Abstract
Chemical activities of hydrophobic substances can determine the windows of environmental conditions over which microbial systems function and the metabolic inhibition of microorganisms by benzene and other hydrophobes can, paradoxically, be reduced by compounds that protect against cellular water stress (Bhaganna et al. in Microb Biotechnol 3:701–716, 2010; Cray et al. in Curr Opin Biotechnol 33:228–259, 2015a). We hypothesized that this protective effect operates at the macromolecule structure–function level and is facilitated, in part at least, by genome-mediated adaptations. Based on proteome profiling of the soil bacterium Pseudomonas putida, we present evidence that (1) benzene induces a chaotrope-stress response, whereas (2) cells cultured in media supplemented with benzene plus glycerol were protected against chaotrope stress. Chaotrope-stress response proteins, such as those involved in lipid and compatible-solute metabolism and removal of reactive oxygen species, were increased by up to 15-fold in benzene-stressed cells relative to those of control cultures (no benzene added). By contrast, cells grown in the presence of benzene + glycerol, even though the latter grew more slowly, exhibited only a weak chaotrope-stress response. These findings provide evidence to support the hypothesis that hydrophobic substances induce a chaotropicity-mediated water stress, that cells respond via genome-mediated adaptations, and that glycerol protects the cell’s macromolecular systems. We discuss the possibility of using compatible solutes to mitigate hydrocarbon-induced stresses in lignocellulosic biofuel fermentations and for industrial and environmental applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Benzene is a noxious hydrophobic substance that is sufficiently soluble to be bioavailable, is highly inhibitory to the majority of microbial species, can bioaccumulate along food chains, and poses a significant risk to animal and human health (Lamm and Grunwald 2006; Kelly et al. 2007; Radniecki et al. 2008; Hamada et al. 2009; Littlejohns et al. 2010). Hydrophobic substances, including benzene, are environmentally ubiquitous and also occur in microbe-driven experimental and industrial processes/systems. These include hydrocarbons derived from plants, oil, agrochemicals (e.g., pesticides), landfill leachate, industrial solvents, and biotechnological substrates and products (Keane et al. 2008; Koopman et al. 2010; Timmis 2010; Womack et al. 2010; Escapa et al. 2012; Cray et al. 2015a). Hydrocarbons—including a considerable number of the substances collectively known as volatile organic compounds—are well known as cellular toxicants or stressors of microbial cells (e.g., Velázquez et al. 2005; Cray et al. 2013a, 2015a; Suryawanshi et al. 2015); those which partition into the hydrophobic domains of cellular macromolecules and membranes tend to disorder macromolecular systems (Sikkema et al. 1995; Bhaganna et al. 2010).Footnote 1
The various activities of solutes, such as water activity, Hofmeister effects, ionic strength, and chaotropicity, can determine the biotic windows of microbial species, the composition of microbial communities, and the extent of the microbial biosphere (Rummel et al. 2014; Ball and Hallsworth 2015; Santos et al. 2015; Stevenson et al. 2015a, b). A number of studies have demonstrated that interactions between solutes with different/opposing chemical activities can redefine, and even extend, the growth windows for microbial cells in saline systems (see Hallsworth et al. 2007; Oren and Hallsworth 2014; Yakimov et al. 2015). Additional studies, of cellular and enzyme activity, provided evidence that protectants against cellular water stress (i.e., compatible solutes such as glycerol) reverse the inhibitory action of soluble chaotropes, such as ethanol, and hydrophobic stressors, such as toluene and benzene (Hallsworth 1998; Joo et al. 2000; Hallsworth et al. 2003a; Bhaganna et al. 2010; Trevisol et al. 2011; Bell et al. 2013; Alves et al. 2015).
Whereas whole-cell studies and macromolecular assays provide evidence for a protective effect of compatible solutes against hydrocarbon stressors including benzene (see above), there is a paucity of information on genome-mediated responses in relation to this aspect of cellular biology. Indeed, relatively little is known about interactions between diverse stress parameters and solute activities in relation to windows for microbial metabolism and cell division (Chin et al. 2010; Harrison et al. 2013), hence the recent interest in the physical and chemical parameters that constrain and/or permit life processes (e.g., Stevenson and Hallsworth 2014; Harrison et al. 2015; Lievens et al. 2015). The role of compatible solutes in microbial ecology, stress biology, and biotechnology has been relatively well studied,Footnote 2 yet aspects of the biophysics and ecophysiology of glycerol remain enigmatic. The aims of the current study were to: (1) see whether microbial stress induced by benzene (which is a paradigm for hydrophobic stressors) and (2) characterize the response of cells to benzene stress versus benzene + glycerol, using proteomic profiling of Pseudomonas putida. This soil bacterium was used as a model system because: it has a well-characterized proteome; hydrocarbons are central to its ecology; chaotropicity-induced stresses were first characterized using this species (Hallsworth et al. 2003b; Bhaganna et al. 2010); of the key roles that pseudomonads play in bioremediation, industrial biocatalysis, and biological control (Timmis 2010; Hassan et al. 2011; Cray et al. 2013a); and, for the strain selected for study (KT2440), it is unable to degrade benzene.
