Abstract
Objectives
Evaluate the performance of a deep learning (DL)–based model for multiple sclerosis (MS) lesion segmentation and compare it to other DL and non-DL algorithms.
Methods
This ambispective, multicenter study assessed the performance of a DL-based model for MS lesion segmentation and compared it to alternative DL- and non-DL-based methods. Models were tested on internal (n = 20) and external (n = 18) datasets from Latin America, and on an external dataset from Europe (n = 49). We also examined robustness by rescanning six patients (n = 6) from our MS clinical cohort. Moreover, we studied inter-human annotator agreement and discussed our findings in light of these results. Performance and robustness were assessed using intraclass correlation coefficient (ICC), Dice coefficient (DC), and coefficient of variation (CV).
Results
Inter-human ICC ranged from 0.89 to 0.95, while spatial agreement among annotators showed a median DC of 0.63. Using expert manual segmentations as ground truth, our DL model achieved a median DC of 0.73 on the internal, 0.66 on the external, and 0.70 on the challenge datasets. The performance of our DL model exceeded that of the alternative algorithms on all datasets. In the robustness experiment, our DL model also achieved higher DC (ranging from 0.82 to 0.90) and lower CV (ranging from 0.7 to 7.9%) when compared to the alternative methods.
Conclusion
Our DL-based model outperformed alternative methods for brain MS lesion segmentation. The model also proved to generalize well on unseen data and has a robust performance and low processing times both on real-world and challenge-based data.
Clinical relevance statement
Our DL-based model demonstrated superior performance in accurately segmenting brain MS lesions compared to alternative methods, indicating its potential for clinical application with improved accuracy, robustness, and efficiency.
Key Points
• Automated lesion load quantification in MS patients is valuable; however, more accurate methods are still necessary.
• A novel deep learning model outperformed alternative MS lesion segmentation methods on multisite datasets.
• Deep learning models are particularly suitable for MS lesion segmentation in clinical scenarios.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Multiple sclerosis (MS) is a chronic immunomediated inflammatory disease of the central nervous system that affects both gray and white matter (WM) [1]. Although the exact cause is unknown, a putative combination of genetic and environmental factors leads to an autoimmune reaction against myelinated axons, resulting in demyelination, gliosis, and neuronal loss. Brain lesions characteristically affect periventricular, juxtacortical, and infratentorial WM, cortical gray matter as well as deep gray structures (i.e., thalamus). Consequently, as a result, persons with MS present variable degrees of cortical and gray nuclei atrophy that exceed the values observed in healthy individuals of the same age [2].
Since its introduction in clinical practice, magnetic resonance imaging (MRI) has proven to be a key study in the evaluation of patients with MS [3]. MRI criteria were first integrated into the diagnostic guidelines for MS in 2001 and evolved in the latest criteria as the leading complementary study to reach diagnosis [4, 5]. MAGNIMS Study Group also emphasizes the importance of MRI to detect the dissemination in space and time of WM lesions and help to rule out alternative diagnoses [6, 7].
Radiological follow-up is mandatory for patients with MS, as asymptomatic radiological lesions are present in an 8–10/1 ratio and are paramount for establishing disease progression and/or treatment failure [8]. Serial brain MRI studies should accurately assess lesion burden, indicating whether disease is stable or progressing. However, lesion load measurement by radiologists is time-consuming and prone to intra- and inter-observer variability and is seldomly obtained in clinical practice. Most radiological reports mention, at best, the number and location of demyelinating plaques, and whether these lesions are new. But a global estimate of the lesion burden is rarely reported.
To overcome this limitation, computer algorithms have been developed for automated WM lesion segmentation and quantification; however, they are not routinely used in radiological clinical practice [9,10,11]. This underutilization may be related to several factors including lack of knowledge of these techniques and how to implement them, lack of resources, or a lack of acceptance.
In the last few years, deep learning (DL)–based methods for automatically segmenting MS lesions have emerged [12]. They tend to be faster and more precise than non-DL-based algorithms and could solve some of the technical problems and challenges found by the latter. We aim to determine if a novel DL-based automated MS lesion load quantification tool (Entelai Neuro) outperforms other DL-based models and non-DL-based methods.
