Abstract
The plant protein ARGONAUTE1 (AGO1) functions in multiple RNA-silencing pathways, including those of microRNAs, key regulators of growth and development. Genetic analysis of ago1 mutants with informative defects has provided valuable insights into AGO1’s biological functions. Tomato encodes two AGO1 homologs (SlAGO1s), but mutants have not been described to date. To analyze SlAGO1s’ involvement in development, we confirmed that both undergo decay in the presence of the Polerovirus silencing suppressor P0 and produce a transgenic responder line (OP:P0HA) that, upon transactivation, expresses P0 C-terminally fused to a hemagglutinin (HA) tag (P0HA) and destabilizes SlAGO1s at the site of expression. By crossing OP:P0HA with a battery of driver lines, constitutive as well as organ- and stage-specific SlAGO1 downregulation was induced in the F1 progeny. Activated plants exhibited various developmental phenotypes that partially overlapped with those of Arabidopsis ago1 mutants. Plants that constitutively expressed P0HA had reduced SlAGO1 levels and increased accumulation of miRNA targets, indicating compromised SlAGO1-mediated silencing. Consistent with this, they exhibited pleiotropic morphological defects and their growth was arrested post-germination. Transactivation of P0HA in young leaf and floral organ primordia dramatically modified corresponding organ morphology, including the radialization of leaflets, petals and anthers, suggesting that SlAGO1s’ activities are required for normal lateral organ development and polarity. Overall, our results suggest that the OP:P0HA responder line can serve as a valuable tool to suppress SlAGO1 silencing pathways in tomato. The suppression of additional SlAGOs by P0HA and its contribution to the observed phenotypes awaits investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
ARGONAUTE1 (AGO1) is a key component of microRNA (miRNA) RNA-induced silencing complexes (RISCs) as well as transgene (Baumberger and Baulcombe 2005) and antiviral (Zhang et al. 2006) RISCs. AGO1 preferentially associates with small RNAs that have a 5′-terminal uridine (Mi et al. 2008), using them as guides to identify and degrade (Tang et al. 2003), or translationally inhibit (Brodersen et al. 2008; Lanet et al. 2009) highly complementary target RNAs. Because most plant miRNAs have a 5′ uridine, AGO1 is considered the most important AGO in the miRNA pathway. Indeed, in ago1 mutants, miRNA accumulation is reduced and their target mRNAs accumulate (Vaucheret et al. 2004). Arabidopsis thaliana encodes 10 AGO proteins, and mutants have been isolated or generated for all of them (Vaucheret 2008). Consistent with the major function played by AGO1 in the miRNA pathway, A. thaliana ago1 null mutants exhibit severe developmental phenotypes, including various defects in organ polarity and occasional growth arrest (Bohmert et al. 1998; Lynn et al. 1999; Kidner and Martienssen 2004, 2005). The ago1-3 and ago1-8 null mutants exhibit narrow thickened rosette leaves, which completely lose their polar identity, and radialized cauline leaves and petals (Bohmert et al. 1998; Lynn et al. 1999). The petal epidermal cell surface in ago1-8 shows abaxial characteristics (Lynn et al. 1999). In contrast, the ago1-10 null mutant exhibits strap-like or radial leaves which undergo clear adaxialization, and radial inflorescence organs (Kidner and Martienssen 2004). Growth arrest occurs in ago1-7 and ago1-8 seedlings upon replacement of the primary shoot apical meristem (SAM) with a single determinate pin-like organ, and in ago1-10 null mutant seedlings due to lack of post-embryonic organs (Lynn et al. 1999; Kidner and Martienssen 2005). In contrast to ago1 mutants, other ago null mutants exhibit no obvious (ago2-6 and ago9) or very limited (ago7/zippy and ago10/pinhead/zwille) developmental defects (Vaucheret 2008). Ler but not Col ecotype ago10 null mutants exhibit, with incomplete penetrance, a severely defective SAM (Lynn et al. 1999; Takeda et al. 2008), whereas ago7 null mutants display premature transition from juvenile to adult vegetative phase, as indicated mainly by precocious appearance of adult leaf traits (Hunter et al. 2003).
Beet western yellows virus (BWYV) viral silencing suppressor (VSR) P0 has been found to inhibit the assembly of small interfering (si) RNA/miRNA RISC effector complexes, ultimately leading to the degradation of AGO1, a key component of these complexes (Baumberger et al. 2007; Bortolamiol et al. 2007; Csorba et al. 2010). Consistent with this, transgenic P0 expression causes pleiotropic developmental phenotypes, including growth arrest as well as enhanced accumulation of several miRNA-target transcripts, mimicking the effects observed in ago1 mutants (Bortolamiol et al. 2007). Furthermore, AtAGO5, AtAGO6, AtAGO2 and AtAGO9 have also been shown to decay in the presence of P0 (Baumberger et al. 2007). Nevertheless, since A. thaliana mutants of these AGOs do not show developmental phenotypes (Vaucheret 2008), the effect of such decay on plant development is likely to be very limited. Thus, the expression of P0 in an organ offers a potent strategy to downregulate AGO1 in that organ, thereby exposing its involvement in organogenesis.
Solanum lycopersicum (tomato) is one of the most important crops in the fresh vegetable market and the food-processing industry, and offers a good model for crop species whose fruit is a fleshy berry (Giovannoni 2001). To date, tomato ago1 mutants have not been identified and the involvement of AGO1 in tomato development has not been investigated. In this study, we address the question of AGO1 functions in tomato development by inducing AGO1 decay via specific P0 expression in developing leaves and flowers. We then dissect the resulting developmental phenotypes to reveal the requirements for AGO1-mediated silencing during tomato vegetative and reproductive development.
Materials and methods
Plant material and growth conditions
Tomato (Solanum lycopersicum L.) cv. M82 (wild-type seeds kindly provided by Dr. D. Zamir, The Hebrew University of Jerusalem, Faculty of Agriculture, Rehovot, Israel) lines 35S:LhG4, FIL:LhG4, OP:GUS (Lifschitz et al. 2006), 650:LhG4, BLS:LhG4, OP:mRFP (Shani et al. 2009) and AP1:LhG4, AP3:LhG4, CRC:LhG4 (Fernandez et al. 2009) have been described elsewhere. The tomato plants were grown under greenhouse conditions with temperatures ranging between 15 and 25 °C in a mixture of tuff-peat mix with nutrients, using 400-ml pots. Germination and seedling growth took place in a growth chamber with a 16-h light period and 8-h dark period (photosynthetic photon flux density: 50–70 μmol m−2 s−1) at a constant temperature of 24 °C. For crosses, closed flowers were emasculated by removal of the petals and stamens and hand-pollinated with the pollen of an appropriate homozygous driver line.
Total RNA extraction
Total RNA was isolated from different tomato tissues with Bio-TRI RNA reagent (Bio-Lab, Jerusalem, Israel) according to the manufacturer’s protocol. After addition of isopropanol, the RNA extract was incubated overnight at −20 °C to enhance the precipitation of low-molecular-weight RNAs. Following an ethanol wash, RNA was resuspended in RNase-free water and kept at −80 °C until use.
