Abstract
Microbes in the coral holobiont play important roles in nitrogen fixation, carbon supply, antibiotic production, mucus recycling, and food supply to maintain homeostasis in corals. However, microbes can also induce coral diseases in response to environmental changes under non-optimal conditions. Therefore, studies of microbial communities are needed to understand the health statuses of corals in response to environmental changes. In this study, we performed 16S rDNA metabarcoding to investigate the bacterial communities in two healthy alcyonacean soft coral species (Sinularia sp. and Litophyton sp.) inhabiting the coast of Weno Island (Chuuk, Micronesia) and in ambient seawater. We identified 18 bacterial phyla, 24 classes, 54 orders, 109 families, and 222 genera associated with the two corals and seawater. The bacterial communities differed in the corals and seawater. The bacterial community in Sinularia sp. was dominated by the genus Spirochaeta in Spirochaetaceae (63.9% relative abundance), followed by Endozoicomonas (10%). In Litophyton sp., the bacterial community also contained Spirochaeta (19.5%) and Endozoicomonas (4.7%), although Cellvibrionaceae (23.7%) was dominant and other groups such as Rhizobiales (11.5%) and Rhodospirillales (8.7%) were evenly distributed. In ambient seawater, the predominant bacteria were Pelagibacter (29.2%), Rhodobacteraceae (15.5%), Prochlorococcus (11.3%), and Vibrio (5.8%), which are distinct from the species in the two coral species. The microbial communities between the two alcyonacean soft corals and seawater were different, and the microbial community differences were coral species-specific.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Coral reefs are among the most biologically diverse ecosystems on Earth, providing niches for numerous marine organisms (Blackall et al. 2015). However, coral reefs and their associated organisms have become increasingly vulnerable to disturbances from climate change and pollution caused by human activities in recent years. Particularly, climate change causes coral bleaching, a stress response in corals, resulting in the loss of their symbiotic algae and causing the corals to turn white (Lesser 2007). In some cases, these stress responses lead to the death of the corals and devastation of coral reefs.
Generally, coral-associated microbes maintain coral health by influencing the structure and function of coral holobionts. This includes adaptation of coral to environmental changes and recycling of nutrients such as sulfur, carbon, and nitrogen compounds (Webster and Reusch 2017; Robbins et al. 2019; Pernice et al. 2020). In addition, the health of the corals depends on their specific bacterial colonization (Morrow et al. 2012; Krediet et al. 2013). In general, bacterial communities associated with corals are among the most diverse and complex communities in the microbial biosphere (Sunagawa et al. 2010; Bourne and Webster 2013; Hernandez-Agreda et al. 2016). Coral-related bacterial communities vary in their response based on intrinsic and extrinsic factors, such as the coral species, habitat, and environmental conditions (Thompson et al. 2014), and differ from those in ambient seawater (Sunagawa et al. 2010; Barott et al. 2011; Roder et al. 2014). Specifically, coral metabolism affects the associated bacterial community structure. Environmental factors, such as geographic location, water depth, nutrient concentration, and temperature, also have significant effects (Apprill et al. 2009; Vega Thurber et al. 2009; Littman et al. 2011). Therefore, the relationship between bacterial communities and their hosts or the environment have been described in terms of bacterial diversity associated with corals (Rohwer et al. 2002), host specificity (Ceh et al. 2011), changes in bacterial communities in response to competition between corals and macroalgae (Vega Thurber et al. 2012), and coral disease (Rohwer et al. 2002; Ceh et al. 2011; Vega Thurber et al. 2012; Wilson et al. 2012). However, changes in the temporal and spatial diversity of bacterial communities in response to environmental changes have not been widely studied. In addition, the composition of the microbial community associated with corals and their health in response to environmental changes and human activities are unclear. Therefore, it is important to investigate the composition of coral-associated microbial communities.
The alcyonacean soft coral genera Litophyton and Sinularia are two of the most widely distributed soft corals (Chen et al. 2012; Abou El-Kassem et al. 2018). They are abundant in the tropical western Pacific regions, including the islands of the Federated States of Micronesia (FSM) (GBIF.org 2020). The FSM are surrounded by coral reefs that support rich biodiversity. Thus, these reefs are beneficial to the Micronesian people, supporting their survival by providing food and revenue from fish sales and tourism. In addition, these corals have drawn increased attention because they contain pharmacological compounds and have potential medicinal properties originating from their various secondary metabolites such as sesquiterpenes, diterpenes, polyhydroxylated steroids, and polyamines (Grote et al. 2008; Chen et al. 2012). Although biodiversity remains high within the FSM reefs, there are threats from a variety of human activities, such as coastal development, pollution, tourism, and fishing. Indeed, a Status Report of 2020 on the FSM (Hall 2020) showed that the coastal areas of Chuuk State, FSM, were more affected by rapid population growth, environmental pollution, and climate change than the other islands around Micronesia, such as the Palau, Kiribati, and Marshall Islands. However, few studies have examined the bacterial community associated with the soft corals Sinularia and Litophyton. Understanding the bacterial associations in these two soft corals may provide insight into the relationship between bacteria and coral in the reefs of Chuuk State.
In this study, we analyzed the relative abundance of bacterial taxa in the microbial communities in two different leather soft corals, Litophyton sp. (Palau green tree leather coral) and Sinularia sp. (smooth leathery soft coral), and in the ambient coastal seawater from the Weno Island, Chuuk State, FSM, using the 16S rDNA metabarcoding approach. Overall, the potential functional roles of common bacteria were explored, and their possible functional roles were assessed based on their occurrence by comparing these with the published reports.
