Abstract
The giant barrel sponge, Xestospongia muta, represents a dominant member of Caribbean reef communities. Recent microsatellite data have revealed the presence of two genetic clusters of X. muta in a monitored population on Conch Reef, Florida Keys, with a reduced abundance of one cluster among the largest individuals. Tracking changes to populations over time and their ecological significance requires rapid identification of each genetic cluster and subsequent studies of biological differences between clusters. Here, we show that single-gene barcoding detected the same intraspecific genetic variation within X. muta from Conch Reef as microsatellite data, with mitochondrial gene sequences (cytochrome c oxidase subunit I, I3-M11 partition) from 54 individuals corresponding to 4 known haplotypes within the two genetic clusters. Remarkably, mapping these haplotypes to barrel sponges worldwide revealed positioning on opposite ends of a global network, despite their sympatric occurrence. Further, we investigated whether differences in symbiotic microbial communities could be detected between the two clusters using next-generation (Illumina) sequencing of 16S rRNA gene amplicons. Both clusters exhibited highly diverse microbial communities, with 12,185 total OTUs spanning 38 bacterial and 3 archaeal phyla, but significant differences in microbial community structure (PERMANOVA; p < 0.001) and diversity (Shannon diversity index; p < 0.01) were detected between the two clusters. As sponges typically exhibit interspecific, but not intraspecific, variability in microbial communities, these findings within a sympatric population provide additional support for ecologically relevant cryptic species of X. muta.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coral reefs represent complex ecological systems, with biodiversity rivaling or surpassing even that of tropical rainforests. Sponges are particularly abundant on Caribbean reefs (Loh and Pawlik 2014), with species richness outnumbering even corals and algae (Diaz and Rützler 2001). As spatial competitors with corals, rapid coral decline has led to concurrent increases in sponge cover on some reefs as sponges colonize newly available space faster than corals (Aronson et al. 2002; McMurray et al. 2010; Bell et al. 2013; Marlow et al. 2019). In the long term, this trend may lead to sponge-dominated reefs replacing coral-dominated reefs (Bell et al. 2013), although this shift will ultimately depend on the causes of coral decline, as sponges may be similarly sensitive to environmental stressors (Powell et al. 2014; reviewed in Pawlik and McMurray 2020).
The increasing abundance of sponges on tropical reefs may amplify the important ecological services these animals provide. Sponges function as sources of three-dimensional habitat for other marine life (Pawlik and Henkel 2005; Seemann et al. 2018; Chak and Rubenstein 2019), feed spongivorous reef fishes, invertebrates, and sea turtles (Pawlik 1983; Van Dam and Diez 1997; Ruzicka and Gleason 2008), and serve to anchor and reinforce reef structures (Wulff and Buss 1979; Diaz and Rützler 2001), all functional roles that may become increasingly important as reef-building corals continue to decline. Sponges also filter vast quantities of water (Reiswig 1974), removing toxic heavy metals and other pollutants from the water column in the process (Cebrian et al. 2007). Perhaps most importantly, sponges are important in nutrient cycling, including the transformation of nitrogen (Hoffmann et al. 2009) and the recirculation of dissolved organic carbon back into trophic systems, a “sponge-loop” that may support the diverse array of life within reefs that are otherwise nutrient-limited (de Goeij et al. 2013).
The giant barrel sponge Xestospongia muta may play an especially important role within Caribbean reef communities. The so-called redwood of the reef can grow to over a meter in height and diameter, and is long-lived, with some individuals estimated to be thousands of years old (McMurray et al. 2008). Xestospongia muta is also one of the most abundant sponge species on Caribbean reefs (Loh and Pawlik 2014), and in the Florida Keys, there is evidence that it is experiencing rapid population growth (McMurray et al. 2015). Three species of Xestospongia are currently recognized worldwide: X. muta, X. testudinaria, and X. bergquistia; however, Swierts et al. (2017) recently have proposed that these species are included among complexes of cryptic species, based on molecular analyses of mitochondrial and nuclear genes that revealed the presence of 17 distinct genotypes. Using microsatellite markers, Deignan et al. (2018) investigated intraspecific variation within a single population of X. muta on Conch Reef, Florida, and detected two distinct, co-occurring genetic clusters (named Cluster 1 and 2). However, the relationship between these sympatric genetic clusters and the genotypes described by Swierts et al. (2017) remains unresolved.
Deignan et al. (2018) further determined that the occurrence of each genetic cluster of X. muta was skewed towards different sponge sizes. Cluster 1 sponges dominated the two largest size classes within the population, while Cluster 2 sponges made up about half of the smallest two size classes. As larger sponge individuals should exhibit far greater reproductive capacity (Uriz et al. 1995; Deignan et al. 2018), Cluster 1 sponges would be expected to dominate at all size classes, and to recruit more heavily into the population. However, the recruitment of Cluster 2 sponges was significantly higher than expected, suggesting a population shift may be occurring on Conch Reef (Deignan et al. 2018). A mechanism to quickly and accurately distinguish between these two genetic clusters would greatly facilitate the tasks of tracking these shifts over time, investigating their ecological significance, and determining the selective forces responsible for this genetic structure shift. Here, we used single-gene sequencing (mitochondrial gene cytochrome c oxidase subunit I, I3-M11 partition) to rapidly identify the two genetic clusters of X. muta previously recovered using time-intensive microsatellite analyses (Deignan et al. 2018), and related these clusters to the global haplotype network described by Swierts et al. (2017).
Sponges also harbor diverse and highly host-specific microbial communities (Taylor et al. 2007; Erwin et al. 2012a; Fiore et al. 2013; Thomas et al. 2016) that remain relatively conserved between individuals of the same species in the same region, but can vary significantly between different species (Taylor et al. 2004; Montalvo and Hill 2011; Easson and Thacker 2014), including congeneric species (Erwin et al. 2012a). Xestospongia muta is a high-microbial abundance (HMA) sponge with a high diversity of microbial symbionts (Montalvo and Hill 2011; Fiore et al. 2013), which have been shown to exhibit regional differences within the Caribbean (Fiore et al. 2013). Recent studies suggest that these differences may be linked to previously undescribed cryptic species in X. muta (Swierts et al. 2017, 2018), and indeed intraspecific genetic differences have been shown to affect microbial community composition in some sympatric sponges (Griffiths et al. 2019). Microbial symbionts represent an integral part of the sponge holobiont; not only do they confer important benefits upon their host, such as supplemental sources of nutrition (Erwin and Thacker 2008), but they have also been implicated in many of the important ecological functions sponges perform, such as the cycling of nitrogen (Fiore et al. 2015). Thus, in addition to the objectives outlined above, we also characterized and compared the symbiotic microbial communities of the two genetic clusters of X. muta to investigate a potential selective force behind host genetic differentiation and shed light on the ecological consequences of the genetic shift occurring within Florida populations of X. muta.
