Abstract
Co-transcriptional histone modifications are a ubiquitous feature of RNA polymerase II (RNAPII) transcription, with profound but incompletely understood effects on gene expression. Unlike the covalent marks found at promoters, which are thought to be instructive for transcriptional activation, these modifications occur in gene bodies as a result of transcription, which has made elucidation of their functions challenging. Here we review recent insights into the regulation and roles of two such modifications: monoubiquitylation of histone H2B at lysine 120 (H2Bub1) and methylation of histone H3 at lysine 36 (H3K36me). Both H2Bub1 and H3K36me are enriched in the coding regions of transcribed genes, with highly overlapping distributions, but they were thought to work largely independently. We highlight our recent demonstration that, as was previously shown for H3K36me, H2Bub1 signals to the histone deacetylase (HDAC) complex Rpd3S/Clr6-CII, and that Rpd3S/Clr6-CII and H2Bub1 function in the same pathway to repress aberrant antisense transcription initiating within gene coding regions. Moreover, both of these histone modification pathways are influenced by protein phosphorylation catalyzed by the cyclin-dependent kinases (CDKs) that regulate RNAPII elongation, chiefly Cdk9. Therefore, H2Bub1 and H3K36me are more tightly linked than previously thought, sharing both upstream regulatory inputs and downstream effectors. Moreover, these newfound connections suggest extensive, bidirectional signaling between RNAPII elongation complexes and chromatin-modifying enzymes, which helps to determine transcriptional outputs and should be a focus for future investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Post-translational modification of histones has long been associated with regulation of RNA polymerase II (RNAPII) transcription. Early, ground-breaking studies identified known transcriptional co-activators or co-repressors as histone acetyltransferases (HATs) or deacetylases (HDACs), respectively (Brownell et al. 1996; Taunton et al. 1996). These factors are recruited at or upstream of gene promoters by DNA-binding transcriptional activators or repressors, and so were quickly established as being instructive for gene expression, such that the acetylation state of promoter-proximal nucleosomes helped determine whether a particular gene was transcribed or not (Kuo et al. 1996, 2000; Kadosh and Struhl 1997; Rundlett et al. 1998). This conceptual framework is still in place today [although it is apparent that non-histone proteins are also important enzymatic targets of “HATs” and “HDACs” (Zhang and Dent 2005)]. This simple picture has been complicated, however, by the discovery of a diverse array of additional histone modifications and the genome-wide profiling of modification patterns by chromatin immunoprecipitation (ChIP). In addition to acting prior to transcriptional activation, many covalent marks are placed on gene-body nucleosomes during transcription, by chromatin-modifying enzymes recruited to elongating RNAPII complexes, and play important but still emerging roles in the transcription cycle. Here we review recent studies that illuminate functions and regulation of these co-transcriptional, covalent histone modifications.
One outcome of these studies is the discovery of a highly conserved, stereotypical pattern of histone modifications within the coding regions of RNAPII transcription units (Li et al. 2007a; Rando 2007; Tanny 2014). A metagene diagram depicting the best-characterized features of the histone modification landscape on typical protein-coding genes is shown in Fig. 1A. Acetylation of multiple histone lysine residues is most highly enriched on nucleosomes immediately surrounding the transcription start site (TSS), consistent with a generally permissive role in transcription initiation. In contrast, site-specific methylation of histone H3 and monoubiquitylation of histone H2B (H2Bub1) are highly enriched within the coding region. Rather than being simply permissive or non-permissive for transcription, these modifications are dependent on ongoing transcription. Moreover, recent evidence indicates that the majority of histone acetylation likewise occurs as a consequence of transcription (Martin et al. 2021). These findings have given rise to a new conceptual framework for understanding the genesis and function of histone modification. The importance of these questions is underscored by the numerous connections that have been uncovered between the relevant histone-modifying enzymes and human disease, most notably cancer (Marsh and Dickson 2019; McDaniel and Strahl 2017).
Regulatory crosstalk between the different modifications is critical for shaping this pattern. For example, H2Bub1 is required for histone H3 lysine 79 methylation (H3K79me) and, to some extent, histone H3 lysine 4 methylation (H3K4me) (Chandrasekharan et al. 2010). Furthermore, H3K4me and H3K36me both regulate histone acetylation in gene coding regions (Buratowski and Kim 2010). Thus, patterning of histone modifications by RNAPII transcription can be a multi-step or combinatorial process, which may serve to expand or fine-tune the repertoire of functional outcomes. Here we focus on new insights from our labs and others into regulatory crosstalk between H3K36me and H2Bub1, two modifications that had been thought to operate largely independently.
Regulation and function of H3K36 methylation
Co-transcriptional deposition of H3K36me is triggered by phosphorylation of the carboxy-terminal domain (CTD) of the RNAPII large subunit Rpb1. The RNAPII CTD is composed of multiple repeats of a heptad motif with the consensus amino-acid sequence YSPTSPS (Jeronimo et al. 2016). It is an important binding target for many transcriptional regulators, and its interactions are regulated by phosphorylation of specific residues within the repeat. For example, phosphorylation of the serine 5 position (pSer5) peaks just downstream of the TSS, whereas phosphorylation of serine 2 (pSer2) is low at the TSS, increases in relative abundance with RNAPII passage through the coding region, and peaks downstream of the polyadenylation signal (PAS) (Fig. 1B). A family of transcription-associated cyclin-dependent kinases (CDKs) catalyze these phosphorylations during transcription. Cdk7 and Cdk9 are primarily implicated in placing pSer5, whereas Cdk9, Cdk12, and Cdk13 phosphorylate the Ser2 position (Sanso and Fisher 2013). The H3K36 methyltransferase Set2 associates directly with the RNAPII CTD phosphorylated at both serine residues (Kizer et al. 2005; Li et al. 2005). In vivo, the pSer2 form of the CTD is specifically required for H3K36 trimethylation (Youdell et al. 2008; Yoh et al. 2008). In yeast, all H3K36me is Set2-dependent; both the di-methyl and tri-methyl forms are enriched within gene coding regions (Fig. 1A) and have overlapping functions (Youdell et al. 2008; Li et al. 2009; DiFiore et al. 2020). Greater functional divergence between the di- and tri-methyl forms has been noted in mammalian cells, in which H3K36me2 is catalyzed by additional methyltransferases (such as NSD1 and NSD2) and can occur independently of RNAPII elongation (Kuo et al. 2011; Weinberg et al. 2019).
H3K36me acts by engaging factors that contain specific “reader” domains (McDaniel and Strahl 2017). H3K36me-specific readers harbor well-characterized methyl-lysine binding domains including PHD fingers, chromodomains, and PWWP domains (Arrowsmith and Schapira 2019). This last class of methyl-lysine binding domain is particularly enriched in H3K36me-binding factors and may be dedicated to H3K36me recognition. H3K36me function has been most extensively studied in budding yeast. Genetic and biochemical analyses have demonstrated that H3K36me is required for the function of Rpd3S, an HDAC complex that acts to maintain low levels of histone acetylation in the coding regions of active genes (Keogh et al. 2005; Carrozza et al. 2005). H3K36me is engaged by the chromodomain of the Eaf3 subunit of Rpd3S (Ruan et al. 2015). Mutations in Rpd3S components or in Set2 lead to hyperacetylation of histones in gene coding regions and also cause transcription to initiate inappropriately from these locations. These aberrant transcription events can result in ectopic sense transcripts that are truncated near their 5’ ends, or in antisense transcripts (Venkatesh et al. 2016; Li et al. 2007b). This phenotype, of aberrant intragenic transcription initiation, is characteristic of mutations thought to perturb chromatin structure during transcription, and was first described in connection with Spt6, an elongation factor and histone chaperone (Kaplan et al. 2003).