Materials and methods
Sample collection and harvest of bacterial communities from two alcyonacean soft corals and ambient seawater
Ten fragments of a single colony in two visually healthy alcyonacean soft coral species, Litophyton sp. and Sinularia sp., were collected from a depth of 3–10 m along the Coast of Weno Island by scuba diving on May 13, 2014 (7°27′22.9″ N, 151°54′21.2″ E, Fig. 1). Each coral subsample was immediately placed in a plastic bag and transported to the ship. On the ship, the coral samples were washed twice with sterilized seawater, placed in sterilized plastic bags, and stored on ice until they were transported to the laboratory. The samples of two alcyonacean soft coral species, Litophyton sp. (No. B_S_HI_00001111) and Sinularia sp. (No. B_S_HI_00001112), were stored in the Library of Marine Samples of the Korea Institute of Ocean Science & Technology. The identification and characterization of these coral species were confirmed using mitochondrial DNA gene cytochrome oxidase I (COI) and phylogenetic analyses (Suppl. Figure 1).
Corals and seawater sampling site, and photographs of the two alcyonacean soft corals. a Red circle shows the location of Chuuk Lagoon in Northeastern Pacific Ocean. b Red circle shows the coral and seawater sampling site (7°27′22.9″ N, 151°54′21.2″ E) near Weno Island in Chuuk Lagoon. c Litophyton sp. d Sinularia sp.
Ambient seawater samples (5 L each, sampling at a depth of 3–10 m using a Niskin water sampler and placed in a sterilized 5-L polyethylene bottle) were collected in duplicate. Bacterial cells on the surface mucus layer of each coral fragment were detached by performing repeated sonication (VibramCell; Sonic & Materials, Inc., Newtown, CT, USA) at 15 A and 4 °C for 5 min in sterilized seawater (Jung et al. 2013). The detached large-sized particles and organic matter were pre-filtrated (or removed) using a 3-μm polycarbonate membrane (TSTP04700; Millipore, Billerica, MA, USA). The bacterial communities were harvested from the flow-through by re-filtering using a 0.2-μm polycarbonate membrane (TTTP04700, Millipore). The filters were washed three times with approximately 50 mL distilled water at 50–60 °C to remove environmental DNA debris (Jung et al. 2018). The bacterial communities from seawater were also harvested using a 0.2-μm polycarbonate membrane. The filters were stored at − 80 °C until genomic DNA (gDNA) extraction.
DNA extraction and 16S rDNA metabarcoding analyses
The filters with the harvested bacterial communities were cut into several pieces for gDNA extraction. gDNA was extracted using a DNeasy PowerSoil Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s instructions; DNA was diluted to a final concentration of 20 ng/μL. The quantity and quality of the total gDNA were determined using a NanoDrop (Nano-MD-NS, SCINCO Ltd., South Korea). The V1–V3 hypervariable regions of the bacterial 16S rDNA genes were amplified using the universal tagged forward (27F: 5′-GAG TTT GAT CMT GGC TCA G-3′) and reverse (518R: 5′-ATT ACC GCG GCT GG-3′) primers (Kim et al. 2016). PCR was performed using ExTaq polymerase (Takara, Shiga, Japan) on a T100 PCR Thermal Cycler (Bio-Rad, Hercules, CA, USA). The PCR protocol consisted of an initial denaturation step at 95 °C for 3 min, followed by 35 cycles of denaturation at 95 °C for 10 s, annealing at 52 °C for 45 s, extension at 72 °C for 1 min, and a final extension step at 72 °C for 5 min. The amplified products were individually purified using a QIAquick PCR Purification Kit (Qiagen). Although we performed duplicate experiments, we attempted to overcome the experimental bias and obtain accurate results by performing three PCRs in distinct tubes and mixing the PCR products to obtain more accurate metabarcoding results (Jung et al. 2018). Prior to obtaining the metabarcoding results, the amplified PCR products were individually purified using an UltraClean PCR Clean-up Kit (MoBio; Folsom, CA, USA). DNA concentrations of the cleaned PCR products were measured using a Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA, USA). Equal concentrations of the PCR products were analyzed using a 454 automated sequencer (GS Junior Sequencing System, Roche, Basel, Switzerland) using titanium chemistry. All sequences obtained from this study have been deposited in the National Center for Biotechnology Information Sequence Read Archive under BioProject accession number PRJNA610452.
Bioinformatics, statistical analyses, and functional annotation of common bacteria
Bioinformatics analyses were performed as described by Kang et al. (2021). The sequences were subjected to a quality check; short sequence reads (length < 150 bp), low-quality sequences (score < 25), singletons and chloroplast sequences, and any non-bacterial ribosome sequences and chimeras were removed using Roche GS20 software (Gontcharova et al. 2010). Using the Basic Local Alignment Search Tool, the sequence reads were compared to the Silva rRNA database (Altschul et al. 1990; Quast et al. 2013). Sequence reads with an E value less than 0.01 were considered as partial 16S rDNA sequences. 16S rDNA sequences with at least 97% similarity were considered for species identification, and their complete taxonomic hierarchy was assigned, which consisted of the phylum, class, order, family, genus, and species.
Alpha diversity indices were estimated by calculating the Chao1, Shannon, and Simpson indices using the R vegan package (Oksanen et al. 2010). All visualizations except Venn diagrams were created in the R ggplot2 package. Venn diagrams were prepared using Venny 2.1.0 (https://bioinfogp.cnb.csic.es/tools/venny/). Operational taxonomic units (OTUs) with an abundance of < 1% in at least one sample were filtered out. The relative abundances of the filtered bacterial OTUs were normalized through logarithmic transformations. Cluster analyses were performed on the data from the filtered bacteria using the group average clustering in the Bray–Curtis similarity method. To test the null hypothesis (no significant difference between the groups discriminated according to the agglomerative clustering analysis), similarities were analyzed with ANOSIM in PRIMER version 6.1.13 (Clarke et al. 2006).
To predict the functional roles of the common bacterial associates, functional annotation of prokaryotic taxa (FAPROTAX) was performed using python collapse_table.py with the normalized OTU table (Louca et al. 2016). The FAPROTAX dataset (available at http://www.zoology.ubc.ca/louca/FAPROTAX) is a manually constructed database based on cultured representatives of marine and freshwater microbiomes. The functional group abundances in each module were calculated by multiplying the calculated values (“function tables”) and the total sum of OTUs belonging to each major module.