Target validation by cleavage-site mapping
Total RNA was extracted from tomato flowers as described above. A modified procedure for RNA ligase-mediated rapid amplification of cDNA ends (5′ RACE) was performed with the GeneRacer Kit (Invitrogen, Carlsbad, CA, USA) as described previously (Talmor-Neiman et al. 2006). Briefly, cDNA was amplified with the GeneRacer-5′ primer and with SlAGO1-1_RACE/SlAGO1-2_RACE primers, followed by nested PCR using GeneRacer-5′-nested primer and SlAGO1-1_RACE_nested/SlAGO1-2_RACE_nested primers, respectively (all primer sequences are given in Supplementary Table S1). The PCR conditions used for both amplification steps were as recommended by the manufacturer. The amplified products were gel-purified, cloned into pGEMT-easy vector (Promega, Madison, WI, USA) and sequenced.
Plasmid construction
For the reporter construct P0HA [P0 C-terminally fused to a hemagglutinin (HA) tag], the coding region of BWYV P0 was PCR-amplified with primers BWYV_P0_SalI-F, which contains a SalI site at the 5′ end, and BWYV_P0_BamHI-HA-R, which contains a BamHI site, a stop codon and 27 bp encoding YPYDVPDYA HA tag. After sequence verification, the amplified fragment was cloned into the SalI/BamHI sites of the OP-TATA-BJ36 shuttle vector between an OP array (Moore et al. 1998) and Agrobacterium tumefaciens octopine synthase terminator (OCS) to generate OP-P0HA. The NotI fragment of the OP-P0HA vector was then mobilized into the binary vector pART27 (Gleave 1992) to generate pART27-OP:P0HA. For the pART27-35S:3xFLAG-SlAGO1-1 construct, the coding region of SlAGO1-1 was PCR-amplified from tomato flower cDNA with the primers FLAG_SlAGO1-1_XhoI-F, which contains a XhoI site at its 5′ end, a start codon and 24 bp encoding DYKDDDDK FLAG tag and SlAGO1-1_XhoI-R, which contains a XhoI site at its 3′ end. The amplified fragments were cloned into the XhoI site of the OP-TATA-BJ36 vector which served as a template for a second PCR with primers 3XFLAG_XhoI-F, which contains a XhoI site at its 5′ end, a start codon and 66 bp encoding DYKDHDGDYKDHDIDYKDDDDK 3XFLAG tag, and SlAGO1-1_XhoI-R. The amplified fragment was cloned into the XhoI site of the pART7 (Gleave 1992) shuttle vector to generate pART7-35S:3xFLAG-SlAGO1-1. After sequence verification, the NotI fragment of the pART7-35S:3xFLAG-SlAGO1-1 vector was mobilized into the binary vector pART27 to generate pART27-35S:3xFLAG-SlAGO1-1. A similar strategy was used to generate the pART27-35S:3xFLAG-SlAGO1-2 construct except that FLAG_SlAGO1-2_XhoI-F and SlAGO1-2_KpnI-R primers were used for the first PCR amplification and 3XFLAG_XhoI-F and SlAGO1-2_KpnI-R for the second PCR amplification.
Agroinfiltration of Nicotiana benthamiana leaves
Agrobacterium tumefaciens strain GV3101 (kindly provided by Dr. A. Gal-On, Volcani Center, Department of Plant Pathology, Bet-Dagan, Israel) cultures harboring the binary plasmids pART27-OP:P0HA, pART27-35S:LhG4, pART27-35S:3xFLAG-SlAGO1-1, and pART27-35S:3xFLAG-SlAGO1-2 were mixed as indicated to a final A600 of 0.4 and infiltrated into young leaves of 3-week-old greenhouse-grown Nicotiana benthamiana (provided by Dr. Gal-On) plants as described previously (Arazi et al. 2005). For Western blot analysis, leaf patches were collected 4 days post-infiltration.
Protein extraction and Western-blot analysis
Total protein extracts were prepared from 100 mg of 3-week-old tomato seedlings or from 100 mg N. benthamiana leaf patches that were ground in 300 μl ESB buffer (75 mM Tris–HCl pH 6.8, 9 M urea, 4.5 % v/v SDS, 7.5 % v/v β-mercaptoethanol). The mixtures were boiled for 10 min and immediately cooled on ice. The cooled homogenates were centrifuged for 10 min at 10,000g and the supernatants were transferred to a new tube, quantified using the Qubit Protein Assay Kit (Invitrogen) and equalized with ESB buffer. After equalization, the extracts were mixed with the appropriate volume of 4× SDS-PAGE loading buffer. Equal volumes of total protein extracts were then fractioned by SDS-PAGE on a 12, 8 or 6 % polyacrylamide gel for detection of P0HA, 3XFLAG-SlAGO1 and SlAGO1s, respectively. The fractionated proteins were electroblotted onto BioTrace NT membranes (Pall, Pensacola, FL, USA) and probed with commercial rabbit anti-HA (Santa-Cruz Biotechnology, Santa Cruz, CA, USA) (1:2,000) polyclonal antibodies or mouse anti-FLAG (Sigma-Aldrich, Rehovot, Israel) (1:2,000) monoclonal antibodies, or commercial rabbit anti-AtAGO1 polyclonal antibodies (Agrisera, Vannes, Sweden) (1:2,000). Antibody–protein interactions were visualized using enhanced chemiluminescence detection (ECL) kit SuperSignal west fempto (Thermo, Rockford, IL, USA) according to the manufacturer’s instructions.
Transformation of tomato plants
The binary vector pART27-OP:P0HA was transformed into tomato cv. M82 as described previously (Stav et al. 2010). Transgenic progeny were selected by germinating sterile seeds on selective medium (1× MS medium, 3 % w/v sucrose, 100 mg/l kanamycin), where only transgenic seedlings developed a branched root system. Further, validation was performed by PCR of genomic DNA with the primer pair BWYV_P0_SalI-F and BWYV_P0_BamHI-HA-R to detect the OP:P0HA transgene. The 35S:LhG4 transgene was detected by pART7-35S-F and LhG4_156-R primers.
Semi- and real-time quantitative RT-PCR analyses
Total RNA was extracted from the tested tissues as described above. Total RNA samples were treated with RNase-free DNase (Fermentas Life Sciences, Vilnius, Lithuania) to eliminate genomic DNA contamination. First-strand cDNA was synthesized from 2 to 2.4 μg of total RNA using the Superscript first-strand synthesis system for RT-PCR kit and an oligo(dT) primer (Invitrogen) following the manufacturer’s instructions. An RT-negative control was used to ensure the absence of genomic DNA template in the samples. For semi-quantitative PCR, 4 μl of 1/4 diluted cDNA was used as template to amplify P0HA and TIP41 with the primer pair BWYV_P0_SalI-F and BWYV_P0_BamHI-HA-R or qRT-TIP41-F and qRT-TIP41-R, respectively. For real-time quantitative PCR, 4 μl of diluted cDNA was used in a 10 μl PCR containing 200 nM of each primer and 5 μl Platinum SYBR Green qPCR SuperMix-UDG (Invitrogen). Primer sequences are listed in Supplementary Table S1. The relative expression levels of SlARF10, SlAP2, SlREV, SlSCL and SlSPL6 transcripts were calculated using the 2−ΔΔCt method normalized to TIP41 as a reference gene; the relative expression levels of GOBLET, SlNAC1, Solyc03g115850, and Solyc06g069710 mRNA were calculated using a two standard curve method normalized to TIP41 as a reference gene.
