Abstract
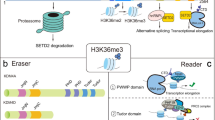
Histone modifications regulate key processes of eukaryotic genomes. Misregulation of the enzymes that place these modifications can lead to disease. An example of this is DOT1L, the enzyme that can mono-, di-, and trimethylate the nucleosome core on lysine 79 of histone H3 (H3K79). DOT1L plays a role in development and its misregulation has been implicated in several cancers, most notably leukemias caused by a rearrangement of the MLL gene. A DOT1L inhibitor is in clinical trials for these leukemias and shows promising results, yet we are only beginning to understand DOT1L’s function and regulation in the cell. Here, we review what happens upstream and downstream of H3K79 methylation. H3K79 methylation levels are highest in transcribed genes, where H2B ubiquitination can promote DOT1L activity. In addition, DOT1L can be targeted to transcribed regions of the genome by several of its interaction partners. Although methylation levels strongly correlate with transcription, the mechanistic link between the two is unclear and probably context-dependent. Methylation of H3K79 may act through recruiting or repelling effector proteins, but we do not yet know which effectors mediate DOT1L’s functions. Understanding DOT1L biology better will help us to understand the effects of DOT1L inhibitors and may allow the development of alternative strategies to target the DOT1L pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Post-translational modifications on histone proteins are involved in many key processes that occur at the chromatin. Whereas many of these modifications are placed on the histones tails that protrude from the nucleosome, some modifications are placed on the globular core of the nucleosome (Jack and Hake 2014). One of these core modifications is methylation on lysine 79 of histone H3 (H3K79) (Fig. 1). H3K79 can be mono-, di-, or tri-methylated by the enzyme disruptor of telomeric silencing-1 (Dot1), first identified in yeast. Dot1 performs this reaction using a catalytic domain that is most related to class-I methyltransferases, rather than the SET domain found in most histone lysine methyltransferases. Both the modification and the Dot1 enzyme are conserved throughout evolution, e.g., in mammals, flies, worms, yeast, and trypanosomes (Feng et al. 2002; Van Leeuwen et al. 2002; Shanower et al. 2005; Janzen et al. 2006; Cecere et al. 2013).
Crystal structure of the nucleosome, with H3K79 indicated in pink. H3K79 is located on the surface of the nucleosome core. H3, blue; H4, green; H2A, yellow and H2B, red. Structure from PDB 2CV5 (Tsunaka et al. 2005)
Dot1-like (DOT1L) is the mammalian H3K79 methyltransferase. DOT1L is important in mouse development, as reviewed by McLean et al. (2014). For example, germ-line Dot1L deletion results in embryonic lethality because of defects in the yolk sack and heart (Jones et al. 2008). At the cellular level, it has been found that inhibiting DOT1L promotes the generation of induced pluripotent stem cells by somatic cell reprogramming (Onder et al. 2012). Recently, this barrier-against-reprogramming function has been linked to a role for DOT1L in aging, acting as a downstream target of NF-κB (Soria-Valles et al. 2015). Finally, DOT1L has been implicated in MLL-rearranged leukemia and other cancers (Bernt and Armstrong 2011; Kim et al. 2012b; Zhang et al. 2014; Cho et al. 2015). H3K79 methylation probably mediates most of DOT1L’s effects, although the androgen receptor is a reported DOT1L substrate too (Yang et al. 2013) and more substrates may still be discovered. In MLL-rearranged leukemia, MLL target genes become hypermethylated on H3K79 and the leukemogenic potential of the MLL fusion protein depends on DOT1L activity (Krivtsov et al. 2008; Guenther et al. 2008; Bernt et al. 2011; Chen and Armstrong 2015). Several highly selective small-molecule inhibitors of DOT1L have been developed, which are all aminonucleosides that bind the co-factor pocket (Daigle et al. 2011; Yu et al. 2012; Daigle et al. 2013). A second-generation inhibitor, EPZ-5676, is currently in clinical trials for the treatment of MLL-rearranged leukemia. The results so far look promising (Stein and Tallman 2015), although the low bioavailability of the drug is a complicating factor (Basavapathruni et al. 2014). Inhibition of proliferation in breast and lung cancer cell lines by the DOT1L inhibitor has also been reported (Kim et al. 2012b; Zhang et al. 2014), and it will be interesting to see whether the DOT1L inhibitor will find more clinical applications.
Although we know of DOT1L’s involvement in important cellular and developmental processes and disease, we do not fully understand the biology of DOT1L and H3K79 methylation at the molecular level. In this review, we discuss the developments in this area. Mono-, di-, and trimethylated H3K79 are found in particular regions of the genome, and we are now beginning to understand how these patterns are established. What happens downstream of H3K79 methylation and whether this is dependent on the precise methylation state (me1/me2/me3) is still poorly understood. We will first describe where in the genome H3K79 methylation is found in mammalian cells. We will then discuss how DOT1L can establish these patterns, assisted by its binding partners and other regulatory pathways. Finally, we will describe what is known so far about the downstream consequences of H3K79 methylation.
H3K79 methylation levels and patterns
Several genome-wide maps of H3K79 methylation have been produced. To put genome-wide maps of H3K79me in perspective, it is useful to first consider the global levels of the different modification states in a cell. In contrast to yeast, where approximately 90 % of all H3K79 is either mono-, di-, or tri-methylated and where H3K79me3 is the dominant form (Van Leeuwen et al. 2002); in mammals most of H3K79 is unmethylated. In a few studies, mass spectrometry was employed to determine the fraction of each methylation state in mammalian cell lines (Jones et al. 2008; Darwanto et al. 2010; Sweet et al. 2010; Leroy et al. 2013; Alabert et al. 2015). Although there is variation between studies and between cell lines (see Supplementary Table 1), H3K79me1 is the dominant methylated form (6–30 %), H3K79me2 is substantially lower (0.1–10 %), and H3K79me3 is present at very low levels (0.1 %) or undetectable.
H3K79me patterns across the genome
Before the studies in mammalian cells were performed, it had already been reported that in flies and budding yeast H3K79me3 was found at coding sequences, with higher levels at expressed genes (Schübeler et al. 2004; Pokholok et al. 2005). The first studies using mammalian cell lines already suggested a similar relationship based on a selected set of genomic loci (Im et al. 2003; Vakoc et al. 2006).
Although H3K79me is relatively understudied compared to several more popular histone marks, we have learned that the pattern of H3K79 methylation is not flat over the genome, nor the same for all genes. In 2008, two important studies reported data sets from chromatin immunoprecipitation (ChIP) experiments followed by a micro-array (ChIP-chip) or sequencing (ChIP-seq), in human cell lines and primary human CD4+ T cells, respectively (Steger et al. 2008; Wang et al. 2008). These studies showed that H3K79 methylation at transcribed genes is a general phenomenon, true for all methylation states. H3K79 methylation is not limited to protein-coding genes, since H3K79me2 has also been found at several enhancers, expressed miRNA genes and at origins of replication (Steger et al. 2008; Suzuki et al. 2011; Fu et al. 2013). Within transcribed genes, there is a trend that the most highly transcribed genes have the highest levels of methylation, as depicted in Fig. 2 (Steger et al. 2008; Andersson et al. 2009; Zhang and Zhang 2011; Wang et al. 2013; Ho et al. 2013). Also, genes with a high elongation rate have higher H3K79me2 levels than slowly transcribed genes (Jonkers et al. 2014; Veloso et al. 2014). Possible explanations for this will be discussed below.
Distribution of the different H3K79 methylation states. An impression of average methylation profiles across +/− 10 kb around the transcription start site of non-, lowly and highly transcribed-genes. All methylation states peak just downstream of the transcription start site (TSS). Patterns based on Deshpande et al. (2014) and other publications mentioned in this review. The percentages refer to the approximate global levels of each methylation state observed in human cells, based on the mass spectrometry data in Supplementary Table 1
Within an expressed gene, the pattern of H3K79 methylation is not flat either (Fig. 2). H3K79me2 and H3K79me3 peak just behind the transcription start site and gradually decline throughout the first intron (Marson et al. 2008; Huff et al. 2010). The distribution of H3K79me1 is broader than that of H3K79me2/me3, but does peak in the same region (Steger et al. 2008; Huff et al. 2010). This is the region of transcriptional transition, behind the H3K4me3 peak found in regions of transcription initiation, and before H3K36me found in regions of transcription elongation. After a gradual decline throughout the first exon and intron, the level of all methylation states drops at internal exons (Huff et al. 2010).
Concerns about ChIP quality
The H3K79 methylation patterns have been determined by ChIP, which comes with certain technical issues. First, in general, the quality of ChIP-based epigenome maps depends on the quality of the antibodies. There are contradictory results on the specificity of several much used antibodies against methylated H3K79, probably due to variation between batches (Jones et al. 2008; Steger et al. 2008; Feng et al. 2010; Egelhofer et al. 2010; FitzGerald et al. 2011; Rothbart et al. 2015). Some of these observations have been collected in the Antibody Validation Database (Egelhofer et al. 2010) and the Histone Antibody Specificity Database (Rothbart et al. 2015), but batch numbers are not routinely reported in publications. Therefore, it will be important to develop new reagents with high specificity and consistent performance, such as monoclonal antibodies or non-antibody-based reagents.
