Abstract
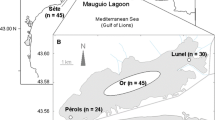
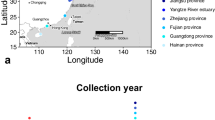
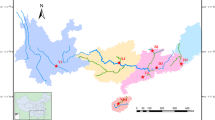
The Japanese eel (Anguilla japonica) is a commercially important fish species in East Asia and its recruitment has been rapidly declining since 1990s. Clarifying the genetic population structure of A. japonica is the basis of multinational cooperation on its management and protection, due to its large distribution range. Gene-associated markers have been proved powerful in delineating fine-scale population genetic structure and spatially varying selection. In the present study, we developed 24 polymorphic gene-associated microsatellite markers including 18 loci associated with the genes under selection in the two North Atlantic eel species (Anguilla anguilla and Anguilla rostrata) and 6 loci based on transcript sequences. A total of 13 geographic populations were sampled across its distribution range, including 11 samples from China (9 from China's mainland and 2 from Taiwan region), and 2 samples from Japan. A total of 416 individuals (mostly glass eels) were collected and genotyped at the 24 microsatellites. All measures of differentiation were accordant with a panmictic scenario (FST=-0.001) in A. japonica. No footprints of spatially varying selection were found, indicating that the selection pattern in A. japonica might be different from that in the two North Atlantic eel species. We suggest that A. japonica should be managed as a single unit and management and conservation efforts must be coordinated at the international level, as overexploitation in any region will decrease its recruitment across the whole distributional range.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Als T D, Hansen M M, Maes G E, Castonguay M, Riemann L, Aarestrup K, Munk P, Sparholt H, Hanel R, Bernatchez L. 2011. All roads lead to home: panmixia of European eel in the Sargasso Sea. Molecular Ecology, 20(7): 1333–1346, https://doi.org/10.1111/j.1365-294X.2011.05011.x.
Antao T, Lopes A, Lopes R J, Beja-Pereira A, Luikart G. 2008. LOSITAN: a workbench to detect molecular adaptation based on a Fst-outlier method. BMC Bioinformatics, 9: 323, https://doi.org/10.1186/1471-2105-9-323.
Arai T. 2014. How have spawning ground investigations of the Japanese eel Anguilla japonica contributed to the stock enhancement?. Reviews in Fish Biology and Fisheries, 24(1): 75–88, https://doi.org/10.1007/s11160-013-9318-6.
Avise J C. 2003. Catadromous eels of the North Atlantic: a review of molecular genetic findings relevant to natural history, population structure, speciation, and phylogeny. In: Aida K, Tsukamoto K, Yamauchi K eds. Eel Biology. Springer, Tokyo. p.31–48, https://doi.org/10.1007/978-4-431-65907-5_3.
Beaumont M A, Nichols R A. 1996. Evaluating loci for use in the genetic analysis of population structure. Proceedings of the Royal Society B: Biological Sciences, 263(1377): 1619–1626, https://doi.org/10.1098/rspb.1996.0237.
Benjamini Y, Yekutieli D. 2001. The control of the false discovery rate in multiple testing under dependency. Annals of Statistics, 29(4): 1165–1188, https://doi.org/10.1214/aos/1013699998.
Chagné D, Chaumeil P, Ramboer A, Collada C, Guevara A, Cervera M T, Vendramin G G, Garcia V, Frigerio J M, Echt C, Richardson T, Plomion C. 2004. Cross-species transferability and mapping of genomic and cDNA SSRs in pines. Theoretical and Applied Genetics, 109(6): 1204–1214, https://doi.org/10.1007/s00122-004-1683-z.
Chan I K K, Chan D K O, Lee S C, Tsukamoto K. 1997. Genetic variability of the Japanese eel Anguilla japonica(Temminck & Schlegel) related to latitude. Ecology of Freshwater Fish, 6(1): 45–49, https://doi.org/10.1111/j.1600-0633.1997.tb00141.x.
Dekker W. 2003. Did lack of spawners cause the collapse of the European eel, Anguilla anguilla?. Fisheries Management and Ecology 10(6): 365–376, https://doi.org/10.1111/j.1365-2400.2003.00352.x.
Evanno G, Regnaut S, Goudet J. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology, 14(8): 2611–2620, https://doi.org/10.1111/j.1365-294X.2005.02553.x.
Excoffier L, Lischer H E L. 2010. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources, 10(3): 564–567, https://doi.org/10.1111/j.1755-0998.2010.02847.x.
Gagnaire P A, Normandeau E, Côté C, Hansen M M, Bernatchez L. 2012. The genetic consequences of spatially varying selection in the Panmictic American Eel(Anguilla rostrata). Genetics, 190(2): 725–703, https://doi.org/10.1534/genetics.111.134825.
Goudet J. 2001. FSTAT version 2.9.3.2. A program to estimate and test gene diversities and fixation indices. Institute of Ecology, Lausanne, Switzerland.