Materials and methods
Bacterial strain, media, culture conditions, and growth-rate determination
Pseudomonas putida strain KT2440 (DSMZ 6125) that has been well characterized at the genome-and proteome level (see Nelson et al. 2002; Heim et al. 2003; Ballerstedt et al. 2007) was obtained from the German Collection of Microorganisms and Cell Cultures (DSMZ, Braunschweig, Germany) and maintained in a liquid minimal mineral salt medium with glucose as the sole carbon substrate and NH4Cl as the sole nitrogen substrate (pH 7.4; for details, see Hartmans et al. 1989). All cultures were grown in 50 ml medium in serum bottles (100 ml, Supelco, Dorset, UK) sealed with aluminum caps with PTFE/butyl septa to avoid volatilization of added stressors, and incubated in a shaking incubator (New Brunswick Innova 44, set at 120 rpm) at 30 °C. Control cultures contained no added stressor or compatible solute. In a second batch of media, benzene was used to supplement media at a concentration (5.2 mM) that induced approximately 50 % growth-rate inhibition relative to the control (see Bhaganna et al. 2010). In the third batch of media, both benzene (5.2 mM) and glycerol (0.52 M) were used to supplement the medium. Glycerol was incorporated prior to autoclaving, whereas benzene was added after media were autoclaved and allowed to cool as described previously (Bhaganna et al. 2010). Whereas water potential is typically used to quantify water availability for wood-rotting fungi or other microbes which can be exposed to high matric potential (e.g. Kashangura et al. 2006), the water activities of culture media were determined for consistency with other studies of bacteria (e.g. Hallsworth et al. 2003b; Stevenson and Hallsworth, 2014; Stevenson et al. 2015a). The water activity values were: control medium, 0.999; benzene-supplemented medium, 0.999; and benzene + glycerol medium, 0.995, determined at 30 °C using a Novasina IC-II water activity machine fitted with an alcohol-resistant humidity sensor and alcohol filter—Novasina, Pfäffikon, Switzerland—as described previously (Hallsworth and Nomura 1999; Stevenson et al. 2015a). Inoculations were carried out through the caps’ septa using disposable syringes (1 ml) fitted with needles. All media were inoculated using cell suspensions from exponentially growing pre-cultures (0.4–0.6 ml) to give a turbidity reading (560 nm) at time zero between 0.1 and 0.15. Samples were taken at hourly or two-hourly intervals, and growth rates during the exponential growth phase were calculated from the slopes of the growth curves. All these experiments were carried out using three biological replicates.
Cell harvesting and protein extraction
Cells were harvested during exponential growth, once the turbidity had reached approximately 0.45, by centrifugation at 4 °C at 10,000 g for 15 min, and then resuspended in phosphate-buffered saline (PBS: 10 mM sodium phosphate buffer, pH 7.4, 138 mM NaCl; Hallsworth et al. 2003b). Harvesting, protein extraction, and proteome analysis were carried out using two technical replicates. Harvested cells were washed twice in PBS, resuspended in PBS buffer, aliquoted in microfuge tubes, pelleted by centrifugation, and then stored at −20 °C overnight as described previously (Hallsworth et al. 2003b). For protein extraction, a cell pellet was allowed to thaw on ice, reswelling solution added, and the suspension sonicated as described by Hallsworth et al. (2003b). The microfuge tubes were allowed to stand at room temperature (15 min), and cell debris and salts were removed as described previously (Hallsworth et al. 2003b; Bobadilla Fazzini et al. 2010). The resulting protein pellets were air-dried and resuspended in 300 ml reswelling solution, and the protein concentration was determined according to Bradford (1976).
Two-dimensional gel electrophoresis and identification of proteins
Two-dimensional gel electrophoresis was carried out according to Görg (1991). Each gel was loaded with proteins extracted from an equivalent cell biomass. Isoelectric focusing was based on the method used by Bobadilla Fazzini et al. (2013): Analytical determinations were made using the Bradford test (150 µg protein mixture) and made up to 300 µl using rehydration solution in the presence of ampholytes/under reducing conditions on ReadyStrip IPG strips (17 cm, pH 3–10). Passive rehydration (2 h, 20 °C) took place on the focusing tray following which samples were covered with silicon oil. Active rehydration was then performed (50 V for 12 h), and isoelectric focusing (10,000 V on Protean® IEF cell; Bio-Rad) was carried out to 75 kWh. Samples were then stored at −70 °C.
The focused strips were then transferred to 1.5-mm-thick gradient sodium dodecyl sulfate (12 %, w/v)–polyacrylamide (15 %, w/v) gels and developed using the IsoDalt system (Amersham Pharmacia Biotech) overnight. The gels were treated with trichloroacetic acid (10 % w/v), stained with Coomassie brilliant blue G-250 (0.1 % w/v) overnight, and then destained using distilled water (Figures S1–S3; Bobadilla Fazzini et al. 2013). The gel images were scanned using a molecular imager GS-800 calibrated densitometer (Bio-Rad), and treatment gels were compared with control gels using Z3 v3.0.7 image-analysis software (Compugen, San Jose, CA) for protein differential expression analysis (Figure S4). Protein identification was carried out using protein spots with differential expression under the experimental treatments; visual analysis confirmed that protein spots on replicate gels did not differ and so spots from one replicate in each case were used for protein identification (Hathout et al. 2002; Hallsworth et al. 2003b). The two replicates from each condition showed high reproducibility by software comparison (our earlier studies also confirm the consistency of biological replicates; Bobadilla Fazzini et al. 2010). For comparative analysis, the relative intensity of each pair of corresponding spots was measured and the data displayed as differential expression values. The data were then checked manually to exclude any error in matching pairs and to judge the significance of the differential analysis by zooming in the spot of interest and checking its incidence in the other set of gels (Hathout et al. 2002). Only matching spots were excised and analyzed. Differentially expressed proteins were selected using a fold-change cutoff: Increases of ≥2.5 were indicative of up-regulation and fold changes of ≤0.5 were indicative of down-regulation. Protein spots (Figure S5) were cut out of gels manually, destained, digested, eluted, and desalted as described by Bobadilla Fazzini et al. (2013): Spots were destained and digested overnight using sequence-grade modified trypsin (Promega); the peptides were eluted/desalted using ZipTip® (Millipore). Samples were loaded along with α-cyano-4-hydroxycinnamic acid matrix for mass spectrometer analysis and analyzed using an Ultraflex II ToF (Bruker Daltonics). Spectra were analyzed using FlexAnalysis 2.0 and Biotools 2.2 software (Bruker Daltonics) and searching carried out on the NCBInr database using Profound version 4.10.5 (Proteometrics). For ESI Q-ToF analysis, 3 µl of sample was directly analyzed after Zip-Tip® elution in a Micromass Q-ToF micro TM mass spectrometer (Waters Corporation) and sequencing analysis carried out using MassLynx Mass Spectrometry Software 4.0 (Waters Corporation). Sequence similarity searching against protein databases was performed using FASTA (European Bioinformatics Institute, Cambridge, UK, at http://www.ebi.ac.uk/fasta33/).