Second, Steger et al. have shown that ChIP for methylated H3K79 is facilitated by protein denaturation by SDS (Steger et al. 2008) and we have observed this using yeast chromatin and other antibodies as well (unpublished data). Since H3K79 is located on the core of the nucleosome rather than on an unstructured histone tail (Fig. 1), it is likely that denaturation is required for the structured part of H3 to resemble the peptide that was used to raise the antibody. Indeed, recently it was found that two antibodies against H3K79me2 could not recognize their PTM in non-denaturing conditions, resulting in a poor ChIP result (Rothbart et al. 2015). This highlights the importance of using denaturing conditions when performing H3K79me ChIP and reporting on the exact ChIP conditions used to allow for meaningful data interpretation and comparison.
H3K79 methylation throughout the cell cycle
The patterns described above are all based on asynchronous populations of cycling cells, but patterns and levels of H3K79 methylation do not remain constant throughout the cell cycle. In S phase, the incorporation of newly synthesized unmethylated histones will lead to a transient drop in global methylation levels. Whereas histone acetylation is generally rapidly re-established, histone methylation takes more time (Barth and Imhof 2010). The relation between cell cycle and H3K79 methylation is not the same for all species and even within human cells different fluctuations have been observed (Feng et al. 2002; Kim et al. 2012b; Fu et al. 2013; Kim et al. 2014; Guppy and McManus 2015). Quantitative mass-spectrometry analyses suggest that overall the levels of H3K79me1 and H3K79me2 vary little throughout the cell cycle and are relatively rapidly re-established on newly synthesized histones (Sweet et al. 2010; Alabert et al. 2015). However, methylation of H3K79 is not restricted to S phase but is a process that can proceed throughout the cell cycle (Sweet et al. 2010; Alabert et al. 2015). Indeed, withdrawal from the cell cycle is accompanied by an increase in H3K79 methylation (Alabert et al. 2015).
H3K79 methylation not only varies throughout mitosis, but also throughout meiosis. An immunofluorescence study showed changes in both the levels and localization of the different methylation states as meiosis advances in spermatocytes (Ontoso et al. 2014). These spatio-temporal patterns may be necessary for proper meiosis, as suggested by studies in mouse oocytes and meiosis in budding yeast (San-Segundo and Roeder 2000; Ontoso et al. 2013; Wang et al. 2014).
DOT1L and its regulation
In the previous section we described how H3K79 methylation is distributed over the genome, but how is this pattern established? DOT1L is the major enzyme responsible for H3K79 methylation; Dot1L KO cells have no detectable H3K79 methylation left (Jones et al. 2008; Feng et al. 2010; Deshpande et al. 2014) and treatment of cells with DOT1L inhibitors greatly reduces H3K79me levels (Daigle et al. 2011). Recently, RE-IIBP has been suggested to possess H3K79 methylation activity too (Woo Park et al. 2015); however, its relative contribution to regulation of H3K79me levels in cells remains to be established. Since DOT1L is the major H3K79 methyltransferase, global H3K79 methylation levels are responsive to changes in DOT1L activity and expression. Global changes in H3K79 methylation through deregulation of DOT1L expression have been observed in cancer cells, upon viral infection and upon physiological aging as well as in aging disorders (Kryczek et al. 2014; O’Connor et al. 2014; Soria-Valles et al. 2015).
To understand the patterns of H3K79me and the regulation of DOT1L, it is important to consider the mechanism of methylation. DOT1L methylates H3K79 using S-adenosylmethionine (SAM) as the methyl donor and producing S-adenosyl-L-homocysteine (SAH) in the process. Such a reaction can be catalyzed in a processive or distributive manner; a processive enzyme can stay bound to its substrate while catalyzing the addition of several methyl groups, while a distributive enzyme must dissociate after each round of methylation. As a result, a distributive enzyme cannot generate the different methylation states independently. In contrast to histone lysine methyltransferases with a SET domain, DOT1L seems to be a distributive enzyme. This is based on its structure (Fig. 3a), in which the active site is occluded and the SAH/SAM exchange seems to require the release of the histone substrate (Min et al. 2003; Sawada et al. 2004; Cheng et al. 2005; Basavapathruni et al. 2012). Interestingly, recently NRMT1 has been identified as another histone methyltransferase with a DOT1L-like fold (Wu et al. 2015). This enzyme’s preference to produce trimethylated states of its substrate lysine on CENP-A could hint at a processive mechanism. However, it can also be explained by the preference of NRMT1 for already methylated substrates (Wu et al. 2015). Whether or not this enzyme is distributive remains to be determined. Characteristic for a distributive enzyme is the shift from higher to lower methylation states upon decreasing the enzyme’s concentration or activity (Frederiks et al. 2008). Indeed, this was observed when changing Dot1’s expression levels in yeast (Frederiks et al. 2008; De Vos et al. 2011). In mammalian cells, a similar experiment has not been reported yet, but at particular loci a shift from H3K79me3 to H3K79me1 was observed upon deletion of the positive regulator AF10 (Deshpande et al. 2014).
hDOT1L structure. a Structure of hDOT1L 1–416, PDB 1NW3 (Min et al. 2003). The methyl donor S-adenosylmethionine (SAM) is shown in its binding pocket, the arrow indicates the predicted binding channel for the H3K79 side chain. The protein backbone is visualized as a ribbon, colored with a gradient from blue to yellow from the N to C terminus. The image was produced using the UCSF Chimera package (Pettersen et al. 2004). b The following domains have been mapped in hDOT1L, from N to C terminus, with the region they span indicated by amino acid numbers: catalytic domain (1–332, Min et al. (2003)), Bat3-interacting domain (361–380, Wakeman et al. (2012)), K-rich patch (390–407, Min et al. (2003)) with a partially overlapping NLS (395–417, Reisenauer et al. (2010)), leucine zipper motif (576–594, Zhang et al. (2004)), CTD binding patch (618–627, Kim et al. (2012a)), three ENL/AF9 binding sites (628–653, 863–878, 877–900, Kuntimaddi et al. (2015)), and two more NLSs (1088–1111, 1164–1171, Reisenauer et al. (2010))
Whereas yeast Dot1 is very active in in vitro methylation reactions, the intrinsic activity of the catalytic domain of human DOT1L is low (Vlaming et al. 2014; Stulemeijer et al. 2015). Given the low number of DOT1L molecules in a cell compared to the high number of histones (Hein et al. 2015), the chance that DOT1L randomly methylates the same histone twice is very small. Moreover, the H3K79 methylation patterns do not completely match the data on DOT1L’s genomic distribution (Kim et al. 2012a), suggesting that DOT1L is activated in particular places. It seems like DOT1L requires other proteins or signals for targeting to and activation at particular regions of the genome. For some regulators, interaction sites in DOT1L have been mapped (Fig. 3b), but the exact mechanisms and relative contributions of the different regulators are not always clear. The different mechanisms of regulating DOT1L to create the observed levels and patterns of H3K79 methylation will be discussed below.
Removing H3K79 methylation
To date, no demethylase for H3K79 has been identified and the stability of H3K79 methylation in HeLa cells suggests that no global demethylase activity is present (Zee et al. 2010; Sweet et al. 2010). However, it is possible that there is demethylase activity in certain cell types or under certain conditions. For example, in fertilized oocytes a sudden loss of H3K79 methylation has been observed (Ooga et al. 2008). Whether this is caused by a demethylase or other mechanisms remains to be established. The distributive DOT1L enzyme does not distinguish between old and new histones when methylating; it can methylate newly synthesized histones as well as histones that it previously methylated. This combined with the lack of demethylase activity, results in an accumulation of H3K79 methylation on old histones (Sweet et al. 2010; De Vos et al. 2011). The accumulation can be counteracted by replacement of methylated histones by unmodified ones, during cell division and by replication-independent histone turnover, as we have reviewed before (Fig. 4a) (Vlaming and van Leeuwen 2012). Thus, factors that affect the rate of cell division or histone turnover will also affect H3K79 methylation levels and patterns across the epigenome, unless the activity of DOT1L is simultaneously regulated.
