Abstract
Environmental factors play a significant role in controlling growth, development and defense responses of plants. Changes in the abiotic environment not only significantly alter the physiological and molecular pathways in plants, but also result in attracting the insect pests that carry a payload of viruses. Invasion of plants by viruses triggers the RNA silencing based defense mechanism in plants. In counter defense the viruses have gained the ability to suppress the host RNA silencing activities. A new paradigm has emerged, with the recognition that plant viruses also have the intrinsic capacity to modulate host plant response to environmental cues, in an attempt to favour their own survival. Thus, plant–virus interactions provide an excellent system to understand the signals in crosstalk between biotic (virus) and abiotic stresses. In this review, we have summarized the basal plant defense responses to pathogen invasion while emphasizing on the role of RNA silencing as a front line of defense response to virus infection. The emerging knowledge indicates overlap between RNA silencing with the innate immune responses during antiviral defense. The suppressors of RNA silencing serve as Avr proteins, which can be recognized by the host R proteins. The defense signals also function in concert with the phytohormones to influence plant responses to abiotic stresses. The current evidence on the role of virus induced host tolerance to abiotic stresses is also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The sessile nature of plants makes them more prone to stress caused by environmental perturbations and attacks by various pathogens. High temperatures exert abiotic stress and can also result in an increase in the incidence of insect vectors that carry with them a load of pathogens. Amongst all pathogens, viruses being obligate parasites can replicate only inside living cells and are responsible for imparting number of diseases in animals and plants. Viruses are present in almost every ecosystem and biological entity and piggy bank on insects for their transmission and spread in plants (Gallet et al. 2018). They invade normal host cells and hijack the cellular machineries for their own replication and multiplication. Virus infected plants often show severe abnormalities like stunting, leaf curling, leaf mosaic pattern, necrosis, chlorosis, wilting etc. (Hull 2009). All these deformities have drastic effects on crop productivity and quality leading to substantial agricultural losses (Dawson and Hilf 1992).
In order to protect themselves, plants have developed sophisticated defense mechanisms, which are both unique and overlapping. Substantial progress has been made in identifying the key stress-responsive genes associated with individual stresses, yet relatively little information is available on the crosstalk in response to combined stresses, specifically the interactions between biotic and abiotic stress responses. The present scenario of climate change is predicted to have a negative impact on biotic stress resistance in plants (Chattopadhyay et al. 2019). Therefore, it is vital to identify the common abiotic stress- and pathogen-specific genes that confer multiple stress tolerance in plants. Here, we discuss the recent advances in understanding the common molecular mechanisms that operate in response to virus infection and abiotic stresses.
Understanding plant immunity
Plants, like all other organisms, have a sophisticated natural defence mechanism by which they are able to distinguish between self and nonself molecules (Wu et al. 2014). Their primary defences can be grouped into two broad categories viz. constitutive and induced. The constitutive defences include the structural barriers like cuticles, thorns, trichomes, cell wall etc. and chemical barriers formed by the deposition of lignin, callose, glucosinolates, saponins (phytoanticipins), terpenoids, phenolics etc. (Freeman and Beattie 2008; Moreira et al. 2012). Initial defense responses also include alkalization of medium due to change in concentrations of H+, K+ and Ca2+ ions across the plasma membrane (Jabs et al. 1997). Elevation of cytoplasmic Ca2+ also serves as a signal for generation of reactive oxygen species (ROS), production of salicylic acid (SA) and closure of stomata (Nomura et al. 2008; Carr et al. 2010).
The induced defenses are activated during infection to orchestrate transcriptional reprogramming using the basal defense systems (Dodds and Rathjen 2010; Tsuda and Katagiri 2010) viz. Pathogen associated molecular pattern triggered immunity (PTI) and effector triggered immunity (ETI). The presence of PTI limits pathogen growth and puts selective pressure on them to invade the host cells (Thomma et al. 2011). Successful pathogens have evolved to produce different types of virulence factors or effectors for suppressing PTI. In retribution, the host have also evolved to produce specific R-proteins that can recognize the effectors and ETI comes into play. In case, the pathogen is able to secrete effectors that are not detected by the R-proteins, they cause effector triggered susceptibility (ETS) and become virulent. This sequence of events is best explained by the Zig-Zag model (Jones and Dangl 2006).
The concept of PTI and ETI is still not clear in case of viral pathogens as they can invade the cells through plasmodesmata connections. However, the viral double stranded RNAs (dsRNAs) are generally paralleled to the PAMPs as they not only trigger the RNA silencing pathways but also activate the PTI signalling. Similarly, the virus encoded RNA silencing suppression activity corresponds to the effectors that are recognized by the R proteins to activate the ETI pathways. The following sections provide a brief overview on the plant immune pathways and their overlap during viral infections has been discussed in RNA silencing shield against virus infection.
PAMP triggered immunity
It is the basal defense response or first line of defense response triggered by the recognition of special molecular patterns on the surface of microbes like viruses, bacteria and fungi. The Pathogen Associated Molecular Patterns (PAMPs) or Microbe Associated Molecular Patterns (MAMPs) are indispensable parts of pathogen (Zhang and Zhou 2010). The PAMPs/MAMPs act as elicitors of defense responses in host cells and are highly conserved across large groups of pathogens (Table 1). The elicitors may be classified as exogenous or endogenous based on their origin (Zipfel 2014). This is best exemplified by cell wall degradation products, as they may be exogenous when plants enzymes like chitinase lead to fungal cell wall degradation or may be endogenous when microbial cellulases lead to the degradation of plant cell wall (Weinberger and Friedlander 2000).
The PAMPs/MAMPs are recognized by pathogen recognition receptors (PRRs), which are normally members of the receptor like kinases (RLKs) and Receptor like protein kinases (RPKs) present on the cell membrane (Goff and Ramonell 2007). PRRs are structurally similar to toll-like receptors (TLRs) that are found in animals (Kopp and Medzhitov 2003). The binding of PAMPs/MAMPs to the PRRs triggers a signalling cascade, which results in rapid oxidative burst due to the production of ROS (Daudi and O’Brien 2012), activation of MAPKs (Schwessinger and Rathjen 2015), formation of callose (Ellinger and Voigt 2014) and expression of defense related genes like those coding for PR proteins (Zhang et al. 2019; Paludan et al. 2021).
It was shown that, binding of Flg22 (Flagellin derived 22; a PAMP) to FLS2 (a PRR) leads to phosphorylation of the kinase domain of FLS2 (Gomez-Gomez et al. 2001). The phosphorylated FLS2 dimerizes with BAK1 (Brassinosteroid insensitive 1 (BRI-1) associated kinase 1) and BIK1 (Botrytis-Induced Kinase 1) to trans-phosphorylate BIK1 (Chinchilla et al. 2007). The addition of phosphate group causes a conformational change and BIK1 is released (Wang et al. 2012). The fls2 mutant Arabidopsis plants are unable to respond to flg22 and this leads to avoidance of PTI (Gomez-Gomez and Boller 2000). In Arabidopsis BIK1 triggers the MAP kinase pathway, which leads to activation of WRKY transcription factors (Pandey and Somssich 2009). The WRKY proteins interact with W-box (TTGACC/T) motif in promoter of defense related genes (Navarro et al. 2006). BAK1 also contributes to antiviral resistance (Kørner et al. 2013) and this is discussed in RNA silencing shield against virus infection.
Effector triggered immunity
Recognition of PAMPs by PRRs puts selection pressure on pathogens to evolve strategies for escaping from host plant detection. The evolution of noneliciting PAMPs is one way by which the pathogen can overcome host resistance. This is best exemplified by Flg15, which is short version of Flg22 and is fully active in tomato while it does not act as elicitor in Arabidopsis (Robatzek et al. 2006). Another strategy involves modification of the PAMPs by post-translational modification to escape from recognition (Taguchi et al. 2009). For example the defense response is higher in response to nonglycosylated Flagellin when compared with the glycosylated one.
The pathogens also produce virulence factors called effectors to evade the PTI (Ranf 2018) and the efficiency of the effector proteins determines the pathogenicity. In bacteria, the effector proteins are produced via type 3 secretion system, from a set of avr genes. The AVR effectors can be detected by the host encoded resistance (R) proteins (Jones and Dangl 2006). Majority of R-proteins contain NB-LRR (NB- nucleotide binding at N-terminal, LRR- leucine rich repeats at C-terminal) domains (de Ronde et al. 2014a). These proteins not only provide an early warning system but also activate further immune signalling. The NB-LRR proteins are very useful for identifying host and nonhost effectors. If the products of dominant allele of avr gene and dominant allele of R gene interact then disease will not occur, due to activation of ETI, which are stronger than PTI and include genes that confer resistance against virus also.
Host resistance response
ETI response is associated with an acquired Host Resistance (HR) response which is a localised mechanism of programmed cell death (PCD) to confine or restrict the virulent pathogen in particular area (Fu and Dong 2013; Lee and Yeom 2015). Phenotypically it is manifested as necrosis and is initiated when the interaction of pathogen-AVR proteins with the host R-proteins. This interaction leads to the production of various kinds of ROS, of which H2O2 is the most stable and common type involved in cell death. The pathogen induced oxidative burst leads to oxidative mutagenesis of DNA and other cellular damage. Many biotrophic pathogens like Puccinia stritiformis try to overcome host generated ROS while the necrotrophic pathogens like Botrytis cinerea utilise ROS for their own benefits. Some pathogenic fungi use hyphal growth to conduct PCD for obtaining nutrition from dead cells while others use it as a direction cue during invasion (Singh et al. 2021).
Local pathogen infection also triggers the SAR (Systemic Acquired Resistance), which provides a general resistance to the whole plant. It is conferred by co-ordinately induced production of SA and the expression of PR proteins. SA inhibits the catalase activity and adds to the elevation of the cellular levels of H2O2. This system also serves as a mechanism to generate long lasting immune memory whose duration may vary with time and is boosted by repeated infections. Transgenic plants that produce excessive amounts of SA degrading enzyme, salicylate hydroxylase and SA mutants were defective in SAR (Fu and Dong 2013).
The SAR also results in transcriptional reprogramming through the NPR1 (Non expressor of PR genes). The overexpression of NPR1 results in broad-spectrum resistance while npr1 mutants are susceptible to infection. The NPR1 domain has 10 conserved cysteine residues, which help in binding to the SA molecule (Noctor et al. 2002; Vanacker et al. 2000). During infection the gradual elevation of SA alters the redox potential and releases the NPR1 monomers which then move into nucleus and bind with TGA transcription factors to induce PR gene expression (Fu et al. 2012). In healthy tissue, SA is conjugated and partitioned in vacuoles and NPR1 is bound by NPR4, which leads to its degradation by 26 proteasome complex.
Non host resistance response
Every plant is not affected by the same pathogen because of the one sided evolution. In this context, Non host resistance (NHR) has emerged as a common and most durable type of resistance. The use of one or more proteins (Table 2) for the detection of effectors of nonadapted pathogens leads to the induction of ETI in NHR (Lee et al. 2017). The NHR acts as a kind of outermost layer, which includes preinvasion and postinvasion responses. The preinvasion response, which can be either passive or active, restricts pathogens from entering into plant while the postinvasive response is the active defense response, which triggers HR.
RNA silencing shield against virus infection
The mechanism of RNAi (RNA interference) or RNA silencing constitutes another line of defense that is activated in response to infections by virus and some bacteria. It helps plants to defend themselves from viral infections and develop memory after reinfection by the same or closely related viruses (Sanan-Mishra et al. 2021). It can operate through post translational gene silencing (PTGS) and transcriptional gene silencing (TGS) (Liu and Chen 2016; Sinha et al. 2017; Song et al. 2019). In PTGS, foreign or aberrant dsRNA is recognized by a microprocessor complex and is converted in to 21–24 nucleotide small RNAs by the action of Dicer-like (DCL) protein (Brant and Budak 2018; Singh et al. 2019). These small RNAs get incorporated in to the RISC (RNA Induced Silencing Complex) and guide it to the target mRNAs in a sequence dependent manner, where it is cleaved by the Argonaute (AGO) protein present in this complex (Sanan-Mishra et al. 2021). In plants, the small RNAs can be further amplified by host encoded RNA dependent RNA polymerases resulting in amplification and spread of the RNA silencing response (Baulcombe 2004; Qiao et al. 2013; Sanan-Mishra et al. 2021). In TGS pathway the small RNAs are incorporated into RITS (RNA Induced Transcriptional Silencing) complex to guide the inactivation of target DNA by methylation and heterochromatinization (Muthamilarasan and Prasad 2013).
The small RNAs, frequently known as noncoding RNAs, have been broadly classified as short-interfering RNAs (siRNAs) and microRNA (miRNAs) based on their biogenesis and function (Table 3). siRNAs are produced from long dsRNAs that may arise either from endogenous sources like transposons, repetitive elements and centromere (Kumar et al. 2014; Chowdhury 2019) or exogenous sources like invading viruses or aberrant inverted repeats. These can be subclassified as repeat-associated siRNA (ra-siRNA), trans-acting siRNA (ta-siRNA), natural-antisense siRNA (nat-siRNA), hetrochromatic siRNA (hc-siRNA) and viral siRNAs (vi-siRNA). It is largely considered that siRNA formation influences resistance to virus infection, suppression of transgene expression and inactivation of transposon action (Sanan-Mishra et al. 2021). The miRNAs are produced endogenously from genome coded hair-pin or stem-loop RNA. They are present as large families and regulate almost every aspect of plant growth, development and response to stress (Waititu et al. 2020). Extensive overlap and redundancy exists between the small RNA pathways for intricately regulating the plant responses.
The role of RNA silencing during virus infection has been well elucidated and the term virus induced gene silencing (VIGS) was specifically coined to explain the phenomenon of recovery from virus infection (Van Kammen 1997). In fact viral vectors have been used to knock down expression of endogenous host genes to assess gene functions and raise virus resistant plants. Infection by viruses triggers the production of local primary small RNAs as well as systemic or transitive secondary small RNAs. In primary silencing, the viral dsRNAs are recognised and processed into siRNAs mainly by the action of DCL4. In secondary silencing, the single stranded viral transcripts are acted upon by RDR6 and SGS3 to produce dsRNA. This process is primed by the primary siRNAs and the dsRNAs thus generated are processed by DCL2 or DCL4 to produce the transitive siRNAs (Sanan-Mishra et al. 2021).