Materials and methods
Sample collection
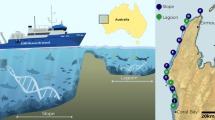
Prior to this study, SCUBA divers identified the location of every Xestospongia muta individual within a single plot (AQS3) on Conch Reef, assigning an x, y coordinate and a unique 3-digit identifier (McMurray et al. 2010). AQS3 (24°56′59″N; 80°27′13 W) is a permanent circular plot measuring 16 m in diameter, located at approximately 20 m depth. The site has been surveyed typically once per annum since 2000, with new recruits identified and tagged, and the status (presence/absence) and size classes of all X. muta individuals assessed annually for change (McMurray et al. 2008, 2010, 2015). Deignan et al. (2018) sampled every X. muta individual in AQS3 for microsatellite analysis. For this study, the same sponge individuals (n = 54) were resampled in June 2017 and classified into genetic Cluster 1 or Cluster 2 based on Deignan et al. (2018). Replicate seawater samples (n = 3; 50 mL) were contemporaneously collected in sterile bottles. Sponge samples were collected with a corer and transferred to separate sterile Whirl–Pak bags at depth. All samples were subsequently stored on ice in an insulated cooler and transported back to the field station (< 16 km away from sampling site). Seawater samples were concentrated onto 0.2-μm membrane filters which were halved and immediately frozen at − 80 °C. Sponge samples were each placed in individual vials and fixed in absolute ethanol before storage at −20 °C. Prior to DNA extraction, a small cube (ca. 2 mm3) from each sponge sample was isolated using sterile techniques. Each cube contained parts of the endosome and ectosome of the sponge, evident by the color change from white to reddish-brown, respectively, and the outside edges of the sponge were specifically avoided. DNA extractions of sponge cubes and seawater filters were performed using the DNeasy Blood and Tissue Kit (Qiagen) following manufacturer protocols.
Haplotype identification of sponge hosts
The I3-M11 partition of the mitochondrial gene cytochrome oxidase I (COI) was sequenced from all sponge samples using the primers C1-Npor2760 and C1-J2165 (Misof et al. 2000; Erpenbeck et al. 2002). PCR amplifications were conducted based on a total volume of 25 μL, consisting of 5 pmol of each primer, 2X MyTaq HS Red Mix (Bioline), and 1 μL (ca. 10 ng) of template DNA on an Eppendorf Mastercycler nexus gradient. The thermocycler program included initial denaturation at 95 °C for 1 min; 35 cycles of 95 °C for 15 s, 45 °C for 15 s, and 72 °C for 10 s; and a final extension at 72 °C for 1 min. Sequencing reactions were then conducted using BigDye Terminator v. 3.1 (Applied Biosystems) and the same primers as above. Sequencing PCR products were purified with BigDye XTerminator (Applied Biosystems) and subsequently sequenced on an AB 3500 genetic analyzer (Applied Biosystems) located at the UNCW Center for Marine Science. Corresponding forward and reverse raw sequences were aligned, cleaned, and trimmed to 534 bp using Geneious version 8.02 (Kearse et al. 2012) to generate consensus sequences. Sequences were then compared to the GenBank database using nucleotide–nucleotide BLAST searches (BLASTn) to confirm host identity based on the highest percent match (Altschul et al. 1990). Final sequences were aligned, and variable sites were identified and compared to known microsatellite clusters (Deignan et al. 2018) and mitochondrial haplotypes of X. muta described previously (López-Legentil and Pawlik 2009; Swierts et al. 2017). All consensus sequences were archived in GenBank with the accession numbers MT773455 to MT773508.
Sequencing of microbial symbionts
Microbial communities of X. muta and seawater samples were characterized through amplification and sequencing of a ca. 300 bp fragment (V4 region) of the 16S ribosomal RNA (rRNA) gene using the universal bacterial/archaeal forward primer 515f and reverse primer 806r (Caporaso et al. 2011). DNA extracts were amplified in an Eppendorf Mastercycler nexus gradient, using a thermocycler program consisting of an initial denaturation at 95 °C for 2 min; 35 cycles of 95 °C for 15 s, 50 °C for 15 s, and 72 °C for 20 s; and a final extension at 72 °C for 2 min. All PCR-viable DNA extracts were sent to Molecular Research LP for amplification, library construction, and multiplexed sequencing of partial 16S rRNA gene sequences on an Illumina MiSeq platform, using the same 515f and 806r primers. All raw sequences were submitted to the Sequence Read Archive of NCBI (accession no. PRJNA645299).
Processing of microbial next-generation sequence data
Raw sequence processing was conducted using the mothur software package (v.1.38.0, Schloss et al. 2009), following a slightly modified version of the Illumina MiSeq SOP pipeline (Kozich et al. 2013). In brief, raw sequences were first quality-filtered to remove low-quality sequences and were then aligned to the SILVA reference database (v132; Pruesse et al. 2007). Putative chimeric sequences were culled via self-reference searches with UChime (Edgar et al. 2011). Based on a naïve Bayesian classifier and bootstrap algorithm for confidence scoring (Wang et al. 2007), sequences were classified using the SILVA reference database (v132; Pruesse et al. 2007). Nontarget sequences (chloroplasts, mitochondria, and eukarya) and singletons were removed from the dataset. Using the opticlust clustering algorithm, all remaining sequences were assigned to operational taxonomic units (OTUs) based on 97% sequence similarity, with the taxonomic classification of OTUs established by majority consensus (Schloss and Westcott 2011). In order to standardize sampling depths (i.e., number of sequence reads) across all samples, each data set (i.e., the microbial community of each sponge individual or seawater sample) was subsampled to the lowest read count (n = 62,700) from the final shared file, and all subsequent data analyses were based on these subsampled data sets. Rarefaction curves (Fig. S1) and coverage calculations (Good’s, average = 98.9%, range = 98.5 to 99.3%) confirmed this sampling depth was sufficient to capture the diversity of each sample’s microbial community.
