Abstract
Objective
To investigate the feasibility of a deep learning–based detection (DLD) system for multiclass lesions on chest radiograph, in comparison with observers.
Methods
A total of 15,809 chest radiographs were collected from two tertiary hospitals (7204 normal and 8605 abnormal with nodule/mass, interstitial opacity, pleural effusion, or pneumothorax). Except for the test set (100 normal and 100 abnormal (nodule/mass, 70; interstitial opacity, 10; pleural effusion, 10; pneumothorax, 10)), radiographs were used to develop a DLD system for detecting multiclass lesions. The diagnostic performance of the developed model and that of nine observers with varying experiences were evaluated and compared using area under the receiver operating characteristic curve (AUROC), on a per-image basis, and jackknife alternative free-response receiver operating characteristic figure of merit (FOM) on a per-lesion basis. The false-positive fraction was also calculated.
Results
Compared with the group-averaged observations, the DLD system demonstrated significantly higher performances on image-wise normal/abnormal classification and lesion-wise detection with pattern classification (AUROC, 0.985 vs. 0.958; p = 0.001; FOM, 0.962 vs. 0.886; p < 0.001). In lesion-wise detection, the DLD system outperformed all nine observers. In the subgroup analysis, the DLD system exhibited consistently better performance for both nodule/mass (FOM, 0.913 vs. 0.847; p < 0.001) and the other three abnormal classes (FOM, 0.995 vs. 0.843; p < 0.001). The false-positive fraction of all abnormalities was 0.11 for the DLD system and 0.19 for the observers.
Conclusions
The DLD system showed the potential for detection of lesions and pattern classification on chest radiographs, performing normal/abnormal classifications and achieving high diagnostic performance.
Key Points
• The DLD system was feasible for detection with pattern classification of multiclass lesions on chest radiograph.
• The DLD system had high performance of image-wise classification as normal or abnormal chest radiographs (AUROC, 0.985) and showed especially high specificity (99.0%).
• In lesion-wise detection of multiclass lesions, the DLD system outperformed all 9 observers (FOM, 0.962 vs. 0.886; p < 0.001).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Chest radiograph is the most common chest imaging modality and is readily available and affordable, with short examination time and low radiation exposure. Because the chest radiograph presents the thorax on a two-dimensional image, overlapping of structures with each other, and sometimes with abnormal lesions, is inevitable. This inherent limitation makes it challenging to detect subtle abnormalities on chest radiograph, even by experienced radiologists [1, 2]. In fact, the sensitivity of chest radiographs for malignant pulmonary nodules had been reported to be 36% to 84% [3,4,5,6]. Moreover, the tremendous number of chest radiographs can make timely reading difficult.
In this regard, there had been large efforts to develop and apply computer-aided detection (CAD) on chest radiograph. Several previous studies have shown the good performance of CAD [7,8,9,10]. Most recently, Nam et al [10] reported that their deep learning–based algorithm, as a second reader, significantly enhanced all 18 physicians’ performance for nodule detection on chest radiograph; in fact, its stand-alone performance was superior to that of 15 of 18 physicians.
However, in addition to nodules, various disease patterns need to be evaluated on chest radiograph. Recent studies have addressed the benefit of automatic detection systems for such purposes [11,12,13,14,15]. Cicero et al [11] showed that with the use of a training dataset that was based on chest radiograph reports, their deep learning–based model diagnosed five categories of abnormalities (i.e., pulmonary edema, effusion, pneumothorax, cardiomegaly, and consolidation) (area under the curve (AUC), 0.85–0.96). Dunnmon et al [12] and Annarumma et al [13] showed a clinically acceptable performance of automated classification and triaging of chest radiographs, in terms of the presence of abnormalities and based on the detection and classification of multiple patterns. Hwang et al [14] also demonstrated that a deep learning–based algorithm outperformed readers in the discrimination of chest radiographs that showed major thoracic diseases and assisted the readers in improving their performance. However, despite the consistent good performance of such algorithm, the accuracy of disease diagnosis remains insufficient (AUC, 0.686), probably because of overlaps in the radiologic findings of major thoracic diseases. Given that the abnormal patterns detected by automatic detection systems are reviewed by radiologists, a more practical approach might be learning abnormal image patterns rather than diseases.
On this premise, in our study, we selected four classes (i.e., nodule/mass, pleural effusion, pneumothorax, and interstitial opacity) that are clinically important and common and do not overlap with each other. We aimed to investigate the feasibility of a deep learning–based detection (DLD) system for multiclass lesions on chest radiograph, in comparison with observers’ performance.