Scanning electron microscopy (SEM), histology, confocal microscopy and fruit analyses
The pattern of Red Fluorescent Protein (RFP) expression was detected by visualization of fresh tissue in an Olympus IX81/FV500 laser-scanning confocal microscope (Olympus Corporation, Tokyo, Japan). For the RFP signal, a 543-nm laser line was used and emission was collected with a BA560IF filter. For chlorophyll autofluorescence, a BA660IF filter was used. For SEM analysis, various tissues were collected and placed in FAA (3.7 % formaldehyde, 5 % acetic acid, 50 % EtOH, by volume) solution until use. Then, the FAA was removed and tissues were washed in an increasing gradient of ethanol (50, 70, 80, 90, 95 and 100 %). Fixed tissues were critical-point-dried, mounted on a copper plate and gold-coated. Samples were viewed in a Jeol JSM-5610 LV microscope (JEOL Ltd, Tokyo, Japan). For histological analyses, tissues were fixed in FAA until use, then dehydrated in increasing concentrations of ethanol, cleared with histoclear and embedded in paraffin. Sections cut by microtome to 10-μm thickness were placed on microscope slides, and stained with 1 % (w/v) Safranin followed by 0.2 % (w/v) Fast Green. Slides were examined under bright-field using a Leica light microscope equipped with a camera. For fruit analysis, immature green fruit at 20–25 days post-anthesis were collected from several independent plants, sliced down the middle longitudinally and scanned at 300 dpi. Fruit parameters were collected by Tomato Analyzer 3.0 software (Rodriguez et al. 2010).
GUS histochemical analysis
GUS assay was performed according to Jefferson et al. (1987). In brief, shortly after harvesting, tomato inflorescences were vacuum-infiltrated for 5 min with GUS assay buffer (1.916 mM X-Gluc, 50 mM NaPO4 buffer pH 7.0, 0.1 mM K3Fe(CN)6, 0.1 mM K4Fe(CN)6, 1 mM EDTA, 20 % methanol) and then incubated overnight at 37 °C. Tissues were washed in an increasing gradient of ethanol (50, 70, 80, 90, 95 and 100 %) for removal of GUS solution and bleaching. Samples were kept in 100 % ethanol.
Results
Tomato encodes two AGO1 homologs that are sensitive to BWYV P0
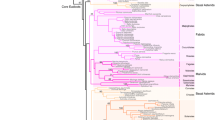
To investigate the roles of AGO1-like proteins during tomato development, we first determined which AGO1-like proteins are encoded by the tomato genome. A BLASTP query of the current version of the publicly available genome [SGN ITAG release 2.3 predicted proteins (SL2.40)] with the A. thaliana AGO1 (AtAGO1) protein sequence revealed two phylogenetically related ORFs which were 81 % identical to each other and highly similar (>90 %) to N. benthamiana AGO1-like proteins (Jones et al. 2006). Accordingly, we named them SlAGO1-1 and SlAGO1-2 (Fig. 1a). SlAGO1-1 (GenBank accession number J×945381) and SlAGO1-2 (GenBank accession number J×945382) are predicted to encode 1,054 aa (117 kDa) and 1152 aa (127 kDa) proteins, respectively. Analysis of published RNA sequencing data (Sato et al. 2012) indicated that the transcripts of both are present in vegetative as well as in reproductive organs, with SlAGO1-2 around 1.4- to 4.0-fold more abundant than SlAGO1-1. Moreover, their expression patterns were quite similar, supporting their functional redundancy (Fig. 1b). The A. thaliana AGO1 is guided to cleavage by ath-miR168 (Rhoades et al. 2002) and this negative regulation is required for AGO1 homeostasis (Vaucheret et al. 2006). Hence, we next asked whether SlAGO1-1 and SlAGO1-2 undergo miR168-guided cleavage. To identify the different members of the miR168 family in tomato, we deep-sequenced the small RNA population in 2-week post-germination M82 seedlings of the 35S:LhG4 driver line (Stav et al. 2010). This sequencing resulted in 4,847,099 redundant small RNA sequences corresponding to 1,671,156 unique sequences (Supplementary Table S2). BLASTN with known miR168 sequences against this data set and the publicly available tomato small RNA sequences (Tomato Functional Genomics Database) revealed a single conserved 21-nt miR168 sequence (sly-miR168a) (Fig. 1c), which is encoded by two independent genomic loci that can fold into a pre-miRNA-like hairpin structure (Fig. 1d). In addition, the specific sly-miR168* strand encoded by each hairpin was identified in our small RNA data set, validating their functionality as sly-miR168 precursors (Fig. 1d). To confirm that SlAGO1-1 and SlAGO1-2 are subjected to sly-miR168 cleavage in vivo, we isolated mRNAs from flowers and performed RLM-RACE to detect their 3′ cleavage products. A single 5′ RACE product of the expected size was amplified for each and the sequence of several inserts revealed that their 5′ ends all terminate at a position that pairs with the 10th sly-miR168 nucleotide, thus indicating targeting by this miRNA (Fig. 1e).
The tomato AGO1 homologs are targeted by sly-miR168. a An unrooted phylogenetic tree based on the A. thaliana AGO proteins and the tomato AGO1-like proteins (arrowheads) was constructed by the neighbor-joining method with 100 bootstrap sampling (MEGA program, version 4.0) (Tamura et al. 2007). b Accumulation of SlAGO1s in different tissues of tomato cv. Heinz based on published RNA-seq data from Sato et al. 2012, Supplementary Table S1. Data are mean ± SE of normalized expression of two independent biological samples. MG mature green, Br breaker. c Nucleotide sequence alignment of A. thaliana (ath), rice (osa), maze (zma) and tomato (sly) miR168 members. Identical nucleotides are shaded in black. d Hairpin secondary structures of sly-miR168 precursors. The position of each precursor in the tomato genome is indicated. Mature miR168 and matching miRNA* sequences are marked by red and blue circles, respectively, and the abundance of the miRNA* in our small RNA data set is indicated below. e Experimental validation of Sly-miR168 cleavage site in SlAGO1-1 and SlAGO1-2 mRNAs by RLM-RACE. Upper panel, ethidium bromide-stained agarose gel showing the 5′ RACE products. The GeneRacer 5′ primer (Adaptor) and SlAGO1 3′ primer (GSP) used for each RACE reaction are indicated above. Lower panel alignment of sly-miR68 with their target mRNAs. The arrows and numbers indicate the positions of cleavage sites inferred from 5′ RACE and fraction of sequenced clones, respectively
It has been suggested that the suppressor of silencing P0 prevents the assembly of small RNA-containing RISCs, resulting in the degradation of AGO1 (Baumberger et al. 2007; Bortolamiol et al. 2007; Csorba et al. 2010). To test the sensitivity of the tomato AGO1-like proteins to P0, plasmids expressing FLAG-tagged SlAGO1-1 and SlAGO1-2 under the 35S promoter were agroinfiltrated into N. benthamiana leaves alone or together with a responder plasmid that, upon transactivation by LhG4 (Moore et al. 1998), expresses P0HA. In addition, a driver plasmid constitutively expressing LhG4 was included in all samples (Fig. 2a). Four days post-infiltration, the expression levels of 3xFLAG-SlAGO1-1/2 and P0HA proteins were analyzed by immunoblotting. This analysis indicate that the level of each 3xFLAG-SlAGO1 protein was significantly lowered in leaf extracts that co-expressed P0HA compared to control extracts that did not express P0HA, demonstrating that SlAGO1-1 and SlAGO1-2 are sensitive to P0-mediated destabilization (Fig. 2b). As SlAGO1-1/2 and AtAGO1 transcripts are cleaved by miR168 and encode similar protein products that are destabilized in the presence of P0, it is likely that these genes are homologs.