Materials and methods
We performed three sequential experiments for this ambispective, multicenter study with MRI coming both from real-life and challenge-based datasets. In experiment 1, we evaluated correlation and spatial agreement between manual operators, which can be seen as an upper bound on expected performance in this task. In experiment 2, we trained, validated, and tested (both with local, external, and challenge-based databases) a DL-based model for automated MS lesion segmentation, comparing its performance to a state-of-the-art and widely adopted DL- and non-DL-based software, nicMSlesions and Lesion Segmentation Tool (LST) respectively [13, 14]. In experiment 3, we assessed the robustness of our DL model by performing repeated MRI scans in 6 patients from our MS clinical cohort, again comparing its performance to nicMSlesions and LST.
This study was approved by our institutional review board (IRB). A waiver was obtained from the IRB for the retrospective arm of this research. Patients involved in the prospective arm (experiment 3) provided written informed consent.
Subjects
Brain MRI data of 20 subjects with MS were selected from our center for experiment 1. For experiment 2, we retrospectively included 295 subjects from our center for model development. Approximately 93% of that dataset (275 subjects) was used for training and validation. The remainder 7% (20 subjects) was retained for internal model testing. For external validation, we included a total of 67 subjects with MS; 18 subjects from an external clinical center in Latin America and 49 subjects corresponding to a European multicentric cohort from the MSSEG challenge dataset [15].
Convenience sampling was used for internal and external dataset building, with an approximately equal distribution of subjects with low, medium, and high MS lesion load, as evaluated by referent neuroradiologists from each center. Low, intermediate, and high lesion load were defined as < 5, 5–15, and > 15 mL respectively. All subjects (n = 53) included in both training and testing datasets from the MSSEG challenge were analyzed with the segmentation algorithms to further evaluate the generalization of the models on unseen data. Four subjects from this dataset were excluded as either their lesion load was equal to 0 mL in the ground truth consensus mask (n = 1) or there were errors on the processing pipeline of LST (n = 3).
For experiment 3, we prospectively included 6 subjects who were scanned 4 times each, during two different visits on two MRI scanners located in our clinical center. Group A included 3 subjects who were scanned twice on the same scanner on the first visit and twice on the other scanner on the second visit. Group B included 3 subjects who were scanned twice on different scanners on each visit. Same-visit scans were separated by 30–60 min, and different-visit scans were separated by 1–3 weeks. On same-visit scans, subjects were allowed to drink water and/or use the restrooms, but they were asked not to leave the MRI facilities.
When available, demographic and clinical data (MS subtype, expanded disability status scale, disease duration, and treatment) were collected from the electronic health records. As several subjects were scanned as outpatients or included from public datasets as previously stated, missing clinical data was tabulated as NA (not available).
MRI
MR images for experiment 1 were acquired on a GE Signa HDxt 3 Tesla (T). In experiment 2, we used MR images from a GE Discovery 750 3 T scanner for training, validation, and internal testing, while real-life external testing was done with MRI acquired on a Siemens Magnetom Prisma 3 T scanner. The challenge-based external testing was done with data proceeding from 4 different MRI scanners: Siemens Verio 3 T, GE Discovery 750 3 T, Siemens Aera 1.5 T, and Philips Ingenia 3 T. Experiment 3 included images acquired on a Philips Achieva 1.5 T and a GE Discovery 750 3 T scanner. The distribution of subjects and MRI scanners used for model testing stratified by lesion load (< 5, 5–15, and > 15 mL) is detailed in Supplementary Table 1.
All protocols included 3D FLAIR and 3D T1-weighted images (sequences parameters are detailed in Supplementary Tables 2 and 3).
Clinical images were downloaded from the Picture Archiving and Communication System (PACS) and converted to Neuroimaging Informatics Technology Initiative (NIfTI) format for post-processing and anonymization. Challenge data was downloaded from the Shanoir-NG (sharing neuroimaging resources next generation) platform (https://shanoir.irisa.fr/) as NIfTI files.
Manual segmentations
Manually performed segmentations were used as ground truth in experiments 1 and 2. Three neuroradiologists (M.M.S., N.I.S., and E.O.S.) with 6, 7, and 6 years in the field, respectively, manually segmented MS lesions from the 3D FLAIR sequences of—all but challenge-derived—MS subjects using ITK-SNAP (http://www.itksnap.org) [16]. Brain T2 hyperintense lesions with radiological characteristics of demyelinating plaques were manually delineated on each slice and binary lesion masks were generated afterwards. Total segmentation time was recorded in minutes.