Materials and methods
This retrospective study was approved by the institutional review board, which waived the requirement for patients’ informed consent. All images were deidentified of patient information.
Dataset for development of the DLD system
We retrospectively collected and anonymized 15,809 chest radiographs from two institutions (Asan Medical Center (institution A) and Seoul National University Bundang Hospital (institution B)). The images from institution A comprised 6119 normal chest radiographs and 4151 chest radiographs with the abnormalities of interest (i.e., nodule/mass, pleural effusion, interstitial opacity, and pneumothorax), whereas the images from institution B comprised 1085 normal chest radiographs and 4454 chest radiographs with the same abnormalities of interest (Table 1).
To build ground truth, two radiologists at institution A outlined the lesions on each half of the abnormal radiographs with reference to computed tomography (CT); in cases without CT for reference, lesions were evaluated and outlined by two radiologists in consensus. In institution A, all cases of nodule/mass and interstitial opacity had reference CT available, and 368 of 1287 pleural effusion cases and 101 of 263 pneumothorax cases had reference CT. In institution B, one radiologist outlined the lesions on all of the abnormal chest radiographs with reference to CT.
Except for 200 radiographs that comprised the test set, all radiographs were randomly split by the frequency of abnormalities into a 9:1 ratio of the training dataset (normal, 6394; abnormal, 7654) to the validation dataset (normal, 710; abnormal, 851) using a stratified sampling that reflected the institutions.
Development of the DLD system
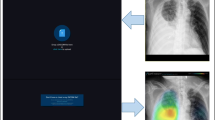
We designed and trained a multitask convolutional neural network (CNN), in which the input comprised the chest radiograph and the outputs were the generated binary classifications (normal/abnormal) and lesion segmentation (Fig. 1). The chest radiographs were resized to a 1024 × 1024 resolution. Then, this image was converted to four images with 512 × 512 resolution; this input passed through the CNN to generate a binary normal/abnormal classification output; the feature maps were used to generate a multiclass lesion segmentation mask, which was a five-channel two-dimensional image, with each channel corresponding to the predicted probability of each pixel being normal or as one of the four abnormalities of interest. We trained the network for 100 epochs, with a learning rate starting from 0.001, divided by 10 upon training loss saturation. After the development of our DLD system, it could analyze one chest radiograph for less than 1 s. Further details are provided in the Supplementary Material.
The multitask learning architecture of the deep convolutional neural network used to generate DLD outputs. We designed and trained a multitask convolutional neural network, which includes chest radiograph as the input and generates both binary classifications (normal/abnormal) and lesion segmentation outputs. The chest radiographs are resized to 1024 × 1024 resolution before the image is converted to four images with 512 × 512 resolution. After the 512 × 512 × 4 input passes through multiple convolution blocks, the feature maps generate a binary normal/abnormal classification output and predict the probability of each pixel being either normal or as one of the four abnormalities of interest
Performance test
The performance test was applied on 200 chest radiographs (100 normal and 100 abnormal). The number of lesions/patients was 83/70 for nodule/mass, 19/10 for interstitial opacity, 11/10 for pleural effusion, and 10/10 for pneumothorax (Table 2). To prepare the test set, chest radiographs were randomly selected from each category, in equal numbers from each hospital. The sizes of the nodules/masses in the test set were measured on chest radiograph by one radiologist who did not participate in the observer test.
At first, the DLD performance was assessed. The DLD classified each lesion detected on chest radiograph and rated the probability of it being a true lesion, which was presented as a continuous value between 0 and 1. DLD-generated annotations were considered true positive (TP) when the centers of the marks that matched the classification were within the ground truth region of interest (ROI).
For the observer performance test, nine readers with varying levels of experience participated. They consisted of seven board-certified radiologists with 2–19 years of experience on chest radiology (observers 1–7) and two radiology residents (observers 8 and 9). The images were evaluated using in-house software that had available options for magnification and adjustment of window settings. The observers were requested to report the detected lesions on chest radiographs, based on the four classes of abnormal findings. Other opacities (e.g., calcified nodule, fibrotic sequelae) that were not included in these categories were indicated to be ignored. The observers placed a 1 cm × 1 cm mark on each perceived lesion on the radiograph and rated the confidence level on a five-point scale: grade 1 (indeterminate) to grade 5 (definite). The readers’ marks were considered TP when the centers of the marks that matched the classification were within the ground truth ROI. Radiographs were defined as normal, when the observers did not report any abnormal finding. Observers were not time limited in the interpretation of chest radiographs.