In addition to promoting Rpd3S function in gene coding regions, H3K36me blocks co-transcriptional histone exchange by regulating the histone chaperone Asf1 or the Isw chromatin remodeling complex; the latter function is mediated by interaction with a PWWP reader domain in Ioc4 (Smolle et al. 2012; Venkatesh et al. 2012). Histone hyperacetylation correlates with increased histone exchange in the set2∆ mutant. Thus, the stabilization of transcribed chromatin by H3K36me may involve both enhanced deacetylase activity and reduced co-transcriptional histone exchange (Venkatesh et al. 2012).
Rpd3S is orthologous to the Sin3B HDAC complex in mammalian cells. A functional link to H3K36me is suggested by the fact that the MRG15 subunit of Sin3B is the ortholog of Eaf3 (Jelinic et al. 2011), although a role for the Sin3B complex in suppression of inappropriate initiation has not been established. H3K36me does participate in an analogous repressive mechanism, however, by recruiting the DNA methyltransferase DNMT3B to direct DNA CpG methylation in transcribed coding regions in embryonic stem cells (Baubec et al. 2015). Mutations that impair H3K36me binding by the DNMT3B PWWP domain lead to the accumulation of adventitious sense transcripts that initiate within gene coding regions (Neri et al. 2017).
Interestingly, Eaf3 is also a component of the HAT complex NuA4. Eaf3 forms a subassembly within this complex (with Eaf5 and the conserved Eaf7 subunit) that promotes its binding to H3K36me-containing nucleosomes (Sathianathan et al. 2016). There is also evidence that an Eaf3/5/7 complex associates with transcribed genes and regulates elongation as a module separate from the acetyltransferase (Rossetto et al. 2014). Other H3K36me-specific reader proteins also positively regulate elongation. For example, H3K36me helps to recruit the HAT complex NuA3, as well as the nucleosome disassembly factors NDF, LEDGF, and HDGF2 (all of which engage H3K36me through PWWP domains), to transcribed coding regions (Gilbert et al. 2014; Flury et al. 2017; Fei et al. 2018; LeRoy et al. 2019). Therefore, H3K36me seems to mediate a complex interplay among factors that make chromatin more or less permissive for transcription. How this balance is achieved at individual genes remains largely unknown.
An additional role for H3K36me, in co-transcriptional mRNA processing, is suggested by interaction of Eaf3 with the Prp45 subunit of the spliceosome (Leung et al. 2019). MRG15 also participates in splicing regulation in mammalian cells, although it acts through the splicing regulator PTB and not through the spliceosome directly (Luco et al. 2010).
Regulation and function of H2B mono-ubiquitylation
H2Bub1 is a dynamic marker of transcribed chromatin that is catalyzed by a complex composed of the E2 ubiquitin conjugating enzyme Rad6 and the E3 ubiquitin ligase Bre1. Rad6 and Bre1 target histone H2B on a conserved C-terminal lysine residue (corresponding to K120 in humans, K123 in S. cerevisiae, or K119 in S. pombe). It is removed during transcription by the de-ubiquitylation (DUB) module of the SAGA (Spt-Ada-Gcn5 acetyltransferase) co-activator complex, the catalytic component of which is Ubp8 (Fuchs and Oren 2014). Co-transcriptional formation of H2Bub1 depends on the activity of a Cdk9/cyclin complex, known in metazoans as positive transcription elongation factor b (P-TEFb), an essential CDK needed for rapid elongation by RNAPII (Tanny 2014; Fuchs et al. 2014; Sanso et al. 2012; Pirngruber et al. 2009). The Cdk9 substrate most clearly linked to H2Bub1 is Spt5, one subunit of a conserved, heterodimeric transcription elongation and processivity factor known in metazoans as the DRB sensitivity-inducing factor (DSIF) (Sanso et al. 2012; Mbogning et al. 2015). Cdk9 phosphorylates Spt5 on its carboxy-terminal repeats (CTRs, which may be functionally analogous to the CTD on the RNAPII large subunit); this form of Spt5 (pSpt5) peaks in abundance just downstream of the TSS, is thought to enhance RNAPII elongation rate, and is distributed throughout the coding region, as is the case for H2Bub1 (Fig. 1A,B) (Sanso et al. 2020; Parua et al. 2018; Cortazar et al. 2019). Another elongation factor, Rtf1, bridges pSpt5 and H2B ubiquitylation enzymes through physical interactions with both pSpt5 and Rad6 (Wier et al. 2013; Mayekar et al. 2013; Van Oss et al. 2016; Mbogning et al. 2013).
The action of H2Bub1 in promoting site-specific methylation of histone H3 on lysine 79, by Dot1, and on lysine 4, by COMPASS/MLL family methyltransferases, has been visualized at atomic resolution in recent cryo-EM structures. The details of these structures have been extensively reviewed elsewhere, but an important theme that emerged from this work is that H2Bub1 acts as an allosteric regulator of both methyltransferase classes, locking them in a conformation that is compatible with activity (Janna et al. 2020; Worden and Wolberger 2019). In vivo, H3K4 tri-methylation (H3K4me3) is enriched around the TSS (Fig. 1A), whereas the di-methyl (H3K4me2) form is distributed more broadly within the coding region. Mono-methylated H3K4 is largely absent from gene coding regions but is a prominent marker of enhancer regions in mammalian cells (Heintzman et al. 2007). H3K4me3 and H3K4me2 marks engage a variety of reader domains; the PHD finger is most often associated with H3K4me recognition, but chromodomains, Tudor domains, and WD40 domains can also have this property (Ruthenburg et al. 2007). H3K4me reader proteins are usually present in large transcriptional regulatory complexes implicated in activation or repression, including chromatin modifiers and general transcription factors (Vermeulen et al. 2010, 2007; Saksouk et al. 2009; Shi et al. 2006; Taverna et al. 2006). As is the case for H3K36me, H3K4me seems generally to pattern factor occupancy or function at transcribed genes, although deciphering how H3K4me-dependent interactions are coordinated at specific genes remains a work in progress. H3K4me can also function to block the association of heterochromatin proteins with chromatin, thus helping to demarcate boundaries between transcriptionally silent heterochromatin and transcriptionally active euchromatin (Ooi et al. 2007; Schuettengruber et al. 2007; Douillet et al. 2020).
In yeast, all H3K4me is catalyzed by a single COMPASS complex whose tri- and di-methyltransferase activity is strictly H2Bub1-dependent. In contrast, multiple methyltransferases contribute to H3K4me in mammalian cells; these enzymes have key roles in regulating gene expression during development (Shilatifard 2012). In these systems, the connection between H2Bub1 and H3K4me is complex, as the H3K4 methyltransferases have differing dependencies on H2Bub1 (Wu et al. 2013; Kwon et al. 2020). In ChIP-seq experiments conducted in mammalian cells, H3K4me is detected at highly expressed genes as a broad peak centered over the transcription start site (TSS), which extends into the coding region (Benayoun et al. 2014). Removal of H2Bub1 specifically decreases peak breadth, rather than peak height (Xie et al. 2017). This would be consistent with a role for H2Bub1 in stimulating H3K4me specifically during early elongation. The COMPASS complex containing the SET1A methyltransferase is required for broad H3K4me3 peaks at these genes and is strongly dependent on H2Bub1, further supporting this regulatory connection (Kwon et al. 2020; Sze et al. 2020).
H3K79 methylation is catalyzed by the Dot1 family of methyltransferases, which are universally H2Bub1-dependent. H3K79 is located on the surface of the nucleosome and, in contrast to other histone methylation marks, H3K79me is not engaged by any known histone modification reader domain. H3K79me is distributed throughout RNAPII transcription units (Fig. 1A) and, like H3K4me, it seems to act by excluding the binding of transcriptional repressors and heterochromatin proteins (Vlaming and Leeuwen 2016).