SlAGO1-1 and SlAGO1-2 are destabilized in the presence of P0HA. a Schematic representation of the binary plasmids used in b. b N. benthamiana leaves were infiltrated with A. tumefaciens mixtures harboring the indicated pART27-35S:3xFLAG-SlAGO1 plasmid and the driver plasmid pART27-35S:LhG4 with (P0HA) or without (LhG4) bacteria harboring the P0HA responder plasmid (pART27-OP-P0HA), and protein samples were taken 4 days post-infiltration. P0HA (upper panel) and 3x-FLAG-SlAGO1 proteins (lower panel) were detected by Western blotting in 5.7 μg total protein extract probed with anti-HA and anti-FLAG commercial antibodies, respectively. 3xFLAG-SlAGO1-1 and 3xFLAG-SlAGO1-2 expression was normalized to an anti-FLAG cross-reacting band (marked by asterisk and levels are indicated at the bottom) that together with Coomassie Blue staining served as a loading control. The positions of molecular-mass standards (kDa) are indicated on the left
Constitutive expression of P0HA arrests seedling growth and increases levels of certain miRNA-targeted mRNAs
The sensitivity of both SlAGO1s to P0 suggested a potential strategy to downregulate both proteins in planta by stable expression of P0HA, thereby revealing developmental processes that depend on AGO1-mediated silencing in tomato. Nevertheless, it has been shown that strong expression of P0 arrests transgenic seedling growth (Bortolamiol et al. 2007). Thus, to stably express P0 in tomato while preventing its possible lethality during transgenic explant regeneration, the two-component OP/LhG4 transactivation system (Moore et al. 1998) was used. Seven independent transgenic tomato plants were regenerated and examined for the presence of the OP:P0HA responder transgene using PCR and genomic DNA (Supplementary Fig. S1). A homozygous responder line OP:P0HA-16, which drove strong P0HA expression upon activation, was then selected for further use (data not shown). To test the effect of constitutive expression of P0HA on tomato, the OP:P0HA-16 (henceforth be called OP:P0HA) plants were crossed (≫) with a 35S:LhG4 driver line in which the LhG4 transgene is under the control of the strong constitutive 35S promoter (Supplementary Fig. S1). Phenotypic analysis of the OP:P0HA responder line indicated that its growth and development are identical to that of the parental M82 tomato (data not shown). In contrast, all the 35S≫P0HA F1 progeny displayed pleiotropic developmental phenotypes. These included lagging germination, crooked hypocotyls and abnormal-looking succulent hairy cotyledons (Fig. 3a–f). In addition, their development was arrested at the true leaf primordium stage (Fig. 3g, h) and 4 weeks post-germination, they shriveled up and died, reminiscent of the phenotype of the A. thaliana 35S:P0 seedlings that lack a functional AGO1 due to strong expression of P0 (Bortolamiol et al. 2007). In agreement, Western blot analysis showed that the P0HA protein accumulates and a specific anti-AtAGO1 cross-reacting ~137 kDa protein band, most likely representing both SlAGO1s, decreases in the abnormal 35S≫P0HA seedlings (Fig. 3i). In A. thaliana, downregulation of AGO1 by P0 differentially modifies the levels of miRNA-target transcripts (Bortolamiol et al. 2007). Consistent with this, quantification of six known miRNA-target mRNAs in 35S≫P0HA versus control seedlings revealed significant upregulation of certain targets (SlARF10, SlSPL6, Solyc03g115850), whereas the levels of others (SlAP2, SlREV, SlSCL) were not different from controls. The pleiotropic developmental phenotypes, lethality, destabilization of SlAGO1 proteins and elevated accumulation of miRNA-target mRNAs strongly suggested that P0HA perturbs the SlAGO1-mediated silencing pathways in 35S≫P0HA seedlings.
Phenotypic and molecular characterization of 35S≫P0HA seedlings. a–f Phenotypes of representative control (35S:LhG4) and 35S≫P0HA 3-week-old seedlings. Arrowheads indicate the first true leaves. Scale bars 1 cm. g, h Control (35S:LhG4) and 35S≫P0HA shoot apices. Scale bars 500 μm. i Immunoblot analysis of P0HA and SlAGO1s in 3-week-old 35S≫P0HA seedlings. P0HA and SlAGO1s’ expression was detected with anti-HA and anti-AtAGO1 polyclonal antibodies, respectively, in equal amounts of total protein extract. Asterisk indicates an anti-HA cross-reacting band which together with Coomassie Blue staining served as a loading control. The positions of molecular-mass standards (kDa) are indicated on the left. j Quantitative RT-PCR analysis of miRNA-target transcripts in 12-day-old seedlings of the indicated genotypes. Primers were designed around the corresponding miRNA’s complementary site. TIP41 expression values were used for normalization. Data are mean ± SE of two biological replicates, each measured in triplicate. Asterisks indicate statistically significant difference as determined by Student’s t test (P ≤ 0.05)
Stage-specific expression of P0HA differentially affects compound leaf development
The severe growth arrest phenotype of the 35S≫P0HA seedlings hampered further analysis of the effects of SlAGO1 downregulation on tomato development beyond germination (Fig. 3). Nevertheless, several leaf- and flower-specific driver lines (promoter:LhG4), which provide different temporal and/or spatial specificities, have been previously characterized (Alvarez et al. 2006; Fernandez et al. 2009; Shalit et al. 2009; Shani et al. 2009). Therefore, the requirement of AGO1-mediated silencing for the different phases of compound leaf development was studied by crossing the OP:P0HA responder line to FIL:LhG4, BLS:LhG4 and 650:LhG4 driver lines, which have been shown to drive comparable expression in different developmental windows and at distinct locations of the leaf primordium (Shani et al. 2009). The FIL (FILAMENTOUS FLOWER) promoter drives expression throughout young leaf primordia soon after they initiate from the SAM, but not in the SAM (Lifschitz et al. 2006; Shani et al. 2009). Both BLS and 650 promoters initiate expression later than FIL (starting from the P4 stage) and drive comparable expression in initiating leaflets, and distal and abaxial domains of the P5 primordia, respectively (Shani et al. 2009). Upon germination, the effect of P0HA expression was notable on FIL≫P0HA and 650≫P0HA cotyledons. The FIL≫P0HA cotyledons were slightly epinastic, and the 650≫P0HA cotyledons were reddish, particularly along the mid vein and on the abaxial side (Supplementary Fig. S2). No obvious phenotypes were observed among BLS≫P0HA cotyledons. In comparison to control plants, FIL≫P0HA plants produced abnormal compound leaves with irregularly arranged leaflets along the rachis that had strap- or filament-like shapes (Fig. 4a), reminiscent of the tentacle-like bladeless leaves of A. thaliana ago1-3 (Bohmert et al. 1998). This contrasts with BLS≫P0HA and 650≫P0HA leaves which showed only slight deviation from the wild-type phenotype. The BLS≫P0HA leaflet blades had a wrinkled instead of smooth surface with patches of dark and pale green, and the 650≫P0HA leaf vasculature had a clear reddish color (Fig. 4a).