Regulation of H3K79 methylation. a Replacement of modified histones by new, unmodified histones effectively removes methylation. b H2Bub is thought to regulate DOT1L by coaching it toward H3K79, the “crash barrier model”. c Proteins proposed to be part of the DOT1L-containing complex Dot1Com. d Other DOT1L regulators. Bat3 may assist DOT1L’s interaction with the nucleosome, c-Myc forms a complex with DOT1L and CBP/p300 and target these proteins to particular DNA sequences, DOT1L interaction with the phosphorylated C-terminal domain (CTD) repeats of RNA Polymerase II may further enhance targeting of H3K79me to transcribed regions
H2Bub cross-talk
The first process found to regulate H3K79 methylation was a conserved trans-histone cross-talk. Ubiquitination of H2B on the nucleosome core promotes methylation of both H3K79 and H3K4 methylation in mammals, yeast, and other species (Sun and Allis 2002; Zhu et al. 2005b). In mammals, ubiquitination on lysine 120 of H2B is carried out by the RNF20/40 E3 ligase complex and the conjugating enzymes RAD6A and RAD6B (Zhu et al. 2005b; Kim et al. 2005; Kim et al. 2009). In many cases, a mono-ubiquitin is recognized by ubiquitin-interacting motifs, but in this case it is more complicated than DOT1L binding to the ubiquitin (McGinty et al. 2008). Although we still do not fully understand the cross-talk on a molecular level, based on observations in vitro, in yeast and in mammalian cells (McGinty et al. 2008; McGinty et al. 2009; Chatterjee et al. 2010; Vlaming et al. 2014; Holt et al. 2015) we and the others proposed the “crash barrier” or “corralling” model (Fig. 4b) (Vlaming et al. 2014; Holt et al. 2015). In this model, ubiquitin folds back onto the nucleosome and forms a barrier to DOT1L, increasing the chance that it is well-positioned for methylating H3K79. There is quite some plasticity in this cross-talk and also ubiquitination on H2BK34 by MSL1/2 can promote the methylation of H3K4 and H3K79 (Wu et al. 2011b).
H2B ubiquitination leads to more H3K79 methylation, but where does the ubiquitination occur? H2BK120 is ubiquitinated co-transcriptionally and H2Bub levels strongly correlate with expression level and RNA polymerase II elongation rate (Minsky et al. 2008; Fuchs et al. 2014). RNF20/40 and MSL1/2 interact with the RNA Polymerase II-associated PAF complex and these proteins are functionally interdependent (Wu et al. 2014). Consistent with the ubiquitin ligases depending on the PAF complex, disruption of the PAF complex decreases H3K79 methylation (Zhu et al. 2005a; Wu et al. 2014). However, along transcribed regions, the activity of DOT1L varies. H2Bub levels drop at intron-exon boundaries, reflecting transcription elongation dynamics (Huff et al. 2010; Fuchs et al. 2014), which is very much like the pattern observed for H3K79 methylation (as described earlier). Since H2B ubiquitination directly promotes H3K79 methylation and the marks occur in roughly the same places, this trans-histone cross-talk and thus, indirectly, transcriptional elongation probably explains much of the H3K79 methylation pattern.
Surprisingly, in two recent publications, an uncoupling of H2B ubiquitination and H3K79 methylation was suggested (Cucinotta et al. 2015; Yao et al. 2015). Mouse fibroblasts lacking the Mediator subunit MED23 show strongly decreased global H2Bub levels while H3K79 methylation is unaffected (Yao et al. 2015). However, in coding regions, where most H3K79 methylation occurs, H3K79me3 is decreased by the deletion of MED23. In budding yeast, uncoupling of global H2Bub and H3K79 methylation levels was observed in specific mutations in the histone H2A protein (Cucinotta et al. 2015). Potentially, other DOT1L regulators could play a role in maintaining H3K79 methylation globally.
Partners in the DOT1L complex
DOT1L has several binding partners that can regulate its activity. Several DOT1L-containing complexes with different compositions have been described. Initially, it was suggested that there was one large complex containing AF4, AF9, AF10, DOT1L, and P-TEFb, a kinase that phosphorylates RNA Pol II in the transition from initiation to elongation (Bitoun et al. 2007; Mueller et al. 2007; Mueller et al. 2009). Later, it was shown that the interactions of AF4 and DOT1L with AF9 were mutually exclusive and there are actually two separate complexes (Yokoyama et al. 2010; Mohan et al. 2010; Park et al. 2010; He et al. 2011; Biswas et al. 2011). The DOT1L-containing complex was named DotCom and contains DOT1L, AF10, and AF9 or ENL, with maybe some other subunits like AF17 and proteins from the Wnt pathway (Fig. 4c) (Mohan et al. 2010; Park et al. 2010; Biswas et al. 2011). Not much work has been done in comparing the activity of DOT1L alone with DOT1L in its complex, but the effect of individual complex subunits on DOT1L has been investigated and will be the topic of the next paragraphs. Notably, AF9, ENL, and AF10 are all frequent translocation partners of MLL1 in MLL-rearranged leukemias (Meyer et al. 2013). Importantly, knockdown of AF10, AF9 and to a lesser extent ENL reduces H3K79 methylation levels (Mueller et al. 2007; Lin et al. 2009; Mohan et al. 2010). This means that they are not only interacting, but that these proteins also regulate DOT1L. In the following paragraphs we will focus on AF10 and AF9/ENL separately.
AF10 was first identified as a fusion partner of MLL in leukemia patients (Chaplin et al. 1995). It contains a region with an octapeptide motif and a leucine zipper (OM-LZ) that is required for the leukemogenic potential of the MLL-AF10 and CALM-AF10 fusion proteins generated by chromosomal translocations (DiMartino et al. 2002; Deshpande et al. 2011). This OM-LZ domain also mediates AF10’s interaction with DOT1L and several other proteins (Okada et al. 2005; Greif et al. 2008). AF10 is necessary for full DOT1L activity; in its absence, H3K79me2 is lost, though H3K79me1 remains (Lin et al. 2009; Deshpande et al. 2014). This could be explained by a role for AF10 in enabling DOT1L to specifically catalyze the second methylation step. However, since DOT1L is likely to be a distributive enzyme, an alternative explanation is that AF10 promotes DOT1L’s general activity or tethers DOT1L to specific regions of the genome, increasing the chance of methylating the same residue twice. Two pieces of evidence support the second idea: (1) AF10 increases the binding of DOT1L to chromatin, which one would expect to have a general positive effect on DOT1L activity (Lin et al. 2009), and (2) at locations in the genome where methylation is very high, there is a shift from high H3K79me3 to high H3K79me1 upon AF10 deletion (Deshpande et al. 2014). This shift happens, for example, on MLL-AF9 target genes, although this could be indirect because also expression of these target genes went down (Deshpande et al. 2014). Very recently, the PZP domain of AF10 was shown to bind to nucleosomes that are unmethylated on H3K27 and this domain was required for full H3K79 methylation (Chen et al. 2015b). Where AF10 sits on the genome has not been studied yet in mammalian cells, but one study showed that C. elegans AF10 homologue ZFP-1 is mostly found in promoter regions of highly expressed genes and shares its target genes with DOT-1.1 (Cecere et al. 2013). This would fit with AF10 binding only in regions devoid of H3K27 methylation (Chen et al. 2015b) and could be a mechanism for DOT1L to be recruited to expressed genes.
Closely related AF9 and ENL were both first discovered as translocation partners of MLL (Nakamura et al. 1993; Rubnitz et al. 1994). They can take the same place in both the DOT1L-containing complex DotCom and the AF4-containing super elongation complex (He et al. 2011). The N terminus of AF9 and ENL contains a YEATS domain, which is important for the targeting of these factors to chromatin (He et al. 2011). This may be because of the interaction of the YEATS domain with PAF1 of the before-mentioned PAF complex (He et al. 2011), or because it can bind nucleosomes with acetylated lysines (Li et al. 2014). AF9 has the highest affinity for H3K9ac, whereas ENL has a higher affinity for H3K27ac (Li et al. 2014). AF9 and ENL also contain an ANC1 homology domain (AHD), retained in MLL fusions, which is generally disordered but can bind AF4 and DOT1L and possibly the Polycomb group protein CBX8 (Zhang et al. 2006b; Tan et al. 2011; Leach et al. 2013; Kuntimaddi et al. 2015). In DOT1L, there are three sites that can bind AF9 (aa 628–653, 863–878, and 877–900, see Fig. 3b) and that may simultaneously bind several AF9s (Kuntimaddi et al. 2015). They all bind AF9 in the same way, but sites 2 and 3 bind with a higher affinity. The same sites in DOT1L have also been reported to interact with AF17, which would direct DOT1L to the cytoplasm and would compete with AF9 binding (Reisenauer et al. 2009). AF9 is important for DOT1L recruitment to chromatin (Li et al. 2014; Kuntimaddi et al. 2015) and there is overlap between H3K9 acetylation and H3K79me3. Moreover, H3K79me3 on AF9 target genes is AF9 dependent (Li et al. 2014), although an indirect effect via transcription cannot be ruled out. Given the biochemical evidence, it is likely that AF9 recruits DOT1L to sites in the genome where H3K9 is acetylated, namely around the transcription start sites of expressed genes (Wang et al. 2008). Interestingly, knockdown of the histone deacetylase SIRT1 leads to an increase in H3K9 acetylation and H3K79 methylation at its target sites (Chen et al. 2015a). If AF9/ENL retains its interaction with the PAF complex when in DotCom, this could also be a mechanism to recruit DOT1L to transcribed regions, where H2Bub would be present to activate DOT1L. Mutations in AF9 that affect DOT1L recruitment differently also have distinct effects on H3K79me2 and H3K79me3 (Kuntimaddi et al. 2015). The D546R mutation in AF9 severely disrupts DOT1L recruitment and reduces both H3K79me2 and H3K79me3, whereas the D544R mutation, which has a limited effect on DOT1L recruitment, affects only H3K79me3. This led to the hypothesis that AF9 increases the residency time of DOT1L so that chances are higher that it reaches the trimethylated state (Kuntimaddi et al. 2015). An interesting notion from the authors is that phosphorylation of DOT1L’s interaction sites (Hornbeck et al. 2012) could disrupt binding to AF9, so the (currently unknown) kinases responsible for this phosphorylation may regulate DOT1L by interfering with the DOT1L-AF9 interaction.
