Abstract
In this report, we summarize recent findings about a role of the outer membrane metabolite channel VDAC/porin in protein import into mitochondria. Mitochondria fulfill key functions for cellular energy metabolism. Their biogenesis involves the import of about 1000 different proteins that are produced as precursors on cytosolic ribosomes. The translocase of the outer membrane (TOM complex) forms the entry gate for mitochondrial precursor proteins. Dedicated protein translocases sort the preproteins into the different mitochondrial subcompartments. While protein transport pathways are analyzed to some detail, only little is known about regulatory mechanisms that fine-tune protein import upon metabolic signaling. Recently, a dual role of the voltage-dependent anion channel (VDAC), also termed porin, in mitochondrial protein biogenesis was reported. First, VDAC/porin promotes as a coupling factor import of carrier proteins into the inner membrane. Second, VDAC/porin regulates the formation of the TOM complex. Thus, the major metabolite channel in the outer membrane VDAC/porin connects protein import to mitochondrial metabolism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mitochondria are known as the powerhouses of the cell. They produce the bulk of cellular energy via oxidative phosphorylation. The respiratory chain complexes generate a proton gradient across the inner membrane, which is used by the ATP synthase to produce ATP. Transport of metabolites across the two surrounding membranes is crucial for oxidative phosphorylation. The voltage-dependent anion channel (VDAC), also termed porin in baker’s yeast Saccharomyces cerevisiae, allows passage of small molecules across the outer membrane. Several specific carrier proteins mediate metabolite transport across the inner membrane (Palmieri and Pierri 2010; Colombini 2012; Krüger et al. 2017; Campo et al. 2017; Checchetto and Szabo 2018; Becker and Wagner 2018). In yeast, many of the proteins involved in oxidative metabolism are upregulated under respiratory growth compared to fermentation. Under these conditions, they represent more than 50% of the total mitochondrial protein content (Morgenstern et al. 2017). Thus, shift from fermentative to respiratory growth involves a tremendous production and import of mitochondrial proteins to ensure full capacity of oxidative phosphorylation (Zhang and Cao 2017; Pascual-Ahuir et al. 2018; Priesnitz and Becker 2018). The mechanisms that regulate mitochondrial protein content in response to metabolic signaling remain poorly understood.
Cytosolic ribosomes produce about 99% of the mitochondrial proteins. Molecular chaperones guide these preproteins to the mitochondrial surface. The translocases of the outer membrane (TOM complex) form the entry gate for the majority of mitochondrial preproteins. The TOM complex consists of seven different subunits: The β-barrel of Tom40 forms the protein-conducting channel and is essential for cell viability. Tom70 and Tom20 are the major receptors for incoming preproteins. The central receptor Tom22 transfers preproteins to Tom40 and is critical for the formation of the mature TOM complex. Finally, the three small Tom subunits Tom5, Tom6 and Tom7 promote assembly and stability of the protein translocase (Neupert and Herrmann 2007; Endo and Yamano 2009; Wiedemann and Pfanner 2017). After passage of the TOM channel, dedicated protein translocases sort preproteins into the mitochondrial subcompartments. There are two major protein sorting pathways into the inner membrane and matrix. The presequence translocase (TIM23 complex) sorts preproteins with a cleavable presequence into the inner membrane and mitochondrial matrix, while the carrier translocases (TIM22 complex) integrates carrier proteins into the inner membrane that lack a cleavable targeting signal. The membrane potential across the inner membrane drives transport of preproteins via both protein import pathways (Neupert and Herrmann 2007; Endo and Yamano 2009; Wiedemann and Pfanner 2017). The TOM and TIM23 complexes form contacts sites to transfer presequence-containing proteins to the inner membrane. Such molecular connection between the carrier translocase and the outer membrane was not reported. Here, small TIM proteins guide the hydrophobic carrier preproteins through the intermembrane space to the TIM22 translocase. How transport steps at both membranes are coordinated remained unknown.