Viral dsRNA act as elicitors
The viral dsRNA, act as elicitors to trigger RNA silencing and PTI. Majority of plant viruses replicate via dsRNA intermediate, which may serve as the principal inducer of the siRNA. The secondary structure or convergent transcription of viral RNAs can also serve as a potent trigger for the RNA silencing pathway. The dsRNAs are recognized and processed into vi-siRNAs by the DCLs for cleaving and inactivating the viral transcripts (Ramesh et al. 2017; Paudel and Sanfaçon 2018; Guo et al. 2019). There are numerous reports to support that dsRNA treatment leads to the protection of plants against viral infection.
Studies using model plant Arabidopsis have shown the induction of PTI response during virus infection (Yang et al. 2010; Zorzatto et al. 2015; Niehl et al. 2016). This pathway involves the receptor kinases like BAK1/SERK3 (Somatic embryogenesis like kinase 3), BKK1/SERK4 and NIK1 (nuclear shuttle protein [NSP] interacting kinase 1), which have capability to form ligand-induced complexes with PRRs and play a central role in PTI (Brustolini et al. 2015; Niehl et al. 2016). The Arabidopsis bak1/serk3 mutants show susceptibility towards different RNA viruses (Kørner et al. 2013) and are impaired in ethylene production (Niehl et al. 2016). This response was independent of viral dsRNA recognition by DCL (Niehl et al. 2016), as it was observed to be elicited in mutants of dcl1 (Vazquez et al. 2006); dcl3 (Xie et al. 2004; Henderson et al. 2006), dcl2 and dcl4 (Deleris et al. 2006; Garcia-Ruiz et al. 2010). The PTI signalling in response to dsRNA also involves the generation of ROS (Lee et al. 2017), induction of defense hormones and the induction of defence gene expression (Zvereva et al. 2016; Nicaise and Candresse 2017) as seen in case of peptidal PAMPs.
Viral dsRNAs are also capable of triggering the MAPK cascade to activate MPK3 and MPK6. The kinases phosphorylate eIF2-alpha to trigger the formation of stress granules (stored preassembled ribosome preinitiation complex) and halt translation (Makinen et al. 2017). Infection with begomovirus was shown to trigger the activation of another SERK related RPK, called NIK1 (Nuclear shuttle protein interacting kinases 1), which leads to the suppression of translational machinery (Zorzatto et al. 2015). This phenomenon was counteracted by NSP, a virus encoded virulence factor, which binds with NIK1 to allow single strand viral DNA into cytoplasm (Zorzatto et al. 2015; Gouveia et al. 2017; Calil and Fontes 2017).
The various proteins participating in small RNA biogenesis and function serve as nodes for the crosstalk with innate immunity signals. RDR1, which is involved in production of secondary vi-siRNAs is induced by SA (Wang et al. 2010), thus implicating its role in the crosstalk. Recently it was shown that DCLs share sequence homology with dsRNA helicases of RIG1 family that act as PRRs in animal systems (Fatyol et al. 2020). It was also shown that DRB4 (dsRNA binding protein 4) interacts with R-proteins to stabilize ETI, in case of Turnip crinkle virus (TCV) infection (Zhu et al. 2013). DRB is an essential component of the RNA silencing machinery that supports the DCL mediated dicing activity. This observation indicated the existence of crosstalk between RNA silencing and R gene pathway. Recently, DRB2 was recognized as a viral invasion sensor molecule during studies on Potato Virus X (PVX) elicited systemic necrosis in Nicotiana benthamiana. (Fatyol et al. 2020). Down regulation of DRB2 rescued virus mediated induction of systemic necrosis, indicating its involvement in virus induced PTI.
It was proposed that two different types of DRBs are expended to select whether the induced immune response will initiate RNA silencing or PTI (Fatyol et al. 2020). An initial response to production of viral dsRNA activates DRB4, which is associated with the RNA silencing pathway as a default process. If RNA silencing fails or gets compromised and there is accumulation of viral dsRNA in the cell, then DRB2 is activated. This observation was supported by accumulation of dsRNA and their interaction with DRB2 in ago2 mutants. DRB2 plays an important role in switching on the PTI response and promoting systemic necrosis of infected tissue (Fatyol et al. 2020).
Virus encoded suppressors as effectors of silencing
The virus encoded RNA silencing suppressors (vRSS) serve as counter measures to the defense system of plants (Burgyán and Havelda 2011; Csorba et al. 2015), suggesting the co-evolution of host and viruses. The vRSS activity is acquired as a secondary function by ordinary viral proteins required for replication and spread like coat protein (CP), movement protein (MP) or proteases. Therefore vRSS show huge diversity in their function, structure, sequence and mode of action (Bivalkar-Mehla et al. 2011; Zhao et al. 2016; Sanan-Mishra et al. 2017). The vRSS can interfere with the host silencing pathway at multiple points to block biogenesis or function of the small RNAs (Karjee et al. 2010; Kumar et al. 2014). The common sites of action are listed in Table 4. For example V2 protein of Tomato yellow leaf curl virus (ToYLCV) binds to SGS3 (a cofactor of RDR6), which leads to the inhibition in dsRNA generation (Glick et al. 2008). Cucumber mosaic virus (CuMV) 2b protein interacts with the PAZ domain of AGO1 to inhibit the RISC activity (Duan et al. 2012) and hence small RNA function.
The vRSS synergistically influence plant responses to multiple pathogens. It has been shown that co-infection of two or more related or unrelated viruses in the same plant results in severe symptom development. The classical example is provided by synergism between PVX and the potyvirus, potato virus Y (PVY). The expression of the PVY encoded vRSS, Hc-Pro, resulted in hyper accumulation of PVX (Pruss et al. 1997). The infection with another potyvirus, Turnip mosaic virus (TuMV), encoded Hc-Pro also allowed ‘nonhost’ bacterial pathogens to grow on Arabidopsis (Navarro et al. 2008). This phenomenon mainly resulted from weakening of the host defence due to suppression of the RNA silencing pathway at multiple steps (Pruss et al. 2004). It was reported that vRSS activity of Hc-Pro of PVY and CP of TCV required the ethylene-inducible transcription factor, RAV2, which normally induces expression of defense genes (Endres et al. 2010). Hc-Pro-transgenic tobacco plants exhibited enhanced resistance to nonviral pathogens and N-mediated resistance to TMV (Nakahara et al. 2012). Another study showed that Hc-Pro interacted with the plant calmodulin-like protein, rgs-CaM and this interaction resulted in Hc-Pro degradation by autophagy. The expression of rgs-CaM is induced after infection with different RNA viruses and it interacts with several other vRSS as well (Pruss et al. 2004).
Virus infection in plants also triggered the R-Avr interactions. Several R proteins like- Rx1/2 (against PVX coat protein), HRT (against TCV coat protein), RCY1 (against CuMV coat protein), Sw-5 (against Tomato spotted wilt virus replicase protein) have been identified (Whitham et al. 1994; Cooley et al. 2000; Takahashi et al. 2001). The movement and replicase proteins of viruses are known to act as Avr factors (Moon and Park 2016). The Cauliflower mosaic virus (CaMV) transactivator (TAV) shows vRSS activity. It interacts with several host proteins that participate in TAV-mediated translation regulation, including TOR (target-of-rapamycin) kinase (Yu et al. 2003). TOR is a negative regulator of autophagy and normally it is inactivated in response to pathogen invasion to promote cell death. TOR-deficient plants are resistant to CaMV, but TAV binding activates TOR to block PCD (Yu et al. 2003).
Similarly, the CC-NB-LRR protein, CYR1 was implicated in resistance to Mungbean yellow mosaic India virus (Maiti et al. 2011). TMV and Tomato mosaic virus, encoded vRSS, p126 was recognized by the N and TM-2 proteins in tobacco and tomato plants, respectively. The activated N protein was re-localized to the nucleus to trigger PCD (Caplan et al. 2008). The vRSS are recognized by the R proteins to elicit the ETI response. The interactions between host plant and invading virus are continuously evolving. Viruses normally encode a few Avr factors and play around by altering their amino acid sequences without disturbing the structure. Thus, individual strains are able to escape from the recognition resulting in occurrence of disease (Zhu et al. 2013).
Role of host miRNAs
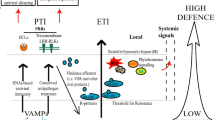
A role for host miRNAs has also been implicated in modulating the PTI and ETI pathways. The vRSS can affect the host miRNA biogenesis and function due to the high overlap and redundancy in the RNA silencing machinery. The plants effectively use this mechanism to their advantage (Fig. 1). It has been shown that repression of miR398 positively regulates PTI and callose deposition (Li et al. 2010) The NBS-LRR genes are regulated by a variety of miRNAs like miR482 binds to the P-loop-NBS-LRR transcripts leading to simultaneous silencing of R-genes (Zhai et al. 2011), while miR472 regulated the coiled-coil-NBS-LRR transcripts (Boccara et al. 2015). The down regulation of these miRNAs during virus invasion activates the NBS-LRR receptors to trigger PTI. The miRNA160a positively modulates callose deposition and it has been shown that the overexpression of miRNA160a in rice, leads to higher accumulation of H2O2 at infection site (Li et al. 2014). There are reports on the operation of similar (miRNA regulated) mechanisms from other organisms as well. The recognition of a bacterial PAMP, flagellin leads to accumulation of miR393, which represses the auxin receptor, AFB resulting in suppression of auxin signalling to enhance plant defense (Navarro et al. 2006). It was also shown that constitutive expression of AFB1 resulted in rapid infection (Navarro et al. 2006). The passenger or star strand of miR393b was shown to control exocytosis of SA-induced PR1 and to redirect the production of secondary metabolites (Zhang et al. 2011a, b). Blumera graminis invasion triggered the production of miR167, miR171, miR408, miR444 and miR138, which are involved in PTI response (Gupta et al. 2012).
Plant immunity is modulated by crosstalk between various cellular components and is regulated through key miRNA and transcription factor nodes. Different receptors (R) like PRRs, or TIR present in the cell membrane and/or cytoplasm help to perceive the biotic, abiotic and hormonal cues from the environment. Recognition of PAMPs by PRRs or RLKs leads to the activation of PTI and ETI pathways. The cellular signals converge on transcription factors (TFs), which in turn regulate the expression of genes coding for various proteins and miRNAs. The NBS-LRR RLKs are negatively regulated by miR472 and miR482. The activation of RLKs triggers a signalling cascade that leads to accumulation of miR393, which interferes with auxin production and its further crosstalk with cytokinin, abscisic acid (ABA), ethylene and Salycylic acid (SA). The hormone signals also influence expression of miRNAs like miR390, miR160, miR167 and miR159 to form complex regulatory loops to control transcription factors and gene expression. Callose deposition is an important consequence of the PTI response and it is positively regulated by miR160 and negatively regulated by miR398 and miR773
Relationship between virus and abiotic stress conditions
Plants are often exposed to multiple environmental factors like even a combination of abiotic and biotic stresses. Several studies have revealed synergism or antagonism between different stress factors indicating the existence of extensive crosstalk between plant responses. The level of virus pathogenicity is positively impacted by variation in the abiotic conditions which enhance plant susceptibility to biotic stresses (Wang et al. 2009; Prasch and Sonnewald 2013; Kissoudis et al. 2016; Sewelam et al. 2016; Kumar et al. 2020). Moreover, reports show that viruses upon infection can also improve the abiotic stress tolerance in plants (Xu et al. 2008; Gorovits et al. 2019; Sinha et al. 2021). Virus infection causes substantial physiological changes in their plant hosts such as reduced stomatal opening, lower transpiration rates and alterations in synthesis and translocation of metabolites (Keller et al. 1989; Matthews and Hull 2002). The lowering of stomatal conductance and transpiration lead to increase in leaf temperatures to minimize the water loss (Woo et al. 2008). The improved water retention seems to mimic the plant responses elicited by abiotic stresses such as drought, salinity and other osmotic stresses.
High temperature stress
High temperatures can significantly affect plant–pathogen interactions, as they are congenial for insect populations that carry with them payloads of pathogenic viruses. Moreover, under high temperatures the host defense responses like RNA silencing are suppressed, which increases plant susceptibility to pathogen infection (Travella et al. 2006; Prasch and Sonnewald 2013). The combination of high temperatures and drought significantly increased the susceptibility of Arabidopsis to TuMV by suppressing the expression of PR and R genes (Prasch and Sonnewald 2013). During Tobacco mosaic virus (TMV) infection, it was shown that under high temperatures there was conformational change in the NB-LRR (N) protein, which perceived the TMV signal. As a result the signal transduction chain that triggered PCD, to restrict virus replication and spread, could not be initiated. Heat-dependent suppression of the HR response and R-gene mediated plant defense responses were also observed during TMV infection in tobacco and Tomato Spotted Wilt Virus (TSWV) infection in tomato (Zhu et al. 2010; Prasch and Sonnewald 2013).
In many cases, abiotic stress has been reported to enhance disease resistance, like heat-induced suppression of replication of Tomato Bushy Stunt Virus (TBSV) in tobacco protoplasts (Jones and Jackson 1990). Therefore, as a counter strategy, certain viral and fungal endophytes confer heat tolerance to the host plant (Márquez et al. 2007).
Generally, pathogenic infections have been found to weaken plant tolerance to abiotic stress. For instance, tomato plants infected with ToYLCV were more susceptible to high temperatures (Anfoka et al. 2016). This increased heat susceptibility has been associated with a down regulation of HSFs and HSPs. However, the mutualistic relationship is best exemplified by high temperature stress tolerance observed in Pigeonpea plants infected with Pigeonpea sterility mosaic virus (PPSMV)-I and -II (Kumar et al. 2017). Similarly, in tomato heat stress responses were induced by ToYLCV infection (Gorovits et al. 2019). It was also reported that CuMV infection enhanced cold tolerance in B. vulgaris (Xu et al. 2008).