Other DOT1L interactors
A few more proteins of different functions have been reported to interact with DOT1L (Fig. 4d). First, Bat3 has been reported to interact with both DOT1L and histone H3 and was suggested to increase DOT1L activity by bringing DOT1L and H3 together (Wakeman et al. 2012). Second, in a very recent publication DOT1L was shown to interact with well-known oncogene c-Myc (Cho et al. 2015). The presence of DOT1L and H3K79me2 at several loci depended on c-Myc, suggesting that DOT1L is targeted to these loci through its interaction with c-Myc. The authors propose that, in the absence of DOT1L, c-Myc interacts with the repressive proteins HDAC1 and DNMT1, but together with DOT1L, it forms an activating complex with histone acetyltransferase CBP/p300. Finally, DOT1L has also been reported to directly interact with RNA Polymerase II phosphorylated on Ser2 and/or Ser5 of its C-terminal domain (Kim et al. 2012a). Of course this could also lead to DOT1L recruitment to transcribed genes. As all DOT1L regulators affect or are affected by transcription, it is difficult to assess which effects on H3K79 methylation are direct and what the relative contributions of each regulator are.
Misregulation in MLL-r leukemias
Above, we have described several mechanisms through which DOT1L can reach higher methylation states in particular regions of the genome. MLL-rearranged leukemias are characterized by aberrant H3K79 methylation patterns (Krivtsov et al. 2008; Bernt et al. 2011). When the C terminus of AF9, ENL or AF10 is fused to MLL1, the fusion partners can directly recruit DOT1L to the MLL target sites, rather than to its normal binding sites (Okada et al. 2005; Kuntimaddi et al. 2015). However, also leukemias in which MLL1 is fused to other proteins, such as AF4, show high H3K79me2 at MLL target loci (Daigle et al. 2011; Deshpande et al. 2013), suggesting that other mechanisms are involved as well. The N terminus of MLL1, which is retained in the oncogenic MLL fusion proteins, directly interacts with the PAF complex and this interaction is required for high H3K79 methylation (Muntean et al. 2010; Milne et al. 2010). Also RNF20, which can be recruited by the PAF complex, was required for high H3K79me2 at MLL-AF9 target genes (Wang et al. 2013). To conclude, several mechanisms are responsible for the high H3K79 methylation at MLL fusion target genes. It will be interesting to determine whether the interactions between DOT1L and AF9/ENL or AF10 (as described above) can be targeted in MLL-r leukemia, like was recently done for the interaction between MLL and Menin (Borkin et al. 2015).
Functions of DOT1L
DOT1L and transcription
Although it is clear that DOT1L is an important enzyme, it is not so clear how DOT1L exerts its functions. What happens downstream of the methylation of H3K79 is relatively understudied. As described, there is a high correlation between transcription elongation rates and H3K79me2, but a causal relationship cannot be assumed. Important lessons can be learned from other histone H3 methylation marks that are associated with active transcription. H3K36me3 and H3K4me2 are examples of marks that correlate with gene transcription although they exert repressive effects within transcribed regions (Margaritis et al. 2012; Smolle and Workman 2013). DOT1L perturbation studies show that there are some examples of genes whose expression depends on DOT1L, such as dystrophin and particular Wnt target genes (Nguyen et al. 2011; Gibbons et al. 2015). On the other hand, the ENaCα gene, encoding a subunit of the epithelial Na + channel, was found to be repressed by H3K79 methylation (Zhang et al. 2006a; Zhang et al. 2007; Reisenauer et al. 2009). Although H3K79 methylation is widespread, there does not seem to be a consistent global effect on transcription upon DOT1L deletion. The number of deregulated genes varies between cell types or studies, ranging from 200 to 2000 (Bernt et al. 2011; Wu et al. 2013; Ho et al. 2013). Interestingly, of these deregulated genes, a majority is upregulated. Whether these effects are all direct and whether the effect on transcription is dependent on chromatin context remains to be determined. One possibility is that H3K79 methylation is a mark of transcriptional memory that may be required to maintain transcription of certain developmentally regulated genes (Kouskouti and Talianidis 2005; Steger et al. 2008; Onder et al. 2012).
In MLL-rearranged leukemia, there is a loss of expression of MLL target genes upon inhibition of DOT1L (Bernt et al. 2011; Daigle et al. 2011). Since many of the MLL fusion proteins can directly recruit DOT1L, DOT1L is thought to be a cause of the upregulation of transcription at these genes. A recent study shed a light on how DOT1L could regulate transcription at these genes. In the absence of DOT1L, there is more SIRT1 present at the MLL target loci (Chen et al. 2015a). Indeed, without SIRT1, MLL-rearranged cells no longer depend on DOT1L for expression of the HoxA cluster and other important target genes (Chen et al. 2015a). However, the situation is more complex because SIRT1 is also a negative regulator of DOT1L, possibly via the recruitment of AF9 (see above). Thus, there seems to be a competition between DOT1L and SIRT1: in the absence of DOT1L, SIRT1’s histone deacetylation activity will lead to silencing, in the absence of SIRT1, DOT1L is unleashed. This is quite similar to Dot1’s function in yeast, where it competes with the silencing protein Sir3 for binding to nucleosomes (Van Leeuwen et al. 2002). In contrast to the competition with Sir3 in yeast (discussed further on), the competition with SIRT1 is not yet understood at the molecular level.
Putting methylated H3K79 into action
Histone PTMs can have direct effects by changing nucleosome structure or dynamics or indirect effects through the binding or repelling of effector proteins (reviewed by Musselman et al. (2012) and Tessarz and Kouzarides (2014), respectively). H3K79 is located on the solvent-accessible surface of the nucleosome core, close to H3-H4 interface. Crystal structures of recombinant nucleosomes with H3K79me0 or an H3K79me2 mimic, showed no changes in the global structure of the nucleosome (Lu et al. 2008). Locally, however, there is a subtle conformational change: when dimethylated, the H3K79 side chain becomes almost completely solvent accessible and opens up a small hydrophobic pocket on the H3-H4 interface. This small change is not expected to have direct effects on nucleosome architecture, so H3K79 methylation likely functions by influencing the binding of proteins to the area of the nucleosome around H3K79 (Lu et al. 2008).
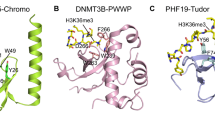
Only a few binders of H3K79 methylation have been proposed: the PWWP domain of hepatoma-derived growth factor 2 (HDGF2) and maybe BRPF1 and BRPF2 (Wu et al. 2011a), the Tudor domain of survival motor neuron protein (SMN) (Sabra et al. 2013) and the tandem Tudor domains of fragile X mental retardation protein (FMRP) and 53BP1 (Huyen et al. 2004; Alpatov et al. 2014). However, most evidence for these binders is based on experiments using methylated peptides. Peptides may not recapitulate the more complex and structured globular domain in the context of the nucleosome, which allows for multiple interactions with different sites (e.g., Armache et al. (2011)). As an example, DOT1L itself does not methylate H3 alone, but requires the histone octamer or whole nucleosome context in order to methylate (McGinty et al. 2008). With the recent advancements in synthesizing histone proteins (Nguyen et al. 2014), it is becoming possible to use chemically defined modified nucleosomes to validate the proposed binders and to discover new binders.
How and in which context the suggested H3K79me readers relay information from chromatin to downstream processes is not well understood. For 53BP1 it is clear that its recruitment to sites of DNA damage is important (Panier and Boulton 2014), but the involvement of H3K79 methylation in its recruitment is less clear. H4K20me2 and not H3K79me has been suggested to be the main histone target of 53BP1 (Botuyan et al. 2006; Tong et al. 2015). It is still possible that H3K79 methylation is important for 53BP1 recruitment under certain circumstances, e.g., when H4K20me2 levels are low (Wakeman et al. 2012). In budding yeast, where H4K20me2 is absent, Rad9(53BP1)/Rad53 checkpoint activation upon damage in the G1 phase does depend on Dot1 (Giannattasio et al. 2005; Wysocki et al. 2005). Whether via 53BP1 recruitment or another mechanism, such as by mediating transcriptional restart as proposed by Oksenych et al. (2013), DOT1L knockdown does increase sensitivity to ionizing radiation and UV (Lin et al. 2009; Wakeman et al. 2012; Oksenych et al. 2013).
