated query consisted of four hydrophobic centers, one H-bond donor atom, and three H-bond acceptor atoms. Six ‘hit’ ligands amongst the 7 molecules were obtained through 3D searches. Furthermore, 100 molecules, including 7 in the training sets and 93 randomly selected from the Maybridge database were used to verify the reliability of the generated model. Seven molecules in the training sets achieved enrichment factors (EFs) (Huang et al. 2006) as an index of screening ability of 10. This indicated that the model was reliable, and can accurately identify gibberellin-like molecules from pools of molecules from the Maybridge database.
Virtual Screening
Virtual screening against the Maybridge database was performed according to the workflow shown in Fig. 1. Prior to virtual screening, the 2D structures of all molecules were converted to 3D structures using SYBYL 7.3. Database screening was performed using the Lipinski’s rule of five (M ≤ 450) (Huang et al. 2006) using SYBYL 7.3 software. DataWarrior was used to determine toxicity, LogS, and the polar surface area (PSA, PSA < 120 Å) of all molecules. All toxic molecules were removed from the database. A total of 3078 molecules resembling the best pharmacophore models were selected and used in the screening process. The molecules were docked onto the binding site of the GID1A receptor using SYBYL7.3. The results were ranked by total scores. To ensure screening accuracy, each conformation obtained from the docking was subjected to clustering analysis, in addition to visual judgments. The top 15% of molecules were selected, amongst which four 1,3-benzodioxole compounds were selected for mechanistic studies.
Binding Mode Analysis
Compounds HTS05467, HTS05309, HTS05461, NRB00180 and GA3 were docked onto the GID1A receptor, and the binding modes were analyzed to elucidate potential molecular mechanisms. The interactions between GA3 and the receptor indicated hydrogen bonds between the ligand and residues SER116 (1.85 Å), TYR127 (1.87 Å), and SER191 (2.74 Å) (Fig. 2a). Water molecules are required for the GA3-GID1A interaction (Murase et al. 2008). In this study, water molecules H2O348, H2O350, and H2O364 formed hydrogen bonds with GA3, whilst other water molecules including H2O380, H2O400, H2O401 created aqueous environments for the complexation of GA3-GID1A-DELLA. These conclusions were consistent with previous studies (Murase et al. 2008; Duan et al. 2013).
Proposed binding modes of GA3 (a) and compound HTS05309 (b) to GID1A (2ZSH). Hydrophobic residues are labeled in green, electrostatic interactions are labeled in red, water molecules are labeled in blue, and unique residues ASP190 were labeled in black. The yellow-dotted lines indicate the hydrogen bonds, purple-dotted lines indicate π–π interactions, and black solid lines indicate π-cation (Color figure online)
Compound HTS05309 could bind to the GID1A receptor (Fig. 2b) through a similar binding mode to GA3. Moreover, the binding conformation of HTS05309 with a linker carrying two aromatic rings in a ‘U-type’ conformation was loaded onto the receptor-binding site. During compound HTS05309 binding, van der Waals interactions contributed to the hydrophobic effects, involving residues PHE27, TYR31and VAL319. Whilst residues, GLY114, SER116, SER127, ASP190, SER191, and ARG244 contributed to the electrostatic interactions, water molecules from H2O348, H2O350, H2O364, H2O400, H2O401, H2O443, and H2O458 played a major role during ligand–receptor binding. The intramolecular π–π interaction, which enhanced the structural stability and affinity, was formed between the two benzene rings of compound HTS05309. The head of the benzene ring of compound HTS05309 was situated in the hydrophobic environment, with the interaction between residues ILE24, PHE27, and TYR31 at the cap region of the DELLA protein stabilizing its structure (Hirano et al. 2007). The other end of compound HTS05309 interacted with residues GIY115, SER116, TYR127, and SER191 in a relatively hydrophilic environment, thereby enhancing the affinity towards the inhibitor, whilst stabilizing the binding site. The intermolecular π-cation interaction that enhances binding affinity, lipophilicity, specificity, and drug design (Yoon et al. 2013) was formed between the benzene ring in compound HTS05309 and an N atom of ARG244 in GID1A. Hydrogen bonding between GIY115 (2.61 Å) or SER191 (2.51 Å) of the GID1A receptor and HTS05309 was also observed. As shown in Table S2, the structure of HTS05309 is similar to that of GA3 with an overlap score of 0.61. The binding score (5.36) to the receptor was comparable to the binding score of GA3 (5.92). It can be speculated that compound HTS05309 acts as a gibberellin functional analog. Compound HTS05309 showed a similar binding mode to GA3 mainly due to the similarity in pharmacophores, competing with GA3 for receptor binding. This finding requires confirmation in protein-binding activity assays, and physiological studies.