Statistical analysis
The performances of the developed DLD system and the observers in classifying chest radiographs as normal or abnormal and in detecting multiclass lesions were evaluated by the area under the receiver operating characteristic (ROC) curve (AUROC) analysis on a per-image basis and by jackknife alternative free-response ROC curve (JAFROC, version 4.2.1; http://www.devchakraborty.com) analysis on a per-lesion basis. The AUROC and the figure of merit (FOM), which was defined as the probability that a lesion was rated higher than the highest-rated non-lesion on normal images [16], were calculated for all classes of abnormalities together and for each individual class of abnormality. In the subgroup analysis, the performance of the DLD was calculated separately for nodule/mass and for the other three abnormal classes (interstitial opacity, pleural effusion, and pneumothorax) combined. The performance of the DLD was compared with that of the pooled observers and each individual observer using pairwise comparison of the ROC curve analysis and JAFROC FOMs. The random-case fixed-reader method was used for each observer and experience-based grouped-observer comparison, and the random-case random-reader method was used for averaged comparisons among all nine observers.
Sensitivity was calculated at a lesion level as the number of TP markings divided by the total number of ground truth lesions. The method of generalized estimating equations was used to compare the sensitivities between the DLD system and the observers. The false-positive (FP) fraction was calculated as the number of FP markings, in which the classification or localization did not match, divided by the total number of chest radiographs. Specificity was calculated at an image level as the number of true negative cases divided by the total number of normal chest radiographs.
ROC analysis was performed using R software (version 3.3.0; http://www.Rproject.org). To account for within-patient correlation, the Dorfman–Berbaum–Metz significance test with Hillis’ improvements was applied using JAFROC, version 4.2.1. Data are presented as mean ± standard deviation. A p value < 0.05 was considered to indicate statistical significance. A p value correction was performed using the Bonferroni method for comparison of the DLD with experience-based observer subgroups by multiplying p values by 2 and for comparison of the DLD with each observer by multiplying p values by 9 [17].
Results
Performance of the DLD system
For the task of classifying chest radiographs as normal or abnormal, the DLD achieved an AUROC of 0.985 and a specificity of 99.0% (99/100). The AUROC of the DLD was 0.971 for nodule/mass and 0.995 for the other three abnormal classes. For the detection and pattern classification tasks for all abnormalities, the DLD achieved an FOM of 0.943 and a sensitivity of 88.6% (109/123), with 0.11 FP fractions. In the subgroup analysis, the DLD showed an FOM of 0.913 with 0.07 FP fractions for nodule/mass and an FOM of 0.995 with 0.05 FP fractions for the other three abnormal classes. The sensitivities were 84.3% (70/83) and 100.0% (40/40) for nodule/mass and the other three abnormal classes, respectively. Among the other three abnormal classes, the DLD detected and accurately classified all pneumothorax lesions without generating any FP annotation (FOM = 1.000). The DLD showed similar high performances for interstitial opacity (FOM 0.997 with a FP fraction of 0.02) and pleural effusion (FOM of 0.998 with a FP fraction of 0.03).
Comparison of performances between the DLD and the observers
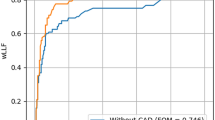
For the task of classifying chest radiographs as normal and abnormal, the nine observers had a pooled AUROC of 0.958 (range, 0.915–0.976) and pooled specificity of 90.7% (816/900). There was no significant difference in pooled performance between the seven radiologists and the two radiology residents (AUROC, 0.958 vs. 0.957, respectively; p = 0.939). Performance was significantly better with the DLD than with the pooled observers (AUROC, 0.985 vs. 0.958, p = 0.001) (Fig. 2).
ROC curves of the DLD and pooled observer performances for all abnormalities on a per-image basis. AUROC analysis showed that the DLD performance for classification as normal or abnormal chest radiograph is significantly higher than the pooled observer performance (AUROC, 0.985 vs. 0.958; p = 0.001)
The superiority of performance of the DLD to that of the pooled observers was consistent for the detection of both nodule/mass (AUROC, 0.971 vs. 0.925, p < 0.001) and the other three abnormal classes (AUROC, 0.995 vs. 0.969, p = 0.001) (Table 3).