Phenotypes of control (FIL:LhG4) and indicated genotype leaves. a Representative fully expanded 3rd leaf. Scale bar 2 cm. b Comparison of representative P2–P6 stages of early leaf development between control and FIL≫P0HA plants. Note the ectopic outgrowth on the abaxial side of the leaflets (arrowhead). The terminal and primary leaflets are indicated by white and black asterisks, respectively. Scale bars 200 μm. c An emerging 3rd leaf’s terminal leaflet blade epidermal patterns. Upper panel global view of the adaxial blade surface around the midvein, scale bars 1 mm. Lower panel magnified views of the abaxial and adaxial blade surfaces around the midvein, scale bars 25 μm. d Light micrographs of transverse sections of fully expanded 3rd leaf terminal leaflet of the indicated genotypes. V vascular bundle, B blade. Scale bars 100 μm
To determine the basis for the FIL≫P0HA leaf phenotype, we first compared the early development of control and FIL≫P0HA leaves. As shown in Fig. 4b, starting from the P2 to P3 stage primordia, a pronounced alteration in FIL≫P0HA primordium morphology was observed, being smaller than the control primordia. At the P4 stage, initiating primary leaflets showed changeable growth orientations, uncovering the reason for the irregular leaflet arrangement observed in mature FIL≫P0HA leaves. At the P6 stage, the primary leaflets were abnormally flattened and developed cylinder-like appendages on their abaxial side that later grew into tentacle-like filaments (Fig. 4b). Quantitative RT-PCR analysis of FIL≫P0HA apices demonstrated a significant increase in selected miRNA-target transcripts, an indication for SlAGO1s downregulation (Supplementary Fig. S3). On the other hand, in young BLS≫P0HA and 650≫P0HA leaves, which initiate P0HA expression later than FIL≫P0HA, no deviating phenotypes were observed in the P1–P5 stage leaf primordia (data not shown). The adaxial–abaxial polarity in the FIL≫P0HA and BLS≫P0HA leaflets was also investigated. The adaxial epidermis of the wild-type smooth terminal leaflet blade (here represented by FIL:LhG4) was characterized by relatively uniform pavement cells, and the abaxial epidermis was characterized by diverse size pavement cells interspersed with stomatal cells (Fig. 4c). The BLS≫P0HA terminal leaflets had a wrinkly instead of smooth blade that was covered with adaxial and abaxial epidermal pavement cells that differed slightly from the wild type (Fig. 4c). In contrast, the epidermal surface of the FIL≫P0HA cylindrical leaflets was not composed of either adaxial or abaxial pavement cells but instead of long rectangular epidermal cells that could not be characterized as adaxial or abaxial (Fig. 4c). A similar epidermal cell phenotype has been previously observed in A. thaliana ago1-3 null mutant cotyledons showing reduced adaxial/abaxial differentiation (Bohmert et al. 1998). Indeed, transverse sections through the FIL≫P0HA leaflets revealed an almost radial structure (Fig. 4d). These leaflets were composed of abnormal vasculature surrounded by uniform parenchyma, mesophyll and epidermal cells, indicating loss of adaxial/abaxial identity, most probably due to SlAGO1 downregulation by P0HA.
Flower-specific expression of P0HA disturbs organogenesis and induces radialization of petals and anthers
Wild-type M82 tomato flowers are composed of four distinct whorls. The perianth contains five or six sepals alternating with five or six yellow petals. The two inner reproductive whorls contain five or six yellow stamens forming a cone, which encloses two fused carpels that develop a multilocular ovary and a protruding style and stigma. To test the effect of suppression of AGO1-mediated silencing on flower development, OP:P0HA plants were crossed with AP1:LhG4, AP3:LhG4 and CRC:LhG4 driver lines that express LhG4 through the well-characterized A. thaliana APETALA1 (AP1), APETALA3 (AP3) and CRABSCLAW (CRC) flower-specific promoters (Jack et al. 1992; Mandel et al. 1992; Alvarez and Smyth 1999; Bowman and Smyth 1999). The detailed expression patterns directed by these promoters in developing tomato flowers were studied by crosses with OP:mRFP (Shani et al. 2009) and OP:GUS (Lifschitz et al. 2006) reporter lines. As shown in Fig. 5, the expression domains of all promoters were quite similar to their known expression in the A. thaliana flower (Jack et al. 1992; Mandel et al. 1992; Bowman and Smyth 1999). The AP1 promoter was initially expressed throughout young floral primordia (Fig. 5b, c), in accordance with its function as a floral meristem identity gene (Mandel et al. 1992). As the buds developed, mRFP expression directed by AP1 was mainly confined to the developing sepals and petals (Fig. 5g, k), in accordance with its function in specifying the identity of these organs (Mandel et al. 1992). In addition, weak mRFP signal in the receptacle and distal part of the second and third whorls was detected. In young floral primordia, the more restricted AP3 promoter showed specific mRFP expression in the distal part of developing sepals (Fig. 5d). In more developed buds, the mRFP signal was detected on the adaxial side of sepals, throughout the developing petals (Fig. 5h), in the stamen vasculature and on the abaxial side of anthers (Fig. 5l, m). The mRFP expression directed by the CRC promoter was limited to the developing carpelloid organs just after the emergence of the carpel primordia (Fig. 5e). At anthesis, the mRFP signal was detected along the style (Fig. 5i) and was restricted to the exocarp (Fig. 5n), slowly declining as the fruit reached the mature green stage (Fernandez et al. 2009). These analyses indicated that the AP1:LhG4, AP3:LhG4 and CRC:LhG4 driver lines can direct specific expression in all four whorls of the tomato flower.