Analysis of microbial community diversity
To compare microbial community diversity between the two sponge clusters (Cluster 1 and Cluster 2), alpha diversity metrics for OTU richness and evenness were calculated within mothur, including the OTU richness indices S (total number of observed OTUs) and Chao 1 (expected number of OTUs), and the Simpson evenness index (E1/D), the inverse Simpson index (D), and the Shannon–Weaver diversity index (H′). Diversity indices for the two genetic clusters of X. muta were statistically compared using analyses of variance (ANOVA) based on a 95% confidence interval.
Analysis of microbial community structure
To compare host specificity and community structure differences between the two sponge clusters, beta diversity metrics for OTU overlap and similarity of microbial symbiont communities were calculated. Venn diagrams were created within mothur to visualize OTU overlap between the two clusters. Within each cluster, core OTU communities, or those OTUs present in all replicate sponges of a genetic cluster, were identified. To compare the microbial communities of each sponge host, Bray–Curtis similarity (BCS) matrices were constructed in PRIMER (version 7.0.13) based on square root transformations of OTU relative abundances and subsequently visualized using multi-dimensional scaling plots. Permutational multivariate analyses of variance (PERMANOVA) were performed to compare the structure of microbial communities of the two clusters and sponges compared to seawater. Permutational multivariate analyses of dispersion (PERMDISP) were performed to verify that all significant PERMANOVA results represented actual structural differences, rather than unequal variability of dispersion between the two clusters. Similarity percentage (SIMPER) analyses were performed to identify the OTUs primarily responsible for the majority (> 50%) of observed differences in microbial communities of the two sponge genetic clusters. To correct for the potential error associated with distance-based metric calculations (Warton et al. 2012), a Metastats test was conducted within mothur using 1000 permutations to determine which SIMPER OTUs exhibit significant differences in relative abundance among groups (White et al. 2009). In addition, we determined whether host sponge size influenced the structure of X. muta microbial communities by classifying size measurements for each sponge into the 5 size classes (I–V) described in McMurray et al. (2010, 2015). PERMANOVA analyses were conducted to determine whether the structure of microbial communities was affected by host size and genetic cluster.
Rare and abundant symbiont community analysis
To determine whether significant differences in community structure also occurred among rarer OTUs, sequence data were further divided into abundant and rare-OTU data partitions, based on a 0.1% cutoff threshold (Fuhrman 2009). This resulted in a cutoff value of 62 sequences, with OTUs containing > 62 sequences considered “abundant,” and OTUs containing ≤ 62 sequences considered “rare”. These abundant and rare data partitions were also tested for differences in microbial community structure through the creation of BCS matrices and PERMANOVA and SIMPER analyses, as described above.
Results
Sponge genetic analysis
Sequences of the I3-M11 partition of the COI gene were obtained for all 54 individuals of X. muta collected from the AQS3 plot. Nucleotide alignments revealed 5 variable sites which corresponded to 4 known haplotypes (López-Legentil and Pawlik 2009; Swierts et al. 2017) and with the two genetic clusters described by Deignan et al. (2018). The majority of sponges grouped in Cluster 1 (77.8%, n = 42) and represented 100% matches for either haplotypes H1/C2 (38.1%, n = 16) or H2/C9 (61.9%, n = 26). The remaining sponges grouped in Cluster 2 (22.2%, n = 12), and represented 100% matches for either haplotypes H3/C5 (33.3%, n = 4) or H4/C8 (66.7%, n = 8; Table 1). Notably, the haplotypes corresponding to Cluster 1 and Cluster 2 sponges were on opposite ends of the global haplotype network (Fig. S2), thus representing highly divergent haplotypes despite their sympatric occurrence.
Symbiotic microbial community composition
For the microbial community assessment, a total of 3,573,900 sequences, consisting of 13,082 distinct OTUs, were obtained from the 57 samples (54 sponge individuals and 3 seawater samples). Seawater bacterioplankton communities represented 2,205 OTUs, with 1,308 of these also present in X. muta microbial communities (Fig. 1). Seawater microbial communities spanned 38 microbial phyla and were dominated by Alphaproteobacteria (66%), Gammaproteobacteria (11%), and Cyanobacteria (7%; Fig. 2). Sponge symbiont communities represented 12,185 OTUs, with the majority of these (89.3%, n = 10,877) unique to sponge hosts and not represented within the bacterioplankton community (Fig. 1). Between the two clusters, 6,225 and 1,030 OTUs were unique to Clusters 1 and 2, respectively, with 4,930 OTUs shared by both clusters. Microbial symbionts of X. muta spanned 37 bacterial and 5 archaeal phyla (Asgardaeota, Crenarchaeota, Euryarchaeota, Nanoarchaeota, and Thaumarchaeota), with Thaumarchaeota present in all sponge samples, and Nanoarchaeota and Euryarcheaota present in 98.1% (n = 53) and 94.4% (n = 51) of all samples for Cluster 1 and 2, respectively. Microbial communities of Cluster 1 sponges included all 37 bacterial and 5 archaeal phyla (Fig. S3) and were dominated by Gammaproteobacteria (16%), Chloroflexi (13%), Thaumarchaeota (13%), and Actinobacteria (10%; Fig. 2). 157 OTUs were core OTUs, present in all Cluster 1 replicates. Of these, only 4 OTUs were core symbionts for Cluster 1 exclusively. Microbial communities of Cluster 2 sponges spanned 31 bacterial phyla and 4 archaeal phyla (Fig. S4) and were dominated by Gammaproteobacteria (18%), Chloroflexi (15%), Thaumarchaeota (11%), and Acidobacteria (8%; Fig. 2). 217 OTUs were found to be part of the core community of Cluster 2, and 64 of these core symbionts were exclusive to Cluster 2 (Table 2).
