Abstract
Understanding how genomic variation gives rise to phenotypic variation is essential for elucidating mechanisms of adaptive evolution, plant and animal breeding, and precision medicine. However, identifying causal links between DNA sequence variants and variation in phenotypes is challenging in human populations, due to large blocks of linkage disequilibrium in the genome and heterogeneous developmental histories, lifestyles, and social and physical environments. Drosophila melanogaster presents a powerful genetic model, since linkage disequilibrium decays rapidly, facilitating assignment of causality to polymorphisms associated with phenotypic variation, and large numbers of individuals can be reared under defined environmental conditions, economically, and without regulatory restrictions. The D. melanogaster Genetic Reference Panel (DGRP), a population of 205 sequenced, inbred wild-derived flies, has enabled genome-wide association studies of morphological, physiological, behavioral, and life history traits, and demonstrated that genetic architectures of complex traits are highly polygenic, sexually dimorphic, and context dependent with extensive sex-, environment-, and genetic background (epistatic) effects. These features together with a modular organization of the transcriptome illustrate a dynamic integrative genetic architecture for complex traits. The complexity of the genetic architectures for complex traits in Drosophila provides important caveats for the interpretation of genetic studies in human populations. Aspects of the genetic underpinnings of complex traits can be represented as simplified gene networks on which human orthologues can be superimposed to provide blueprints for subsequent studies on analogous traits in human populations. Fundamental principles of the genetic architectures of Drosophila complex traits are likely applicable across phyla, from the DGRP to human populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Genetic studies on human populations: trials and tribulations
Understanding the relationship between genotypic variation and variation in complex and quantitative trait phenotypes in human populations remains an ongoing challenge (Mackay 2014; Boyle et al. 2017). Yet, being able to make causal links between DNA variants and variation in organismal traits will result in a major leap forward to ultimately enable predicting both healthy phenotypes (e.g., lifespan, body weight) and susceptibility to disease.
Rare diseases that occur with high incidence within families or inbred populations and with high penetrance and that arise from single mutations with large effects can be subjected to classical linkage analysis in pedigrees to identify the causal DNA variant. However, most traits and common diseases have a complex genetic basis and their manifestation depends on multiple segregating genes, and interactions between them and environmental factors. Analyses of such traits require population-based association mapping approaches.
The classic measure that partitions phenotypic variation in genetic and environmental variance is the heritability (Falconer and Mackay 1996). The narrow sense heritability (h 2) is the portion of additive genetic variance that contributes to the total phenotypic variance that can be transmitted from parents to offspring. This measure of heritability is important for animal and plant breeding as it can be readily estimated from correlations among relatives and predicts response to artificial selection. The broad sense heritability (H 2) represents the fraction of phenotypic variance in a population that can be attributed to all sources of genetic variation, including both additive and non-additive (dominance, epistatic) genetic variance. High values of H 2 provide favorable scenarios for genome-wide association (GWA) analyses. However, even when H 2 is substantial, establishing causality of candidate polymorphisms to the phenotype in human GWA studies is challenging due to large blocks of linkage disequilibrium (LD)—the correlation in allele frequencies between polymorphic sites—in the human genome (Reich et al. 2001). Furthermore, genotype by environment interactions may result in lack of reproducibility between populations, when associations that are significant in a population in one environment cannot be resolved in a population under different environmental conditions. Gene–gene interactions (epistasis) may also result in failure to reproduce associations in different populations with different allele frequencies of causal loci (Mackay 2014, 2015). Moreover, when GWA analyses on individuals with vastly different genetic backgrounds are combined, spurious associations may be observed due to admixture (ethnic stratification; Pritchard and Rosenberg 1999). In addition, often the causal relationship between the trait and genes of unknown function or without biological context remains enigmatic.
The structure of the human genome, which contains regions of linkage disequilibrium, and differences in developmental history, lifestyle, and social and physical environments, requires phenotyping very large populations to resolve significant effects of candidate polymorphisms on quantitative traits (e.g., height; Wood et al. 2014) and susceptibility to common diseases (e.g., Zeggini et al. 2008). Such large population studies, often with tens of thousands of subjects, have identified many candidate genes that contain single nucleotide polymorphisms (SNPs) associated with common diseases, such as diabetes (Gaulton et al. 2015; Mohlke and Boehnke 2015) or cardiovascular disease (CARDIoGRAMplusC4D Consortium et al. 2013) as well as a vast number of copy number variants associated with schizophrenia (Stefansson et al. 2008, 2009). The concept of “common variants for common diseases” (Reich and Lander 2001), however, needs to be re-evaluated, as many rare variants with relatively large effects may cumulatively contribute to a significant fraction of disease risk within a population (Keinan and Clark 2012; Bomba et al. 2017).
Whereas certain human parameters, such as height, can be quantified precisely (Wood et al. 2014), many other phenotypes are difficult to quantify consistently. For example, evaluation of propensity to use alcohol or drugs often relies on self-reported questionnaires, and alcohol use and addiction may be confounded by psychiatric disorders or stress conditions. In addition, different investigators may use different criteria for alcohol use and addiction (Morozova et al. 2014). Furthermore, criminalization of illegal drug use makes it difficult to recruit subjects for GWA studies to explore genetic variants that may predispose to substance abuse. Genetic susceptibility to toxic exposure (e.g., heavy metals) is also hard to quantify as it is often confounded by exposure to multiple toxicants, and toxic effects may become evident long after the initial exposure. Similarly, cumulative effects of oxidative stress, for example, due to exposure to pesticides, which may lead to neurological disorders such as Parkinson’s disease (Dardiotis et al. 2013), are challenging to quantify. In addition, many human disease phenotypes, e.g., autism, are heterogeneous and present a spectrum of manifestations and severity, which may not necessarily arise from the same genetic risk factors (Chahrour et al. 2016). Finally, most human studies on genetic disease risk focus on extreme states, i.e., presence of disease, whereas clinical manifestation may result from surpassing a threshold of normal trait distribution within the population. For example, sociopathic aggression represents an extreme along the spectrum from assertiveness to shyness. Where to draw the threshold in this case is not unambiguous, but may be influenced by sociocultural boundaries.
Despite significant advances in studies on the genetic underpinnings of complex traits in human populations, both in health and disease, the challenges and impediments inherent in these studies illustrate a compelling need for comparative studies in model organisms. Studies on the genetic architectures of complex traits in Drosophila melanogaster have provided many insights with translational potential for human population genetics.
Drosophila: an advantageous model for the genetic dissection of complex traits
Many of the constraints encountered in human population studies can be overcome in the D. melanogaster genetic model system. We can inbreed flies, thus enabling strict control over the genetic background, and we can rear virtually unlimited numbers of genetically identical individuals. Drosophila has a 2-week generation interval under standard laboratory conditions and can be grown rapidly under well-controlled environmental conditions, without regulatory restrictions and at relatively low cost. We can readily substitute chromosomes, generate transgenic flies, and use CRISPR/Cas9 technology for gene editing, which allows us to create “designer” genotypes (Bassett and Liu 2014). About 75% of human disease-associated genes have a Drosophila ortholog (Reiter et al. 2001); and a wide range of morphological, physiological, behavioral, and life history traits, many with relevance to analogous human traits, can be quantified precisely. Hence, evolutionary conservation of fundamental biological principles empowers translational inferences across phyla, including humans.
The genetic architectures of complex traits can be explored by large-scale mutational screens, conventional quantitative trait locus (QTL) linkage mapping approaches, and GWA analyses. Large collections of Drosophila stocks with transposon insertions, such as P-element insertions, which often target promoter regions, or piggyBac inserts, which disrupt genes without promoter bias, are publicly available for mutational screens (Zhai et al. 2003; Thibault et al. 2004; Bellen et al. 2011). Transposon-based mutagenesis screens have shown that complex traits present large mutational targets, which are affected by mutations in many genes, often representing a substantial fraction of the genome, implicating extensive pleiotropy (Sambandan et al. 2006; Mensch et al. 2008; Magwire et al. 2010; Zwarts et al. 2011).
Whereas mutational screens identify genes that contribute to manifestation of the trait under study, linkage and association mapping analyses, respectively, identify a subset of those genes that harbor polymorphisms that contribute to variation in the phenotype. Genome-wide studies can identify candidate genes that contribute to phenotypic variation, while subsequent mutational analyses can provide evidence that disruption of the gene indeed affects the trait and provide a focus for detailed mechanistic studies. Thus, these approaches are complementary. In Drosophila, GWA analyses became possible with the development of the D. melanogaster Genetic Reference Panel (DGRP; Mackay et al. 2012; Huang et al. 2014).
The D. melanogaster Genetic Reference Panel (DGRP)
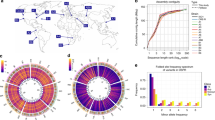
The DGRP is a population of 205 inbred wild-derived lines, which were generated by subjecting the offspring of individual gravid females collected from the Farmer’s Market in Raleigh, North Carolina, to 20 generations of full sib inbreeding (Mackay et al. 2012; Huang et al. 2014). Inbreeding minimizes genetic variation among individuals within each line, while genetic variation between the lines reflects the variation in the population from which they were derived (Fig. 1a). The lines were sequenced to high coverage (average ~ 27-fold) and a total of 4,565,215 naturally occurring molecular variants were identified, including 3,976,011 high-quality single or multiple nucleotide polymorphisms; 169,053 polymorphic insertions; 293,363 polymorphic deletions; and 125,788 polymorphic microsatellites. The vast majority of the DGRP lines are genetically unrelated, except for a few lines which may have resulted from random sampling of related individuals from the population (Fig. 1b). Residual heterozygosity in the lines is largely due to segregating polymorphic inversions, which represent islands of diversity within otherwise homozygous genomes. In addition, 53% of the DGRP lines are infected with the maternally transmitted endosymbiotic bacterium, Wolbachia pipientis (Huang et al. 2014).

Modified from Mackay et al. (2012)
The Drosophila melanogaster Genetic Reference Panel. a Diagram of the derivation of DGRP lines from a natural population. b Genomic relatedness among DGRP lines. The distribution of the relationship between all DGRP lines and the reference sequence is displayed as a box plot. From Huang et al. (2014). c Decay of LD (average R-squared) in bp as a function of physical distance.
