Abstract
Diseases of marine organisms, including sponges on coral reefs, are being reported with increasing frequency worldwide. Aplysina Red Band Syndrome (ARBS) occurs across the Caribbean basin, predominantly affecting Aplysina cauliformis, one of the most common reef sponges in this region. Existing ARBS lesions and their effects on the sponge holobiont have been documented, yet little is known about the biochemical and microbial changes associated with the onset of infection. Due to the transmissible nature of ARBS, infection can be induced and monitored through sponge-to-sponge direct contact. Nine-day contact experiments with healthy-diseased and healthy-healthy sponge pairings were conducted in the Bahamas in January and July to compare individual sponges sampled initially and at one of three successive time points. Temporal changes in bacterial assemblages and photosymbiont abundance (via concentrations of chlorophyll a), and concentrations of total protein, heat shock protein 70, and major secondary metabolites that may correspond with disease onset were characterized. All healthy sponges in contact with diseased sponges developed ARBS by day 9 in January and by day 6 in July, suggesting that observed changes in the holobiont corresponded with the development of ARBS. The concentrations of several major secondary metabolites, as well as heat shock protein 70 and chlorophyll a, changed significantly in samples of visibly healthy tissue from initially healthy sponges that became diseased. In contrast, the composition of the associated bacterial community changed in all attached sponges over time. These results suggest that infection with ARBS elicits rapid responses by the sponge holobiont, providing a model system in which to investigate immune responses in an early metazoan.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diseases of marine organisms have been reported with increasing frequency in recent decades, contributing to a dramatic decline in marine ecosystem services worldwide (e.g., Ward and Lafferty 2004; Mydlarz et al. 2006; Webster 2007; Hewson et al. 2014; Lafferty and Hofmann 2016; Precht et al. 2016; Walton et al. 2018). To date, studies of diseases affecting sessile marine invertebrates have primarily been descriptive, and predominantly focused on scleractinian corals due to their roles as “ecosystem engineers” (Jones et al. 1997). However, in many cases, the physiological and ecological impacts of marine diseases are poorly understood. For example, research on coral diseases has concentrated largely on describing their physical manifestations, outbreak characteristics, relationships with other environmental stressors, and identifying putative etiologic agents. As a result, modes of transmission and physiological impacts on individuals in the early stages of pathogenesis for most marine diseases remain unknown, in part due to the difficulties of identifying transmission pathways in the ocean, particularly when pathogens have not been identified or are unculturable. As might be expected in the marine environment, modes of disease transmission typically include water-borne, vector-borne and/or direct contact (reviewed in Shore and Caldwell 2019).
Sponges represent an immense source of biodiversity and biomass on coral reefs (van Soest et al. 2012), and provide numerous ecosystem services, ranging from habitat and food for other organisms to hosting diverse microbial assemblages that serve a plethora of functional roles in terms of nutrient cycling (Diaz and Rützler 2001; Wulff 2012; de Goeij et al. 2013), and producing a rich source of secondary metabolites with both ecological and potential pharmaceutical applications (Mehbub et al. 2014; Paul et al. 2019). Whereas sponge diseases have not been as well described as coral diseases, they are also being reported worldwide with increasing frequency (reviewed by Webster 2007; Luter and Webster 2017). However, little is known about how sponge diseases are transmitted between individuals or how the holobiont responds to disease. Because sponges do not leave a skeletal record when they die, in the absence of an observed epizootic event, sponge mortality due to disease tends to be patchy and easily overlooked. Nonetheless, sublethal effects of sponge diseases can affect their ecological interactions, as well as the ecosystem services they provide.
Aplysina Red Band Syndrome (ARBS) is a disease affecting Caribbean sponges within the genus Aplysina, particularly Aplysina cauliformis, the purple rope sponge (Olson et al. 2006). ARBS is characterized by the presence of an initial rust to red-colored circular lesion that spreads outward along the sponge branch, leaving behind spongin fibers devoid of live tissue that become colonized by epibiotic algae and microorganisms (Olson et al. 2006; Gochfeld et al. 2019). The characteristic red coloration of the lesion is due to the presence of the filamentous cyanobacterium Leptolyngbya sp. (Olson et al. 2006, 2014; Gochfeld et al. 2019) although transmission studies with a sponge-derived isolate failed to establish disease (Olson et al. 2014). Of the sponge diseases described to date (Webster 2007; Luter and Webster 2017), only ARBS has been shown to be readily transmissible via direct contact in situ (Easson et al. 2013) and experimentally, with signs of disease in healthy sponges placed in contact with diseased sponges visible within 7 days (Olson et al. 2006). Water-borne transmission of ARBS has also been documented, although it occurs less frequently (Olson et al. 2006; Easson et al. 2013). The rapid and reliable contact-driven transmission of ARBS to healthy sponges provides an unprecedented opportunity to characterize responses of the sponge holobiont during the course of pathogenesis.
ARBS has diverse sublethal effects on the sponge holobiont, including changes to the composition of the associated bacterial community, reductions in the number of cyanobacterial photosymbionts, changes in host cell densities and cellular composition, reduced protein content, changes in secondary metabolite profiles, reduced growth rates, and an increased probability of breakage during storm events (Gochfeld et al. 2012a, b, 2019; Easson et al. 2013; Olson et al. 2014). However, these effects have been described in sponges with established ARBS lesions and do not provide information regarding impacts of the initiation of ARBS infection on the sponge holobiont. To better understand the process of disease onset (i.e., before and once early signs of disease are apparent), we performed contact transmission experiments to characterize the composition of sponge-associated bacterial communities, and to measure the concentrations of chlorophyll a (as a proxy for the sponges’ cyanobacterial symbionts), soluble protein, heat shock protein 70 (as a marker for sponge stress: López-Legentil et al. 2008; Fan et al. 2013), and major secondary metabolites in A. cauliformis exposed to ARBS for different periods of time, relative to ARBS-affected and healthy in situ sponges. Seasonal differences in rates of disease transmission and its impacts were also evaluated. These results demonstrate how rapidly changes occur within the sponge holobiont as disease becomes established and create a baseline for additional investigations into marine invertebrate responses to pathogenesis.
Materials and methods
Experimental design and sample collection
To test for seasonal differences in rate of disease transmission and impacts on the sponge holobiont, transmission experiments were conducted in July 2010 and January 2011. Temperature loggers (HOBO, Onset Computer Corp.) were deployed at the experimental site and data were collected every 10 min for the duration of each experiment. Based on initial results and available resources, some of the time points and measured variables were revised between experiments.
On day 0, scissors were used to cut approximately 25 cm long branches from visibly healthy (i.e., no signs of discoloration or damage) A. cauliformis individuals at a depth of 5 m on North Norman’s reef (23° 47.388′ N, 76° 08.273′W), Exuma Cays, Bahamas. Approximately half of each healthy sponge was placed into an individually labeled 1 gal re-sealable plastic bag for transport to the wet laboratory at the Perry Institute for Marine Science (PIMS) on Lee Stocking Island for collection of initial (Day 0) samples. The other half of each healthy sponge branch was cable-tied to either a tagged ‘in situ Healthy’ A. cauliformis or in contact with the red band on a tagged ‘in situ ARBS’ affected A. cauliformis (Fig. 1a, b). All diseased sponges were photographed with a Canon PowerShot D20 underwater digital camera prior to attaching the healthy sponge, and all sponge pairs were photographed following attachment.