Characterization of the expression directed by flower-specific driver lines. a–m Confocal microscope images of developing tomato flowers of controls (OP:mRFP) and the indicated genotypes. a–d Stage 1–3 buds. Stages are according to Xiao et al. (2009). b Whole-mount GUS staining of AP1≫GUS inflorescence meristem (marked by an arrow) and stage 2 bud. e–h Longitudinal sections of stage 5 buds. i The upper part of the style at anthesis. j, n Longitudinal sections of ovary at anthesis. k Longitudinal section of stage 8 bud. Transverse (l) and longitudinal (m) sections of anther at anthesis. The red and blue colors represent mRFP and chloroplast fluorescence signals, respectively. Arrowheads indicate the position of the petal primordia and asterisks represents the position of the anther primordia. S sepal, C carpel, O ovary, A anther, P petal, ST stigma, VB vascular bundle, AL anther locule, OV ovule, PG pollen grains, PC pericarp. All scale bars 200 μm
The AP1≫P0HA, AP3≫P0HA and CRC≫P0HA plants showed a wild-type phenotype during vegetative growth (data not shown). However, they displayed dramatic floral phenotypes within the expression domains of each promoter. Expressing P0HA under control of the AP1 promoter resulted in the development of smaller flowers with deformed organs in all four whorls, in agreement with AP1 expression in young floral primordia (Fig. 6). The sepals were more succulent and frequently failed to enclose the developing flower (Fig. 6). In the second whorl, most of the flowers produced smaller needle-like petals. The third whorl contained deformed filament-like anthers, some of which were fused (Fig. 6). The fourth whorl contained an abnormal pistil composed of an enlarged ovule-less ovary and numerous styles either tipped or not tipped with stigmatic tissue (Fig. 6). The defects in the reproductive whorls of AP1≫P0HA flowers led to sterility. Notwithstanding, very rarely, parthenocarpic fruit did develop (Supplementary Fig. S4). Consistent with the AP3 expression pattern (Fig. 5), expressing P0HA using this promoter resulted in abnormal phenotypes only in the second and third whorls, while the sepals showed an almost normal phenotype and the pistils developed normally, like the wild-type controls (Fig. 6). The AP3≫P0HA petals were filament-like, but unlike the AP1≫P0HA petals, their proximal region was flat and encircled the reproductive whorls. The AP3≫P0HA stamens were deformed and occasionally fused as in the AP1≫P0HA flowers (Fig. 6). Moreover, these plants set fruit that contained viable seeds (Supplementary Fig. S4). The filament-like phenotypes of petals and anthers suggested that they suffer from an alteration in organ polarity. Histological analyses of AP1≫P0HA and AP3≫P0HA stage 6 buds revealed radial petals compared to the control and an excessive number of abnormal anthers, some of which were fused, radial or both (Fig. 7a–c). Anther radialization was more frequent in AP3≫P0HA buds, which is in agreement with the expression of AP3 promoter in anthers (Fig. 5). SEM examination of AP1≫P0HA and AP3≫P0HA stage 6 buds supported the above observations, showing filament-like petals and stamens (Fig. 7d–f). In the wild-type tomato flower at anthesis, the petal’s adaxial epidermis is composed of cylinder-shaped cells while the abaxial epidermis consists of pavement cells (Fig. 7g). In contrast, the surfaces of the AP1≫P0HA and AP3≫P0HA filamentous petals were composed of elongated rectangular epidermal cells with no recognizable abaxial or adaxial identity (Fig. 7h, i). A similar loss of cellular identity and phenotype was observed in the epidermal cells on the outer surface of the FIL≫P0HA needle-like leaflets (Fig. 4c), suggesting that P0HA expression during petal growth, as in leaflets, disturbed the establishment of proper adaxial–abaxial identity. Quantitative RT-PCR analysis of AP1≫P0HA buds demonstrated a significant increase in certain miRNA-target transcripts, validating the perturbation of the miRNA pathway by expressed P0HA (Supplementary Fig. S3). Expression of P0HA under the control of the carpel-specific CRC promoter resulted in the development of conic instead of round ovaries (Fig. 8a), with no obvious effect on the more outer flower whorls (data not shown). Accordingly, a change in the shape of developed CRC≫P0HA immature young fruits was observed. These fruits were significantly more triangular at their distal part than the round control fruits (Fig. 8a, b). As the fruits matured, the differences between CRC≫P0HA fruits and controls become less obvious, most likely due to the diminished expression of P0HA following anthesis (Fig. 8c).
Phenotypes of AP1≫P0HA and AP3≫P0HA flowers. Abnormalities caused by flower-specific P0HA expression. The upper panel presents the whole flower, the middle panel presents whorls two and three, and the lower panel presents the pistil. P petal, A anther, O ovary, ST style. Stigma-less styles are indicated by arrowheads. Scale bars 1 mm
Histological and microscopic analyses of AP1≫P0HA and AP3≫P0HA flowers. a–c Transverse sections of 2 mm buds of controls (AP1:LhG4) and the indicated genotypes. Insets show representative petals at a higher magnification, scale bars 100 μm. d–f Scanning electron micrographs of 2-mm buds. The sepals were removed to expose the developing petals and anthers. g–i Scanning electron micrographs of the petal epidermis of the wild type and the indicated genotypes, scale bars 50 μm. S sepal, P petal, A anther, O ovary
Modification of tomato fruit shape by carpel-specific expression of P0HA. a Represented phenotypes of control (35S:LhG4) and CRC≫P0HA ovaries at anthesis (upper panel) and immature green fruit 20 days post-anthesis (dpa) (lower panel). b Quantification of Fruit shape triangle in 20–25 dpa fruit (n = 16 for 35S:LhG4 and n = 25 for CRC≫P0HA). Error bars represent ± SD. Asterisk represents statistically significant difference as determined by Student’s t test (P ≤ 0.05). c Semi-quantitative analysis of P0HA transcript in total RNA extracted from the indicated CRC≫P0HA reproductive tissues. Specificity of primers used is indicated on the right. P0HA expression was normalized to TIP41 mRNA and levels are indicated below the panel. The number of cycles of amplification is indicated in parentheses. Non-template control (NTC); P a plasmid that harbors the corresponding gene
Discussion
We previously demonstrated that strong expression of Tomato bushy stunt virus VSR P19 in tomato can be achieved by expressing it via the OP/LhG4 transactivation system that separates transformation from transgene expression (Stav et al. 2010). Here, we successfully utilized the same approach to transgenically express the VSR P0 in tomato. P0 has been suggested to suppress silencing by inhibiting the formation of siRNA/miRNA-RISCs (Csorba et al. 2010), thus leading to AGO1 degradation (Baumberger et al. 2007; Bortolamiol et al. 2007). AGO1 is an essential component of miRNA-RISC (Baumberger et al. 2007), the effector complex of miRNAs which play key roles in plant development (Chen 2009). Accordingly, our data associate reduced SlAGO1 levels due to P0HA accumulation in 35S≫P0HA tomato seedlings with pleiotropic morphological phenotypes, including post-germination growth arrest. Such post-embryonic growth arrest is reminiscent of a fraction of the A. thaliana loss-of-function ago1 mutant seedlings that developed a single determinate pin-like organ instead of leaves (Kidner and Martienssen 2005; Lynn et al. 1999). A common phenotype of FIL≫P0HA leaves and AP1/AP3≫P0HA flowers was the development of filament-like, almost radialized leaflets and petals, respectively, indicating compromised adaxial–abaxial polarity, an organ defect that has been previously reported in several A. thaliana ago1 loss-of-function and hypomorphic mutants (Bohmert et al. 1998; Lynn et al. 1999; Kidner and Martienssen 2004). In addition, similar to A. thaliana ago1 mutants, 35S≫P0HA seedlings, FIL≫P0HA leaves and AP1≫P0HA flowers accumulated increased levels of certain miRNA target transcripts, establishing a correlation between their morphological defects and perturbation of the endogenous miRNA pathway. Together these data suggest that via expression of P0HA through transactivation, we were able to mimic ago1 mutant phenotypes in tomato. Nevertheless, as both SlAGO1-1 and SlAGO1-2 tomato homologs are sensitive to P0HA, it is highly likely that the observed phenotypes are the result of downregulation of both proteins; the contribution of each could not be determined by our current analysis. In addition, since other AGOs have been shown to destabilize in the presence of P0 (Baumberger et al. 2007), and ago10 mutant seedlings are also growth-arrested, exhibiting either an empty apex, pinhead structure or solitary leaf (Moussian et al. 1998; Lynn et al. 1999), we cannot rule out the possibility that P0-mediated degradation of tomato AGO10-like protein occurred in 35S≫P0HA seedlings and contributed to the growth-arrest phenotype.
