Abstract
Key message
A chalcone synthase gene ( PaCHS ) was isolated and functionally characterized from liverwort. The ectopic expression of PaCHS in Marchantia paleacea callus raised the flavonoids content.
Abstract
Chalcone synthase (CHS; EC 2.3.1.74) is pivotal for the biosynthesis of flavonoid and anthocyanin pigments in plants. It produces naringenin chalcone by condensing one p-coumaroyl- and three malonyl-coenzyme A thioesters through a polyketide intermediate that is cyclized by intramolecular Claisen condensation. Although CHSs of higher plants have been extensively studied, enzyme properties of the CHSs in liverworts have been scarcely characterized. In this study, we report the cloning and characterization of CHS (designated as PaCHS) from the liverwort Plagiochasma appendiculatum. The gene product was 60–70 % identical with chalcone synthases from other species, and contained the characteristic conserved Cys-His-Asn catalytic triad. The recombinant PaCHS was able to catalyze p-coumaroyl-CoA and malonyl-CoA to generate naringenin in vitro. Heterologously expressed PaCHS protein showed similar kinetic properties to those of higher plant CHS. The ectopic expression of PaCHS in Marchantia paleacea callus raised the content of the total flavonoids. These results suggested that PaCHS played a key role in the flavonoids biosynthesis in liverworts. Furthermore, when the thallus of P. appendiculatum was treated with abiotic stress inducers methyl jasmonate, salicylic acid and abscisic acid, PaCHS expression was enhanced. This is the first time that a CHS in liverworts has been functionally characterized.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Flavonoids are abundant and widely distributed plant secondary metabolites in vascular plants, but also have been reported in early and primitive taxa, such as bryophytes (Feld et al. 2003; Wang et al. 2013) and horsetails (Oh et al. 2004). In plants, the flavonoids have different and important roles, such as providing floral pigments in color signatures, antibiotics, UV protectants and insect repellents (Markham 1988). In addition to their important physiological roles in plants, flavonoids have been demonstrated to show significant pharmacological activities (Agrawal 2011), elevating their importance as nutritional components in our diet.
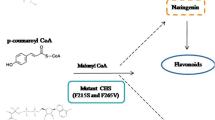
The biosynthesis of flavonoids starts with the condensation of one molecule of p-coumaryl CoA with three molecules of malonyl-CoA through the action of chalcone synthase (CHS) to generate naringenin chalcone, the common precursor for the flavonoids synthesis (Fig. 1). Chalcone synthases (CHSs) are the well-known representatives of the type III polyketide synthase (PKS) superfamily, which catalyze iterative decarboxylative condensations of malonyl unit with a CoA-linked starter molecule to generate backbones of a variety of plant secondary metabolites including chalcones, stilbenes, benzophenones, biphenyls and bibenzyls with notable structural and bioactive diversity (Yu and Jez 2008). CHSs are homodimeric enzymes comprising a pair of 40–45 kDa subunits, in which each monomer utilizes a Cys-His-Asn catalytic triad within a buried active site cavity, as revealed by the crystal structure of Medicago sativa chalcone synthase (Ferrer et al. 1999). In the past several years, the enzyme has been widely studied using genetic, biochemical and molecular approaches (Abe and Morita 2010). So far, more than 900 type III PKS genes have been reported in databases (NCBI: http://www.ncbi.nlm.gov/), and more than 20 type III PKSs have been cloned and characterized from plants and bacteria (Abe and Morita 2010).
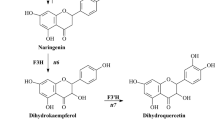
A proposed synthetic pathway for the flavonoids in liverworts. The pathway involves the conversion of 4-coumaroyl-CoA and malonyl-CoA to naringenin chalcone by CHS (chalcone synthase). PAL phenylalanine ammonia lyase, C4H cinnamic acid 4 hydroxylase, 4CL 4-coumarate coenzyme A ligase, CHI Chalcone isomerase, F3H flavanone 3β-hydroxylase, FNS flavone synthase, FLS flavonol synthase, F3′H flavonoid 3′-hydroxylase
As simple flavonoids (flavanones, flavones and flavanols) are found in bryophytes, it is thought that CHS first appeared in bryophytes during the early evolution of land plants. Hence, characterization of the CHS-superfamily enzymes in bryophytes would provide a basis for comparing the biochemical feature of CHSs in vascular and the basal land plants and for speculation on how the flavonoid biosynthetic pathways evolved throughout the history of plants (Harashima et al. 2004). A CHS gene (PpCHS) has been cloned from the gametophores of Physcomitrella patens, and the heterologously expressed PpCHS protein showed similar enzymatic characteristics as compared to higher plant CHSs (Jiang et al. 2006). Flavonoids have been isolated from liverworts (Asakawa 1995), and the CHS enzyme activity was also demonstrated in crude extracts of Marchantia polymorpha (Fischer et al. 1995). However, as yet, only one CHS-like gene was isolated from the liverwort, Marchantia paleacea, and its gene product (presumably a CHS) was shown to be expressed light dependently in photoautotrophic cells (Harashima et al. 2004). The details of the enzyme properties of the CHS in liverworts remain unclear prior to the present investigation.