Portions of initially healthy sponges were attached to a in situ healthy or b in situ ARBS-affected sponges using cable ties. Portions of the attached sponges were sampled initially, and final samples of each pair of attached and in situ sponges were collected at a later date (days 3, 6, or 9 in July; days 1, 3 or 9 in January). When sponge pairs were separated in the laboratory following 1–9 days of contact, the point of contact c for pairs of healthy sponges displayed raised edges where the sponges began to grow together, whereas d initially healthy sponges attached to ARBS-affected sponges began to develop a red band around the margin of contact and displayed necrotic tissue
In the laboratory, the collected half of each sponge that was attached to an in situ healthy or in situ ARBS-affected sponge (referred to as Attached Healthy-Initial or Attached ARBS-Initial, respectively) was photographed in water, and several pie-shaped wedges from small cross-sections (~ 1 mm thick) cut from each branch were preserved in cryovials containing 1.8 ml RNAlater™ for analysis of bacterial communities. During the January experiment, subsamples for analysis of heat shock protein 70 (Hsp70) concentrations were collected and placed into microcentrifuge tubes containing Hsp sample buffer (see below). In both experiments, the remainder of each sponge branch was cut into three pieces, and each piece was wrapped in aluminum foil, placed into a labeled plastic bag, and frozen at − 40 °C until concentrations of chlorophyll a, soluble protein, and secondary metabolites, respectively, were determined. Samples were transported to the University of Alabama (bacterial communities) or University of Mississippi (chlorophyll a, protein, Hsp70, and secondary metabolites) for analysis.
In July 2010, 7 pairs from each sample condition group (Attached Healthy-Final–in situ Healthy and Attached ARBS-Final–in situ ARBS) were randomly selected for collection after 3, 6, and 9 days of contact (total of 42 sponges: 21 attached and 21 in situ). However, in the Bahamas, there are two recognized morphologies of A. cauliformis (Zea et al. 2014), one characterized as a thin purple (“lilac creeping”) form that does not release an exudate when handled, and a thicker (“brown erect”) morph that releases a blue exudate upon handling (Gochfeld, pers. obs.). Because differences in chemistry have been found between the two morphs (Puyana et al. 2015; Stockton 2016), all sponges of the brown erect morph were removed from subsequent analyses for the July experiment, resulting in different numbers of sponges sampled at each time point and reducing the statistical power for this experiment (n = 4–7 sponges per sample group; Supplemental Table 1). In January 2011, 8 pairs from each sample condition group were randomly selected for collection after 1, 3, and 9 days of contact (total of 48 sponges). Only the lilac creeping morph was used in the January experiment (n = 7–8 sponges per sample group with minor losses due to wave action; Supplemental Table 1).
After obtaining in situ photographs, each cable-tied pair of sponges was removed from the reef and placed into a labeled re-sealable plastic bag for transport to PIMS. In the laboratory, each sponge pair was photographed in a shallow water bath while still in contact and following careful separation. From visibly healthy tissue near (~ 1 cm distance) the point of contact, samples were collected from the attached sponges (referred to as Attached Healthy-Final or Attached ARBS-Final) and preserved as described previously. This experimental design generated two samples from the same individual sponges, one taken just prior to attachment to an in situ sponge (Attached Healthy-Initial or Attached ARBS-Initial) and one after 1–9 days of attachment (Attached Healthy-Final or Attached ARBS-Final). At the same time that the final sample was taken from the attached sponges, samples of visibly healthy tissue from the in situ Healthy and in situ ARBS-affected sponges were also collected ~ 1 cm from the point of contact, and preserved as described above. All samples included portions of the outer pinacoderm and interior mesohyl.
DNA extraction and PCR for bacterial community analyses
Using the FastDNA Spin Kit for Soil (MP Biomedicals), total genomic DNA was extracted from RNAlater™ preserved sponge samples following the manufacturer’s protocol, with the lysis step extended to an overnight incubation at 37 °C with shaking. To amplify the 16S rRNA gene by polymerase chain reaction (PCR), universal bacterial primers 8F (5’-AGAGTTTGATCMTGGCTCAG-3’; Edwards et al. 1989) with a fluorescent S-hexachlorofluorescein (HEX) label and 1392R (5’-ACGGGCGGTGTGTACA-3’; Lane 1991) were used to amplify an approximately 1385 bp region of the gene. Each reaction was run in triplicate and consisted of 2 U PerfectTaq (5-Prime), 1X PerfectTaq buffer, 1.25 mM Mg(OAc)2, 0.06 mM deoxynucleoside triphosphates, 0.8 μg bovine serum albumin (BSA), 25 pmol of each primer, and sterile deionized water to a final volume of 100 μl. PCR reaction conditions were 85 °C for 5 min, 35 cycles of 94 °C for 45 s, 62 °C for 90 s, and 72 °C for 90 s, followed by a final extension step at 72 °C for 10 min. Negative reagent controls without added template DNA were run with all reactions. PCR products were checked for quality and appropriate size by gel electrophoresis on 1.0% agarose gels containing GelRed™ (Biotium). PCR products were individually cleaned with the E.Z.N.A. Cycle Pure Kit (Omega Bio-Tek) following the manufacturer’s protocol before triplicate products were pooled for downstream analyses.
Terminal restriction fragment length polymorphism analysis
A Nano-drop 2000c spectrophotometer (Thermo Scientific) was used to determine the concentration of DNA in the pooled PCR products. Digestion reactions were carried out using 100 ng of cleaned PCR product, 2 U of the restriction endonuclease HaeIII (New England BioLabs), 1X enzyme buffer, and sterile deionized water to a total volume of 50 μl. Following an 8-h incubation at 37 °C, enzymes were inactivated by incubation at 80 °C for 30 min. Digestion products were precipitated overnight in 100% ethanol at -20 °C prior to being centrifuged at 16,000 × g for 15 min. The resulting pellets were rinsed in 500 μl of ice-cold 70% ethanol, centrifuged for another 15 min, and dried in a Jouan RC1022 centrivac (Thermo Scientific). Pellets were resuspended in 10 µl of deionized formamide and 0.5 µl of 6-carboxytetramethylrhodamine size standard (Applied Biosystems) prior to being analyzed on an ABI 310 Genetic Analyzer with a 50 cm capillary array (Applied Biosystems).
Using the Peak Scanner v1.0 analysis software (Applied Biosystems), terminal restriction fragment (T-RF) lengths were determined by the Local Southern size-calling algorithm. Peaks below 50 fluorescence units were removed as background noise, and peaks smaller than 100 bp and larger than 500 bp were also removed to avoid uncertainties associated with fragment size determination (reviewed in Schütte et al. 2008). Using T-REX analysis software (Culman et al. 2009), the resulting profiles were binned at a 0.5 clustering threshold prior to statistical analyses.
Chlorophyll a analysis
To assess photosynthetic potential of the sponges’ cyanobacterial photosymbionts, chlorophyll a was measured in frozen foil-wrapped samples from both experiments using methods described in Erwin and Thacker (2007), except that sponges were lyophilized prior to extraction (Gochfeld et al. 2012a). Briefly, 0.25 g wet weight of sponge tissue was extracted in 10 ml of 90% acetone for 18 h at 4˚C in the dark. Extracts were then transferred to quartz cuvettes, and the absorbance of each extract was quantified at 750, 664, 647 and 630 nm on an Agilent 8453 UV spectrophotometer. Chlorophyll a concentrations were calculated using formulas from Parsons et al. (1984) and standardized to the dry mass of the extracted sponge to obtain µg chlorophyll a mg−1 sponge tissue.
Protein analysis
Soluble protein concentration in frozen samples was measured as an indicator of sponge holobiont condition for both experiments. The Bradford (1976) assay was used, in which 10 g of lyophilized sponge tissue was extracted in 5 ml of 1 M NaOH for 18 h, after which 100 µl of each sample was added to a test tube with 5 ml of Quick Start™ Bradford Dye Reagent (Bio-Rad). The absorbance of each sample was measured at 595 nm using a BioPhotometer 6131 (Eppendorf) and plotted against a standard curve using BSA. Protein concentrations were standardized to the dry weight of each sponge sample to obtain µg protein mg−1 sponge tissue.