Although we understand mitochondrial protein sorting in detail, only little is known how protein import is adapted to mitochondrial metabolism. Two recent studies identified an unexpected role of the major isoform of yeast VDAC/porin, Por1, in protein transport into mitochondria, linking protein import to mitochondrial metabolism (Ellenrieder et al. 2019; Sakaue et al. 2019).
Porin promotes import of carrier proteins
Baker’s yeast contains two isoforms of VDAC/porin: Por1 is present in a few hundred thousand copies, while only small amounts of Por2 have been detected per cell (Morgenstern et al. 2017). Por1 associates with the mature TOM complex in isolated mitochondria (Müller et al. 2016). However, whether it plays a role in protein import into mitochondria remained unknown. In a recent study, our group showed that loss of Por1 impaired selectively the transport of carrier proteins, while the import of presequence-containing precursors via the TIM23 complex remained unaffected (Ellenrieder et al. 2019). To reveal how Por1 mediates import of carrier proteins, single transport steps of this import pathway were analyzed (Ryan et al. 1999). Surprisingly, transport of carrier proteins across the outer membrane is only mildly affected in the absence of Por1, whereas the transfer of the preproteins from the TOM complex to the carrier translocase of the inner membrane is impaired (Ellenrieder et al. 2019). How does Por1 promote transfer of carrier proteins across the intermembrane space to the inner membrane? The anion selectivity of the Por1 channel appears not to be critical for its role in protein import. Por1 point mutants with changed ion selectivity were fully competent to drive import of carrier proteins. Instead, Por1 interacts with carrier precursors that are bound by small TIM proteins in the intermembrane space. Furthermore, it is associated with the carrier translocase to form a novel contact site between the outer and inner mitochondrial membrane (Ellenrieder et al. 2019). Altogether, Por1 acts as a coupling factor to coordinate outer and inner membrane transport steps of the carrier pathway (Fig. 1a).
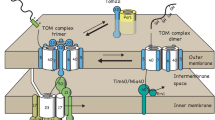
Porin plays a dual role in mitochondrial protein biogenesis. a The major VDAC/porin isoform in yeast, Por1, stimulates import of carrier proteins into the inner mitochondrial membrane. Cytosolic chaperones like Hsp70 and Hsp90 guide precursors of carrier proteins to the Tom70 receptor of the translocase of the outer membrane (TOM complex). After passage of the TOM channel, small TIM chaperones (Tim9/Tim10) guide these precursors to the carrier translocase (TIM22 complex) for insertion into the inner membrane. Por1 acts as a coupling factor to promote transfer of carrier precursors to the inner membrane. It is associated with the carrier precursor bound to the small TIM chaperones in the intermembrane space. Por1 further interacts with the TOM complex and the carrier translocase to spatially coordinate preprotein transport steps. b Por1 modulates the formation of the TOM complex. Por1 binds to unassembled Tom22 to control its integration into the TOM complex. In addition to Por1, Tom6 stabilizes the interaction of Tom40 and Tom22 with the translocase. The association of Tom22 and Tom40 is a critical step for the formation of the mature TOM complex. OM outer membrane, IM inner membrane, IMS intermembrane space
Porin regulates the assembly of the TOM complex
In a parallel study, Sakaue et al. (2019) reported that Por1 regulates the formation of the TOM complex. The TOM complex contains two to three pores. The central receptor Tom22 connects two Tom40 pores and is, therefore, critical for the assembly of the TOM complex (Künkele et al. 1998; Model et al. 2002; Becker et al. 2011; Shiota et al. 2015; Bausewein et al. 2017; Makki et al. 2019). Por1 binds to Tom22 that is not integrated into the TOM translocase (Sakaue et al. 2019). Two scenarios are feasible that can lead to the pool of free Tom22. First, the pool represents newly imported Tom22 that is not yet assembled in the TOM complex. Second, a portion of endogenous TOM complex could dissociate into a dimeric pore complex and free Tom22, which could facilitate the