MSSEG challenge data included seven manual segmentation masks (performed by different trained experts split over the three acquisition sites) and a consensus ground truth segmentation, built with the LOP STAPLE algorithm [17]. We analyzed whether the performance of these external annotators was similar to ours by comparing spatial agreement between them.
Non-DL-based automated MS lesion quantification
Non-DL-based automated MS lesion load quantification was performed using lesion growth algorithm (LGA) and lesion prediction algorithm (LPA) as implemented in LST version 3.0.0 (http://www.statistical-modelling.de/lst.html) [14, 18]. A more detailed description of the software is provided in the supplementary material.
DL-based automated MS lesion quantification
NicMSlesions
NicMSlesions toolbox [13, 19] is based on a cascade of two 3D patchwise convolutional neural networks (https://github.com/sergivalverde/nicMSlesions). A more detailed description of the software is provided in the supplementary material.
Entelai Neuro
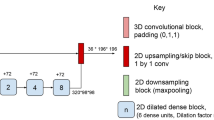
In this study, we used Entelai Neuro, a commercial segmentation software based on an adaptation of a fully convolutional densely connected network (Tiramisu architecture) following Oguz et al strategy for MS lesion quantification [20, 21]. Entelai Neuro uses 2.5D stacked slices independently for each orthogonal MR view (axial, sagittal, and coronal), which provide global context along the in-plane direction as well as local context in the out-of-plane direction by considering stacks of 3 contiguous slices. Differently from standard 2D convolutional models, the 2.5D stacked inputs provide local 3D context by concatenating multimodal neighboring slices along the channel dimension. At the same time, it substantially reduces the number of parameters when compared with 3D convolutional models which employ 3D kernels and has access to more samples for training by taking stacked data from three orientations. The model architecture is composed of an initial 2D convolutional layer, followed by 5 densely connected plus transitional steps blocks, and the symmetrical upsampling path, as depicted in Fig. 1. For more details about the Transition Down/Up modules and Dense Blocks, see [20]. Each densely connected block is an iterative concatenation of previous feature maps. This idea is founded on the observation that a feed-forward network composed of layers that are directly connected to all other layers can improve both accuracy and ease of training [22]. We also employed skip connections between the downsampling and upsampling paths to recover fine-grained information and avoid the smoothing effect caused by encoder-decoder architectures with bottleneck. We used focal loss function for training [23].
Detailed architecture for the DL segmentation model. The 2.5D input image is first processed by a standard 2D convolution, followed by a downsampling path which interleaves dense blocks (blue), concatenation modules (yellow), and transition down (orange) modules. The upsampling module recovers the original image resolution by combining transition up (green) modules, dense blocks, and concatenation modules. A more detailed description is included in “DL-based automated MS lesion quantification”
The model was implemented in Pytorch. We trained the model with 275 subjects, in an 80–10–10% (train-validation-test) scheme. All subjects contained 3D T1 and 3D FLAIR sequences as well as manually segmented MS lesion masks. Images were scaled to 1 mm spacing and FLAIR sequences were co-registered to T1 using rigid transformation (only rotation and translation operations permitted). We used the pre-trained model as the initialization of network parameters. Only one model was trained without an ensemble. We used the standard grid-search algorithm for hyper-parameter tunning and reported the results for the best-performing version, which is in fact the one used by the commercial version of Entelai Neuro.
Statistical analysis
To obtain a concurrent estimate of consistency and agreement between volumes derived from the different observers or methods, we computed intraclass correlation coefficients (ICC) [24]. The ICC assesses reproducibility between repeated measures within one subject by comparing the variability between the repeated measures with the total variability of the data (Supplementary Fig. 1A). A strong correlation would confirm a good consistency between techniques. ICCs were computed automatically specifying a two-way mixed-effect model.
To assess spatial agreement, we used DC between segmentations generated by the different human operators (or the consensus among them) and the output binary segmentations generated by the different software, using manual segmentations as ground truth [25]. DC is defined as two times the area of the intersection of A and B, divided by the sum of the areas of A and B (Supplementary Fig. 1B).