Heat shock protein analysis
Changes in the concentration of Hsp70 were compared among sample groups in the January 2011 experiment, as outlined in Sarkis et al. (2005). A small (~ 1 g) piece of each sponge was removed using a sterile razor blade and immediately homogenized in 2 ml of Hsp sample buffer (125 mM Tris pH 6.8, 10% glycerol, 2% sodium dodecyl sulfate (SDS)) in a microcentrifuge tube. Samples were boiled for 3 min and then spun in a microcentrifuge at 500 rpm for 10 min. The supernatant was transferred to a clean tube and spun a second time. The final supernatant was diluted to a 1:2 ratio of sample to fresh Hsp buffer and frozen for transport. Total protein concentration was measured using the DC Protein Assay (Bio-Rad) with BSA standards, and sample protein content was standardized for use in the assay. Samples were boiled for 3 min and then separated by SDS–polyacrylamide gel electrophoresis and stained with SYPRO Tangerine protein gel stain (Molecular Probes). Separated proteins were transferred to nitrocellulose and incubated with monoclonal anti-Hsp70 clone BRM-22 antibodies (Sigma-Aldrich). Bands were visualized by chemiluminescence (LumiGLO®, Cell Signaling Technology) using a Chemidoc MP Imaging System (Bio-Rad), and mean gray pixel intensity of the bands, which represents relative Hsp70 concentration, was compared using the ImageJ program (Schneider et al. 2012).
Secondary metabolite profiles
Frozen A. cauliformis samples from both experiments were lyophilized and ground, and 30 mg of powdered sponge tissue was extracted three times in 15 mL methanol. The extracts were combined, filtered, dried and weighed. Extracts were re-dissolved to a concentration of 5 mg mL−1 in methanol and filtered through a 0.45-μm polytetrafluoroethylene (PTFE) filter. Chemical profiles of each extract were generated by injecting 10 μl of filtered extract onto an analytical high-performance liquid chromatography (HPLC) system (Waters Alliance 2695) coupled to a photodiode array detector (Waters 2998). Analysis was performed on a Phenomenex 5 μm Luna C18 250 × 4.6 mm column, using a gradient system of 0.05% trifluoroacetic acid (TFA) in acetonitrile and 0.05% TFA in water, following methods in Gochfeld et al. (2012b). Chromatograms were extracted at 254 nm, and the areas under the curve for 5 well-resolved peaks were integrated. Peak retention times and identities (from Gochfeld et al. 2012b) are 12.7 min (aplysamine-1), 14.3 min, 15.5 min, 19.7 min, and 34.8 min (fistularin-3) for peaks 1–5, respectively.
Data analysis
Using the vegan and RVAideMemoire packages (Hervé 2019; Oksanen et al. 2018; R Core Team 2019) in R version 3.5.2, relative abundance transformations of the normalized T-RFLP peak area data from T-REX were created for bacterial communities. Bray–Curtis dissimilarity matrices of the transformed data were generated in R (vegdist) and examined for homogeneity of variance (permdisp), followed by ANOVA to compare differences by sample condition (attached healthy, attached ARBS, in situ Healthy, in situ ARBS), time (initial, final), and time by sample interactions for each collection day. Subsequently, permutational analyses of variance (PERMANOVA; adonis) and pairwise comparisons of PERMANOVA (pw.permanova) results were performed for each collection day. Adjusted p-values were determined for pairwise comparisons using false discovery rates. The function metaMDS, with a Wisconsin transformation of the Bray–Curtis dissimilarity matrix, was used to generate non-metric multidimensional scaling (nMDS) plots to visualize the bacterial communities associated with the samples in two-dimensional space.
For the July 2010 and January 2011 experiments, chlorophyll a and protein concentration were compared among attached sponge samples using repeated measures ANOVAs (RMANOVA), after testing for normality using Shapiro–Wilk tests, on the initial and final samples for each sample condition group (attached to an in situ Healthy or an in situ ARBS-affected sponge). Individual sponges were sampled initially (Day 0) and at one final time point (Days 3, 6, or 9 in July 2010; Days 1, 3, or 9 in January 2011). Due to this paired sample design, a separate RMANOVA was performed for sponges sampled on each collection day. To maximize the ecological context of our results (e.g., Smith 2020), we provided effect sizes of the responses. For the in situ healthy and ARBS-affected sponges, concentrations were compared using a one-way analysis of covariance (ANCOVA) with sample condition (healthy or diseased) as the main factor and collection day (i.e., number of days of attachment) as the covariate. Hsp70 band intensity in the January 2011 experiment was analyzed similarly.
To assess secondary metabolite composition in both experiments, the areas under the curve for the 5 peaks integrated from the chemical profiles of attached and in situ sponges were analyzed as described for the bacterial community analysis (see above). Additionally, a similarity percentage analysis (simper in the vegan package) was run for all sponges to determine the contribution of each of the five peaks to the dissimilarity between the groups. For the in situ sponges only, a one-way ANCOVA was used to compare areas under the curve for each of the 5 peaks, with sample condition (healthy or diseased) as the factor and collection day as the covariate.
Results
Disease transmission
In the July experiment, 3 of the 7 initially healthy sponges (42.9%) attached to a diseased sponge showed visible signs of disease transmission (e.g., red discoloration and tissue necrosis; Fig. 1d) after 3 days of contact, while 100% exhibited disease signs by day 6. Considering that all of the sponges in contact with ARBS showed signs of the disease by day 6 in July, and to better characterize disease initiation and progression with varying water temperature (average ambient water temperatures were 28 °C in July and 24 °C in January), 8 pairs were collected after 1 day of contact in January. None of these sponges showed signs of ARBS, although even in January, 50% of the sponges attached to diseased sponges had developed ARBS after 3 days of contact, and 100% were diseased after 9 days of contact. No evidence of ARBS was observed in either member of the healthy-healthy pairs at any time point in either experiment, although even healthy sponges exhibited a reaction to contact with conspecifics (Fig. 1c).
Bacterial community analyses
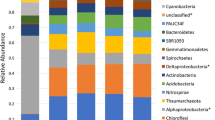
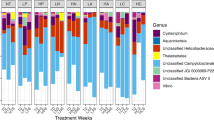
Contact between a healthy sponge and another sponge, whether healthy or ARBS-affected, resulted in changes to the composition of the bacterial community, with significant differences among sample condition for days 3 and 9 in July and at all time points in January (PERMANOVA; Supplemental Table 2, P < 0.05). Pairwise PERMANOVA comparisons showed that the communities in the attached sponges rapidly changed to reflect those in the in situ sponges to which they were attached, with the Attached Healthy-Final–in situ Healthy and the Attached ARBS-Final–in situ ARBS comparisons not significantly different from each other at all time points (Table 1). Similarly, the Attached ARBS-Initial–in situ ARBS comparisons were significantly different on day 9 in July and at all time points in January and the Attached Healthy-Initial–in situ Healthy comparisons were significantly different on days 3 and 9 in July and day 9 in January (Table 1). However, in contrast to the differences between bacterial communities in individual ARBS-affected and healthy sponges that were reported in Olson et al. (2014), none of the in situ Healthy–in situ ARBS comparisons were significantly different. Shifts in the composition of the sponge-associated bacterial communities over time can be seen in the nMDS plots, but no clear pattern was evident (Supplemental Fig. 1). A number of the permdisp analyses showed heterogeneity of variance within the bacterial communities among sample groups (Table 1; Supplemental Table 2), reflecting the intraspecific variability previously reported by Olson et al. (2014) using similar T-RFLP approaches.
Chlorophyll a content
Chlorophyll a concentrations in visually healthy tissue differed significantly between in situ Healthy and in situ ARBS-affected sponges in both experiments, with lower concentrations in diseased sponges (ANCOVA; July: df = 1, F = 6.36, P = 0.017; January: df = 1, F = 6.33, P = 0.016; Supplemental Fig. 2a, b). In both experiments, for the sponges attached to either an in situ healthy or diseased sponge for 9 days, there was a significant time (initial, final) by sample condition interaction (RMANOVA; July: df = 1, F = 13.405, P = 0.008; January: df = 1, F = 16.2, P = 0.002; Supplemental Table 3). Sponges attached to in situ healthy sponges for 9 days exhibited a mean increase in chlorophyll a of 26.7% in July and 13.8% in January, while those attached to in situ diseased sponges exhibited a mean decrease in chlorophyll a concentration of 11.1% in July and 12.5% in January (Fig. 2a, b).