For the tasks of detection and pattern classification for all abnormalities, the pooled FOM of the nine observers was 0.849, with an average FP fraction of 0.19. There was no significant difference in performance between the seven radiologists and the two radiology residents (FOM, 0.851 vs. 0.839, respectively, p = 0.661). The performance was significantly better with the DLD than with the pooled observers (FOM, 0.943 vs. 0.849, p < 0.001). The DLD showed better performance than all nine observers (FOM range, 0.817–0.880; p < 0.001 to 0.006) (Fig. 3).
The DLD consistently outperformed the pooled observers in the detection of both nodule/mass (FOM, 0.913 vs. 0.847, p < 0.001) and the other three abnormal classes (FOM, 0.995 vs. 0.843, p < 0.001). The DLD showed significantly better performance than 6 of 9 observers in nodule/mass detection (FOM range, 0.804–0.873; p < 0.001 to 0.282) (Figs. 4 and 5). For the detection of each class of the other three abnormal classes, the pooled observers had FOMs of 0.983 for pneumothorax, 0.756 for interstitial opacity, and 0.883 for pleural effusion (Table 4).
A 27-mm adenocarcinoma in the right lower lobe in a 56-year-old woman. a The nodule in the right lower lung zone (arrow) is faintly visible and overlaps with a rib shadow on the original chest radiograph. None of the observers detected the lesion. b The corresponding computed tomography (CT) image reveals a 27-mm nodule in the right lower lobe. c The DLD generates a true-positive mark on the lesion
A 16-mm biopsy-proven benign nodule in the left lower lobe in a 62-year-old woman. a The well-defined nodule overlaps with a left cardiac border (arrow) on the original chest radiograph. b The corresponding CT image reveals a 16-mm nodule in the left lower lobe. c The DLD system and all of the observers except one failed to detect the nodule. The lesion was confused with vessels and heart border
In terms of sensitivity and FP fraction, the pooled sensitivities of observers were 73.2% (810/1107) for all abnormalities, 75.4% (563/747) for nodule/mass, and 70.8% (255/360) for the other three abnormal classes. Compared with the observers, the DLD exhibited significantly higher sensitivities for the detection of all abnormalities (p < 0.001), nodule/mass (p = 0.003), and the other three abnormal classes (p < 0.001) (Fig. 6). The average FP fractions for the observers were 0.19 (range, 0.09–0.28) for all abnormalities, 0.11 (range, 0.05–0.28) for nodule/mass, and 0.08 (range, 0.03–0.15) for the other three abnormal classes.
Biopsy-proven usual interstitial pneumonia in a 52-year-old man. a Bilateral fine reticular opacities in basal lung zones on the original chest radiograph. Only two of the observers annotated true-positive marks on the bilateral lesions. b, c The corresponding CT images reveal pleura-based reticular opacities associated with traction bronchiectasis in both lower lobes. d Although the left-side mark has a somewhat wider range, the DLD generated true-positive marks on the bilateral lesions
Discussion
In our study, we demonstrated that the DLD system accurately classified chest radiographs with four abnormal classes as normal or abnormal. Furthermore, our system detected lesions with correct pattern classification, outperforming 9 of 9 observers.
Except for the system described by Hwang et al [14], previous CAD systems mainly focused on nodule or mass detection and, therefore, had limitations in utility for clinical practice. The main challenge for the detection of multiclass lesions on chest radiographs is the varying sizes and extents of lesions; a wide view is needed for large lesions, such as pleural effusion and pneumothorax, while preserving a high resolution for subtle lesions, such as nodule or opacity. To handle this conflicting requirement, our deep learning–based model exploited inverse pixel shuffling, which effectively expanded the field of view of the model without loss of resolution. Another challenge in the development of a DLD is reducing FP lesion detections. We used two types of loss for training (i.e., lesion segmentation and binary normal/abnormal classification) and effectively facilitated the suppression of FP lesion detection.
In terms of nodule/mass detection, the FOM of our DLD system (0.913) showed better performance than that by the observers (average FOM, 0.847 (0.804–0.873)). Compared with the observers, our system achieved higher sensitivity (84.3% vs. 75.4%) and smaller FP fractions (0.07 vs. 0.11). Our results are similar with those of Nam et al [10], who indicated that the deep learning–based automatic detection (DLAD) algorithm outperformed physicians. Notably, in that study, the FOMs of the DLAD algorithm and the board-certified radiologists (0.870 and 0.821, respectively) were slightly lower, compared with ours. This difference can be explained by the fact that the mean size of the lesions was greater in our study than in their study (3.7 cm vs. 2.6 cm), because our test set included masses, in addition to nodules.