Here, we describe one CHS present in liverwort P. appendiculatum, which condensed p-coumaroyl-CoA as starter CoA with malonyl-CoA to form naringenin. Both the structure and biochemical function of this CHS is similar with that of the higher plants’ CHS. To comprehensively investigate the CHS-like genes in liverworts, the biochemical feature of other CHS-like genes of liverwort (PaSTCS1 and MpSTCS2) in the genbank database was also characterized. Ectopic expression of PaCHS increased the flavonoids content in M. paleacea callus. The expression of PaCHS was induced when the thallus of P. appendiculatum was treated with abiotic stress inducers methyl jasmonate (MeJA), abscisic acid (ABA) and salicylic acid (SA).
Materials and methods
Reagents
p-Coumaroyl-CoA was synthesized following Stöckigt and Zerk (Stöckigt and Zenk 1974). All other reagents were purchased from Sigma-Aldrich (St. Louis, USA) unless otherwise stated.
Plant material and phytohormone treatment
Plagiochasma appendiculatum thallus was propagated asexually by gemmae and sexually, and was maintained in a greenhouse held at 25 °C with a 12-h photoperiod. Two-month-old thallus of P. appendiculatum was treated by 1 mM ABA, 1 mM MeJA or 1 mM SA for 0, 6, 12, 24, 36 and 48 h. M. paleacea callus was maintained at 22 ± 1 °C, 12 h photoperiod on Murashige and Skoog (1962) medium supplemented with 1 mg L−1 2,4-D and sub-cultured every 2 weeks (Cheng et al. 2013; Wang et al. 2013). All samples were immediately frozen in liquid nitrogen and then stored at −80 °C until further use.
DNA and RNA isolation and gene expression analysis
Genomic DNA was extracted from fresh P. appendiculatum thallus according to the CTAB method (Porebski et al. 1997). Total RNA was extracted from the thallus using the modified CTAB method (Gambino et al. 2008). The DNA and RNA were quantified with Eppendorf BioPhotometer plus (Eppendorf, Germany).
For the gene expression analysis, first-strand cDNA was synthesized according to the manufacturer’s instructions of RevertAid™ First Strand cDNA Synthesis Kit (MBI, USA). The gene transcription level in the untreated (control) and 1 mM ABA-, 1 mM MeJA- or 1 mM SA-treated P. appendiculatum thallus was quantified with gene-specific primers (PaCHS-RTF and PaCHS-RTR) (Table 1) utilizing the Mastercycler ep realplex RealTime PCR System (Eppendorf, Germany). All samples were evaluated in three independent experiments. The elongation factor from P. appendiculatum, selected as a reference gene, was amplified employing the PaelongationF and PaelongationR primers (Sun et al. 2013).
CHS cloning, recombinant protein expression and purification
The PaCHS open reading frame (ORF) was amplified from the sequenced cDNA clones (Cheng et al. 2013) using primers PaCHSF/R (Table 1), meanwhile, the genomic DNA sequence was obtained with the same primers from genomic DNA (Table 1). Primers PaSTCS1F/R and MpSTCS2F/R were used to amplify the corresponding ORFs of PaSTCS1 and MpSTCS2, and then the ORFs were introduced into the pET32a plasmid and transferred into Escherichia coli BL21 (DE3) cells to obtain heterologous expression. The recombinant protein was induced by the addition of 0.5 mM IPTG and purified as described previously (Sun et al. 2013). Protein concentrations were determined using the Bradford reagent (Beyotime, Shanghai, China) with BSA employed as a standard. The resulting soluble and purified proteins were subjected to denaturing SDS-PAGE (12 % acrylamide). Standard molecular weight markers (10–170 kDa, Fermantas) were applied to estimate the Mr of the heterologously expressed proteins, following their Coomassie blue R250 staining.
Enzymatic assay and kinetics
Enzyme assays were performed at 35 °C for 30 min in 250 μl 100 mM potassium phosphate buffer (pH 7.0) containing 50 μM p-coumaroyl-CoA, 56 μM malonyl-CoA and 10 μg protein extracted from the recombinant E. coli cells; as a control, protein was also prepared from E. coli carrying an empty pET32a plasmid. The reactions were terminated by adding 25 μl of 10 % glacial acetic acid, then extracted twice with 250 μl ethyl acetate and centrifuged at 12,000×g for 10 min. After the solvent had been removed under vacuum, the residue was dissolved in 50 μl methanol. Analysis of the enzymatic products was performed using HPLC (1260 Infinity Binary LC system, Agilent Technologies, USA) and HPLC–MS. The latter employed an Agilent 1100 system (Agilent Technologies, CA) equipped with a multi-wavelength diode array detector and an ESI mass spectrometer. Samples were separated through a reversed-phase C18-silica column (XDB-18, 5 μm; Agilent) with a flow rate of 0.8 mL/min. The eluents were water containing 0.2 % glacial acetic acid (A) and methanol (B); the ratio of B rose from 50 to 85 % over the first 21 min, and then was reduced to 50 % for a further 5 min. Standard solutions of reference compounds were used for calibration.
For optimum temperature determination, incubations were carried out at a constant pH of 7.0 while varying the temperatures (20, 30, 35, 40, 50, and 60 °C). The assay mixture was the same as described before. For pH optimum, assays were performed at 35 °C with a series of potassium phosphate buffer with different pHs (100 mM; pH 6.0, 6.2, 6.5, 6.7, 6.9, 7.0, 7.2, 7.4, 7.6 and 8.0). For all data points, including pH and temperature optimum determinations, three parallels were done.