Microbial community composition averaged for the three sources: Xestospongia muta genetic clusters 1 and 2, and seawater. Phylum-level classifications are shown except for Proteobacteria, which are divided into major classes (Alpha-, Delta-, and Gammaproteobacteria). “Other taxa” represents less abundant taxa and includes: AncK6, unclassified archaea, Asgardaeota, unclassified bacteria, Calditrichaeota, Chlamydiae, Crenarchaeota, Dadabacteria, Deinococcus-Thermus, Dependentiae, Elusimicrobia, Entotheonellaeota, Epsilonbacteraeota, FCPU426, Fibrobacteres, Firmicutes, Fusobacteria, Kiritimatiellaeota, Latescibacteria, LCP-89, Lentisphaerae, Margulisbacteria, Marinimicrobia (SAR406_clade), Modulibacteria, Nanoarchaeaeota, Nitrospinae, Omnitrophicaeota, Patescibacteria, Planctomycetes, unclassified Proteobacteria, Schekmanbacteria, Spirochaetes, Tenericutes, Verrucomicrobia, and Zixibacteria
Symbiont diversity
To determine whether differences in microbial community diversity of the two genetic clusters of X. muta were statistically significant, alpha diversity metrics were calculated by cluster. Of the alpha diversity metrics investigated, the Shannon diversity index indicated that the genetic cluster of the host sponge had a significant effect on microbial community diversity (ANOVA, F1,52 = 8.407, p < 0.01) and the inverse Simpson evenness index indicated that there were significant differences in the evenness of microbial communities across the two sponge genetic clusters (ANOVA, F1,52 = 6.561, p < 0.05). Other indices detected no significant differences in microbial community alpha diversity between the clusters (ANOVA, p > 0.05; Table 3).
Symbiont community structure
Microbial communities clustered significantly in response to source [sponge or seawater; PERMANOVA pseudo-F = 14.217, p(MC) < 0.001; Fig. S5]. Pairwise comparisons further revealed that when grouped by host-sponge genetic cluster, both clusters exhibited significant differences in microbial community structure compared to seawater [t = 4.964 and 4.856 for clusters 1 and 2, respectively, p(MC) = 0.001 for both clusters]. Within microbial communities associated with X. muta, a significant difference in community structure was also detected between the two sponge clusters [PERMANOVA pseudo-F = 3.466, p = 0.001; Table 4; Fig. 3], with host sponge cluster alone explaining 26.7% of the observed variation. The structure of microbial communities from sponges within each genetic cluster were on average 60.7% and 60.9% similar to other sponges within the same genetic cluster, for Clusters 1 and 2, respectively. Microbial communities of sponges within different clusters exhibited 58.2% structural similarity on average. However, no significant differences in dispersion were detected between clusters (PERMDISP, F1,52 = 0.532, p = 0.589) for the overall data partition (Table 4). SIMPER analyses revealed that for the overall data partition, a difference in the relative abundance of a proportionally small number of OTUs (n = 370; 3.04% of all sponge-associated OTUs) were responsible for the majority (50%) of the observed differences between the microbial communities of the two sponge clusters (Table S1). PERMANOVA analyses revealed no significant effect of sponge size on sponge microbial community structure (PERMANOVA, pseudo-F = 1.042, p = 0.335), a significant effect of host sponge genetic cluster (PERMANOVA, pseudo-F = 2.718, p = 0.001), and no significant interaction effect between sponge size and genetic cluster (PERMANOVA, pseudo-F = 0.890, p = 0.83).
Non-metric multi-dimensional scaling plot based on Bray–Curtis similarity matrices of microbial communities in Xestospongia muta genetic clusters 1 (green triangles) and 2 (red triangles). The spatial grouping of all replicate samples by genetic cluster indicates a high degree of host specificity and distinct structural divisions between Cluster 1 and Cluster 2 microbial communities
Rare and abundant symbiont communities
To determine whether differences in OTU relative abundance impacted observed differences in overall microbial community structure, OTUs were sorted into abundant (> 0.1% relative abundance) and rare (≤ 0.1%) partitions. Partitioning resulted in 660 abundant OTUs, representing 98.4% of all microbial sequences, and 12,422 rare OTUs (1.6% of all microbial sequences). Analyses of both the abundant and rare data partitions indicated significant differences in microbial community structure between the two X. muta genetic clusters (PERMANOVA, pseudo-F = 4.993, p = 0.001 for the abundant data partition, and pseudo-F = 1.573, p = 0.002 for the rare data partition; Table 4). SIMPER analyses revealed that for the abundant data partition, microbial communities in sponges within Clusters 1 and 2 exhibited 72.7% and 72.9% similarity, respectively. Between the two clusters, sponges were on average 69.9% similar, with the majority of differences between the two clusters stemming from variation in the relative abundance of just 108 OTUs (0.89% of all sponge-associated OTUs). In contrast, for the rare data set, microbial communities within Cluster 1 and 2 sponges were 12.2% and 14.1% similar on average respectively, with an average similarity between the two clusters of 11.7%, driven primarily (50% contribution) by variation in relative abundance of 1,917 OTUs (15.7% of all sponge-associated OTUs). However, a significant difference in dispersion (PERMDISP, F1,52 = 38.338, p = 0.003) was detected within microbial communities between the two sponge genetic clusters for the rare data partition, but not the abundant data partition (PERMDISP, F1,52 = 0.180, p = 0.721; Table 4).
Discussion
Previous work with microsatellite loci by Deignan et al. (2018) revealed the presence of two genetic clusters of X. muta within a single monitored plot on Conch Reef off the Florida Keys. Here, using the same sponge individuals as the 2018 study, we determined that the I3-M11 partition of the mitochondrial COI gene detects the same fine-scale intraspecific genetic variation and exhibits 100% clustering agreement with the microsatellite data (Deignan et al. 2018). In order to investigate potential selective forces that may contribute to this variation and the genetic shift described in Deignan et al. (2018), we examined whether differences existed between the sponge-associated microbial communities of two genetic clusters. We detected significant differences in microbial community structure and diversity between Cluster 1 and Cluster 2 sponges, which given the high host specificity of sponge symbionts, provides additional support for the cryptic speciation of X. muta proposed by Swierts et al. (2017) and possibly maintained by multiple reproductive peaks and variability in spawning synchronicity (Neely and Butler 2020). Taxonomic descriptions and species delineations are now required to formalize the molecular and microbiological data for cryptic speciation in X. muta.