Verification of Binding-Free Energy
The free binding energy of a 10 ns molecular dynamic simulation of compound HTS05309 and GA3 was assessed using AMBER14 software. As shown in Table 1, HTS05309 binds to the GID1A receptor with a total binding energy of − 47.53 kcal/mol, which is remarkably lower than that of GA3 (− 39.13 kcal/mol). This indicates that HTS05309 has a stronger receptor-binding affinity than GA3. Furthermore, energy decomposition assessments showed that van der Waals and electrostatic interactions were the major contributors to the binding affinities. The binding energy of compound HTS05309 was dictated by van der Waals interactions (− 48.20 kcal/mol) which were twofold higher than the electrostatic interactions (− 23.86 kcal/mol). As previously described (Chen et al. 2018), non-polar solvation exerts positive effects on the binding energy, while polar solvation has negative effects, in agreement with the results of this study (Table 1).
To explore the contribution of each individual residue, the energy decomposition of ligand–residue pairs was determined. The decomposing energy of key-binding residues between GA3 and compound HTS05309 are illustrated in Fig. 3. The contribution rates of the major residues for GA3 and compound HTS05309 are listed in Table S3. In agreement with our binding mode analysis, GA3-receptor binding was dominated by Van der Waals interactions involving LE24, PHE27, TYR31, and VAL319 (Duan et al. 2013), with a contribution rate of 50.76%, compared to the electrostatic interactions of SER116, SER191, and ARG244 (Murase et al. 2008) that contributed only 9.85% (Table S3).
ILE24, PHE27, TYR31, GLY115, HIS119, VAL239, VAL319, TYR322, and LEU323 interact with GA3 and HTS05309 through van der Waals forces (Table S3) in agreement with our binding mode results. The contribution rate of ILE24 in the cap region to GA3 binding was fivefold higher than HTS05309 (0.87%). However, the contribution of PHE27 to GA3 binding (13.09%) was lower than HTS05309 (17.61%) as HTS05309 was closer to PHE27. TYR31 was found to interact with GA3 and HTS05309 with almost equal van der Waals forces. GLY115 was unique and stabilized compound HTS05309 through H-bond interactions (Fig. 3). Van der Waals interactions were not involved in GA3 binding, enhancing the binding affinity. The other 3 residues HIS119, VAL239, and TYR322 also interacted with GA3 and HTS05309 with equal contribution rates of approximately 11%. It was noteworthy that the contribution of VAL239 to the binding was higher than that of HIS119 and TYR322. Residue VAL319 reported to be important for the formation of the GA3-GID1A-DELLA complex (Murase et al. 2008), also interacts with HTS05309 through van der Waals interactions. Compound HTS05309 binds to the GID1A receptor with a van der Waals force of − 48.20 kcal/mol, which is close to that of GA3 (− 44.42 kcal/mol). This indicates that HTS05309 and GA3 have equal binding affinities for van der Waals interactions.
Other residues including SER116, SER127, ASP190, SER191, and ARG244 interacted with GA3 and HTS05309 through electrostatic interactions (Table S3), in agreement with our binding mode analysis. However, despite hydrogen bond formation between GA3 and SER116 (1.85 Å), the contribution of SER116 to the GA3-GID1A-DELLA (3.55%) interaction remained lower than that of HTS05309-GID1A-DELLA (5.52%), most likely due to a loss of polarity. ASP190 has a negative effect on the receptor interaction between GA3 (− 10.92%) and HTS05309 (− 4.23%) due to effects of polarity that may offset the electrostatic interactions. ASP190 may thus be important in the ligand-target interaction, and should be considered during drug design. Hydrogen bonds between SER191 and the ligand were observed in the crystal structure of GA3-GID1A-DELLA and the docking of HTS05309-GID1A-DELLA complexes. Its contribution to the GA3-GID1A-DELLA complex was 1.64-fold higher than the HTS05309-GID1A-DELLA complex. ARG244 equally interacts with GA3 and HTS05309 (8.19% and 9.01%, respectively) and has a smaller effect on the polarity interaction. Overall, although the interaction modes of HTS05309 and GA3 were similar, polar interactions lead to differences in the total free binding energy. For GA3 and compound HTS05309, the differences were reflected by the contribution of PHE27 and ASP190.