H2Bub1 also has roles that are independent of downstream histone methylation. In vitro, H2Bub1 directly stimulates a nucleosome sliding activity of the chromatin remodeling enzyme Chd1, but inhibits remodeling activity by ISWI family enzymes (Levendosky et al. 2016; Dann et al. 2017). The SWI/SNF and Ino80 chromatin remodeling complexes have also been shown to respond to H2Bub1, although whether these effects are direct or indirect is not known (Shema-Yaacoby et al. 2013; Segala et al. 2016). H2Bub1 directly influences the co-transcriptional nucleosome assembly and disassembly activities of FACT (facilitates chromatin transcription), a nucleosome reorganizing complex and chaperone for H2A-H2B dimers (Pavri et al. 2006; Murawska et al. 2020). Together, these findings suggest that H2Bub1 is deeply involved in regulating structural transitions in chromatin that accompany elongation by RNAPII; indeed, yeast mutants lacking H2Bub1 have defects in genic chromatin structure, which are not due to loss of H3K4me or H3K79me (Murawska et al. 2020; Batta et al. 2011). The proximity of the consensus H2B ubiquitylation site to the “acidic patch” on the nucleosome surface, an important interaction site for several nucleosome-binding factors (including COMPASS and Dot1), suggests a common mechanism through which H2Bub1 may modulate these interactions (Fig. 2)(Worden and Wolberger 2019; Dann et al. 2017). There is evidence for H2Bub1 roles in other aspects of RNAPII elongation, notably mRNA splicing and export, although the relevant mechanisms have not been determined (Vitaliano-Prunier et al. 2012; Moehle et al. 2012). H2Bub1 may also promote elongation as a feedback regulator of Cdk9. In both S. pombe and mammalian systems, H2Bub1 and Cdk9 regulate one another: inactivation of Cdk9 leads to diminished H2Bub1, while mutations in the H2B ubiquitylation pathway impair phosphorylation of Cdk9 substrates such as Spt5 (Sanso et al. 2012; Wu et al. 2014). How this mutual dependence influences effects of H2Bub1 on co-transcriptional chromatin transitions and gene expression remains to be determined.
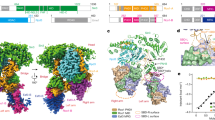
Interaction surfaces for Set2, Dot1, and COMPASS/MLL on the nucleosome. Sites of interaction (circles) and relevant histone modification sites (red dots) are highlighted on a model of the nucleosome crystal structure (PDB 1KX5). In this view, COMPASS/MLL and Set2 methylate the histone H3 monomer on the left side of the dyad axis (coloured red), whereas Dot1 methylates the histone H3 monomer on the right (coloured brown)
Evidence for crosstalk in H3K36me and H2Bub1 pattern formation
Crosstalk between H3K36me and H2Bub1 pathways would be consistent with the partly overlapping distributions of the two marks in transcribed genes (Fig. 1A). In general, the two have been considered to be functionally complementary but independent modifications acting during transcription elongation. Results of recent studies contain hints that the two pathways are in fact interconnected. The cryo-EM structures of Set2 bound to unmodified and H2Bub1-containing nucleosomes indicate that Set2 binding is positioned by H2Bub1 (Bilokapic and Halic 2019). Set2 engages the nucleosome where the DNA entering and exiting the nucleosome overlap (Fig. 2); in fact, Set2 binding displaces roughly one helical turn of DNA at one end from the histone octamer. Set2 stabilizes this partially unwrapped state through electrostatic interactions with histone H3 and with DNA. These interactions involve the catalytic SET domain of Set2 and position the H3 tail for K36 methylation. This binding mode is distinct from that observed for established H2Bub1-dependent methyltransferases Dot1 and COMPASS/MLL, which relies on the “acidic patch” region and does not impinge on histone-DNA interactions (Fig. 2). Nonetheless, in Set2-nucleosome structures that included ubiquitin attached to H2B via disulfide linkage (to mimic authentic H2Bub1), a subset of density maps detected a defined position for ubiquitin in which its C-terminal ß strand was in close proximity to the AWS (associated with SET) domain of Set2. The AWS domain is immediately N-terminal to the SET domain in H3K36 methyltransferases related to Set2. Biochemical experiments suggested this interaction could be functional, as ubiquitin attachment modestly enhanced Set2 activity towards nucleosome substrates in vitro. To date, no structure–function analysis to ascertain the importance of AWS-domain residues for the stimulatory effect of H2Bub1, or for Set2 function in vivo, has been reported, leaving the physiological relevance of the H2Bub1 effect uncertain. However, these results offer intriguing hints that H2Bub1 can influence nucleosome interactions with distinct classes of chromatin-modifying enzymes.
Signaling between H2Bub1 and H3K36me has been demonstrated to proceed in the reverse direction in the fission yeast S. pombe. That communication is mediated by the S. pombe NuA3 complex, which is a major histone H3 acetyltransferase previously implicated in transcription elongation. NuA3 is recruited to transcribed coding regions by a PWWP reader domain interaction with H3K36me (Gilbert et al. 2014; Flury et al. 2017). The key finding linking H3K36me to H2Bub1 is that NuA3 also acetylates the H2Bub1-specific E3 ligase Brl1, thereby enhancing its activity toward H2B (Flury et al. 2017). Acetylation of Brl1 was found to negatively regulate RNAi-mediated heterochromatin formation. Heterochromatin comprises constitutively repressed chromatin regions that harbor conserved molecular hallmarks: methylation of histone H3 at lysine 9, association of HP1 orthologs, and low levels of RNAPII. In S. pombe, pericentric heterochromatin formation also requires small non-coding RNAs generated by the RNAi pathway, akin to piRNA-directed silencing in the mammalian germline (Castel and Martienssen 2013). Small RNAs normally act to establish heterochromatin only within previously established heterochromatin domains; loss of Brl1 acetylation was found to relax this restriction (Flury et al. 2017). Brl1 acetylation occurred at a lysyl residue that is distant from the catalytic domain; mutation of this site decreased H2Bub1 levels in vivo by ~ twofold. It is unclear whether this modest reduction in H2Bub1, or a non-enzymatic function of Brl1 that is compromised upon loss of Brl1 acetylation, is responsible for the observed phenotype of ectopic heterochromatin formation. Further studies investigating the role of this residue in regulating Brl1 activity or protein–protein interactions will be needed to resolve this issue. Nevertheless, these data suggest that NuA3 supports a mutually reinforcing function of H3K36me and H2Bub1 during transcription elongation.
Evidence for crosstalk in H3K36me and H2Bub1 function
Our recent findings now provide evidence for a shared function of H3K36me and H2Bub1 in regulation of aberrant antisense transcription by the Rpd3S HDAC complex (Sanso et al. 2020). We identified a role for H2Bub1 in suppressing aberrant antisense transcription in S. pombe by strand-specific RNA-seq analysis. We then performed genetic epistasis analysis to relate this function to those of other known, negative regulators of antisense transcription. We focused on Set2 and Rpd3S (Clr6-CII in S. pombe) because of their previously characterized roles in antisense suppression. We also included the CHD family chromatin remodeling factor Hrp3 because it has been implicated in antisense regulation (through a pathway distinct from Clr6-CII) and suggested as a potential target of H2Bub1 (Levendosky et al. 2016; Hennig et al. 2012; Pointner et al. 2012; Shim et al. 2012). The single mutants htb1-K119R (lacking the ubiquitylation site on H2B), set2∆, cph1∆ (lacking Cph1, a Clr6-CII subunit) or hrp3∆ each displayed increased antisense transcription, as measured by strand-specific RT-qPCR at candidate genes. Combining htb1-K119R with each of the other mutations led to two different phenotypic outcomes with respect to antisense transcript levels. In the htb1-K119R hrp3∆ double mutant, there was an additive effect of the two mutations, arguing that the two individual mutations affect antisense transcription through different pathways. In contrast, antisense levels in the set2∆ htb1K119R and cph1∆ htb1-K119R double mutants were similar to those in the single set2∆ and cph1∆ mutants, indicating epistasis and suggesting that H2Bub1 regulates antisense transcription through the same pathway as Set2 or Clr6-CII. ChIP-seq analysis in the htb1-K119R mutant indicated that association of the Clr6-CII complex with transcribed coding regions was impaired genome-wide (by about twofold) in the absence of H2Bub1. This effect was unlikely to be an indirect result of changes in H3K36me, because no effects on Set2 occupancy or H3K36me levels were detected by ChIP-qPCR at select target genes. These results suggest that H2Bub1, like H3K36me, promotes the function of the Rpd3S/Clr6-CII complex.