Determining the causes of the ongoing shift in Conch Reef X. muta populations (Deignan et al. 2018) and cascading impacts on coral reef ecosystems requires additional study facilitated by the molecular tool developed herein. The ability of single-gene sequencing to identify Cluster 1 and Cluster 2 sponge individuals will expedite the tracking of population shifts among X. muta and the study of physiological and ecological differentiation among lineages. Our results also allow the integration of microsatellite data (Deignan et al. 2018) with global investigations of barrel sponge diversity (Swierts et al. 2017). Importantly, the haplotypes corresponding to Cluster 1 and Cluster 2 sponges were on opposite ends of a global haplotype network despite their sympatric occurrence, when mapped against the mitochondrial network developed by Swierts et al. (2017) for giant barrel sponges. Thus, the haplotypes within Cluster 1 and Cluster 2 sponges were found to be more similar to haplotypes from the Indian Ocean, Red Sea, and the Indo-Pacific than they were to each other. We hypothesize that this high degree of genetic divergence in sympatric populations of X. muta underlies important physiological and ecological differences between lineages.
Assessing such ecological differences will require stratifying individuals of X. muta by lineage to determine whether variability in specific traits (e.g., chemical defenses, Loh and Pawlik 2014; reproductive timing, Neely and Butler 2020) relate to cluster status. Gross morphology is currently unable to differentiate lineages of X. muta, as sponges within Clusters 1 and 2 at AQS3 exhibited no consistent morphological differences related to surface roughness. Previous work suggested that some surface morphologies may have a genetic basis in X. muta, as sponges that possessed smooth surfaces versus those with pronounced digitate extensions were assigned to different haplotypes (H4 and H3, respectively; López-Legentil and Pawlik 2009). These haplotypes correspond to C5 and C8 (Swierts et al. 2017), and both represented Cluster 2 sponges. However, other sponge individuals with the same two haplotypes exhibited the most common morphology (rough surface) shared by Cluster 1 sponges (haplotypes C2 and C9, Swierts et al. 2017). Further, annual time-series photographs of individual sponges on Conch Reef over a decade have revealed changes from smooth to rough surfaces, or the opposite, over time (J Pawlik pers. obs.). Accordingly, no morphological differences in surface features consistently differentiate the Cluster 1 and Cluster 2 genetic lineages. Microbial community characterization can help distinguish between morphologically indistinguishable cryptic sponge lineages (Chambers et al. 2013; Gloeckner et al. 2013; Cuvelier et al. 2014). In addition to possessing host-specific microbial symbionts, sponges form highly stable microbial associations that often remain consistent across broad spatial (Hentschel et al. 2002; Pita et al. 2013b) and temporal scales (Friedrich et al. 2001; Lee et al. 2006; Pita et al. 2013b; Erwin et al. 2015), and in spite of fluctuating or otherwise stressful environmental conditions (Friedrich et al. 2001; López-Legentil et al. 2010; Erwin et al. 2012b; Simister et al. 2012; Pita et al. 2013a). Accordingly, the stability and host specificity of sponge microbial associations suggest their potential usefulness as a tool for delineating between cryptic species, especially in the absence of observable morphological differences.
The detection of distinct microbial communities in each genetic cluster of X. muta also provides first insights into biological differences between host lineages. Structurally distinct microbiomes may have ecological consequences for sponge populations if they provide functional contributions to their hosts. The large diversity of symbionts within the microbial communities detected herein, and the varied functional potential described previously for symbionts in X. muta (Fiore et al. 2015), suggest that at least some of these microbial symbionts confer benefits upon their sponge hosts. Here, we focus on putative benefits of the most abundant core OTUs, as the sheer abundance and universal distribution of these OTUs among all the sponges sampled suggest that they play critical roles in the overall function of their host. Indeed, increases in symbiont abundance have been linked to enhanced host function in other marine invertebrates (Goffredi et al. 2007). The top 25 most abundant core microbiome OTUs presented here were also present within the seawater samples, though in extremely low abundance. This finding provides further support for X. muta sourcing at least some of its symbionts via horizontal acquisition (Schmitt et al. 2008), subsequently amplifying rare members of the environmental microbial “seed bank” (Lynch and Neufeld 2015).
Within the core microbiome, the most common archaeal symbionts detected were all within the family Nitrosopumilaceae, known sponge symbionts (Zhang et al. 2014) and ammonia oxidizers (Könneke et al. 2005). Several species of Nitrosopumilus have been described as capable of utilizing urea as an ammonia source for energy generation and biomass production (Qin et al. 2017). Another top OTU in the core microbiome of both clusters was classified within the genus Nitrospira of the bacterial phylum Nitrospirae, representing known bacterial nitrifiers (Van Kessel et al. 2015). Further, OTUs of the family Woesceiaceae and specifically of the genus JTB255 have putative links to denitrification (Mußmann et al. 2017). These three OTUs correspond to microbes directly involved with the nitrogen cycle, providing further evidence of the important role sponges may play in this biogeochemical cycle (Hoffmann et al. 2009; Fiore et al. 2015). Other abundant OTUs included cyanobacterial symbionts that may provide fixed carbon to their sponge hosts through photosynthesis (Erwin and Thacker 2008) and Poribacteria, which are well-known sponge symbionts (Fieseler et al. 2004) that have putative links to organic phosphorous consumption (Kamke et al. 2014) and vitamin B12 synthesis (Siegl et al. 2011). Finally, Myxococcales are known sponge symbionts that produce abundant antibiotically active secondary metabolites (Hoffmann et al. 2013), that may confer anti-fouling or other defense capabilities to the sponge holobiont.