The peptide mass fingerprints obtained were identified using a P. putida (KT2440) protein database that was constructed in conjunction with a genome sequencing study (Nelson et al. 2002). Proteins were classified according to their presumed primary metabolic function as described by Hallsworth et al. (2003b), Volkers et al. (2006), and Bhaganna et al. (2010).
Results
Proteomic profiling of benzene-stressed P. putida cells
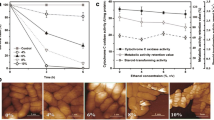
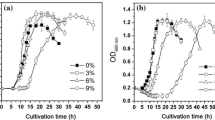
As expected, bacterial cultures in benzene-supplemented media (5.2 mM benzene) were inhibited relative to the control cultures that had no benzene added (by 46 %, based on a comparison of exponential-phase growth rates, see Figs. 1, 2a). Analysis of the proteome profiles for benzene-stressed cells revealed changes in a total of 23 proteins, 15 of which were up-regulated and eight did not change significantly relative to those in the control; there were no down-regulated proteins (Table 1). These proteins were listed according to their primary metabolic function within two overall categories: those that are known descriptors for chaotropicity-mediated stress and those known to be general stress adaptations/responses (Table 1). There was a low level of general stress response: there was no apparent up-regulation of proteins involved in protein synthesis and there were only three proteins involved in energy metabolism that were up-regulated by ≥fourfold (Table 1). By contrast, there were increases (≥twofold) of ~10 proteins that, collectively, act as descriptors of chaotropicity-mediated stress. These include proteins associated with lipid metabolism and membrane composition, protein stabilization, compatible-solute transport and metabolism, and nucleic-acid protection/synthesis/repair, as well as those involved in responses to oxidative-stress (see Hallsworth 1998; Hallsworth et al. 2003b; Alves et al. 2015; Ball and Hallsworth 2015; Cray et al. 2015a). Furthermore, there was a tenfold to 15-fold up-regulation of five of these proteins (involved in compatible-solute transport and oxidative-stress response) relative to the control, see Fig. 2.
Growth curves of P. putida (30 °C) cultured in a defined liquid minimal mineral salt medium (see “Materials and methods” section) without any stressor (control; black line), supplemented with benzene (5.2 mM; red line), and benzene + glycerol (5.2 mM and 0.52 M, respectively; blue line). Stress tolerance assays were carried out in triplicate; standard deviations are shown
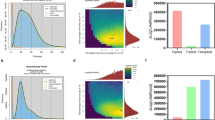
Expression profiling of protein descriptors for chaotrope stress in exponential growth-phase P. putida cells cultured in media supplemented with a benzene + glycerol (5.2 mM and 0.52 M, respectively; 60 % growth-rate inhibition relative to the control—see dashed line) and b benzene (5.2 mM; 46 % growth-rate inhibition—see dashed line). The color coding denotes primary protein function as indicated on the x-axis; columns show fold changes relative to the control except for pale-red and pale-green columns which indicate proteins not present in control gels (plotted arbitrarily at the twofold level). All experiments were carried out in triplicate
Proteomic profiling of benzene + glycerol-grown cells
The high level of growth-rate inhibition observed in glycerol-protected, benzene-grown cultures, i.e., 60 % relative to the control (Figs. 1, 2a), was consistent with the intense stress-response of these cells. Among the 16 up-regulated proteins, there were 12 that were up-regulated between approximately four- and 70-fold that are involved primarily in protein synthesis and energy metabolism (Table 2). In contrast to the strong up-regulation of the protein descriptors for chaotrope stress in benzene-supplemented cultures (Table 1), glycerol-protected benzene-stressed cells only showed a weak level of chaotrope-stress response: despite the higher level of growth-rate inhibition, the up-regulation of chaotrope-stress descriptor proteins was limited to between ~twofold and fivefold, except for two proteins (DNA-binding stress protein PP1210 and amino acid ABC transporter, periplasmic amino acid-binding protein PputGB1 5004) that were up-regulated by ~sevenfold (Fig. 2a).
Discussion
Whereas the P. putida cells cultured in benzene (5.2 mM)-supplemented media were inhibited, their growth rate was nevertheless >50 % that of the control cultures, indicating that these cells were not close to the point of failure (for P. putida strain KT 2440, system failure occurs at ~9 mM benzene; Bhaganna et al. 2010).Footnote 3 This observation is consistent with the weakness of the stress response observed (Table 1). Proteins up-regulated in the presence of benzene (current study) are consistent with those up-regulated in the presence of chaotropic solutes such as ethanol, phenol, and benzyl alcohol (Hallsworth et al. 2003b). These included fadB1X (involved in lipid metabolism) and catalase/peroxidase HPI (removal of reactive oxygen species). Chaotropic substances induce oxidative-stress responses, regardless of microbial species, as a consequence of a breakdown in homeostasis (e.g., Alexandre et al. 2001; Russo et al. 2001; Tam et al. 2006; Wierckx et al. 2008; Ball and Hallsworth 2015; Cray et al. 2015a). Chaotropes cannot cause oxidative damage to cellular components directly, and it is likely that chaotrope-induced oxidative-stress results from disordering of cellular membranes and consequent disruption of electron transport processes that causes a release of free radicals (resulting in lipid peroxidation; Zager 1996; Russo et al. 2001; Hallsworth et al. 2003b; Albano 2006; Kavazis et al. 2009). In the present study, the katG-encoded, catalase/peroxidase HPI was up-regulated 15.1-fold in cells from the benzene-only treatment (Table 1), but not detected in cells treated with benzene + glycerol (Table 2). Up-regulation of this enzyme has previously been observed in P. putida cells exposed to phenol, pyruvate (Kurbatov et al. 2006), and the chlorophenoxy herbicides such as 2,4-dichlorophenoxyacetic acid, 2-(2,4-dichlorophenoxy) propanoic acid, and 2,4-dichlorophenol (Benndorf et al. 2006). The apparent lack of catalase/peroxidase HPI in P. putida cells from the benzene + glycerol treatment may be a consequence of the reduced level of benzene stress (see Bhaganna et al. 2010) and/or the ability of glycerol to scavenge reactive oxygen species (Smirnoff and Cumbes 1989).