The robustness (repeatability and reproducibility) of repeated measures was assessed using the within-subject DC and coefficient of variation (CV) for each method. CV may be defined as the ratio of the standard deviation of a number of measurements to the arithmetic mean (Supplementary Fig. 1C) [26]. A software is robust if its output is consistently accurate even if one or more of the input variables are changed. For robustness estimation, four variables were defined: same-scanner same-visit (SSSV), same-scanner different-visit (SSDV), different-scanner same-visit (DSSV), and different-scanner different-visit (DSDV).
All statistical analyses were performed using R version 4.2.0. Group comparisons between methods were tested using the Kruskal–Wallis rank test, and in case of significant differences, post hoc paired analysis was performed using the Wilcoxon rank-sum test. A p < 0.05 was considered statistically significant. The Checklist for Artificial Intelligence in Medical Imaging (CLAIM) was used for reporting in this study [27].
Results
Subjects
Demographic and main clinical data from all subjects included in this study are summarized in Table 1.
Correlation and agreement between manual segmentations
Test–retest reliability between manual segmentations achieved an ICC of 0.89 (95% CI 0.73–0.95, p < 0.001) between observer 1 and 2, 0.89 (95% CI 0.73–0.95, p < 0.001) between observer 2 and 3, and 0.99 (95% CI 0.97–0.99, p < 0.001) between observer 1 and 3 (Supplementary Fig. 2). We found a median DC for all lesion sizes and all observers of 0.63. Spatial agreement was lower between observers on subjects with low lesion load (less than 5 mL) with a median DC of 0.55 compared to the agreement between observers on subjects with medium lesion load (between 5 and 15 mL) with a median DC of 0.63 and high lesion load (more than 15 mL) with a median DC of 0.67. The median DC between observers categorized by lesion load size is summarized in Supplementary Table 4. DC differences between observer couples were nonsignificant. It took observers a mean time of 44 ± 33 min to segment each subject (17 ± 12 min on low, 44 ± 24 min on medium, and 69 ± 28 min on high lesion load subjects).
The seven annotators that performed the segmentations for the MSSEG challenge dataset had a similar performance with a median DC (range) of 0.68 (0.52–0.84), confirming the expertise of the raters and quality of the ground truth masks (Supplementary Fig. 3).
Correlation and agreement between DL- and non-DL-based automated segmentations
The correlation was higher between ground truth and Entelai Neuro both in the internal and external datasets (both ICC 0.96, 95% CI 0.89–0.98) when compared to nicMSlesions (internal ICC 0.84, 95% CI 0.63–0.94 and external ICC 0.64, 95% CI 0.26–0.85), LGA (internal ICC 0.01, 95% CI − 0.42–0.44 and external ICC 0.78, 95% CI 0.50–0.91), and LPA algorithms (internal ICC 0.93, 95% CI 0.83–0.97 and external ICC 0.78, 95% CI 0.51–0.91). The correlation was also higher between the consensus ground truth and Entelai Neuro on the challenge dataset (ICC 0.86, 95% CI 0.76–0.92) when compared to nicMSlesions (ICC 0.55, 95% CI 0.33–0.72), LGA (ICC 0.45, 95% CI 0.20–0.65), and LPA algorithms (ICC 0.83, 95% CI 0.72–0.90). Correlation results are graphed in Supplementary Figs. 4, 5, and 6.
On the internal dataset, spatial agreement between Entelai Neuro segmentation masks and ground truth was higher than LGA, LPA, and nicMSlesions (median DC 0.73 vs 0.41, 0.57, and 0.53 respectively, p < 0.05). On the external dataset, Entelai Neuro maintained a higher spatial agreement with ground truth compared to LGA, LPA, and nicMSlesions (median DC 0.66 vs 0.48, 0.30, and 0.43 respectively, p < 0.05). Moreover, Entelai Neuro performance on the external dataset showed no statistically significant difference when compared to the performance on the internal dataset (median DC 0.73 vs 0.66, p = 0.093). Finally, on the challenge dataset, Entelai Neuro also had a higher spatial agreement with ground truth compared to LGA, LPA, and nicMSlesions (median DC 0.70 vs 0.53, 0.48, and 0.58 respectively, p < 0.05). Spatial agreement results are summarized in Tables 2, 3, and 4, Figs. 2, 3, 4, and 5, and Supplementary Fig. 7.