The kinetic parameters were determined using seven different substrate concentrations, and the second substrate was the saturated point with a concentration of 400 μM. The experiment was done three times with 2 μg purified enzyme in a final volume of 250 μl of 100 mM potassium phosphate buffer at 35 °C for 10 min. The amount of reaction product was calculated according to the standard calibration curve. Then kinetic parameters were obtained by Michaelis–Menten equation analysis with Graphpad Prism 5 software.
Sequence alignment and bioinformatic analyses
Deduced CHS polypeptide sequences were aligned with those of other known plant type III PKSs using DNAMAN software (Version 5.2.2, Lynnon Biosoft, Canada). SWISS-MODEL was used to construct the three-dimensional (3D) structural model of the PaCHS-product analog complexes (Arnold et al. 2006), the resulting models were visualized using PyMOL (http://www.pymol.org/citing). A phylogenetic tree was constructed using MEGA 5.0 software, applying the neighbor-joining method (Tamura et al. 2007).
Transformation of PaCHS gene into M. paleacea callus
The full length PaCHS cDNA was amplified by PCR with PaCHSPS-F/R primers, and subcloned into the plant binary vector pSTART under the control of the CaMV 35S promoter. The resulting construct was then introduced into Agrobacterium tumefaciens strain GV3101 via the freeze/thaw method. A 1 mL aliquot of an overnight A. tumefaciens culture was inoculated into 100 mL yeast extract peptone medium supplemented with 50 mg L−1 gentamycin, 50 mg L−1 kanamycin and 50 mg L−1 rifamycin. The cultures were shaken (200 rpm) at 28 °C until the OD600 had reached 1.5–2.0. The cells were harvested by centrifugation (4,000×g, 5 min) and re-suspended in 50 mL liquid MS medium supplemented with 100 μM acetosyringone (Sigma-Aldrich, St Louis, MO, USA). The cells were added to M. paleacea callus which was maintained on MS medium under a 16-h photoperiod at 22 °C for 7 day, the culture was held for 1 h with agitation (110 rpm) and then left to develop at 22 °C in the dark for 72 h. The calli were subsequently washed four times in sterile, deionized water, and transferred to a MS medium containing 50 mg L−1 kanamycin and 200 mg L−1 cefotaxime to select for transformed cells. After a further 2–3 weeks, surviving calli were transferred to a MS medium containing 100 mg L−1 kanamycin and 50 mg L−1 cefotaxime, on which they were sub-cultured at two weekly intervals. RNA was extracted from transgenic and wild-type calli using modified CTAB method (Gambino et al. 2008) and then reverse transcripted to cDNA which was used as the template for real-time RT-PCR, employing as primers PaCHS-RTF/R (Table 1). A fragment of the M. paleacea gene encoding an elongation factor gene was used as the internal reference sequence, and amplified using primers Mpa Elongation F/R (Table 1).
Chemical component analysis
Transgenic and non-transgenic calli were snap frozen in liquid nitrogen and ground to a fine powder. To analyze flavonoid content, a ~0.1 g powder was mixed with 1 mL of methanol containing 1 % HCl and ultrasonicated for 1 h. The extract (two parts) was thoroughly mixed with two parts of chloroform and one part of deionized water, followed by centrifugation at 13,000 rpm for 10 min, the supernatant was applied to measure the A340 values using a U-3900 spectrophotometer (Hitachi High-Tech, Japan) (Muir et al. 2001).
Results
Isolation of CHS cDNAs from liverwort
Two unigenes, referred to as PaSTCS1 and PaCHS from a sequenced normalized cDNA library of P. appendiculatum thallus (Cheng et al. 2013) showed high sequence homology with genes encoding plant type III PKSs. Using their sequences as a query string, it was then possible to identify M. polymorpha homologue, namely MpSTCS2 (Accession no., AAW30010). The two P. appendiculatum CHS-like sequences have been submitted to GenBank as accessions KF724393 and KF724394. The full length MpSTCS2 cDNA was derived from thallus mRNA. The largest open reading frame (ORF) of the PaSTCS1 open reading frame (ORF) was 1,182 bp, and that of PaCHS 1,236 bp, so their predicted translation products comprised, respectively, 393 and 411 residues, with a calculated molecular mass of, respectively, 42.7 kDa and 44.4 kDa. The length of the MpSTCS2 ORF was 1,179 bp, encoding polypeptides of length 392 residues, with a calculated molecular mass of 44.4 kDa. A sequence alignment indicated that PaCHS and MpSTCS2 were highly homologous to one another with 89 % identity, while the PaSTCS1 shared less than 60 % identity with the PaCHS and MpSTCS2 (Fig. 2). The deduced peptide sequences of PaCHS, MpSTCS2 and PaSTCS1 polypeptides were only 60–70 % homologous with other known CHS sequences, but retained the catalytic triad Cys180–His319–Asn352 (numbering with PaCHS), which is a characteristic of plant type III PKSs (Fig. 2). Another highly conserved CHS signature sequence, G388FGPG (Suh et al. 2000) were also found in them (Fig. 2). Most of the active site residues identified in M. sativa (Thr132, Ser133, Met137, Gly163, Thr194, Gly211, Gly216, Ile254, Ser338 and Pro375), along with the CHS “gatekeeper” Phe215 and Phe265 (residue numbers refer to M. sativa CHS) (Jez et al. 2002) were also present in the liverwort CHSs (Fig. 2). These results suggest that PaCHS, MpSTCS2 and PaSTCS1 are members of the CHS-superfamily enzymes.