The mechanistic linkage between H2Bub1 and Rpd3S/Clr6-CII has yet to be elucidated. Although loss of H2Bub1 reduced Clr6-CII occupancy on chromatin, we did not find evidence for increased histone H3 acetylation levels on the candidate genes we examined. This may reflect acetylation-site specificity or gene specificity of the H2Bub1 effect. When combined with inhibition of Cdk9, however, loss of H2Bub1 led to more dramatic, genome-wide decreases in Clr6-CII recruitment to gene bodies, relative to RNAPII occupancy, and synergistically increased levels of histone H3 acetylation on select genes we analyzed. These interactions roughly mirrored the combinatorial effects of H2Bub1 loss and Cdk9 inhibition on antisense suppression: (1) increases in the numbers of genes affected (i.e., those with increased antisense transcription) and (2) enhanced severity of the antisense de-repression detected at individual loci.
Alternatively, H2Bub1 may play a role in augmenting a non-enzymatic, chromatin-stabilizing role for the HDAC complex in antisense suppression (Chen et al. 2012). Further investigation will be required to test these possibilities. These results present an interesting contrast with H3K36me, which was found to promote Rpd3S HDAC activity while having little impact on its association with chromatin (Drouin et al. 2010; Govind et al. 2010). Thus, the two modifications may act on a shared target in different ways. Detailed biochemical and structural studies have shown that H3K36me affects Rpd3S activity directly; whether this is true for H2Bub1 remains to be determined. Intriguingly, the mammalian Sin3B subunit MRG15 is reported to bind directly to ubiquitylated histones (Wu et al. 2011). Our epistasis results are consistent with H2Bub1 and H3K36me both acting to suppress antisense transcription through Clr6-CII, but we cannot exclude the existence of an alternative pathway. Previous work in S. pombe suggests that functions of Set2 and Clr6-CII in antisense suppression are overlapping but distinct (Nicolas et al. 2007). The nature of the putative Set2-dependent but Clr6-CII-independent pathway is not known, but links between Set2 and histone-exchange mechanisms or acetyltransferases point to other, potentially shared functions of H3K36me and H2Bub1 in antisense regulation.
Another recent study also documented a role for H2Bub1 in antisense suppression (Murawska et al. 2020). This work focused on a potential role for H2Bub1 in regulating the FACT nucleosome reorganizing complex, which had emerged from previous work in budding yeast and in vitro. FACT and H2Bub1 were required for antisense suppression at different genomic loci, but a dual loss-of-function mutant had an effect similar to that of FACT ablation alone. It was therefore suggested that H2Bub1 loss leads to increased antisense transcription through an aberrant activity of FACT. However, correction of aberrant FACT activity could also reflect the effect of H2Bub1 on Rpd3S/Clr6-CII, as genetic interactions in budding yeast suggest that H3K36me and the Rpd3S complex act in opposition to FACT (Biswas et al. 2006; Stevens et al. 2011). Deciphering how Rpd3S/Clr6-CII and FACT mediate the interplay between H2Bub1 and H3K36me requires further investigation.
H2Bub1 and H3K36me both affect regional gene silencing in S. pombe
H2Bub1 and H3K36me are abundant in the actively transcribed, euchromatic regions of the genome and are excluded from or depleted in pericentric and telomeric heterochromatin. However, both H2Bub1 and H3K36me regulate heterochromatin, likely through indirect but conserved mechanisms that are beginning to emerge. The reduced NuA3 localization to gene coding regions caused by loss of Set2 activity in S. pombe allows the complex to bind to chromatin promiscuously, leading to some association with heterochromatin. This leads to inappropriate acetylation (of unknown targets) and destabilizes the transcriptionally repressed state (Flury et al. 2017; Georgescu et al. 2020). A similar sequestration mechanism involving H3K36me has been shown to regulate heterochromatin formation during development in the nematode C. elegans (Cabianca et al. 2019).
In S. pombe, Set2 is also important for repression of highly condensed, subtelomeric chromatin domains (termed “knobs”) that are ~ 50 kilobases away from the telomere and distinct from telomeric heterochromatin (Matsuda et al. 2015). These domains are not associated with H3K9 methylation or HP1 and instead require the shugoshin ortholog Sgo2 for their establishment or maintenance (Matsuda et al. 2015; Tashiro et al. 2016). Shugoshin normally functions to protect sister chromatid cohesion at centromeres; its function at subtelomeres is not yet clear. Given that H3K36me is not abundant at subtelomeric chromatin it is likely to exert its function indirectly. Interestingly, sequestering of NuA3 only partially accounts for Set2-dependent repression in these regions (Georgescu et al. 2020). This suggests that H3K36me may sequester additional factors to prevent their association with subtelomeric chromatin domains.
Loss of H2Bub1 enhances transcriptional repression in and around S. pombe heterochromatin. This is accompanied by an increase in H3K9 trimethylation within heterochromatin domains, as well as invasion of H3K9me from pericentric regions into the central core of the centromere (Zofall and Grewal 2007; Sadeghi et al. 2014). We and Murawska et al. both found that subtelomeric “knob” regions were also hyper-repressed in H2Bub1 mutants, despite the fact that these regions harbor relatively low levels of H2Bub1 in wild-type cells (Fig. 3A)(Murawska et al. 2020). This hyper-repression was not due to inappropriate spread of telomeric heterochromatin, as it was maintained in a double mutant with sir2∆ (Fig. 3B). By analogy to the sequestration function of H3K36me, we hypothesize a related function for H2Bub1 that, when disrupted, leads to aberrant action of a repressive factor in heterochromatin or subtelomeric regions. Evidence in support of this type of mechanism includes the finding that reduced FACT function partially alleviates hyper-repression at subtelomeres. We found that the cph1∆ and alp13∆ mutations that impair Clr6-CII function (Alp13 is the ortholog of Eaf3 and MRG15) have a similar effect—relief of hyper-repression—at the candidate subtelomeric genes that we tested (Fig. 3C). However, there is as yet no clear-cut evidence from ChIP experiments that loss of H2Bub1 leads to redistribution of Clr6-CII or FACT, as is clearly the case for NuA3 when H3K36me is removed. Therefore, another factor regulated by H2Bub1 may be involved in hyper-repression at heterochromatin and subtelomeric regions.