It should be noted that lines that survived inbreeding have been purged of highly deleterious alleles and selection may have occurred for epistatic interactions among variants that protect fitness during inbreeding. Also, long-term maintenance in the laboratory may give rise to adaptation to the laboratory environment. Despite these limitations, the DGRP provides a rich collection of natural variants that have survived the sieve of natural selection and are a treasure trove of genetic variation that can be harnessed for GWA studies. All traits measured in the DGRP to date show extensive phenotypic variation that far exceeds variation observed among standard laboratory inbred strains or recombinant inbred lines. Since individuals within each line are genetically virtually identical, the same genotypes can be measured repeatedly, which enables accurate quantification of phenotypes. Since phenotypic measurements can be spread over time, environmental noise can be randomized, further minimizing error in phenotypic estimates. Furthermore, since the DGRP is a publicly available resource, different laboratories can measure and correlate phenotypes on the same genotypes. Table 1 lists GWA analyses performed to date using the DGRP.
One example of the translational potential between flies and humans is a study which examined genome-wide variation in gene expression with phenotypic variation in alcohol sensitivity across 40 DGRP lines followed by validation of candidate genes through transposon-mediated mutagenesis (Morozova et al. 2009). One gene that emerged as a particularly interesting candidate gene was Men, which encodes malic enzyme. Malic enzyme represents a metabolic switch between energy production and lipid biosynthesis. Since excessive alcohol consumption in people results in fatty liver syndrome, a subsequent study in the Framingham Heart Study Offspring Cohort population focused on the cytoplasmic Malic Enzyme 1 (ME1) gene and identified polymorphisms associated with variation in alcohol consumption, which would not have been otherwise detected in a GWA study in this limited size population (Morozova et al. 2009).
Another example of how studies on the DGRP can guide investigations on human disease-associated genes is the recent identification of candidate modifier genes in a Drosophila model for retinitis pigmentosa (Chow et al. 2016).
Statistical considerations for GWA analyses
Large LD blocks in the human genome enable the use of tagging SNPs for GWA studies (Reich et al. 2001; Ke et al. 2004). In contrast to the human genome, LD in the D. melanogaster genome decays rapidly with physical distance, on average within a few hundred base pairs (Fig. 1c; Mackay et al. 2012). The advantage of low LD is that associated SNPs are likely causal or very near the causal variant. However, low LD prevents the use of tagging SNPs and mandates whole genome sequences for GWA analyses. Because the X chromosome experiences a lower effective population size, LD decays more slowly on the X chromosome than on the autosomes, and LD is notably increased in areas of reduced recombination, i.e., in the vicinity of autosomal telomeres and centromeres, and in polymorphic inversions, where recombination is suppressed (Mackay et al. 2012). Furthermore, due to the restricted population size of the DGRP, variants with minor allele frequencies of less than 5% may exhibit elevated long-range LD, since such rare variants will be in perfect LD with all variants that are private to a particular DGRP line. Thus, polymorphisms with minor allele frequencies of less than 5% should be excluded from GWA analyses.
The gold standard for genome-wide statistical significance in human GWA studies is a Bonferroni-adjusted threshold for multiple testing. GWA analyses in the DGRP are confined to only 205 lines with several million tests. Consequently, we seldom observe SNPs that show associations with P values that exceed a strict Bonferroni-corrected threshold. However, quantile–quantile plots (Q–Q plots) can indicate deviations from linearity between the distributions of observed and predicted values, indicating enrichment of true positive associations at P values which are orders of magnitude below the Bonferroni threshold. For most GWA analyses in the DGRP, we can identify approximately several hundred polymorphisms associated with phenotypic variation at an empirical threshold of P < 10− 5, supported by Q–Q plots (Weber et al. 2012; Swarup et al. 2013; Harbison et al. 2013; Morozova et al. 2015; Shorter et al. 2015; Garlapow et al. 2015; Carbone et al. 2016). Since all polymorphisms in the DGRP are known, those polymorphisms represent the top SNPs and/or insertions–deletions, which are associated with variation in the trait under study.
Mutational analyses or targeted RNAi experiments can be used to assess whether candidate genes that harbor such associated polymorphisms themselves affect the trait when their expression is disrupted. Such validation on a sample of candidate genes for which viable mutants are available establishes an empirical false discovery rate at the gene level, albeit not at the level of molecular variants. Previous studies have shown that we can validate about 60–80% of candidate genes using this approach (Weber et al. 2012; Swarup et al. 2013; Harbison et al. 2013; Morozova et al. 2015; Shorter et al. 2015; Garlapow et al. 2015; Carbone et al. 2016). Within pleiotropic genes different SNPs can be associated with variation in specific organismal traits (Carbone et al. 2006; Wang et al. 2010).
We can assess to what extent candidate genes with associated polymorphisms can be organized in interaction networks based on known genetic or physical interactions, and we can estimate the likelihood that a similar size network would emerge if the same number of genes were chosen at random. Here, the P value for significance of the network is not encumbered by multiple test considerations. Moreover, genes that form networks show very low empirical false discovery rates and validate at a high rate when subjected to mutational analysis (Fig. 2). However, only a restricted subset of genes with associated polymorphisms can be interconnected to form a network, although some of the remaining genes can sometimes be implicated in similar biological processes represented by the network through gene ontology analyses.
Genetic networks for variation in olfactory behavior. a A network of interactions among candidate genes associated with variation in response to the odorant benzaldehyde among DGRP lines. Candidate genes are indicated by rectangles, missing genes (i.e., genes without significant associations) by triangles, and metabolites by circles. Components of the network associated with distinct interconnected cellular processes are highlighted by the colored backgrounds. b Validation of the connectivity of the predicted network. Transposon insertion in the Pkc53e locus results in a two-fold increase in gene expression accompanied by aberrant olfactory behavior. qRT-PCR of transcripts of genes connected in the network shown in panel a in a Pkc53e P{MiET1}-insertion mutant show increased expression levels compared to control, corroborating their functional connectivity. Error bars indicate SEM. *0.01 ≤ P ≤ 0.05; **0.001 ≤ P ≤ 0.01; ***0.0001 ≤ P ≤ 0.001. c An extended network of candidate genes associated with variation in responses to 14 structurally different odorants based on GWA and extreme QTL mapping analyses. Note the absence of missing genes in this network.
Extreme QTL mapping
We can increase the power of GWA analyses by generating advanced intercross populations (AIP) from DGRP lines. There are several strategies for generating such AIPs. We can select a small number of lines with extreme phenotypes and cross them for many generations (Swarup et al. 2013; Shorter et al. 2015); we can select a random number of DGRP lines (Huang et al. 2012; Morozova et al. 2015; Carbone et al. 2016); or, we can select DGRP lines that are genetically unrelated with minimal residual heterozygosity, and free of inversions and infection of the symbiotic bacterium, W. pipientis (Garlapow et al. 2017). The lines can be crossed in a round-robin crossing design (Fig. 3) or a diallel crossing scheme (Griffing 1956) to generate a base population. Many generations of intercrossing in large population sizes to minimize loss of diversity due to drift will result through recombination in a virtually unlimited number of unique genotypes. We can now select and pool individuals with extreme phenotypes, subject the pools to bulk DNA sequencing, and identify alleles that differentially segregate among the phenotypic extremes (Fig. 3). This experimental approach, known as “extreme QTL mapping” (Ehrenreich et al. 2010), resembles case–control studies in human genetics, where allele frequencies associated with disease status are contrasted with those among unaffected age-matched individuals.
The advantages of the AIP design are that alleles that are present at low frequencies in the DGRP and may have large phenotypic effects, but cannot be analyzed by conventional GWA due to the risk of spurious LD, will be present at higher frequencies in the AIP base population, and hence their effects can be assessed. The diversity of genotypes generated through intercrossing increases statistical power, provided the pools for bulk DNA sequencing contain sufficiently large numbers of individuals. Indeed, extreme QTL mapping studies using DGRP-derived AIPs can resolve trait-associated variants that surpass a Bonferroni threshold for multiple tests. It is noted, however, that generating an AIP results in loss of polymorphisms that are not represented among the lines used to generate the base population. Nevertheless, AIPs derived from ~ 40 DGRP lines still captured a substantial amount of the genetic variation present in the DGRP (Huang et al. 2012).
From flies to human genetics: common concepts
The transcriptional niche
Genes do not act in isolation, but form part of functional expression networks. Introduction of a mutation that alters gene expression can result in altered transcript abundances of a suite of genes in addition to that of the target gene (Anholt et al. 2003). Analyses of correlations among transcripts of 40 DGRP lines reared under standard growth conditions showed a modular organization of the transcriptome, in which 10,096 genetically variable transcripts could be clustered in 241 modules, such that the genetic correlation of transcripts within each module was maximized and the genetic correlation of transcripts between modules minimized (Ayroles et al. 2009). This modular organization, however, is dynamic and changes under different environmental conditions (Zhou et al. 2012). Analyses of correlated transcript abundances with variation in gene expression of a focal gene define a group of correlated transcripts, the size of which depends on the statistical correlation threshold. We have designated such a modular array around a focal gene as the gene’s “transcriptional niche” (Fig. 4a; Arya et al. 2010). Gene ontology analyses of transcriptional niches can provide functional contexts to genes of unknown function or indicate pleiotropy of the focal gene (Fig. 4b, c).
The transcriptional niche. a Gene X gives rise to transcript X the abundance of which is correlated with that of other transcripts, indicated by circles of which the color shades indicate different statistical levels of correlation. b Different segments of the transcriptional niche of gene X can be recruited to give rise to different phenotypes, indicating that the transcriptional niche can serve as a conduit for pleiotropy. c The transcriptional niche is plastic and can change, for example, as a function of developmental stage, accompanied by a switch in its relationship to the organismal phenotype, as illustrated in this diagram, where purple circles indicate transcripts that are uniquely associated with the transcriptional niche of gene X in adulthood. d Transcriptional niches of different genes can overlap and interact to modify the organismal phenotypes associated with genes X and Y and give rise to different phenotypes, here illustrated as phenotype Z
The concept of the transcriptional niche is of relevance to human genetics, but often underappreciated. Identification of a SNP associated with a disease phenotype does not necessarily mean that the gene harboring this SNP is directly causal to the phenotype, since its effect may be mediated through a shift in its transcriptional niche, such that modulation of expression of a different gene may actually be causal to the phenotype (Fig. 4d).
Epistasis, an inconvenient truth
Early QTL mapping studies on D. melanogaster recombinant inbred lines provided evidence for epistasis (Long et al. 1995; Leips and Mackay 2000; Dilda and Mackay 2002; Montooth et al. 2003). Subsequent studies on olfactory behavior, sleep, and waking activity, using chromosome substitution lines in which different chromosomes extracted from DGRP lines were introduced in a common genetic background with and without single P-element insertional mutations, provided evidence for suppressing epistasis, i.e., epistatic interactions tend to suppress the effects of new mutations (Yamamoto et al. 2009; Swarup et al. 2012). However, it remained unclear how prominent the contribution of epistasis is to the genetic architecture of complex traits.