Alignment of the deduced amino acid sequence of P. appendiculatum CHS with other CHS-superfamily enzymes. Multiple sequence alignment was calculated with the DNAMAN package. Black shading shows amino acid identities, red and blue shading shows amino acid with different similarity. The catalytic residues conserved in the plant type III PKS (Cys164, His303 and Asn336, numbering of MsCHS) are indicated with diamond, the highly conserved CHS signature sequence, G388FGPG are boxed with red line. The abbreviations for species and accession numbers are: AtCHS (Arabidopsis thaliana, AAB35812), PpCHS (P. patens, ABB84527), MpCHSLK1 (M. paleacea, AB119056), MsCHS (M. sativa, AAA02824), MpSTCS1 (M. polymorpha, AAW30009), MpSTCS2 (M. polymorpha, AAW30010) and PaSTCS1 (P. appendiculatum, AHY39237) (color figure online)
Three-dimensional model of PaCHS
To characterize the structural properties of PaCHS, a homology model was generated from the native crystal structure of M. sativa CHS–naringenin complexes (1CGK) using the SWISS-MODEL server (Arnold et al. 2006; Ferrer et al. 1999), which shared 64 % identity with the query sequence. PaCHS monomer presented similar folding patterns to those of the M. sativa CHS X-ray structures, which consisted of two main structural domains. The naringenin located at the active cavity of the PaCHS monomer (Fig. 3a). The active site residues of PaCHS possess similar position with those from M. sativa CHS, in particular, the catalytic triad (Cys180, His319 and Asn352) and the highly conserved Phe231 define the active site (residue numbers refer to PaCHS) (Fig. 3b). As earlier reports have been noted, the Cys180 may serve as a nucleophile and play the attachment function through CoA molecules. His319 most likely acts as a general base during the generation of a nucleophilic thiolate anion from Cys180. Phe231 and Asn352 may function in the decarboxylation reaction. Phe281 separates the coumaroyl binding site from the cyclization pocket and may function as a mobile steric gate during successive rounds of polyketide elongation (Ferrer et al. 1999).
Homology model of the PaCHS-product analog complexes. a The CHS–naringenin complex viewed down the CoA-binding tunnel. b The key residues are highlighted and labeled. Bonds are colored by atom type with hydrogen bonds depicted as dashed lines. Naringenin and protein carbon atoms are colored green and cyan, respectively. Bonds are colored by atom type with hydrogen bonds depicted as dashed cylinders (color figure online)
Other active residues including Thr148, Ser149, Thr210, Thr213, Gly232, Ile270, Leu279, Ser338, and Pro391, which may participate in CoA-binding, coumaroyl-binding and/or cyclization process, are all conserved in their position (Fig. 3b). These residues interact with the naringenin molecule primarily via van der Waals contacts. However, the carbonyl oxygen of Gly232 hydrogen bonds to the phenolic oxygen of naringenin, and the hydroxyl of Thr213 interacts with the carbonyl of naringenin derived from coumaroyl-CoA.
Genomic organization of the PaCHS gene
A 1,700 bp PCR fragment was obtained from the genomic DNA of P. appendiculatum with primers PaCHSF/R, and the sequence has been submitted to genebank as accession KM506763. Comparing the cDNA and genomic DNA sequences of PaCHS indicate that this gene contains one 464 bp intron and two exons (Fig. 4a), the gene model of PaCHS and several other CHS genes harboring one intron was shown in Fig. 4. Several previous studies showed that most CHS-superfamily genes contained an intron in a conserved position starting in an Cys codon, splitting Cys in a highly conserved position (Harashima et al. 2004). Interestingly, the intron of PaCHS gene inserted after a Cys codon, locating between exon 1 and exon 2, which is different among these CHS genes (Fig. 4b).
Genome structure of PaCHS and other CHS-superfamily genes. a Schematic representation of genome structure. The black arrow and gray box indicate the exon and intron, respectively. b Nucleotide and amino acid sequences flanking the intron in PaCHS and other CHSs. The accession number of each gene is as follows: PpCHS (P. patens, DQ275627), MpSTCS2 (M. polymorpha, AY847706), PsCHS (Pinus sylvestris, X60754), MpCHSLKG1 (M. paleacea, AB119057), AtCHS (A. thaliana, S80554), PtCHSA (Petunia hybrida, X14591), HmSTCS2 (Hydrangea macrophylla, AF456446), MpSTCS1 (M. polymorpha, AY847705)
Phylogenetic analysis
A phylogenetic tree was created by the neighbor-joining method using PaCHS, MpSTCS2, PaSTCS1 and other CHS-superfamily polypeptide sequences, with the E. coli β-ketoacyl-synthase III as an outgroup. In the phylogenetic analysis based on 1,000 replicates, the phylogeny tree occurred from the E. coli β-ketoacyl-synthase III and diverged into several major groups. The CHS-like genes PaCHS and MpSTCS2 from liverworts and PpCHS from moss grouped closer and occured early from the spanning tree root, while the other CHS-like genes PaSTCS1, MpSTCS1, MpCHSLK1 and LcCHSL from liverworts got together and occurred just after the first branch. Then CHSs from pteridophytes appeared next to the CHSs of liverworts. Then the CHSs and STSs from gymnosperms and angiosperms formed a large cluster (Fig. 5). The position of the CHS proteins in the phylogenetic tree is consistent with their evolutionary position.