Drought stress
Virus infected plants show differential responses to water deficit conditions (Bergès et al. 2018). In several cases infection with viruses like CuMV, TMV, Tobacco rattle virus (TRV) or Brome mosaic virus (BMV) caused accumulation of antioxidants and osmoprotectants in host plants (Cucurbita pepo, Nicotiana tabacum and N. benthamiana, Solanum lycopersicum, Beta vulgaris) which delayed the onset of drought symptoms (Xu et al. 2008). Similar observations were reported from wheat plants infected with Barley yellow dwarf virus (BYDV) (Davis et al. 2015).
Generally, viruses induce stomatal closure that have been shown to enhance drought tolerance via Abscisic acid (ABA) or SA mediated pathways (Westwood et al. 2013). It was reported that transgenic Arabidopsis plants expressing 2b protein of CuMV showed increased drought tolerance via ABA-mediated signalling pathway (Westwood et al. 2013). The virus infection resulted in altered morphology of the roots and decreased stomatal permeability and water loss thereby stimulating drought tolerance (Westwood et al. 2013). Similarly Rice tungro spherical virus (RTSV) infected rice plants showed a delayed response to drought stress due to increased leaf hydration, less negative leaf water potential and enhanced stomatal conductance (Grimmer et al. 2012; Encabo et al. 2020). The C4 protein encoded by ToYLCV enhanced drought tolerance in Nicotiana benthamiana and tomato through an ABA-independent mechanism (Corrales‐Gutierrez et al. 2020).
Salt stress
Salinity stress constitutes one of the major abiotic stress, which creates adverse conditions for plant growth and seed germination. High salt concentration in soil causes ionic and osmotic stress resulting in cellular damage, reduction in turgor pressure and subsequently stomatal closure (Miller et al. 2010). It was shown that salinity stress favoured spread of bacterial and viral pathogens within plants to enhance infection ( Kissoudis et al. 2016; Varela et al. 2019). Cowpea plants become more susceptible to Cowpea severe mosaic virus (CpSMV) infection when exposed to salinity stress (200 mM NaCl) either simultaneously or 24 h prior to CPSMV inoculation (Varela et al. 2019). Similarly, it was reported that Nicotiana benthamiana plants subjected to osmotic, salt and wounding stresses became more susceptible to Potato virus A (Suntio and Mäkinen 2012). Studies in our lab have shown that overexpression of insect Flock House Virus B2 protein, in tobacco and rice enhances tolerance to salt stress by increasing stomatal conductance, photosynthetic efficiency and proline accumulation (Sinha et al. 2021).
Molecular mechanisms of crosstalk
Pathogenic plant viruses are parasitic symbionts as they depend on host resources for their reproduction, transmission and infection (Islam et al. 2017; Takahashi et al. 2019). During this process, they exert a strong influence on the vegetative and reproductive performance of plants by directly or indirectly stimulating the defense responses, disrupting the phytohormone accumulation and signals and altering the expression of transcription factors (Ma and Ma 2016). The main modes of crosstalk include the recognition by generic set of receptors to generate signalling cascades that culminate on transcription factors to alter gene expression. During this process the phytohormone profiles are also modulated and they respond antagonistically or synergistically to regulate response to stresses (Checker et al. 2018).
Receptors
The PRRs comprise a diverse family of cell surface-localized receptors (Frescatada-Rosa et al. 2015), including the RLKs and RPKs (Goff et al. 2007). RLKs belong to a gene superfamily, which contains 610 and 1131 members in Arabidopsis and rice, respectively (Gish and Clark 2011). They are classified into 44 subfamilies including LRRs, lectins, epidermal growth factor-like repeats, lysine motifs, self-compatibility domain and wall associated kinase (Frescatada-Rosa et al. 2015). The LRRs are known play essential roles in regulating growth and development, grain yield, hormone perception, initiating PTI responses and adaptation to abiotic stresses (Osakabe et al. 2013; Macho and Zipfel 2014; Zou et al. 2015). The RLKs contain extracellular ligand-binding domains that can perceive a wide range of PAMPs and abiotic stresses signals (Fig. 1). The ligand binding triggers phosphorylation of their intracellular serine/threonine kinase domains to activate downstream intracellular signalling cascades (Zipfel 2014; Trdá et al. 2015). In Arabidopsis a cytosolic cysteine-rich RLK, ARCK1, negatively regulates ABA and osmotic stress signal transduction (Tanaka et al. 2012). When the kinase activity is inactivated by mutation the plants show higher sensitivity to ABA and osmotic stress. CRK36, is another member of the RLK family that interacts with ARCK1 to fine-tune plant responses to abiotic stresses via hormone signalling (Tanaka et al. 2012). Studies also showed the involvement of GbRLK in response to both biotic and abiotic stresses. It mediated the reduction in rate of water loss and increased sensitivity to ABA to confer tolerance to salinity and drought stress and resistance to Verticillium wilt in transgenic cotton and Arabidopsis lines (Zhao et al. 2013; Jun et al. 2015).
Phytohormones
Phytohormones like Auxin, Cytokinins (CKs), Gibberellins (GA), ABA and Ethylene (ET) are involved in regulating various aspects of plant growth, development, reproduction and defense in response to virus infection (Adie et al. 2007; Tamaoki et al. 2013). Thus, extensive crosstalk and interactions operate between the diverse phytohormonal pathways to regulate plant physiology in response to virus attack (Pacifici et al. 2015). Role for several vRSS has been implicated in the development of disease symptoms by altering the hormone signals (Wang et al. 2012). Arabidopsis plants ectopically expressing TuMV silencing suppressor HC-Pro or Beat curly type virus (BCTV) encoded C4 protein show developmental leaf abnormalities as seen during virus infection (Kasschau et al. 2003; Mills-Lujan and Deom 2010). The vRSS, P6, encoded by CaMV when over-expressed in Arabidopsis plants resulted in stunting, chlorosis and banding of vein in leaves by interrupting multiple hormonal pathways (Rodriguez et al. 2014).
The ABA, ET, SA and JA comprise core group of defense-related phytohormones that are produced in response to both, biotic and abiotic stresses (Kissoudis et al. 2016; Schwessinger et al. 2015). Biotic stress and the resistance inducing compounds, also trigger primed state of defense. In Arabidopsis plants primed by BABA (beta-aminobutyric acid) or inoculation of an avirulent Pseudomonas syringae strain PstavrRpt2, the progeny showed higher accumulation of defense related transcripts. This also made the progeny plants more resistant to the virulent pathogens. If additional priming treatment, was provided to the already primed progenies they exhibited much stronger defense response (Slaughter et al. 2012). Chromatin immunoprecipitation analysis revealed the involvement of epigenetic regulation in this process. H3K9 acetylation of SA inducible promoters lead to permissive transcription while methylation at H3K27 of JA inducible promoters lead to repression of JA marker genes like PDF1.2. It is hypothesized that trans-generational SAR is transmitted by hypomethylated genes to direct the priming of SA dependent defenses in subsequent generations (Luna et al. 2012).
Auxin, gibberellin and cytokinin
Auxin is the most important phytohormone that regulates different aspects of plant growth and development. GA is also considered a major plant growth hormone known to stimulate stem elongation, flowering and germination. The CKs are tightly linked to auxin in promoting lateral organ development. Auxin and CK act in an antagonistic manner to regulate cell division and differentiation. Auxin supports cell division and maintenance of the apical meristem while CK promotes cell differentiation by repressing the polar auxin transport. It has been reported that SHY2 acts as a central molecule to negatively regulate Aux/IAA signalling (Calderon Villalobos et al. 2012). Its expression is induced by CK but the protein is degraded by auxin (Ioio et al. 2008). GA also participates in SHY2 regulation by repressing the CK-dependent transcriptional activation of SHY2 (Moubayidin et al. 2010).
Symptoms generated during virus infection resemble the phenotypic disorders seen in plant mutants having disrupted hormone signalling like leaf curling, stunting, mosaic and mottle or gibberellin deficiency like leaf darkening and stunting (Schaller et al. 2015). It has been shown that auxin and GA functions are directly interrupted by viral components (Jin et al. 2016). The replication proteins of TMV interact with the Aux/IAA family members to disrupt the signalling pathway, which results in development of disease symptoms such as leaf curling (Collum et al. 2016). The interference with phloem specific Aux/IAA prevents their translocation to the nucleus resulting in inhibition of the Auxin responsive TFs (ARFs). This reduces the auxin levels in the phloem and facilitates enhanced virus loading and their systemic movement (Collum et al. 2016). Similarly Rice dwarf virus (RDV) also induces symptoms like stunting in the infected rice plants, which can be recued by exogenous application of GA (Zhu et al. 2005). It was reported that the symptoms appear due to direct interaction between the RDV-P2, an outer capsid protein and the rice IAA10 and entkaurene oxidase, a key enzyme in GA biosynthesis (Hayashi et al. 2010; Satoh et al. 2011). P7-2 capsid protein encoded by Rice black streaked dwarf virus (RBSDV) interacts with GID2 (Gibberellin Insensitive Dwarf 2) to lower the levels of GA (Huang et al. 2017). GA can modulate the levels of SA and JA/ET depending upon the biotrophic or necrotrophic nature of infection (Robert-Seilaniantz et al. 2007). The CKs also work synergistically with SA to stimulate defense responses, through activation of Arabidopsis response regulators (ARRs) The ARRs can bind to promoters to induce the expression of genes, such as PR1, PR2, PR5, SID1 and SID2 which are involved in plant defense and SA biosynthesis (Choi et al. 2010). The exogenous application of CK, dihydrozeatin was shown to reduce accumulation of White clover mosaic potexvirus (WCMV).
Ethylene, salicylic acid and jasmonic acid
Ethylene (ET) is a pleiotropic hormone, which promotes fruit ripening and organ senescence. It is also involved in plant defense (Graham et al. 2011; Pieterse et al. 2012). Mutants of ET biosynthesis such as acs1, ein2 and erf106 were resistant to TMV-cg infection, but exogenous application of ACC increased the susceptibility of treated plants. TMV infection in Nicotiana tabacum, lead to the formation of lesions where ACC and/or ET accumulated. However, if ACC was sprayed before the infection, it prevented the formation of necrotic areas (de Laat and van Loon 1983: Ohtsubo et al. 1999; Alazem and Lin 2015; Lovato et al. 2019). In Phaseolus vulgaris the application of ACC reduced the virus (WCIMV) titres (Clarke et al. 1999). The ET pathways show synergistic interaction with ABA. During CMV infection, the levels of ABA and ET were increased and this resulted in suppression of hypocotyl elongation (Aharoni et al. 1977).
Salicylic acid (SA) and Jasmonic acid (JA) are majorly responsible for the plant’s reaction to biotic stresses. SA is the main phytohormone responsible for induction of SAR, which is considered as a type of long lasting immune memory. Antagonistic relationship has been observed for SA- and JA/ET-mediated defense pathways during virus infection. Exogenous application of methyl jasmonate to TMV resistant tobacco decreases resistance and facilitates systemic movement of virus (Robert-Seilaniantz et al. 2011). Depletion of endogenous SA- or disruption of SA-signalling pathway leads to the impairment of defense response and susceptibility to viral infection (Yang et al. 2015).
SA acts through NPR1 (Nonexpressor of PR genes 1), which is the key regulator of transcriptional reprogramming (a primed memory) during SAR. It has been hypothesized that in healthy tissue, when there is no accumulation of SA and NPR1 exists as an inactive oligomer bound to NPR4/NPR3 and is degraded by 26 proteasome complex. Upon infection, the gradual elevation of SA alters the cellular redox potential and this causes release of NPR1 due to reduction of its disulfide bonds. The NPR1 monomers are post-translationally modified in the presence of SA and are targeted to the nucleus, where they interact with TGA transcription factors to induce PR gene expression. The SA primed removal of NPR1 also elicits necrosis as NPR1 acts as a negative regulator of PCD (Fu.et al. 2012). The npr1 mutants are susceptible to diseases due to failed SAR while the over expression of NPR1 gives broad spectrum resistance. Accumulation of SA decreases salicylate hydrolase by the use of NahG transgene that negates resistance provided by the potato Ny-1 R-gene against Potato virus Y (Baebler et al. 2014). Viral proteins like CaMV encoded P6 protein and TMV replication protein inhibit SA-dependent defense response by causing mis-localization of the inactive form of NPR1 to the nucleus thereby disrupting the SA-signalling pathway and making the plants more susceptible to pathogens (Love et al. 2012; Wang et al. 2012). In Arabidopsis thaliana, TMV-Cg coat protein (CgCP) was shown to suppress SA-signalling pathway by stabilizing DELLA proteins that negatively regulate and represses SA-mediated defense response, thus acting antagonistically between SA and JApathways (Rodriguez et al. 2014).
The activation of PTI pathways also triggers JA mediated defense response (Shi et al. 2019). Activation of JA depends on regulation of a F-box protein, COI1 (Coronatine insensitive 1). JA acts as the mobile signal through MYC and ERF (Ethylene response factor) pathways to trigger biosynthesis of various chemicals to counter pathogen invasion (Berrocal-lobo et al. 2002; Kazan 2015). The ERF-TF branch in JA pathway is cross-influenced by ET and SA signals. It has been observed that at low concentrations the SA and JA show synergistic effects on expression of defense marker genes, but at higher concentrations their effect is antagonistic (Mur et al. 2006). The JA signals also inhibit DELLA repressor IAA59 and HY5 to interfere with the GA, auxin and light signalling pathways (Hou et al. 2016).