When results from GWA analyses in the DGRP on startle behavior, recovery time from a chill-induced coma, and resistance to starvation stress were compared to the results from extreme QTL mapping experiments in DGRP-derived AIPs, SNPs identified by extreme QTL mapping were different from those identified by GWA analysis for all three traits (Huang et al. 2012). However, gene ontology and network analyses revealed that candidate genes implicated by both analyses converged on the same biological processes. Pairwise analyses of epistasis revealed extensive networks of epistatic interactions connecting results from the two analyses (Huang et al. 2012). A similar result emerged from studies on the genetic underpinnings of olfactory behavior, where combined GWA and extreme QTL mapping analyses implicated a network of genes associated with neural development and function as a substrate for variation in the behavioral phenotype (Swarup et al. 2013). This observation was extended and further confirmed by a GWA analysis that identified epistatic partners which interact with transposon-tagged mutants of two neurogenic genes, Sema-5c and neuralized, both of which have large effects on olfactory behavior (He et al. 2016). Similarly, transgenic expression of a misfolded mutant of human preproinsulin, proposed as a model for neonatal diabetes mellitus, causes morphological defects in adult flies; when crossed to different DGRP lines, F1 offspring displays a continuous range of morphological phenotypes, indicating genetic background-dependent modulation of the effect of the transgene (Park et al. 2014). Epistatic interactions between SNPs identified through GWA analysis and SNPs identified through extreme QTL mapping were also identified and validated in a study on variation in aggression (Shorter et al. 2015). These studies, along with others (Sanjuán and Elena 2006; Corbett-Detig et al. 2013; Chow et al. 2016), show convincingly that epistasis is a prevalent determinant of the genetic architecture of complex traits.
Lack of replication of associated polymorphisms between GWA studies in the DGRP and extreme QTL mapping analyses in the AIP is most likely due to the different frequencies of causal molecular variants in the two populations; note that this will almost always occur since the AIP populations are derived from a small subset of DGRP lines. A hallmark of epistatic interactions is that the effect of a focal locus on a quantitative trait depends on the allele frequencies at the interacting locus/loci (Fig. 5; Mackay 2015). This is not true for additive interactions among loci. Epistatic interactions are also sensitive to environmental conditions and are likely to affect the composition of the transcriptional niches of interacting partners, causing higher-order ripple effects. Thus, epistasis might be a confounding factor for the interpretation of GWA studies in human populations. For example, well-executed studies with sufficient statistical power to replicate associated variants in an independent population often fail to replicate the initial findings. However, lack of replication is actually expected in the presence of epistasis when populations have different allele frequencies at the genotyped loci (Greene et al. 2009; Moskvina et al. 2011; Mackay 2015). Disease-associated polymorphisms that are ignored because they do not replicate in different populations may nevertheless contribute important disease risk to members of the population in which they could be identified.
Epistasis depends on allele frequencies of interacting partners. a, b Absence of epistatic effects between alleles of locus B and alleles of locus A. c Differential epistatic effects of alleles B1 and B2 with alleles of locus A and d the dependence of the effect of locus A on the frequency of the B1 allele. e Antagonistic epistatic interactions between the B1 (enhancing epistasis) and B2 (suppressing epistasis) alleles with locus A and f the resulting dependence of the effect of locus A on the frequency of the B1 allele
Although epistasis has been recognized in genetic studies on human populations (Webber 2017), it is often dismissed in analyses, because assessing pairwise epistatic interactions genome-wide is subject to a huge multiple testing penalty such that only interactions with very large effects could be detected. However, genetic variants uncovered in extensively studied human traits (a notable example is height; Wood et al. 2014) account for only a small fraction of the heritability (Manolio et al. 2009). Epistasis is one factor that can contribute to this “missing heritability” (Zuk et al. 2012).
Sexually dimorphic architecture of complex traits
Virtually every complex trait examined to date by mutational analyses, QTL analysis, GWA analysis, or extreme QTL mapping shows extensive genetic variation in sexual dimorphism, often with little overlap between trait-associated polymorphic markers of males and females (Fig. 6; Weber et al. 2012; Swarup et al. 2013; Harbison et al. 2013; Morozova et al. 2015; Garlapow et al. 2015; Zhou et al. 2016; Carbone et al. 2016). This is perhaps not surprising since extensive sexual dimorphism and genetic variation in sexual dimorphism is also apparent in genome-wide transcript abundances across the DGRP (Ayroles et al. 2009; Huang et al. 2015). In addition to inherent sex bias, environmental or genetic effects on gene expression may show sex specificity or sexually antagonistic modulation.

Adapted from Harbison et al. (2013)
Sexual dimorphism is prevalent among phenotypes measured in the DGRP. The example of sexual dimorphism shown here is phenotypic variation in sleep duration. Blue bars in the histograms in panels a and c denote male line means and purple bars denote female line means. The difference in line means between males and females (male–female) is shown in b and d. a Night sleep. b Male–female night sleep. c Day sleep. d Male–female day sleep.
It is well known that disease susceptibility varies between men and women in human populations. It is, therefore, essential to include sex as a variable in analyses of GWA data (Golden and Voskuhl 2017). Failure to do so can reduce statistical power if a DNA variant affects a phenotype in one sex only or can lead to the erroneous conclusion that a particular variant poses a disease risk in both sexes. Considerations of sexual dimorphism are, therefore, essential for the development of genetic information-based precision medicine.
Genes and the environment
Phenotypic plasticity is the ability of a genotype to give rise to different phenotypes under different environmental conditions (Fig. 7). Studies in which an AIP derived from 40 DGRP lines was subjected to a wide range of different exposures showed that most of the Drosophila transcriptome is robust in the face of environmental changes, while the environmentally sensitive transcriptome includes genetically variable transcripts associated with detoxification, metabolism, proteolysis, heat shock proteins, and transcriptional regulation (Zhou et al. 2012).
Phenotypic plasticity and genotype by environment interaction. The diagram shows schematic reaction norms for two genotypes in two environments (Env). Phenotypic plasticity is evident when the reaction norms are parallel, i.e., the two genotypes respond similarly to a change in environment. Genotype by environment interaction is evident when the genotypes respond to different extents or in opposite directions to environmental change, or when one genotype responds to environmental change, whereas another is unaffected
Genotype by environment interaction is due to genetic variation in phenotypic plasticity, i.e., different genotypes respond differently in response to environmental changes (Fig. 7). Some examples of genotype by environment interactions have been documented in studies of human populations. A classic study by Caspi et al. (2002) showed that a variable number tandem repeat polymorphism at the promoter of the MAOA gene was associated with violent behavior only in individuals who had experienced maltreatment as children. Furthermore, this effect was sex-specific and only observed in males. Similarly, polymorphisms in the promoter of the serotonin transporter gene have been associated with depression, dependent on stressful life experiences (Caspi et al. 2003; Rocha et al. 2015). These well-executed studies have not been universally corroborated (Munafò et al. 2009), potentially due to epistasis or differences in environmental sensitivity of allelic effects in different populations, as mentioned earlier. A well-established example of genotype by environment interactions is the observation that the effects of polymorphisms in immune response genes, including members of the interleukin family, associated with risk for asthma, are dependent on previous exposure to allergens (Sordillo et al. 2015; Bønnelykke and Ober 2016; Li et al. 2016).
Genotype by environment interaction is notably evident upon exposure to toxins. The Drosophila model is eminently suitable as a model system for population-based large-scale toxicogenomic studies. Several studies have focused on the genetic factors that underlie individual variation in susceptibility to heavy metals, such as cadmium, lead, and methylmercury (Akins et al. 1992; Hirsch et al. 2012). Metallothioneins are small cysteine-rich proteins that can bind ingested toxic heavy metals, notably cadmium, and contribute to their long persistence following exposure (Isani and Carpene 2014). In Drosophila, levels of metallothionein contribute to differences in sensitivity to cadmium between different strains (Gill et al. 1989), but cadmium sequestration by metallothionein is not the only factor that is responsible for cadmium resistance (Nguyen et al. 2014), indicating that other mechanisms contribute to cadmium toxicity.
Flies reared on low concentrations of lead acetate show changes in courtship, fecundity, and locomotor activity (Hirsch et al. 2003). A study on recombinant inbred lines, which used expression microarrays to identify cis-eQTL (i.e., where a local gene controls its own transcription) and trans-eQTL (i.e., where the activity of distant genes controls transcription) that were differentially expressed among control and lead-exposed flies, identified a co-regulated ensemble of 33 lead-induced genes, many of which are associated with neurodevelopment (Ruden et al. 2009). A study on lead sensitivity across the DGRP showed similar results. DGRP flies showed different effects on development time and adult locomotion when reared on medium supplemented with lead acetate, and gene ontology and network analyses showed enrichment of genes associated with early development and function of the nervous system (Zhou et al. 2016). Observations in the Drosophila model can guide studies of lead toxicity in children, which leads to neurological and cognitive deficits (Canfield et al. 2003; Jakubowski 2011; Liu and Lewis 2014; McDermott et al. 2014), by suggesting candidate pathways or cellular mechanisms.
A subsequent extreme QTL mapping study on a DGRP-derived AIP identified allelic variants associated with sensitivity to lead and cadmium exposure (Zhou et al. 2017). This study revealed genetic networks on which human counterparts of Drosophila genes could be superimposed. Human genes previously implicated in heavy metal toxicity could be placed in biological context along with identification of novel targets for heavy metal toxicity. This study showed that evolutionary conservation of fundamental biological processes enables Drosophila to serve as a translational model for toxicogenomics studies.
Methylmercury, a toxic heavy metal that can accumulate in seafood, has detrimental effects on the developing nervous system by interfering with the Notch receptor pathway (Bland and Rand 2006; Alattia et al. 2011; Engel et al. 2012). A GWA study on susceptibility to methylmercury exposure of DGRP lines identified candidate genes involved in muscle and neuromuscular development, and pupae exposed to methylmercury showed disrupted development of indirect flight muscle (Montgomery et al. 2014). In addition to effects on neural development, these observations implicate effects on muscle development as a consequence of methylmercury exposure. Glutamate cysteine ligase provides protection against methylmercury and overexpression of this enzyme in muscle-rescued eclosion of flies reared on methylmercury-supplemented medium. Mutations in kirre, a myogenic gene identified as a candidate gene associated with variation in methylmercury sensitivity, modulated eclosion rates upon exposure to methylmercury (Montgomery et al. 2014).