Phylogenetic relationships among CHS-superfamily enzymes. The sequences were aligned with clustalW with default settings, and the tree was developed with MEGA 4.0 program, specifically employing the neighbor-joining method. The numbers at each fork represent bootstrap percentages estimated from 1,000 replicates. E. coli β-ketoacyl-(acyl carrier protein) synthase III (KAS III, BAA35899) served as an outgroup. The accession numbers used in the analysis are listed as follows: H. macrophylla (STCS, AAN76182; CHS, BAA32732), Arachis hypogaea (STS, BAA78617; CHS, AAO32821), M. sativa CHS (P30074), Glycine max CHS (ABB30178), Petroselinum crispum CHS (CAA24779), Vitis vinifera CHS (CAA53583), Gerbera hybrid CHS (CAA86218), A. thaliana CHS (AT5G13930), Zea mays CHS (CAA42764), Picea mariana CHS (AAF35890), Equisetum arvense CHS (BAA89501), Psilotum nudum (CHS, BAA87922; STS, BAA87924), P. patens CHS (ABB84527), Huperzia serrata CHS (ABI94386), Cannabis sativa CHS (AAL92879), Ginkgo biloba (AAT68477), Pinus strobus(STS, CAA87012; CHS, CAA06077), P. sylvestris STS (CAA43165), Rheum palmatum CHS (ABB13607), Rheum tataricum STS(AAP13782), Sorbus aucuparia CHS (ABB89213), P. hybrida CHSA (CAA32731), Hypericum androsaemum CHS (AAG30295), Hypericum perforatum CHS (AAL67805), Rubus idaeus CHS (AAK15174), Chrysosplenium americanum CHS (AAB54075), Scutellaria baicalensis CHS (BAA23373), M. paleacea CHSLK1 (BAD42329), M. polymorpha (STCS1, AAW30009; STCS2, AAW30010), P. appendiculatum STCS1 (AHY39237), Lunularia cruciata CHSL (ABF67971), Selaginella moellendorffii CHSL (XP_002965309)
Biochemical characterization of recombinant PaCHS in vitro
The putative PaCHS ORF was digested with BamHI and XhoI and ligated to pET32a vector for heterologously expression in E. coli, which produced the PaCHS recombinant protein with Trx-tag, His-tag and S-tag. As shown in Fig. 6, the purified recombinant proteins were analyzed by SDS-PAGE with relative molecular mass of about 70 kDa (including the tag). Incubation of PaCHS with p-coumaroyl-CoA and malonyl-CoA produced a product which shared the same HPLC retention time as naringenin (Fig. 7a, b). Its negative electron spray ionization (ESI)-ion mass spectrum resolved a molecular ion [M-H]− at m/z of 271. The profile of fragmented form of this [M-H]− and MS2 species was identical to that generated from naringenin standard (Fig. 7c–f). The majority daughter ions at m/z 151 indicated the induction of fission of the C-ring via the retro Diels–Alder reaction, while the m/z 177 ion reflected the loss of the B-ring (Fig. 7g). Thus, PaCHS was able to use p-coumaryol-CoA as a substrate to generate naringenin (Fig. 7). The same experiments above were done for MpSTCS2 and PaSTCS1, and the results showed that the MpSTCS2 recombinant protein was able to use p-coumaryol-CoA as a substrate to generate naringenin chalcone, but PaSTCS1 was not (Fig. S1–S3).
In vitro activity of recombinant PaCHS presented with p-coumaroyl-CoA and malonyl-CoA as substrate. HPLC chromatograms of the products in the reactions of PaCHS recombinant protein and the empty control (a), naringenin standard (b). c, d The mass spectrometric spectrum of reaction products. e, f The mass spectrometric spectrum of the naringenin standard. g The putative splitting way of naringenin on MS2
Monitoring the catalytic activity of the enzyme with different temperature and pH value revealed that the optimal temperature for PaCHS was 35 °C, and the optimum pH was 7.4. Under optimum pH and temperature, using p-coumaryol-CoA and malonyl-CoA as substrates, the PaCHS recombinant enzyme showed an apparent K m of 11.7 ± 3.8 µM (n = 3, mean ± standard error) and k cat of 0.0129 s−1 for p-coumaryol-CoA and K m of 23.76 ± 3.3 µM and k cat of 0.0129 s−1 for malonyl-CoA, which were similar to the values of CHSs from other higher plants (Jiang et al. 2006).
PaCHS expression in response to abiotic elicitors
Expression of the CHS gene can be stimulated by various biotic and abiotic elicitors including light, infection, mechanical wounding and plant hormones (Harashima et al. 2004; Jeong et al. 2004; Junghans et al. 1993). To characterize the transcription pattern of PaCHS in response to different abiotic elicitors (MeJA, ABA and SA), the expression levels of the PaCHS gene were measured using real-time PCR. As shown in Fig. 8a, PaCHS transcripts in the P. appendiculatum thallus were significantly induced by MeJA treatment, initially increasing gradually from 6 to 12 h, then reaching the highest levels at 24 h with about a fourfold increase in comparison to the control. Transcript levels gradually declined from 36 h. In the thallus treated with ABA, PaCHS transcription enhanced after 12 h and reached the highest level at 24 h with about a fourfold increase comparing with the control followed by sharp decline, with approximately 20 % to the control levels at 48 h (Fig. 8b). For the SA-treated sample, the PaCHS was markedly induced, with about twofold higher than the control at 6 h, then with fourfold increase at 12 h and 24 h. Transcript levels then decreased from 36 h (Fig. 8c).