The JAZ–MYC complex comprises another main branch of the JA signalling pathway. In healthy tissues, when JA levels are low, JAZ is expressed which acts as a repressor of defense genes by binding to MYC2 (Pavwels et al. 2010). In presence of JA, MYC2 is released from the JAZ repressor complex to activate defense related genes and SA production. It has been shown that foliar spray of meJA reduced RBSDV infection in rice (He et al. 2020). The JAZ–MYC complex is one of the common targets of plant viruses to suppress plant defenses. The β-C1 protein produced by begomovirus satellites and ToYLCV encoded proteins interact with MYC2 (Li et al. 2019) while 2b protein of CuMV interacts with JAZ to suppress the activity of defense genes (Wu et al. 2017). The β-C1 also controls WRKY20, which are expressed in phloem to control SA induced immunity (Zhao et al. 2019a, b), as SA receptors, NPR3 and NPR4, show positive effects on JA signalling by inhibiting JAZ receptors (Liu et al. 2016).
Abscisic acid
ABA is predominantly involved in abiotic stress adaptation, but it acts as a negative regulator of pathogen resistance by inhibiting SA, JA and ethylene activated transcription of plant genes required for pathogen resistance. The ABA responses are manifested through Ca2+ gradients and MAPK cascades coupled to ROS signals to form flexible feedback loops. The first response of ABA during both biotic and abiotic stress is stomatal closure allowing reduction in water loss and maintaining beneficial water potential. As a secondary response, ABA signals induce callose deposition at the site of pathogen invasion, by inhibiting the activity of callose degrading enzyme, β-1,3 glucanase (Mauch-Mani and Mauch 2005). It has been elucidated that ABA and ROS can function in a positive amplification loop to control stomatal function and gene expression. ABA also acts by regulating the RNA silencing machinery. The dcl-11, hen1, dcl2, dcl3 and dcl4 mutants showed hypersensitivity towards ABA while aba1-5 mutants showed increased concentration of AGO1 (Li et al. 2013a, b). Moreover, ABA responsive elements (ARE) are present in their promoter region of miR168a, which targets the AGO1 (Li et al. 2013a, b).
It was shown that exogenous application of ABA or drought stress increased the susceptibility of plants to several Pseudomonas syringae and other bacterial strains (Fan et al. 2011). This effect was mediated Nine cis-Epoxycarotenoid Dioxygenase 5 (NCED5). The overexpression of NCED5 lead to a > twofold increase in ABA content, which increased the pathogen susceptibility (Jiang et al. 2010; Fan et al. 2011; Choi et al. 2013). During invasion by TMV and Bamboo mosaic virus (BaMV), ABA and SA are induced simultaneously (Alazem and Lin 2015) to increase plant resistance, but actually, ABA shows strong antagonism with SA. ABA mutants, nced3 and aba2-1 show reduced titres of BaMV and CMV (Alazem and Lin 2015).
Transcription factors
The response to biotic and abiotic stimuli involves massive transcriptional reprogramming that is intricately coordinated through the various signalling cascades that converge on the transcription factors (TFs) like MYBs, NAC, AP2 and so on. Most of these TFs also serve as critical miRNA-regulated nodes in phytohormone pathways thus facilitating cross regulations (Fig. 1).
The Auxin responsive ARFs have antiviral functions and they primarily act by targeting the viral proteins to SCF complex for proteasomal degradation (Qin et al. 2020). The vRSS like Hc-Pro, P19 and P15 deregulate miR167 to enhance suppression of ARFs (Jay et al. 2011). During PVY infection, the levels of GA are lowered to cause accumulation of miR167 to enhance suppression of ARFs (Kriznik et al. 2017).
The MYB-TF encoding botrytis susceptible 1 (bos1) gene was shown to confer resistance to pathogens and tolerance to salt, drought and oxidative stresses by regulating the JA and GA signalling pathways (Mengiste et al. 2003).
The NACs represent a large plant-specific family of TFs that play a crucial role in plant development and response to stress (Hernandez and Sanan-Mishra 2017). One of the NAC member, Arabidopsis thaliana activating factor1 (ATAF1) is a stimulus-dependent regulator of ABA biosynthesis. Its expression is induced in response to salinity and drought stress in an ABA-independent manner (Lu et al. 2007; Wu et al. 2009). ATAF1 knockout mutants have high levels of several stress responsive genes such as COR47, ERD10, KIN1, RD22 and RD29A (Lu et al. 2007; Wu et al. 2009) and pathogenesis related (PR) genes, PR-1 and PR-5 (Lu et al. 2007). Therefore, ATAF1 acts as a critical node in regulating the antagonistic interplay between ABA- and JA or ET-signaling pathways. It was also shown that ATAF1 acts as an activator of senescence-promoting TF, ORESARA1 (AtNAC092) and repressor of MYB-TF GOLDEN2-LIKE1 (Garapati et al. 2015). Similarly, OsNAC6, which shows high similarity to genes in the ATAF subfamily, was shown to be a positive regulator of plant response to biotic and abiotic stresses (Nakashima et al. 2007). TMV replication protein targets degradation of ATAF2 to regulate basal defense responses (Wang et al. 2009). Expression of OsNAC6 was induced by cold, drought, high salinity, wounding, ABA, JA and blast disease in rice (Ohnishi et al. 2005; Nakashima et al. 2007; Takasaki et al. 2010). The expression of OsNAC5 was induced under drought, cold, high salinity, ABA and methyl-JA to increase the expression of stress-responsive genes, such as OsLEA3 (Takasaki et al. 2010).
The heat shock TFs (HSFs) also play a central role in response to multiple abiotic and biotic stresses via ABA-dependent signalling pathways (Chung et al. 2013; Xue et al. 2015). Scientific evidence suggested that HSFA2 and HSFA4A are responsive to oxidative stress signals during heat and pathogen attack (Scarpeci et al. 2008). In Arabidopsis, AtHsfA6a transcript was induced under high salinity and dehydration conditions (Hwang et al. 2014). HsfA3 was associated with response to drought and salt stress (Li et al. 2013a, b), while HSF4 was induced by salinity and/or osmotic stress and fungal infection (Sham et al. 2015).
Summary and perspectives
Being sessile organisms, plants have evolved a plethora of complex overlapping response pathways, which enable them to survive a combination of environmental (biotic and abiotic) stresses. The different phytohormones are known to interact for coordinating plant growth in response to environmental changes. The plant immune system is activated by complex interactions between the pathogenic elicitor or effectors and host receptors molecules. Experimental evidence also highlights the presence of substantial overlap with the response to abiotic stresses, particularly heat, drought and salinity. Thus, exposure to biotic stress, like virus infection or pathogen attack can also give cross tolerance to abiotic stresses and vice versa. It is apparent that participation of ROS signals, MAP kinase cascades and phytohormones are common responses to both biotic and abiotic stresses. There is also recognition for the role of receptor proteins as key initiators of the crosstalk. The various signalling cascades converge on TFs which positively or negatively regulate gene expression to reprogramme the genetic machinary for increasing plant tolerance to subsequent spell of one or the other stress. The TF nodes are fine tuned by a variety of small RNAs that play a significant role in crosstalk. The cross interactions between TFs and small RNAs generate a series of regulatory nodes that enable plants to rapidly respond to diverse environmental challenges and coordinate these responses with developmental programs for endurance and survival. However, the complete understanding of crosstalk between the cellular components that facilitate plant survival in response to diverse stresses is still limited. In depth studies are required to identify the central hubs controlling the plant defense mechanisms and stress signaling interactions. The intricacies of interactions between the TFs and small RNAs involved in the regulation of stress- and hormone responsive genes need to be elucidated. The availability of high throughput sequencing, bioinformatics tools and advances in imaging technologies will support the genome-wide transcriptome and proteome studies to provide insights into the molecular components and pathways operating in response to a variety of stresses. Another challenge will be to dissect the common and specific pathways, with respect to each stress condition, which prime the plants and provide a crosslink to stress adaptations. This will help in designing effective strategies to negate the effect of stresses on plant growth and productivity.
References
Adenot X, Elmayan T, Lauressergues D, Boutet S, Bouché N, Gasciolli V, Vaucheret H (2006) DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr Biol 16(9):927–932
Adie BA, Pérez-Pérez J, Pérez-Pérez MM, Godoy M, Sánchez-Serrano JJ, Schmelz EA, Solano R (2007) ABA is an essential signal for plant resistance to pathogens affecting JA biosynthesis and the activation of defenses in Arabidopsis. Plant Cell 19(5):1665–1681
Aharoni N, Blumenfeld A, Richmond AE (1977) Hormonal activity in detached lettuce leaves as affected by leaf water content. Plant Physiol 59(6):1169–1173
Alazem M, Lin NS (2015) Roles of plant hormones in the regulation of host–virus interactions. Mol Plant Pathol 16(5):529–540
Albert I, Böhm H, Albert M, Feiler CE, Imkampe J, Wallmeroth N, Brancato C, Raaymakers TM, Oome S, Zhang H (2015) An RLP23–SOBIR1–BAK1 complex mediates NLP-triggered immunity. Nat Plants 1:15140
Allen E, Howell M (2010) MiRNAs in the biogenesis of trans-acting siRNAs in higher plants. Semin Cell Develop Biol 21:798–804
An C, Wang C, Mou Z (2017) The Arabidopsis Elongator complex is required for non host resistance against the bacterial pathogens Xanthomonas citri subsPcitri and Pseudomonas syringae pV Phaseolicola NPS3121. New Phytol 214(3):1245–1259
Anfoka G, Moshe A, Fridman L, Amrani L, Rotem O, Kolot M, Zeidan M, Czosnek H, Gorovits R (2016) Tomato yellow leaf curl virus infection mitigates the heat stress response of plants grown at high temperatures. Sci Rep 6:19715
Azevedo J, Garcia D, Pontier D, Ohnesorge S, Yu A, Garcia S, Braun L, Bergdoll M, Hakimi MA, Lagrange T, Voinnet O (2010) Argonaute quenching and global changes in Dicer homeostasis caused by a pathogen-encoded GW repeat protein. Genes Dev 24(9):904–915
Baebler Š, Witek K, Petek M, Stare K, Tušek-Žnidarič M, Pompe-Novak M, Renaut J, Szajko K, Strzelczyk-Żyta D, Marczewski W, Morgiewicz K, Gruden K, Hennig J (2014) Salicylic acid is an indispensable component of the Ny-1 resistance-gene-mediated response against Potato virus Y infection in potato. J Exp Bot 65(4):1095–1109
Baulcombe DC (2004) RNA silencing in plants. Nature 431:356–363
Bergès SE, Vile D, Vazquez-Rovere C, Blanc S, Yvon M, Bédiée A, Rolland G, Dauzat M, van Munster M (2018) Interactions between drought and plant genotype change epidemiological traits of Cauliflower mosaic virus. Front Plant Sci 9:703
Berrocal-Lobo M, Molina A, Solano R (2002) Constitutive expression of ETHYLENE-RESPONSE-FACTOR1 in Arabidopsis confers resistance to several necrotrophic fungi. Plant J 29(1):23–32
Bivalkar-Mehla S, Vakharia J, Mehla R, Abreha M, Kanwar JR, Tikoo A, Chauhan A (2011) Viral RNA silencing suppressors (RSS): novel strategy of viruses to ablate the host RNA interference (RNAi) defense system. Virus Res 155(1):1–9
Boccara M, Sarazin A, Thiébeauld O, Jay F, Voinnet O, Navarro L, Colot V (2015) The Arabidopsis miR472-RDR6 silencing pathway modulates PAMP- and effector-triggered immunity through the post-transcriptional control of disease resistance genes. PLoS Pathog 11(4):e1004814
Borsani O, Zhu J, Verslues PE, Sunkar R, Zhu JK (2005) Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 123(7):1279–1291
Bortolamiol D, Pazhouhandeh M, Marrocco K, Genschik P, Ziegler-Graff V (2007) The Polerovirus F box protein P0 targets ARGONAUTE1 to suppress RNA silencing. Curr Biol 17(18):1615–1621
Brant EJ, Budak H (2018) Plant small non-coding RNAs and their roles in biotic stresses. Front Plant Sci 9:1038
Brown GD, Taylor PR, Reid DM, Willment JA, Williams DL, Martinez-Pomares L, Wong SY, Gordon S (2002) Dectin-1 is a major beta-glucan receptor on macrophages. J Exp Med 196(3):407–412