General stress responses are characterized by an increased requirement for energy generation and an up-regulation of proteins involved in protein synthesis (Park et al. 2001; Hallsworth et al. 2003b; Veláquez et al. 2005; Domínguez-Cuevas et al. 2006). In the current study, P. putida cells up-regulated the sugar transport protein ABC transporter PP1015 under benzene stress (Table 1), but not in the benzene + glycerol treatment (Table 2). This is consistent with the increased energy required and increased sugar uptake observed in P. putida cells exposed to toluene stress (Segura et al. 2005). Segura et al. (2005) found additional evidence of the need to increase energy generation in response to toluene, as two enzymes of the tricarboxylic acid cycle were induced: dihydrolipoamide dehydrogenase (E3 component of 2-oxoglutarate dehydrogenase complex) and succinyl coenzyme A synthetase. Additional studies, of a transporter-deficient P. putida mutant, indicated that the ABC transporter is implicated in tolerance to toluene shock (at 1 %, v/v) (Kim et al. 1998); this study also demonstrated that an increased uptake of metabolites supports the high energy demands. F1F0–ATP synthase, up-regulated in benzene-stressed cells, is known to play roles in energy generation as well as proton translocation (Len et al. 2004). This protein may therefore be required by the cell in response to a breakdown in trans-bilayer proton gradients induced by the chaotropicity of benzene.
The periplasmic amino acid-binding protein PP3593 was up-regulated 2.2-fold in P. putida cells exposed to benzene (Table 1), but not those in the benzene + glycerol treatment (Table 2). Conversely, there was up-regulation of the periplasmic amino acid-binding proteins PputGB1 5004 and PputGB1_3991 in cells exposed to benzene + glycerol (Table 2), but not in the benzene-only treatment (Table 1). The general amino acid ABC transporter, periplasmic binding protein aapj, was up-regulated under both benzene and benzene + glycerol treatments (Tables 1, 2). However, the fold increase was notably larger under benzene stress (Table 1). Up-regulation of proteins involved in the transport and metabolism of amino acid has similarly been observed in cells of P. putida exposed to sublethal concentrations of phenol, pyruvate, toluene, tetracycline, and benzoate (Santos et al. 2004; Segura et al. 2005; Kurbatov et al. 2006; Yun et al. 2006, 2011). Enhanced uptake of amino acids via ABC transport proteins may relate to both accumulation of compatible solutes (see below) and the efflux of stressors/toxic metabolites in P. putida KT2440 (Nelson et al. 2002; Dos Santos et al. 2004).
Studies of the stress metabolism of diverse microbial species demonstrate that tolerance of diverse cellular stresses is enhanced by up-regulation of proteins involved in protein stabilization (e.g., heat-shock proteins and chaperonins), import or synthesis and retention of compatible solutes, and by actively transporting protectants to the extracellular matrix in order to protect the exterior of the plasma membrane (Eleutherio et al. 1993; Hocking 1993; Albertyn et al. 1994; Kets et al. 1996a; Cray et al. 2015a). Whereas P. putida is known to produce high levels of protein stabilization proteins in response to chaotropic solutes and hydrophobic stressors, as do other bacterial species and eukaryotic microbes (Hallsworth et al. 2003b; Bhaganna et al. 2010; Cray et al. 2015a), this response was not so evident in strain KT2440 under benzene stress at the benzene concentrations and under the conditions used in the current study. Both amino acids and carbohydrates can act as compatible solutes in P. putida, e.g., glycine, betaine, glutamate, N-acetylglutaminylglutamine amide, glycerol, mannitol, and trehalose (see Kets et al. 1996b; Joo et al. 2000; Cray et al. 2013a). Glycerol, which is well known for its properties as a cryoprotectant, osmolyte, and protectant of macromolecular structures under solute-induced stresses (see Brown 1990; Hallsworth et al. 1998, 2003b; Hubalek 2003), has protective activity against benzene (though genome-mediated responses were not investigated; Bhaganna et al. 2010) hence the selection of glycerol as the focus of the current study.
At molar concentrations, glycerol can itself induce cellular inhibition, and this may relate to viscosity (see “Materials and methods” section; Chirife et al. 1984; Wyatt et al. 2015a, 2015 b), water activity reduction, and its own chaotropic activity (see Williams and Hallsworth 2009; Cray et al. 2013b; Alves et al. 2015). Benzene + glycerol-grown cells in the current study were more inhibited than those exposed to benzene alone (see Figs. 1, 2), but the relatively low concentration of glycerol used to supplemented culture media (i.e., 0.52 M) was an order of magnitude lower than would be needed for glycerol to behave in a chaotropic manner (Williams and Hallsworth 2009; Alves et al. 2015). Whereas a background level of chaotropicity-mediated benzene stress was evident in cultures from benzene + glycerol media, the overall level of these chaotrope-stress protein descriptors was 200–300 % lower than those observed in the benzene-stressed culture (see Fig. 2b).