Results
Bacterial communities in the two alcyonacean soft corals (Litophyton sp. and Sinularia sp.) and ambient seawater were identified using 16S rDNA metabarcoding. A mean of 6437 reads was retrieved after quality trimming (Table 1), among which 6108 were detected in ambient seawater samples, whereas 7233 and 5971 reads were found in Litophyton sp. and Sinularia sp., respectively. The mean number of observed OTUs with 97% quantified reads was lowest for Sinularia sp. (270), followed by Litophyton sp. (327) and ambient seawater (384). Chao 1, Shannon diversity, and Simpson evenness are presented in Fig. 2a; ambient seawater samples had the highest alpha diversity index (Chao1: mean 597; Shannon: 3.9; Simpson: 0.9), whereas Sinularia sp. had the lowest index (Chao 1: mean 443; Shannon: 2.1; Simpson: 0.6). The Venn diagram indicated that bacterial species in ambient seawater, Litophyton sp., and Sinularia sp. represented 519, 424, and 531 taxa, respectively; in all samples, common bacteria overlapped in 39 taxa (3.3% of the total number of bacteria) (Fig. 2b). Sinularia sp. and Litophyton sp. showed the largest OTU overlap of 178 taxa and Litophyton sp., and ambient seawater showed the smallest OTU overlap of 68 taxa.
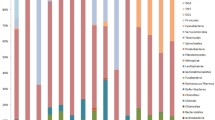
Taxonomic classification revealed 18 bacterial phyla, 24 classes, 54 orders, 109 families, and 222 genera. Except for the most dominant Proteobacteria, which were assigned at the class level, the other bacterial groups were classified at the phylum level (Fig. 3). The next most dominant phylum was Cyanobacteria (13.3%), followed by Bacteroidetes (3.8%). The bacteria associated with Sinularia sp. were dominated by the genus Spirochaeta in Spirochaetaceae, with a mean of 63.2% relative abundance, followed by Gammaproteobacteria (13.0%), Alphaproteobacteria (6.1%), and Cyanobacteria (3.4%). In Litophyton sp., the associated bacteria differed from those of Sinularia sp. and included Gammaproteobacteria (32.1%), Alphaproteobacteria (26.6%), Spirochaetes (19.7%), and Tenericutes (6.2%). In seawater, Alphaproteobacteria and Gammaproteobacteria belonging to the phylum Proteobacteria were dominant, with relative abundances of 57.9% and 22.4%, respectively.
Common bacterial OTUs displaying a mean relative abundance of > 1% in the two corals and ambient seawater included 36 taxa. The mean of the total relative abundance of common OTUs was 85.7% in Sinularia sp., 84.0% in Litophyton sp., and 69.4% in ambient seawater. The bacterial communities from the two corals were clustered with 35% similarity and contained four common bacterial OTUs (OTU1161, OTU1162, OTU1002, and OTU1008); the first two OTUs (Spirochaeta) were assigned to Spirochaetaceae, and the other two OTUs to Endozoicomonas. Interestingly, the common bacterial associates showing > 1% relative abundance were not common between the corals and seawater (Fig. 4). Spirochaeta and Endozoicomonas were predominant in Sinularia sp. with relative abundances of 63.9% and 10.0%, respectively. In addition to the two bacterial groups that were common between the two corals, Litophyon sp. had other associates, such as Cellvibrionaceae (23.7%), Rhizobiales (11.5%), and Rhodospirillales (8.7%). In ambient seawater, the predominant bacterial communities completely differed from those in the two coral species, namely Pelagibacter in SAR11 (29.2%), Rhodobacteraceae (15.5%), Prochlorococcus in Cyanobacteria (11.3%), Vibrio (5.8%), Alteromonadaceae (3.8%), and Oceanospirillaceae (2.5%).
Thirty-six common bacterial OTUs were identified using the pipeline of FAPROTAX to evaluate the potential functional differences among samples (Fig. 5). The functional profiles of bacteria in Sinularia sp. were mainly associated with heterotrophy such as aerobic chemoheterotrophy and chemoheterotrophy, autotrophy such as in Cyanobacteria, oxygenic photoautotrophy and phototrophy, carbon cycling such as fermentation, and nitrogen cycling such as nitrogen fixation. Particularly, chemoheterotrophy and fermentation were the most prominent functions, averaging 48.4% and 41.8% of the total functional categories, respectively. In Litophyton sp., the functional categories predominant were heterotrophy (52.4%; chemoheterotrophy, aerobic chemoheterotrophy, and photoheterotrophy), carbon cycling associated with fermentation (9.0%), nitrogen cycling (24.9%; nitrogen fixation, ureolysis, nitrogen reduction, nitrate respiration, and nitrogen respiration), and autotrophy (1.5%). In ambient seawater, the functional groups of bacteria were autotrophy (33.0%; photoautotrophy, phototrophy, oxygenic phototrophy, Cyanobacteria, and phototrophy), heterotrophy (69.0%; chemoheterotrophy, aerobic chemoheterotrophy, and fermentation), carbon cycling (1.2%; hydrocarbon degradation and aliphatic non-methane hydrocarbon degradation), and nitrogen cycling (11.2%; nitrogen reduction, nitrate respiration, and nitrogen respiration).