exchange of damaged subunits and support import of a subset of preproteins (Shiota et al. 2015; Sakaue et al. 2019). Por1 regulates the amount of free Tom22 and its assembly into the TOM complex. Upon loss of Por1, the unassembled pool of Tom22 is diminished and its integration into the mature TOM complex is enhanced (Sakaue et al. 2019; Ellenrieder et al. 2019). Tom6 stabilizes the Tom22–Tom40 association within the mature TOM complex (Alconada et al. 1995; Dekker et al. 1998; Sherman et al. 2005; Becker et al. 2011). In the absence of Tom6, the TOM complex dissociates and the unassembled Tom22 accumulates at Por1. Remarkably, cell cycle-dependent phosphorylation of Tom6 stabilizes the TOM complex and in turn decreases the Por1-bound Tom22 pool (Harbauer et al. 2014; Sakaue et al. 2019). Parallel loss of Por1 and Tom6 enables Tom22 to assemble into the TOM complex. However, this alternative TOM complex is not fully functional in importing preproteins (Sakaue et al. 2019). These findings illustrate that Por1 and Tom6 control the balance between the fully assembled TOM complex and its Tom22-free dimeric pore form (Sakaue et al. 2019). We speculate that the binding of Tom22 to Por1 represents a control point to prevent improper assembly of Tom22 into the TOM complex.
Perspectives
Protein transport pathways into mitochondria have been characterized in detail. Nevertheless, only little is known how protein transport is regulated (Priesnitz and Becker 2018). Previously, two major mechanisms have been reported. First, phosphorylation of TOM subunits by cytosolic kinases modulates mitochondrial protein biogenesis during the cell cycle and upon metabolic signaling (Schmidt et al. 2011; Geberth et al. 2013; Harbauer et al. 2014). Second, breakdown of the membrane potential across the inner membrane serves as a signal for mitochondrial damage. Under these conditions, preprotein import is impaired and stress response pathways are induced (Nargund et al. 2012; Lazarou et al. 2012). Two recent studies reported a dual role of Por1 in mitochondrial protein biogenesis. Por1 promotes import of carrier proteins and regulates the assembly of the TOM complex (Ellenrieder et al. 2019; Sakaue et al. 2019). The identified role of Por1 as a coupling factor in the carrier import pathway points to the exciting possibility that binding of protein translocases to partner proteins like Por1 represents a third mode to control protein import into mitochondria. The abundance of subunits of mitochondrial protein translocases remains largely unaffected upon respiratory growth, while the content of Por1 is raised about up to threefold (Morgenstern et al. 2017). Consequently, increased coupling by Por1 could stimulate import of carrier proteins under respiratory growth conditions to coordinate transport capacities for metabolites across the outer and inner membrane. Further observations support the view of a close link between mitochondrial protein biogenesis and metabolism. Respiratory chain complexes and the ADP/ATP carrier are associated with the TIM23 complex to support membrane potential-dependent import steps (van der Laan et al. 2006; Mehnert et al. 2014). In addition, the TIM22 translocase shares one subunit with the succinate dehydrogenase of the respiratory chain (Gebert et al. 2011). In human mitochondria, the carrier translocase is linked via its subunit acylglycerol kinase to lipid metabolism (Vukotic et al. 2017; Kang et al. 2017). Furthermore, outer membrane protein sorting is connected to organelle contact sites, which are important for lipid exchange between mitochondria and the endoplasmic reticulum or the vacuole (Kornmann et al. 2009; Lahiri et al. 2014; Murley et al. 2015; Elbaz-Alon et al. 2015; Ellenrieder et al. 2016; González-Montoro et al. 2018). Finally, the iron cluster assembly machinery is associated with the respiratory chain supercomplexes, which could play a role to adjust iron sulfur formation to the increasing demands during respiratory growth (Böttinger et al. 2018). Based on these examples, we propose that molecular coupling of protein translocases to partner proteins is an important mode to fine-tune protein biogenesis in response to metabolic state of mitochondria.