The identification of miRNAs has revealed a previously unappreciated post-transcriptional regulatory layer that plays key roles in various aspects of vegetative and reproductive development as can be deduced from the severe developmental defects seen in A. thaliana mutants such as carpel factory/dicer-like1 (caf/dcl1) (Garcia 2008) and argonaute1 (ago1) (Bohmert et al. 1998; Vaucheret et al. 2004), both of which are major players in miRNA-mediated silencing. In light of this, and the severe 35S≫P0HA phenotype, it is reasonable to assume that the use of as yet unidentified ago1 null mutants to study AGO1 functions in later stages of tomato development will be either not feasible, because mutant tomato plants will not reach that stage, or potentially biased due to major developmental aberrations that have already taken place. On the other hand, the transactivated P0 approach enabled us to suppress SlAGO1 at a particular developmental stage or in a particular organ without disturbing prior development or unrelated organs, thus revealing the involvement of SlAGO1-mediated silencing in that particular organ or developmental stage. For example, we specifically suppressed SlAGO1 activity in the 2nd to 3rd (AP3≫P0HA) or 4th (CRC≫P0HA) flower whorls without disturbing other whorls. In addition, stage-specific SlAGO1 suppression during leaf maturation resulted in either a disordered leaf or subtle leaf phenotypes depending on the timing and expression domains of P0HA. Our results suggest that P0HA expression throughout the leaf primordium as soon as it emerges from the SAM (FIL≫P0HA) dramatically modifies leaf morphology, in turn suggesting that the primary morphogenesis phase, which determines the basic leaf structure, requires SlAGO1-mediated silencing for its normal function. Indeed, regulation of the transcription factors LANCEOLATE and GOBLET by miR319 and miR164, respectively, during this phase, has been shown to be essential for tomato compound-leaf development (Ori et al. 2007; Berger et al. 2009). Later FIL-comparable P0HA expression in the initiating leaflets of the P4 primordia (BLS≫P0HA) and at the distal end of the P5 leaf primordia (650≫P0HA) resulted in wrinkled blades and reddish vasculature, respectively, indicating that these domains are less sensitive than initiating primordia to AGO1-suppression. Taking into account that P0HA might suppress additional SlAGOs, the OP:P0HA responder line can be used to suppress SlAGO1 and unravel the involvement of AGO1-mediated silencing in any target tissue or developmental stage, provided the corresponding tomato driver line is available. Recently, comparative analyses of transgenic A. thaliana plants expressing three VSRs and a hypomorphic dcl1 mutant revealed novel endogenous targets of miRNA and trans-acting-siRNA, as well as several upregulated miRNA targets. Of the latter, the increase in AUXIN RESPONSE FACTOR 8 (ARF8) was found to be responsible for the abnormal leaf phenotypes exhibited by the VSR transgenic plants (Jay et al. 2011). Utilizing OP:P0HA reporter plants, a similar strategy will be employed in future studies to uncover putative upregulated endogenous targets of SlAGO1-mediated silencing, which may be associated with the described abnormal tomato phenotypes.
Abbreviations
- AGO1:
-
ARGONAUTE1
- miRNA:
-
MicroRNA
- PO:
-
Polerovirus silencing suppressor
- Tomato:
-
Solanum lycopersicum
References
Alvarez J, Smyth DR (1999) CRABS CLAW and SPATULA, two Arabidopsis genes that control carpel development in parallel with AGAMOUS. Development 126:2377–2386
Alvarez JP, Pekker I, Goldshmidt A, Blum E, Amsellem Z, Eshed Y (2006) Endogenous and synthetic microRNAs stimulate simultaneous, efficient, and localized regulation of multiple targets in diverse species. Plant Cell 18:1134–1151
Arazi T, Talmor-Neiman M, Stav R, Riese M, Huijser P, Baulcombe DC (2005) Cloning and characterization of micro-RNAs from moss. Plant J 43:837–848
Baumberger N, Baulcombe DC (2005) Arabidopsis ARGONAUTE1 is an RNA Slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci USA 102:11928–11933
Baumberger N, Tsai CH, Lie M, Havecker E, Baulcombe DC (2007) The Polerovirus silencing suppressor P0 targets ARGONAUTE proteins for degradation. Curr Biol 17:1609–1614
Berger Y, Harpaz-Saad S, Brand A, Melnik H, Sirding N, Alvarez JP, Zinder M, Samach A, Eshed Y, Ori N (2009) The NAC-domain transcription factor GOBLET specifies leaflet boundaries in compound tomato leaves. Development 136:823–832
Bohmert K, Camus I, Bellini C, Bouchez D, Caboche M, Benning C (1998) AGO1 defines a novel locus of Arabidopsis controlling leaf development. EMBO J 17:170–180
Bortolamiol D, Pazhouhandeh M, Marrocco K, Genschik P, Ziegler-Graff V (2007) The Polerovirus F box protein P0 targets ARGONAUTE1 to suppress RNA silencing. Curr Biol 17:1615–1621
Bowman JL, Smyth DR (1999) CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 126:2387–2396
Brodersen P, Sakvarelidze-Achard L, Bruun-Rasmussen M, Dunoyer P, Yamamoto YY, Sieburth L, Voinnet O (2008) Widespread translational inhibition by plant miRNAs and siRNAs. Science 320:1185–1190
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44
Csorba T, Lozsa R, Hutvagner G, Burgyan J (2010) Polerovirus protein P0 prevents the assembly of small RNA-containing RISC complexes and leads to degradation of ARGONAUTE1. Plant J 62:463–472
Fernandez AI, Viron N, Alhagdow M, Karimi M, Jones M, Amsellem Z, Sicard A, Czerednik A, Angenent G, Grierson D et al (2009) Flexible tools for gene expression and silencing in tomato. Plant Physiol 151:1729–1740
Garcia D (2008) A miRacle in plant development: role of microRNAs in cell differentiation and patterning. Semin Cell Dev Biol 19:586–595
Giovannoni J (2001) Molecular biology of fruit maturation and ripening. Annu Rev Plant Physiol Plant Mol Biol 52:725–749
Gleave AP (1992) A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Hunter C, Sun H, Poethig RS (2003) The Arabidopsis heterochronic gene ZIPPY is an ARGONAUTE family member. Curr Biol 13:1734–1739
Jack T, Brockman LL, Meyerowitz EM (1992) The homeotic gene APETALA3 of Arabidopsis thaliana encodes a MADS box and is expressed in petals and stamens. Cell 68:683–697
Jay F, Wang Y, Yu A, Taconnat L, Pelletier S, Colot V, Renou JP, Voinnet O (2011) Misregulation of AUXIN RESPONSE FACTOR 8 underlies the developmental abnormalities caused by three distinct viral silencing suppressors in Arabidopsis. PLoS Pathog 7:e1002035
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jones L, Keining T, Eamens A, Vaistij FE (2006) Virus-induced gene silencing of Argonaute genes in Nicotiana benthamiana demonstrates that extensive systemic silencing requires Argonaute1-like and Argonaute4-like genes. Plant Physiol 141:598–606
Kidner CA, Martienssen RA (2004) Spatially restricted microRNA directs leaf polarity through ARGONAUTE1. Nature 428:81–84