Brustolini OJB, Machado JPB, Condori-Apfata JA, Coco D, Deguchi M, Loriato VAP, Pereira WA, Alfenas-Zerbini P, Zerbini FM, Inoue-Nagata AK, Santos AA, Chory J, Silva FF, Fontes EPB (2015) Sustained NIK-mediated antiviral signalling confers broad-spectrum tolerance to begomoviruses in cultivated plants. Plant Biotechnol J 13(9):1300–1311
Brutus A, Sicilia F, Macone A, Cervone F, De Lorenzo G (2010) A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. Proc Natl Acad Sci USA 107(20):9452–9457
Buchmann RC, Asad S, Wolf JN, Mohannath G, Bisaro DM (2009) Geminivirus AL2 and L2 proteins suppress transcriptional gene silencing and cause genome-wide reductions in cytosine methylation. J Virol 83(10):5005–5013
Burgyán J, Havelda Z (2011) Viral suppressors of RNA silencing. Trends Plant Sci 16(5):265–272
Calderón Villalobos LI, Lee S, De Oliveira C, Ivetac A, Brandt W, Armitage L, Sheard LB, Tan X, Parry G, Mao H, Zheng N, Napier R, Kepinski S, Estelle M (2012) A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nat Chem Biol 8(5):477–485
Calil IP, Fontes EPB (2017) Plant immunity against viruses: antiviral immune receptors in focus. Ann Bot 119(5):711–723
Caplan JL, Mamillapalli P, Burch-Smith TM, Czymmek K, Dinesh-Kumar SP (2008) Chloroplastic protein NRIP1 mediates innate immune receptor recognition of a viral effector. Cell 132(3):449–462
Carr JP, Lewsey MG, Palukaitis P (2010) Signaling in induced resistance. Adv Virus Res 76:57–121
Chattopadhyay C, Birah A, Jalali BL (2019) Climate change: impact on biotic stresses afflicting crop plants. In: Peshin R, Dhawan A (eds) Natural resource management: ecological perspectives. Sustainability in plant and crop protection. Springer, Charm, pp 133–146
Checker VG, Kushwaha HR, Kumari P, Yadav S (2018) Role of phytohormones in plant defense: signaling and cross talk. In: Singh A, Singh I (eds) Molecular aspects of plant–pathogen interaction. Springer, Singapore, pp 159–184
Chellappan P, Xia J, Zhou X, Gao S, Zhang X, Coutino G, Vazquez F, Zhang W, Jin H (2010) siRNAs from miRNA sites mediate DNA methylation of target genes. Nucl Acids Res 38(20):6883–6894
Chinchilla D, Zipfel C, Robatzek S, Kemmerling B, Nürnberger T, Jones JD, Felix G, Boller T (2007) A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 448:497–500
Choi SE, Kemper JK (2013) Regulation of SIRT1 by MicroRNAs. Mol Cell 36:385–392
Choi J, Huh SU, Kojima M, Sakakibara H, Paek KH, Hwang I (2010) The cytokinin-activated transcription factor ARR2 promotes plant immunity via TGA3/NPR1-dependent salicylic acid signaling in Arabidopsis. Dev Cell 19:284–295
Chowdhury FT, Shohan MUS, Islam T, Mimu TT, Palit P (2019) A therapeutic approach against leishmania donovani by predicting RNAi molecules against the surface protein, gp63. Cur Bioinforma 14:541–550
Chung E, Kim KM, Lee JH (2013) Genome-wide analysis and molecular characterization of heat shock transcription factor family in Glycine max. J Genet Genomics 40(3):127–135
Clarke SF, McKenzie MJ, Burritt DJ, Guy PL, Jameson PE (1999) Influence of white clover mosaic potexvirus infection on the endogenous cytokinin content of bean. Plant Physiol 120:547–552
Collins NC, Thordal-Christensen H, Lipka V, Bau S, Kombrink E, Qiu JL, Hückelhoven R, Stein M, Freialdenhoven A, Somerville SC, Schulze-Lefert P (2003) SNARE-protein-mediated disease resistance at the plant cell wall. Nature 425:973–977
Collum TD, Padmanabhan MS, Hsieh YC, Culver JN (2016) Tobacco mosaic virus-directed reprogramming of auxin/indole acetic acid protein transcriptional responses enhances virus phloem loading. Proc Natl Acad Sci USA 113(19):E2740-2749
Cooley MB, Pathirana S, Wu HJ, Kachroo P, Klessig DF (2000) Members of the Arabidopsis HRT/RPP8 family of resistance genes confer resistance to both viral and oomycete pathogens. Plant Cell 12(5):663–676
Corrales-Gutierrez M, Medina-Puche L, Yu Y, Wang L, Ding X, Luna AP, Bejarano ER, Castillo AG, Lozano-Duran R (2020) The C4 protein from the geminivirus Tomato yellow leaf curl virus confers drought tolerance in Arabidopsis through an ABA-independent mechanism. Plant Biotechnol J 18(5):1121–1123
Csorba T, Kontra L, Burgyán J (2015) Viral silencing suppressors: tools forged to fine-tune host-pathogen coexistence. Virology 479–480:85–103
Daudi A, O’Brien JA (2012) Detection of hydrogen peroxide by dab staining in arabidopsis Leaves. Bio Protoc 2(18):e263
Davis TS, Bosque-Perez NA, Foote NE, Magney T, Sanford DE (2015) Environmentally dependent host–pathogen and vector–pathogen interactions in the Barley yellow dwarf virus pathosystem. J Appl Ecol 52:1392–1401
Dawson WO, Hilf ME (1992) Host-Range determinants of plant viruses. Annu Rev Plant Physiol Plant Mol Biol 43:527–555
de Laat AMM, van Loon LC (1983) The relationship between stimulated ethylene production and symptom expression in virus-infected tobacco leaves. Physiol Plant Pathol 22(2):261–273
de Ronde D, Butterbach P, Kormelink R (2014a) Dominant resistance against plant viruses. Front Plant Sci 5:307
de Ronde D, Pasquier A, Ying S, Butterbach P, Lohuis D, Kormelink R (2014b) Analysis of Tomato spotted wilt virus NSs protein indicates the importance of the N-terminal domain for avirulence and RNA silencing suppression. Mol Plant Pathol 15(2):185–195
Deleris A, Gallego-Bartolome J, Bao J, Kasschau KD, Carrington JC, Voinnet O (2006) Hierarchical action and inhibition of plant Dicer-like proteins in antiviral defense. Science 313(5783):68–71
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant–pathogen interactions. Nat Rev Genet 11(8):539–548
Douchkov D, Lueck S, Hensel G, Kumlehn J, Rajaraman J, Johrde A, Doblin MS, Beahan CT, Kopischke M, Fuchs R, Lipka V, Niks RE, Bulone V, Chowdhury J, Little A, Burton RA, Bacic A, Fincher GB, Schweizer P (2016) The barley (Hordeum vulgare) cellulose synthase-like D2 gene (HvCslD2) mediates penetration resistance to host-adapted and nonhost isolates of the powdery mildew fungus. New Phytol 212(2):421–433
Du Z, Chen A, Chen W, Liao Q, Zhang H, Bao Y, Roossinck MJ, Carr JP (2014) Nuclear-cytoplasmic partitioning of cucumber mosaic virus protein 2b determines the balance between its roles as a virulence determinant and an RNA-silencing suppressor. J Virol 88(10):5228–5241
Duan CG, Fang YY, Zhou BJ, Zhao JH, Hou WN, Zhu H, Ding SW, Guo HS (2012) Suppression of Arabidopsis ARGONAUTE1-mediated slicing, transgene-induced RNA silencing, and DNA methylation by distinct domains of the Cucumber mosaic virus 2b protein. Plant Cell 24(1):259–274
Ellinger D, Voigt CA (2014) Callose biosynthesis in Arabidopsis with a focus on pathogen response: what we have learned within the last decade. Ann Bot 114(6):1349–1358
Encabo JR, Macalalad-Cabral RJA, Matres JMK, Coronejo SCTP, Jonson GB, Kishima Y, Henry A, Choi IR (2020) Infection with an asymptomatic virus in rice results in a delayed drought response. Funct Plant Biol 47(3):239–249
Endres MW, Gregory BD, Gao Z, Foreman AW, Mlotshwa S, Ge X, Pruss GJ, Ecker JR, Bowman LH, Vance V (2010) Two plant viral suppressors of silencing require the ethylene-inducible host transcription factor RAV2 to block RNA silencing. PLoS Pathog 6(1):e1000729
Fan J, Crooks C, Creissen G, Hill L, Fairhurst S, Doerner P, Lamb C (2011) Pseudomonas sax genes overcome aliphatic isothiocyanate-mediated non-host resistance in Arabidopsis. Science 331(6021):1185–1188
Fátyol K, Fekete KA, Ludman M (2020) Double-stranded-RNA-binding protein 2 participates in antiviral defense. J Virol 94(11):e00017-20
Fei Q, Xia R, Meyers BC (2013) Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell 25(7):2400–2415
Freeman BC, Beattie GA (2008) An overview of plant defenses against pathogens and herbivores. The Plant Health Instructor. https://doi.org/10.1094/PHI-I-2008-0226-01
Frescatada-Rosa M, Robatzek S, Hannah K (2015) Should I stay or should I go? Traffic control for plant pattern recognition receptors. Curr Opin Plant Biol 28:23–29
Frye CA, Tang D, Innes RW (2001) Negative regulation of defense responses in plants by a conserved MAPKK Kinase. Proc Natl Acad Sci USA 98:373–378
Fu ZQ, Dong X (2013) Systemic acquired resistance: turning local infection into global defense. Annu Rev Plant Biol 64:839–863
Fu ZQ, Yan S, Saleh A, Wang W, Ruble J, Oka N, Mohan R, Spoel SH, Tada Y, Zheng N, Dong X (2012) NPR3 and NPR4 are receptors for the immune signal salicylic acid in plants. Nature 486(7402):228–232
Gallet R, Fabre F, Thébaud G, Sofonea MT, Sicard A, Blanc S, Michalakis Y (2018) Small bottleneck size in a highly multipartite virus during a complete infection cycle. J Virol 92(14):e00139-e218
Garapati P, Xue GP, Munné-Bosch S, Balazadeh S (2015) Transcription factor ATAF1 in arabidopsis promotes senescence by direct regulation of key chloroplast maintenance and senescence transcriptional cascades. Plant Physiol 168(3):1122–1139
Garcia-Ruiz H, Takeda A, Chapman EJ, Sullivan CM, Fahlgren N, Brempelis KJ, Carrington JC (2010) Arabidopsis RNA-dependent RNA polymerases and dicer-like proteins in antiviral defense and small interfering RNA biogenesis during Turnip Mosaic Virus infection. Plant Cell 22(2):481–496
Giner A, Lakatos L, García-Chapa M, López-Moya JJ, Burgyán J (2010) Viral protein inhibits RISC activity by argonaute binding through conserved WG/GW motifs. PLoS Pathog 6(7):e1000996
Gish LA, Clark SE (2011) The RLK/Pelle family of kinases. Plant J 66(1):117–127
Glick E, Zrachya A, Levy Y, Mett A, Gidoni D, Belausov E, Citovsky V, Gafni Y (2008) Interaction with host SGS3 is required for suppression of RNA silencing by tomato yellow leaf curl virus V2 protein. Proc Natl Acad Sci USA 105(1):157–161
Goff KE, Ramonell KM (2007) The role and regulation of receptor-like kinases in plant defense. Gene Regul Syst Bio 1:167–175
Gómez-Gómez L, Boller T (2000) FLS2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol Cell 5(6):1003–1011
Gómez-Gómez L, Bauer Z, Boller T (2001) Both the extracellular leucine-rich repeat domain and the kinase activity of FLS2 are required for flagellin binding and signaling in Arabidopsis. Plant Cell 13(5):1155–1163
Gorovits R, Sobol I, Altaleb M, Czosnek H, Anfoka G (2019) Taking advantage of a pathogen: understanding how a virus alleviates plant stress response. Phytopathol Res 1:1–6
Gouveia BC, Calil IP, Machado JP, Santos AA, Fontes EP (2017) Immune receptors and co-receptors in antiviral innate immunity in plants. Front Microbiol 7:2139
Graham J, Hackett CA, Smith K, Woodhead M, MacKenzie K, Tierney I, Cooke D, Bayer M, Jennings N (2011) Towards an understanding of the nature of resistance to Phytophthora root rot in red raspberry. Theor Appl Genet 123(4):585–601
Grimmer MK, John Foulkes M, Paveley ND (2012) Foliar pathogenesis and plant water relations: a review. J Exp Bot 63:4321–4331
Guleria P, Mahajan M, Bhardwaj J, Yadav SK (2011) Plant small RNAs: biogenesis, mode of action and their roles in abiotic stresses. Genomics Proteomics Bioinformatics 9(6):183–199
Guo Z, Li Y, Ding S-W (2019) Small RNA-based antimicrobial immunity. Nat Rev Immunol 19:31–44
Gupta OP, Permar V, Koundal V, Singh UD, Parveen S (2012) MicroRNA regulated defense responses in Triticum aestivum L. during Puccinia gramminis f.sp. tritici infection. Mol Biol Rep 39:817–824
Hayashi K, Horie K, Hiwatashi Y, Kawaide H, Yamaguchi S, Hanada A, Nakashima T, Nakajima M, Mander LN, Yamane H, Hasebe M, Nozaki H (2010) Endogenous diterpenes derived from ent-kaurene, a common gibberellin precursor, regulate protonema differentiation of the moss Physcomitrella patens. Plant Physiol 153(3):1085–1097
He L, Chen X, Yang J, Zhang T, Li J, Zhang S, Zhong K, Zhang H, Chen J, Yang J (2020) Rice black-streaked dwarf virus-encoded P5–1 regulates the ubiquitination activity of SCF E3 ligases and inhibits jasmonate signaling to benefit its infection in rice. New Phytol 225(2):896–912
Hendelman A, Kravchik M, Stav R, Zik M, Lugassi N, Arazi T (2013) The developmental outcomes of P0-mediated ARGONAUTE destabilization in tomato. Planta 237(1):363–377
Henderson IR, Zhang X, Lu C, Johnson L, Meyers BC, Green PJ, Jacobsen SE (2006) Dissecting Arabidopsis thaliana DICER function in small RNA processing, gene silencing and DNA methylation patterning. Nat Genet 38(6):721–725
