Abstract
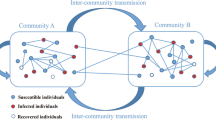
This paper investigates the effects of the community structure of a network on the spread of an epidemic. To this end, we first establish a susceptible–infected–recovered (SIR) model in a two-community network with an arbitrary joint degree distribution. The network is formulated as a probability generating function. We also obtain the sufficient conditions for disease outbreak and extinction, which involve the first-order and second-order moments of the degree distribution. As an example, we then study the effect of community structure on epidemic spread in a complex network with a Poisson joint degree distribution. The numerical solutions of the SIR model well agree with stochastic simulations based on the Monte Carlo method, confirming that the model is reliable and accurate. Finally, by strengthening the community structure in the simulation, i.e. fixing the total degree distribution and reducing the number ratio of the external edges, we can increase or decrease the final cumulative epidemic incidence depending on the transmissibility of the virus between humans and the community structure at that point. Why community structure can affect disease dynamics in a complicated way is also discussed. In any case, for large-scale epidemics, strengthening the community structure to reduce the size of disease is undoubtedly an effective way.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Control of epidemic spreading has always attracted interest in the biological mathematics field. However, many models describing the spread of an infection assume a randomly mixed population, in which contact between any two individuals is equally likely (Ball et al. 2013; Chris 2005). This assumption is strong and in many cases unrealistic. Epidemics on contact networks provide a more realistic alternative. In general, a contact network represents the possible connections through which an infection can transmit. Classical dynamic models of epidemics consider the impact of disease factors on the spread of an infectious disease. Therefore, much research on infectious disease transmission in complex populations has focused on understanding the implications of network structures on the epidemic processes. Recently, Kiss et al. (2017) have shown a large pool of epidemic models on networks, ranging from exact and stochastic to approximate differential equation models.
Recent works have shown that community structure is a universal feature of contact networks. The community structure describes the division of the network nodes into groups with dense connections among the group members, but sparser connections among the groups. The community may be classmates, friends, co-workers and club members. Research on the community structures of networks has yielded rich results. Such research has focused on the features and excavation of communities, or epidemic propagation in a network. Present results on the former are focused on community detection (Newman and Girvan 2004; Traud et al. 2009; Wayne 1977) and community growth models (Jin et al. 2001). Orman et al. (2013) considered the relationship between community structure and clustering coefficient. They discovered that even a network with near-zero transitivity can form a clearly defined community structure. Epidemic propagation through networks has focused on the effect of community structure on epidemic propagation. Various simulation studies (Huang and Li 2007; Liu and Hu 2005; Sun and Gao 2007) have revealed that networks without community structure are more robust than those with community structure. For details, see Huang and Li (2007), Wu and Liu (2008), Yan et al. (2007). Rowthorn et al. (2009) and Salathé and Jones (2010) discuss the control of diseases in metapopulations and in networks with community structure, respectively. Furthermore, the latter shows that community structure will lower the final size and peak prevalence under certain parameter conditions.
Disease spread through complex networks can be modeled by various approaches, such as mean field equations (Gross et al. 2006; Luo et al. 2014; Wang et al. 2010), the moment closure model (Eames 2008; House and Keeling 2010), a branching process approximation (Ball 1983; Ball and Neal 2008; Neal 2007), the effective degree model (Ma et al. 2013), bond percolation theory (Gleeson 2009; Gleeson et al. 2010; Miller 2009a; Newman 2003b; Wang et al. 2012) and a generating function-based model (Miller 2009b, 2011; Volz 2008). House and Keeling (2011) introduce the generating function into the approximation formula, and get a low-dimensional model with a small number of dynamical variables. Some of these methods have been extended to disease spread through complex networks with community structure.