Drosophila has also proven to be a valuable model to study genetic variation in susceptibility to environmental oxidative stress agents, notably the highly toxic herbicide paraquat, which has been implicated as a causative agent for Parkinson’s disease, characterized by degeneration of dopaminergic neurons. Paraquat neurotoxicity in Drosophila also affects dopaminergic neurons (Martin et al. 2014) and expression of the dopamine receptor (Cassar et al. 2015). However, GWA studies on DGRP lines show that the genetic architecture that determines variation in susceptibility to paraquat and other oxidative stressors is highly polygenic. There is extensive variation in sensitivity to acute exposure to two oxidative stress-inducing agents, paraquat and menadione sodium bisulfite with little overlap of SNPs associated with variation in sensitivity to these compounds (Weber et al. 2012). GWA analyses of DGRP lines exposed to chronic oxidative stress induced by menadione sodium bisulfite revealed a network of candidate genes on which human orthologues could be superimposed. These candidate genes were associated with inositol triphosphate signaling and synaptic transmission, intermediary metabolism, signaling by NGF, EGFR, and Rho GTPases, and DNA replication (Jordan et al. 2012). These studies further underscore the translational potential of the Drosophila model as they highlight molecular processes that are conserved across phyla.
Orthologous networks
As mentioned earlier, we can assemble candidate genes identified through GWA analyses of the DGRP and/or extreme QTL mapping analyses of DGRP-derived AIPs into networks (Fig. 2). By superimposing human orthologues on these Drosophila genetic networks, we can build a translational blueprint to target candidate genes for subsequent focused studies in human populations (Fig. 8; Jordan et al. 2012; Zhou et al. 2016, 2017; Carbone et al. 2016). This approach is based on the principle of evolutionary conservation of fundamental biological processes and has several advantages. First, the Drosophila network provides a functional biological context for its human orthologue. Second, association of a gene with a trait in Drosophila, especially if it is a hub gene in the network, increases the chances for discovering an association of its orthologue with an analogous phenotype in a human population. Third, by identifying candidate genes with human orthologues through genome-wide screens in Drosophila, one can focus a subsequent association study on a single orthologue, which greatly reduces the multiple testing problem and, thus, increases statistical power.
A genetic network for susceptibility to lead exposure. The network was derived from candidate genes identified in GWA analyses for development time, viability, and activity of flies exposed to lead acetate. Yellow square boxes indicate candidate genes associated with any of these traits, while gray ovals represent computationally recruited intermediate genes. Blue font indicates genes with human orthologues. Note the extent by which human orthologues can be superimposed on their Drosophila counterparts to identify potential human candidate genes that may contribute to susceptibility to lead toxicity. From Zhou et al. (2016)
Conclusion
GWA studies in Drosophila highlight the importance of sex-, environment-, and genetic background-dependent (epistatic) effects, modularity of gene expression, pleiotropy, and interactions among these parameters, which give rise to a dynamic integrative genetic architecture for complex traits. Understanding the genetic mechanisms that lead to the manifestation of genotype by environment interactions is critical for human health and developing genetics-informed precision medicine. Elucidating these mechanisms requires comprehensive empirical and computational approaches which integrate DNA sequence variation with variation at the levels of the transcriptome, proteome, and metabolome, while accounting for the complexity and dynamics of epistatic interactions, genome–environment interactions, and plasticity of transcriptional niches. D. melanogaster is the most favorable model organism for the pursuit of such a systems genetics endeavor. Principles derived from studies on this powerful genetic model system are universal and apply across phyla, from the DGRP to human populations.
References
Akhund-Zade J, Bergland AO, Crowe SO, Unckless RL (2017) The genetic basis of natural variation in Drosophila (Diptera: Drosophilidae) virgin egg retention. J Insect Sci 17:pii 5
Akins JM, Schroeder JA, Brower DL, Aposhian HV (1992) Evaluation of Drosophila melanogaster as an alternative animal for studying the neurotoxicity of heavy metals. Biometals 5:111–120
Alattia JR, Kuraishi T, Dimitrov M, Chang I, Lemaitre B, Fraering PC (2011) Mercury is a direct and potent γ-secretase inhibitor affecting Notch processing and development in Drosophila. FASEB J 25:2287–2295
Anholt RRH, Dilda CL, Chang S, Fanara JJ, Kulkarni NH, Ganguly I, Rollmann SM, Kamdar KP, Mackay TFC (2003) The genetic architecture of odor-guided behavior in Drosophila: epistasis and the transcriptome. Nat Genet 35:180–184
Appel M, Scholz CJ, Müller T, Dittrich M, König C, Bockstaller M, Oguz T, Khalili A, Antwi-Adjei E, Schauer T, Margulies C, Tanimoto H, Yarali A (2015) Genome-wide association analyses point to candidate genes for electric shock avoidance in Drosophila melanogaster. PLoS ONE 10:e0126986
Arya GH, Weber AL, Wang P, Magwire MM, Serrano Negron YL, Mackay TFC, Anholt RRH (2010) Natural variation, functional pleiotropy and transcriptional networks of Odorant binding protein genes in Drosophila melanogaster. Genetics 186:1475–1485
Arya GH, Magwire MM, Huang W, Serrano-Negron YL, Mackay TFC, Anholt RRH (2015) The genetic basis for variation in olfactory behavior in Drosophila melanogaster. Chem Senses 40:233–243
Ayroles JF, Carbone MA, Stone EA, Jordan KW, Lyman RF, Magwire MM, Rollmann SM, Duncan LH, Lawrence F, Anholt RRH, Mackay TFC (2009) Systems genetics of complex traits in Drosophila melanogaster. Nat Genet 41:299–307
Ayroles JF, Buchanan SM, O’Leary C, Skutt-Kakaria K, Grenier JK, Clark AG, Hartl DL, de Bivort BL (2015) Behavioral idiosyncrasy reveals genetic control of phenotypic variability. Proc Natl Acad Sci USA 112:6706–6711
Bassett AR, Liu JL (2014) CRISPR/Cas9 and genome editing in Drosophila. J Genet Genomics 41:7–19
Battlay P, Schmidt JM, Fournier-Level A, Robin C (2016) Genomic and transcriptomic associations identify a new insecticide resistance phenotype for the selective sweep at the Cyp6g1 locus of Drosophila melanogaster. G3 6:2573–2581
Bellen HJ, Levis RW, He Y, Carlson JW, Evans-Holm M, Bae E, Kim J, Metaxakis A, Savakis C, Schulze KL, Hoskins RA, Spradling AC (2011) The Drosophila gene disruption project: progress using transposons with distinctive site specificities. Genetics 188:731–743
Bland C, Rand MD (2006) Methylmercury induces activation of Notch signaling. Neurotoxicology 27:982–991
Bomba L, Walter K, Soranzo N (2017) The impact of rare and low-frequency genetic variants in common disease. Genome Biol 18:77
Bønnelykke K, Ober C (2016) Leveraging gene-environment interactions and endotypes for asthma gene discovery. J Allergy Clin Immunol 137:667–679
Bou Sleiman MS, Osman D, Massouras A, Hoffmann AA, Lemaitre B, Deplancke B (2015) Genetic, molecular and physiological basis of variation in Drosophila gut immunocompetence. Nat Commun 6:7829
Boyle EA, Li YI, Pritchard JK (2017) An expanded view of complex traits: from polygenic to omnigenic. Cell 169:1177–1186
Brown EB, Layne JE, Zhu C, Jegga AG, Rollmann SM (2013) Genome-wide association mapping of natural variation in odour-guided behaviour in Drosophila. Genes Brain Behav 12:503–515
Bushnell PJ, Ward WO, Morozova TV, Oshiro WM, Lin MT, Judson RS, Hester SD, McKee JM, Higuchi M (2017) Editor’s highlight: genetic targets of acute toluene inhalation in Drosophila melanogaster. Toxicol Sci 156:230–239
Canfield RL, Henderson CR Jr, Cory-Slechta DA, Cox C, Jusko TA, Lanphear BP (2003) Intellectual impairment in children with blood lead concentrations below 10 microg per deciliter. N Engl J Med 348:1517–1526
Carbone MA, Jordan KW, Lyman RF, Harbison ST, Leips J, Morgan TJ, DeLuca M, Awadalla P, Mackay TFC (2006) Phenotypic variation and natural selection at catsup, a pleiotropic quantitative trait gene in Drosophila. Curr Biol 16:912–919
Carbone MA, Yamamoto A, Huang W, Lyman RA, Meadors TB, Yamamoto R, Anholt RRH, Mackay TFC (2016) Genetic architecture of natural variation in visual senescence in Drosophila. Proc Natl Acad Sci USA 113:E6620-E6629