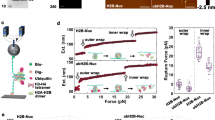
H2Bub1 preferentially activates genes in subtelomeric “knob” domains. A Fraction of H2Bub1-activated genes (defined by RNA-seq in ref 49) within the indicated genomic intervals. Significant enrichment within subtelomeric regions was assessed by hypergeometric test. B Expression of the knob gene aes1+ or the heterochomatic gene tlh1+ was determined by RT-qPCR as in ref 49 in the indicated strains; values in the wild-type strain were set to 1. Asterisks denote significant differences from wild-type (n = 3; unpaired t test with Bonferroni correction; p < 0.05). C Expression of the indicated knob genes was determined by RT-qPCR in the indicated strains and normalized to act1+ expression as in ref 49. Asterisks denote significant differences from wild-type (n = 4; unpaired t-test with Bonferroni correction; p < 0.05). Primer sequences are available upon request
Perspectives
We are still far from a complete understanding of the interplay between chromatin structure and the transcription machinery. A defining feature of both H2Bub1 and H3K36me is that they are placed predominantly or exclusively during the act of transcription and are thus stringently regulated by the factors that govern RNAPII elongation. Indeed, the enzymes most intimately linked to the relevant histone-modifying activities are the CDKs that phosphorylate Rpb1, Spt5 and other components of the transcriptional machinery to coordinate the RNAPII cycle. Cdk9, the rate-limiting kinase for RNAPII elongation in metazoans and fission yeast (Jonkers et al. 2014; Booth et al. 2018), appears to play a central role in regulating H2Bub1 (and H3K4me) and H3K36me in S. pombe, but other transcriptional CDKs clearly contribute (Sanso et al. 2012; Mbogning et al. 2015). These functions of Cdk9 in regulating chromatin modification have been conserved in metazoans, but prominent roles in these pathways have also been ascribed to Cdk7, the kinase component of the initiation factor TFIIH, which in mammalian cells is also the activating kinase for Cdk9, Cdk12 and Cdk13 (Pirngruber et al. 2009; Larochelle et al. 2012; Ebmeier et al. 2017; Rimel et al. 2020). Just as H2Bub1 and H3K36me appear to influence each other, signaling between the CDKs and histone modifications such as H2Bub1 and H3K36me is bidirectional, with examples of crosstalk and feedback that have not been fully explained. For example, despite the mutual dependence of H2Bub1 and pSpt5 in vivo, loss-of-function mutations in the H2Bub1 and Cdk9 pathways can have opposing, additive or synergistic effects on downstream events including antisense transcription (Sanso et al. 2012, 2020). Moreover, the two pathways collaborate to govern recruitment and function of Clr6-CII genome-wide, but also appear to work independently to suppress unscheduled antisense transcription at specific gene sets, possibly through Clr6-CII-independent mechanisms.
Here we have summarized evidence for regulatory and functional crosstalk between H2Bub1 and H3K36me that highlights the extent to which events during transcription are interconnected. Signaling between H2Bub1 and H3K36me is apparently unique in that it is bidirectional, suggesting a mutually reinforcing relationship. Determining the mechanistic basis for that relationship will require addressing several key questions: (1) What is the significance of non-histone acetylation by NuA3 and perhaps other acetyltransferase complexes? (2) Do H2Bub1 and H3K36me act in combination to engage shared targets such as Rpd3S/Clr6-CII? (3) How are the sequestering functions of these modifications coordinated to balance their regulatory effects at transcribed genes and at heterochromatin? Answers to these questions will have broad implications for understanding the regulation and function of co-transcriptional histone modification. Further investigation is also needed to unravel the communication between the RNAPII elongation complex and histone modification pathways, which appears to be conserved in evolution and relevant to human disease processes; the CDKs implicated in this signaling have recently emerged as potential anti-cancer drug targets (Parua and Fisher 2020).
References
Arrowsmith CH, Schapira M (2019) Targeting non-bromodomain chromatin readers. Nat Struct Mol Biol 26:863–869
Batta K, Zhang Z, Yen K, Goffman DB, Pugh BF (2011) Genome-wide function of H2B ubiquitylation in promoter and genic regions. Genes Dev 25:2254–2265
Baubec T, Colombo DF, Wirbelauer C, Schmidt J, Burger L, Krebs AR, Akalin A, Schubeler D (2015) Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature 520:243–247
Benayoun BA, Pollina EA, Ucar D, Mahmoudi S, Karra K, Wong ED, Devarajan K, Daugherty AC, Kundaje AB, Mancini E, Hitz BC, Gupta R, Rando TA, Baker JC, Snyder MP, Cherry JM, Brunet A (2014) H3K4me3 breadth is linked to cell identity and transcriptional consistency. Cell 158:673–688
Bilokapic S, Halic M (2019) Nucleosome and ubiquitin position Set2 to methylate H3K36. Nat Commun 10:3795
Biswas D, Dutta-Biswas R, Mitra D, Shibata Y, Strahl BD, Formosa T, Stillman DJ (2006) Opposing roles for Set2 and yFACT in regulating TBP binding at promoters. Embo J 25:4479–4489
Booth GT, Parua PK, Sanso M, Fisher RP, Lis JT (2018) Cdk9 regulates a promoter-proximal checkpoint to modulate RNA polymerase II elongation rate in fission yeast. Nat Commun 9:543
Brownell JE, Zhou J, Ranalli T, Kobayashi R, Edmondson DG, Roth SY, Allis CD (1996) Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 84:843–851
Buratowski S, Kim T (2010) The role of cotranscriptional histone methylations. Cold Spring Harb Symp Quant Biol 75:95–102
Cabianca DS, Munoz-Jimenez C, Kalck V, Gaidatzis D, Padeken J, Seeber A, Askjaer P, Gasser SM (2019) Active chromatin marks drive spatial sequestration of heterochromatin in C. elegans nuclei. Nature 569:734–739
Carrozza MJ, Li B, Florens L, Suganuma T, Swanson SK, Lee KK, Shia WJ, Anderson S, Yates J, Washburn MP, Workman JL (2005) Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123:581–592
Castel SE, Martienssen RA (2013) RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Genet 14:100–112
Chandrasekharan MB, Huang F, Sun ZW (2010) Histone H2B ubiquitination and beyond: regulation of nucleosome stability, chromatin dynamics and the trans-histone H3 methylation. Epigenetics 5:460–468
Chen XF, Kuryan B, Kitada T, Tran N, Li JY, Kurdistani S, Grunstein M, Li B, Carey M (2012) The Rpd3 core complex is a chromatin stabilization module. Curr Biol 22:56–63
Cortazar MA, Sheridan RM, Erickson B, Fong N, Glover-Cutter K, Brannan K, Bentley DL (2019) Control of RNA pol II speed by PNUTS-PP1 and Spt5 dephosphorylation facilitates termination by a “sitting duck torpedo” mechanism. Mol Cell 76:896–908