The mean field equations and moment closure model were extended to a two-community network by Peng et al. (2013), Tunc and Shaw (2014) and Zhou and Liu (2009). Peng et al. (2013) and Zhou and Liu (2009) studied a susceptible–infected–susceptible (SIS) epidemiological model in a two-community network with a static and mobile agent, respectively. Both studies concluded that epidemic spreading in a community can be sustained through contact with another community. Tunc and Shaw (2014) studied the effects of community structure on epidemic spreading in an adaptive network. They showed that an epidemic can change the community structure. The household structure, as a special kind of community structure, has also been studied. Using mean field equations, Liu et al. (2004) concluded that some simple geometrical quantities of networks crucially affect the infection property of infectious diseases. Ball et al. (1997) reported that household structure exerts an amplification effect that permits a population-scale outbreak at very low levels of global transmission. Ball et al. (2009, 2010) derived the disease threshold parameters by branching-process approximations. However, these models do not describe the disease dynamics. In an effective degree SIR model, Ma et al. (2013) derived expressions for the disease threshold parameters. They showed that households can accelerate or decelerate the disease dynamics depending on the variance of the inter-household degree distribution.
All of the above epidemic models in a network with community structure assume homogeneous mixing. In 2013, Koch et al. (2013) extended the Miller network SIR model (Miller 2011). They modeled the spread of disease in complex two-community networks in the special case of independence between the inter- and intra-community degrees, obtaining an 8 dimensional model. They proved that random edge removal, from either group or between the groups, always decreases the basic reproduction number. In the same year, Miller and Volz (2013) extended the Miller network SIR model (Miller 2011) to model disease spread in complex networks with n communities and an arbitrary joint degree distribution (a 2-community network yielded a 4- dimensional model). They proved that for the same distribution of within and between-group partnerships in two populations, varying the correlations of the within and between-group partnerships will alter the course of an epidemic.
The present paper extends the Volz network SIR model (Volz 2008) to disease spreading in two-community complex networks with an arbitrary joint degree distribution. We obtain a 12 dimensional model. As the Volz (2008) and Miller (2011) models are consistent, our model is consistent with the model of Miller and Volz (2013) with \(n=2\). Although the model of Miller and Volz (2013) has fewer dimensions than our model, the basic reproduction number of their model requires finding the root of a quartic equation, which greatly complicates the disease threshold analysis. Instead, our model obtains the disease threshold conditions analytically, which involve the high-order moments of the degree distribution.
Furthermore, although the model of Miller and Volz (2013) assumes the same distribution of within and between-group partnerships in both populations, which implies the same community structure in the two populations, the total degree distribution in this process is also changed. Consequently, their work ignores the impact of community structure on the epidemic dynamics. In Koch et al. (2013), the random edge removal alters not only the community structure, but also the total degree distribution. Therefore, they cannot discern whether their dynamical behaviors are altered by changing the total degree distribution or by changing the community structure. In this paper, we investigate the effect of community structure on the propagation of a SIR epidemic in a network with a specific degree distribution. The total degree distribution is fixed, and the community structure is altered by adjusting the number ratio of the internal and external edges. Two main conclusions emerged from the study. First, by strengthening the community structure (i.e, fixing the total degree distribution while reducing the number ratio of the external edges), we can increase or decrease the final cumulative epidemic incidence depending on the transmissibility of the virus between humans and the community structure of the network at that point. Second, disease transmission is most obviously affected by the community structure when the human-to-human transmissibility of the virus is near the threshold.
The remainder of this paper is organized as follows. Section 2 introduces the definitions and measurements of community structures and the generation of complex networks with community structure. Section 3 develops our low-dimensional model of SIR-epidemic percolation in a two-community network, and obtains the epidemic threshold beyond which the disease outbreaks in both communities. The main conclusions of the study are demonstrated in an example, and the accuracy of the model and theorems are confirmed in a simulation study. In Sect. 4, the effects of community structure on disease spread are explored in additional simulations. Section 5 summarizes our present findings and attempts to explain why community structure can affect disease dynamics in a complicated way.