H3K79 methylation can also function by disrupting interactions. In yeast, the silencing protein Sir3 binds to unmethylated nucleosomes and this interaction is disrupted by methyl groups on H3K79 (Van Leeuwen et al. 2002; Ehrentraut et al. 2011; Armache et al. 2011). It seems that the interaction can be disrupted by all methylation states, meaning there is a functional redundancy (Frederiks et al. 2008). The consequence of blocking the interaction with Sir3 is that the silencing complex is concentrated on the parts of the genome without H3K79 methylation, namely the telomeres and the silent-mating type locus. In line with the yeast data, MEFs lacking DOT1L show reduced levels of heterochromatic marks at centromeres and telomeres (Jones et al. 2008). It is still unclear whether this occurs via a similar competition mechanism.
So, H3K79 methylation is expected to act through promoting or preventing the binding of effector proteins, but how these would mediate DOT1L’s described functions is currently not understood. Also, it is unknown whether mono-, di-, and trimethylated H3K79 attract or repel specific proteins, as has been shown for the different methylation states of several other histone lysines (Wagner et al. 2014). Methyl-state specific binders may still be found, but it is also possible that the different methylation states are redundant, as has been suggested for yeast silencing (Frederiks et al. 2008). In that case, higher methylation states would merely reflect longer DOT1L residency time. Longer residency time could make the methylation more robust, i.e., fewer histones will remain without methyl groups, and the higher methylation states could be a by-product.
Concluding remarks
The conserved enzyme DOT1L methylates H3K79 and is important in development, reprogramming, and cancer. In recent years, progress has been made in understanding how DOT1L is regulated to methylate mainly in transcribed ORFs. H2B ubiquitination and DOT1L’s interactors, including those in the DOT1L complex, both contribute to the pattern of H3K79 methylation, by activating DOT1L or increasing its residency time at transcriptionally active regions of the genome. Several of these factors are deregulated in MLL-rearranged leukemia, underscoring the central role of the DOT1L-H3K79me pathway in controlling cell function. Targeting DOT1L regulators may become an alternative to DOT1L inhibitors as a way to modulate DOT1L activity in disease. Given the importance of DOT1L in normal development (McLean et al. 2014) and the fact that a large fraction of MLL-rearranged leukemia patients are infants (Meyer et al. 2013), alternative treatment options could become very important.
Our mechanistic understanding of downstream effects of H3K79 methylation is still limited. There are hypotheses based on genome-wide correlations, but the next challenge will be to move beyond correlations and establish the function of H3K79 methylation. There are a few strong cases where DOT1L seems to be causally involved in transcriptional regulation, but the underlying molecular mechanism remains to be determined. Establishing how the methylation state of histone H3K79 is relayed into action in the cell will be important to fully understand the contribution of H3K79me to control of the genome.
References
Alabert C, Barth TK, Reverón-Gómez N et al (2015) Two distinct modes for propagation of histone PTMs across the cell cycle. Genes Dev 29:585–90. doi:10.1101/gad.256354.114
Alpatov R, Lesch BJ, Nakamoto-Kinoshita M et al (2014) A chromatin-dependent role of the fragile X mental retardation protein FMRP in the DNA damage response. Cell 157:869–81. doi:10.1016/j.cell.2014.03.040
Andersson R, Enroth S, Rada-Iglesias A et al (2009) Nucleosomes are well positioned in exons and carry characteristic histone modifications. Genome Res 19:1732–41. doi:10.1101/gr.092353.109
Armache K-J, Garlick JD, Canzio D et al (2011) Structural basis of silencing: Sir3 BAH domain in complex with a nucleosome at 3.0 a resolution. Science 334:977–982. doi:10.1126/science.1210915
Barth TK, Imhof A (2010) Fast signals and slow marks: the dynamics of histone modifications. Trends Biochem Sci 35:618–26. doi:10.1016/j.tibs.2010.05.006
Basavapathruni A, Jin L, Daigle SR et al (2012) Conformational adaptation drives potent, selective and durable inhibition of the human protein methyltransferase DOT1L. Chem Biol Drug Des 80:971–80. doi:10.1111/cbdd.12050
Basavapathruni A, Olhava EJ, Daigle SR et al (2014) Nonclinical pharmacokinetics and metabolism of EPZ-5676, a novel DOT1L histone methyltransferase inhibitor. Biopharm Drug Dispos 35:237–52. doi:10.1002/bdd.1889
Bernt KM, Armstrong SA (2011) A role for DOT1L in MLL -rearranged leukemias. Epigenomics 3:667–670. doi:10.2217/epi.11.98
Bernt KM, Zhu N, Sinha AU et al (2011) MLL-rearranged leukemia is dependent on aberrant H3K79 methylation by DOT1L. Cancer Cell 20:66–78. doi:10.1016/j.ccr.2011.06.010
Biswas D, Milne TA, Basrur V et al (2011) Function of leukemogenic mixed lineage leukemia 1 (MLL) fusion proteins through distinct partner protein complexes. Proc Natl Acad Sci U S A 108:15751–15756. doi:10.1073/pnas.1111498108
Bitoun E, Oliver PL, Davies KE (2007) The mixed-lineage leukemia fusion partner AF4 stimulates RNA polymerase II transcriptional elongation and mediates coordinated chromatin remodeling. Hum Mol Genet 16:92–106. doi:10.1093/hmg/ddl444
Borkin D, He S, Miao H et al (2015) Pharmacologic inhibition of the menin-MLL interaction blocks progression of MLL leukemia in vivo. Cancer Cell 27:589–602. doi:10.1016/j.ccell.2015.02.016
Botuyan MV, Lee J, Ward IM et al (2006) Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127:1361–73. doi:10.1016/j.cell.2006.10.043
Cecere G, Hoersch S, Jensen MB et al (2013) The ZFP-1(AF10)/DOT-1 complex opposes H2B ubiquitination to reduce Pol II transcription. Mol Cell 50:894–907. doi:10.1016/j.molcel.2013.06.002
Chaplin T, Ayton P, Bernard OA et al (1995) A novel class of zinc finger/leucine zipper genes identified from the molecular cloning of the t(10;11) translocation in acute leukemia. Blood 85:1435–41
Chatterjee C, McGinty RK, Fierz B, Muir TW (2010) Disulfide-directed histone ubiquitylation reveals plasticity in hDot1L activation. Nat Chem Biol 6:267–9. doi:10.1038/nchembio.315
Chen C-W, Armstrong SA (2015) Targeting DOT1L and HOX gene expression in MLL-rearranged leukemia and beyond. Exp Hematol 43:673–684. doi:10.1016/j.exphem.2015.05.012
Chen C-W, Koche RP, Sinha AU et al (2015a) DOT1L inhibits SIRT1-mediated epigenetic silencing to maintain leukemic gene expression in MLL-rearranged leukemia. Nat Med 21:335–343. doi:10.1038/nm.3832
Chen S, Yang Z, Wilkinson AW, et al (2015b) The PZP Domain of AF10 Senses Unmodified H3K27 to Regulate DOT1L-Mediated Methylation of H3K79. Mol Cell 60:319–327. doi:10.1016/j.molcel.2015.08.019
Cheng X, Collins RE, Zhang X (2005) Structural and sequence motifs of protein (histone) methylation enzymes. Annu Rev Biophys Biomol Struct 34:267–94. doi:10.1146/annurev.biophys.34.040204.144452
Cho M-H, Park J-H, Choi H-J et al (2015) DOT1L cooperates with the c-Myc-p300 complex to epigenetically derepress CDH1 transcription factors in breast cancer progression. Nat Commun 6:7821. doi:10.1038/ncomms8821
Cucinotta CE, Young AN, Klucevsek KM, Arndt KM (2015) The nucleosome acidic patch regulates the H2B K123 monoubiquitylation cascade and transcription elongation in saccharomyces cerevisiae. PLoS Genet 11:e1005420. doi:10.1371/journal.pgen.1005420
Daigle SR, Olhava EJ, Therkelsen CA et al (2013) Potent inhibition of DOT1L as treatment of MLL-fusion leukemia. Blood 122:1017–1025. doi:10.1182/blood-2013-04-497644
Daigle SR, Olhava EJ, Therkelsen CA et al (2011) Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 20:53–65. doi:10.1016/j.ccr.2011.06.009