Biological Activity Assays
To evaluate the activity of the lead compound, HTS05467, HTS05309, HTS05461 and NRB00180, were synthesized in two-steps (Platzer et al. 2016; Yan et al. 2016) as shown in Fig. 4. Firstly, benzyl bromide with substituents was reacted with thioglycolic acid. Using Et3N as an acid-binding agent, the thioether was then reacted with benzo[d] [1,3] dioxol-5-amine under the catalysis of DMAP and EDCI, producing the target products. The details in synthesis are shown in supplementary material.
Root lengths were used as indicators of HTS05467, HTS05309, HTS05461 and NRB00180 activity against the growth of WT and mutant A. thaliana (Table 2). In control group, GA3 had no effect, whilst uniconazole led to significant inhibition. As indicated by the root length, HTS05467, HTS05309, HTS05461 and NRB00180 had a significant inhibition growth effects on WT plants, which is similar with that of uniconazole. Compounds HTS05467, HTS05309, HTS05461 and NRB00180 had no significant effects on WT plants at low concentration, but exhibited considerable effects at high concentrations (Liang et al. 2018). At 0.1 and 10 μmol L−1 (Table 2), the effects of compound HTS05461 on WT plants were similar with uniconazole, and higher than that of compounds HTS05309 (10% and 11%) and NRB00180 (15% and 16%). At 10 μmol L−1, compounds HTS05309 and NRB00180 significantly inhibited WT plants (25% and 37%, respectively) whilst HTS05467 and HTS05461 had no significant effects. At 100 μmol L−1, all compounds significantly inhibited WT plants with inhibition rates of 89%, 100%, 76%, and 73% for compounds HTS05467, HTS05309, HTS05461 and NRB00180, respectively. Compounds HTS05467, HTS05309, HTS05461 and NRB00180 had strong inhibitory effects on mutant plants. In general, compounds HTS05467, HTS05309, HTS05461 and NRB00180 had significant inhibitory effects on both WT and mutant type. In particular, compound HTS05309 was similar with that of uniconazole, which exhibited 100% inhibition at 100 μmol L−1.
The results showed that the lead compounds inhibited plant growth, which may be due to the common pharmacophore characteristics between the promoters and inhibitors. To validate these findings, four commercially available growth retardants, namely uniconazole, paclobutrazol, prohexadione calcium and trinexapac-ethyl were collected to establish the pharmacophore model (Fig. 5). The compounds were screened though flexible screening. Upon comparison, the main difference was that the inhibitor model lacked a symmetric hydrogen bond acceptor, and the only hydrogen bond donor was hydrophobic. Inhibitors and accelerator models are more comparable with overlapping pharmacodynamics. Thus, the model can successfully screen growth retardants.
Promoter and retardant pharmacophore characteristics. a Promoter model using AC94377 as an example. b Characteristics of the inhibitor simulation using uniconazole as an example. Cyan spheres: hydrophobic centers (HY); red spheres: donor atoms (DA); green spheres: acceptor atoms (AA); yellow-dotted lines: differences between the models (Color figure online)
Discussion
Phthalimides have been shown to act as competitive inhibitors of gibberellin receptors, of which AC94377 and GA3 have comparable effects (Jiang et al. 2017) and bind to the active sites of GA4 receptors. Our findings suggests that AC94377 competes with gibberellin for receptor binding due to its high-structural similarity, with both compounds entering the active pocket of the receptor, with similar modes of action. The crystal structure of GA3-GID1A-DELLA, docking models, dynamic simulations, and combined free decomposition indicated that residues TYR127, SER116, and SER191 contribute important electrostatic interactions and hydrogen bonding during GA3 and GID1A binding, whilst non-polar residues ILE24, PHE27, TYR31 and VAL319 contribute hydrophobic effects. Important residues during the binding of GA3 to GID1A have been previously characterized (Murase et al. 2008; Duan et al. 2013). ASP190 negatively effects the interaction due to polarity effects offsetting the electrostatic interactions, which may be of importance during ligand-target drug design.