Dann GP, Liszczak GP, Bagert JD, Muller MM, Nguyen UTT, Wojcik F, Brown ZZ, Bos J, Panchenko T, Pihl R, Pollock SB, Diehl KL, Allis CD, Muir TW (2017) ISWI chromatin remodellers sense nucleosome modifications to determine substrate preference. Nature 548:607–611
DiFiore JV, Ptacek TS, Wang Y, Li B, Simon JM, Strahl BD (2020) Unique and shared roles for histone H3K36 methylation states in transcription regulation functions. Cell Rep 31:107751
Douillet D, Sze CC, Ryan C, Piunti A, Shah AP, Ugarenko M, Marshall SA, Rendleman EJ, Zha D, Helmin KA, Zhao Z, Cao K, Morgan MA, Singer BD, Bartom ET, Smith ER, Shilatifard A (2020) Uncoupling histone H3K4 trimethylation from developmental gene expression via an equilibrium of COMPASS, polycomb and DNA methylation. Nat Genet 52:615–625
Drouin S, Laramee L, Jacques PE, Forest A, Bergeron M, Robert F (2010) DSIF and RNA polymerase II CTD phosphorylation coordinate the recruitment of Rpd3S to actively transcribed genes. PLoS Genet 6:e1001173
Ebmeier CC, Erickson B, Allen BL, Allen MA, Kim H, Fong N, Jacobsen JR, Liang K, Shilatifard A, Dowell RD, Old WM, Bentley DL, Taatjes DJ (2017) Human TFIIH kinase CDK7 regulates transcription-associated chromatin modifications. Cell Rep 20:1173–1186
Fei J, Ishii H, Hoeksema MA, Meitinger F, Kassavetis GA, Glass CK, Ren B, Kadonaga JT (2018) NDF, a nucleosome-destabilizing factor that facilitates transcription through nucleosomes. Genes Dev 32:682–694
Flury V, Georgescu PR, Iesmantavicius V, Shimada Y, Kuzdere T, Braun S, Buhler M (2017) The histone acetyltransferase Mst2 protects active chromatin from epigenetic silencing by acetylating the ubiquitin ligase Brl1. Mol Cell 67:294–307
Fuchs G, Oren M (2014) Writing and reading H2B monoubiquitylation. Biochim Biophys Acta 1839:694–701
Fuchs G, Hollander D, Voichek Y, Ast G, Oren M (2014) Cotranscriptional histone H2B monoubiquitylation is tightly coupled with RNA polymerase II elongation rate. Genome Res 24:1572–1583
Georgescu PR, Capella M, Fischer-Burkart S, Braun S (2020) The euchromatic histone mark H3K36me3 preserves heterochromatin through sequestration of an acetyltransferase complex in fission yeast. Microb Cell 7:80–92
Gilbert TM, McDaniel SL, Byrum SD, Cades JA, Dancy BC, Wade H, Tackett AJ, Strahl BD, Taverna SD (2014) A PWWP domain-containing protein targets the NuA3 acetyltransferase complex via histone H3 lysine 36 trimethylation to coordinate transcriptional elongation at coding regions. Mol Cell Proteomics 13:2883–2895
Govind CK, Qiu H, Ginsburg DS, Ruan C, Hofmeyer K, Hu C, Swaminathan V, Workman JL, Li B, Hinnebusch AG (2010) Phosphorylated Pol II CTD recruits multiple HDACs, including Rpd3C(S), for methylation-dependent deacetylation of ORF nucleosomes. Mol Cell 39:234–246
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, Wang W, Weng Z, Green RD, Crawford GE, Ren B (2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet 39:311–318
Hennig BP, Bendrin K, Zhou Y, Fischer T (2012) Chd1 chromatin remodelers maintain nucleosome organization and repress cryptic transcription. EMBO Rep 13:997–1003
Janna A, Davarinejad H, Joshi M, Couture JF (2020) Structural paradigms in the recognition of the nucleosome core particle by histone lysine methyltransferases. Front Cell Dev Biol 8:600
Jelinic P, Pellegrino J, David G (2011) A novel mammalian complex containing Sin3B mitigates histone acetylation and RNA polymerase II progression within transcribed loci. Mol Cell Biol 31:54–62
Jeronimo C, Collin P, Robert F (2016) The RNA polymerase II CTD: the increasing complexity of a low-complexity protein domain. J Mol Biol 428:2607–2622
Jonkers I, Kwak H, Lis JT (2014) Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. Elife 3:e02407
Kadosh D, Struhl K (1997) Repression by Ume6 involves recruitment of a complex containing Sin3 corepressor and Rpd3 histone deacetylase to target promoters. Cell 89:365–371
Kaplan CD, Laprade L, Winston F (2003) Transcription elongation factors repress transcription initiation from cryptic sites. Science 301:1096–1099
Keogh MC, Kurdistani SK, Morris SA, Ahn SH, Podolny V, Collins SR, Schuldiner M, Chin K, Punna T, Thompson NJ, Boone C, Emili A, Weissman JS, Hughes TR, Strahl BD, Grunstein M, Greenblatt JF, Buratowski S, Krogan NJ (2005) Cotranscriptional set2 methylation of histone H3 lysine 36 recruits a repressive Rpd3 complex. Cell 123:593–605
Kizer KO, Phatnani HP, Shibata Y, Hall H, Greenleaf AL, Strahl BD (2005) A novel domain in Set2 mediates RNA polymerase II interaction and couples histone H3 K36 methylation with transcript elongation. Mol Cell Biol 25:3305–3316
Kuo MH, Brownell JE, Sobel RE, Ranalli TA, Cook RG, Edmondson DG, Roth SY, Allis CD (1996) Transcription-linked acetylation by Gcn5p of histones H3 and H4 at specific lysines. Nature 383:269–272
Kuo MH, vom Baur E, Struhl K, Allis CD (2000) Gcn4 activator targets Gcn5 histone acetyltransferase to specific promoters independently of transcription. Mol Cell 6:1309–1320
Kuo AJ, Cheung P, Chen K, Zee BM, Kioi M, Lauring J, Xi Y, Park BH, Shi X, Garcia BA, Li W, Gozani O (2011) NSD2 links dimethylation of histone H3 at lysine 36 to oncogenic programming. Mol Cell 44:609–620
Kwon M, Park K, Hyun K, Lee JH, Zhou L, Cho YW, Ge K, Skalnik DG, Muir TW, Kim J (2020) H2B ubiquitylation enhances H3K4 methylation activities of human KMT2 family complexes. Nucleic Acids Res 48:5442–5456
Larochelle S, Amat R, Glover-Cutter K, Sanso M, Zhang C, Allen JJ, Shokat KM, Bentley DL, Fisher RP (2012) Cyclin-dependent kinase control of the initiation-to-elongation switch of RNA polymerase II. Nat Struct Mol Biol 19:1108–1115
LeRoy G, Oksuz O, Descostes N, Aoi Y, Ganai RA, Kara HO, Yu JR, Lee CH, Stafford J, Shilatifard A, Reinberg D (2019) LEDGF and HDGF2 relieve the nucleosome-induced barrier to transcription in differentiated cells. Sci Adv 5:eaay3068
Leung CS, Douglass SM, Morselli M, Obusan MB, Pavlyukov MS, Pellegrini M, Johnson TL (2019) H3K36 methylation and the chromodomain protein Eaf3 are required for proper cotranscriptional spliceosome assembly. Cell Rep 27:3760–3769
Levendosky RF, Sabantsev A, Deindl S, Bowman GD (2016) The Chd1 chromatin remodeler shifts hexasomes unidirectionally. Elife. https://doi.org/10.7554/eLife.21356
Li M, Phatnani HP, Guan Z, Sage H, Greenleaf AL, Zhou P (2005) Solution structure of the Set2-Rpb1 interacting domain of human Set2 and its interaction with the hyperphosphorylated C-terminal domain of Rpb1. Proc Natl Acad Sci USA 102:17636–17641
Li B, Carey M, Workman JL (2007a) The role of chromatin during transcription. Cell 128:707–719
Li B, Gogol M, Carey M, Pattenden SG, Seidel C, Workman JL (2007b) Infrequently transcribed long genes depend on the Set2/Rpd3S pathway for accurate transcription. Genes Dev 21:1422–1430