The data obtained are consistent with the hypotheses that hydrocarbons induce cellular water stress, that the stress mechanism is mediated via chaotropicity, and that glycerol protects the cell’s macromolecular machinery via genome-wide responses/adaptations (see also Bhaganna et al. 2010; McCammick et al. 2010). Comparative analyses of the membrane lipid composition during growth with benzene and benzene + glycerol are now needed to shed further light on mechanisms of benzene tolerance. Evidence suggests that compatible solutes such as glycerol can mitigate competitive interactions between microbes which involve chaotropic antimicrobials, and reduce chaotropicity-induced stresses that occur during lignocellulosic and other fermentations (Bhaganna et al. 2010; Cray et al. 2013a, 2015a, 2015b; Alves et al. 2015). For instance, lignin breakdown products include chaotropic substances such as syringaldehyde and vanillin as well as hydrophobic stressors such as (cinnamaldehyde), see Cray et al. (2015a). Compatible solutes can mitigate stresses induced by such substances in yeast and fungi as well as bacteria (Bhaganna et al. 2010; Alves et al. 2015; Cray et al. 2015a). Further work is needed to elucidate the temporal sequence of molecular responses to hydrocarbon-induced water stress, apply biophysical techniques to quantify changes in the non-covalent interactions of macromolecular systems, investigate glycerol-mediated protection against hydrophobes in fungal systems (Cray et al. 2015a; Rangel et al. 2015b), and determine whether the kosmotropic and/or protectant activities of biologically relevant substances can be utilized to minimize hydrocarbon-induced stresses in microbial systems used for biocatalysis, biotransformation, bioremediation, and other industrial and environmental applications.
Notes
In other words, there was an apparent cessation of biotic activity (NB stressor concentrations at which metabolic activity and cell division cease and at which death occurs can differ; see Santos et al. 2015).
References
Albano E (2006) Alcohol, oxidative stress and free radical damage. Proc Nutr Soc 67:278–290
Albertyn J, Hohmann S, Thevelein JM, Prior BA (1994) Gpd1, which encodes glycerol-3-phosphate dehydrogenase, is essential for growth under osmotic-stress in Saccharomyces-cerevisiae, and its expression is regulated by the high-osmolarity glycerol response pathway. Mol Cell Biol 14:4135–4144
Alexandre H, Ansanav-Galeote V, Dequin S, Blondin B (2001) Global gene expression during short-term ethanol stress in Saccharomyces cerevisiae. FEBS Lett 498:98–103
Alves FL, Stevenson A, Baxter E, Gillion JLM, Hejazi F, Hayes S, Morrison IEG, Prior BA, McGenity TJ, Rangel DEN, Magan N, Timmis KN, Hallsworth JE (2015) Concomitant osmotic and chaotropicity-induced stresses in Aspergillus wentii: compatible solutes determine the biotic window. Curr Genet 61:457–477
Ball P, Hallsworth JE (2015) Water structure and chaotropicity: their uses, abuses, and implications for biology. Phys Chem Chem Phys 17:8297–8305
Ballerstedt H, Volkers RJM, Mars AE, Hallsworth JE et al (2007) Genomotyping of Pseudomonas putida strains using P. putida KT2440-based high-density DNA microarrays: implications for transcriptomics studies. Appl Microbiol Biotechnol 75:1133–1142
Bell AN, Magill E, Hallsworth JE, Timson DJ (2013) Effects of alcohols and compatible solutes on the activity of β-galactosidase. Appl Biochem Biotechnol 169:786–794
Benndorf D, Thiersch M, Loffhagen N, Kunath C, Harms H (2006) Pseudomonas putida KT2440 responds specifically to chlorophenoxy herbicides and their initial metabolites. Proteomics 6:3319–3329
Bhaganna P, Volkers RJM, Bell ANW, Kluge K, Timson DJ, McGrath JW, Ruijssenaars HJ, Hallsworth JE (2010) Hydrophobic substances induce water stress in microbial cells. Microb Biotechnol 3:701–716
Bobadilla Fazzini RA, Preto MJ, Quintas AC, Bielecka A, dos Santos VA (2010) Consortia modulation of the stress response: proteomic analysis of single strain versus mixed culture. Environ Microbiol 12:2436–2449
Bobadilla Fazzini RA, Skindersoe ME, Bielecki P, Puchałka J, Givskov M, Martins dos Santos VA (2013) Protoanemonin: a natural quorum sensing inhibitor that selectively activates iron starvation response. Environ Microbiol 15:111–120
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brown AD (1990) Microbial water stress physiology. Principles and perspectives. Wiley, Chichester
Chin JP, Megaw J, Magill CL, Nowotarski K, Williams JP, Bhaganna P, Linton M, Patterson MF, Underwood GJC, Mswaka AY, Hallsworth JE (2010) Solutes determine the temperature windows for microbial survival and growth. Proc Natl Acad Sci USA 107:7835–7840
Chirife J, Favetto G, Fontán CF (1984) Microbial growth at reduced water activities: some—physicochemical properties of compatible solutes. J Appl Microbiol 56:259–268
Cray JA, Bell ANW, Bhaganna P, Mswaka AY, Timson DJ, Hallsworth JE (2013a) The biology of habitat dominance; can microbes behave as weeds? Microb Biotechnol 6:453–492
Cray JA, Russell JT, Timson DJ, Singhal RS, Hallsworth JE (2013b) A universal measure of chaotropicity and kosmotropicity. Environ Microbiol 15:287–296
Cray JA, Bhaganna P, Singhal RS, Patil SV, Saha D, Chakraborty R, Iwaguchi S, Timson DJ, Hallsworth JE (2014) Chaotropic and hydrophobic stress mechanisms of antifungal substances. In: Dehne HW, Deising HB, Fraaije B, Gisi U, Hermann D, Mehl A, Oerke EC, Russell PE, Stammler G, Kuck KH, Lyr H (eds) Modern fungicides and antifungal compounds, vol VII. Deutsche Phytomedizinische Gesellschaft, Braunschweig. ISBN: 978-3-941261-13-6