Significant structural differences between microbial communities of each cluster were also evident and may manifest in functional consequences that contribute to the observed genetic shift in host sponge populations. Importantly, these microbiome differences were not a function of sponge size, a potential confounding variable as Cluster 1 sponges skew towards larger sizes (Deignan et al. 2018). The majority (> 50%) of microbial community dissimilarity between genetic clusters of X. muta could be explained by variation in the relative abundance of a proportionally small number of OTUs, including all of the top 25 most abundant universal core OTUs described above. For instance, a top core OTU within Cluster 1 sponges corresponded to a proteobacterial species within the genus Albidovulum (OTU00030), a thermophilic lineage most often described in hot spring environmental samples (Travassos et al. 2002; Yin et al. 2012). While this particular OTU was not even within the top 50 OTUs for Cluster 2, Cluster 2 sponges did possess a different Albidovulum lineage (OTU00050) within their top 50 most abundant core OTUs, suggesting possible co-divergence of X. muta lineages and Albidovulum symbionts. Although thermophilic symbionts have been proposed to provide some temperature tolerance capabilities in other marine invertebrate hosts (Lee et al. 2008), such benefits have been suggested to result from physical insulation rather than increased host tolerance (Lee et al. 2008). Another major driver of differences between the two clusters was an OTU within the genus Cenarchaeum, a Thaumarchaeota lineage (OTU00083). This genus currently has only 3 described species, with the best-studied being the psychrophilic C. symbiosum, which is thought to associate exclusively with a single sponge species (Preston et al. 1996), and may benefit the host sponge through B-vitamin synthesis (Hallam et al. 2006; Fan et al. 2012). Genes associated with B-vitamin production have been detected in X. muta metagenomes (Fiore et al. 2015), and this particular symbiont exhibited a greater average abundance in Cluster 2 sponges. Variable functional contributions to host physiology may differentially impact fitness between lineages, thereby representing a potential selective force behind the apparent genetic shift occurring on Caribbean reefs that seems to paradoxically favor the recruitment of Cluster 2 sponges. Further, these population shifts may have cascading impacts on community-level processes like nutrient cycling and other important sponge contributions to ecosystem functioning. Additional studies validating functional predictions from taxonomic data (e.g., metatranscriptomic comparisons of the two clusters) and testing hypothesized relationships between symbiont function and host fitness are required to determine the ecological relevance of this apparent population shift.
In summary, we found that single-gene sequencing of the I3-M11 partition of the mitochondrial COI gene easily differentiates between the two genetic clusters defined by Deignan et al. (2018) using microsatellite markers, and thus represents an important molecular tool for future ecological studies of X. muta populations (e.g., correlating reproductive timing and host genotype). We mapped these genetic clusters to opposite ends of a global barrel sponge network, revealing high haplotype divergence despite their sympatric occurrence. We also found that the symbiotic microbial communities of X. muta exhibit significant differences in community structure and diversity between, but not within, these two genetic clusters. We detected numerous OTUs that represent well-known sponge symbionts and that may potentially provide a multitude of benefits to their host. These lineage-specific differences in the structure and putative functionality of microbial communities provide preliminary evidence for a microbial-mediated driver of genetic shifts in X. muta populations currently occurring on Florida reefs.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Aronson RB, Precht WF, Toscano MA, Koltes KH (2002) The 1998 bleaching event and its aftermath on a coral reef in Belize. Mar Biol 141:435–447
Bell JJ, Davy SK, Jones T, Taylor MW, Webster NS (2013) Could some coral reefs become sponge reefs as our climate changes? Glob Chang Biol 19:2613–2624
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci 108:4516–4522
Cebrian E, Uriz MJ, Turon X (2007) Sponges as biomonitors of heavy metals in spatial and temporal surveys in northwestern Mediterranean: Multispecies comparison. Environ Toxicol Chem 26:2430–2439
Chak ST, Rubenstein DR (2019) Social transitions in sponge-dwelling snapping shrimp. Curr Opin Insect Sci 34:33–39
Chambers K, Padovan A, Alvarez B, Gibb K (2013) Microbial signatures can help distinguish moon sponges (family Tetillidae) from Darwin Harbour, Australia. Mar Freshw Res 64:716–725
Cuvelier ML, Blake E, Mulheron R, McCarthy PJ, Blackwelder P, Thurber RLV, Lopez JV (2014) Two distinct microbial communities revealed in the sponge Cinachyrella. Front Microbiol 5
de Goeij JM, Van Oevelen D, Vermeij MJA, Osinga R, Middelburg JJ, De Goeij AFPM, Admiraal W (2013) Surviving in a marine desert: The sponge loop retains resources within coral reefs. Science (80- ) 342:108–110
Deignan LK, Pawlik JR, López-Legentil S (2018) Evidence for shifting genetic structure among Caribbean giant barrel sponges in the Florida Keys. Mar Biol 165
Diaz MC, Rützler K (2001) Sponges: An essential component of Caribbean coral reefs. Bull Mar Sci 69:535–546
Easson CG, Thacker RW (2014) Phylogenetic signal in the community structure of host-specific microbiomes of tropical marine sponges. Front Microbiol 5
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Erpenbeck D, Breeuwer JAJ, Van der Velde HC, Van Soest RWM (2002) Unravelling host and symbiont phylogenies of halichondrid sponges (Demospongiae, Porifera) using a mitochondrial marker. Mar Biol 141:377–386
Erwin PM, Thacker RW (2008) Phototrophic nutrition and symbiont diversity of two Caribbean sponge-cyanobacteria symbioses. Mar Ecol Prog Ser 362:139–147
Erwin PM, López-Legentil S, González-Pech R, Turon X (2012a) A specific mix of generalists: Bacterial symbionts in Mediterranean Ircinia spp. FEMS Microbiol Ecol 79:619–637