CARDIoGRAMplusC4D Consortium, Deloukas P, Kanoni S, Willenborg C, Farrall M, Assimes TL, Thompson JR, Ingelsson E, Saleheen D, Erdmann J, Goldstein BA, Stirrups K, König IR, Cazier JB, Johansson A, Hall AS, Lee JY, Willer CJ, Chambers JC, Esko T, Folkersen L, Goel A, Grundberg E, Havulinna AS, Ho WK, Hopewell JC, Eriksson N, Kleber ME, Kristiansson K, Lundmark P, Lyytikäinen LP, Rafelt S, Shungin D, Strawbridge RJ, Thorleifsson G, Tikkanen E, Van Zuydam N, Voight BF, Waite LL, Zhang W, Ziegler A, Absher D, Altshuler D, Balmforth AJ, Barroso I, Braund PS, Burgdorf C, Claudi-Boehm S, Cox D, Dimitriou M, Do R, DIAGRAM Consortium, CARDIOGENICS Consortium, Doney AS, El Mokhtari N, Eriksson P, Fischer K, Fontanillas P, Franco-Cereceda A, Gigante B, Groop L, Gustafsson S, Hager J, Hallmans G, Han BG, Hunt SE, Kang HM, Illig T, Kessler T, Knowles JW, Kolovou G, Kuusisto J, Langenberg C, Langford C, Leander K, Lokki ML, Lundmark A, McCarthy MI, Meisinger C, Melander O, Mihailov E, Maouche S, Morris AD, Müller-Nurasyid M, MuTHER Consortium, Nikus K, Peden JF, Rayner NW, Rasheed A, Rosinger S, Rubin D, Rumpf MP, Schäfer A, Sivananthan M, Song C, Stewart AF, Tan ST, Thorgeirsson G, van der Schoot CE, Wagner PJ, Wellcome Trust Case Control Consortium, Wells GA, Wild PS, Yang TP, Amouyel P, Arveiler D, Basart H, Boehnke M, Boerwinkle E, Brambilla P, Cambien F, Cupples AL, de Faire U, Dehghan A, Diemert P, Epstein SE, Evans A, Ferrario MM, Ferrières J, Gauguier D, Go AS, Goodall AH, Gudnason V, Hazen SL, Holm H, Iribarren C, Jang Y, Kähönen M, Kee F, Kim HS, Klopp N, Koenig W, Kratzer W, Kuulasmaa K, Laakso M, Laaksonen R, Lee JY, Lind L, Ouwehand WH, Parish S, Park JE, Pedersen NL, Peters A, Quertermous T, Rader DJ, Salomaa V, Schadt E, Shah SH, Sinisalo J, Stark K, Stefansson K, Trégouët DA, Virtamo J, Wallentin L, Wareham N, Zimmermann ME, Nieminen MS, Hengstenberg C, Sandhu MS, Pastinen T, Syvänen AC, Hovingh GK, Dedoussis G, Franks PW, Lehtimäki T, Metspalu A, Zalloua PA, Siegbahn A, Schreiber S, Ripatti S, Blankenberg SS, Perola M, Clarke R, Boehm BO, O’Donnell C, Reilly MP, März W, Collins R, Kathiresan S, Hamsten A, Kooner JS, Thorsteinsdottir U, Danesh J, Palmer CN, Roberts R, Watkins H, Schunkert H, Samani NJ (2013) Large-scale association analysis identifies new risk loci for coronary artery disease. Nat Genet 45:25–33
Caspi A, McClay J, Moffitt TE, Mill J, Martin J, Craig IW, Taylor A, Poulton R (2002) Role of genotype in the cycle of violence in maltreated children. Science 297:851–854
Caspi A, Sugden K, Moffitt TE, Taylor A, Craig IW, Harrington H, McClay J, Mill J, Martin J, Braithwaite A, Poulton R (2003) Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science 301:386–389
Cassar M, Issa AR, Riemensperger T, Petitgas C, Rival T, Coulom H, Iché-Torres M, Han KA, Birman S (2015) A dopamine receptor contributes to paraquat-induced neurotoxicity in Drosophila. Hum Mol Genet 24:197–212
Chahrour M, O’Roak BJ, Santini E, Samaco RC, Kleiman RJ, Manzini MC (2016) Current perspectives in autism spectrum disorder: from genes to therapy. J Neurosci 36:11402–11410
Chaston JM, Dobson AJ, Newell PD, Douglas AE (2016) Host genetic control of the microbiota mediates the Drosophila nutritional phenotype. Appl Environ Microbiol 82:671–679
Chow CY, Wolfner MF, Clark AG (2013a) Using natural variation in Drosophila to discover previously unknown endoplasmic reticulum stress genes. Proc Natl Acad Sci USA 110:9013–9018
Chow CY, Wolfner MF, Clark AG (2013b) Large neurological component to genetic differences underlying biased sperm use in Drosophila. Genetics 193:177–185
Chow CY, Kelsey KJ, Wolfner MF, Clark AG (2016) Candidate genetic modifiers of retinitis pigmentosa identified by exploiting natural variation in Drosophila. Hum Mol Genet 25:651–659
Corbett-Detig RB, Zhou J, Clark AG, Hartl DL, Ayroles JF (2013) Genetic incompatibilities are widespread within species. Nature 504:135–137
Dardiotis E, Xiromerisiou G, Hadjichristodoulou C, Tsatsakis AM, Wilks MF, Hadjigeorgiou GM (2013) The interplay between environmental and genetic factors in Parkinson’s disease susceptibility: the evidence for pesticides. Toxicology 307:17–23
Dembeck LM, Böröczky K, Huang W, Schal C, Anholt RRH, Mackay TFC (2015a) Genetic architecture of natural variation in cuticular hydrocarbon composition in Drosophila melanogaster. eLife 4:e09861
Dembeck LM, Huang W, Magwire MM, Lawrence F, Lyman RF, Mackay TFC (2015b) Genetic architecture of abdominal pigmentation in Drosophila melanogaster. PLoS Genet 11:e1005163
Dilda CL, Mackay TFC (2002) The genetic architecture of Drosophila sensory bristle number. Genetics 162:1655–1674
Dobson AJ, Chaston JM, Newell PD, Donahue L, Hermann SL, Sannino DR, Westmiller S, Wong AC, Clark AG, Lazzaro BP, Douglas AE (2015) Host genetic determinants of microbiota-dependent nutrition revealed by genome-wide analysis of Drosophila melanogaster. Nat Commun 6:6312
Durham MF, Magwire MM, Stone EA, Leips J (2014) Genome-wide analysis in Drosophila reveals age-specific effects of SNPs on fitness traits. Nat Commun 5:1–8
Ehrenreich IM, Torabi N, Jia Y, Kent J, Martis S, Shapiro JA, Gresham D, Caudy AA, Kruglyak L (2010) Dissection of genetically complex traits with extremely large pools of yeast segregants. Nature 464:1039–1042
Engel GL, Delwig A, Rand MD (2012) The effects of methylmercury on Notch signaling during embryonic neural development in Drosophila melanogaster. Toxicol In Vitro 26:485–492
Falconer DS, Mackay TFC (1996) Introduction to quantitative genetics, 4th edn. Longman, Burnt Mill
Gaertner BE, Ruedi EA, McCoy LJ, Moore JM, Wolfner MF, Mackay TFC (2015) Heritable variation in courtship patterns in Drosophila melanogaster. G3 5:531–539
Garlapow ME, Huang W, Yarboro MT, Peterson KR, Mackay TFC (2015) Quantitative genetics of food intake in Drosophila melanogaster. PLoS ONE 10:e0138129
Garlapow ME, Everett LJ, Zhou S, Gearhart AW, Fay KA, Huang W, Morozova TV, Arya GH, Turlapati L, St Armour G, Hussain YN, McAdams SE, Fochler S, Mackay TFC (2017) Genetic and genomic response to selection for food consumption in Drosophila melanogaster. Behav Genet 47:227–243
Gaulton KJ, Ferreira T, Lee Y, Raimondo A, Mägi R, Reschen ME, Mahajan A, Locke A, Rayner NW, Robertson N, Scott RA, Prokopenko I, Scott LJ, Green T, Sparso T, Thuillier D, Yengo L, Grallert H, Wahl S, Frånberg M, Strawbridge RJ, Kestler H, Chheda H, Eisele L, Gustafsson S, Steinthorsdottir V, Thorleifsson G, Qi L, Karssen LC, van Leeuwen EM, Willems SM, Li M, Chen H, Fuchsberger C, Kwan P, Ma C, Linderman M, Lu Y, Thomsen SK, Rundle JK, Beer NL, van de Bunt M, Chalisey A, Kang HM, Voight BF, Abecasis GR, Almgren P, Baldassarre D, Balkau B, Benediktsson R, Blüher M, Boeing H, Bonnycastle LL, Bottinger EP, Burtt NP, Carey J, Charpentier G, Chines PS, Cornelis MC, Couper DJ, Crenshaw AT, van Dam RM, Doney AS, Dorkhan M, Edkins S, Eriksson JG, Esko T, Eury E, Fadista J, Flannick J, Fontanillas P, Fox C, Franks PW, Gertow K, Gieger C, Gigante B, Gottesman O, Grant GB, Grarup N, Groves CJ, Hassinen M, Have CT, Herder C, Holmen OL, Hreidarsson AB, Humphries SE, Hunter DJ, Jackson AU, Jonsson A, Jørgensen ME, Jørgensen T, Kao WH, Kerrison ND, Kinnunen L, Klopp N, Kong A, Kovacs P, Kraft P, Kravic J, Langford C, Leander K, Liang L, Lichtner P, Lindgren CM, Lindholm E, Linneberg A, Liu CT, Lobbens S, Luan J, Lyssenko V, Männistö S, McLeod O, Meyer J, Mihailov E, Mirza G, Mühleisen TW, Müller-Nurasyid M, Navarro C, Nöthen MM, Oskolkov NN, Owen KR, Palli D, Pechlivanis S, Peltonen L, Perry JR, Platou CG, Roden M, Ruderfer D, Rybin D, van der Schouw YT, Sennblad B, Sigurðsson G, Stančáková A, Steinbach G, Storm P, Strauch K, Stringham HM, Sun Q, Thorand B, Tikkanen E, Tonjes A, Trakalo J, Tremoli E, Tuomi T, Wennauer R, Wiltshire S, Wood AR, Zeggini E, Dunham I, Birney E, Pasquali L, Ferrer J, Loos RJ, Dupuis J, Florez JC, Boerwinkle E, Pankow JS, van Duijn C, Sijbrands E, Meigs JB, Hu FB, Thorsteinsdottir U, Stefansson K, Lakka TA, Rauramaa R, Stumvoll M, Pedersen NL, Lind L, Keinanen-Kiukaanniemi SM, Korpi-Hyövälti E, Saaristo TE, Saltevo J, Kuusisto J, Laakso M, Metspalu A, Erbel R, Jöcke KH, Moebus S, Ripatti S, Salomaa V, Ingelsson E, Boehm BO, Bergman RN, Collins FS, Mohlke KL, Koistinen H, Tuomilehto J, Hveem K, Njølstad I, Deloukas P, Donnelly PJ, Frayling TM, Hattersley AT, de Faire U, Hamsten A, Illig T, Peters A, Cauchi S, Sladek R, Froguel P, Hansen T, Pedersen O, Morris AD, Palmer CN, Kathiresan S, Melander O, Nilsson PM, Groop LC, Barroso I, Langenberg C, Wareham NJ, Gloyn AL, Altshuler D, Boehnke M, Teslovich TM, McCarthy MI, Morris AP, DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium (2015) Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat Genet 47:1415–1425