Cray JA, Stevenson A, Ball P, Bankar SB, Eleutherio ECA, Ezeji TC, Singhal RS, Thevelein JM, Timson DJ, Hallsworth JE (2015a) Chaotropicity: a key factor in product tolerance of biofuel-producing microorganisms. Curr Opin Biotechnol 33:228–259
Cray JA, Houghton JD, Cooke LR, Hallsworth JE (2015b) A simple inhibition coefficient for quantifying potency of biocontrol agents against plant-pathogenic fungi. Biol Control 81:93–100
Domínguez-Cuevas P, González-Pastor J-E, Marqués S, Ramos JL, de Lorenzo V (2006) Transcriptional tradeoff between metabolic and stress-response programs in Pseudomonas putida KT2440 cells exposed to toluene. J Biol Chem 281:11981–11991
Dos Santos VAP, Heim S, Moore ERB, Strätz M, Timmis KN (2004) Insights into the genomic basis of niche specificity of Pseudomonas putida KT2440. Environ Microbiol 6:1264–1286
Eleutherio ECA, Dearaujo PS, Panek AD (1993) Role of the trehalose carrier in dehydration resistance of Saccharomyces cerevisiae. Biochim Biophys Acta 1156:263–266
Escapa IF, García JL, Bühler B, Blank LM, Prieto MA (2012) The polyhydroxyalkanoate metabolism controls carbon and energy spillage in Pseudomonas putida. Environ Microbiol 14:1049–1063
Görg A (1991) Two-dimensional electrophoresis. Nature 349:545–546
Hallsworth JE (1998) Ethanol-induced water stress in yeast. J Ferment Bioeng 85:125–137
Hallsworth JE, Magan N (1994) Improved biological control by changing polyols/trehalose in conidia of entomopathogens. Brighton Crop Protection Conference 1994—Pests and Diseases, vol 3. BCPC Publications, Farnham, pp 1091–1096
Hallsworth JE, Magan N (1995) Manipulation of intracellular glycerol and erythritol enhances germination of conidia at low water availability. Microbiol SGM 29:7–13
Hallsworth JE, Nomura Y (1999) A simple method to determine the water activity of ethanol-containing samples. Biotechnol Bioeng 62:242–245
Hallsworth JE, Nomura Y, Iwahara M (1998) Ethanol-induced water stress and fungal growth. J Ferment Bioeng 86:451–456
Hallsworth JE, Prior BA, Nomura Y, Iwahara M, Timmis KN (2003a) Compatible solutes protect against chaotrope (ethanol)-induced, nonosmotic water stress. Appl Environ Microb 69:7032–7034
Hallsworth JE, Heim S, Timmis KN (2003b) Chaotropic solutes cause water stress in Pseudomonas putida. Environ Microbiol 5:1270–1280
Hallsworth JE, Yakimov MM, Golyshin PN, Gillion JLM, D’Auria G, Alves FL, La Cono V, Genovese M, McKew BA, Harris G, Guiliano L, Timmis KN, McGenity TJ (2007) Limits of life in MgCl2-containing environments: chaotropicity defines the window. Environ Microbiol 9:803–813
Hamada T, Maeda Y, Matsuda H, Sameshima Y, Sameshima Y, Honda K, Omasa T, Kato J, Ohtake H (2009) Effect of cell-surface hydrophobicity on bacterial conversion of water-immiscible chemicals in two-liquid-phase culture systems. J Biosci Bioeng 108:116–120
Harrison JP, Gheeraert N, Tsigelnitskiy D, Cockell CS (2013) The limits for life under multiple extremes. Trends Microbiol 21:204–212
Harrison JP, Hallsworth JE, Cockell CS (2015) Reduction of the temperature sensitivity of Halomonas hydrothermalis by iron starvation combined with microaerobic conditions. Appl Environ Microbiol 81:2156–2162
Hartmans S, Smits JP, van der Werf MJ, Volkering F, de Bont JAM (1989) Metabolism of styrene oxide and 2-phenylethanol in the styrene-degrading Xanthobacter strain 124X. Appl Environ Microbiol 55:2850–2855
Hassan MN, Afghan S, Hafeez FY (2011) Biological control of red rot in sugarcane by native pyoluteorin-producing Pseudomonas putida strain NH-50 under field conditions and its potential modes of action. Pest Manag Sci 67:1147–1154
Hathout Y, Riordan K, Gehrmann M, Fenselau C (2002) Differential protein expression in the cytosol fraction of an MCF-7 breast cancer cell line selected for resistance towards Melphalan. J Proteom Res 1:435–442
Heim S, Ferrer M, Heuer H, Regenhardt D, Nimtz M, Timmis KN (2003) Proteome reference map of Pseudomonas putida strain KT2440 for genome expression profiling: distinct responses of KT2440 and Pseudomonas aeruginosa strain PAO1 to iron deprivation and a new form of superoxide dismutase. Environ Microbiol 5:1257–1269
Hocking AD (1993) Responses of xerophilic fungi to changes in water activity. In: Jennings DH (ed) Stress Tolerance of Fungi. Marcel Decker, New York, pp 233–256
Hubalek Z (2003) Protectants used in the cryopreservation of microorganisms. Cryobiol 46:205–229
Joo WH, Shin YS, Lee Y, Park SM, Jeong KY, Seo JY, Park JU (2000) Intracellular changes of trehalose content in toluene tolerant Pseudomonas sp. BCNU 171 after exposure to toluene. Biotechnol Lett 22:1021–1024
Kar JR, Hallsworth JE, Singhal RS (2015) Glycine betaine-mediated protection of peas (Pisum sativum L.) during blanching and frozen storage. J Sci Food Agric. doi:10.1080/10942912.2015.1099043
Kar JR, Hallsworth JE, Singhal RS (2015) Fermentative production of glycine betaine and trehalose from acid whey using Actinopolyspora halophila (MTCC 263). Environ Technol Innov 3:68–76
Kashangura C, Hallsworth JE, Mswaka, AY (2006) Phenotypic diversity amongst strains of Pleurotus sajor-caju: implications for cultivation in arid environments. Mycol Res 110: 312–317