Discussion
Generally, coral-associated bacteria communities play crucial roles in coral holobionts and are essential for coral biology by performing complementary metabolism. Therefore, identifying and studying these communities can provide insights into the interactions between coral and microbiota and predict the coral health condition in response to environmental changes. In this study, we compared the bacterial communities in two co-occurring and ecologically important tropical alcyonacean soft coral species, Sinularia sp. and Litophyton sp., with those in ambient seawater. We found that the microbial communities between the two alcyonacean soft corals and seawater were different and observed microbial community differences in coral species-specific (Frias-Lopez et al. 2002; Apprill et al. 2009; Ochsenkühn et al. 2018). Specifically, the microbial community in Sinularia sp. was dominated by the genus Spirochaeta in Spirochaetaceae, followed by Endozoicomonas. In Litophyton sp., the microbial community was dominated by Spirochaeta and Endozoicomonas, but other associates, such as Cellvibrionaceae, Rhizobiales, and Rhodospirillales, were also common. However, in ambient seawater, the predominant bacteria were Pelagibacter, Rhodobacteraceae, Prochlorococcus, and Vibrio, which are distinct from the species in the two alcyonacean coral species. Particularly, Spirochaetaceae was the predominant taxon in approximately half the Sinularia sp. and was present in Litophyton sp. but not in ambient seawater. Generally, some bacteria, such as Spirochaetaceae and Endozoicomonas in Sinularia sp., are frequently found in various corals worldwide, whereas Cellvibrionaceae, Rhizobiales, and Rhodospirillales bacteria in the Litophyton sp. are rarely detected near other corals (Lesser et al. 2018; Weber and Apprill 2020; Rosales et al. 2020). Indeed, previous studies reported that Spirochaetaceae is associated with various corals in different regions, such as the Muricea coral species (Holm and Heidelberg 2016), Anthothela coral species from deep sea (Lawler et al. 2016), Lophelia pertusa from cold water (Kellogg et al. 2009), gorgonians from deep sea (Gray et al. 2011), and Corallium rubrum (van de Water et al. 2016), suggesting that Spirochaetaceae are commonly found in coral reefs including Sinularia sp. and Litophyton sp. Although Spirochaetaceae are a coral symbiont found across different coral species, regions, and water depths, we could not confirm the specific environmental factors that determine the occurrence of these bacteria as a coral association.
Additionally, previous studies reported that Spirochaetes are likely associated with the breakdown of lignocellulose and nitrogen fixation in termite guts (Lilburn et al. 2001; Brune 2014). Although Spirochaetaceae was found in two alcyonacean soft corals, the role of Spirochaetaceae remains unclear, as they have been found in diseased and healthy corals (Frias-Lopez et al. 2002; Sekar et al. 2008; Closek et al. 2014; Ainsworth et al. 2015). Therefore, functional prediction using FAPROTAX and the database of metagenomics of bacterial community-identified ecosystem functions were searched, although the database is not exhaustive. The putative roles of the identified common bacteria are summarized in Table 2. Recently, FAPROTAX and metagenomic sequencing have been developed as powerful tools for predicting metabolic and ecological relevant functions of bacterial communities from 16S rRNA gene sequencing data (Jung et al. 2021; Sansupa et al. 2021). In this study, the functional profiles revealed that differences in the bacterial communities between the two corals and ambient seawater can be attributed to nitrogen functions in coral bacteria and phototrophic functions in seawater.
Generally, distinct bacterial communities in corals may play functional roles in sustaining nutrient cycling and promoting coral acclimatization to environmental changes (Reshef et al. 2006; Ziegler et al. 2017; Bang et al. 2018). The roles of bacterial associations in coral have been estimated in terms of their contribution to symbiont functions, including nitrogen (Rohwer et al. 2002; Lesser et al. 2004; Siboni et al. 2008; Lema et al. 2012; Rädecker et al. 2015), sulfur (Raina et al. 2013; Zhang et al. 2015), and carbon cycling (Baker et al. 2015; van de Water et al. 2018). In addition, they protect the host coral from potential pathogens in ambient seawater by producing antibiotics (Reshef et al. 2006; Ritchie 2006; Shnit-Orland and Kushmaro 2009). Indeed, previous studies reported that coral-associated microbial communities are involved in nitrogen fixation, carbon supply, antibiotic production, mucus recycling, and food supply in corals (Rohwer et al. 2002; Lesser et al. 2004; Wild et al. 2004; Ritchie 2006). For example, the functional characteristics of Spirochetes as coral associations have been studied in the context of nitrogen (Lilburn et al. 2001; Kimes et al. 2010) and carbon fixation (Baker et al. 2015), as well as in chemotactic responses to chemical stimulants (Charon et al. 1992). Recently, van de Water et al. (2016) reported that Spirochetes were abundant in red coral Corallium rubrum and may have a symbiotic function similar to nitrogen-fixing bacteria in tropical regions, indicating that Spirochaetaceae are involved in nitrogen fixation, carbon supply, antibiotic production, mucus recycling, and food supply in corals, which may have important roles in the holobiont health of the two soft corals.
Spirochaetaceae and Endozoicomonas species were found to be common bacterial associates in our two soft corals and are frequently found in shallow waters globally. They are considered as conserved “core” microbial associates (van de Water et al. 2016; Neave et al. 2017; Brener-Raffalli et al. 2018; Kellogg 2019). The core bacteria in coral holobiont may contribute to dimethylsulfoniopropionate (DMSP) breakdown in the sulfur cycle (Raina et al. 2009, 2010). However, this functional role in Endozoicomonas has been questioned because of the lack of genes related to DMSP metabolic pathways (Neave et al. 2017). Recently, Tandon et al. (2020) corroborated the functional role of Endozoicomonas in sulfur cycles by confirming the presence of DMSP degradation-related genes. A genomic study of Endozoicomonas revealed functional genes related to the transport of molecules that potentially influence the transfer of organic molecules between diverse animal hosts and the bacteria (Neave et al. 2014, 2017). Therefore, Endozoicomonas bacteria species play important roles in the coral sulfur cycle in the two alcyonacean soft corals. In addition to Spirochaetaceae and Endozoicomonas, Litophyton sp. also has other common bacterial associates, including Cellvibrionaceae, Rhizobiales, and Rhodospirillales. Cellvibrionaceae are marine bacteria found in surface seawater (Lucena et al. 2020) and as a symbiont in the gills of mollusks (Spring et al. 2015; Brito et al. 2018). Cellvibrionaceae has also been reported from other corals, such as Porites astreoides (Weber and Apprill 2020) and stony coral (Rosales et al. 2020), although they were not abundant. Most species in this family possess a large variety of polysaccharide-degrading enzymes (Spring et al. 2015; Lucena et al. 2020). The abundant presence of Cellvibrionaceae in Litophyton sp. may contribute to carbon cycling through polysaccharide degradation. Rhizobiales and Rhodospirillales are diazotrophic bacteria that contribute to nitrogen fixation (Lodwig et al. 2003; Lesser et al. 2018). These bacteria are frequently found in other corals in a high relative abundance (Lesser et al. 2018; Rosales et al. 2020). Widely known as diazotrophs, the functional role of Rhizobiales and Rhodospirillales in corals may be related to nitrogen cycling (Lema et al. 2012; Olson and Lesser 2013). In addition, Rhizobiales are potential causative agents for stony coral tissue loss disease (Rosales et al. 2020). However, the Rhizobiales in Litophyton sp. may play functional roles in nitrogen cycling and not as disease-causing agents, as we observed no lesions in this study. Taken together, the dominant Spirochaetaceae bacteria may contribute to nitrogen and carbon cycling and Endozoicomonas to sulfur cycling in the coral host Sinularia sp. In addition, in Litophyton sp., Spirochaetaceae, Rhizobiales, and Rhodospirillales may play functional roles in nitrogen fixation, Endozoicomonas in sulfur cycling, and Cellvibrionaceae in carbon cycling.