Hernández Y, Sanan-Mishra N (2017) miRNA mediated regulation of NAC transcription factors in plant development and environment stress response. Plant Gene 11:190–198
Hiruma K, Takano Y (2011) Roles of EDR1 in non-host resistance of Arabidopsis. Plant Signal Behav 6(11):1831–1833
Hou S, Wang X, Chen D, Yang X, Wang M, Turrà D, Di Pietro A, Zhang W (2014) The secreted peptide PIP1 amplifies immunity through receptor-like kinase 7. PLoS Pathog 10(9):e1004331
Hou X, Rivers J, León P, McQuinn RP, Pogson BJ (2016) Synthesis and function of apocarotenoid signals in plants. Trends Plant Sci 21:792–803
Huang X, Zhang X, Gong Z, Yang S, Shi Y (2017) ABI4 represses the expression of type-A ARRs to inhibit seed germination in Arabidopsis. Plant J 89:354–365
Hull R (2009) Comparative Plant Virology. Elsevier, Amsterdam
Hwang SM, Kim DW, Woo MS, Jeong HS, Son YS, Akhter S, Choi GJ, Bahk JD (2014) Functional characterization of Arabidopsis HsfA6a as a heat-shock transcription factor under high salinity and dehydration conditions. Plant Cell Environ 37:1202–1222
Ioio RD, Nakamura K, Moubayidin L, Perilli S, Taniguchi M, Morita MT, Aoyama T, Costantino P, Sabatini S (2008) A genetic framework for the control of cell division and differentiation in the root meristem. Science 322(5906):1380–1384
Ishiga Y, Uppalapati SR, Gill US, Huhman D, Tang Y, Mysore KS (2015) Transcriptomic and metabolomic analyses identify a role for chlorophyll catabolism and phytoalexin during Medicago nonhost resistance against Asian soybean rust. Sci Rep 5:13061
Islam W (2017) Management of plant virus diseases: farmer’s knowledge and our suggestions. Hosts Viruses 4:5–20
Jabs T, Tschöpe M, Colling C, Hahlbrock K, Scheel D (1997) Elicitor-stimulated ion fluxes and O2− from the oxidative burst are essential components in triggering defense gene activation and phytoalexin synthesis in parsley. Proc Natl Acad Sci USA 94(9):4800–4805
Jamous RM, Boonrod K, Fuellgrabe MW, Ali-Shtayeh MS, Krczal G, Wassenegger M (2011) The helper component-proteinase of the Zucchini yellow mosaic virus inhibits the Hua Enhancer 1 methyltransferase activity in vitro. J Gen Virol 92(Pt 9):2222–2226
Jay F, Wang Y, Yu A, Taconnat L, Pelletier S, Colot V, Renou JP, Voinnet O (2011) Misregulation of Auxin Response Factor 8 underlies the developmental abnormalities caused by three distinct viral silencing suppressors in Arabidopsis. PLoS Pathog 7(5):e1002035
Jiang CJ, Shimono M, Sugano S, Kojima M, Yazawa K, Yoshida R, Inoue H, Hayashi N, Sakakibara H, Takatsuji H (2010) Abscisic acid interacts antagonistically with salicylic acid signaling pathway in rice-Magnaporthe grisea interaction. Mol Plant Microbe Interact 23(6):791–798
Jin L, Qin Q, Wang Y, Pu Y, Liu L, Wen X, Ji S, Wu J, Wei C, Ding B, Li Y (2016) Rice dwarf virus P2 protein hijacks auxin signaling by directly targeting the rice OsIAA10 protein, enhancing viral infection and disease development. PLoS Pathog 12(9):e1005847
Johnson C, Kasprzewska A, Tennessen K, Fernandes J, Nan GL, Walbot V, Sundaresan V, Vance V, Bowman LH (2009) Clusters and superclusters of phased small RNAs in the developing inflorescence of rice. Genome Res 19(8):1429–1440
Jones JD, Dangl JL (2006) The plant immune system. Nature 444(7117):323–329
Jones RW, Jackson AO (1990) Replication of sonchus yellow net virus in infected protoplasts. Virol 179(2):815–820
Jun Z, Zhang Z, Gao Y, Zhou L, Fang L, Chen X, Ning Z, Chen T, Guo W, Zhang T (2015) Overexpression of GbRLK, a putative receptor-like kinase gene, improved cotton tolerance to Verticillium wilt. Sci Rep 5:15048
Kaku H, Nishizawa Y, Ishii-Minami N, Akimoto-Tomiyama C, Dohmae N, Takio K, Minami E, Shibuya N (2006) Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc Natl Acad Sci USA 103:11086–11091
Kang L, Li J, Zhao T, Xiao F, Tang X, Thilmony R, He S, Zhou JM (2003) Interplay of the Arabidopsis nonhost resistance gene NHO1 with bacterial virulence. Proc Natl Acad Sci USA 100:3519–3524
Karjee S, Sanan-Mishra N, Mukherjee SK (2010) Viral Suppressors of RNA Silencing in Plants. Pest Technol 4(1):1–13
Karran RA, Sanfaçon H (2014) Tomato ringspot virus coat protein binds to ARGONAUTE 1 and suppresses the translation repression of a reporter gene. Mol Plant Microbe Interact 27(9):933–943
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 4(2):205–217
Katiyar-Agarwal S, Morgan R, Dahlbeck D, Borsani O, Villegas A Jr, Zhu JK, Staskawicz BJ, Jin H (2006) A pathogen-inducible endogenous siRNA in plant immunity. Proc Natl Acad Sci USA 103(47):18002–18007
Kazan K (2015) Diverse roles of jasmonates and ethylene in abiotic stress tolerance. Trends Plant Sci 20(4):219–229
Keller P, Lüttge U, Wang X-C, Büttner G (1989) Influence of rhizomania disease on gas exchange and water relations of a susceptible and a tolerant sugar beet variety. Physiol Mol Plant Pathol 34:379–392
Kissoudis C, Sunarti S, van de Wiel C, Visser RG, van der Linden CG, Bai Y (2016) Responses to combined abiotic and biotic stress in tomato are governed by stress intensity and resistance mechanism. J Exp Bot 67(17):5119–5132
Kopp E, Medzhitov R (2003) Recognition of microbial infection by Toll-like receptors. Curr Opin Immunol 15(4):396–401
Kørner CJ, Klauser D, Niehl A, Domínguez-Ferreras A, Chinchilla D, Boller T, Heinlein M, Hann DR (2013) The immunity regulator BAK1 contributes to resistance against diverse RNA viruses. Mol Plant Microbe Interact 26(11):1271–1280
Križnik M, Petek M, Dobnik D, Ramšak Ž, Baebler Š, Pollmann S, Kreuze JF, Žel J, Gruden K (2017) Salicylic acid perturbs sRNA-gibberellin regulatory network in immune response of potato to potato virus y infection. Front Plant Sci 8:2192
Kumar V, Anand A, Mukherjee SK, Sanan-Mishra N (2014) Engineering viral suppressors of RNA silencing: requirement and applications. In: Reddy DVR, Ananda P (eds) Genetically Engineered Crops in Developing Countries. P Lava Kumar, G Lobenstein, C Kameswara Rao, Studium Press, LLC, Houston, USA, Kumar, pp 309–332
Kumar S, Tanti B, Patil BL, Mukherjee SK, Sahoo L (2017) RNAi-derived transgenic resistance to Mungbean yellow mosaic India virus in cowpea. PLoS ONE 12(10):e0186786
Kumar S, Purkyastha S, Roy C, Ranjan T, Ranjan RD (2020) Genes for different abiotic stresses tolerance in wheat. In: Hossain A (ed) Plant Stress Physiol. Intech Open
Lakatos L, Csorba T, Pantaleo V, Chapman EJ, Carrington JC, Liu YP, Dolja VV, Calvino LF, López-Moya JJ, Burgyán J (2006) Small RNA binding is a common strategy to suppress RNA silencing by several viral suppressors. EMBO J 25(12):2768–2780
Landeo-Ríos Y, Navas-Castillo J, Moriones E, Cañizares MC (2016) The p22 RNA silencing suppressor of the Crinivirus Tomato chlorosis virus is dispensable for local viral replication but important for counteracting an antiviral RDR6-mediated response during systemic infection. Viruses 8(7):182
Langenbach C, Campe R, Schaffrath U, Goellner K, Conrath U (2013) UDP-glucosyltransferase UGT84A2/BRT1 is required for Arabidopsis nonhost resistance to the Asian soybean rust pathogen Phakopsora pachyrhizi. New Phytol 198(2):536–545
Lee HA, Yeom SI (2015) Plant NB-LRR proteins: tightly regulated sensors in a complex manner. Briefings Functional Genomics 14(4):233–242
Lee HA, Lee HY, Seo E, Lee J, Kim SB, Oh S, Choi E, Choi E, Lee SE, Choi D (2017) Current understandings of plant nonhost resistance. Mol Plant Microbe Interact 30(1):5–15
Li C, Zhang B (2016) MicroRNAs in control of plant development. J Cell Physiol 231(2):303–313
Li Y, Zhang Q, Zhang J, Wu L, Qi Y, Zhou JM (2010) Identification of microRNAs involved in pathogen-associated molecular pattern-triggered plant innate immunity. Plant Physiol 152(4):2222–2231
Li Z, Zhang L, Wang A, Xu X, Li J (2013b) Ectopic overexpression of SlHsfA3, a heat stress transcription factor from tomato, confers increased thermotolerance and salt hypersensitivity in germination in transgenic Arabidopsis. PLoS ONE 8(1):e54880
Li W, Wang L, Sheng X, Yan C, Zhou R, Hang J, Yin P, Yan N (2013a) Molecular basis for the selective and ABA-independent inhibition of PP2CA by PYL13. Cell Res 23:1369–1379
Li Y, Lu YG, Shi Y, Wu L, Xu YJ, Huang F, Guo XY, Zhang Y, Fan J, Zhao JQ, Zhang HY, Xu PZ, Zhou JM, Wu XJ, Wang PR, Wang WM (2014) Multiple rice microRNAs are involved in immunity against the blast fungus Magnaporthe oryzae. Plant Physiol 164(2):1077–1092
Li N, Han X, Feng D, Yuan D, Huang LJ (2019) Signaling crosstalk between salicylic acid and ethylene/jasmonate in plant defense: do we understand what they are whispering? Int J Mol Sci 20(3):671
Lipka V, Dittgen J, Bednarek P, Bhat R, Wiermer M, Stein M, Landtag J, Brandt W, Rosahl S, Scheel D, Llorente F, Molina A, Parker J, Somerville S, Schulze-Lefert P (2005) Pre- and post invasion defenses both contribute to non host resistance in Arabidopsis. Science 310(5751):1180–1183
Liu L, Chen X (2016) RNA quality control as a key to suppressing RNA silencing of endogenous genes in plants. Mol Plant 9(6):826–836
Lovato A, Pignatti A, Vitulo N, Vandelle E, Polverari A (2019) Inhibition of virulence-related traits in Pseudomonas syringae pv. actinidiae by gunpowder green tea extracts. Frontiers Microbiol 10:2362
Love AJ, Geri C, Laird J, Carr C, Yun BW, Loake GJ, Tada Y, Sadanandom A, Milner JJ (2012) Cauliflower mosaic virus protein P6 inhibits signaling responses to salicylic acid and regulates innate immunity. PLoS ONE 7(10):e47535
Lu M, Tang X, Zhou JM (2001) Arabidopsis NHO1 is required for general resistance against Pseudomonas bacteria. Plant Cell 13:437–447
Lu PL, Chen NZ, An R, Su Z, Qi BS, Ren F, Chen J, Wang XC (2007) A novel drought-inducible gene, ATAF1, encodes a NAC family protein that negatively regulates the expression of stress-responsive genes in Arabidopsis. Plant Mol Biol 63(2):289–305
Luna E, Bruce TJ, Roberts MR, Flors V, Ton J (2012) Next-generation systemic acquired resistance. Plant Physiol 158(2):844–853
Ma KW, Ma W (2016) Phytohormone pathways as targets of pathogens to facilitate infection. Plant Mol Biol 91(6):713–725
Macho AP, Zipfel C (2014) Plant PRRs and the activation of innate immune signaling. Mol Cell 54(2):263–272
Maiti S, Basak J, Kundagrami S, Kundu A, Pal A (2011) Molecular marker-assisted genotyping of mungbean yellow mosaic India virus resistant germplasms of mungbean and urdbean. Mol Biotechnol 47(2):95–104
Mäkinen K, Lõhmus A, Pollari M (2017) Plant RNA regulatory network and RNA granules in virus infection. Frontiers Plant Sci 8:2093
Márquez LM, Redman RS, Rodriguez RJ, Roossinck MJ (2007) A virus in a fungus in a plant: three-way symbiosis required for thermal tolerance. Science 315(5811):513–515
Matthews REF, Hull R (2002) Matthews’ plant virology. Gulf professional publishing
Mauch-Mani B, Mauch F (2005) The role of abscisic acid in plant–pathogen interactions. Curr Opin Plant Biol 8(4):409–414
Mengiste T, Chen X, Salmeron J, Dietrich R (2003) The BOTRYTIS SUSCEPTIBLE1 gene encodes an R2R3MYB transcription factor protein that is required for biotic and abiotic stress responses in Arabidopsis. Plant Cell 15(11):2551–2565
Mérai Z, Kerényi Z, Kertész S, Magna M, Lakatos L, Silhavy D (2006) Double-stranded RNA binding may be a general plant RNA viral strategy to suppress RNA silencing. J Virol 80(12):5747–5756
Miller G, Suzuki N, Ciftci-Yilmaz S, Mittler R (2010) Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ 33(4):453–467
Mills-Lujan K, Deom CM (2010) Geminivirus C4 protein alters Arabidopsis development. Protoplasma 239(1–4):95–110
Molnár A, Csorba T, Lakatos L, Várallyay É, Lacomme C, Burgyán J (2005) Plant virus-derived small interfering RNAs originate predominantly from highly structured single-stranded viral RNAs plant virus-derived small interfering RNAs originate predominantly from highly structured single-stranded viral RNAs. J Virol 79:7812–7818
Moon JY, Park JM (2016) Cross-talk in viral defense signaling in plants. Frontiers Microbiol 7:2068
Moreau M, Degrave A, Vedel R, Bitton F, Patrit O, Renou JP, Barny MA, Fagard M (2012) EDS1 contributes to nonhost resistance of Arabidopsis thaliana against Erwinia amylovora. Mol Plant-Microbe Interact 25:421–430
Moreira X, Zas R, Sampedro L (2012) Differential allocation of constitutive and induced chemical defenses in pine tree juveniles: a test of the optimal defense theory. PLoS ONE 7(3):e34006