Responses of Aplysina cauliformis attached to healthy or ARBS-affected sponges in contact experiments performed in July and January. Concentration (mean ± SE) of chlorophyll a in a July and b January, soluble protein in c July and d January, and e relative concentration of heat shock protein 70 in January. At each time point, initial and final values are from the same sponge (n = 5–8 pairs per sample condition per day). Repeated measures ANOVA results reported in text
Soluble protein concentrations
Protein concentrations in in situ Healthy and in situ ARBS-affected sponges did not vary uniformly across the two experiments. In July, there were no significant differences between in situ healthy and diseased sponges (ANCOVA; df = 1, F = 0.023, P = 0.88), but there were in January (ANCOVA; df = 1, F = 15.384, P = 0.0003; Supplemental Fig. 2c,d), although day was a significant covariate in January (P = 0.03). Protein concentration in attached sponges also did not vary uniformly with sample condition. A significant increase in protein was observed in attached sponges collected on days 6 and 9 in July (RMANOVA; day 6: df = 1, F = 12.085, P = 0.007; day 9: df = 1, F = 6.375, P = 0.036; Fig. 2c), but after 9 days of contact in January, all attached sponges showed decreases in protein concentration (RMANOVA; df = 1, F = 7.482, P = 0.019; Fig. 2d, Supplemental Table 4). In July, sponges attached to in situ healthy sponges for 6 days exhibited a mean increase in protein concentration of 10.8% while those attached for 9 days increased by 13.8% and protein concentration in sponges attached to in situ diseased sponges increased by 9.5% on day 6 and 3.6% by day 9 (Fig. 2c). In contrast, sponges attached to in situ healthy sponges for 9 days in January exhibited a mean decrease in protein concentration of 11.4% while those attached to in situ diseased sponges exhibited a 19.7% decrease in protein concentration (Fig. 2d).
Heat shock protein 70 concentration
In the January experiment, relative concentration of Hsp70 differed significantly between in situ healthy and diseased sponges (ANCOVA; df = 1, F = 19.799, P = < 0.001; Supplemental Fig. 2e). There was also a significant interaction between sample condition and the covariate of day in the in situ sponges, in which Hsp70 concentration in the in situ healthy sponges increased with the duration of attachment to another sponge (ANCOVA; df = 1, F = 5.740, P = 0.02). RMANOVA for the attached sponges also showed a significant time by sample condition interaction for Hsp70 concentrations at 9 days (P = 0.05; Fig. 2e; Supplemental Table 5). Relative Hsp70 concentrations in attached sponges were comparable to those of in situ diseased sponges after 1 and 3 days of attachment but there was a dramatic increase by day 9, with a much greater increase in those sponges attached to diseased (11.5-fold) relative to healthy (3.1-fold) sponges (Fig. 2e). This resulted in a 3.7 times greater increase in Hsp70 in sponges attached to diseased individuals compared to those attached to healthy individuals. On day 9, Hsp70 concentrations were 2.6-fold higher in the attached sponges than in the in situ healthy sponges to which they were attached, and 7.4-fold higher in attached sponges than in the in situ ARBS-affected sponges to which they were attached.
Chemical metabolite profiles
Significant differences in secondary metabolite concentrations were found between the in situ Healthy and in situ ARBS-affected sponges for each of the 5 metabolites, with peaks 1 (aplysamine-1), 4, and 5 (fistularin-3) occurring in higher concentrations in healthy sponges and peaks 2 and 3 higher in diseased individuals (SIMPER Results: Supplemental File 1; Fig. 3; Supplemental Fig. 3). Within in situ sponges, different metabolites exhibited different responses to being in contact with a conspecific for up to 9 days (Fig. 3; Supplemental Fig. 3). For the attached sponges in July, PERMANOVA showed significant time x sample interactions on secondary metabolite profiles after 6 and 9 days (Supplemental Table 6). In January, secondary metabolite profiles of attached sponges varied between initial and final time points on all collection days, and with sample condition on days 3 and 9 (Supplemental Table 6). By day 9 in both experiments, pairwise PERMANOVA indicated significant changes in the Attached ARBS-Final samples compared to Attached ARBS-Initial samples and in situ healthy sponges (Table 2). Concentrations of metabolites 2 and 3 were again higher in sponges attached to diseased individuals, while metabolites 4 and fistularin-3 (peak 5) were higher in those attached to healthy sponges (Fig. 3; Supplemental Fig. 4).
Nonmetric multidimensional scaling plots of secondary metabolite profiles from Aplysina cauliforms samples collected after 9 days in contact transmission experiments performed in July (a) and January (b). Length and directionality of the vectors indicate the concentration of individual metabolites (peaks) within samples, with peaks 1 (aplysamine-1), 4, and 5 (fistularin-3) more prevalent in healthy sponges and peaks 2 and 3 in ARBS-affected sponges
Discussion
ARBS is a highly infectious disease readily transmissible by contact (Olson et al. 2006; Easson et al. 2013), providing a unique model system to investigate responses of marine invertebrates to pathogenesis. Numerous impacts of this disease on the A. cauliformis holobiont have been identified and include changes ranging from the molecular to organismal levels (Gochfeld et al. 2012a, b; Easson et al. 2013; Olson et al. 2014; Gochfeld et al. 2019), but like most marine diseases, studies have focused on individuals with established lesions, whereas little is known about changes that occur in the holobiont during the early stages of pathogenesis. To date, ARBS is the only sponge disease for which a mode of transmission has been confirmed, and contact with an affected sponge results in transmission within just a few days (Olson et al. 2006) and can also occur through incidental in situ contact due to wave and current action (Easson et al. 2013). Following 9 days of contact in both July and January, all previously healthy sponges placed in contact with in situ ARBS-affected sponges developed ARBS lesions. Because disease transmission occurred so rapidly in the July experiment (i.e., 43% infected by day 3), the timing for the January experiment was altered in an attempt to identify changes in the holobiont that might occur during earlier stages of pathogenesis, even though water temperatures are lower in January. Many microorganisms grow more rapidly at warmer temperatures, suggesting the potential for slower transmission in January. However, although no new ARBS infections were visible after 1 day of contact, even in January, transmission occurred in 50% of sponges attached to in situ ARBS-affected sponges after 3 days of contact, and in 100% after 9 days of contact.
Initially healthy sponge holobionts responded rapidly to disease onset, with significant changes detected in the secondary metabolite profiles, a significant decrease in the concentration of chlorophyll a, and a significant increase in relative Hsp70 concentration within 1–9 days of contact with an ARBS-affected sponge. Although individuals of A. cauliformis appear to respond to simply being in contact with a conspecific, regardless of whether it is healthy or diseased, some changes appear to be disease-related. All samples were collected from visibly healthy tissue of the attached and in situ sponges, indicating that infection with ARBS broadly impacted the organism, which is in contrast with results for diseases of Aplysina aerophoba (Webster et al. 2008) and Agelas tubulata (Deignan et al. 2018), where responses to disease were localized to the disease lesion.
Sponge associated bacteria have been shown to contribute to the functioning of their host, including by aiding in digestion or serving as a source of food (Godefroy et al. 2019), synthesizing bioactive molecules for chemical defense (e.g., Unson et al. 1994; Schmidt et al. 2000; Flatt et al. 2005; Wilson et al. 2014), and participating in nutrient acquisition (e.g., Wilkinson 1983; Freeman and Thacker, 2011; Freeman et al., 2013; Rix et al. 2020) and cycling (e.g., Taylor et al. 2007; Bayer et al. 2008; Freeman and Thacker 2011; Hentschel et al. 2012; de Goeij et al. 2013), as well as in detoxification of metals (Santos-Gandelman et al. 2014; Keren et al. 2015, 2017). As a result of these myriad functions, substantial changes to the composition of associated bacterial communities likely have cascading effects on the functions of the sponge holobiont. Although the bacterial communities associated with A. cauliformis are known to be extremely diverse (Easson and Thacker 2014; Olson et al. 2014), only the translocation of photosynthate from the cyanobacterium Synechococcus spongiarum has been demonstrated in this species (Freeman et al. 2013), leaving the functions of the remaining members of its microbial community largely unknown. Using high-throughput sequencing approaches, intraspecific dissimilarity in the composition of bacterial communities in healthy A. cauliformis was reported to be relatively low (Easson and Thacker 2014; Thomas et al. 2016; Freeman et al. 2020), suggesting that changes to community composition could be important from a functional perspective.