Li B, Jackson J, Simon MD, Fleharty B, Gogol M, Seidel C, Workman JL, Shilatifard A (2009) Histone H3 lysine 36 dimethylation (H3K36me2) is sufficient to recruit the Rpd3s histone deacetylase complex and to repress spurious transcription. J Biol Chem 284:7970–7976
Luco RF, Pan Q, Tominaga K, Blencowe BJ, Pereira-Smith OM, Misteli T (2010) Regulation of alternative splicing by histone modifications. Science 327:996–1000
Marsh DJ, Dickson KA (2019) Writing histone monoubiquitination in human malignancy-the role of RING finger E3 ubiquitin ligases. Genes (basel) 10:67
Martin BJE, Brind’Amour J, Kuzmin A, Jensen KN, Liu ZC, Lorincz M, Howe LJ (2021) Transcription shapes genome-wide histone acetylation patterns. Nat Commun 12:210
Matsuda A, Chikashige Y, Ding DQ, Ohtsuki C, Mori C, Asakawa H, Kimura H, Haraguchi T, Hiraoka Y (2015) Highly condensed chromatins are formed adjacent to subtelomeric and decondensed silent chromatin in fission yeast. Nat Commun 6:7753
Mayekar MK, Gardner RG, Arndt KM (2013) The recruitment of the Saccharomyces cerevisiae Paf1 complex to active genes requires a domain of Rtf1 that directly interacts with the Spt4-Spt5 complex. Mol Cell Biol 33:3259–3273
Mbogning J, Nagy S, Page V, Schwer B, Shuman S, Fisher RP, Tanny JC (2013) The PAF complex and Prf1/Rtf1 delineate distinct Cdk9-dependent pathways regulating transcription elongation in fission yeast. PLoS Genet 9:e1004029
Mbogning J, Page V, Burston J, Schwenger E, Fisher RP, Schwer B, Shuman S, Tanny JC (2015) Functional interaction of Rpb1 and Spt5 C-terminal domains in co-transcriptional histone modification. Nucleic Acids Res 43:9766–9775
McDaniel SL, Strahl BD (2017) Shaping the cellular landscape with Set2/SETD2 methylation. Cell Mol Life Sci 74:3317–3334
Moehle EA, Ryan CJ, Krogan NJ, Kress TL, Guthrie C (2012) The yeast SR-like protein Npl3 links chromatin modification to mRNA processing. PLoS Genet 8:e1003101
Murawska M, Schauer T, Matsuda A, Wilson MD, Pysik T, Wojcik F, Muir TW, Hiraoka Y, Straub T, Ladurner AG (2020) The chaperone FACT and histone H2B ubiquitination maintain S. pombe genome architecture through genic and subtelomeric functions. Mol Cell 77:501–513
Neri F, Rapelli S, Krepelova A, Incarnato D, Parlato C, Basile G, Maldotti M, Anselmi F, Oliviero S (2017) Intragenic DNA methylation prevents spurious transcription initiation. Nature 543:72–77
Nicolas E, Yamada T, Cam HP, Fitzgerald PC, Kobayashi R, Grewal SI (2007) Distinct roles of HDAC complexes in promoter silencing, antisense suppression and DNA damage protection. Nat Struct Mol Biol 14:372–380
Ooi SK, Qiu C, Bernstein E, Li K, Jia D, Yang Z, Erdjument-Bromage H, Tempst P, Lin SP, Allis CD, Cheng X, Bestor TH (2007) DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448:714–717
Parua PK, Fisher RP (2020) Dissecting the Pol II transcription cycle and derailing cancer with CDK inhibitors. Nat Chem Biol 16:716–724
Parua PK, Booth GT, Sanso M, Benjamin B, Tanny JC, Lis JT, Fisher RP (2018) A Cdk9-PP1 switch regulates the elongation-termination transition of RNA polymerase II. Nature 558:460–464
Pavri R, Zhu B, Li G, Trojer P, Mandal S, Shilatifard A, Reinberg D (2006) Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125:703–717
Pirngruber J, Shchebet A, Schreiber L, Shema E, Minsky N, Chapman RD, Eick D, Aylon Y, Oren M, Johnsen SA (2009) CDK9 directs H2B monoubiquitination and controls replication-dependent histone mRNA 3’-end processing. EMBO Rep 10:894–900
Pointner J, Persson J, Prasad P, Norman-Axelsson U, Stralfors A, Khorosjutina O, Krietenstein N, Svensson JP, Ekwall K, Korber P (2012) CHD1 remodelers regulate nucleosome spacing in vitro and align nucleosomal arrays over gene coding regions in S. pombe. EMBO J 31:4388–4403
Rando OJ (2007) Global patterns of histone modifications. Curr Opin Genet Dev 17:94–99
Rimel JK, Poss ZC, Erickson B, Maas ZL, Ebmeier CC, Johnson JL, Decker TM, Yaron TM, Bradley MJ, Hamman KB, Hu S, Malojcic G, Marineau JJ, White PW, Brault M, Tao L, DeRoy P, Clavette C, Nayak S, Damon LJ, Kaltheuner IH, Bunch H, Cantley LC, Geyer M, Iwasa J, Dowell RD, Bentley DL, Old WM, Taatjes DJ (2020) Selective inhibition of CDK7 reveals high-confidence targets and new models for TFIIH function in transcription. Genes Dev 34:1452–1473
Rossetto D, Cramet M, Wang AY, Steunou AL, Lacoste N, Schulze JM, Cote V, Monnet-Saksouk J, Piquet S, Nourani A, Kobor MS, Cote J (2014) Eaf5/7/3 form a functionally independent NuA4 submodule linked to RNA polymerase II-coupled nucleosome recycling. EMBO J 33:1397–1415
Ruan C, Lee CH, Cui H, Li S, Li B (2015) Nucleosome contact triggers conformational changes of Rpd3S driving high-affinity H3K36me nucleosome engagement. Cell Rep 10:204–215
Rundlett SE, Carmen AA, Suka N, Turner BM, Grunstein M (1998) Transcriptional repression by UME6 involves deacetylation of lysine 5 of histone H4 by RPD3. Nature 392:831–835
Ruthenburg AJ, Allis CD, Wysocka J (2007) Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol Cell 25:15–30
Sadeghi L, Siggens L, Svensson JP, Ekwall K (2014) Centromeric histone H2B monoubiquitination promotes noncoding transcription and chromatin integrity. Nat Struct Mol Biol 21:236–243
Saksouk N, Avvakumov N, Champagne KS, Hung T, Doyon Y, Cayrou C, Paquet E, Ullah M, Landry AJ, Cote V, Yang XJ, Gozani O, Kutateladze TG, Cote J (2009) HBO1 HAT complexes target chromatin throughout gene coding regions via multiple PHD finger interactions with histone H3 tail. Mol Cell 33:257–265
Sanso M, Fisher RP (2013) Pause, play, repeat: CDKs push RNAP II’s buttons. Transcription 4:146–152
Sanso M, Lee KM, Viladevall L, Jacques PE, Page V, Nagy S, Racine A, St Amour CV, Zhang C, Shokat KM, Schwer B, Robert F, Fisher RP, Tanny JC (2012) A positive feedback loop links opposing functions of P-TEFb/Cdk9 and histone H2B ubiquitylation to regulate transcript elongation in fission yeast. PLoS Genet 8:e1002822
Sanso M, Parua PK, Pinto D, Svensson JP, Page V, Bitton DA, MacKinnon S, Garcia P, Hidalgo E, Bahler J, Tanny JC, Fisher RP (2020) Cdk9 and H2Bub1 signal to Clr6-CII/Rpd3S to suppress aberrant antisense transcription. Nucleic Acids Res 48:7154–7168
Sathianathan A, Ravichandran P, Lippi JM, Cohen L, Messina A, Shaju S, Swede MJ, Ginsburg DS (2016) The Eaf3/5/7 subcomplex stimulates NuA4 interaction with methylated histone H3 Lys-36 and RNA polymerase II. J Biol Chem 291:21195–21207
Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G (2007) Genome regulation by polycomb and trithorax proteins. Cell 128:735–745
Segala G, Bennesch MA, Pandey DP, Hulo N, Picard D (2016) Monoubiquitination of histone H2B blocks eviction of histone variant H2A.Z from inducible enhancers. Mol Cell 64:334–346