Gill HJ, Nida DL, Dean DA, England MW, Jacobson KB (1989) Resistance of Drosophila to cadmium: biochemical factors in resistant and sensitive strains. Toxicology 56:315–321
Golden LC, Voskuhl R (2017) The importance of studying sex differences in disease: the example of multiple sclerosis. J Neurosci Res 95:633–643
Greene CS, White BC, Moore JH (2009) Sensible initialization using expert knowledge for genome-wide analysis of epistasis using genetic programming. Genet Evol Comput Conf 2009:1289–1296
Griffing B (1956) Concept of general and specific combining ability in relation to diallel crossing systems. Aust J Biol Sci 9:463–493
Grubbs N, Leach M, Su X, Petrisko T, Rosario JB, Mahaffey JW (2013) New components of Drosophila leg development identified through genome wide association studies. PLoS ONE 8:e60261
Harbison ST, McCoy LJ, Mackay TFC (2013) Genome-wide association study of sleep in Drosophila melanogaster. BMC Genomics 14:281
He X, Zhou S, St Armour GE, Mackay TFC, Anholt RRH (2016) Epistatic partners of neurogenic genes modulate Drosophila olfactory behavior. Genes Brain Behav 15:280–290
Hirsch HV, Mercer J, Sambaziotis H, Huber M, Stark DT, Torno-Morley T, Hollocher K, Ghiradella H, Ruden DM (2003) Behavioral effects of chronic exposure to low levels of lead in Drosophila melanogaster. Neurotoxicology 24:435–442
Hirsch HV, Lnenicka G, Possidente D, Possidente B, Garfinkel MD, Wang L, Lu X, Ruden DM (2012) Drosophila melanogaster as a model for lead neurotoxicology and toxicogenomics research. Front Genet 3:68
Horváth B, Betancourt AJ, Kalinka AT (2016) A novel method for quantifying the rate of embryogenesis uncovers considerable genetic variation for the duration of embryonic development in Drosophila melanogaster. BMC Evol Biol 16:200
Howick VM, Lazzaro BP (2017) The genetic architecture of defence as resistance to and tolerance of bacterial infection in Drosophila melanogaster. Mol Ecol 26:1533–1546
Huang W, Richards S, Carbone MA, Zhu D, Anholt RRH, Ayroles JF, Duncan L, Jordan KW, Lawrence F, Magwire MM, Warner CB, Blankenburg K, Han Y, Javaid M, Jayaseelan J, Jhangiani SN, Muzny D, Ongeri F, Perales L, Wu YQ, Zhang Y, Zou X, Stone EA, Gibbs RA, Mackay TFC (2012) Epistasis dominates the genetic architecture of Drosophila quantitative traits. Proc Natl Acad Sci USA 109:15553–15559
Huang W, Massouras A, Inoue Y, Peiffer J, Ràmia M, Tarone AM, Turlapati L, Zichner T, Zhu D, Lyman RF, Magwire MM, Blankenburg K, Carbone MA, Chang K, Ellis LL, Fernandez S, Han Y, Highnam G, Hjelmen CE, Jack JR, Javaid M, Jayaseelan J, Kalra D, Lee S, Lewis L, Munidasa M, Ongeri F, Patel S, Perales L, Perez A, Pu L, Rollmann SM, Ruth R, Saada N, Warner C, Williams A, Wu YQ, Yamamoto A, Zhang Y, Zhu Y, Anholt RRH, Korbel JO, Mittelman D, Muzny DM, Gibbs RA, Barbadilla A, Johnston JS, Stone EA, Richards S, Deplancke B, Mackay TFC (2014) Natural variation in genome architecture among 205 Drosophila melanogaster Genetic Reference Panel lines. Genome Res 24:1193–1208
Huang W, Carbone MA, Magwire MM, Peiffer JA, Lyman RF, Stone EA, Anholt RRH, Mackay TFC (2015) Genetic basis of transcriptome diversity in Drosophila melanogaster. Proc Natl Acad Sci USA 112:E6010-E6019
Hunter CM, Huang W, Mackay TFC, Singh ND (2016) The genetic architecture of natural variation in recombination rate in Drosophila melanogaster. PLoS Genet 12:e1005951
Isani G, Carpene E (2014) Metallothioneins, unconventional proteins from unconventional animals: a long journey from nematodes to mammals. Biomolecules 4:435–457
Ivanov DK, Escott-Price V, Ziehm M, Magwire MM, Mackay TFC, Partridge L, Thornton JM (2015) Longevity GWAS using the Drosophila Genetic Reference Panel. J Gerontol Ser A Biol Sci Med Sci 70:1470–1478
Jakubowski M (2011) Low-level environmental lead exposure and intellectual impairment in children–the current concepts of risk assessment. Int J Occup Med Environ Health 24:1–7
Jordan KW, Craver KL, Magwire MM, Cubilla CE, Mackay TFC, Anholt RRH (2012) Genome-wide association for sensitivity to chronic oxidative stress in Drosophila melanogaster. PLoS ONE 7:e38722
Jumbo-Lucioni P, Bu S, Harbison ST, Slaughter JC, Mackay TFC, Moellering DR, De Luca M (2012) Nuclear genomic control of naturally occurring variation in mitochondrial function in Drosophila melanogaster. BMC Genomics 13:659
Katzenberger RJ, Chtarbanova S, Rimkus SA, Fischer JA, Kaur G, Seppala JM, Swanson LC, Zajac JE, Ganetzky B, Wassarman DA (2015) Death following traumatic brain injury in Drosophila is associated with intestinal barrier dysfunction. eLife 4:e04790
Ke X, Durrant C, Morris AP, Hunt S, Bentley DR, Deloukas P, Cardon LR (2004) Efficiency and consistency of haplotype tagging of dense SNP maps in multiple samples. Hum Mol Genet 13:2557–2565
Keinan A, Clark AG (2012) Recent explosive human population growth has resulted in an excess of rare genetic variants. Science 336:740–743
Leips J, Mackay TFC (2000) Quantitative trait loci for life span in Drosophila melanogaster: interactions with genetic background and larval density. Genetics 155:1773–1788
Li L, Li Y, Zeng XC, Li J, Du XY (2016) Role of interleukin-4 genetic polymorphisms and environmental factors in the risk of asthma in children. Genet Mol Res 15:gmr15048873
Liu J, Lewis G (2014) Environmental toxicity and poor cognitive outcomes in children and adults. J Environ Health 76:130–138
Long AD, Mullaney SL, Reid LA, Fry JD, Langley CH, Mackay TFC (1995) High resolution mapping of genetic factors affecting abdominal bristle number in Drosophila melanogaster. Genetics 139:1273–1291
Mackay TFC (2014) Epistasis and quantitative traits: using model organisms to study gene-gene interactions. Nat Rev Genet 15:22–33
Mackay TFC (2015) Epistasis for quantitative traits in Drosophila. Methods Mol Biol 1253:47–70
Mackay TFC, Richards S, Stone EA, Barbadilla A, Ayroles JF, Zhu D, Casillas S, Han Y, Magwire MM, Cridland JM, Richardson MF, Anholt RRH, Barrón M, Bess C, Blankenburg KP, Carbone MA, Castellano D, Chaboub L, Duncan L, Harris Z, Javaid M, Jayaseelan JC, Jhangiani SN, Jordan KW, Lara F, Lawrence F, Lee SL, Librado P, Linheiro RS, Lyman RF, Mackey AJ, Munidasa M, Muzny DM, Nazareth L, Newsham I, Perales L, Pu LL, Qu C, Ràmia M, Reid JG, Rollmann SM, Rozas J, Saada N, Turlapati L, Worley KC, Wu YQ, Yamamoto A, Zhu Y, Bergman CM, Thornton KR, Mittelman D, Gibbs RA (2012) The Drosophila melanogaster Genetic Reference Panel. Nature 482:173–178
Magwire MM, Yamamoto A, Carbone MA, Roshina NV, Symonenko AV, Pasyukova EG, Morozova TV, Mackay TFC (2010) Quantitative and molecular genetic analyses of mutations increasing Drosophila life span. PLoS Genet 6:e1001037
Magwire MM, Fabian DK, Schweyen H, Cao C, Longdon B, Bayer F, Jiggins FM (2012) Genome-wide association studies reveal a simple genetic basis of resistance to naturally coevolving viruses in Drosophila melanogaster. PLoS Genet 8:e1003057
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A, Cho JH, Guttmacher AE, Kong A, Kruglyak L, Mardis E, Rotimi CN, Slatkin M, Valle D, Whittemore AS, Boehnke M, Clark AG, Eichler EE, Gibson G, Haines JL, Mackay TFC, McCarroll SA, Visscher PM (2009) Finding the missing heritability of complex diseases. Nature 461:747–753
Martin CA, Barajas A, Lawless G, Lawal HO, Assani K, Lumintang YP, Nunez V, Krantz DE (2014) Synergistic effects on dopamine cell death in a Drosophila model of chronic toxin exposure. Neurotoxicology 44:344–351
McDermott S, Bao W, Tong X, Cai B, Lawson A, Aelion CM (2014) Are different soil metals near the homes of pregnant women associated with mild and severe intellectual disability in children? Dev Med Child Neurol 56:888–897
Mensch J, Lavagnino N, Carreira VP, Massaldi A, Hasson E, Fanara JJ (2008) Identifying candidate genes affecting developmental time in Drosophila melanogaster: pervasive pleiotropy and gene-by-environment interaction. BMC Dev Biol 8:78
Mitchell CL, Latuszek CE, Vogel KR, Greenlund IM, Hobmeier RE, Ingram OK, Dufek SR, Pecore JL, Nip FR, Johnson ZJ, Ji X, Wei H, Gailing O, Werner T (2017) α-Amanitin resistance in Drosophila melanogaster: a genome-wide association approach. PLoS ONE 12:e0173162
Mohlke KL, Boehnke M (2015) Recent advances in understanding the genetic architecture of type 2 diabetes. Hum Mol Genet 24:R85-R92
Montgomery SL, Vorojeikina D, Huang W, Mackay TFC, Anholt RRH, Rand MD (2014) Genome-wide association analysis of tolerance to methylmercury toxicity in Drosophila implicates myogenic and neuromuscular developmental pathways. PLoS ONE 9:e110375
Montooth KL, Marden JH, Clark AG (2003) Mapping determinants of variation in energy metabolism, respiration and flight in Drosophila. Genetics 165:623–635