Mott GA, Thakur S, Smakowska E, Wang PW, Belkhadir Y, Desveaux D, Guttman DS (2016) Genomic screens identify a new phytobacterial microbe-associated molecular pattern and the cognate Arabidopsis receptor-like kinase that mediates its immune elicitation. Genome Biol 17:98
Moubayidin L, Perilli S, Ioio RD, Di Mambro R, Costantino P, Sabatini S (2010) The rate of cell differentiation controls the Arabidopsis root meristem growth phase. Curr Biol 20(12):1138–1143
Mur LA, Kenton P, Atzorn R, Miersch O, Wasternack C (2006) The outcomes of concentration-specific interactions between salicylate and jasmonate signaling include synergy, antagonism, and oxidative stress leading to cell death. Plant Physiol 140(1):249–262
Muthamilarasan M, Prasad M (2013) Plant innate immunity: an updated insight into defense mechanism. J Biosci 38(2):433–449
Nakahara KS, Masuta C, Yamada S, Shimura H, Kashihara Y, Wada TS, Meguro A, Goto K, Tadamura K, Sueda K, Sekiguchi T, Shao J, Itchoda N, Matsumura T, Igarashi M, Ito K, Carthew RW, Uyeda I (2012) Tobacco calmodulin-like protein provides secondary defense by binding to and directing degradation of virus RNA silencing suppressors. Proc Natl Acad Sci USA 109(25):10113–10118
Nakashima K, Tran LS, Van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi-Shinozaki K (2007) Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J 51(4):617–630
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O, Jones JD (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312(5772):436–439
Navarro L, Bari R, Achard P, Lisón P, Nemri A, Harberd NP, Jones JD (2008) DELLAs control plant immune responses by modulating the balance of jasmonic acid and salicylic acid signaling. Curr Biol 18(9):650–655
Nicaise V, Candresse T (2017) Plum pox virus capsid protein suppresses plant pathogen-associated molecular pattern (PAMP)-triggered immunity. Mol Plant Pathol 18(6):878–886
Niehl A, Wyrsch I, Boller T, Heinlein M (2016) Double-stranded RNAs induce a pattern-triggered immune signaling pathway in plants. New Phytol 211(3):1008–1019
Noctor G, Gomez L, Vanacker H, Foyer CH (2002) Interactions between biosynthesis, compartmentation and transport in the control of glutathione homeostasis and signalling. J Exp Bot 53(372):1283–1304
Nomura H, Komori T, Kobori M, Nakahira Y, Shiina T (2008) Evidence for chloroplast control of external Ca2+-induced cytosolic Ca2+ transients and stomatal closure. Plant J 53(6):988–998
Ohnishi T, Sugahara S, Yamada T, Kikuchi K, Yoshiba Y, Hirano HY, Tsutsumi N (2005) OsNAC6, a member of the NAC gene family, is induced by various stresses in rice. Genes Genet Syst 80(2):135–139
Ohtsubo N, Mitsuhara I, Koga M, Seo S, Ohashi Y (1999) Ethylene promotes the necrotic lesion formation and basic PR gene expression in TMV-infected tobacco. Plant Cell Physiol 40(8):808–817
Osakabe Y, Yamaguchi-Shinozaki K, Shinozaki K, Tran LS (2013) Sensing the environment: key roles of membrane-localized kinases in plant perception and response to abiotic stress. J Exp Bot 64:445–458
Pacifici E, Polverari L, Sabatini S (2015) Plant hormone cross-talk: the pivot of root growth. J Expt Bot 66(4):1113–1121
Paludan SR, Pradeu T, Masters SL, Mogensen TH (2021) Constitutive immune mechanisms: mediators of host defence and immune regulation. Nat Rev Immunol 21:137–150
Pandey SP, Somssich IE (2009) The role of WRKY transcription factors in plant immunity. Plant Physiol 150(4):1648–1655
Paudel DB, Sanfaçon H (2018) Exploring the diversity of mechanisms associated with plant tolerance to virus infection. Front Plant Sci 9:1575
Pauwels L, Barbero GF, Geerinck J, Tilleman S, Grunewald W, Pérez AC, Chico JM, Bossche RV, Sewell J, Gil E, García-Casado G, Witters E, Inzé D, Long JA, De Jaeger G, Solano R, Goossens A (2010) NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464(7289):788–791
Pieterse CM, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees SC (2012) Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 28:489–521
Prasch CM, Sonnewald U (2013) Simultaneous application of heat, drought, and virus to Arabidopsis plants reveals significant shifts in signaling networks. Plant Physiol 162(4):1849–1866
Pruitt RN, Schwessinger B, Joe A, Thomas N, Liu F, Albert M, Robinson MR, Chan LJ, Luu DD, Chen H, Bahar O, Daudi A, De Vleesschauwer D, Caddell D, Zhang W, Zhao X, Li X, Heazlewood JL, Ruan D, Majumder D, Chern M, Kalbacher H, Midha S, Patil PB, Sonti RV, Petzold CJ, Liu CC, Brodbelt JS, Felix G, Ronald PC (2015) The rice immune receptor XA21 recognizes a tyrosine-sulfated protein from a Gram-negative bacterium. Sci Adv 1(6):e1500245
Pruss GJ, Ge X, Shi XM, Carrington JC, Bowman Vance V (1997) Plant viral synergism: the potyviral genome encodes a broad-range pathogenicity enhancer that transactivates replication of heterologous viruses. Plant Cell 9(6):859–868
Pruss GJ, Lawrence CB, Bass T, Li QQ, Bowman LH, Vance V (2004) The potyviral suppressor of RNA silencing confers enhanced resistance to multiple pathogens. Virol 320(1):107–120
Qiao Y, Liu L, Xiong Q, Flores C, Wong J, Shi J, Wang X, Liu X, Xiang Q, Jiang S, Zhang F, Wang Y, Judelson HS, Chen X, Ma, (2013) Oomycete pathogens encode RNA silencing suppressors. Nature Genet 45:330–333
Qin Q, Li G, Jin L, Huang Y, Wang Y, Wei C, Xu Z, Yang Z, Wang H, Li Y (2020) Auxin response factors (ARFs) differentially regulate rice antiviral immune response against rice dwarf virus. PLoS Pathog 16(12):e1009118
Ramesh SV, Williams S, Kappagantu M, Mitter N, Pappu HR (2017) Transcriptome-wide identification of host genes targeted by tomato spotted wilt virus-derived small interfering RNAs. Virus Res 238:13–23
Ranf S (2018) Pattern recognition receptors—Versatile genetic tools for engineering broad-spectrum disease resistance in crops. Agronomy 8(8):134
Ranf S, Gisch N, Schäffer M, Illig T, Westphal L, Knirel YA, Sánchez-Carballo PM, Zähringer U, Hückelhoven R, Lee J (2015) A lectin S-domain receptor kinase mediates lipopolysaccharide sensing in Arabidopsis thaliana. Nat Immunol 16:426–433
Rawlings RA, Krishnan V, Walter NG (2011) Viral RNAi suppressor reversibly binds siRNA to outcompete Dicer and RISC via multiple turnover. J Mol Biol 408(2):262–276
Robatzek S, Chinchilla D, Boller T (2006) Ligand-induced endocytosis of the pattern recognition receptor FLS2 in Arabidopsis. Genes Dev 20(5):537–542
Robert-Seilaniantz A, Navarro L, Bari R, Jones JD (2007) Pathological hormone imbalances. Curr Opin Plant Biol 10:372–379
Robert-Seilaniantz A, Grant M, Jones JD (2011) Hormone crosstalk in plant disease and defense: more than just jasmonate-salicylate antagonism. Annu Rev Phytopathol 49:317–343
Rodriguez MC, Conti G, Zavallo D, Manacorda CA, Asurmendi S (2014) TMV-Cg Coat Protein stabilizes DELLA proteins and in turn negatively modulates salicylic acid-mediated defense pathway during Arabidopsis thaliana viral infection. BMC Plant Biol 14(1):1–17
Ron M, Avni A (2004) The receptor for the fungal elicitor ethylene-inducing xylanase is a member of a resistance-like gene family in tomato. Plant Cell 16(6):1604–1615
Ruiz-Ferrer V, Voinnet O (2009) Roles of plant small RNAs in biotic stress responses. Annu Rev Plant Biol 60:485–510
Sanan-Mishra N, Chakraborty S, Gupta D, Mukherjee SK (2017) RNAi suppressors: biology and mechanisms. In: Rajewsky N, Jurga S, Barciszewski J (eds) Plant epigenetics RNA technologies. Springer, Charm
Sanan-Mishra N, Jailani AAK, Mandal B, Mukherjee SK (2021) Secondary siRNAs in plants: biosynthesis, various functions and applications in virology. Front Plant Sci 12:610283
Satoh K, Shimizu T, Kondoh H, Hiraguri A, Sasaya T, Choi IR, Omura T, Kikuchi S (2011) Relationship between symptoms and gene expression induced by the infection of three strains of Rice dwarf virus. PLoS ONE 6(3):e18094
Saur IM, Kadota Y, Sklenar J, Holton NJ, Smakowska E, Belkhadir Y, Zipfel C, Rathjen JP (2016) NbCSPR underlies age-dependent immune responses to bacterial cold shock protein in Nicotiana benthamiana. Proc Natl Acad Sci USA 113(12):3389–3394
Scarpeci TE, Zanor MI, Valle EM (2008) Investigating the role of plant heat shock proteins during oxidative stress. Plant Signaling Behav 3(10):856–857
Schaller GE, Bishopp A, Kieber JJ (2015a) The yin-yang of hormones: cytokinin and auxin interactions in plant development. Plant Cell 27(1):44–63
Schwessinger B, Rathjen JP (2015) Changing SERKs and priorities during plant life. Trends Plant Sci 20(9):531–533
Sewelam N, Kazan K, Schenk PM (2016) Global plant stress signaling: reactive oxygen species at the cross-road. Front Plant Sci 7:187
Sham A, Moustafa K, Al-Ameri S, Al-Azzawi A, Iratni R, AbuQamar S (2015) Identification of Arabidopsis candidate genes in response to biotic and abiotic stresses using comparative microarrays. PLoS ONE 10(5):e0125666
Shen R, Pan S, Qi S, Lin X, Cheng S (2010) Epigenetic repression of microRNA-129-2 leads to overexpression of SOX4 in gastric cancer. Biochem Biophys Res Commun 394(4):1047–1052
Shi X, Preisser EL, Liu B, Pan H, Xiang M, Xie W, Wang S, Wu Q, Li C, Liu Y, Zhou X, Zhang Y (2019) Variation in both host defense and prior herbivory can alter plant–vector–virus interactions. BMC Plant Biol 19(1):556
Shimizu T, Nakano T, Takamizawa D, Desaki Y, Ishii-Minami N, Nishizawa Y, Minami E, Okada K, Yamane H, Kaku H, Shibuya N (2010) Two LysM receptor molecules, CEBiP and OsCERK1, cooperatively regulate chitin elicitor signaling in rice. Plant J 64:204–214
Singh A, Manjunath LE, Kundu P, Sahoo S, Das A, Suma HR, Fox PL, Eswarappa SM (2019) Let-7a-regulated translational readthrough of mammalian AGO1 generates a microRNA pathway inhibitor. EMBO J 38(16):e100727
Singh Y, Nair AM, Verma PK (2021) Surviving the odds: From perception to survival of fungal phytopathogens under host-generated oxidative burst. Plant Commun 2(3):100142
Sinha KV, Anand A, Mukherjee SK, Sanan-Mishra N (2017) RNAi based strategies for enhancing plant resistance to virus infection. In: Datta A, Fakruddin M, Iqbal HMN, Abraham J (eds) Advances in biotechnology chapter. Open Access eBooks, USA
Sinha KV, Das SS, Sanan-Mishra N (2021) Overexpression of a RNA silencing suppressor, B2 protein encoded by Flock House virus, in tobacco plants results in tolerance to salt stress. Phytoparasitica 49:299–316
Slaughter A, Daniel X, Flors V, Luna E, Hohn B, Mauch-Mani B (2012) Descendants of primed Arabidopsis plants exhibit resistance to biotic stress. Plant Physiol 158(2):835–843
Song X, Li Y, Cao X, Qi Y (2019) MicroRNAs and their regulatory roles in plant–environment interactions. Annu Rev Plant Biol 70:489–525
Stein M, Dittgen J, Sanchez-Rodriguez C, Hou BH, Molina A, Schulze-Lefert P, Lipka V, Somerville S (2006) Arabidopsis PEN3/PDR8, an ATP binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration. Plant Cell 18:731–746
Sumit R, Sahu BB, Xu M, Sandhu D, Bhattacharyya MK (2012) Arabidopsis nonhost resistance gene PSS1 confers immunity against an oomycete and a fungal pathogen but not a bacterial pathogen that cause diseases in soybean. BMC Plant Biol 12(1):1–15
Sun W, Cao Y, Jansen Labby K, Bittel P, Boller T, Bent AF (2012) Probing the Arabidopsis flagellin receptor: FLS2-FLS2 association and the contributions of specific domains to signaling function. Plant Cell 24(3):1096–1113
Sun YW, Tee CS, Ma YH, Wang G, Yao XM, Ye J (2015) Attenuation of histone methyltransferase KRYPTONITE-mediated transcriptional gene silencing by geminivirus. Sci Rep 5:16476
Suntio T, Mäkinen K (2012) Abiotic stress responses promote Potato virus A infection in Nicotiana benthamiana. Molecular Plant Pathol 13(7):775–784
Taguchi F, Suzuki T, Takeuchi K, Inagaki Y, Toyoda K, Shiraishi T, Ichinose Y (2009) Glycosylation of flagellin from Pseudomonas syringae pv. tabaci 6605 contributes to evasion of host tobacco plant surveillance system. Physiol Mol Plant Pathol 74(1):11–17
Takahashi H, Suzuki M, Natsuaki K, Shigyo T, Hino K, Teraoka T, Hosokawa D, Ehara Y (2001) Mapping the virus and host genes involved in the resistance response in Cucumber mosaic virus-infected Arabidopsis thaliana. Plant Cell Physiol 42:340–347
Takahashi H, Fukuhara T, Kitazawa H, Kormelink R (2019) Virus latency and the impact on plants. Frontiers Microbiol 10:2764
Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K, Nakashima K (2010) The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Genet Genomics 284(3):173–183