References
Alconada A, Kübrich M, Moczko M, Hönlinger A, Pfanner N (1995) The mitochondrial receptor complex: the small subunit Mom8b/Isp6 supports association of receptors with the general insertion pore and transfer preproteins. Mol Cell Biol 15:6196–6205
Bausewein T, Mills DJ, Langer JD, Nitschke B, Nussberger S, Kühlbrandt W (2017) Cryo-EM structure of the TOM core complex from Neurospora crassa. Cell 170:693–700
Becker T, Wenz LS, Thornton N, Stroud DA, Meisinger C, Wiedemann N, Pfanner N (2011) Biogenesis of mitochondria: dual role of Tom7 in modulating assembly of the preprotein translocase of the outer membrane. J Mol Biol 405:113–124
Becker T, Wagner R (2018) Mitochondrial outer membrane channels: emerging diversity in transport processes. BioEssays 40(7):1800013
Böttinger L, Mårtensson CU, Song J, Zufall N, Wiedemann N, Becker T (2018) Respiratory chain supercomplexes associate with the cysteine desulfurase complex of the iron-sulfur cluster assembly machinery. Mol Biol Cell 29:776–785
Campo ML, Peixoto PM, Martinez-Caballero S (2017) Revisiting trends on mitochondrial mega-channels for the import of proteins and nucleic acids. J Bioenerg Biomembr 49:75–99
Checchetto V, Szabo I (2018) Novel channels of the outer membrane of mitochondria: recent discoveries change our view. BioEssays 40:1700232
Colombini M (2012) VDAC structure, selectivity, and dynamics. Biochim Biophys Acta 1818:1457–1465
Dekker PJ, Ryan MT, Brix J, Müller H, Hönlinger A, Pfanner N (1998) Preprotein translocase of the outer mitochondrial membrane: molecular dissection and assembly of the general import pore complex. Mol Cell Biol 18:6515–6524
Elbaz-Alon Y, Eisenberg-Bord M, Shinder V, Stiller SB, Shimoni E, Wiedemann N, Geiger T, Schuldiner M (2015) Lam6 regulates the extent of contacts between organelles. Cell Rep 12:7–17
Ellenrieder L, Opaliński Ł, Becker L, Krüger V, Mirus O, Straub SP, Ebell K, Flinner N, Stiller SB, Guiard B, Meisinger C, Wiedemann N, Schleiff E, Wagner R, Pfanner N, Becker T (2016) Separating mitochondrial protein assembly and endoplasmic reticulum tethering by selective coupling of Mdm10. Nat Commun 7:13021
Ellenrieder L, Dieterle MP, Doan KN, Mårtensson CU, Floerchinger A, Campo ML, Pfanner N, Becker T (2019) Dual role of mitochondrial porin in metabolite transport across the outer membrane and protein transfer to the inner membrane. Mol Cell 73:1056–1065
Endo T, Yamano K (2009) Multiple pathways for mitochondrial protein traffic. Biol Chem 390:723–730
Gebert N, Gebert M, Oeljeklaus S, von der Malsburg K, Stroud DA, Kulawiak B, Wirth C, Zahedi RP, Dolezal P, Wiese S, Simon O, Schulze-Specking A, Truscott KN, Sickmann A, Rehling P, Guiard B, Hunte C, Warscheid B, van der Laan M, Pfanner N, Wiedemann N (2011) Dual function of Sdh3 in the respiratory chain and TIM22 protein translocase of the mitochondrial inner membrane. Mol Cell 44:811–818
Gerbeth C, Schmidt O, Rao S, Harbauer AB, Mikropoulou D, Opalińska M, Guiard B, Pfanner N, Meisinger C (2013) Glucose-induced regulation of protein import receptor Tom22 by cytosolic and mitochondria-bound kinases. Cell Metab 18:578–587