The GALAHAD method can accurately construct pharmacodynamics using external macro definition files to assign features, whilst allowing the donor and acceptor to overlap (Abrahamian et al. 2003). Richmond et al. (2006). showed that the ligand pharmacophore generated by GALAHAD resembles that of the crystal structure of its receptor. Moreover, the identification of new features can be beneficial for drug modifications, and so the GALAHAD method was used to establish accurate pharmacophore models for screening using gibberellins and its functional analogs. Virtual screening employs multiple pathways, including Lipinski’s rule of five (M ≤ 450) (Huang et al. 2006), physicochemical properties, toxicity, LogS, polar surface areas (PSA, PSA < 120 Å), pharmacodynamics of rifalazil and target-based drug design. A total of four 1,3-benzodioxole compounds were screened from the Maybridge database. The binding modes of the lead compounds were similar to GA3, whilst the binding activity of the compounds increased. The most significant difference between the binding mode of lead compounds and GA3 was the contribution of ASP190. Compared with GA3, the polar interactions and collision between the ligands and target were enhanced, thus increasing binding affinity. Thus, on the premise of retaining interactions with key residues, reducing the polar interaction and collision of the ligands with ASP190 enhances the affinity between the ligand and its target, which is conducive to improved activity. Screening and bioactivity determinations based on the pharmacophore model and target revealed that the model was a valid screening method that could exclude molecules that did not match the structure and reduces the number of compounds entering the next stage. Screening based on these targets could lead to the identification of drugs based on the binding mode (hydrogen bonds, hydrophobicity and hydrophilicity).
In summary, the 1,3-benzodioxoles HTS05467, HTS05309, HTS05461 and NRB00180 were filtered out from the Maybridge database through a multiple-screening strategy. Their potential interaction mechanisms were studied and lead compounds were synthesized to determine their effects on WT and mutant Arabidopsis root lengths. The 1,3-benzodioxole compounds, and compounds HTS05467, HTS05309, HTS05461 and NRB00180 exhibited inhibitory effects on A. thaliana growth (inhibition rates of 89%, 100%, 76% and 73%, respectively). Compound HTS05309 was comparable to uniconazole in terms of activity. Quantitative experiments further demonstrated that the 1,3-benzodioxole compounds inhibited A. thaliana growth. The inhibitor model was constructed to reveal this phenomenon, and indicated that the retardant or inhibitor model and accelerator model have similar pharmacophore characteristics, ultimately leading to the production of an inhibitory lead compound. The discovery of gibberellin inhibitors is beneficial to improving plant resistance to lodging and stress stimuli, ultimately increasing the biological yield.