Kidner CA, Martienssen RA (2005) The role of ARGONAUTE1 (AGO1) in meristem formation and identity. Dev Biol 280:504–517
Lanet E, Delannoy E, Sormani R, Floris M, Brodersen P, Crete P, Voinnet O, Robaglia C (2009) Biochemical evidence for translational repression by Arabidopsis microRNAs. Plant Cell 21:1762–1768
Lifschitz E, Eviatar T, Rozman A, Shalit A, Goldshmidt A, Amsellem Z, Alvarez JP, Eshed Y (2006) The tomato FT ortholog triggers systemic signals that regulate growth and flowering and substitute for diverse environmental stimuli. Proc Natl Acad Sci USA 103:6398–6403
Lynn K, Fernandez A, Aida M, Sedbrook J, Tasaka M, Masson P, Barton MK (1999) The PINHEAD/ZWILLE gene acts pleiotropically in Arabidopsis development and has overlapping functions with the ARGONAUTE1 gene. Development 126:469–481
Mandel MA, Gustafson-Brown C, Savidge B, Yanofsky MF (1992) Molecular characterization of the Arabidopsis floral homeotic gene APETALA1. Nature 360:273–277
Mi S, Cai T, Hu Y, Chen Y, Hodges E, Ni F, Wu L, Li S, Zhou H, Long C et al (2008) Sorting of small RNAs into Arabidopsis Argonaute complexes is directed by the 5′ terminal nucleotide. Cell 133:116–127
Moore I, Galweiler L, Grosskopf D, Schell J, Palme K (1998) A transcription activation system for regulated gene expression in transgenic plants. Proc Natl Acad Sci USA 95:376–381
Moussian B, Schoof H, Haecker A, Jurgens G, Laux T (1998) Role of the ZWILLE gene in the regulation of central shoot meristem cell fate during Arabidopsis embryogenesis. EMBO J 17:1799–1809
Ori N, Cohen AR, Etzioni A, Brand A, Yanai O, Shleizer S, Menda N, Amsellem Z, Efroni I, Pekker I et al (2007) Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat Genet 39:787–791
Rhoades MW, Reinhart BJ, Lim LP, Burge CB, Bartel B, Bartel DP (2002) Prediction of plant microRNA targets. Cell 110:513–520
Rodriguez GR, Moyseenko JB, Robbins MD, Morejon NH, Francis DM, van der Knaap E (2010) Tomato analyzer: a useful software application to collect accurate and detailed morphological and colorimetric data from two-dimensional objects. J Vis Exp. doi:10.3791/1856
Sato S, Tabata S, Hirakawa H, Asamizu E, Shirasawa K, Isobe S, Kaneko T, Nakamura Y, Shibata D, Aoki K et al (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Shalit A, Rozman A, Goldshmidt A, Alvarez JP, Bowman JL, Eshed Y, Lifschitz E (2009) The flowering hormone florigen functions as a general systemic regulator of growth and termination. Proc Natl Acad Sci USA 106:8392–8397
Shani E, Burko Y, Ben-Yaakov L, Berger Y, Amsellem Z, Goldshmidt A, Sharon E, Ori N (2009) Stage-specific regulation of Solanum lycopersicum leaf maturation by class 1 KNOTTED1-LIKE HOMEOBOX proteins. Plant Cell 21:3078–3092
Stav R, Hendelman A, Buxdorf K, Arazi T (2010) Transgenic expression of tomato bushy stunt virus silencing suppressor P19 via the pOp/LhG4 transactivation system induces viral-like symptoms in tomato. Virus Genes 40:119–129
Takeda A, Iwasaki S, Watanabe T, Utsumi M, Watanabe Y (2008) The mechanism selecting the guide strand from small RNA duplexes is different among Argonaute proteins. Plant Cell Physiol 49:493–500
Talmor-Neiman M, Stav R, Frank W, Voss B, Arazi T (2006) Novel micro-RNAs and intermediates of micro-RNA biogenesis from moss. Plant J 47:25–37
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tang G, Reinhart BJ, Bartel DP, Zamore PD (2003) A biochemical framework for RNA silencing in plants. Genes Dev 17:49–63
Vaucheret H (2008) Plant ARGONAUTES. Trends Plant Sci 13:350–358
Vaucheret H, Vazquez F, Crete P, Bartel DP (2004) The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev 18:1187–1197
Vaucheret H, Mallory AC, Bartel DP (2006) AGO1 homeostasis entails coexpression of MIR168 and AGO1 and preferential stabilization of miR168 by AGO1. Mol Cell 22:129–136
Xiao H, Radovich C, Welty N, Hsu J, Li D, Meulia T, van der Knaap E (2009) Integration of tomato reproductive developmental landmarks and expression profiles, and the effect of SUN on fruit shape. BMC Plant Biol 9:49
Zhang X, Yuan YR, Pei Y, Lin SS, Tuschl T, Patel DJ, Chua NH (2006) Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute1 cleavage activity to counter plant defense. Genes Dev 20:3255–3268
Acknowledgments
We thank Véronique Ziegler-Graff, Institut de Biologie Moléculaire des Plantes du CNRS, Strasbourg, France, for the P0 plasmid. We thank Hanita Zemach and Eduard Belausov for excellent technical assistance in histology and microscopy, respectively. This work was supported by a grant from the Chief Scientist of the Israel Ministry of Agriculture and Rural Development no. 256-0828-10 to T.A. This is contribution no. 116/2012 series, from the Agricultural Research Organization, the Volcani Center, Bet Dagan 50250, Israel.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2012_1778_MOESM3_ESM.tif
Supplementary Figure S3.Transgenic plant validation. a Schematic representation of the binary constructs used for transgenic expression of P0HA under the OP promoter and LhG4 under the 35S promoter. Restriction enzymes used for cloning are indicated. Eight copies of the lac operator sequence linked to a minimal Cauliflower mosaic virus (CaMV) 35S promoter (OP-TATA), cauliflower mosaic virus 35S promoter (35S), octopine synthase terminator (3’ OCS), selectable kanamycin-resistance marker (nptII), nopaline synthase promoter (pNOS), nopaline synthase terminator (3’ NOS). RB – right border; LB – left border. b Genomic DNA PCR analysis of representative tomato OP:P0HA reporter lines and 35S:LhG4 driver line. Specificity of primers used is indicated on the right. (TIFF 361 kb)
425_2012_1778_MOESM4_ESM.tif
Supplementary Figure S4.Effects of leaf-specific P0HA expression on cotyledon development. Control (FIL:LhG4) and P0HA-expressing genotypes are indicated. Scale bar = 5 mm. (TIFF 524 kb)
425_2012_1778_MOESM5_ESM.tif
Supplementary Figure S5.Specific P0HA expression increases the levels of miRNA-target transcripts in the expressing tissue. Quantitative RT-PCR analysis of selected miRNA-target transcripts from the indicated genotypes in apices collected from a 1-month-old seedling (a) and 2–3 mm buds (b). Primers were designed around the corresponding miRNA complementary site. TIP41 expression values were used for normalization. Data are means ± SE of two biological replicates, each measured in triplicate. Asterisks indicate statistically significant difference compared to control as determined by Student’s t test (P ≤ 0.05). (TIFF 68 kb)
425_2012_1778_MOESM6_ESM.tif
Supplementary Figure S6.Cross sections of red fruit of the control (OP:P0HA) and indicated genotypes. Scale bar = 1 mm. (TIFF 1525 kb)
Rights and permissions
About this article
Cite this article
Hendelman, A., Kravchik, M., Stav, R. et al. The developmental outcomes of P0-mediated ARGONAUTE destabilization in tomato. Planta 237, 363–377 (2013). https://doi.org/10.1007/s00425-012-1778-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-012-1778-8