Olson et al. (2014) showed that the bacterial communities in visibly healthy tissue associated with 10 healthy and 10 ARBS-affected A. cauliformis were significantly different using T-RFLP methods. Although comparable differences in the composition of bacterial communities within visibly healthy tissues from the in situ healthy and ARBS-affected sponges were not detected in the transmission experiments described here, likely due to a generalized stress response from being in contact with a conspecific, the present study demonstrated the speed with which the bacterial community in attached sponges alters, with significant changes beginning after only 1 day of contact with a diseased sponge (Table 1). Shifts in the composition of the microbial community prior to or during disease progression have been recognized for a number of sponge diseases, including those that are not transmissible via contact (Webster et al. 2008; Luter et al. 2010; Angermeier et al. 2011, 2012; Stabili et al. 2012; Sweet et al. 2015; Deignan et al. 2018), suggesting that microbial dysbioses may be a hallmark for diseases of marine sponges. However, in the present study, significant changes in the bacterial community composition were found in all attached sponges, regardless of the condition of the sponge to which they were attached, which confirms that dysbiosis can occur with multiple types of stressors and does not necessarily reflect disease onset. Similarly, alterations in the composition of associated microbial communities were found for some sponge species exposed to other environmental stressors, such as varying seawater pH (Ribes et al. 2016; Kandler et al. 2018), temperature (Fan et al. 2013; Ramsby et al. 2018; Vargas et al. 2020), and a combination of pH and temperature (Lesser et al. 2016). More work is needed to understand these complex environmental interactions, as shifts in microbiome composition with environmental stress appear variable across species (e.g., Erwin et al. 2012; Pita et al. 2013).
Some sponges obtain a large portion of their energy and carbon budgets from their photosymbionts (Wilkinson 1983; Cheshire and Wilkinson 1991), including A. cauliformis (70–77% of carbon; Freeman and Thacker 2011). Chlorophyll a concentrations were found to be significantly correlated with the density of cyanobacterial symbionts in A. cauliformis (Freeman and Thacker 2011; Easson et al. 2014). In the current study, significant decreases in chlorophyll a concentration with the initiation of disease indicate either a loss of cyanobacterial symbionts or a decline in the concentration of chlorophyll a per cyanobacterial cell (as shown by Freeman and Thacker 2011 for shaded A. cauliformis). Thylakoid membranes of S. spongiarum in ARBS-affected sponges exhibit disintegration (Gochfeld et al. 2019), which would reduce chlorophyll a concentration, as previously reported for ARBS-affected sponges with established lesions (Gochfeld et al. 2012a). Because some species of Aplysina did not switch feeding strategies to heterotrophy even when energy acquisition from their photosymbionts was compromised (as observed in the congeneric Aplysina fulva; Freeman and Thacker 2011), this loss of photopigments would ultimately negatively affect energy acquisition by the sponge, likely resulting in the observed decreased growth rates in diseased sponges (Gochfeld et al. 2012a). The newly infected sponges in the present study exhibited significant decreases in chlorophyll a concentrations in visibly healthy tissue after only 9 days of contact in both summer and winter, further indicating that impacts of disease on the sponge-associated microbial communities occur rapidly and are functionally important.
ARBS-affected A. cauliformis have been previously reported to have reduced concentrations of soluble protein compared to healthy conspecifics (Gochfeld et al. 2012a). Significant declines in soluble protein concentration were found in all attached sponges after 9 days in the January experiment, regardless of whether they were attached to a healthy or ARBS-affected sponge, suggesting that the stress of being attached to another sponge may be sufficient to reduce protein production, at least in winter. In contrast, by days 6 and 9 in the July experiment, protein concentration increased significantly in all attached sponges. Seasonal differences were also evident in overall protein concentrations, which were approximately twice as high in July as in January, regardless of sample condition, and may be driving the variable responses in protein concentration. Easson et al. (2014) reported that cyanobacterial density was significantly positively correlated with soluble protein concentrations in A. cauliformis, suggesting that if a decline in the number of cyanobacterial cells with disease onset was the mechanism responsible for the observed decrease in chlorophyll a concentration, other responses of the holobiont may temporarily be able to offset these losses of cyanobacterially derived protein. This may indicate that different proteins in the various components of the holobiont respond differently to disease onset or contact with a conspecific, with increases in the concentrations of some proteins, decreases in others, and some remaining stable, confounding measurements of total soluble protein concentrations. The different protein concentrations and contrasting responses to disease onset detected across seasons suggest that total protein concentration may not be a useful metric to evaluate holobiont response to disease.
To examine the concentration of a single protein, heat shock protein 70 (Hsp70), a highly conserved stress protein involved in the folding and unfolding of proteins and in minimizing intracellular aggregations of proteins was used as a measure of generalized stress response. Hsp70 exhibits a higher level of expression under stressful conditions (e.g., increases in temperature, UV radiation, or presence of heavy metals) in many organisms (reviewed in Aguilar-Camacho and McCormack 2017). Recently, Pita et al. (2018) demonstrated that Hsp70 expression increased in two sponges that were exposed to a cocktail of microbe-derived molecules, suggesting that it also has a role in innate immunity. Significantly higher relative concentrations of Hsp70 were observed by day 9 in sponges attached to both healthy and ARBS-affected sponges, indicative of a generalized stress response. However, relative Hsp70 concentration in sponges attached to ARBS-affected sponges was approximately 3.7 times higher than in those attached to healthy sponges, suggesting that while being attached to another sponge may be stressful, the initiation of ARBS magnified that response dramatically, potentially inducing innate immune responses. Similar increases in Hsp70 gene expression were reported in Rhopaloeides odorabile sponges after exposure to thermal stress (Fan et al. 2013) and in Xestospongia muta affected by sponge orange band disease and elevated water temperature (López-Legentil et al. 2008), indicating that this protein is a useful marker to examine generalized stress responses in sponges. Average water temperature in January, when our Hsp70 measurements were made, was 24° C, substantially lower than the temperatures reported to increase Hsp70 gene expression in sponges (> 30° C; López-Legentil et al. 2008; Fan et al. 2013), suggesting that this response was due to stress caused by disease initiation and/or contact with conspecific sponges, rather than temperature.
Sponges in the genus Aplysina produce a diversity of brominated secondary metabolites, many of which provide antipredator (Pawlik et al. 1995; Gochfeld et al. 2012b), allelopathic (Pawlik et al. 2007), and antimicrobial (Kelly et al. 2005; Gochfeld et al. 2012b) protection for the host sponge. It has been suggested that some of these compounds may be rapidly transformed by enzymes in response to tissue damage, resulting in the formation of more bioactive products (Teeyapant and Proksch 1993; Thoms et al. 2006), although this does not appear to be the case for Caribbean Aplysina species (Puyana et al. 2003). Some secondary metabolites of Aplysina spp. may represent inducible defenses, which require up-regulation of proteins in the sponge or its associated microbes and take longer to respond to a specific stressor (Gochfeld et al. 2012b). The secondary metabolites in Aplysina spp. are hypothesized to be produced by the sponge, although there may be some collaboration from microbial associates (Sacristán-Soriano et al. 2011). Shifts in both the secondary metabolite profiles and composition of the associated bacterial communities following contact with a diseased sponge support the potential for a connection, but do not address this question directly.