Expression characteristics of PaCHS in response to MeJA (a) and ABA (b) and SA (c) at different time points (0, 6, 12, 24, 36, 48 h). Elongation factor was used as endogenous control and the results were shown as the mean of three replicate reactions with standard deviations, *p < 0.05, **p < 0.01 according to Student’s t test
The constitutive expression of PaCHS in M. paleacea enhanced flavonoids production
To further dissect the in vivo function, PaCHS was successfully constitutively expressed in M. paleacea callus using Agrobacterium-mediated genetic transformation (Fig. 9a–e). The expression cassettes were assembled in the pSTART binary vector (Fig. 9a). The M. paleacea calli maintained on MS medium under a 16-h photoperiod at 22 °C for 7 d were used as the transformation target (Fig. 9b). The 7-day-old calli were co-cultivated with Agrobacterium in the presence of acetosyringone (Fig. 9c). After co-cultivation for 3 days in the dark, the calli were transferred to MS medium with kanamycin and cefotaxime for 2–3 weeks, which led to the formation of kanamycin-resistant clones, whereas kanamycin-sensitive cells developed into chlorotic cell clumps (Fig. 9d). The surviving calli were sub-cultured every 2 weeks in the MS medium containing kanamycin and cefotaxime (Fig. 9e). DNA and RNA from transgene lines and non-transformed callus were extracted for further analyses. PCR were performed to identify the transgene clones using the forward primer 35SF which located at the binary vector and the reward primer PaCHS-RTR in Table 1. The results showed that PaCHS has been inserted into the genome of M. paleacea (data not shown). To monitor transgene expression in M. paleacea callus, RT-qPCR was applied to analyze the transcription abundance of the wild-type and 3 independent transgenic lines randomly selected from kanamycin-resistant callus (Fig. 9f). Analysis of the flavonoid content of the transgenic and non-transgenic material showed that the heterologous expression of PaCHS was associated with an enhanced accumulation of flavonoids (Fig. 9g).
Heterologous expression of PaCHS in M. paleacea. a The transgene cassette used for the heterologous expression of PaCHS. b Callus used as the transformation recipient. c Callus co-cultured with A. tumefaciens harboring PaCHS. d Post co-culturing callus exposed to kanamycin and cefotaxime selection. e Transgenic callus growing on a medium containing 100 μg mL−1 kanamycin. f PaCHS transcript abundance in three independent transgenic lines by RT-PCR. g Total flavonoid content in callus expressing PaCHS and wild-type callus. Values represent the mean ± SD measured from at least three biological replicates. Statistical analysis was performed using the Student’s t test. *Significant difference at p < 0.05
Discussion
Chalcone synthases (CHSs), the well-known representatives of the type III polyketide synthase (PKS) superfamily, catalyze the condensation of p-coumaroyl-CoA and three malonyl-CoA molecules to form the naringenin chalcone, which is the first committed step in the flavonoids pathway of plants, leading to the biosynthesis of flavonoids, isoflavonoids, and anthocyanins (Oh et al. 2004). Chalcone synthases have been extensively studied in higher plants. While flavonoids are ubiquitous in vascular plants, they have also been found in liverworts and mosses (Markham 1988), which are the earliest diverging extant lineages of land plants. However, as yet, only one CHS gene from the moss P. patens was functionally characterized and the enzyme properties of the CHSs in liverworts have not been characterized prior to the present investigation although a CHS-like gene sequence was identified from M. paleacea (Harashima et al. 2004). Here, we characterized one plant type III PKS in the liverwort P. appendiculatum. Sequence analysis and homology modeling revealed that PaCHS shares characteristics with many other CHSs and possessed a conserved Cys–His–Asn catalytic triad in an internal active site (Figs. 2, 3). The biochemical function analysis indicated that PaCHS catalyze p-coumaroyl-CoA and malonyl-CoA to form naringenin and showed the similar kinetics parameters with the CHSs in higher plants and P. patents (Jiang et al. 2006). These experimental data suggested that both the structure and biochemical function of CHSs in liverworts are conserved with the higher plants. We also characterized the other CHS-like genes of liverwort in the genbank database. The results indicated that MpSTCS2 showed the same catalytic activity with PaCHS, while PaSTCS1 did not show any chalcone-naringenin formation activity (Fig. S2-S3). From the phylogenetic analysis, PaSTCS1 from P. appendiculatum, MpSTCS1 and MpCHSLK1 from M. paleacea were clustered in one clade, while PaCHS and MpSTCS2 formed a distinct cluster with PpCHS, which were consistent with their chalcone-naringenin formation activity of these genes. Our results indicated that the CHS appeared earlier than the split of the different bryophyte lineage. The identification of other CHS-like genes in liverworts is undertaken, which will shed light on the original and evolution of CHS genes in plants.