Morgante F, Sørensen P, Sorensen DA, Maltecca C, Mackay TFC (2015) Genetic architecture of micro-environmental plasticity in Drosophila melanogaster. Sci Rep 5:9785
Morozova TV, Ayroles JF, Jordan KW, Duncan LH, Carbone MA, Lyman RF, Stone EA, Govindaraju DR, Ellison CR, Mackay TFC, Anholt RRH (2009) Alcohol sensitivity in Drosophila: translational potential of systems genetics. Genetics 83:733–745
Morozova TV, Mackay TFC, Anholt RRH (2014) Genetics and genomics of alcohol sensitivity. Mol Genet Genomics 289:253–269
Morozova TV, Huang W, Pray VA, Whitham T, Anholt RRH, Mackay TFC (2015) Polymorphisms in early neurodevelopmental genes affect natural variation in alcohol sensitivity in adult Drosophila. BMC Genomics 16:865
Moskvina V, Craddock N, Müller-Myhsok B, Kam-Thong T, Green E, Holmans P, Owen MJ, O’Donovan MC (2011) An examination of single nucleotide polymorphism selection prioritization strategies for tests of gene-gene interaction. Biol Psychiatry 70:198–203
Munafò MR, Durrant C, Lewis G, Flint J (2009) Gene X environment interactions at the serotonin transporter locus. Biol Psychiatry 65:211–219
Najarro MA, Hackett JL, Macdonald SJ (2017) Loci contributing to boric acid toxicity in two reference populations of Drosophila melanogaster. G3 7:1631–1641
Nelson CS, Beck JN, Wilson KA, Pilcher ER, Kapahi P, Brem RB (2016) Cross-phenotype association tests uncover genes mediating nutrient response in Drosophila. BMC Genomics 17:867
Nguyen AH, Altomare LE, McElwain MC (2014) Decreased accumulation of cadmium in Drosophila selected for resistance suggests a mechanism independent of metallothionein. Biol Trace Elem Res 160:245–249
Park SY, Ludwig MZ, Tamarina NA, He BZ, Carl SH, Dickerson DA, Barse L, Arun B, Williams CL, Miles CM, Philipson LH, Steiner DF, Bell GI, Kreitman M (2014) Genetic complexity in a Drosophila model of diabetes-associated misfolded human proinsulin. Genetics 196:539–555
Pritchard JK, Rosenberg NA (1999) Use of unlinked genetic markers to detect population stratification in association studies. Am J Hum Genet 65:220–228
Reich DE, Lander ES (2001) On the allelic spectrum of human disease. Trends Genet 17:502–510
Reich DE, Cargill M, Bolk S, Ireland J, Sabeti PC, Richter DJ, Lavery T, Kouyoumjian R, Farhadian SF, Ward R, Lander ES (2001) Linkage disequilibrium in the human genome. Nature 411:199–204
Reiter LT, Potocki L, Chien S, Gribskov M, Bier E (2001) A systematic analysis of human disease-associated gene sequences in Drosophila melanogaster. Genome Res 11:1114–1125
Rocha TB, Hutz MH, Salatino-Oliveira A, Genro JP, Polanczyk GV, Sato JR, Wehrmeister FC, Barros FC, Menezes AM, Rohde LA, Anselmi L, Kieling C (2015) Gene-environment interaction in youth depression: replication of the 5-HTTLPR moderation in a diverse setting. Am J Psychiatry 172:978–985
Rohde PD, Gaertner B, Ward K, Sørensen P, Mackay TFC (2017) Genomic analysis of genotype-by-social environment interaction for Drosophila melanogaster aggressive behavior. Genetics 206:1969–1984
Ruden DM, Chen L, Possidente D, Possidente B, Rasouli P, Wang L, Lu X, Garfinkel MD, Hirsch HV, Page GP (2009) Genetical toxicogenomics in Drosophila identifies master-modulatory loci that are regulated by developmental exposure to lead. Neurotoxicology 30:898–914
Sambandan D, Yamamoto A, Fanara JJ, Mackay TFC, Anholt RRH (2006) Dynamic genetic interactions determine odor-guided behavior in Drosophila melanogaster. Genetics 174:1349–1363
Sanjuán R, Elena SF (2006) Epistasis correlates to genomic complexity. Proc Natl Acad Sci USA 103:14402–14405
Shorter J, Couch C, Huang W, Carbone MA, Peiffer J, Anholt RRH, Mackay TFC (2015) Genetic architecture of natural variation in Drosophila melanogaster aggressive behavior. Proc Natl Acad Sci USA 112:E3555-E3563
Sordillo JE, Kelly R, Bunyavanich S, McGeachie M, Qiu W, Croteau-Chonka DC, Soto-Quiros M, Avila L, Celedón JC, Brehm JM, Weiss ST, Gold DR, Litonjua AA (2015) Genome-wide expression profiles identify potential targets for gene-environment interactions in asthma severity. J Allergy Clin Immunol 136:885–892
Srivastav SP, Kelleher ES (2017) Paternal induction of hybrid dysgenesis in Drosophila melanogaster is weakly correlated with both P-element and hobo element dosage. G3 7:1487–1497
Stefansson H, Rujescu D, Cichon S, Pietiläinen OP, Ingason A, Steinberg S, Fossdal R, Sigurdsson E, Sigmundsson T, Buizer-Voskamp JE, Hansen T, Jakobsen KD, Muglia P, Francks C, Matthews PM, Gylfason A, Halldorsson BV, Gudbjartsson D, Thorgeirsson TE, Sigurdsson A, Jonasdottir A, Jonasdottir A, Bjornsson A, Mattiasdottir S, Blondal T, Haraldsson M, Magnusdottir BB, Giegling I, Möller HJ, Hartmann A, Shianna KV, Ge D, Need AC, Crombie C, Fraser G, Walker N, Lonnqvist J, Suvisaari J, Tuulio-Henriksson A, Paunio T, Toulopoulou T, Bramon E, Di Forti M, Murray R, Ruggeri M, Vassos E, Tosato S, Walshe M, Li T, Vasilescu C, Mühleisen TW, Wang AG, Ullum H, Djurovic S, Melle I, Olesen J, Kiemeney LA, Franke B, GROUP, Sabatti C, Freimer NB, Gulcher JR, Thorsteinsdottir U, Kong A, Andreassen OA, Ophoff RA, Georgi A, Rietschel M, Werge T, Petursson H, Goldstein DB, Nöthen MM, Peltonen L, Collier DA, St Clair D, Stefansson K (2008) Large recurrent microdeletions associated with schizophrenia. Nature 455:232–236
Stefansson H, Ophoff RA, Steinberg S, Andreassen OA, Cichon S, Rujescu D, Werge T, Pietiläinen OP, Mors O, Mortensen PB, Sigurdsson E, Gustafsson O, Nyegaard M, Tuulio-Henriksson A, Ingason A, Hansen T, Suvisaari J, Lonnqvist J, Paunio T, Børglum AD, Hartmann A, Fink-Jensen A, Nordentoft M, Hougaard D, Norgaard-Pedersen B, Böttcher Y, Olesen J, Breuer R, Möller HJ, Giegling I, Rasmussen HB, Timm S, Mattheisen M, Bitter I, Réthelyi JM, Magnusdottir BB, Sigmundsson T, Olason P, Masson G, Gulcher JR, Haraldsson M, Fossdal R, Thorgeirsson TE, Thorsteinsdottir U, Ruggeri M, Tosato S, Franke B, Strengman E, Kiemeney LA, Genetic Risk Outcome in Psychosis (GROUP), Melle I, Djurovic S, Abramova L, Kaleda V, Sanjuan J, de Frutos R, Bramon E, Vassos E, Fraser G, Ettinger U, Picchioni M, Walker N, Toulopoulou T, Need AC, Ge D, Yoon JL, Shianna KV, Freimer NB, Cantor RM, Murray R, Kong A, Golimbet V, Carracedo A, Arango C, Costas J, Jönsson EG, Terenius L, Agartz I, Petursson H, Nöthen MM, Rietschel M, Matthews PM, Muglia P, Peltonen L, St Clair D, Goldstein DB, Stefansson K, Collier DA (2009) Common variants conferring risk of schizophrenia. Nature 460:744–747
Swarup S, Harbison ST, Hahn LE, Morozova TV, Yamamoto A, Mackay TFC, Anholt RRH (2012) Extensive epistasis for olfactory behavior, sleep and waking activity in Drosophila melanogaster. Genet Res 94:9–20
Swarup S, Huang W, Mackay TFC, Anholt RRH (2013) Analysis of natural variation reveals neurogenetic networks for Drosophila olfactory behavior. Proc Natl Acad Sci USA 110:1017–1022
Takahara B, Takahashi KH (2015) Genome-wide association study on male genital shape and size in Drosophila melanogaster. PLoS ONE 10:e0132846
Thibault ST, Singer MA, Miyazaki WY, Milash B, Dompe NA, Singh CM, Buchholz R, Demsky M, Fawcett R, Francis-Lang HL, Ryner L, Cheung LM, Chong A, Erickson C, Fisher WW, Greer K, Hartouni SR, Howie E, Jakkula L, Joo D, Killpack K, Laufer A, Mazzotta J, Smith RD, Stevens LM, Stuber C, Tan LR, Ventura R, Woo A, Zakrajsek I, Zhao L, Chen F, Swimmer C, Kopczynski C, Duyk G, Winberg ML, Margolis J (2004) A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nat Genet 36:283–287
Turner TL, Miller PM, Cochrane VA (2013) Combining genome-wide methods to investigate the genetic complexity of courtship song variation in Drosophila melanogaster. Mol Biol Evol 30:2113–2120
Unckless RL, Rottschaefer SM, Lazzaro BP (2015a) A genome-wide association study for nutritional indices in Drosophila. G3 5:417–425
Unckless RL, Rottschaefer SM, Lazzaro BP (2015b) The complex contributions of genetics and nutrition to immunity in Drosophila melanogaster. PLoS Genet 11:e1005030
Vaisnav M, Xing C, Ku HC, Hwang D, Stojadinovic S, Pertsemlidis A, Abrams JM (2014) Genome-wide association analysis of radiation resistance in Drosophila melanogaster. PLoS ONE 9:e104858
Vonesch SC, Lamparter D, Mackay TFC, Bergmann S, Hafen E (2016) Genome-wide analysis reveals novel regulators of growth in Drosophila melanogaster. PLoS Genet 12:e1005616
Wang P, Lyman RF, Mackay TFC, Anholt RRH (2010) Natural variation in odorant recognition among odorant-binding proteins in Drosophila melanogaster. Genetics 184:759–767
Wang JB, Lu H-L, St. Leger RJ (2017) The genetic basis for variation in resistance to infection in the Drosophila melanogaster Genetic Reference Panel. PLoS Pathog 13:e1006260
Webber C (2017) Epistasis in neuropsychiatric disorders. Trends Genet 33:256–265