Differences in chemical profiles between healthy and diseased individuals of A. aerophoba and A. cauliformis have been previously demonstrated (Webster et al. 2008; Gochfeld et al. 2012b), as have feeding deterrent and antibacterial activities of extracts and isolated metabolites from A. cauliformis (Gochfeld et al. 2012b). Gochfeld et al. (2012b) reported that secondary metabolite profiles did not vary significantly in tissue samples collected over a range of distances (1–8 cm) from the ARBS lesion, indicative of a systemic, rather than localized, response to this disease. In concordance with the results of Gochfeld et al. (2012b), the concentration of certain metabolites (e.g., peak 1 = aplysamine-1, peak 5 = fistularin-3) were higher in in situ healthy sponges than in in situ diseased conspecifics, while others (peaks 2, 3) showed the opposite trend (Fig. 3; Supplemental Fig. 3,4). Whereas specific metabolites occurred in different concentrations in the in situ healthy and diseased sponges in the present study, the overall secondary metabolite profiles of the in situ sponges did not differ statistically at any time point in July, although they did differ on days 3 and 9 in January. In addition to contact-dependent effects, another possible explanation for the absence of consistent differences in the chemical profiles of the in situ sponges in this study is the smaller sample size (n = 4–8 for each condition at each time point) in this study, and the lower number of peaks quantified (n = 5) in comparison to the 10 sponges per condition and 15 peaks quantified in Gochfeld et al. (2012b). Nonetheless, by day 9 in both January and July, the secondary metabolite profiles of the Attached ARBS-Final sponges and the in situ ARBS-affected sponges were not significantly different (Fig. 3; Supplemental Table 6), whereas the Attached ARBS-Initial profiles grouped with the other sponges that remained healthy (Fig. 3). These patterns were largely driven by peaks 2 and 3 in the in situ ARBS and newly ARBS-infected sponges, whereas aplysamine-1 and fistularin-3 (peaks 1 and 5, respectively), and to a lesser extent peak 4, were associated with healthy sponges. Both qualitative and quantitative differences in secondary metabolite profiles were observed when comparing healthy A. cauliformis and healthy tissue adjacent to advanced lesions in diseased individuals (Gochfeld et al. 2012b), whereas in this study of early lesion development, only quantitative differences were observed. Secondary metabolite profiles within apparently healthy tissue from diseased X. muta individuals revealed changes relative to healthy sponges, although the number of metabolites declined in discolored tissue adjacent to the lesion and were absent within the lesion itself (Angermeier et al. 2011).
A holistic view of the sampled metrics in the current study, which include changes within the symbionts (bacterial community, chlorophyll a) and the host sponge (Hsp70), as well as physiological measurements that likely encompass both compartments of the holobiont (soluble protein, secondary metabolites), suggests that ARBS has a systemic effect on the A. cauliformis holobiont. Although the combination of these metrics provides strong evidence for an ARBS-related stress response on the sponge holobiont, further research could investigate specific aspects of the impact of ARBS in more detail. For example, the T-RFLP approach used in this study is a cost-effective mechanism to investigate changes in bacterial communities and was previously shown to be a useful tool for investigating ARBS impact (Olson et al. 2014). However, this method does not provide fine taxonomic data on what microbial members are specifically impacted by ARBS, nor does it provide sufficient resolution to identify a potential etiologic agent(s) of this transmissible disease. Future research should employ next-generation sequencing approaches to capture the full community with a taxon-level view of potential changes related to ARBS infection.
In concordance with results from studies of diseases affecting other sessile marine invertebrates (e.g., corals and soft corals: Pantos et al. 2003; Gil-Agudelo et al. 2006; Pantos and Bythell 2006; Sunagawa et al. 2009; Meyer et al. 2019) but in contrast to other studies of sponge diseases (Webster et al. 2008; Angermeier et al. 2011, 2012; Deignan et al. 2018), A. cauliformis appears to mount a systemic response to ARBS infection. Although ARBS results in increased breakage (Easson et al. 2013), this is often a chronic disease that is not always fatal, and in many cases, the lesion stops growing and the active red band appears to senesce (Gochfeld et al. 2012a; Easson et al. 2013), suggesting that either the holobiont can mount an effective immune response or an environmental factor limits the duration and extent of infection. ARBS-affected sponges have increased densities and activity of archaeocytes, which contain many engulfed bacteria and cyanobacteria in various stages of digestion (Gochfeld et al. 2019), and along with the production of bioactive secondary metabolites (Gochfeld et al. 2012b) represent an active innate immune response. The present study demonstrated that ARBS transmits rapidly via contact, in some cases as quickly as 1–3 days, and significantly affects concentrations of chlorophyll a, Hsp70 and secondary metabolites in visibly healthy tissue. Similar impacts have been described from established lesions, and it remains to be seen what happens within the sponge when lesions senesce. In addition, reallocation of resources that occurs upon disease initiation and subsequent tissue damage may increase susceptibility of the sponge to other stressors, creating the potential for additional loss of sponge biomass on impacted reefs.
References
Aguilar-Camacho JM, McCormack, GP (2017) Molecular responses of sponges to climate change. In: Carballo JL, Bell JJ (eds) Climate change, ocean acidification and sponges Springer, Cham, Switzerland, pp 79–104
Angermeier H, Kamke J, Abdelmohsen UR, Krohne G, Pawlik JR, Lindquist NL, Hentschel U (2011) The pathology of sponge orange band disease affecting the Caribbean barrel sponge Xestospongia muta. FEMS Microbiol Ecol 75:218–230
Angermeier H, Glöckner V, Pawlik JR, Lindquist NL, Hentschel U (2012) Sponge white patch disease affecting the Caribbean sponge Amphimedon compressa. Dis Aquat Org 99:95–102
Bayer K, Schmitt S, Hentschel U (2008) Physiology, phylogeny and in situ evidence for bacterial and archaeal nitrifiers in the marine sponge Aplysina aerophoba. Environ Microbiol 10:2942–2955
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cheshire AC, Wilkinson CR (1991) Modelling the photosynthetic production by sponges on Davies Reef, Great Barrier Reef. Mar Biol 109:13–18
Culman SW, Bukowski R, Gauch HG, Cadillo-Quiroz H, Buckley DH (2009) T-REX: software for the processing and analysis of T-RFLP data. BMC Bioinformatics 10:171. https://doi.org/10.1186/1471-2105-10-171
De Goeij JM, van Oevelen D, Vermeij MJA, Osinga R, Middelburg JJ, de Goeij AFPM, Admiraal W (2013) Surviving in a marine desert: the sponge loop retains resources within coral reefs. Science 342:108–110
Deignan LK, Pawlik JR, Erwin PM (2018) Agelas wasting syndrome alters prokaryotic symbiont communities of the Caribbean brown tube sponge, Agelas tubulata. Microb Ecol 76:459–466
Diaz CM, Rützler K (2001) Sponges: an essential component of Caribbean coral reefs. Bull Mar Sci 69:535–546
Easson CG, Thacker RW (2014) Phylogenetic signal in the community structure of host-specific microbiomes of tropical marine sponges. Front Microbiol 5:532
Easson CG, Slattery M, Momm HG, Olson JB, Thacker RW, Gochfeld DJ (2013) Exploring individual- to population-level impacts of disease on coral reef sponges: using spatial analysis to assess the fate, dynamics, and transmission of Aplysina red band syndrome (ARBS). PLoS ONE. 8: e79976