Previous investigation indicated that a conserved gene structure is a common feature of the plant-specific type III PKS genes in flowering plants including both the angiosperms and gymnosperms. Most plant-specific type III PKS genes studied so far contain a single intron (Durbin et al. 2000). Antirrhinum majus CHS gene was reported a second intron in exon 2 in an earlier research (Sommer and Saedler 1986). Recently, two PKSs encoded by three-intron genes from Polygonum cuspidatum was characterized (Ma et al. 2009). Notably, the P. patens CHS gene family exhibits an unusual collection of genes in terms of exon–intron architectures in that there are seven intronless genes and two genes with two introns as well as five with one intron (Koduri et al. 2010). Here we reported PaCHS obtained from liverwort containing one intron, similar to many PKS III genes previously reported. However, the position of the intron of PaCHS is located after the third nucleotide of the Cys76-encoding triplets (numbering of P. appendiculatum), which is different from other CHSs. The intron is inserted between the first and second nucleotides of the Cys-encoding triplets in other CHS-superfamily genes (Harashima et al. 2004). The subtle differences of the intron position between the liverwort P. appendiculatum and other plants did not affect the biochemical function.
Flavonoids play an important role in plant defense, and CHS as the gatekeeper of flavonoid biosynthesis plays an important role in regulating the pathway. There have been a great number of reports that CHS gene expression in different plants was induced by various environmental stresses. For example, induction of CHS gene expression by UV light, wounding, elicitor and temperature changing have been reported in various plants (Christie and Jenkins 1996; Jeong et al. 2004; Junghans et al. 1993). In the present investigation, PaCHS transcripts were measured after MeJA, ABA or SA treatment. The PaCHS transcripts responded effectively after MeJA treatment and exhibited consistent trend with other plants that under MeJA stress (Lei et al. 2010). The PaCHS expression was also increased after ABA and SA treatment. These results indicated that the PaCHS might contribute to the defense of abiotic stresses in P. appendiculatum.
As an essential component of functional genomics, transformation techniques have been developed in M. polymorpha, the model plant of liverworts. Cells of young thalli grown from gemmae and immature thalli could be readily transformed by particle bombardment (Chiyoda et al. 2008; Takenaka et al. 2000) and Agrobacterium-mediated genetic transformation (Ishizaki et al. 2008). In M. polymorpha, genetic transformation has facilitated gene functional analyses via overexpression and gene silencing (Ishizaki et al. 2008). However, no successful transformation method has been reported in other liverworts. In the present investigation, we established a high efficiency Agrobacterium-mediate transformation method using rapid growing M. paleacea callus. In our protocol, calli developed from spores were directly infected by Agrobacterium and selected on kanamycin containing medium and acetosyringone was required during the infection and co-culture steps, which indicated that virulence gene activity induced by acetosyringone is essential for Agrobacterium-mediated T-DNA transfer into M. paleacea callus. The efficient Agrobacterium-mediated transformation using M. paleacea callus provided a useful method to investigate the gene function of liverworts. Using this efficient transformation method, we successfully transformed PaCHS into M. paleacea callus. The total flavonoids content was increased in the transgenic callus, which indicated that the in vivo function of PaCHS was consistent with that of in vitro. The expression level of PaCHS was lower in the transgene line two than other two lines as shown in Fig. 9f, which would lead to the lower flavonoids synthesis in transgenic line two. In addition, there would be some more complicated regulatory mechanism for the flavonoids synthesis, leading to the differences among the transgenic lines.
The present study cloned and functionally characterized a CHS from the liverwort P. appendiculatum. This is the first time the roles of CHS in liverworts have been functionally characterized both in vitro and in vivo. The results show that the CHS isolated from the basal land plants, liverworts, has similar structure and function with that of higher plants. Our results add a new gene to the flavonoids biosynthetic pathway and provide new insight into the evolution of flavonoids biosynthesis.
Author contribution statement
Ai-Xia Cheng and Hong-Xiang Lou designed the research. Hai-Na Yu and Lei Wang performed most of the experiments. Bin Sun and Shuai Gao contributed to the substrate synthesis experiment. Ai-Xia Cheng and Hai-Na Yu wrote the paper. All authors discussed the results and edited the article and approved the final manuscript.