Erwin PM, Pita L, López-Legentil S, Turon X (2012b) Stability of sponge-associated bacteria over large seasonal shifts in temperature and irradiance. Appl Environ Microbiol 78:7358–7368
Erwin PM, Coma R, López-Sendino P, Serrano E, Ribes M (2015) Stable symbionts across the HMA-LMA dichotomy: Low seasonal and interannual variation in sponge-associated bacteria from taxonomically diverse hosts. FEMS Microbiol Ecol 91
Fan LL, Reynolds D, Liu M, Stark M, Kjelleberg S, Webster NS, Thomas T (2012) Functional equivalence and evolutionary convergence in complex communities of microbial sponge symbionts. Proc Natl Acad Sci U S A 109:E1878–E1887
Fieseler L, Horn M, Wagner M, Hentschel U (2004) Discovery of the novel candidate phylum “Poribacteria” in marine sponges. Appl Environ Microbiol 70:3724–3732
Fiore CL, Jarett JK, Lesser MP (2013) Symbiotic prokaryotic communities from different populations of the giant barrel sponge, Xestospongia muta. Microbiologyopen 2:938–952
Fiore CL, Labrie M, Jarett JK, Lesser MP (2015) Transcriptional activity of the giant barrel sponge, Xestospongia muta Holobiont: Molecular evidence for metabolic interchange. Front Microbiol 6
Friedrich AB, Fischer I, Proksch P, Hacker J, Hentschel U (2001) Temporal variation of the microbial community associated with the Mediterranean sponge Aplysina aerophoba. FEMS Microbiol Ecol 38:105–113
Fuhrman JA (2009) Microbial community structure and its functional implications. Nature 459:193–199
Gloeckner V, Hentschel U, Ereskovsky AV, Schmitt S (2013) Unique and species-specific microbial communities in Oscarella lobularis and other Mediterranean Oscarella species (Porifera: Homoscleromorpha). Mar Biol 160:781–791
Goffredi SK, Johnson SB, Vrijenhoek RC (2007) Genetic diversity and potential function of microbial symbionts associated with newly discovered species of Osedax polychaete worms. Appl Environ Microbiol 73:2314–2323
Griffiths SM, Antwis RE, Lenzi L, Lucaci A, Behringer DC, Butler IV MJ, Preziosi RF (2019) Host genetics and geography influence microbiome composition in the sponge Ircinia campana. J Anim Ecol
Hallam SJ, Mincer TJ, Schleper C, Preston CM, Roberts K, Richardson PM, DeLong EF (2006) Pathways of carbon assimilation and ammonia oxidation suggested by environmental genomic analyses of marine Crenarchaeota. PLoS Biol 4:520–536
Hentschel U, Hopke J, Horn M, Friedrich AB, Wagner M, Hacker J, Moore BS (2002) Molecular evidence for a uniform microbial community in sponges from different oceans. Appl Environ Microbiol 68:4431–4440
Hoffmann F, Radax R, Woebken D, Holtappels M, Lavik G, Rapp HT, Schläppy ML, Schleper C, Kuypers MMM (2009) Complex nitrogen cycling in the sponge Geodia barretti. Environ Microbiol 11:2228–2243
Hoffmann T, Müller S, Nadmid S, Garcia R, Müller R (2013) Microsclerodermins from terrestrial myxobacteria: An intriguing biosynthesis likely connected to a sponge symbiont. J Am Chem Soc 135:16904–16911
Kamke J, Rinke C, Schwientek P, Mavromatis K, Ivanova N, Sczyrba A, Woyke T, Hentschel U (2014) The candidate phylum Poribacteria by single-cell genomics: New insights into phylogeny, cell-compartmentation, eukaryote-like repeat proteins, and other genomic features. PLoS One 9
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA (2005) Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437:543–546
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120
Lee OO, Lau SCK, Qian PY (2006) Consistent bacterial community structure associated with the surface of the sponge Mycale adhaerens Bowerbank. Microb Ecol 52:693–707
Lee CK, Cary SC, Murray AE, Daniel RM (2008) Enzymic approach to eurythermalism of Alvinella pompejana and its episymbionts. Appl Environ Microbiol 74:774–782
Loh T-L, Pawlik JR (2014) Chemical defenses and resource trade-offs structure sponge communities on Caribbean coral reefs. Proc Natl Acad Sci 111:4151–4156
López-Legentil S, Pawlik JR (2009) Genetic structure of the Caribbean giant barrel sponge Xestospongia muta using the I3–M11 partition of COI. Coral Reefs 28:157–165
López-Legentil S, Erwin PM, Pawlik JR, Song B (2010) Effects of sponge bleaching on ammonia-oxidizing Archaea: Distribution and relative expression of ammonia monooxygenase genes associated with the barrel sponge Xestospongia muta. Microb Ecol 60:561–571
Lynch MDJ, Neufeld JD (2015) Ecology and exploration of the rare biosphere. Nat Rev Microbiol 13:217–229
Marlow J, Schönberg CHL, Davy SK, Haris A, Jompa J, Bell JJ (2019) Bioeroding sponge assemblages: the importance of substrate availability and sediment. J Mar Biol Assoc United Kingdom 99:343–358
McMurray SE, Blum JE, Pawlik JR (2008) Redwood of the reef: Growth and age of the giant barrel sponge Xestospongia muta in the Florida Keys. Mar Biol 155:159–171
McMurray SE, Henkel TP, Pawlik JR (2010) Demographics of increasing populations of the giant barrel sponge Xestospongia muta in the Florida Keys. Ecology 91:560–570
McMurray SE, Finelli CM, Pawlik JR (2015) Population dynamics of giant barrel sponges on Florida coral reefs. J Exp Mar Bio Ecol 473:73–80
Misof B, Erpenbeck D, Sauer KP (2000) Mitochondrial gene fragments suggest paraphyly of the genus Panorpa (Mecoptera, Panorpidae). Mol Phylogenet Evol 17:76–84
Montalvo NF, Hill RT (2011) Sponge-associated bacteria are strictly maintained in two closely related but geographically distant sponge hosts. Appl Environ Microbiol 77:7207–7216
Mußmann M, Pjevac P, Krüger K, Dyksma S (2017) Genomic repertoire of the Woeseiaceae/JTB255, cosmopolitan and abundant core members of microbial communities in marine sediments. ISME J 11:1276–1281
Neely KL, Butler CB (2020) Seasonal, lunar, and diel patterns in spawning by the giant barrel sponge, Xestospongia mutaCoral Reefs https://doi.org/10.1007/s00338-020-02009-2
Pawlik JR (1983) A Sponge-Eating Worm from Bermuda: Branchiosyllis oculata (Polychaeta, Syllidae). Mar Ecol 4:65–79
Pawlik TP, Henkel JR (2005) Habitat use by sponge-dwelling brittlestars. Mar Biol 146:301–313
Pawlik JR, McMurray SE (2020) The Emerging Ecological and Biogeochemical Importance of Sponges on Coral Reefs. Ann Rev Mar Sci 12:315–337
Pita L, Turon X, López-Legentil S, Erwin PM (2013) Host rules: Spatial stability of bacterial communities associated with marine sponges (Ircinia spp.) in the Western Mediterranean Sea. FEMS Microbiol Ecol 86:268–276