We found that the microbial communities in ambient seawater significantly differ from those in the two corals, as they were dominated by Pelagibacter, Prochlorococcus, Rhodobacteraceae, and Vibrio bacteria. The bacteria in the ambient seawater of Chuuk, FSM, were similar to those in our previous studies (Suh et al. 2014). Generally, Proteobacteria, Bacteroidetes, and Cyanobacteria are common bacteria in the ocean environment, and the most common genera are Pelagibacter, Roseovarius, Prochlorococcus, and Vibrio. In particular, Pelagibacter and Roseovarius are most abundant in the oligotrophic ocean environment (Morris et al. 2002). Prochlorococcus is the smallest solitary Cyanobacteria and most abundant picophytoplankton in tropical waters (Partensky et al. 1999), dominating nutrient-depleted tropical regions (Biller et al. 2015). The occurrence of these oligotrophic bacterial associates matches well with the oligotrophic seawater conditions in Chuuk, suggesting that seawater-associated bacteria in Chuuk are widely distributed in the ocean environment, particularly in nutrient-depleted tropical regions. We also confirmed that Vibrio spp. is common in seawater. Generally, Vibrio species are widely distributed in aquatic environments and have been extensively studied in several coral diseases (Mouchka et al. 2010; Sweet et al. 2014). Therefore, its presence in the ambient seawater indicates that the corals can be infected by opportunistic outbreaks of these potential pathogenic bacteria following environmental changes caused by anthropogenic pollution and climate changes. However, these potential pathogenic bacteria are only rarely present in the corals, possibly because of the defense mechanisms used by corals (van de Water et al. 2018). In general, coral metabolites play a role in defending against infection by potential pathogenic bacteria via inhibition of the growth and attachment of pathogens (Gochfeld et al. 2015). In addition, the absence of Vibrio sp. in the corals’ microbiome may result from the defense mechanism, such as shedding of the surface mucus layer, despite the presence of potentially pathogenic Vibrio sp. in ambient seawater. Taken together, the bacteria community may play an important role in the defense mechanisms in the two alcyonacean soft corals.
In conclusion, the two alcyonacean soft corals (Litophyton sp. and Sinularia sp.) from Chuuk, Micronesia, have different microbial associates, and the most common bacterial associates are related to the biogeochemical functional roles of the coral holobiont. This study improves the understanding of the composition and functions of the bacterial communities in the two corals and provides a foundation for further investigating the health status of the two corals in response to environmental changes in the Chuuk State.
References
Abou El-Kassem LT, Hawas UW, El-Desouky SK, Al-Farawati R (2018) Sesquiterpenes from the Saudi Red Sea: Litophyton arboreum with their cytotoxic and antimicrobial activities. Z Naturforsch C J Biosci 73:9–14
Ainsworth TD, Krause L, Bridge T, Torda G, Raina J-B, Zakrzewski M, Gates RD, Padilla-Gamiño JL, Spalding HL, Smith C, Woolsey ES, Bourne DG, Bongaerts P, Hoegh-Guldberg O, Leggat W (2015) The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J 9:2261–2274
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Apprill A, Marlow HQ, Martindale MQ, Rappe MS (2009) The onset of microbial associations in the coral Pocillopora meandrina. ISME J 3:685–699
Baker BJ, Lazar CS, Teske AP, Dick GJ (2015) Genomic resolution of linkages in carbon, nitrogen, and sulfur cycling among widespread estuary sediment bacteria. Microbiome 3:14
Bang C, Dagan T, Deines P, Dubilier N, Duschl WJ, Fraune S, Hentschel U, Hirt H, Hülter N, Lachnit T, Picazo D, Pita L, Pogoreutz C, Rädecker N, Saad MM, Schmitz RA, Schulenburg H, Voolstra CR, Weiland-Bräuer N, Ziegler M, Bosch TCG (2018) Metaorganisms in extreme environments: do microbes play a role in organismal adaptation? Zoology 127:1–19
Barott KL, Rodriguez-Brito B, Janouskovec J, Marhaver KL, Smith JE, Keeling P, Rohwer FL (2011) Microbial diversity associated with four functional groups of benthic reef algae and the reef-building coral Montastraea annularis. Environ Microbiol 13:1192–1204
Biller SJ, Berube PM, Lindell D, Chisholm SW (2015) Prochlorococcus: the structure and function of collective diversity. Nat Rev Microbiol 13:13–27
Blackall LL, Wilson B, van Oppen MJH (2015) Coral—the world’s most diverse symbiotic ecosystem. Mol Ecol 24:5330–5347
Bourne DG, Webster NS (2013) Coral reef bacterial communities. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The Prokaryotes. Springer, Berlin, Heidelberg, pp 163–187
Brener-Raffalli K, Clerissi C, Vidal-Dupiol J, Adjeroud M, Bonhomme F, Pratlong M, Aurelle D, Mitta G, Toulza E (2018) Thermal regime and host clade, rather than geography, drive Symbiodinium and bacterial assemblages in the scleractinian coral Pocillopora damicornis sensu lato. Microbiome 6:39