References
Abe I, Morita H (2010) Structure and function of the chalcone synthase superfamily of plant type III polyketide synthases. Nat Prod Rep 27:809–838
Agrawal A (2011) Pharmacological activities of flavonoids: a review. Int J Pharm Sci Nanotechnol 4:1394–1398
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinform 22:195–201
Asakawa Y (1995) Chemical constituents of the bryophytes. In: progress in the chemistry of organic natural products. Springer, vienna, Berline, pp 1–562
Cheng A, Wang L, Sun Y, Lou H (2013) Identification and expression analysis of key enzymes of the terpenoids biosynthesis pathway of a liverwort Plagiochasma appendiculatum by EST analysis. Acta Physiol Plant 35:107–118
Chiyoda S, Ishizaki K, Kataoka H, Yamato KT, Kohchi T (2008) Direct transformation of the liverwort Marchantia polymorpha L. by particle bombardment using immature thalli developing from spores. Plant Cell Rep 27:1467–1473
Christie JM, Jenkins GI (1996) Distinct UV-B and UV-A/blue light signal transduction pathways induce chalcone synthase gene expression in Arabidopsis cells. Plant Cell 8:1555–1567
Durbin ML, McCaig B, Clegg MT (2000) Molecular evolution of the chalcone synthase multigene family in the morning glory genome In: Plant Molecular Evolution. Springer, Netherlands, Berline, pp 79–92
Feld H, Zapp J, Becker H (2003) Secondary metabolites from the liverwort Tylimanthus renifolius. Phytochemistry 64:1335–1340
Ferrer J-L, Jez JM, Bowman ME, Dixon RA, Noel JP (1999) Structure of chalcone synthase and the molecular basis of plant polyketide biosynthesis. Nat Struct Mol Biol 6:775–784
Fischer S, Böttcher U, Reuber S, Anhalt S, Weissenböck G (1995) Chalcone synthase in the liverwort Marchantia polymorpha. Phytochemistry 39:1007–1012
Gambino G, Perrone I, Gribaudo I (2008) A rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochem Anal 19:520–525
Harashima S, Takano H, Ono K, Takio S (2004) Chalcone synthase-like gene in the liverwort, Marchantia paleacea var. diptera. Plant Cell Rep 23:167–173
Ishizaki K, Chiyoda S, Yamato KT, Kohchi T (2008) Agrobacterium-mediated transformation of the haploid liverwort Marchantia polymorpha L., an emerging model for plant biology. Plant Cell Physiol 49:1084–1091
Jeong ST, Goto-Yamamoto N, Kobayashi S, Esaka M (2004) Effects of plant hormones and shading on the accumulation of anthocyanins and the expression of anthocyanin biosynthetic genes in grape berry skins. Plant Sci 167:247–252
Jez JM, Bowman ME, Noel JP (2002) Expanding the biosynthetic repertoire of plant type III polyketide synthases by altering starter molecule specificity. Proc Natl Acad Sci USA 99:5319–5324
Jiang C, Schommer CK, Kim SY, Suh D-Y (2006) Cloning and characterization of chalcone synthase from the moss, Physcomitrella patens. Phytochemistry 67:2531–2540
Junghans H, Dalkin K, Dixon RA (1993) Stress responses in alfalfa (Medicago sativa L.). 15. Characterization and expression patterns of members of a subset of the chalcone synthase multigene family. Plant Mol Biol 22:239–253
Koduri PH, Gordon GS, Barker EI, Colpitts CC, Ashton NW, Suh D-Y (2010) Genome-wide analysis of the chalcone synthase superfamily genes of Physcomitrella patens. Plant Mol Biol 72:247–263
Lei W, Tang S-H, Luo K-M, Sun M (2010) Molecular cloning and expression profiling of a chalcone synthase gene from hairy root cultures of Scutellaria viscidula Bunge. Genet Mol Biol 33:285–291
Ma L-Q, Guo Y-W, Gao D-Y, Ma D-M, Wang Y-N, Li G-F, Liu B-Y, Wang H, Ye H-C (2009) Identification of a Polygonum cuspidatum three-intron gene encoding a type III polyketide synthase producing both naringenin and p-hydroxybenzalacetone. Planta 229:1077–1086
Markham KR (1988) Distribution of flavonoids in the lower plants and its evolutionary significance. In: the flavonoids. Springer, US, Berline, pp 427–468
Muir SR, Collins GJ, Robinson S, Hughes S, Bovy A, De Vos CR, van Tunen AJ, Verhoeyen ME (2001) Overexpression of petunia chalcone isomerase in tomato results in fruit containing increased levels of flavonols. Nat Biotechnol 19:470–474
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Oh H, Kim D-H, Cho J-H, Kim Y-C (2004) Hepatoprotective and free radical scavenging activities of phenolic petrosins and flavonoids isolated from Equisetum arvense. J Ethnopharmacol 95:421–424
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Rep 15:8–15
Sommer H, Saedler H (1986) Structure of the chalcone synthase gene of Antirrhinum majus. Mol Gen Genet 202:429–434
Stöckigt J, Zenk M (1974) Chemical syntheses and properties of hydroxycinnamoyl-coenzyme A derivatives. Z Naturforsch Sect C Biosci 30:352–358
Suh D, Fukuma K, Kagami J, Yamazaki Y, Shibuya M, Ebizuka Y, Sankawa U (2000) Identification of amino acid residues important in the cyclization reactions of chalcone and stilbene synthases. Biochem J 350:229–235
Sun Y, Wu Y, Zhao Y, Han X, Lou H, Cheng A (2013) Molecular cloning and biochemical characterization of two cinnamyl alcohol dehydrogenases from a liverwort Plagiochasma appendiculatum. Plant Physiol Biochem 70:133–141
Takenaka M, Yamaoka S, Hanajiri T, Shimizu-Ueda Y, Yamato KT, Fukuzawa H, Ohyama K (2000) Direct transformation and plant regeneration of the haploid liverwort Marchantia polymorpha L. Transgenic Res 9:179–185
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Wang L, Wang L-N, Zhao Y, Lou H-X, Cheng A-X (2013) Secondary metabolites from Marchantia paleacea calluses and their allelopathic effects on Arabidopsis seed growth. Nat Prod Res 27:274–276
Yu O, Jez JM (2008) Nature’s assembly line: biosynthesis of simple phenylpropanoids and polyketides. Plant J 54:750–762
Acknowledgments
We are grateful for the support of the National Natural Science Foundation of China (No. 30925038 and 31170280).
Conflict of interest
The authors declare that there is no conflict of interest in the present investigation.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by Zeng-Yu Wang.
H.-N. Yu and L. Wang are contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, HN., Wang, L., Sun, B. et al. Functional characterization of a chalcone synthase from the liverwort Plagiochasma appendiculatum . Plant Cell Rep 34, 233–245 (2015). https://doi.org/10.1007/s00299-014-1702-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-014-1702-8