Pita L, Erwin PM, Turon X, López-Legentil S (2013a) Till death do us part: Stable sponge-bacteria associations under thermal and food shortage stresses. PLoS One 8
Powell A, Smith DJ, Hepburn LJ, Jones T, Berman J, Jompa J, Bell JJ (2014) Reduced diversity and high sponge abundance on a sedimented Indo-Pacific reef system: Implications for future changes in environmental quality. PLoS One 9
Preston CM, Wu KY, Molinski TF, DeLong EF (1996) A psychrophilic crenarchaeon inhabits a marine sponge: Cenarchaeum symbiosum gen. nov., sp. nov. Proc Natl Acad Sci U S A 93:6241–6246
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glöckner FO (2007) SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196
Qin W, Heal KR, Ramdasi R, Kobelt JN, Martens-Habbena W, Bertagnolli AD, Amin SA, Walker CB, Urakawa H, Könneke M, Devol AH, Moffett JW, Armbrust EV, Jensen GJ, Ingalls AE, Stahl DA (2017) Nitrosopumilus maritimus gen. nov., sp. nov., Nitrosopumilus cobalaminigenes sp. nov., Nitrosopumilus oxyclinae sp. nov., and Nitrosopumilus ureiphilus sp. nov., four marine ammonia-oxidizing archaea of the phylum Thaumarchaeot. Int J Syst Evol Microbiol 67:5067–5079
Reiswig HM (1974) Water transport, respiration and energetics of three tropical marine sponges. J Exp Mar Bio Ecol 14:231–249
Ruzicka R, Gleason DF (2008) Latitudinal variation in spongivorous fishes and the effectiveness of sponge chemical defenses. Oecologia 154:785–794
Schloss PD, Westcott SL (2011) Assessing and improving methods used in operational taxonomic unit-based approaches for 16S rRNA gene sequence analysis. Appl Environ Microbiol 77:3219–3226
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Schmitt S, Angermeier H, Schiller R, Lindquist N, Hentschel U (2008) Molecular microbial diversity survey of sponge reproductive stages and mechanistic insights into vertical transmission of microbial symbionts. Appl Environ Microbiol 74:7694–7708
Seemann J, Yingst A, Altieri AH, Edgar GJ, Stuart-Smith RD (2018) The importance of sponges and mangroves in supporting fish communities on degraded coral reefs in Caribbean Panama. PeerJ 6:e4455
Siegl A, Kamke J, Hochmuth T, Piel J, Richter M, Liang C, Dandekar T, Hentschel U (2011) Single-cell genomics reveals the lifestyle of Poribacteria, a candidate phylum symbiotically associated with marine sponges. ISME J 5:61–70
Simister R, Taylor MW, Tsai P, Webster N (2012) Sponge-microbe associations survive high nutrients and temperatures. PLoS One 7
Swierts T, Peijnenburg KTCA, de Leeuw CA, Breeuwer JAJ, Cleary DFR, de Voogd NJ (2017) Globally intertwined evolutionary history of giant barrel sponges. Coral Reefs 36:933–945
Swierts T, Cleary DFR, de Voogd NJ (2018) Prokaryotic communities of Indo-Pacific giant barrel sponges are more strongly influenced by geography than host phylogeny. FEMS Microbiol Ecol 94
Taylor MW, Schupp PJ, Dahllöf I, Kjelleberg S, Steinberg PD (2004) Host specificity in marine sponge-associated bacteria, and potential implications for marine microbial diversity. Environ Microbiol 6:121–130
Taylor MW, Radax R, Steger D, Wagner M (2007) Sponge-associated microorganisms: Evolution, ecology, and biotechnological potential. Microbiol Mol Biol Rev 71:295–347
Thomas T, Moitinho-Silva L, Lurgi M, Björk JR, Easson C, Astudillo-García C, Olson JB, Erwin PM, López-Legentil S, Luter H, Chaves-Fonnegra A, Costa R, Schupp PJ, Steindler L, Erpenbeck D, Gilbert J, Knight R, Ackermann G, Victor Lopez J, Taylor MW, Thacker RW, Montoya JM, Hentschel U, Webster NS (2016) Diversity, structure and convergent evolution of the global sponge microbiome. Nat Commun 7:1–12
Travassos P, Albuquerque L, Santos J, Silva MT, Nobre MF, da Costa MS, Rainey FA, Wait R, Empadinhas N (2002) Albidovulum inexpectatum gen. nov., sp. nov., a nonphotosynthetic and slightly thermophilic bacterium from a marine hot spring that is very closely related to members of the photosynthetic genus Rhodovulum. Appl Environ Microbiol 68:4266–4273
Uriz MJ, Turon X, Becerro MA, Galera J, Lozano J (1995) Patterns of resource allocation to somatic, defensive, and reproductive functions in the Mediterranean encrusting sponge Crambe crambe (Demospongiae, Poecilosclerida). Mar Ecol Prog Ser 124:159–170
Van Dam RP, Diez CE (1997) Predation by hawksbill turtles on sponges at Mona Island, Puerto Rico. Proceeding 8th Int Coral Reef Symp 2:1421–1426
Van Kessel MAHJ, Speth DR, Albertsen M, Nielsen PH, Op Den Camp HJM, Kartal B, Jetten MSM, Lücker S (2015) Complete nitrification by a single microorganism. Nature 528:555–559
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Warton DI, Wright ST, Wang Y (2012) Distance-based multivariate analyses confound location and dispersion effects. Methods Ecol Evol 3:89–101
White JR, Nagarajan N, Pop M (2009) Statistical methods for detecting differentially abundant features in clinical metagenomic samples. PLoS Comput Biol 5:e1000352
Wulff J, Buss LW (1979) Do sponges help hold coral reefs together? Nature 281:474–475
Yin D, Xiao J, Ao J, Ai C, Chen X (2012) Albidovulum xiamenense sp. nov., a moderately thermophilic bacterium from a terrestrial hot spring. Int J Syst Evol Microbiol 62:1609–1612
Zhang F, Pita L, Erwin PM, Abaid S, López-Legentil S, Hill RT (2014) Symbiotic archaea in marine sponges show stability and host specificity in community structure and ammonia oxidation functionality. FEMS Microbiol Ecol 90:699–707
Acknowledgements
This research was supported by a Grant from the National Science Foundation (NSF), Biological Oceanography Program, Number OCE-1558580. We thank the staff of FIU’s Aquarius Reef Base in Key Largo, Florida for their logistical support with fieldwork. We also thank S. McMurray and M. Valentine for their assistance with specimen collection. All research within the Florida Keys National Marine Sanctuary was conducted under Permit FKNMS-2009-126-A1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Topic Editor Carly Kenkel
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Evans, J.S., López-Legentil, S., Pawlik, J.R. et al. Molecular detection and microbiome differentiation of two cryptic lineages of giant barrel sponges from Conch Reef, Florida Keys. Coral Reefs 40, 853–865 (2021). https://doi.org/10.1007/s00338-021-02089-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00338-021-02089-8