Tamaoki D, Seo S, Yamada S, Kano A, Miyamoto A, Shishido H, Miyoshi S, Taniguchi S, Akimitsu K, Gomi K (2013) Jasmonic acid and salicylic acid activate a common defense system in rice. Plant Signal Behav 8(6):e24260
Tanaka H, Osakabe Y, Katsura S, Mizuno S, Maruyama K, Kusakabe K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) Abiotic stress-inducible receptor-like kinases negatively control ABA signaling in Arabidopsis. Plant J 70(4):599–613
Thomma BP, Nürnberger T, Joosten MH (2011) Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell 23(1):4–15
Travella S, Klimm TE, Keller B (2006) RNA interference-based gene silencing as an efficient tool for functional genomics in hexaploid bread wheat. Plant Physiol 142(1):6–20
Trdá L, Boutrot F, Claverie J, Brulé D, Dorey S, Poinssot B (2015) Perception of pathogenic or beneficial bacteria and their evasion of host immunity: pattern recognition receptors in the frontline. Front Plant Sci 6:219
Trinks D, Rajeswaran R, Shivaprasad PV, Akbergenov R, Oakeley EJ, Veluthambi K, Hohn T, Pooggin MM (2005) Suppression of RNA silencing by a geminivirus nuclear protein, AC2, correlates with transactivation of host genes. J Virol 79:2517–2527
Tsuda K, Katagiri F (2010) Comparing signaling mechanisms engaged in pattern-triggered and effector-triggered immunity. Curr Opin Plant Biol 13(4):459–465
van Kammen A (1997) Virus-induced gene silencing in infected and transgenic plants. Trends Plant Sci. https://doi.org/10.1016/S1360-1385(97)01128-X
Vanacker H, Carver TL, Foyer CH (2000) Early H2O2 accumulation in mesophyll cells leads to induction of glutathione during the hyper-sensitive response in the barley-powdery mildew interaction. Plant Physiol 123:1289–1300
Várallyay E, Havelda Z (2013) Unrelated viral suppressors of RNA silencing mediate the control of ARGONAUTE1 level. Mol Plant Pathol 14(6):567–575
Várallyay É, Oláh E, Havelda Z (2014) Independent parallel functions of p19 plant viral suppressor of RNA silencing required for effective suppressor activity. Nuc Acids Res 42(1):599–608
Varela ALN, Oliveira JTA, Komatsu S, Silva RGG, Martins TF, Souza PFN, Lobo AKM, Vasconcelos IM, Carvalho FEL, Silveira JAG (2019) A resistant cowpea (Vigna unguiculata [L.] Walp.) genotype became susceptible to cowpea severe mosaic virus (CPSMV) after exposure to salt stress. J Proteomics 194:200–217
Vaucheret H (2006) Post-transcriptional small RNA pathways in plants: mechanisms and regulations. Genes Dev 20:759–771
Vázquez MA, Caballero-Pérez J, Vielle-Calzada JP (2006) A family of microRNAs present in plants and animals. Plant Cell 18(12):3355–3369
Waititu JK, Zhang C, Liu J, Wang H (2020) Plant Non-Coding RNAs: origin, biogenesis, mode of action and their roles in abiotic stress. Intl J Mol Sci 21(21):8401
Wang XQ, Yang PF, Liu Z, Liu WZ, Hu Y, Chen H, Kuang TY, Pei ZM, Shen SH, He YK (2009) Exploring the mechanism of Physcomitrella patens desiccation tolerance through a proteomic strategy. Plant Physiol 149(4):1739–1750
Wang L, Wang D, Bai Y, Huang Y, Fan M, Liu J, Li Y (2010) Spatially complex neighboring relationships among grassland plant species as an effective mechanism of defense against herbivory. Oecologia 164(1):193–200
Wang K, Senthil-Kumar M, Ryu CM, Kang L, Mysore KS (2012) Phytosterols play a key role in plant innate immunity against bacterial pathogens by regulating nutrient efflux into the apoplast. Plant Physiol 158:1789–1802
Wang L, Albert M, Einig E, Fürst U, Krust D, Felix G (2016) The pattern-recognition receptor CORE of Solanaceae detects bacterial cold-shock protein. Nat Plants 2:16185
Weinberger F, Friedlander M (2000) Endogenous and exogenous elicitors of a hypersensitive response in Gracilaria conferta (Rhodophyta). J Appl Phycology 12:139–145
Westerink N, Brandwagt BF, de Wit PJ, Joosten MH (2004) Cladosporium fulvum circumvents the second functional resistance gene homologue at the Cf-4 locus (Hcr9-4E ) by secretion of a stable avr4E isoform. Mol Microbiol 54(2):533–545
Westwood JH, McCann L, Naish M, Dixon H, Murphy AM, Stancombe MA, Bennett MH, Powell G, Webb AA, Carr JP (2013) A viral RNA silencing suppressor interferes with abscisic acid-mediated signalling and induces drought tolerance in Arabidopsis thaliana. Mol Plant Pathol 14(2):158–170
Whitham S, Dinesh-Kumar SP, Chol D, Hehl R, Corr C, Baker B (1994) The product of the tobacco mosaic virus resistance gene N: similarity to toll and the interleukin-1 receptor. Cell 78:1101–1115
Wong JEMM, Gysel K, Birkefeldt TG, Vinther M, Muszyński A, Azadi P, Laursen NS, Sullivan JT, Ronson CW, Stougaard J, Andersen KR (2020) Structural signatures in EPR3 define a unique class of plant carbohydrate receptors. Nat Commun 11(1):3797
Woo NS, Badger MR, Pogson BJ (2008) A rapid, non-invasive procedure for quantitative assessment of drought survival using chlorophyll fluorescence. Plant Methods 4:27
Wu Y, Deng Z, Lai J, Zhang Y, Yang C, Yin B, Zhao Q, Zhang L, Li Y, Yang C, Xie Q (2009) Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res 19:1279–1290
Wu L, Mao L, Qi Y (2012) Roles of dicer-like and argonaute proteins in TAS-derived small interfering RNA-triggered DNA methylation. Plant Physiol 160(2):990–999
Wu L, Chen H, Curtis C, Fu ZQ (2014) Go in for the kill: How plants deploy effector-triggered immunity to combat pathogens. Virulence 5(7):710–721
Wu D, Qi T, Li WX, Tian H, Gao H, Wang J, Ge J, Yao R, Ren C, Wang XB, Liu Y, Kang L, Ding SW, Xie D (2017) Viral effector protein manipulates host hormone signaling to attract insect vectors. Cell Res 27(3):402–415
Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, Zilberman D, Jacobsen SE, Carrington JC (2004) Genetic and functional diversification of small RNA pathways in plants. Plos Biol 2(5):E104
Xie F, Jones DC, Wang Q, Sun R, Zhang B (2015) Small RNA sequencing identifies miRNA roles in ovule and fibre development. Plant Biotechnol J 13(3):355–369
Xu P, Chen F, Mannas JP, Feldman T, Sumner LW, Roossinck MJ (2008) Virus infection improves drought tolerance. New Phytol 180:911–921
Xu J, Wang G, Wang J, Li Y, Tian L, Wang X, Guo W (2017) The lysin motif-containing proteins, Lyp1, Lyk7 and LysMe3, play important roles in chitin perception and defense against Verticillium dahliae in cotton. BMC Plant Biol 7(1):148
Xue GP, Drenth J, McIntyre CL (2015) TaHsfA6f is a transcriptional activator that regulates a suite of heat stress protection genes in wheat (Triticum aestivum L) including previously unknown Hsf targets. J Exp Bot 66:1025–1039
Yang H, Gou X, He K, Xi D, Du J, Lin H, Li J (2010) BAK1 and BKK1 in Arabidopsis thaliana confer reduced susceptibility to Turnip crinkle virus. Eur J Plant Pathol 127:149–156
Yang YX, Ahammed GJ, Wu C, Fan SY, Zhou YH (2015) Crosstalk among jasmonate, salicylate and ethylene signaling pathways in plant disease and immune responses. Curr Protein Pept ScI 16(5):450–461
Yu D, Fan B, MacFarlane SA, Chen Z (2003) Analysis of the involvement of an inducible Arabidopsis RNA-dependent RNA polymerase in antiviral defense. Mol Plant-Microbe Interact 16(3):206–216
Zhai J, Jeong DH, De Paoli E, Park S, Rosen BD, Li Y, González AJ, Yan Z, Kitto SL, Grusak MA, Jackson SA, Stacey G, Cook DR, Green PJ, Sherrier DJ, Meyers BC (2011) MicroRNAs as master regulators of the plant NB-LRR defense gene family via the production of phased, trans-acting siRNAs. Genes Dev 25(23):2540–2553
Zhang B, Wang Q (2015) MicroRNA-based biotechnology for plant improvement. J Cell Physiol 230:1–15
Zhang J, Zhou JM (2010) Plant immunity triggered by microbial molecular signatures. Mol Plant 3(5):783–793
Zhang X, Zhao H, Gao S, Wang WC, Katiyar-Agarwal S, Huang HD, Raikhel N, Jin H (2011a) Arabidopsis Argonaute 2 regulates innate immunity via miRNA393(∗)-mediated silencing of a Golgi-localized SNARE gene, MEMB12. Mol Cell 42(3):356–366
Zhang Z, Chen H, Huang X, Xia R, Zhao Q, Lai J, Teng K, Li Y, Liang L, Du Q, Zhou X, Guo H, Xie Q (2011b) BSCTV C2 attenuates the degradation of SAMDC1 to suppress DNA methylation-mediated gene silencing in Arabidopsis. Plant Cell 23(1):273–288
Zhang X, Xia J, Lii YE, Barrera-Figueroa BE, Zhou X, Gao S, Lu L, Niu D, Chen Z, Leung C, Wong T, Zhang H, Guo J, Li Y, Liu R, Liang W, Zhu JK, Zhang W, Jin H (2012) Genome-wide analysis of plant nat-siRNAs reveals insights into their distribution, biogenesis and function. Genome Biol 13(3):R20
Zhang YJ, Guo L, Gonzales PK, Gendron TF, Wu Y, Jansen-West K, O’Raw AD, Pickles SR, Prudencio M, Carlomagno Y, Gachechiladze MA, Ludwig C, Tian R, Chew J, De Ture M, Lin WL, Tong J, Daughrity LM, Yue M, Song Y, Andersen JW, Castanedes-Casey M, Kurti A, Datta A, Antognetti G, McCampbell A, Rademakers R, Oskarsson B, Dickson DW, Kampmann M, Ward ME, Fryer JD, Link CD, Shorter J, Petrucelli L (2019) Heterochromatin anomalies and double-stranded RNA accumulation underlie C9orf72 poly(PR) toxicity. Science 363(6428):eaav2606
Zhao J, Liu Q, Zhang H, Jia Q, Hong Y, Liu Y (2013) The rubisco small subunit is involved in tobamovirus movement and Tm-22-mediated extreme resistance. Plant Physiol 161(1):374–383
Zhao L, Huang Y, Hu J, Zhou H, Adeleye AS, Keller AA (2016) 1H NMR and GC–MS based metabolomics reveal defense and detoxification mechanism of cucumber plant under nano-Cu stress. Environ Sci Technol 50(4):2000–2010
Zhao T, Liu W, Zhao Z, Yang H, Bao Y, Zhang D, Wang Z, Jiang J, Xu Y, Zhang H, Li J, Chen Q, Xu X (2019b) Transcriptome profiling reveals the response process of tomato carrying Cf-19 and Cladosporium fulvum interaction. BMC Plant Biol 19(1):572
Zhao D, Yu Y, Shen Y, Liu Q, Zhao Z, Sharma R, Reiter RJ (2019a) Melatonin synthesis and function: evolutionary history in animals and plants. Front Endocrinol 10:249
Zhu S, Gao F, Cao X, Chen M, Ye G, Wei C, Li Y (2005) The rice dwarf virus P2 protein interacts with ent-kaurene oxidases in vivo, leading to reduced biosynthesis of gibberellins and rice dwarf symptoms. Plant Physiol 139(4):1935–1945
Zhu Y, Qian WQ, Hua J (2010) Temperature modulates plant defense responses through NB-LRR proteins. PLoS Pathog 6:e1000844
Zhu QH, Fan L, Liu Y, Xu H, Llewellyn D, Wilson I (2013) miR482 regulation of NBS-LRR defense genes during fungal pathogen infection in cotton. PLoS ONE 8(12):e84390
Zipfel C (2014) Plant pattern-recognition receptors. Trends Immunol 35(7):345–351
Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JD, Boller T, Felix G (2006) Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125(4):749–760
Zorzatto C, Machado JP, Lopes KV, Nascimento KJ, Pereira WA, Brustolini OJ, Reis PA, Calil IP, Deguchi M, Sachetto-Martins G, Gouveia BC, Loriato VA, Silva MA, Silva FF, Santos AA, Chory J, Fontes EP (2015) NIK1-mediated translation suppression functions as a plant antiviral immunity mechanism. Nature 520(7549):679–682
Zou X, Qin Z, Zhang C, Liu B, Liu J, Zhang C, Lin C, Li H, Zhao T (2015) Over-expression of an S-domain receptor-like kinase extracellular domain improves panicle architecture and grain yield in rice. J Exp Bot 66(22):7197–7209
Zvereva AS, Golyaev V, Turco S, Gubaeva EG, Rajeswaran R, Schepetilnikov MV, Srour O, Ryabova LA, Boller T, Pooggin MM (2016) Viral protein suppresses oxidative burst and salicylic acid-dependent autophagy and facilitates bacterial growth on virus-infected plants. New Phytol 211:1020–1034
Acknowledgements
The study was supported by research grants to N.S-M received from NSM from Department of Biotechnology (DBT), Government of India (No. BT/PR14346/AGill/103/1006/2018). S.K.S. acknowledges S.E.R.B. for support as a Distinguished Fellow. K.V.S. received fellowship from DBT. A.R. was supported by Indian science academies' summer research fellowship programme.
Author information
Authors and Affiliations
Contributions
SKS and NSM conceived the idea and designed the concept. The manuscript draft was prepared by contributions of AR and KVS. The manuscript draft was edited and proof read by SKS and NSM. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that they have no conflict of interest.
Additional information
Editorial responsibility: Aryadeep Roychoudhury
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rahman, A., Sinha, K.V., Sopory, S.K. et al. Influence of virus–host interactions on plant response to abiotic stress. Plant Cell Rep 40, 2225–2245 (2021). https://doi.org/10.1007/s00299-021-02718-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02718-0