Kavazis AN, Talbert EE, Smuder AJ, Hudson MB, Nelson WB, Powers SK (2009) Mechanical ventilation induces diaphragmatic mitochondrial dysfunction and increased oxidant production. Free Rad Biol Med 46:842–850
Keane A, Lau PCK, Ghoshal S (2008) Use of a whole-cell biosensor to assess the bioavailability enhancement of aromatic hydrocarbon compounds by nonionic surfactants. Biotechnol Bioeng 99:86–98
Kelly BC, Ikonomou MG, Blair JD, Morin AE, Gobas FAPC (2007) Food web-specific biomagnification of persistent organic pollutants. Science 317:236–239
Kets EPW, de Bont JAM, Heipieper HJ (1996a) Physiological response of Pseudomonas putida S12 subjected to reduced water activity. FEMS Microbiol Lett 139:133–137
Kets EPW, Galinski EA, de Wit M, de Bont JAM, Heipieper HJ (1996b) Mannitol, a novel bacterial compatible solute in Pseudomonas putida S12. J Bacteriol 178:6665–6670
Kim K, Lee S, Lee K, Lim D (1998) Isolation and characterization of toluene-sensitive mutants from the toluene-resistant bacterium Pseudomonas putida GM73. J Bacteriol 180(14):3692–3696
Koopman F, Wierckx N, de Winde JH, Ruijssenaars HJ (2010) Identification and characterization of the furfural and 5-(hydroxymethyl)furfural-degradation pathways of Cupriavidus basilensis HMF14. Proc Natl Acad Sci USA 107:4919–4924
Kurbatov L, Albrecht D, Herrmann H, Petruschka L (2006) Analysis of the proteome of Pseudomonas putida KT2440 grown on different sources of carbon and energy. Environ Microbiol 8:466–478
Lamm SH, Grunwald HW (2006) Benzene exposure and hematotoxicity. Science 312:998
Len ACL, Harty DWS, Jacques NA (2004) Stress-responsive proteins are upregulated in Streptococcus mutans during acid tolerance. Microbiology 150:1339–1351
Lievens B, Hallsworth JE, Pozo MI, Belgacem ZB, Stevenson A, Willems KA, Jacquemyn H (2015) Microbiology of sugar-rich environments: diversity, ecology, and system constraints. Environ Microbiol 17:278–298
Littlejohns JV, McAuley KB, Daugulis AJ (2010) Model for a solid-liquid stirred tank two-phase partitioning bioscrubber for the treatment of BTEX. J Hazard Mat 175:872–882
McCammick EM, Gomase VS, Timson DJ, McGenity TJ, Hallsworth JE (2010) Water–hydrophobic compound interactions with the microbial cell. In: Timmis KN (ed) Handbook of hydrocarbon and lipid microbiology—hydrocarbons, oils and lipids: diversity, properties and formation, vol 2. Springer, New York, pp 1451–1466
Nelson KE, Paulsen IT, Weinel C, Dodson RJ, Hilbert H et al (2002) Complete genome sequence and comparative analysis of the metabolically versatile Pseudomonas putida KT2440. Environ Microbiol 4:799–808
Oren A, Hallsworth JE (2014) Microbial weeds in saline habitats: the enigma of the weed-like Haloferax mediterranei. FEMS Microbiol Lett 359:134–142
Park SH, Oh KH, Kim CK (2001) Adaptive and cross-protective responses of Pseudomonas sp. DJ-12 to several aromatics and other stress shocks. Curr Microbiol 43:176–181
Radniecki TS, Dolan ME, Semprini L (2008) Physiological and transcriptional responses of Nitrosomonas europaea to toluene and benzene inhibition. Environ Sci Technol 42:4093–4098
Rangel DEN, Braga GUL, Fernandes ÉKK, Keyser CA, Hallsworth JE, Roberts DW (2015a) Stress tolerance and virulence of insect-pathogenic fungi are determined by environmental conditions during conidial formation. Curr Genet 61:383–404
Rangel DEN, Alder-Rangel A, Dadachova E, Finlay RD, Kupiec M, Dijksterhuis J, Braga GUL, Corrochano LM, Hallsworth JE (2015b) Fungal stress biology: a preface to the Fungal Stress Responses special edition. Curr Genet 61:231–238
Rummel JD, Beaty DW, Jones MA, Bakermans C, Barlow NG, Boston PJ, Chevrier VF, Clark BC, de Vera JPP, Gough RV, Hallsworth JE, Head JW, Hipkin VJ, Kieft TL, McEwen AS, Mellon MT, Mikucki JA, Nicholson WL, Omelon CR, Peterson R, Roden EE, Sherwood Lollar B, Tanaka KL, Viola D, Wray JJ (2014) A new analysis of Mars “Special Regions”: findings of the second MEPAG Special Regions Science Analysis Group (SR-SAG2). Astrobiology 14:887–968
Russo A, Palumbo M, Scifo C, Cardile V, Barcellona ML, Renis M (2001) Ethanol-induced oxidative stress in rat astrocytes: role of HSP70. Cell Biol Toxicol 17:153–168
Santos PM, Benndorf D, Sá-Correia I (2004) Insights into Pseudomonas putida KT2440 response to phenol-induced stress by quantitative proteomics. Proteomics 4:2640–2652
Santos R, Stevenson A, de Carvalho CCCR, Grant IR, Hallsworth JE (2015) Extraordinary solute stress tolerance contributes to the environmental tenacity of mycobacteria. Environ Microbiol Rep 7:746–764
Sauer T, Galinski EA (1998) Bacterial milking: a novel bioprocess for production of compatible solutes. Biotechnol Bioeng 57:306–313
Segura A, Godoy P, Van Dillewijn P, Hurtado A, Arroyo N, Santacruz S, Ramos JL (2005) Proteomic analysis reveals the participation of energy- and stress-related proteins in the response of Pseudomonas putida DOT-T1E to toluene. J Bacteriol 187:5937–5945
Sikkema J, de Bont JAM, Poolman B (1995) Mechanisms of membrane toxicity of hydrocarbons. Microbiol Rev 59:201–222
Smirnoff N, Cumbes QJ (1989) Hydroxyl radical scavenging activity of compatible solutes. Phytochemistry 28:1057–1060
Stevenson A, Hallsworth JE (2014) Water and temperature relations of soil Actinobacteria. Environ Microbiol Rep 6:744–755