Shema-Yaacoby E, Nikolov M, Haj-Yahya M, Siman P, Allemand E, Yamaguchi Y, Muchardt C, Urlaub H, Brik A, Oren M, Fischle W (2013) Systematic identification of proteins binding to chromatin-embedded ubiquitylated H2B reveals recruitment of SWI/SNF to regulate transcription. Cell Rep 4:601–608
Shi X, Hong T, Walter KL, Ewalt M, Michishita E, Hung T, Carney D, Pena P, Lan F, Kaadige MR, Lacoste N, Cayrou C, Davrazou F, Saha A, Cairns BR, Ayer DE, Kutateladze TG, Shi Y, Cote J, Chua KF, Gozani O (2006) ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442:96–99
Shilatifard A (2012) The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem 81:65–95
Shim YS, Choi Y, Kang K, Cho K, Oh S, Lee J, Grewal SI, Lee D (2012) Hrp3 controls nucleosome positioning to suppress non-coding transcription in eu- and heterochromatin. EMBO J 31:4375–4387
Smolle M, Venkatesh S, Gogol MM, Li H, Zhang Y, Florens L, Washburn MP, Workman JL (2012) Chromatin remodelers Isw1 and Chd1 maintain chromatin structure during transcription by preventing histone exchange. Nat Struct Mol Biol 19:884–892
Stevens JR, O’Donnell AF, Perry TE, Benjamin JJ, Barnes CA, Johnston GC, Singer RA (2011) FACT, the Bur kinase pathway, and the histone co-repressor HirC have overlapping nucleosome-related roles in yeast transcription elongation. PLoS ONE 6:e25644
Sze CC, Ozark PA, Cao K, Ugarenko M, Das S, Wang L, Marshall SA, Rendleman EJ, Ryan CA, Zha D, Douillet D, Chen FX, Shilatifard A (2020) Coordinated regulation of cellular identity-associated H3K4me3 breadth by the COMPASS family. Sci Adv 6:eaaz4764
Tanny JC (2014) Chromatin modification by the RNA polymerase II elongation complex. Transcription 5:e988093
Tashiro S, Handa T, Matsuda A, Ban T, Takigawa T, Miyasato K, Ishii K, Kugou K, Ohta K, Hiraoka Y, Masukata H, Kanoh J (2016) Shugoshin forms a specialized chromatin domain at subtelomeres that regulates transcription and replication timing. Nat Commun 7:10393
Taunton J, Hassig CA, Schreiber SL (1996) A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 272:408–411
Taverna SD, Ilin S, Rogers RS, Tanny JC, Lavender H, Li H, Baker L, Boyle J, Blair LP, Chait BT, Patel DJ, Aitchison JD, Tackett AJ, Allis CD (2006) Yng1 PHD finger binding to H3 trimethylated at K4 promotes NuA3 HAT activity at K14 of H3 and transcription at a subset of targeted ORFs. Mol Cell 24:785–796
Van Oss SB, Shirra MK, Bataille AR, Wier AD, Yen K, Vinayachandran V, Byeon IL, Cucinotta CE, Heroux A, Jeon J, Kim J, VanDemark AP, Pugh BF, Arndt KM (2016) The histone modification domain of Paf1 complex subunit Rtf1 directly stimulates H2B ubiquitylation through an interaction with Rad6. Mol Cell 64:815–825
Venkatesh S, Smolle M, Li H, Gogol MM, Saint M, Kumar S, Natarajan K, Workman JL (2012) Set2 methylation of histone H3 lysine 36 suppresses histone exchange on transcribed genes. Nature 489:452–455
Venkatesh S, Li H, Gogol MM, Workman JL (2016) Selective suppression of antisense transcription by Set2-mediated H3K36 methylation. Nat Commun 7:13610
Vermeulen M, Mulder KW, Denissov S, Pijnappel WW, van Schaik FM, Varier RA, Baltissen MP, Stunnenberg HG, Mann M, Timmers HT (2007) Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131:58–69
Vermeulen M, Eberl HC, Matarese F, Marks H, Denissov S, Butter F, Lee KK, Olsen JV, Hyman AA, Stunnenberg HG, Mann M (2010) Quantitative interaction proteomics and genome-wide profiling of epigenetic histone marks and their readers. Cell 142:967–980
Vitaliano-Prunier A, Babour A, Herissant L, Apponi L, Margaritis T, Holstege FC, Corbett AH, Gwizdek C, Dargemont C (2012) H2B ubiquitylation controls the formation of export-competent mRNP. Mol Cell 45:132–139
Vlaming H, van Leeuwen F (2016) The upstreams and downstreams of H3K79 methylation by DOT1L. Chromosoma 125:593–605
Weinberg DN, Papillon-Cavanagh S, Chen H, Yue Y, Chen X, Rajagopalan KN, Horth C, McGuire JT, Xu X, Nikbakht H, Lemiesz AE, Marchione DM, Marunde MR, Meiners MJ, Cheek MA, Keogh MC, Bareke E, Djedid A, Harutyunyan AS, Jabado N, Garcia BA, Li H, Allis CD, Majewski J, Lu C (2019) The histone mark H3K36me2 recruits DNMT3A and shapes the intergenic DNA methylation landscape. Nature 573:281–286
Wier AD, Mayekar MK, Heroux A, Arndt KM, VanDemark AP (2013) Structural basis for Spt5-mediated recruitment of the Paf1 complex to chromatin. Proc Natl Acad Sci USA 110:17290–17295
Worden EJ, Wolberger C (2019) Activation and regulation of H2B-Ubiquitin-dependent histone methyltransferases. Curr Opin Struct Biol 59:98–106
Wu J, Chen Y, Lu LY, Wu Y, Paulsen MT, Ljungman M, Ferguson DO, Yu X (2011) Chfr and RNF8 synergistically regulate ATM activation. Nat Struct Mol Biol 18:761–768
Wu L, Lee SY, Zhou B, Nguyen UT, Muir TW, Tan S, Dou Y (2013) ASH2L regulates ubiquitylation signaling to MLL: trans-regulation of H3 K4 methylation in higher eukaryotes. Mol Cell 49:1108–1120
Wu L, Li L, Zhou B, Qin Z, Dou Y (2014) H2B ubiquitylation promotes RNA Pol II processivity via PAF1 and pTEFb. Mol Cell 54:920–931
Xie W, Nagarajan S, Baumgart SJ, Kosinsky RL, Najafova Z, Kari V, Hennion M, Indenbirken D, Bonn S, Grundhoff A, Wegwitz F, Mansouri A, Johnsen SA (2017) RNF40 regulates gene expression in an epigenetic context-dependent manner. Genome Biol 18:32
Yoh SM, Lucas JS, Jones KA (2008) The Iws1:Spt6:CTD complex controls cotranscriptional mRNA biosynthesis and HYPB/Setd2-mediated histone H3K36 methylation. Genes Dev 22:3422–3434
Youdell ML, Kizer KO, Kisseleva-Romanova E, Fuchs SM, Duro E, Strahl BD, Mellor J (2008) Roles for Ctk1 and Spt6 in regulating the different methylation states of histone H3 lysine 36. Mol Cell Biol 28:4915–4926
Zhang K, Dent SY (2005) Histone modifying enzymes and cancer: going beyond histones. J Cell Biochem 96:1137–1148
Zofall M, Grewal SI (2007) HULC, a histone H2B ubiquitinating complex, modulates heterochromatin independent of histone methylation in fission yeast. J Biol Chem 282:14065–14072
Acknowledgements
We thank members of the Tanny and Fisher labs for review of the manuscript and helpful discussions.
Funding
This work was supported by a Canadian Institutes of Health Research Grant MOP-130362 to JCT, Natural Sciences and Engineering Council of Canada Grant RGPIN 03661-15 to JCT, and NIH Grant R35GM127289 to RPF.
Author information
Authors and Affiliations
Contributions
DP and VP performed experiments indicated in Fig. 3. JCT and RPF wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Communicated by Michael Polymenis.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pinto, D., Pagé, V., Fisher, R.P. et al. New connections between ubiquitylation and methylation in the co-transcriptional histone modification network. Curr Genet 67, 695–705 (2021). https://doi.org/10.1007/s00294-021-01196-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-021-01196-x