Brito TL, Campos AB, Bastiaan von Meijenfeldt FA, Daniel JP, Ribeiro GB, Silva GGZ, Wilke DV, de Moraes DT, Dutilh BE, Meirelles PM, Trindade-Silva AE (2018) The gill-associated microbiome is the main source of wood plant polysaccharide hydrolases and secondary metabolite gene clusters in the mangrove shipworm Neoteredo reynei. PLoS ONE 13:e0200437
Brune A (2014) Symbiotic digestion of lignocellulose in termite guts. Nat Rev Microbiol 12:168–180
Ceh J, Van Keulen M, Bourne DG (2011) Coral-associated bacterial communities on Ningaloo Reef, Western Australia. FEMS Microbiol Ecol 75:134–144
Charon NW, Greenberg EP, Koopman MBH, Limberger RJ (1992) Spirochete chemotaxis, motility, and the structure of the spirochetal periplasmic flagella. Res Microbiol 143:597–603
Chen W-t, Li Y, Guo Y-w (2012) Terpenoids of Sinularia soft corals: chemistry and bioactivity. Acta Pharm Sin B 2:227–237
Clarke KR, Clarke K, Gorley K, Clarke K, Gorley R (2006) PRIMER v6: user manual/tutorial
Closek CJ, Sunagawa S, DeSalvo MK, Piceno YM, DeSantis TZ, Brodie EL, Weber MX, Voolstra CR, Andersen GL, Medina M (2014) Coral transcriptome and bacterial community profiles reveal distinct Yellow Band Disease states in Orbicella faveolata. Isme j 8:2411–2422
Frias-Lopez J, Zerkle AL, Bonheyo GT, Fouke BW (2002) Partitioning of Bacterial Communities between Seawater and Healthy, Black Band Diseased, and Dead Coral Surfaces. Appl Environ Microbiol 68:2214
GBIF.org (2020) GBIF Home Page
Gochfeld DJ, Ankisetty S, Slattery M (2015) Proteomic profiling of healthy and diseased hybrid soft corals Sinularia maxima × S. polydactyla. Dis Aquat Organ 116:133–141
Gontcharova V, Youn E, Wolcott RD, Hollister EB, Gentry TJ, Dowd SE (2010) Black Box Chimera Check (B2C2): A windows-based software for batch depletion of chimeras from bacterial 16S rRNA gene datasets. Open Microbiol J 4:47–52
Gray MA, Stone RP, McLaughlin MR, Kellogg CA (2011) Microbial consortia of gorgonian corals from the Aleutian islands. FEMS Microbiol Ecol 76:109–120
Grote D, Dahse H-M, Seifert K (2008) Furanocembranoids from the soft corals Sinularia asterolobata and Litophyton arboreum. Chem Biodivers 5:2449–2456
Hall I (2020) The Federated States of Micronesia: Sixth national report to the convention on biological diversity. United Nations Environment Programme and the Global Environment Facility, The Federated States of Micronesia
Hernandez-Agreda A, Leggat W, Bongaerts P, Ainsworth TD (2016) The microbial signature provides insight into the mechanistic basis of coral success across reef habitats. mBio 7:e00560-00516
Holm JB, Heidelberg KB (2016) Microbiomes of Muricea californica and M. fruticosa: Comparative analyses of two co-occurring eastern Pacific octocorals. Front Microbiol 7:917
Jung SW, Kang Y-H, Baek SH, Lim D, Han M-S (2013) Biological control of Stephanodiscus hantzschii (Bacillariophyceae) blooms in a field mesocosm by the immobilized algicidal bacterium Pseudomonas fluorescens HYK0210-SK09. J Appl Phycol 25:41–50
Jung SW, Kang D, Kim HJ, Shin HH, Park JS, Park SY, Lee TK (2018) Mapping distribution of cysts of recent dinoflagellate and Cochlodinium polykrikoides using next-generation sequencing and morphological approaches in South Sea. Korea Sci Rep 8:7011
Jung SW, Kang J, Park JS, Joo HM, Suh SS, Kang D, Lee TK, Kim HJ (2021) Dynamic bacterial community response to Akashiwo sanguinea (Dinophyceae) bloom in indoor marine microcosms. Sci Rep 11:6983
Kang J, Park JS, Jung SW, Kim H-J, Joo HM, Kang D, Seo H, Kim S, Jang M-C, Lee K-W, Jin OhS, Lee S, Lee T-K (2021) Zooming on dynamics of marine microbial communities in the phycosphere of Akashiwo sanguinea (Dinophyta) blooms. Mol Ecol 30:207–221
Kellogg CA (2019) Microbiomes of stony and soft deep-sea corals share rare core bacteria. Microbiome 7:90
Kellogg CA, Lisle JT, Galkiewicz JP (2009) Culture-independent characterization of bacterial communities associated with the cold-water coral Lophelia pertusa in the northeastern Gulf of Mexico. Appl Environ Microbiol 75:2294–2303
Kim HJ, Jung SW, Lim D-I, Jang M-C, Lee T-K, Shin K, Ki J-S (2016) Effects of temperature and nutrients on changes in genetic diversity of bacterioplankton communities in a semi-closed bay, South Korea. Mar Pollut Bull 106:139–148
Kimes NE, Van Nostrand JD, Weil E, Zhou J, Morris PJ (2010) Microbial functional structure of Montastraea faveolata, an important Caribbean reef-building coral, differs between healthy and yellow-band diseased colonies. Environ Microbiol 12:541–556
Krediet CJ, Ritchie KB, Paul VJ, Teplitski M (2013) Coral-associated micro-organisms and their roles in promoting coral health and thwarting diseases. Proc Royal Soc 280:20122328
Lawler SN, Kellogg CA, France SC, Clostio RW, Brooke SD, Ross SW (2016) Coral-associated bacterial diversity is conserved across two deep-sea Anthothela species. Front Microbiol 7:458
Lema KA, Willis BL, Bourne DG (2012) Corals form characteristic associations with symbiotic nitrogen-fixing bacteria. Appl Environ Microbiol 78:3136