Stevenson A, Cray JA, Williams JP, Santos R, Sahay R, Neuenkirchen N, McClure CD, Grant IR, Houghton JDR, Quinn JP, Timson DJ, Patil SV, Singhal RS, Anton J, Dijksterhuis J, Hocking AD, Lievens B, Rangel DEN, Voytek MA, Gunde-Cimerman N, Oren A, Timmis KN, McGenity TJ, Hallsworth JE (2015a) Is there a common water-activity limit for the three domains of life? ISME J 9:1333–1351
Stevenson A, Burkhardt J, Cockell CS, Cray JA, Dijksterhuis J, Fox-Powell M, Kee TP, Kminek G, McGenity TJ, Timmis KN, Timson DJ, Voytek MA, Westall F, Yakimov MM, Hallsworth JE (2015b) Multiplication of microbes below 0.690 water activity: implications for terrestrial and extraterrestrial life. Environ Microbiol 17:257–277
Suryawanshi RK, Patil CD, Borase HP, Narkhede CP, Stevenson A, Hallsworth JE, Patil SV (2015) Towards an understanding of bacterial metabolites prodigiosin and violacein and their potential for use in commercial sunscreens Internat. J Cosmet Sci 37:98–107
Tam LT, Eymann C, Albrecht D, Sietmann R, Schauer F, Hecker M, Antelmann H (2006) Differential gene expression in response to phenol and catechol reveals different metabolic activities for the degradation of aromatic compounds in Bacillus subtilis. Environ Microbiol 8:1408–1427
Timmis KN (ed) (2010) Handbook of hydrocarbon and lipid microbiology—hydrocarbons, oils and lipids: diversity, properties and formation, vol 2. Springer, New York
Trevisol ET, Panek AD, Mannarino SC, Eleutherio EC (2011) The effect of trehalose on the fermentation performance of aged cells of Saccharomyces cerevisiae. Appl Microbiol Biotechnol 90:697–704
Velázquez F, Parro V, de Lorenzo V (2005) Inferring the genetic network of m-xylene metabolism through expression profiling of the xyl genes of Pseudomonas putida mt-2. Mol Microbiol 57:1557–1569
Volkers RJM, de Jong AL, Hulst AG, van Baar BLM, de Bont JS, Wery J (2006) Chemostat-based proteomic analysis of toluene-affected Pseudomonas putida S12. Environ Microbiol 8:1674–1679
Wierckx NJP, Ballerstedt H, de Bont JAM, de Winde JH, Ruijssernaars HJ, Wery J (2008) Transcriptome analysis of a phenol-producing Pseudomonas putida S12 construct: genetic and physiological basis for improved production. J Bacteriol 190:2822–2830
Williams JP, Hallsworth JE (2009) Limits of life in hostile environments; no limits to biosphere function? Environ Microbiol 11:3292–3308
Womack AM, Bohannan BJM, Green JL (2010) Biodiversity and biogeography of the atmosphere. Phil Trans R Soc B 365:3645–3653
Wyatt TT, Golovina EA, van Leeuwen MR, Hallsworth JE, Wösten HAB, Dijksterhuis J (2015a) A decrease in bulk water and mannitol and accumulation of trehalose and trehalose-based oligosaccharides define a two-stage maturation process towards extreme stress resistance in ascospores of Neosartorya fischeri (Aspergillus fischeri). Environ Microbiol 17:283–294
Wyatt TT, van Leeuwen MR, Gerwig GJ, Golovina EA, Hoekstra FA, Kuenstner EJ, Palumbo EA, Snyder NL, Visagie C, Verkennis A, Hallsworth JE, Kamerling JP, Wösten HAB, Dijksterhuis J (2015b) Functionality and prevalence of trehalose-based oligosaccharides as novel compatible solutes in ascospores of Neosartorya fischeri (Aspergillus fischeri) and other fungi. Environ Microbiol 17:395–411
Yakimov MM, Lo Cono V, La Spada G, Bortoluzzi G, Messina E, Smedile F, Werner J, Teeling H, Borghini M, Ferrer M, Cray JA, Hallsworth JE, Golyshin PN, Giuliano L (2015) Microbial community of seawater-brine interface of the deep-sea brine Lake Kryos as revealed by recovery of mRNA are active below the chaotropicity limit of life. Environ Microbiol 17:364–382
Yun SH, Kim YH, Joo EJ, Choi JS, Sohn JH, Kim SI (2006) Proteome analysis of cellular response of Pseudomonas putida KT2440 to tetracycline stress. Curr Microbiol 53:95–101
Yun SH, Park GW, Kim JY, Kwon SO, Choi CW, Leem SH, Kwon KH, Yoo JS, Lee C, Kim S, Kim SI (2011) Proteomic characterization of the Pseudomonas putida KT2440 global response to a monocyclic aromatic compound by iTRAQ analysis and 1DE-MudPIT. J Proteomics 74:620–628
Zager RA (1996) Mitochondrial free radical production induces lipid peroxidation during myohemoglobinuria. Kidney Internat 49:741–751
Acknowledgments
This project was carried out within the research program of the Kluyver Centre for Genomics of Industrial Fermentation which is part of the Netherlands Genomics Initiative/Netherlands Organization for Scientific Research, the EU-funded MIFRIEND and LINDANE projects, a Beaufort Marine Research Award for Marine Biodiscovery (Marine Institute, Ireland), and a Biotechnology and Biological Sciences Research Council-funded project (UK; Grant No. BBF0034711) which is part of the Pseudomonas putida Systems Biology of Microorganisms project (PSYSMO). We are grateful for useful discussions with Giuseppe Albano (Edinburgh University, UK), Philip G. Hamill, Luke Morton, Allen Y. Mswaka, Andrew Stevenson, and Dave J. Timson, (Queen’s University Belfast), Victor de Lorenzo (Centro Nacional de Biotecnología, CSIC, Spain), Terry J. McGenity and Jen Waring (University of Essex), Mary Palfreyman (Outwood Grange College, UK), and Kenneth N. Timmis (Helmholtz Centre for Infection Research, Germany).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no financial/commercial conflicts of interest.
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhaganna, P., Bielecka, A., Molinari, G. et al. Protective role of glycerol against benzene stress: insights from the Pseudomonas putida proteome. Curr Genet 62, 419–429 (2016). https://doi.org/10.1007/s00294-015-0539-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-015-0539-1