Easson CG, Slattery M, Baker DM, Gochfeld DJ (2014) Complex ecological associations: competition and facilitation in a sponge-algal interaction. Mar Ecol Prog Ser 507:153–167
Edwards U, Rogall T, Blöcker H, Emde M, Böttger EC (1989) Isolation and direct complete nucleotide determination of entire genes: characterization of a gene coding for 16S ribosomal RNA. Nucl Acid Res 19:7843–7853
Erwin PM, Pita L, López-Legentil S, Turon X (2012) Stability of sponge-associated bacteria over large seasonal shifts in temperature and irradiance. Appl Environ Microbiol 78:7358–7368
Erwin PM, Thacker RW (2007) Phylogenetic analyses of marine sponges within the order Verongida: a comparison of morphological and molecular data. Invert Biol 126:220–234
Fan L, Liu M, Simister R, Webster NS, Thomas T (2013) Marine microbial symbiosis heats up: the phylogenetic and functional response of a sponge holobiont to thermal stress. ISME J 7:991–1002
Flatt PM, Gautschi JT, Thacker RW, Musafija-Girt M, Crews P, Gerwick WH (2005) Identification of the cellular site of polychlorinated peptide biosynthesis in the marine sponge Dysidea (Lamellodysidea) herbacea and symbiotic cyanobacterium Oscillatoria spongeliae by CARD-FISH analysis. Mar Biol 147:761–774
Freeman CJ, Easson CG, Matterson KO, Thacker RW, Baker DM, Paul VJ (2020) Microbial symbionts and ecological divergence of Caribbean sponges: a new perspective on an ancient association. ISME J 14:1571–1583
Freeman CJ, Thacker RW (2011) Complex interactions between marine sponges and their symbiotic microbial communities. Limnol Oceanogr 56:1577–1586
Freeman CJ, Thacker RW, Baker DM, Fogel ML (2013) Quality or quantity: is nutrient transfer driven more by symbiont identity and productivity than by symbiont abundance? ISME J 7:1116–1125
Gil-Agudelo DL, Myers C, Smith GW, Kim K (2006) Changes in the microbial communities associated with Gorgonia ventalina during aspergillosis infection. Dis Aquat Org 69:89–94
Gochfeld DJ, Easson CG, Freeman CJ, Thacker RW, Olson JB (2012a) Disease and nutrient enrichment as potential stressors on the Caribbean sponge Aplysina cauliformis and its bacterial symbionts. Mar Ecol Prog Ser 456:101–111
Gochfeld DJ, Kamel HN, Olson JB, Thacker RW (2012b) Trade-offs in defensive metabolite production but not ecological function in healthy and diseased sponges. J Chem Ecol 38:451–462
Gochfeld DJ, Diaz MC, Renegar DA, Olson JB (2019) Histological and ultrastructural features of Aplysina cauliformis affected by Aplysina red band syndrome. Invert Biol. 138: e12247
Godefroy N, Le Goff E, Martinand-Mari C, Belkhir K, Vacelet J, Baghdiguian S (2019) Sponge digestive system diversity and evolution: filter feeding to carnivory. Cell Tissue Res 377:341–351
Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nature Rev Microbiol 10:641–654
Hervé M (2019) RVAideMemoire: Testing and Plotting Procedures for Biostatistics R package version 09–73. https://CRANR-projectorg/package=RVAideMemoire
Hewson I, Button JB, Gudenkauf BM, Miner B, Newton AL, Gaydos JK, Wynne J, Groves CL, Hendler G, Murray M, Fradkin S, Breitbart M, Fahsbender E, Lafferty KD, Kilpatrick AM, Miner CM, Raimondi P, Lahner L, Friedman CS, Daniels S, Haulena M, Marliave J, Burge CA, Eisenlord ME, Harvell CD (2014) Densovirus associated with sea-star wasting disease and mass mortality. Proc Natl Acad Sci 111:17278–17283
Jones CG, Lawton JH, Shachak M (1997) Positive and negative effects of organisms as physical ecosystem engineers. Ecol 78:1946–1957
Kandler NM, Wahab MAA, Noonan SHC, Bell JJ, Davy SK, Webster NS, Luter HM (2018) In situ responses of the sponge microbiome to ocean acidification. FEMS Microbiol Ecol 94:2018,fiy205 [doi: https://doi.org/10.1093/femsec/fiy205]
Kelly SR, Garo E, Jensen PR, Fenical W, Pawlik JR (2005) Effects of Caribbean sponge secondary metabolites on bacterial surface colonization. Aquatic Microb Ecol 40:191–203
Keren R, Lavy A, Mayzel B, Ilan M (2015) Culturable associated bacteria of the sponge Theonella swinhoei show tolerance to high arsenic concentrations. Front Microbiol 6:154
Keren R, Mayzel B, Lavy A, Polishchuk I, Levy D, Fakra SC, Pokroy B, Ilan M (2017) Sponge-associated bacteria mineralize arsenic and barium on intracellular vesicles. Nat Comm 8:14393
Lafferty KD, Hofmann EE (2016) Marine disease impacts, diagnosis, forecasting, management and policy. Phil Trans Roy Soc b: Biol Sci 371:1689.
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley and Sons, Chichester, pp 115–175
Lesser MP, Fiore C, Slattery M, Zaneveld J (2016) Climate change stressors destabilize the microbiome of the Caribbean barrel sponge, Xestospongia muta. J Exp Mar Biol Ecol 475:11–18
López-Legentil S, Song B, McMurray SE, Pawlik JR (2008) Bleaching and stress in coral reef ecosystems: hsp70 expression by the giant barrel sponge Xestospongia muta. Mol Ecol 17:1840–1849
Luter HM, Webster NS (2017) Sponge disease and climate change. In: Carballo JL, Bell JJ (eds) Climate change, ocean acidification and sponges. Springer, Cham, Switzerland, pp 411–428
Luter HM, Whalan S, Webster NS (2010) Prevalence of tissue necrosis and brown spot lesions in a common marine sponge. Mar Freshwater Res 61:484–489
Mehbub MF, Lei J, Franco C, Zhang W (2014) Marine sponge derived natural products between 2001 and 2010: trends and opportunities for discovery of bioactives. Mar Drugs 12:4539–4577
Meyer JL, Castellanos-Gell J, Aeby GS, Häse CC, Ushijima B, Paul VJ (2019) Microbial community shifts associated with the ongoing Stony Coral Tissue Loss Disease outbreak on the Florida reef tract. Front Microbiol 10:2244.
Mydlarz LD, Jones LE, Harvell CD (2006) Innate immunity, environmental drivers, and disease ecology of marine and freshwater invertebrates. Ann Rev Ecol Evol System 37:251–288
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2018) vegan: Community Ecology Package R package version 25–2. https://CRANR-projectorg/package=vegan
Olson JB, Gochfeld DJ, Slattery M (2006) Aplysina red band syndrome: a new threat to Caribbean sponges. Dis Aquat Org 71:163–168
Olson JB, Thacker RW, Gochfeld DJ (2014) Molecular community profiling reveals impacts of time, space, and disease status on the bacterial community associated with the Caribbean sponge Aplysina cauliformis. FEMS Microbiol Ecol 87:268–279
Pantos O, Bythell JC (2006) Bacterial community structure associated with white band disease in the elkhorn coral Acropora palmata determined using culture-independent 16S rRNA techniques. Dis Aquat Org 69:79–88
Pantos O, Cooney RP, Le Tissier MDA, Barer MR, O’Donnell AG, Bythell JC (2003) The bacterial ecology of a plague-like disease affecting the Caribbean coral Montastrea annularis. Environ Microbiol 5:370–382
Parsons TR, Maita Y, Lalli CM (1984) A manual of chemical and biological methods for seawater analysis. Pergamon Press, New York
Paul VJ, Freeman CJ, Agarwal V (2019) Chemical ecology of marine sponges: new opportunities through “-omics.” Int Comp Biol 59:765–776
Pawlik JR, Chanas B, Toonen RJ, Fenical W (1995) Defenses of Caribbean sponges against predatory reef fish. I. Chemical Deterrency Mar Ecol Prog Ser 127:183–194
Pawlik JR, Steindler L, Henkel TP, Beer S, Ilan M (2007) Chemical warfare on coral reefs: sponge metabolites differentially affect coral symbiosis in situ. Limnol Oceanogr 52:907–911
Pita L, Erwin PM, Turon X, López-Legentil S (2013) Till death do us part: Stable sponge-bacteria associations under thermal and food shortage stresses. PLoS ONE. https://doi.org/10.1371/journal.pone.0080307
Pita L, Hoeppner MP, Ribes M, Hentschel U (2018) Differential expression of immune receptors in two marine sponges upon exposure to microbial-associated molecular patterns. Sci Rep 8:16081.