Gutierrez M V, Patto M C V, Huguet T, Cubero J I, Moreno M T, Torres A M. 2005. Cross-species amplification of Medicago truncatula microsatellites across three major pulse crops. Theoretical and Applied Genetics, 110(7): 1210–1217, https://doi.org/10.1007/s00122-005-1951-6.
Han Y S, Hung C L, Liao Y F, Tzeng W N. 2010. Population genetic structure of the Japanese eel Anguilla japonica: panmixia at spatial and temporal scales. Marine Ecology Progress Series, 401: 221–232, https://doi.org/10.3354/meps08422.
Hauser L, Carvalho G R. 2008. Paradigm shifts in marine fisheries genetics: ugly hypotheses slain by beautiful facts. Fish and Fisheries, 9(4): 333–362, https://doi.org/10.1111/j.1467-2979.2008.00299.x.
Hemmer-Hansen J, Therkildsen N O, Pujolar J M. 2014. Population genomics of marine fishes: next-generation prospects and challenges. Biological Bulletin, 227(2): 117–132, https://doi.org/10.1086/BBLv227n2p117.
Henkel C V, Dirks R P, de Wijze D L, Minegishi Y, Aoyama J, Jansen H J, Turner B, Knudsen B, Bundgaard M, Hvam K L, Boetzer M, Pirovano W, Weltzien F A, Dufour S, Tsukamoto K, Spaink H P, van den Thillart G E E J M. 2012. First draft genome sequence of the Japanese eel, Anguilla japonica. Gene, 511(2): 195–201, https://doi.org/10.1016/j.gene.2012.09.064.
Holsinger K E, Weir B S. 2009. Genetics in geographically structured populations: defining, estimating and interpreting F ST. Nature Reviews Genetics, 10(9): 639–650, https://doi.org/10.1038/nrg2611.
Hsu H Y, Chen S H, Cha Y R, Tsukamoto K, Lin C Y, Han Y S. 2015. De novo assembly of the whole transcriptome of the wild embryo, preleptocephalus, leptocephalus, and glass eel of Anguilla japonica and deciphering the digestive and absorptive capacities during early development. PLoS One, 10(9): e0139105, https://doi.org/10.1371/journal.pone.0139105.
Ishikawa S, Aoyama J, Tsukamoto K, Nishida M. 2001. Population structure of the Japanese eel Anguilla japonica as examined by mitochondrial DNA sequencing. Fisheries Science, 67(2): 246–253, https://doi.org/10.1046/j.1444-2906.2001.00227.x.
Itakura H, Kitagawa T, Miller M J, Kimura S. 2015. Declines in catches of Japanese eels in rivers and lakes across Japan: have river and lake modifications reduced fishery catches?. Landscape and Ecological Engineering, 11(1): 147–160, https://doi.org/10.1007/s11355-014-0252-0.
Joost S, Bonin A, Bruford M W, Després L, Conord C, Erhardt G, Taberlet P. 2007. A spatial analysis method(SAM) to detect candidate loci for selection: towards a landscape genomics approach to adaptation. Molecular Ecology, 16(18): 3955–3969, https://doi.org/10.1111/j.1365-294X.2007.03442.x.
Li Y L, Liu J X. 2018. StructureSelector: a web-based software to select and visualize the optimal number of clusters using multiple methods. Molecular Ecology Resources, 18(1): 176–177, https://doi.org/10.1111/1755-0998.12719.
Liewlaksaneeyanawin C, Ritland C E, El-Kassaby Y A, Ritland K. 2004. Single-copy, species-transferable microsatellite markers developed from loblolly pine ESTs. Theoretical and Applied Genetics, 109(2): 361–369, https://doi.org/10.1007/s00122-004-1635-7.
Liu J X, Avise J C. 2011. High degree of multiple paternity in the viviparous Shiner Perch, Cymatogaster aggregata, a fish with long-term female sperm storage. Marine Biology, 158(4): 893–901, https://doi.org/10.1007/s00227-010-1616-0.
Milano I, Babbucci M, Cariani A, Atanassova M, Bekkevold D, Carvalho G R, Espiñeira M, Fiorentino F, Garofalo G, Geffen A J, Hansen J H, Helyar S J, Nielsen E E, Ogden R, Patarnello T, Stagioni M, Consortium F, Tinti F, Bargelloni L. 2014. Outlier SNP markers reveal fine-scale genetic structuring across European hake populations(Merluccius merluccius). Molecular Ecology, 23(1): 118–135, https://doi.org/10.1111/mec.12568.
Minegishi Y, Aoyama J, Yoshizawa N, Tsukamoto K. 2012. Lack of genetic heterogeneity in the Japanese eel based on a spatiotemporal sampling. Coastal Marine Science, 35(1): 269–276.
Pashley C H, Ellis J R, McCauley D E, Burke J M. 2006. EST databases as a source for molecular markers: lessons from Helianthus. Journal of Heredity, 97(4): 381–388, https://doi.org/10.1093/jhered/esl013.
Peakall R, Smouse P E. 2006. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology Notes, 6(1): 288–295, https://doi.org/10.1111/j.1471-8286.2005.01155.x.