Lesser MP (2007) Coral reef bleaching and global climate change: Can corals survive the next century? Proc Natl Acad Sci USA 104:5259–5260
Lesser MP, Mazel CH, Gorbunov MY, Falkowski PG (2004) Discovery of symbiotic nitrogen-fixing cyanobacteria in corals. Science 305:997–1000
Lesser MP, Morrow KM, Pankey SM, Noonan SHC (2018) Diazotroph diversity and nitrogen fixation in the coral Stylophora pistillata from the Great Barrier Reef. ISME J 12:813–824
Lilburn TG, Kim KS, Ostrom NE, Byzek KR, Leadbetter JR, Breznak JA (2001) Nitrogen fixation by symbiotic and free-living Spirochetes. Science 292:2495
Littman R, Willis BL, Bourne DG (2011) Metagenomic analysis of the coral holobiont during a natural bleaching event on the Great Barrier Reef. Environ Microbiol Rep 3:651–660
Lodwig EM, Hosie AHF, Bourdès A, Findlay K, Allaway D, Karunakaran R, Downie JA, Poole PS (2003) Amino-acid cycling drives nitrogen fixation in the legume–Rhizobium symbiosis. Nature 422:722–726
Louca S, Parfrey LW, Doebeli M (2016) Decoupling function and taxonomy in the global ocean microbiome. Science 353:1272–1277
Lucena T, Arahal DR, Sanz-Sáez I, Acinas SG, Sánchez O, Aznar R, Pedrós-Alió C, Pujalte MJ (2020) Thalassocella blandensis gen. nov., sp. nov., a novel member of the family Cellvibrionaceae. Int J Syst Evol Microbiol 70:1231–1239
Morris RM, Rappé MS, Connon SA, Vergin KL, Siebold WA, Carlson CA, Giovannoni SJ (2002) SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420:806–810
Morrow KM, Moss AG, Chadwick NE, Liles MR (2012) Bacterial associates of two Caribbean coral species reveal species-specific distribution and geographic variability. Appl Environ Microbiol 78:6438–6449
Mouchka ME, Hewson I, Harvell CD (2010) Coral-associated bacterial assemblages: current knowledge and the potential for climate-driven impacts. Integr Comp Biol 50:662–674
Neave MJ, Michell CT, Apprill A, Voolstra CR (2014) Whole-genome sequences of three symbiotic Endozoicomonas bacteria. Genome Announc 2:e00802-00814
Neave MJ, Michell CT, Apprill A, Voolstra CR (2017) Endozoicomonas genomes reveal functional adaptation and plasticity in bacterial strains symbiotically associated with diverse marine hosts. Sci Rep 7:40579
Ochsenkühn MA, Schmitt-Kopplin P, Harir M, Amin SA (2018) Coral metabolite gradients affect microbial community structures and act as a disease cue. Commun Biol 1:184
Oksanen J, Blanchet FG, Kindt R, Legendre P, O’hara R, Simpson GL, Solymos P, Stevens MHH, Wagner H (2010) Vegan: Community ecology package. R package version 1.17–4. URL http://CRANR-project.org/package=vegan
Olson ND, Lesser MP (2013) Diazotrophic diversity in the Caribbean coral, Montastraea cavernosa. Arch Microbiol 195:853–859
Partensky F, Hess WR, Vaulot D (1999) Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63:106–127
Pernice M, Raina J-B, Rädecker N, Cárdenas A, Pogoreutz C, Voolstra CR (2020) Down to the bone: the role of overlooked endolithic microbiomes in reef coral health. ISME J 14:325–334
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596
Rädecker N, Pogoreutz C, Voolstra CR, Wiedenmann J, Wild C (2015) Nitrogen cycling in corals: the key to understanding holobiont functioning? Trends Microbiol 23:490–497
Raina J-B, Tapiolas D, Willis BL, Bourne DG (2009) Coral-associated bacteria and their role in the biogeochemical cycling of sulfur. Appl Environ Microbiol 75:3492–3501
Raina J-B, Dinsdale EA, Willis BL, Bourne DG (2010) Do the organic sulfur compounds DMSP and DMS drive coral microbial associations? Trends Microbiol 18:101–108
Raina J-B, Tapiolas DM, Forêt S, Lutz A, Abrego D, Ceh J, Seneca FO, Clode PL, Bourne DG, Willis BL, Motti CA (2013) DMSP biosynthesis by an animal and its role in coral thermal stress response. Nature 502:677–680
Reshef L, Koren O, Loya Y, Zilber-Rosenberg I, Rosenberg E (2006) The coral probiotic hypothesis. Environ Microbiol 8:2068–2073
Ritchie KB (2006) Regulation of microbial populations by coral surface mucus and mucus-associated bacteria. Mar Ecol Prog Ser 322:1–14
Robbins SJ, Singleton CM, Chan CX, Messer LF, Geers AU, Ying H, Baker A, Bell SC, Morrow KM, Ragan MA, Miller DJ, Forêt S, Ball E, Beeden R, Berumen M, Aranda M, Ravasi T, Bongaerts P, Hoegh-Guldberg O, Cooke I, Leggat B, Sprungala S, Fitzgerald A, Shang C, Lundgren P, Fyffe T, Rubino F, van Oppen M, Weynberg K, Robbins SJ, Singleton CM, Xin Chan C, Messer LF, Geers AU, Ying H, Baker A, Bell SC, Morrow KM, Ragan MA, Miller DJ, Foret S, Voolstra CR, Tyson GW, Bourne DG, Voolstra CR, Tyson GW, Bourne DG, ReFuGe C (2019) A genomic view of the reef-building coral Porites lutea and its microbial symbionts. Nat Microbiol 4:2090–2100
Roder C, Arif C, Daniels C, Weil E, Voolstra CR (2014) Bacterial profiling of white plague disease across corals and oceans indicates a conserved and distinct disease microbiome. Mol Ecol 23:965–974
Rohwer F, Seguritan V, Azam F, Knowlton N (2002) Diversity and distribution of coral-associated bacteria. Mar Ecol Prog Ser 243:1–10