Example subject from the internal dataset (35-year-old male) with original 3D FLAIR sequence (first column) and manual (second column), LGA (third column), LPA (fourth column), nicMSlesions (fifth column), and Entelai Neuro (sixth column) segmentations masks over axial (first row), sagittal (second row), and coronal (third row) multiplanar reformatted 3D FLAIR sequence
Robustness of brain lesion segmentation
Entelai Neuro maintained a higher DC in all four variables (SSSV, SSDV, DSSV, and DSDV) ranging from 0.82 to 0.90. Differences were statistically significant in all four variables when compared to LGA and nicMSlesions and in all-but-one variable (SSSV) when compared to LPA. The intra-method spatial agreement was worst in the DSDV group followed by DSSV in every method. Also, we found that Entelai Neuro had the lowest CV among all methods, ranging from 0.7 to 7.9%. Robustness experiment results are summarized in Table 5, Fig. 6, and Supplementary Table 5.
Processing time
The mean processing time for each segmentation was 48 ± 40 min for the manual operator, 21 ± 4 min for LPA, 54 ± 9 min for LGA, 126 ± 221 min for nicMSlesions, and 2.3 ± 0.08 min for Entelai Neuro.
Discussion
Here, we showed that MS lesion manual segmentation is prone to inter-observer variability. We then validated a generalizable and robust DL-based software for MS lesion segmentation that uses 3D T1 and FLAIR sequences as an input. The model was fully trained with real-world clinical data. We tested the model on both internal and external datasets from Latin American and European centers, including images acquired in different models of all three major MRI scanner vendors (GE, Philips, and Siemens). The model, which is fully automated, not only outperformed other MS lesion segmentation software, but also maintains its performance on unseen data.
Manual segmentation of MS lesions is a task in which even trained neuroradiologists do not achieve very high agreement. We obtained median DC of 0.63 and 0.68 on clinical- and challenge-based datasets, respectively, similar to what was found by other authors [28]. Thus, as manual segmentations are commonly used as ground truth for the evaluation of automated methods, their performances are far from perfect. We believe the accuracy in terms of DC achieved by Entelai Neuro in this setting is highly competitive.
It is worth noting that there is only a moderate degree of correlation (ICC = 0.45) between LPA and ground truth on the external challenge-based dataset and a very low degree of correlation (ICC = 0.01) between LGA and ground truth on the internal dataset. This is due to the presence of outliers which have extreme lesion load volumes in LPA and LGA masks respectively. After visually inspecting these segmentation masks, we identified errors which segmented normal-appearing gray and WM. As it is known, outliers can have a very large effect on the line of best fit and the correlation coefficient, as happens in this case. That is why, spatial correlation metrics (like DC) are preferred when evaluating the performance of segmentation models.
Several AI-based solutions have been proposed, developed, and tested for the MS lesion segmentation task and have been extensively reviewed [9, 10, 29,30,31]. However, most of them were not tested on real clinical scenarios. Based on the recommendations for MS protocol harmonization [32], we chose to use standard practice 3D unenhanced MRI sequences (T1 and FLAIR) as input, instead of a 2D or multichannel approach [33]. We also opted to test the performance of this novel model both with real-world and challenge-based data, as data from challenges tends to be highly curated and controlled [15]. Differently from previous studies which only use images obtained in Europe and the USA, here we also included clinical datasets captured in Latin America.
Due to its recent success in computer vision, medical image analysis, and brain lesion segmentation, we chose to use supervised DL algorithms for this task [34,35,36]. As DL architectures become more mature, they gradually outperform previous state-of-the-art classical ML algorithms.
Our model had a median DC (interquartile range) of 0.70 (0.57–0.77) on 87 subjects (derived from internal and external datasets), similar to other published DL algorithms for MS lesion segmentation like DeepLesionBrain, nicMSlesions, DeepMedic, and Tiramisu with 2.5D stacked slices [19, 20, 37]. DeepLesionBrain, which is based on a large group of compact 3D CNNs, includes data augmentation and hierarchical specialization learning to reduce dependency with respect to training data specificity, reporting a DC of 0.66 on a series of cross-dataset experiments [37]. NicMSlesions latest version, which is based on a cascade of two 3D patchwise CNNs, achieved a DC of 0.58 on a challenge dataset [13]. NicMSlesions performance was also validated on 14 subjects with MS [38] achieving a mean DC between 0.49 and 0.66, depending on whether the model was used as default, was optimized, and/or trained. By directly comparing model performance, we are able to ascertain that our model outperformed nicMSlesions both in the internal and external datasets. Tiramisu with 2.5D stacked slices is based on a fully convolutional densely connected network. This approach is the one that is more similar to ours, and the best-performing variant of their model obtained a DC of 0.69 on a challenge dataset [20].