References
Abrahamian E, Fox PC, Naerum L, Christensen IT, Thøgersen H, Clark RD (2003) Efficient generation, storage, and manipulation of fully flexible pharmacophore multiplets and their use in 3-D similarity searching. J Chem Inf Comput Sci 43:458–468. https://doi.org/10.1021/ci025595r
Adriansen E, Odgaard P (1997) Residues of paclobutrazol and uniconazole in nutrient solutions from ebb and flood irrigation of pot plants. Sci Horic 69:73–83. https://doi.org/10.1016/S0304-4238(96)00982-X
Chen X, Gan Q, Feng CG, Liu X, Zhang Q (2018) Virtual screening of novel and selective inhibitors of PTP1B over TCPTP using a bidentate inhibition strategy. J Chem Inf Model 58:837–847. https://doi.org/10.1021/acs.jcim.8b00040
Clark RD, Strizhey A, Leonard JM, Blake JF, Matthew JB (2002) Consensus scoring for ligand/protein interactions. J Mol Graph Model 20:281–295. https://doi.org/sci-hub.tw/10.1016/s1093-3263(01)00125-5
Duan HX, Li DL, Liu HC, Liang DS, Yang XL (2013) Computational insight into novel molecular recognition mechanism of different bioactive GAs and the Arabidopsis receptor GID1A. J Mol Model 19:4613–4624. https://doi.org/10.1007/s00894-013-1971-0
Garcia-Martinez JL, Gil J (2002) light regulation of gibberellin biosynthesis and mode of action. J Plant Growth Regul 20:354–368. https://doi.org/10.1007/s003440010033
Hauvermale AL, Ariizumi T, Steber CM (2014) The roles of the GA receptors GID1a, GID1b, and GID1c in Sly1-independent GA signaling. Plant Signal Behav 9:2125–2139. https://doi.org/10.4161/psb.28030
Hirano K, Asano K, Tsuji H, Kawamura M, Mori H (2010) Characterization of the molecular mechanism underlying gibberellin perception complex formation in rice. Plant Cell 22:2680–2696. https://doi.org/10.1105/tpc.110.075549
Hirano K, Nakajima M, Asano K, Nishiyama T, Sakakibara H, Kojima M, Katoh E, Xiang H, Tanahashi T, Hasebe M (2007) The GID1-mediated gibberellin perception mechanism is conserved in the lycophyte selaginella moellendorffii but not in the bryophyte physcomitrella patens. Plant Cell 19:3058–3079. https://doi.org/10.1105/tpc.107.051524
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006) Comparison of multiple AMBER force fields and development of improved protein backbone parameters. Proteins 65:712–725. https://doi.org/10.1002/prot.21123
Huang N, Shoichet BK, Irwin JJ (2006) Benchmarking sets for molecular docking. J Med Chem 49:6789–6819. https://doi.org/10.1021/jm0608356
Jain AN (2003) Surflex: fully automatic flexible molecular docking using a molecular similarity-based a search engine. J Med Chem 46:499–511. https://doi.org/10.1021/jm020406h
Jiang K, Otani M, Shimotakahara H, Yoon JM, Park SH, Miyaji T, Nakano T, Nakamura H, Nakajima M, Asami T (2017) Substituted phthalimide AC94377 is a selective agonist of the gibberellin receptor GID1. Plant Physiol 133:825–835. https://doi.org/10.1104/pp.16.00937
Kelder J, Grootenhuis PDJ, Bayada DM, Delbressine LPC, Ploemen JP (1999) Polar molecular surface as a dominating determinant for oral absorption and brain penetration of drugs. Pharm Res 16:1514–1519. https://doi.org/10.1023/a:1015040217741
Liang ZB, Xi QL (2018) π-Cation interactions in molecular recognition: perspectives on pharmaceuticals and pesticides. J Agric Food Chem 66:3315–3323. https://doi.org/10.1021/acs.jafc.8b00758
Matsuoka M (2003) Gibberellin signaling: how do plant cells respond to ga signals? J Plant Growth Regul 22:123–125. https://doi.org/10.1007/s00344-003-0039-2
Murase K, Hirano Y, Sun T, Hakoshima T (2008) Gibberellin-induced DELLA recognition by the gibberellin receptor GID1. Nature 456:459–464. https://doi.org/10.1038/nature07519
Nakajima M, Shimada A, Takashi Y, Kim YC, Park SH, Ueguchi-Tanaka M, Suzuki H, Katoh E, Iuchi S, Kobayashi M, Maeda T, Matsuoka M, Yamaguchi I (2006) Identification and characterization of Arabidopsis gibberellin receptors. Plant J 46:880–889. https://doi.org/10.1111/j.1365-313x.2006.02748.x
Ozga JA, Reinecke DM (2003) Hormonal interactions in fruit development. J Plant Growth Regul 22:73–81. https://doi.org/10.1007/s00344-003-0024-9
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) 'Green revolution' genes encode mutant gibberellin response modulators. Nature 400:256–261. https://doi.org/10.1038/22307