In the study by Nam et al [10], one of the main strengths of their model was the markedly decreased FP fraction, compared with that of the previously reported conventional image processing–based CAD (0.02–0.34 vs. 0.9–3.3) [18,19,20,21]. Similarly, our model showed a reduced FP fraction (0.07) for nodule/mass detection, which was even lower than that in observers (0.11). Considering that high FP fraction of CAD may result in additional observer’s FP fractions and increase reading time, FP reduction in our study could show a technical advance of CAD and the possibility of its application to real practice. We believed that our system has the potential to help radiologists in clinical practice. To confirm our results, external validation or a prospective study is warranted.
Our DLD system had a significantly better performance, compared with that of the observers, and the difference in the FOM between the two was larger for the other three abnormal classes than for nodule/mass (0.152 vs. 0.066). This result was contrary to our expectation that the detection of the other three abnormal classes was easier than that of nodule/mass lesions and would be less different between the DLD and the radiologists. In the per-class analysis, there was no significant difference (p = 0.245) for pneumothorax, but there were significant differences for interstitial opacity (p < 0.001) and pleural effusion (p = 0.001). We assumed that readers were well trained and experienced to detect pneumothorax, which is a clinically important disease that requires attention, but were relatively less sensitive to report interstitial opacity and pleural effusion, unless these lesions were definite or severe. In this respect, our system may encourage readers to scrutinize less noticeable lesions by providing marks for suspicious lesions.
On the basis of accurate detection of multiclass lesions, our system showed high performance on the binary classification as normal or abnormal chest radiographs (AUROC, 0.985). These results suggested that the DLD can automatically prioritize abnormal chest radiographs, thereby improving the workflow and efficiency in the radiology department. Moreover, the feasibility of our system in detecting multiclass lesions can promote broader application of CAD on chest radiograph, which could be accelerated by covering additional disease patterns such as rib fractures, pulmonary tuberculosis, and cardiomegaly.
Our study had several limitations. First, it did not include temporal or external validation. However, we obtained datasets from two medical centers, using different reconstruction methods; therefore, the limitations of a single-center study may have been overcome to some extent. Further validation, such as external or a prospective study, is warranted before our model can be used more generally. Second, as our study was a feasibility study, we did not determine whether our system can be an effective CAD; our future work will expound on this. Nevertheless, we successfully demonstrated that our DLD system was feasible and had better performance, compared with that of the radiologists. Third, because our data were collected separately from disease-positive and disease-negative subjects on the basis of radiologic reports in a case-controlled manner, which is referred to as convenience sampling [22], spectrum bias may have been introduced. Therefore, we tried to include a wide spectrum of disease severity in the training set, which is expected to offset the spectrum bias to some extent. Fourth, the test set had a much higher prevalence of disease pattern than that in real clinical practice. Furthermore, the study condition, in which the readers were focused on only few and fixed disease patterns, may not have reflected the actual clinical situation. However, the disease patterns addressed in our study are clinically important and common. Therefore, we believe that our study closely resembled routine practice, compared with the other studies that dealt with nodules alone.
In conclusion, the DLD system showed the potential for detection of lesions and pattern classification on chest radiographs, performing normal/abnormal classifications and achieving high diagnostic performance.