Rosales SM, Clark AS, Huebner LK, Ruzicka RR, Muller EM (2020) Rhodobacterales and Rhizobiales are associated with stony coral tissue loss disease and its suspected sources of transmission. Front Microbiol 11:681
Sansupa C, Wahdan SF, Hossen S, Disayathanoowat T, Wubet T, Purahong W (2021) Can we use functional annotation of prokaryotic taxa (FAPROTAX) to assign the ecological functions of soil bacteria? Appl Sci 11:688
Sekar R, Kaczmarsky LT, Richardson LL (2008) Microbial community composition of black band disease on the coral host Siderastrea siderea from three regions of the wider Caribbean. Mar Ecol Prog Ser 362:85–98
Shnit-Orland M, Kushmaro A (2009) Coral mucus-associated bacteria: A possible first line of defense. FEMS Microbiol Ecol 67:371–380
Siboni N, Ben-Dov E, Sivan A, Kushmaro A (2008) Global distribution and diversity of coral-associated Archaea and their possible role in the coral holobiont nitrogen cycle. Environ Microbiol 10:2979–2990
Spring S, Scheuner C, Göker M, Klenk H-P (2015) A taxonomic framework for emerging groups of ecologically important marine gammaproteobacteria based on the reconstruction of evolutionary relationships using genome-scale data. Front Microbiol 6:281
Suh SS, Park M, Hwang J, Lee S, Chung Y, Lee TK (2014) Distinct patterns of marine bacterial communities in the South and North Pacific Oceans. J Microbiol 52:834–841
Sunagawa S, Woodley CM, Medina M (2010) Threatened corals provide underexplored microbial habitats. PLoS ONE 5:e9554
Sweet MJ, Croquer A, Bythell JC (2014) Experimental antibiotic treatment identifies potential pathogens of white band disease in the endangered Caribbean coral Acropora cervicornis. Proc Royal Soc 281:20140094
Tandon K, Lu C-Y, Chiang P-W, Wada N, Yang S-H, Chan Y-F, Chen P-Y, Chang H-Y, Chiou Y-J, Chou M-S, Chen W-M, Tang S-L (2020) Comparative genomics: Dominant coral-bacterium Endozoicomonas acroporae metabolizes dimethylsulfoniopropionate (DMSP). ISME J 14:1290–1303
Thompson JR, Rivera HE, Closek CJ, Medina M (2014) Microbes in the coral holobiont: Partners through evolution, development, and ecological interactions. Front Cell Infect Microbiol 4:176
van de Water JA, Melkonian R, Junca H, Voolstra CR, Reynaud S, Allemand D, Ferrier-Pages C (2016) Spirochaetes dominate the microbial community associated with the red coral Corallium rubrum on a broad geographic scale. Sci Rep 6:27277
van de Water JAJM, Allemand D, Ferrier-Pagès C (2018) Host-microbe interactions in octocoral holobionts - recent advances and perspectives. Microbiome 6:64
Vega Thurber R, Willner-Hall D, Rodriguez-Mueller B, Desnues C, Edwards RA, Angly F, Dinsdale E, Kelly L, Rohwer F (2009) Metagenomic analysis of stressed coral holobionts. Environ Microbiol 11:2148–2163
Vega Thurber R, Burkepile DE, Correa AM, Thurber AR, Shantz AA, Welsh R, Pritchard C, Rosales S (2012) Macroalgae decrease growth and alter microbial community structure of the reef-building coral Porites astreoides. PLoS ONE 7:e44246
Weber L, Apprill A (2020) Diel, daily, and spatial variation of coral reef seawater microbial communities. PLoS ONE 15:e0229442
Webster NS, Reusch TBH (2017) Microbial contributions to the persistence of coral reefs. ISME J 11:2167–2174
Wiegel JKW (2015) Xanthobacter Bergey's manual of systematics of archaea and bacteria. Wiley, Chichester, pp 1–22
Wild C, Huettel M, Klueter A, Kremb SG, Rasheed MYM, Jørgensen BB (2004) Coral mucus functions as an energy carrier and particle trap in the reef ecosystem. Nature 428:66–70
Wilson B, Aeby GS, Work TM, Bourne DG (2012) Bacterial communities associated with healthy and Acropora white syndrome-affected corals from American Samoa. FEMS Microbiol Ecol 80:509–520
Zhang Y, Ling J, Yang Q, Wen C, Yan Q, Sun H, Van Nostrand JD, Shi Z, Zhou J, Dong J (2015) The functional gene composition and metabolic potential of coral-associated microbial communities. Sci Rep 5:16191
Ziegler M, Seneca FO, Yum LK, Palumbi SR, Voolstra CR (2017) Bacterial community dynamics are linked to patterns of coral heat tolerance. Nat Commun 8:14213
Acknowledgements
The genomic DNA samples of corals and seawaters were obtained from the Library of Marine Samples, Korea Institute of Ocean Science & Technology, Republic of Korea. This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science & ICT (NRF-2017M3A9E4072753) and entitled “Development of technology for mass production of useful marine bioproducts” by the Research Program of Korea Institute of Ocean Science and Technology (PE99922).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
There are no conflicts of interests to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Topic Editor Morgan S. Pratchett
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Park, J.S., Han, J., Suh, SS. et al. Characterization of bacterial community structure in two alcyonacean soft corals (Litophyton sp. and Sinularia sp.) from Chuuk, Micronesia. Coral Reefs 41, 563–574 (2022). https://doi.org/10.1007/s00338-021-02176-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-021-02176-w