González-Montoro A, Auffarth K, Hönscher C, Bohnert M, Becker T, Warscheid B, Regiorri F, van der Laan M, Fröhlich F, Ungermann C (2018) Vps39 interacts with Tom40 to establish one of two functionally distinct vacuole-mitochondria contact sites. Dev Cell 45:621–636
Harbauer AB, Opalińska M, Gerbeth C, Herman JS, Rao S, Schönfisch B, Guiard B, Schmidt O, Pfanner N, Meisinger C (2014) Cell cycle-dependent regulation of mitochondrial preprotein translocase. Science 346:1109–1113
Kang Y, Stroud DA, Baker MJ, De Souza DP, Frazier AE, Liem M, Tull D, Mathivanan S, McConville MJ, Thorburn DR, Ryan MT, Stojanovski D (2017) Sengers syndrome-associated mitochondrial acylglycerol kinase is a subunit of the human TIM22 protein import complex. Mol Cell 67:457–475
Kornmann B, Currie E, Collins CR, Schuldiner M, Nunnari J, Weissman JS, Walter P (2009) An ER-mitochondria tethering complex revealed by a synthetic biology screen. Science 325:477–481
Krüger V, Becker T, Becker L, Montilla-Martinez M, Ellenrieder L, Vögtle FN, Meyer H, Ryan M, Wiedemann N, Warscheid B, Pfanner N, Wagner R, Meisinger C (2017) Identification of new channels by systematic analysis of the mitochondrial outer membrane. J Cell Biol 216:3485–3495
Künkele KP, Heins S, Dembowski M, Nargang FE, Benz R, Thieffry M, Walz J, Lill R, Nussberger S, Neupert W (1998) The preprotein translocation channel of the outer membrane of mitochondria. Cell 93:1009–1019
Lahiri S, Chao JT, Tavassoli S, Wong AK, Choudhary V, Young BP, Loewen CJ, Prinz WA (2014) A conserved endoplasmic reticulum membrane protein complex (EMC) facilitates phospholipid transfer from the ER to mitochondria. PLoS Biol 12:e1001969
Lazarou M, Jin SM, Kane LA, Youle RJ (2012) Role of PINK1 binding to the TOM complex and alternate intracellular membranes in recruitment and activation of the E3 ligase Parkin. Dev Cell 22:320–333
Makki A, Rada P, Žárský V, Kereïche S, Kováčik L, Novotný M, Jores T, Rapaport D, Tachezy J (2019) Triplet-pore structure of a highly divergent TOM complex of hydrogenosomes in Trichomonas vaginalis. PLoS Biol 17:e3000098
Mehnert CS, Rampelt H, Gebert M, Oeljeklaus S, Schrempp SG, Kochbeck L, Guiard B, Warscheid B, van der Laan M (2014) The mitochondrial ADP/ATP carrier associates with the inner membrane presequence translocase in a stoichiometric manner. J Biol Chem 289:27352–27362
Model K, Prinz T, Ruiz T, Rademacher M, Krimmer T, Kühlbrandt W, Pfanner N, Meisinger C (2002) Protein translocase of the outer mitochondrial membrane: role of import receptors in the structural organization of the TOM complex. J Mol Biol 316:657–666
Morgenstern M, Stiller SB, Lübbert P, Peikert CD, Dannenmaier S, Drepper F, Weill U, Höß P, Feuerstein R, Gebert M, Bohnert M, van der Laan M, Schuldiner M, Schütze C, Oeljeklaus S, Pfanner N, Wiedemann N, Warscheid B (2017) Definition of a high-confidence mitochondrial proteome at quantitative scale. Cell Rep 19:2836–2852
Müller CS, Bildl W, Haupt A, Ellenrieder L, Becker T, Hunte C, Fakler B, Schulte U (2016) Cryo-slicing blue native-mass spectrometry (csBN-MS): a novel technology for high-resolution complexome profiling. Mol Cell Proteomics 15:669–681