Abbreviations
- AUC:
-
Area under the curve
- AUROC:
-
Area under the receiver operating characteristic curve
- CAD:
-
Computer-aided detection
- DLD:
-
Deep learning–based detection
- FOM:
-
Figure of merit
- FP:
-
False positive
- JAFROC:
-
Jackknife alternative free-response receiver operating characteristic curve
- ROC:
-
Receiver operating characteristic
- TP:
-
True positive
References
de Hoop B, Schaefer-Prokop C, Gietema HA et al (2010) Screening for lung cancer with digital chest radiography: sensitivity and number of secondary work-up CT examinations. Radiology 255:629–637
Kundel HL (1981) Predictive value and threshold detectability of lung tumors. Radiology 139:25–29
Quekel LG, Kessels AG, Goei R, van Engelshoven JMA (2001) Detection of lung cancer on the chest radiograph: a study on observer performance. Eur J Radiol 39:111–116
Toyoda Y, Nakayama T, Kusunoki Y, Iso H, Suzuki T (2008) Sensitivity and specificity of lung cancer screening using chest low-dose computed tomography. Br J Cancer 98:1602–1607
Li F, Arimura H, Suzuki K et al (2005) Computer-aided detection of peripheral lung cancers missed at CT: ROC analyses without and with localization. Radiology 237:684–690
Gavelli G, Giampalma E (2000) Sensitivity and specificity of chest X-ray screening for lung cancer: review article. Cancer 89:2453–2456
Bley TA, Baumann T, Saueressig U et al (2008) Comparison of radiologist and CAD performance in the detection of CT-confirmed subtle pulmonary nodules on digital chest radiographs. Invest Radiol 43:343–348
Kasai S, Li F, Shiraishi J, Doi K (2008) Usefulness of computer-aided diagnosis schemes for vertebral fractures and lung nodules on chest radiographs. AJR Am J Roentgenol 191:260–265
Li F, Hara T, Shiraishi J, Engelmann R, MacMahon H, Doi K (2011) Improved detection of subtle lung nodules by use of chest radiographs with bone suppression imaging: receiver operating characteristic analysis with and without localization. AJR Am J Roentgenol 196:W535–W541
Nam JG, Park S, Hwang EJ et al (2019) Development and validation of deep learning-based automatic detection algorithm for malignant pulmonary nodules on chest radiographs. Radiology 290:218–228
Cicero M, Bilbily A, Colak E et al (2017) Training and validating a deep convolutional neural network for computer-aided detection and classification of abnormalities on frontal chest radiographs. Invest Radiol 52:281–287
Dunnmon JA, Yi D, Langlotz CP, Re C, Rubin DL, Lungren MP (2019) Assessment of convolutional neural networks for automated classification of chest radiographs. Radiology 290:537–544
Annarumma M, Withey SJ, Bakewell RJ, Pesce E, Goh V, Montana G (2019) Automated triaging of adult chest radiographs with deep artificial neural networks. Radiology. https://doi.org/10.1148/radiol.2018180921:180921
Hwang EJ, Park S, Jin KN et al (2019) Development and validation of a deep learning-based automated detection algorithm for major thoracic diseases on chest radiographs. JAMA Netw Open 2:e191095
Park S, Lee SM, Kim N et al (2019) Application of deep learning-based computer-aided detection system: detecting pneumothorax on chest radiograph after biopsy. Eur Radiol. https://doi.org/10.1007/s00330-019-06130-x
Chakraborty DP (2006) Analysis of location specific observer performance data: validated extensions of the jackknife free-response (JAFROC) method. Acad Radiol 13:1187–1193
Bender R, Lange S (2001) Adjusting for multiple testing—when and how? J Clin Epidemiol 54:343–349
Schalekamp S, van Ginneken B, Koedam E et al (2014) Computer-aided detection improves detection of pulmonary nodules in chest radiographs beyond the support by bone-suppressed images. Radiology 272:252–261
Novak RD, Novak NJ, Gilkeson R, Mansoori B, Aandal GE (2013) A comparison of computer-aided detection (CAD) effectiveness in pulmonary nodule identification using different methods of bone suppression in chest radiographs. J Digit Imaging 26:651–656
Dellios N, Teichgraeber U, Chelaru R, Malich A, Papageorgiou IE (2017) Computer-aided detection fidelity of pulmonary nodules in chest radiograph. J Clin Imaging Sci 7:8–8
Schalekamp S, van Ginneken B, Karssemeijer N, Schaefer-Prokop CM (2014) Chest radiography: new technological developments and their applications. Semin Respir Crit Care Med 35:3–16
Park SH, Han K (2018) Methodologic guide for evaluating clinical performance and effect of artificial intelligence technology for medical diagnosis and prediction. Radiology 286:800–809
Funding
This study has received funding from the Industrial Strategic Technology Development Program (10072064, Development of Novel Artificial Intelligence Technologies To Assist Imaging Diagnosis of Pulmonary, Hepatic, and Cardiac Diseases and Their Integration into Commercial Clinical PACS Platforms), which is funded by the Ministry of Trade Industry and Energy (MI, South Korea).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Sang Min Lee.
Conflict of interest
The authors declare that they have no conflict of interest.
Statistics and biometry
The statistician of our institution (Seon Ok Kim) kindly provided statistical advice for this manuscript.
Informed consent
Written informed consent was waived by the institutional review board.
Ethical approval
Institutional review board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study
• multicenter study
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 18 kb)
Rights and permissions
About this article
Cite this article
Park, S., Lee, S.M., Lee, K.H. et al. Deep learning-based detection system for multiclass lesions on chest radiographs: comparison with observer readings. Eur Radiol 30, 1359–1368 (2020). https://doi.org/10.1007/s00330-019-06532-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-019-06532-x