Differently from most studies proposing DL-based models for MS lesion segmentation which evaluate their models in laboratory conditions, our paper focuses on measuring model robustness in real clinical datasets, with repeatability and reproducibility exceeding traditional artificial intelligence-based software. We also tested the model using a wide range of lesion load volumes, as MS patients have different disease duration and burden, and segmentation models should perform similarly in all of them. This issue is particularly important in patients with low lesion load, where most algorithms tend to show degraded performance.
This work has limitations that need to be taken into consideration. First, although several scanner vendors and models, field strengths, and sequence variations were tested, we are far from evaluating the algorithm in all possible clinical scenarios. Second, although we included a total of 87 subjects—adding both internal and external datasets—the sample size of the testing group is modest. However, as previously stated, we included subjects with a wide range of lesion load (to contemplate the different forms of MS and spectrum of patients with this disease), coming from Latin America and Europe. Finally, we did not evaluate the clinical validation or workflow integration of the model; we plan to address this issue in future works.
Conclusion
Automated lesion segmentation in MS is reliable and shows good agreement with manual segmentation. Entelai Neuro, a DL-based method, outperformed non-DL-based and DL-based methods in the task of MS lesion segmentation on real-world and challenged-derived data. The model also proved to generalize well on unseen data and has a robust performance.
Abbreviations
- CV:
-
Coefficient of variation
- DC:
-
Dice coefficient
- DL:
-
Deep learning
- DSDV:
-
Different-scanner different-visit
- DSSV:
-
Different-scanner same-visit
- ICC:
-
Intraclass correlation coefficient
- LGA:
-
Lesion growth algorithm
- LPA:
-
Lesion prediction algorithm
- LST:
-
Lesion segmentation tool
- MS:
-
Multiple sclerosis
- SSDV:
-
Same-scanner different-visit
- SSSV:
-
Same-scanner same-visit
- WM:
-
White matter
References
Reich DS, Lucchinetti CF, Calabresi PA (2018) Multiple sclerosis. N Engl J Med 378:169–180. https://doi.org/10.1056/NEJMra1401483
Rodríguez Murúa S, Farez MF, Quintana FJ (2022) The immune response in multiple sclerosis. Annu Rev Pathol 17:121–139. https://doi.org/10.1146/annurev-pathol-052920-040318
Young IR, Hall AS, Pallis CA et al (1981) Nuclear magnetic resonance imaging of the brain in multiple sclerosis. Lancet 318:1063–1066. https://doi.org/10.1016/S0140-6736(81)91273-3
McDonald WI, Compston A, Edan G et al (2001) Recommended diagnostic criteria for multiple sclerosis: guidelines from the International Panel on the diagnosis of multiple sclerosis. Ann Neurol 50:121–127. https://doi.org/10.1002/ana.1032
Thompson AJ, Banwell BL, Barkhof F et al (2018) Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. Lancet Neurol 17:162–173. https://doi.org/10.1016/S1474-4422(17)30470-2
Filippi M, Rocca MA, Ciccarelli O et al (2016) MRI criteria for the diagnosis of multiple sclerosis: MAGNIMS consensus guidelines. Lancet Neurol 15:292–303. https://doi.org/10.1016/S1474-4422(15)00393-2
on behalf of the MAGNIMS study group, Geraldes R, Ciccarelli O et al (2018) The current role of MRI in differentiating multiple sclerosis from its imaging mimics. Nat Rev Neurol 14:199–213. https://doi.org/10.1038/nrneurol.2018.14
Gasperini C, Prosperini L, Tintoré M et al (2019) Unraveling treatment response in multiple sclerosis: a clinical and MRI challenge. Neurology 92:180–192. https://doi.org/10.1212/WNL.0000000000006810
Mortazavi D, Kouzani AZ, Soltanian-Zadeh H (2012) Segmentation of multiple sclerosis lesions in MR images: a review. Neuroradiology 54:299–320. https://doi.org/10.1007/s00234-011-0886-7