Darwanto A, Curtis MP, Schrag M et al (2010) A modified “cross-talk” between histone H2B Lys-120 ubiquitination and H3 Lys-79 methylation. J Biol Chem 285:21868–76. doi:10.1074/jbc.M110.126813
De Vos D, Frederiks F, Terweij M et al (2011) Progressive methylation of ageing histones by Dot1 functions as a timer. EMBO Rep 12:956–62. doi:10.1038/embor.2011.131
Deshpande AJ, Chen L, Fazio M et al (2013) Leukemic transformation by the MLL-AF6 fusion oncogene requires the H3K79 methyltransferase Dot1l. Blood 121:2533–2541. doi:10.1182/blood-2012-11-465120
Deshpande AJ, Deshpande A, Sinha AU et al (2014) AF10 regulates progressive H3K79 methylation and HOX gene expression in diverse AML subtypes. Cancer Cell 26:896–908. doi:10.1016/j.ccell.2014.10.009
Deshpande AJ, Rouhi A, Lin Y et al (2011) The clathrin-binding domain of CALM and the OM-LZ domain of AF10 are sufficient to induce acute myeloid leukemia in mice. Leukemia. doi:10.1038/leu.2011.153
DiMartino JF, Ayton PM, Chen EH et al (2002) The AF10 leucine zipper is required for leukemic transformation of myeloid progenitors by MLL-AF10. Blood 99:3780–5. doi:10.1182/blood.V99.10.3780
Egelhofer TA, Minoda A, Klugman S et al (2010) An assessment of histone-modification antibody quality. Nat Struct Mol Biol 18:91–93. doi:10.1038/nsmb.1972
Ehrentraut S, Hassler M, Oppikofer M et al (2011) Structural basis for the role of the Sir3 AAA+ domain in silencing: interaction with Sir4 and unmethylated histone H3K79. Genes Dev 25:1835–46. doi:10.1101/gad.17175111
Feng Q, Wang H, Ng HH et al (2002) Methylation of H3-Lysine 79 is mediated by a new family of HMTases without a SET domain. Curr Biol 12:1052–1058. doi:10.1016/S0960-9822(02)00901-6
Feng Y, Yang Y, Ortega MM et al (2010) Early mammalian erythropoiesis requires the Dot1L methyltransferase. Blood 116:4483–91. doi:10.1182/blood-2010-03-276501
FitzGerald J, Moureau S, Drogaris P et al (2011) Regulation of the DNA damage response and gene expression by the Dot1L histone methyltransferase and the 53Bp1 tumour suppressor. PLoS One 6:e14714. doi:10.1371/journal.pone.0014714
Frederiks F, Tzouros M, Oudgenoeg G et al (2008) Nonprocessive methylation by Dot1 leads to functional redundancy of histone H3K79 methylation states. Nat Struct Mol Biol 15:550–7. doi:10.1038/nsmb.1432
Fu H, Maunakea AK, Martin MM et al (2013) Methylation of histone H3 on lysine 79 associates with a group of replication origins and helps limit DNA replication once per cell cycle. PLoS Genet 9:e1003542. doi:10.1371/journal.pgen.1003542
Fuchs G, Hollander D, Voichek Y et al (2014) Cotranscriptional histone H2B monoubiquitylation is tightly coupled with RNA polymerase II elongation rate. Genome Res 24:1572–83. doi:10.1101/gr.176487.114
Giannattasio M, Lazzaro F, Plevani P, Muzi-Falconi M (2005) The DNA damage checkpoint response requires histone H2B ubiquitination by Rad6-Bre1 and H3 methylation by Dot1. J Biol Chem 280:9879–86. doi:10.1074/jbc.M414453200
Gibbons GS, Owens SR, Fearon ER, Nikolovska-Coleska Z (2015) Regulation of Wnt signaling target gene expression by the histone methyltransferase DOT1L. ACS Chem Biol 10:109–14. doi:10.1021/cb500668u
Greif PA, Tizazu B, Krause A et al (2008) The leukemogenic CALM/AF10 fusion protein alters the subcellular localization of the lymphoid regulator Ikaros. Oncogene 27:2886–2896. doi:10.1038/sj.onc.1210945
Guenther MG, Lawton LN, Rozovskaia T et al (2008) Aberrant chromatin at genes encoding stem cell regulators in human mixed-lineage leukemia. Genes Dev 22:3403–8. doi:10.1101/gad.1741408
Guppy BJ, McManus KJ (2015) Mitotic accumulation of dimethylated lysine 79 of histone h3 is important for maintaining genome integrity during mitosis in human cells. Genetics 199:423–33. doi:10.1534/genetics.114.172874
He N, Chan CK, Sobhian B et al (2011) Human polymerase-associated factor complex (PAFc) connects the super elongation complex (SEC) to RNA polymerase II on chromatin. Proc Natl Acad Sci U S A 108:E636–45. doi:10.1073/pnas.1107107108
Hein MY, Hubner NC, Poser I et al (2015) A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163:712–723. doi:10.1016/j.cell.2015.09.053
Ho L-L, Sinha A, Verzi M et al (2013) DOT1L-mediated H3K79 methylation in chromatin is dispensable for Wnt pathway-specific and other intestinal epithelial functions. Mol Cell Biol 33:1735–1745. doi:10.1128/MCB.01463-12
Holt MT, David Y, Pollock S et al (2015) Identification of a functional hotspot on ubiquitin required for stimulation of methyltransferase activity on chromatin. Proc Natl Acad Sci U S A 112:10365–10370. doi:10.1073/pnas.1504483112
Hornbeck PV, Kornhauser JM, Tkachev S et al (2012) PhosphoSitePlus: a comprehensive resource for investigating the structure and function of experimentally determined post-translational modifications in man and mouse. Nucleic Acids Res 40:261–270. doi:10.1093/nar/gkr1122
Huff JT, Plocik AM, Guthrie C, Yamamoto KR (2010) Reciprocal intronic and exonic histone modification regions in humans. Nat Struct Mol Biol 17:1495–9. doi:10.1038/nsmb.1924
Huyen Y, Zgheib O, Ditullio RA et al (2004) Methylated lysine 79 of histone H3 targets 53BP1 to DNA double-strand breaks. Nature 432:406–11. doi:10.1038/nature03114
Im H, Park C, Feng Q et al (2003) Dynamic regulation of histone H3 methylated at lysine 79 within a tissue-specific chromatin domain. J Biol Chem 278:18346–52. doi:10.1074/jbc.M300890200
Jack APM, Hake SB (2014) Getting down to the core of histone modifications. Chromosoma 123:355–371. doi:10.1007/s00412-014-0465-x
Janzen CJ, Hake SB, Lowell JE, Cross GAM (2006) Selective di- or trimethylation of histone H3 lysine 76 by two DOT1 homologs is important for cell cycle regulation in Trypanosoma brucei. Mol Cell 23:497–507. doi:10.1016/j.molcel.2006.06.027
Jones B, Su H, Bhat A et al (2008) The histone H3K79 methyltransferase Dot1L is essential for mammalian development and heterochromatin structure. PLoS Genet 4:e1000190. doi:10.1371/journal.pgen.1000190
Jonkers I, Kwak H, Lis JT (2014) Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. Elife 3:e02407. doi:10.7554/eLife.02407
Kim J, Guermah M, McGinty RK et al (2009) RAD6-mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137:459–71. doi:10.1016/j.cell.2009.02.027
Kim J, Hake SB, Roeder RG (2005) The human homolog of yeast BRE1 functions as a transcriptional coactivator through direct activator interactions. Mol Cell 20:759–70. doi:10.1016/j.molcel.2005.11.012
Kim S-K, Jung I, Lee H et al (2012a) Human histone H3K79 methyltransferase DOT1L methyltransferase binds actively transcribing RNA polymerase II to regulate gene expression. J Biol Chem 287:39698–39709. doi:10.1074/jbc.M112.384057
Kim W, Choi M, Kim J-E (2014) The histone methyltransferase Dot1/DOT1L as a critical regulator of the cell cycle. Cell Cycle 13:726–38. doi:10.4161/cc.28104
Kim W, Kim R, Park G et al (2012b) Deficiency of H3K79 histone methyltransferase Dot1-like protein (DOT1L) inhibits cell proliferation. J Biol Chem 287:5588–99. doi:10.1074/jbc.M111.328138
Kouskouti A, Talianidis I (2005) Histone modifications defining active genes persist after transcriptional and mitotic inactivation. EMBO J 24:347–57. doi:10.1038/sj.emboj.7600516
Krivtsov AV, Feng Z, Lemieux ME et al (2008) H3K79 methylation profiles define murine and human MLL-AF4 leukemias. Cancer Cell 14:355–68. doi:10.1016/j.ccr.2008.10.001
Kryczek I, Lin Y, Nagarsheth N et al (2014) IL-22(+)CD4(+) T cells promote colorectal cancer stemness via STAT3 transcription factor activation and induction of the methyltransferase DOT1L. Immunity 40:772–84. doi:10.1016/j.immuni.2014.03.010
Kuntimaddi A, Achille NJ, Thorpe J et al (2015) Degree of recruitment of DOT1L to MLL-AF9 defines level of H3K79 di- and tri-methylation on target genes and transformation potential. Cell Rep 11:808–820. doi:10.1016/j.celrep.2015.04.004
Leach BI, Kuntimaddi A, Schmidt CR et al (2013) Leukemia fusion target AF9 is an intrinsically disordered transcriptional regulator that recruits multiple partners via coupled folding and binding. Structure 21:176–183. doi:10.1016/j.str.2012.11.011