Petracek PD, Silverman FP, Greene DW (2013) A history of commercial plant growth regulators in apple production. HortScience 38:937–942. https://doi.org/10.21273/HORTSCI.38.5.937
Platzer S, Kar M, Leymaa R, Chiba S, Jirsa Roller AF, Krachler R, MacFarlane DR, Kandioller W, Keppler BK (2016) Task-specific thioglycolate ionic liquids for heavy metal extraction: synthesis, extraction efficacies and recycling properties. J Hazard Mater 324:241–249. https://doi.org/10.1016/j.jhazmat.2016.10.054
Rademacher W (2015) Plant growth regulators: backgrounds and uses in plant production. J Plant Growth Regul 34:845–872. https://doi.org/10.1007/s00344-015-9541-6
Richmond NJ, Abrams CA, Wolohan PR, Abrahamian E, Willett P, Clark RD (2006) GALAHAD: 1. Pharmacophore identification by hypermolecular alignment of ligands in 3D. J Comput Aid Mol Des 20:567–587. https://doi.org/10.1007/s10822-006-9082-y
Sander T, Freyss J, Von KM, Rufener C (2015) DataWarrior: an open-source program for chemistry aware data visualization and analysis. J Chem Inf Model 55:460–473. https://doi.org/10.1021/ci500588j
Suttle JC, Hultstr JF (1987) Physiological studies of a synthetic gibberellin-like bioregulator. Plant Physiol 84:1068–1073. https://doi.org/10.1104/pp.84.4.1068
Suttle JC, Hultstr JF, Tanaka FS (1992) The biological activities of five azido N-substituted phthalimides: potential photoaffinity reagents for gibberellin receptors. Plant Growth Regul 11:311–318. https://doi.org/10.1007/bf00024570
Tian H, Xu YR, Liu SJ, Jin DS, Zhang JJ, Duan LS, Tan WM (2017) Synthesis of gibberellic acid derivatives and their effects on plant growth. Molecules 22:694–703. https://doi.org/10.3390/molecules22050694
Ueguchi-Tanaka M, Ashikari M, Nakajima M, Itoh H, Katoh E, Kobayashi M, Chow TY, Hsing YI, Hsing H, Yamaguchi I, Matsuoka M (2005) Gibberellin insensitive dwarf1 encodes a soluble receptor for gibberellin. Nature 437:693–698. https://doi.org//10.1038/nature04028
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25:1157–1174. https://doi.org/10.1002/jcc.20035
Yalpani N, Suttle JC, Hultstr JF, Rodaway SJ (1989) Competition for in vitro [3H] gibberellin A4 binding in cucumber by substituted phthalimides: comparison with in vivo gibberellin-like activity. Plant Physiol 91:823–828. https://doi.org/10.1104/pp.91.3.823
Yan JB, Li SH, Gu M, Yao RF, Li YW, Chen J, Yang M, Nan FJ, Xie DX (2016) Endogenous bioactive jasmonate is composed of a set of (+)-7-iso-JA-amino scid conjugates. Plant Physiol 172:2154–2164. https://doi.org/10.1104/pp.16.00906
Yoon JY, Nakajima M, Mashiguchi K, Park SH, Otania M, Asami T (2013) Chemical screening of an inhibitor for gibberellin receptors based on a yeast two-hybrid system. Bioorg Med Chem Lett 23:1096–1098. https://doi.org/10.1016/j.bmcl.2012.12.007
Zhu YL, Beroza P, Artis DR (2014) Including explicit water molecules as part of the protein structure in MM/PBSA calculations. J Chem Inf Model 54:462–469. https://doi.org/10.1021/ci4001794
Acknowledgements
This work was financially supported by the National Key Research and Development Program of China (2017YFD0201300) and the National Natural Science Foundation of China (31872850).
Author information
Authors and Affiliations
Contributions
Jine Wang carried out drug calculation, biological activity experiments and manuscript writing; Zhikun Yang carried out the synthesis of lead compounds, biological activity experiments and chapter writing; Hongxia Duan helped to carry out experimental analysis and control the writing progress and review of manuscripts; Liusheng Duan confirmed the research content, controlled the progress of the test and the quality of the manuscript writing; Weiming Tan conceived and designed experiments, and controlled the writing progress and review of manuscripts.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, J., Yang, Z., Duan, H. et al. Computational-Aided Discovery of Novel 1,3-Benzodioxole Plant Growth Retardants. J Plant Growth Regul 39, 888–896 (2020). https://doi.org/10.1007/s00344-019-10030-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-019-10030-1