Murley A, Sarsa RD, Toulmay A, Yamada J, Prinz WA, Nunnari J (2015) Ltc1 is an ER-localized sterol transporter and a component of ER-mitochondria and ER-vacuole contacts. J Cell Biol 209:539–548
Nargund AM, Pellegrino MW, Fiorese CJ, Baker BM, Haynes CM (2012) Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337:587–590
Neupert W, Herrmann JM (2007) Translocation of proteins into mitochondria. Annu Rev Biochem 76:723–749
Palmieri F, Pierri CL (2010) Mitochondrial metabolite transport. Essays Biochem 47:37–52
Pascual-Ahuir A, Manzanares-Estreder Timón-Gómez A, Proft M (2018) Ask yeast how to burn your fats: lessons learned from the metabolic adaption to salt stress. Curr Genet 64:63–69
Priesnitz C, Becker T (2018) Pathways to balance mitochondrial translation and protein import. Genes Dev 32:1285–1296
Ryan MT, Müller H, Pfanner N (1999) Functional staging of ADP/ATP carrier translocation across the outer mitochondrial membrane. J Biol Chem 274:20619–20627
Sakaue H, Shiota T, Ishizaka N, Kawano S, Tamura Y, Tan KS, Imai K, Motono C, Hirokawa T, Taki K, Miyata N, Kuge O, Lithgow T, Endo T (2019) Porin associates with Tom22 to regulate the mitochondrial protein gate assembly. Mol Cell 73:1044–1055
Schmidt O, Harbauer AB, Rao S, Eyrich B, Zahedi RP, Stojanovski D, Schönfisch B, Guiard B, Sickmann A, Pfanner N, Meisinger C (2011) Regulation of mitochondrial protein import by cytosolic kinases. Cell 144:227–239
Sherman EL, Go NE, Nargang FE (2005) Functions of the small proteins in the TOM complex of Neurospora crassa. Mol Biol Cell 16:4172–4182
Shiota T, Imai K, Qiu J, Hewitt VL, Tan K, Shen HH, Sakiyama N, Fukasawa Y, Hayat S, Kamiya M, Elofsson A, Tomii K, Horton P, Wiedemann N, Pfanner N, Lithgow T, Endo T (2015) Molecular architecture of the active mitochondrial protein gate. Science 349:1544–1548
van der Laan M, Wiedemann N, Mick DU, Guiard B, Rehling P, Pfanner N (2006) A role for Tim21 in membrane potential-dependent preprotein sorting in mitochondria. Curr Biol 16:2271–2276
Vukotic M, Nolte H, König T, Saita S, Ananjew M, Krüger M, Tatsuta T, Langer T (2017) Acylglycerol kinase mutated in Sengers syndrome is a subunit of the TIM22 protein translocase in mitochondria. Mol Cell 67:471–490
Wiedemann N, Pfanner N (2017) Mitochondrial machineries for protein import and assembly. Annu Rev Biochem 86:685–714
Zhang N, Cao L (2017) Starvation signals in yeast are integrated to coordinate metabolic reprogramming and stress responses to ensure longevity. Curr Genet 63:839–843
Acknowledgements
The work was supported by grants of the Deutsche Forschungsgemeinschaft (BE 4679/2–2), Research Training Group 278002225/RTG 2202 and Germany’s Excellence Strategy (CIBSS-EXC-2189—Project ID 390939984). Work included in this study has also been performed in partial fulfillment of the requirements for the doctoral thesis of K.N.D. at the University of Freiburg.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Doan, K.N., Ellenrieder, L. & Becker, T. Mitochondrial porin links protein biogenesis to metabolism. Curr Genet 65, 899–903 (2019). https://doi.org/10.1007/s00294-019-00965-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-00965-z