García-Lorenzo D, Francis S, Narayanan S et al (2013) Review of automatic segmentation methods of multiple sclerosis white matter lesions on conventional magnetic resonance imaging. Med Image Anal 17:1–18. https://doi.org/10.1016/j.media.2012.09.004
Gryska E, Schneiderman J, Björkman-Burtscher I, Heckemann RA (2021) Automatic brain lesion segmentation on standard magnetic resonance images: a scoping review. BMJ Open 11:e042660. https://doi.org/10.1136/bmjopen-2020-042660
Zeng C, Gu L, Liu Z, Zhao S (2020) Review of deep learning approaches for the segmentation of multiple sclerosis lesions on brain MRI. Front Neuroinformatics 14:610967. https://doi.org/10.3389/fninf.2020.610967
Valverde S, Salem M, Cabezas M et al (2019) One-shot domain adaptation in multiple sclerosis lesion segmentation using convolutional neural networks. NeuroImage Clin 21:101638. https://doi.org/10.1016/j.nicl.2018.101638
Schmidt P, Gaser C, Arsic M et al (2012) An automated tool for detection of FLAIR-hyperintense white-matter lesions in Multiple Sclerosis. Neuroimage 59:3774–3783. https://doi.org/10.1016/j.neuroimage.2011.11.032
Commowick O, Istace A, Kain M et al (2018) Objective evaluation of multiple sclerosis lesion segmentation using a data management and processing infrastructure. Sci Rep 8. https://doi.org/10.1038/s41598-018-31911-7
Yushkevich PA, Piven J, Hazlett HC et al (2006) User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31:1116–1128. https://doi.org/10.1016/j.neuroimage.2006.01.015
Akhondi-Asl A, Hoyte L, Lockhart ME, Warfield SK (2014) A logarithmic opinion pool based STAPLE algorithm for the fusion of segmentations with associated reliability weights. IEEE Trans Med Imaging 33:1997–2009. https://doi.org/10.1109/TMI.2014.2329603
Schmidt P (2017) Bayesian inference for structured additive regression models for large-scale problems with applications to medical imaging. Text.PhDThesis, Ludwig-Maximilians-UniversitätMünchen
Valverde S, Cabezas M, Roura E et al (2017) Improving automated multiple sclerosis lesion segmentation with a cascaded 3D convolutional neural network approach. Neuroimage 155:159–168. https://doi.org/10.1016/j.neuroimage.2017.04.034
Zhang H, Valcarcel AM, Bakshi R et al (2019) Multiple sclerosis lesion segmentation with Tiramisu and 2.5D Stacked Slices. In: Shen D, Liu T, Peters TM et al (eds) Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. Springer International Publishing, Cham, pp 338–346
Jégou S, Drozdzal M, Vazquez D, et al (2017) The one hundred layers tiramisu: Fully convolutional densenets for semantic segmentation. In Proceedings of the IEEE conference on computer vision and pattern recognition workshops, pp 11–19
Huang G, Liu Z, Van Der Maaten L, Weinberger KQ (2017) Densely connected convolutional networks. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE, Honolulu, HI, pp 2261–2269
Lin T-Y, Goyal P, Girshick R, et al (2017) Focal loss for dense object detection. In: 2017 IEEE International Conference on Computer Vision (ICCV). IEEE, Venice, pp 2999–3007
Shrout PE, Fleiss JL (1979) Intraclass correlations: uses in assessing rater reliability. Psychol Bull 86:420–428. https://doi.org/10.1037/0033-2909.86.2.420
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302. https://doi.org/10.2307/1932409
Hendricks WA, Robey KW (1936) The sampling distribution of the coefficient of variation. Ann Math Stat 7:129–132. https://doi.org/10.1214/aoms/1177732503
Mongan J, Moy L, Kahn CE (2020) Checklist for Artificial Intelligence in Medical Imaging (CLAIM): a guide for authors and reviewers. Radiol Artif Intell 2:e200029. https://doi.org/10.1148/ryai.2020200029
Egger C, Opfer R, Wang C et al (2017) MRI FLAIR lesion segmentation in multiple sclerosis: does automated segmentation hold up with manual annotation? NeuroImage Clin 13:264–270. https://doi.org/10.1016/j.nicl.2016.11.020
Danelakis A, Theoharis T, Verganelakis DA (2018) Survey of automated multiple sclerosis lesion segmentation techniques on magnetic resonance imaging. Comput Med Imaging Graph 70:83–100. https://doi.org/10.1016/j.compmedimag.2018.10.002