Precht WF, Gintert BE, Robbart ML, Fura R, van Woesik R (2016) Unprecedented disease-related coral mortality in southeastern Florida. Sci Rep 6:31374.
Puyana M, Fenical W, Pawlik JR (2003) Are there activated chemical defenses in sponges of the genus Aplysina from the Caribbean? Mar Ecol Prog Ser 246:127–135
Puyana M, Pawlik J, Blum J, Fenical W (2015) Metabolite variability in Caribbean sponges of the genus Aplysina. Rev Bras Farmacogn 25:592–599
Ramsby BD, Hoogenboom MO, Whalan S, Webster NS (2018) Elevated seawater temperature disrupts the microbiome of an ecologically important bioeroding sponge. Mol Ecol 27:2124–2137
Ribes M, Calvo E, Movilla J, Logares R, Coma R, Pelejero C (2016) Restructuring of the sponge microbiome favors tolerance to ocean acidification. Envr Micro Rep 8:536–544
Rix L, Ribes M, Coma R, Jahn MT, de Goeij JM, van Oevelen D, Escrig S, Meibom A, Hentschel U (2020) Heterotrophy in the earliest gut: a single-cell view of heterotrophic carbon and nitrogen assimilation in sponge-microbe symbioses. ISME J 14:2554–2567
Sacristán-Soriano O, Banaigs B, Casamayor EO, Becerro MA (2011) Exploring the links between natural products and bacterial assemblages in the sponge Aplysina aerophoba. Appl Environ Microbiol 77:862–870
Santos-Gandelman JF, Giambiagi-deMarval M, Muricy G, Barkay T, Laport MS (2014) Mercury and methylmercury detoxification potential by sponge-associated bacteria. Antonie Van Leeuwenhoek 106:585–590
Sarkis S, Boettcher A, Ueda N, Hohn C (2005) A simple transport procedure for juvenile calico scallops, Argopecten gibbus (Linnaeus, 1758). J Shellfish Res 24:377–380
Schmidt EW, Obraztsova AY, Davidson SK, Faulkner DJ, Haygood MG (2000) Identification of the antifungal peptide-containing symbiont of the marine sponge Theonella swinhoei as a novel δ-proteobacterium, “Candidatus Entotheonella palauensis”. Mar Biol 136:969–977
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Schütte UME, Abdo Z, Bent SJ, Shyu C, Williams CJ, Pierson JD, Forney LJ (2008) Advances in the use of terminal restriction fragment length polymorphism (T-RFLP) analysis of 16S rRNA genes to characterize microbial communities. Appl Microbiol Biotechnol 80:365–380
Shore A, Caldwell JM (2019) Modes of coral disease transmission: how do diseases spread between individuals and among populations? Mar Biol 166:45.
Smith EP (2020) Ending reliance on statistical significance will improve environmental inference and communication. Estuaries Coasts 43:1–6
Stabili L, Cardone F, Alifano P, Tredici SM, Piraino S, Corriero G, Gaino E (2012) Epidemic mortality of the sponge Ircinia variabilis (Schmidt, 1962) associated to proliferation of a Vibrio bacterium. Invert Microbiol 64:802–813
Stockton S (2016) Variability in antibacterial chemical defenses in Caribbean sponges of the genus Aplysina. Honors Thesis, University of Mississippi
Sunagawa S, DeSantis TZ, Piceno YM, Brodie EL, DeSalvo MK, Voolstra CR, Weil E, Andersen GL, Medina M (2009) Bacterial diversity and White Plague Disease-associated community changes in the Caribbean coral Montastraea faveolata. ISME J 3:512–521
Sweet M, Bulling M, Cerrano C (2015) A novel sponge disease caused by a consortium of microorganisms. Coral Reefs 34:871–883
Taylor MW, Radax R, Steger D, Wagner M (2007) Sponge-associated microorganisms: evolution, ecology, and biotechnological potential. Microbiol Mol Biol Rev 71:295–347
Team RDC (2019) R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2011 URL: https://www. R-project Org
Teeyapant R, Proksch P (1993) Biotransformation of brominated compounds in the marine sponge Verongia aerophoba- evidence for an induced chemical defense? Naturwissenschaften 80:369–370
Thomas T, Moitinho-Silva L, Lurgi M, Björk JR, Easson C, Astudillo-García C, Olson JB, Erwin PM, López-Legentil S, Luter H, Chaves-Fonnegra A, Costa R, Schupp PJ, Steindler L, Erpenbeck D, Gilbert J, Knight R, Ackermann G, Lopez JV, Taylor MW, Thacker RW, Montoya JM, Hentschel U, Webster NS (2016) Diversity, structure and convergent evolution of the global sponge microbiome. Nature Comm 7:11870. https://doi.org/10.1038/ncomms11870
Thoms C, Ebel R, Proksch P (2006) Activated chemical defense in Aplysina sponges revisited. J Chem Ecol 32:97–123
Unson MD, Holland ND, Faulkner DJ (1994) A brominated secondary metabolite synthesized by the cyanobacterial symbiont of a marine sponge and accumulation of the crystalline metabolite in the sponge tissue. Mar Biol 119:1–11
Van Soest RWM, Boury-Esnault N, Vacelet J, Dohrmann M, Erpenbeck D, De Voogd NJ, Santodomingo N, Vanhoorne B, Kelly M, Hooper JNA (2012) Global diversity of sponges (Porifera). PLoS ONE. 7: e35105
Vargas S, Leiva L, Wörheide G (2020) Short-term exposure to high-temperature water causes a shift in the microbiome of the common aquarium sponge Lendenfeldia chondrodes. Microb Ecol. https://doi.org/10.1007/s00248-020-01556-z
Walton CJ, Hayes NK, Gilliam DS (2018) Impacts of a regional, multi-year, multi-species coral disease outbreak in southeast Florida. Front Mar Sci. 8: 323
Ward JR, Lafferty KD (2004) The elusive baselines of marine disease: are diseases in ocean ecosystems increasing? PLoS Biol. https://doi.org/10.1371/journal.pbio.0020120
Webster NS (2007) Sponge disease: a global threat? Environ Microbiol 9:1363–1375
Webster NS, Xavier JR, Freckelton M, Motti CA, Cobb R (2008) Shifts in microbial and chemical patterns within the marine sponge Aplysina aerophoba during a disease outbreak. Envr Microbiol 10:3366–3376
Wilkinson CR (1983) Net primary productivity in coral reef sponges. Science 219:410–412
Wilson MC, Mori T, Rückert C, Uria AR, Helf MJ, Takada K, Gernert C, Steffens UAE, Heycke N, Schmitt S, Rinke C, Helfrich EJN, Brachmann AO, Gurgui C, Wakimoto T, Kracht M, Crüsemann M, Hentschel U, Abe I, Matsunaga S, Kalinowski J, Takeyama H, Piel J (2014) An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 506:58–62
Wulff J (2012) Ecological interactions and the distribution, abundance, and diversity of sponges. Adv Mar Biol 61:273–344
Zea S, Henkel TF, Pawlik JR (2014) The Sponge Guide: a picture guide to Caribbean sponges. 3rd ed. Available online at www.spongeguide.org
Acknowledgements
We thank M Ansley, S Criddle, S Lee, D Lee, C Williams, and Dr. JL Stevens for their help with the laboratory analyses, and Drs. E Hunkin and M Slattery for their help in the field. Dr. E Mueller and the Perry Institute of Marine Science provided logistical support. Samples were collected under Bahamas Department of Marine Resources Scientific Research Permit MAF/FIS/1,12,46A,79. Funding was provided by National Science Foundation grants OCE-1214303 to DJG and JBO, OCE-0727833 to JBO and OCE-0727996 to DJG and a National Institute of Undersea Science and Technology grant to DJG.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Topic Editor Carly Kenkel
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Olson, J.B., Easson, C.G. & Gochfeld, D.J. Temporal changes in the sponge holobiont during the course of infection with Aplysina Red Band Syndrome. Coral Reefs 40, 1211–1226 (2021). https://doi.org/10.1007/s00338-021-02126-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-021-02126-6