Weber AL, Khan GF, Magwire MM, Tabor CL, Mackay TFC, Anholt RRH (2012) Genome-wide association for oxidative stress resistance in Drosophila melanogaster. PLoS ONE 7:e34745
Wood AR, Esko T, Yang J, Vedantam S, Pers TH, Gustafsson S, Chu AY, Estrada K, Luan J, Kutalik Z, Amin N, Buchkovich ML, Croteau-Chonka DC, Day FR, Duan Y, Fall T, Fehrmann R, Ferreira T, Jackson AU, Karjalainen J, Lo KS, Locke AE, Mägi R, Mihailov E, Porcu E, Randall JC, Scherag A, Vinkhuyzen AA, Westra HJ, Winkler TW, Workalemahu T, Zhao JH, Absher D, Albrecht E, Anderson D, Baron J, Beekman M, Demirkan A, Ehret GB, Feenstra B, Feitosa MF, Fischer K, Fraser RM, Goel A, Gong J, Justice AE, Kanoni S, Kleber ME, Kristiansson K, Lim U, Lotay V, Lui JC, Mangino M, Mateo Leach I, Medina-Gomez C, Nalls MA, Nyholt DR, Palmer CD, Pasko D, Pechlivanis S, Prokopenko I, Ried JS, Ripke S, Shungin D, Stancáková A, Strawbridge RJ, Sung YJ, Tanaka T, Teumer A, Trompet S, van der Laan SW, van Setten J, Van Vliet-Ostaptchouk JV, Wang Z, Yengo L, Zhang W, Afzal U, Arnlöv J, Arscott GM, Bandinelli S, Barrett A, Bellis C, Bennett AJ, Berne C, Blüher M, Bolton JL, Böttcher Y, Boyd HA, Bruinenberg M, Buckley BM, Buyske S, Caspersen IH, Chines PS, Clarke R, Claudi-Boehm S, Cooper M, Daw EW, De Jong PA, Deelen J, Delgado G, Denny JC, Dhonukshe-Rutten R, Dimitriou M, Doney AS, Dörr M, Eklund N, Eury E, Folkersen L, Garcia ME, Geller F, Giedraitis V, Go AS, Grallert H, Grammer TB, Gräßler J, Grönberg H, de Groot LC, Groves CJ, Haessler J, Hall P, Haller T, Hallmans G, Hannemann A, Hartman CA, Hassinen M, Hayward C, Heard-Costa NL, Helmer Q, Hemani G, Henders AK, Hillege HL, Hlatky MA, Hoffmann W, Hoffmann P, Holmen O, Houwing-Duistermaat JJ, Illig T, Isaacs A, James AL, Jeff J, Johansen B, Johansson Å, Jolley J, Juliusdottir T, Junttila J, Kho AN, Kinnunen L, Klopp N, Kocher T, Kratzer W, Lichtner P, Lind L, Lindström J, Lobbens S, Lorentzon M, Lu Y, Lyssenko V, Magnusson PK, Mahajan A, Maillard M, McArdle WL, McKenzie CA, McLachlan S, McLaren PJ, Menni C, Merger S, Milani L, Moayyeri A, Monda KL, Morken MA, Müller G, Müller-Nurasyid M, Musk AW, Narisu N, Nauck M, Nolte IM, Nöthen MM, Oozageer L, Pilz S, Rayner NW, Renstrom F, Robertson NR, Rose LM, Roussel R, Sanna S, Scharnagl H, Scholtens S, Schumacher FR, Schunkert H, Scott RA, Sehmi J, Seufferlein T, Shi J, Silventoinen K, Smit JH, Smith AV, Smolonska J, Stanton AV, Stirrups K, Stott DJ, Stringham HM, Sundström J, Swertz MA, Syvänen AC, Tayo BO, Thorleifsson G, Tyrer JP, van Dijk S, van Schoor NM, van der Velde N, van Heemst D, van Oort FV, Vermeulen SH, Verweij N, Vonk JM, Waite LL, Waldenberger M, Wennauer R, Wilkens LR, Willenborg C, Wilsgaard T, Wojczynski MK, Wong A, Wright AF, Zhang Q, Arveiler D, Bakker SJ, Beilby J, Bergman RN, Bergmann S, Biffar R, Blangero J, Boomsma DI, Bornstein SR, Bovet P, Brambilla P, Brown MJ, Campbell H, Caulfield MJ, Chakravarti A, Collins R, Collins FS, Crawford DC, Cupples LA, Danesh J, de Faire U, den Ruijter HM, Erbel R, Erdmann J, Eriksson JG, Farrall M, Ferrannini E, Ferrières J, Ford I, Forouhi NG, Forrester T, Gansevoort RT, Gejman PV, Gieger C, Golay A, Gottesman O, Gudnason V, Gyllensten U, Haas DW, Hall AS, Harris TB, Hattersley AT, Heath AC, Hengstenberg C, Hicks AA, Hindorff LA, Hingorani AD, Hofman A, Hovingh GK, Humphries SE, Hunt SC, Hypponen E, Jacobs KB, Jarvelin MR, Jousilahti P, Jula AM, Kaprio J, Kastelein JJ, Kayser M, Kee F, Keinanen-Kiukaanniemi SM, Kiemeney LA, Kooner JS, Kooperberg C, Koskinen S, Kovacs P, Kraja AT, Kumari M, Kuusisto J, Lakka TA, Langenberg C, Le Marchand L, Lehtimäki T, Lupoli S, Madden PA, Männistö S, Manunta P, Marette A, Matise TC, McKnight B, Meitinger T, Moll FL, Montgomery GW, Morris AD, Morris AP, Murray JC, Nelis M, Ohlsson C, Oldehinkel AJ, Ong KK, Ouwehand WH, Pasterkamp G, Peters A, Pramstaller PP, Price JF, Qi L, Raitakari OT, Rankinen T, Rao DC, Rice TK, Ritchie M, Rudan I, Salomaa V, Samani NJ, Saramies J, Sarzynski MA, Schwarz PE, Sebert S, Sever P, Shuldiner AR, Sinisalo J, Steinthorsdottir V, Stolk RP, Tardif JC, Tönjes A, Tremblay A, Tremoli E, Virtamo J, Vohl MC, Electronic Medical Records and Genomics (eMEMERGEGE) Consortium, MIGen Consortium, PAGEGE Consortium, LifeLines Cohort Study, Amouyel P, Asselbergs FW, Assimes TL, Bochud M, Boehm BO, Boerwinkle E, Bottinger EP, Bouchard C, Cauchi S, Chambers JC, Chanock SJ, Cooper RS, de Bakker PI, Dedoussis G, Ferrucci L, Franks PW, Froguel P, Groop LC, Haiman CA, Hamsten A, Hayes MG, Hui J, Hunter DJ, Hveem K, Jukema JW, Kaplan RC, Kivimaki M, Kuh D, Laakso M, Liu Y, Martin NG, März W, Melbye M, Moebus S, Munroe PB, Njølstad I, Oostra BA, Palmer CN, Pedersen NL, Perola M, Pérusse L, Peters U, Powell JE, Power C, Quertermous T, Rauramaa R, Reinmaa E, Ridker PM, Rivadeneira F, Rotter JI, Saaristo TE, Saleheen D, Schlessinger D, Slagboom PE, Snieder H, Spector TD, Strauch K, Stumvoll M, Tuomilehto J, Uusitupa M, van der Harst P, Völzke H, Walker M, Wareham NJ, Watkins H, Wichmann HE, Wilson JF, Zanen P, Deloukas P, Heid IM, Lindgren CM, Mohlke KL, Speliotes EK, Thorsteinsdottir U, Barroso I, Fox CS, North KE, Strachan DP, Beckmann JS, Berndt SI, Boehnke M, Borecki IB, McCarthy MI, Metspalu A, Stefansson K, Uitterlinden AG, van Duijn CM, Franke L, Willer CJ, Price AL, Lettre G, Loos RJ, Weedon MN, Ingelsson E, O’Connell JR, Abecasis GR, Chasman DI, Goddard ME, Visscher PM, Hirschhorn JN, Frayling TM (2014) Defining the role of common variation in the genomic and biological architecture of adult human height. Nat Genet 46:1173–1186
Yamamoto A, Anholt RRH, Mackay TFC (2009) Epistatic interactions attenuate mutations affecting startle behaviour in Drosophila melanogaster. Genet Res (Camb) 91:373–382
Zeggini E, Scott LJ, Saxena R, Voight BF, Marchini JL, Hu T, de Bakker PI, Abecasis GR, Almgren P, Andersen G, Ardlie K, Boström KB, Bergman RN, Bonnycastle LL, Borch-Johnsen K, Burtt NP, Chen H, Chines PS, Daly MJ, Deodhar P, Ding CJ, Doney AS, Duren WL, Elliott KS, Erdos MR, Frayling TM, Freathy RM, Gianniny L, Grallert H, Grarup N, Groves CJ, Guiducci C, Hansen T, Herder C, Hitman GA, Hughes TE, Isomaa B, Jackson AU, Jørgensen T, Kong A, Kubalanza K, Kuruvilla FG, Kuusisto J, Langenberg C, Lango H, Lauritzen T, Li Y, Lindgren CM, Lyssenko V, Marvelle AF, Meisinger C, Midthjell K, Mohlke KL, Morken MA, Morris AD, Narisu N, Nilsson P, Owen KR, Palmer CN, Payne F, Perry JR, Pettersen E, Platou C, Prokopenko I, Qi L, Qin L, Rayner NW, Rees M, Roix JJ, Sandbaek A, Shields B, Sjögren M, Steinthorsdottir V, Stringham HM, Swift AJ, Thorleifsson G, Thorsteinsdottir U, Timpson NJ, Tuomi T, Tuomilehto J, Walker M, Watanabe RM, Weedon MN, Willer CJ, Wellcome Trust Case Control Consortium, Illig T, Hveem K, Hu FB, Laakso M, Stefansson K, Pedersen O, Wareham NJ, Barroso I, Hattersley AT, Collins FS, Groop L, McCarthy MI, Boehnke M, Altshuler D (2008) Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat Genet 40:638–645
Zhai RG, Hiesinger PR, Koh TW, Verstreken P, Schulze KL, Cao Y, Jafar-Nejad H, Norga KK, Pan H, Bayat V, Greenbaum MP, Bellen HJ (2003) Mapping Drosophila mutations with molecularly defined P element insertions. Proc Natl Acad Sci USA 100:10860–10865
Zhou S, Campbell TG, Stone EA, Mackay TFC, Anholt RRH (2012) Phenotypic plasticity of the Drosophila transcriptome. PLoS Genet 8:e1002593
Zhou S, Morozova TV, Hussain YN, Luoma SE, McCoy L, Yamamoto A, Mackay TFC, Anholt RRH (2016) The genetic basis for variation in sensitivity to lead toxicity in Drosophila melanogaster. Environ Health Perspect 124:1062–1070
Zhou S, Luoma SE, St Armour GE, Thakkar E, Mackay TFC, Anholt RRH (2017) A Drosophila model for toxicogenomics: genetic variation in susceptibility to heavy metal exposure. PLoS Genet 13:e1006907
Zuk O, Hechter E, Sunyaev SR, Lander ES (2012) The mystery of missing heritability: genetic interactions create phantom heritability. Proc Natl Acad Sci USA 109:1193–1198
Zwarts L, Magwire MM, Carbone MA, Versteven M, Herteleer L, Anholt RRH, Callaerts P, Mackay TFC (2011) Complex genetic architecture of Drosophila aggressive behavior. Proc Natl Acad Sci USA 108:17070–17075
Zwarts L, Vanden Broeck L, Cappuyns E, Ayroles JF, Magwire MM, Vulsteke V, Clements J, Mackay TFC, Callaerts P (2015) The genetic basis of natural variation in mushroom body size in Drosophila melanogaster. Nat Commun 6:10115
Acknowledgements
Work in the authors’ laboratories is supported by NIH Grants GM059469, AA016560, GM076083, and AG043490.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Anholt, R.R.H., Mackay, T.F.C. The road less traveled: from genotype to phenotype in flies and humans. Mamm Genome 29, 5–23 (2018). https://doi.org/10.1007/s00335-017-9722-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-017-9722-7