Zhang H, Oguz I (2021) Multiple sclerosis lesion segmentation-a survey of supervised cnn-based methods. In Brainlesion: Glioma, Multiple Sclerosis, Stroke and Traumatic Brain Injuries: 6th International Workshop, BrainLes 2020, Held in Conjunction with MICCAI 2020, Lima, Peru, October 4, 2020, Revised Selected Papers, Part I, vol 6. Springer International Publishing, pp 11–29
Kaur A, Kaur L, Singh A (2021) State-of-the-art segmentation techniques and future directions for multiple sclerosis brain lesions. Arch Comput Methods Eng 28:951–977. https://doi.org/10.1007/s11831-020-09403-7
De Stefano N, Battaglini M, Pareto D et al (2022) MAGNIMS recommendations for harmonization of MRI data in MS multicenter studies. NeuroImage Clin 34:102972. https://doi.org/10.1016/j.nicl.2022.102972
Shiee N, Bazin P-L, Ozturk A et al (2010) A topology-preserving approach to the segmentation of brain images with multiple sclerosis lesions. Neuroimage 49:1524–1535. https://doi.org/10.1016/j.neuroimage.2009.09.005
Krizhevsky A, Sutskever I, Hinton GE (2012) Imagenet classification with deep convolutional neural networks. In: Advances in Neural Information Processing Systems
Chan H-P, Samala RK, Hadjiiski LM, Zhou C (2020) Deep learning in medical image analysis. In: Lee G, Fujita H (eds) Deep Learning in Medical Image Analysis. Springer International Publishing, Cham, pp 3–21
Akkus Z, Galimzianova A, Hoogi A et al (2017) Deep learning for brain MRI segmentation: state of the art and future directions. J Digit Imaging 30:449–459. https://doi.org/10.1007/s10278-017-9983-4
Kamraoui RA, Ta V-T, Tourdias T et al (2022) DeepLesionBrain: towards a broader deep-learning generalization for multiple sclerosis lesion segmentation. Med Image Anal 76:102312. https://doi.org/10.1016/j.media.2021.102312
Weeda MM, Brouwer I, de Vos ML et al (2019) Comparing lesion segmentation methods in multiple sclerosis: input from one manually delineated subject is sufficient for accurate lesion segmentation. NeuroImage Clin 24:102074. https://doi.org/10.1016/j.nicl.2019.102074
Acknowledgements
The authors gratefully acknowledge the support of NVIDIA Corporation with the donation of a Titan XP GPU used for this research. EF, DFS, and DS were supported by Fundación Sadosky.
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Hernán Chaves.
Conflict of Interest
The listed authors declare relationships with the following companies:
• Diego Fernández Slezak: is CTO and co-founder of Entelai.
• Diego E. Shalom: has received stipends as a scientific advisor from Entelai.
• Enzo Ferrante: has received stipends as a scientific advisor from Entelai.
• Pilar Ananía: Entelai employee.
• Felipe Kitamura: consultant for MD.ai and employed by DASA.
• Hernán Chaves: has received stipends as a medical advisor from Entelai.
• Jorge Correale: received stipends from Biogen, Merck, Novartis, Roche, Bayer, Sanofi-Genzyme, Gador, Raffo, Bristol Myers Squibb, and Janssen.
• María Mercedes Serra: has received stipends as a medical advisor from Entelai.
• Martín Elías Costa: Entelai employee.
• Mauricio Franco Farez: is CEO and co-founder of Entelai.
Statistics and Biometry
One of the authors (Mauricio Franco Farez) has significant statistical expertise.
Informed Consent
Written informed consent was waived by the Institutional Review Board.
Ethical Approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
No study subjects or cohorts have been previously reported.
Methodology
• prospective and retrospective
• diagnostic and observational study
• multicenter study
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chaves, H., Serra, M.M., Shalom, D.E. et al. Assessing robustness and generalization of a deep neural network for brain MS lesion segmentation on real-world data. Eur Radiol 34, 2024–2035 (2024). https://doi.org/10.1007/s00330-023-10093-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-10093-5