Peakall R, Smouse P E. 2012. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics, 28(19): 2537–2539, https://doi.org/10.1093/bioinformatics/bts460.
Pritchard J K, Stephens M, Donnelly P. 2000. Inference of population structure using multilocus genotype data. Genetics, 155(2): 945–959.
Pujolar J M, Jacobsen M W, Als T D, Frydenberg J, Munch K, Jónsson B, Jian J B, Cheng L, Maes G E, Bernatchez L, Hansen M M. 2014. Genome-wide single-generation signatures of local selection in the panmictic European eel. Molecular Ecology, 23(10): 2514–2528, https://doi.org/10.1111/mec.12753.
Rellstab C, Gugerli F, Eckert A J, Hancock A M, Holderegger R. 2015. A practical guide to environmental association analysis in landscape genomics. Molecular Ecology, 24(17): 4348–4370, https://doi.org/10.1111/mec.13322.
Rousset F. 2008. GENEPOP' 007: a complete reimplementation of the GENEPOP software for Windows and Linux. Molecular Ecology Resources, 8(1): 103–106, https://doi.org/10.1111/j.1471-8286.2007.01931.x.
Russello M A, Kirk S L, Frazer K K, Askey P J. 2012. Detection of outlier loci and their utility for fisheries management. Evolutionary Applications, 5(1): 39–52, https://doi.org/10.1111/j.1752-4571.2011.00206.x.
Sang T K, Chang H Y, Chen C T, Hui C F. 1994. Population structure of the Japanese eel, Anguilla japonica. Molecular Biology and Evolution, 11(2): 250–260.
Tesch F W. 1977. The Eel: Biology and Management of Anguillid Eels. Chapman & Hall, London. p.81–240.
Tesch F W. 2003. The Eel. Blackwell Science, Oxford. p.119–212, https://doi.org/10.1002/9780470995389.
Tseng M C, Tzeng W N, Lee S C. 2006. Population genetic structure of the Japanese eel Anguilla japonica in the northwest Pacific Ocean: evidence of non-panmictic populations. Marine Ecology Progress Series, 308: 221–230, https://doi.org/10.3354/meps308221.
Tsukamoto K, Otake T, Mochioka N, Lee T W, Fricke H, Inagaki T, Aoyama J, Ishikawa S, Kimura S, Miller M J, Hasumoto H, Oya M, Suzuki Y. 2003. Seamounts, new moon and eel spawning: the search for the spawning site of the Japanese eel. Environmental Biology of Fishes, 66(3): 221–229
Tsukamoto K. 1990. Recruitment mechanism of the eel, Anguilla japonica, to the Japanese Coast. Journal of Fish Biology, 36(5): 659–671, https://doi.org/10.1111/j.1095-8649.1990.tb04320.x.
Tsukamoto K. 1992. Discovery of the spawning area for Japanese Eel. Nature, 356: 789–791, https://doi.org/10.1038/356789a0.
Tsukamoto K. 2006. Spawning of eels near a seamount. Nature, 439: 929–929, https://doi.org/10.1038/439929a.
Tsukamoto K. 2013. Advances in Aquaculture for the production of artificial glass eel to help conserve Anguillid Eel Populations Worldwide. The University of Tokyo, Japan: Presented at the World Fisheries Congress, Edinburgh.
Tzeng W N. 1990. Relationship between growth rate and age at recruitment of Anguilla japonica elvers in a Taiwan estuary as inferred from otolith growth increments. Marine Biology, 107(1): 75–81, https://doi.org/10.1007/BF01313244.
Ulrik M G, Pujolar J M, Ferchaud A L, Jacobsen M W, Als T D, Gagnaire P A, Frydenberg J, Bøcher P K, Jónsson B, Bernatchez L, Hansen M M. 2014. Do North Atlantic eels show parallel patterns of spatially varying selection?. BMC Evolutionary Biology, 14: 138, https://doi.org/10.1186/1471-2148-14-138.
Van Oosterhout C, Hutchinson W F, Wills D P M, Shipley P. 2004. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes, 4(3): 535–538. https://doi.org/10.1111/j.1471-8286.2004.00684.x.
Acknowledgment
We thank Dr. LI Yulong for his help in data analysis. Thanks Dr. LIU Bingjian, Dr. Yokogawa Koji, and Dr. HAN Yusan from National Taiwan University, China, for their help in collecting the samples.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Natural Science Foundation of China (NSFC) (No. 41676137) and the Creative Team Project of the Laboratory for Marine Ecology and Environmental Science, Qingdao National Laboratory for Marine Science and Technology (No. LMEES-CTSP-2018-1)
Rights and permissions
About this article
Cite this article
Yu, L., Liu, Y. & Liu, J. Gene-associated microsatellite markers confirm panmixia and indicate a different pattern of spatially varying selection in the endangered Japanese eel Anguilla japonica. J. Ocean. Limnol. 38, 1572–1583 (2020). https://doi.org/10.1007/s00343-020-0048-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00343-020-0048-z