Leroy G, Dimaggio PA, Chan EY et al (2013) A quantitative atlas of histone modification signatures from human cancer cells. Epigenetics Chromatin 6:20. doi:10.1186/1756-8935-6-20
Li Y, Wen H, Xi Y et al (2014) AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell 159:558–571. doi:10.1016/j.cell.2014.09.049
Lin Y-H, Kakadia PM, Chen Y et al (2009) Global reduction of the epigenetic H3K79 methylation mark and increased chromosomal instability in CALM-AF10-positive leukemias. Blood 114:651–8. doi:10.1182/blood-2009-03-209395
Lu X, Simon MD, Chodaparambil JV et al (2008) The effect of H3K79 dimethylation and H4K20 trimethylation on nucleosome and chromatin structure. Nat Struct Mol Biol 15:1122–4. doi:10.1038/nsmb.1489
Margaritis T, Oreal V, Brabers N et al (2012) Two distinct repressive mechanisms for histone 3 lysine 4 methylation through promoting 3’-end antisense transcription. PLoS Genet 8:e1002952. doi:10.1371/journal.pgen.1002952
Marson A, Levine SS, Cole MF et al (2008) Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134:521–33. doi:10.1016/j.cell.2008.07.020
McGinty RK, Kim J, Chatterjee C et al (2008) Chemically ubiquitylated histone H2B stimulates hDot1L-mediated intranucleosomal methylation. Nature 453:812–6. doi:10.1038/nature06906
McGinty RK, Köhn M, Chatterjee C et al (2009) Structure-activity analysis of semisynthetic nucleosomes: mechanistic insights into the stimulation of Dot1L by ubiquitylated histone H2B. ACS Chem Biol 4:958–68. doi:10.1021/cb9002255
McLean CM, Karemaker ID, van Leeuwen F (2014) The emerging roles of DOT1L in leukemia and normal development. Leukemia 28:2131–8. doi:10.1038/leu.2014.169
Meyer C, Hofmann J, Burmeister T et al (2013) The MLL recombinome of acute leukemias in 2013. Leukemia 27:2165–2176. doi:10.1038/leu.2013.135
Milne TA, Kim J, Wang GG et al (2010) Multiple interactions recruit MLL1 and MLL1 fusion proteins to the HOXA9 locus in leukemogenesis. Mol Cell 38:853–63. doi:10.1016/j.molcel.2010.05.011
Min J, Feng Q, Li Z et al (2003) Structure of the catalytic domain of human DOT1L, a Non-SET domain nucleosomal histone methyltransferase. Cell 112:711–723. doi:10.1016/S0092-8674(03)00114-4
Minsky N, Shema E, Field Y et al (2008) Monoubiquitinated H2B is associated with the transcribed region of highly expressed genes in human cells. Nat Cell Biol 10:483–8. doi:10.1038/ncb1712
Mohan M, Herz H-M, Takahashi Y-H et al (2010) Linking H3K79 trimethylation to Wnt signaling through a novel Dot1-containing complex (DotCom). Genes Dev 24:574–89. doi:10.1101/gad.1898410
Mueller D, Bach C, Zeisig D et al (2007) A role for the MLL fusion partner ENL in transcriptional elongation and chromatin modification. Blood 110:4445–54. doi:10.1182/blood-2007-05-090514
Mueller D, García-Cuéllar M-P, Bach C et al (2009) Misguided transcriptional elongation causes mixed lineage leukemia. PLoS Biol 7:e1000249. doi:10.1371/journal.pbio.1000249
Muntean AG, Tan J, Sitwala K et al (2010) The PAF complex synergizes with MLL fusion proteins at HOX loci to promote leukemogenesis. Cancer Cell 17:609–21. doi:10.1016/j.ccr.2010.04.012
Musselman CA, Lalonde M-E, Côté J, Kutateladze TG (2012) Perceiving the epigenetic landscape through histone readers. Nat Struct Mol Biol 19:1218–27. doi:10.1038/nsmb.2436
Nakamura T, Alder H, Gu Y et al (1993) Genes on chromosomes 4, 9, and 19 involved in 11q23 abnormalities in acute leukemia share sequence homology and/or common motifs. Proc Natl Acad Sci U S A 90:4631–5. doi:10.1073/pnas.90.10.4631
Nguyen AT, Xiao B, Neppl RL et al (2011) DOT1L regulates dystrophin expression and is critical for cardiac function. Genes Dev 25:263–274. doi:10.1101/gad.2018511
Nguyen UTT, Bittova L, Müller MM et al (2014) Accelerated chromatin biochemistry using DNA-barcoded nucleosome libraries. Nat Methods 11:834–840. doi:10.1038/nmeth.3022
O’Connor CM, DiMaggio PA, Shenk T, Garcia BA (2014) Quantitative proteomic discovery of dynamic epigenome changes that control human cytomegalovirus (HCMV) infection. Mol Cell Proteomics 13:2399–2410. doi:10.1074/mcp.M114.039792
Okada Y, Feng Q, Lin Y et al (2005) hDOT1L links histone methylation to leukemogenesis. Cell 121:167–78. doi:10.1016/j.cell.2005.02.020
Oksenych V, Zhovmer A, Ziani S et al (2013) Histone methyltransferase DOT1L drives recovery of gene expression after a genotoxic attack. PLoS Genet 9:e1003611. doi:10.1371/journal.pgen.1003611
Onder TT, Kara N, Cherry A et al (2012) Chromatin-modifying enzymes as modulators of reprogramming. Nature 483:598–602. doi:10.1038/nature10953
Ontoso D, Acosta I, van Leeuwen F et al (2013) Dot1-dependent histone H3K79 methylation promotes activation of the Mek1 meiotic checkpoint effector kinase by regulating the Hop1 adaptor. PLoS Genet 9:e1003262. doi:10.1371/journal.pgen.1003262
Ontoso D, Kauppi L, Keeney S, San-Segundo PA (2014) Dynamics of DOT1L localization and H3K79 methylation during meiotic prophase I in mouse spermatocytes. Chromosoma 123:147–164. doi:10.1007/s00412-013-0438-5
Ooga M, Inoue A, Kageyama S et al (2008) Changes in H3K79 methylation during preimplantation development in mice. Biol Reprod 78:413–24. doi:10.1095/biolreprod.107.063453
Panier S, Boulton SJ (2014) Double-strand break repair: 53BP1 comes into focus. Nat Rev Mol Cell Biol 15:7–18. doi:10.1038/nrm3719
Park G, Gong Z, Chen J, Kim J-E (2010) Characterization of the DOT1L network: implications of diverse roles for DOT1L. Protein J 29:213–223. doi:10.1007/s10930-010-9242-8
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–12. doi:10.1002/jcc.20084
Pokholok DK, Harbison CT, Levine S et al (2005) Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122:517–27. doi:10.1016/j.cell.2005.06.026
Reisenauer MR, Anderson M, Huang L et al (2009) AF17 competes with AF9 for binding to Dot1a to up-regulate transcription of epithelial Na + channel alpha. J Biol Chem 284:35659–69. doi:10.1074/jbc.M109.038448
Reisenauer MR, Wang SW, Xia Y, Zhang W (2010) Dot1a contains three nuclear localization signals and regulates the epithelial Na + channel (ENaC) at multiple levels. Am J Physiol Renal Physiol 299:F63–76. doi:10.1152/ajprenal.00105.2010
Rothbart SB, Dickson BM, Raab JR et al (2015) An interactive database for the assessment of histone antibody specificity. Mol Cell 59:502–511. doi:10.1016/j.molcel.2015.06.022
Rubnitz J, Morrissey J, Savage P, Cleary M (1994) ENL, the gene fused with HRX in t(11;19) leukemias, encodes a nuclear protein with transcriptional activation potential in lymphoid and myeloid cells. Blood 84:1747–1752
Sabra M, Texier P, El Maalouf J, Lomonte P (2013) The Tudor protein survival motor neuron (SMN) is a chromatin-binding protein that interacts with methylated lysine 79 of histone H3. J Cell Sci 126:3664–77. doi:10.1242/jcs.126003
San-Segundo PA, Roeder GS (2000) Role for the silencing protein Dot1 in meiotic checkpoint control. Mol Biol Cell 11:3601–3615. doi:10.1091/mbc.11.10.3601
Sawada K, Yang Z, Horton JR et al (2004) Structure of the conserved core of the yeast Dot1p, a nucleosomal histone H3 lysine 79 methyltransferase. J Biol Chem 279:43296–306. doi:10.1074/jbc.M405902200
Schübeler D, MacAlpine DM, Scalzo D et al (2004) The histone modification pattern of active genes revealed through genome-wide chromatin analysis of a higher eukaryote. Genes Dev 18:1263–71. doi:10.1101/gad.1198204
Shanower GA, Muller M, Blanton JL et al (2005) Characterization of the grappa gene, the Drosophila histone H3 lysine 79 methyltransferase. Genetics 169:173–84. doi:10.1534/genetics.104.033191
Smolle M, Workman JL (2013) Transcription-associated histone modifications and cryptic transcription. Biochim Biophys Acta 1829:84–97. doi:10.1016/j.bbagrm.2012.08.008
Soria-Valles C, Osorio FG, Gutiérrez-Fernández A et al (2015) NF-κB activation impairs somatic cell reprogramming in ageing. Nat Cell Biol 17:1004–13. doi:10.1038/ncb3207
Steger DJ, Lefterova MI, Ying L et al (2008) DOT1L/KMT4 recruitment and H3K79 methylation are ubiquitously coupled with gene transcription in mammalian cells. Mol Cell Biol 28:2825–39. doi:10.1128/MCB.02076-07
Stein EM, Tallman MS (2015) Mixed lineage rearranged leukaemia: pathogenesis and targeting DOT1L. Curr Opin Hematol 22:92–6. doi:10.1097/MOH.0000000000000123
Stulemeijer IJE, De Vos D, van Harten K et al (2015) Dot1 histone methyltransferases share a distributive mechanism but have highly diverged catalytic properties. Sci Rep 5:9824. doi:10.1038/srep09824
Sun Z-W, Allis CD (2002) Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418:104–8. doi:10.1038/nature00883
Suzuki H, Takatsuka S, Akashi H et al (2011) Genome-wide profiling of chromatin signatures reveals epigenetic regulation of microRNA genes in colorectal cancer. Cancer Res 71:5646–5658. doi:10.1158/0008-5472.CAN-11-1076
Sweet SMM, Li M, Thomas PM et al (2010) Kinetics of re-establishing H3K79 methylation marks in global human chromatin. J Biol Chem 285:32778–86. doi:10.1074/jbc.M110.145094
Tan J, Jones M, Koseki H et al (2011) CBX8, a polycomb group protein, is essential for MLL-AF9-induced leukemogenesis. Cancer Cell 20:563–575. doi:10.1016/j.ccr.2011.09.008
Tessarz P, Kouzarides T (2014) Histone core modifications regulating nucleosome structure and dynamics. Nat Rev Mol Cell Biol 15:703–708. doi:10.1038/nrm3890
Tong Q, Cui G, Botuyan MV et al (2015) Structural plasticity of methyllysine recognition by the tandem tudor domain of 53BP1. Structure 23:312–21. doi:10.1016/j.str.2014.11.013
Tsunaka Y, Kajimura N, Tate S, Morikawa K (2005) Alteration of the nucleosomal DNA path in the crystal structure of a human nucleosome core particle. Nucleic Acids Res 33:3424–3434. doi:10.1093/nar/gki663
Vakoc CR, Sachdeva MM, Wang H, Blobel GA (2006) Profile of histone lysine methylation across transcribed mammalian chromatin. Mol Cell Biol 26:9185–95. doi:10.1128/MCB.01529-06
Van Leeuwen F, Gafken PR, Gottschling DE (2002) Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 109:745–756. doi:10.1016/S0092-8674(02)00759-6
Veloso A, Kirkconnell KS, Magnuson B et al (2014) Rate of elongation by RNA polymerase II is associated with specific gene features and epigenetic modifications. Genome Res 24:896–905. doi:10.1101/gr.171405.113
Vlaming H, van Leeuwen F (2012) Crosstalk between aging and the epigenome. Epigenomics 4:5–7. doi:10.2217/epi.11.113
Vlaming H, van Welsem T, de Graaf EL et al (2014) Flexibility in crosstalk between H2B ubiquitination and H3 methylation in vivo. EMBO Rep 15:1077–1084. doi:10.15252/embr.201438793
Wagner T, Robaa D, Sippl W, Jung M (2014) Mind the methyl: methyllysine binding proteins in epigenetic regulation. ChemMedChem 9:466–83. doi:10.1002/cmdc.201300422
Wakeman TP, Wang Q, Feng J, Wang X-F (2012) Bat3 facilitates H3K79 dimethylation by DOT1L and promotes DNA damage-induced 53BP1 foci at G1/G2 cell-cycle phases. EMBO J 31:2169–81. doi:10.1038/emboj.2012.50
Wang E, Kawaoka S, Yu M et al (2013) Histone H2B ubiquitin ligase RNF20 is required for MLL-rearranged leukemia. Proc Natl Acad Sci U S A 110:3901–6. doi:10.1073/pnas.1301045110
Wang X, Gao W, Ma X et al (2014) Dot1L mediated histone H3 lysine79 methylation is essential to meiosis progression in mouse oocytes. Neuro Endocrinol Lett 35:523–30
Wang Z, Zang C, Rosenfeld JA et al (2008) Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40:897–903. doi:10.1038/ng.154
Woo Park J, Kim K-B, Kim J-Y et al (2015) RE-IIBP methylates H3K79 and induces MEIS1-mediated apoptosis via H2BK120 ubiquitination by RNF20. Sci Rep 5:12485. doi:10.1038/srep12485
Wu H, Chen L, Zhang X et al (2013) Aqp5 is a new transcriptional target of Dot1a and a regulator of Aqp2. PLoS One 8:e53342. doi:10.1371/journal.pone.0053342
Wu H, Zeng H, Lam R et al (2011a) Structural and histone binding ability characterizations of human PWWP domains. PLoS One 6:e18919. doi:10.1371/journal.pone.0018919
Wu L, Li L, Zhou B et al (2014) H2B ubiquitylation promotes RNA Pol II processivity via PAF1 and pTEFb. Mol Cell 54:920–31. doi:10.1016/j.molcel.2014.04.013
Wu L, Zee BM, Wang Y et al (2011b) The RING finger protein MSL2 in the MOF complex is an E3 ubiquitin ligase for H2B K34 and is involved in crosstalk with H3 K4 and K79 methylation. Mol Cell 43:132–44. doi:10.1016/j.molcel.2011.05.015
Wu R, Yue Y, Zheng X, Li H (2015) Molecular basis for histone N-terminal methylation by NRMT1. Genes Dev 29:2337–42. doi:10.1101/gad.270926.115
Wysocki R, Javaheri A, Allard S et al (2005) Role of Dot1-dependent histone H3 methylation in G1 and S phase DNA damage checkpoint functions of Rad9. Mol Cell Biol 25:8430–43. doi:10.1128/MCB.25.19.8430-8443.2005
Yang L, Lin C, Jin C et al (2013) lncRNA-dependent mechanisms of androgen-receptor-regulated gene activation programs. Nature 500:598–602. doi:10.1038/nature12451
Yao X, Tang Z, Fu X, et al. (2015) The mediator subunit MED23 couples H2B mono-ubiquitination to transcriptional control and cell fate determination. EMBO J e201591279. doi: 10.15252/embj.201591279
Yokoyama A, Lin M, Naresh A et al (2010) A higher-order complex containing AF4 and ENL family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription. Cancer Cell 17:198–212. doi:10.1016/j.ccr.2009.12.040
Yu W, Chory EJ, Wernimont AK et al (2012) Catalytic site remodelling of the DOT1L methyltransferase by selective inhibitors. Nat Commun 3:1288. doi:10.1038/ncomms2304
Zee BM, Levin RS, Xu B et al (2010) In vivo residue-specific histone methylation dynamics. J Biol Chem 285:3341–50. doi:10.1074/jbc.M109.063784
Zhang L, Deng L, Chen F et al (2014) Inhibition of histone H3K79 methylation selectively inhibits proliferation, self-renewal and metastatic potential of breast cancer. Oncotarget 5:10665–10677. doi:10.18632/oncotarget.2496
Zhang W, Hayashizaki Y, Kone BC (2004) Structure and regulation of the mDot1 gene, a mouse histone H3 methyltransferase. Biochem J 377:641–51. doi:10.1042/BJ20030839
Zhang W, Xia X, Jalal DI et al (2006a) Aldosterone-sensitive repression of ENaCalpha transcription by a histone H3 lysine-79 methyltransferase. Am J Physiol Cell Physiol 290:C936–46. doi:10.1152/ajpcell.00431.2005
Zhang W, Xia X, Reisenauer MR et al (2006b) Dot1a-AF9 complex mediates histone H3 Lys-79 hypermethylation and repression of ENaCalpha in an aldosterone-sensitive manner. J Biol Chem 281:18059–68. doi:10.1074/jbc.M601903200
Zhang W, Xia X, Reisenauer MR et al (2007) Aldosterone-induced Sgk1 relieves Dot1a-Af9-mediated transcriptional repression of epithelial Na + channel alpha. J Clin Invest 117:773–83. doi:10.1172/JCI29850
Zhang Z, Zhang MQ (2011) Histone modification profiles are predictive for tissue/cell-type specific expression of both protein-coding and microRNA genes. BMC Bioinformatics 12:155. doi:10.1186/1471-2105-12-155
Zhu B, Mandal SS, Pham A-D et al (2005a) The human PAF complex coordinates transcription with events downstream of RNA synthesis. Genes Dev 19:1668–73. doi:10.1101/gad.1292105
Zhu B, Zheng Y, Pham A-D et al (2005b) Monoubiquitination of human histone H2B: the factors involved and their roles in HOX gene regulation. Mol Cell 20:601–11. doi:10.1016/j.molcel.2005.09.025
Acknowledgments
We thank Tessy Korthout for critical reading of the manuscript. This work was supported by the Netherlands Organisation for Scientific Research [NWO-VICI-016.130.627 to FVL] and the Dutch Cancer Society [KWF NKI2014-7232 and NKI2009-4511 to FVL].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Human and animal rights
The article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(XLSX 11 kb)
Rights and permissions
About this article
Cite this article
Vlaming, H., van Leeuwen, F. The upstreams and downstreams of H3K79 methylation by DOT1L. Chromosoma 125, 593–605 (2016). https://doi.org/10.1007/s00412-015-0570-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-015-0570-5