2 Complex networks with community structure
2.1 Definitions and measurement of the community structure
Consider a closed population of N individuals. The population is described as a simple network \(G = (V,E)\), where \(V=\{v_1,v_2,\cdots ,v_N\}\) is the set of nodes representing the individuals and E is the set of edges representing the connections between the individuals. The network is simple, meaning that at most one edge connects two individuals. We assume also that contacts are symmetric, that is, if an edge \((v_1, v_2)\in E\) connects \(v_1\) to \(v_2\), then an edge also connects \(v_2\) to \(v_1\). Although the network is undirected (i.e. any two neighboring vertices can infect each other), we wish to keep track of who infects who. Therefore, for each edge \((v_1, v_2)\in E\), we define two arcs as the ordered pairs \((v_1, v_2)\) and \((v_2, v_1)\). The first and second elements in the ordered pair \((v_1, v_2)\) are frequently called the ego and the alter, respectively (Volz 2008).
The N network nodes are divided into two groups, one with \(N_1\) nodes, the other with \(N_2\) nodes, where \(N_1+N_2=N\). We call the first and second groups the first and second community, respectively. Let \(P_l(k,j)\) (\(l=1,2\)) be the probability that a node in the lth community has k intra-community neighbors and j inter-community neighbors. Then the probability generating functions (PGFs) of the joint degree distribution of the first and second community are given by
respectively, where \(\sum \nolimits _{k,j}P_l(k,j)=1\), \(l=1,2\). For convenience, let
As the two communities share common external edges, the balance condition is given by
We denote by square brackets [ \(]^{in}_l\) and [ \(]^{out}_l\) the expected numbers of any particular internal and external contacts with the lth community, respectively, and use the following notations:
and
For convenience, we apply the following notations
for \(l=1,2\). Thus, the expected numbers of internal and external degrees of the lth community are
respectively. The internal, external and mixed second moments of the lth community are respectively given by
Furthermore, the average internal and external redundancies along an internal edge in the lth community are
respectively. Similarly, the average internal and external redundancies along an external edge in the lth community are
respectively. Furthermore, the PGF of the total degree distribution of the whole network is
We now introduce the quality function Q proposed by Newman (2003a), which measures the strength of the community structure in a complex network:
where \(e_{ij}\) is the fraction of links in the network that connect nodes in communities i and j, and \(a_i=\sum \nolimits _{j}e_{ij}\) represents the fraction of edges connected to vertices in community i. Q approximates zero in a random network, and one in a network with a strong community structure. In real-world networks, Q typically falls within the range 0.3–0.7 (Newman 2003a). Higher values are rare. As an example, Fig. 1 shows a two-community network with \(Q=0.314\).
From Eqs. (3), (4) and (8), we have
Substituting equation (9) into (8), we obtain
Thus the strength Q of the community structure in a complex network depends on the parameters \(G_l,G^l\), \(l=1,2\).
A random network with two communities. Community A consists of 100 nodes (black circles) with an average intra-community degree \([k]^{in}_1=4\). Community B consists of 100 nodes (red circles) with \([k]^{in}_2=5\). The average inter-community degrees of Communities A and B are \( [j]^{out}_1=1\) and \( [j]^{out}_2=1\), respectively (color figure online)
2.2 Generation of complex networks with community structure
Stochastic simulations were performed on a two-community contacting network with a specified joint degree distribution. The network was generated by the Configuration Model (Molloy and Reed 1995). This process, a variation of the CM model proposed by Volz (2008), is briefly described below:
Each node is assigned to a single community. The nodes are apportioned according to the sizes of the two communities. To each node \(v_l\) in community l (\(l=1,2\)), assign a joint degree \((\delta (v_l), \zeta (v_l))\) from the joint degree distribution \(P_l(k,j)\). For all nodes in each community \(l=1,2\), generate new sets \(X_l\) of “half-edges” with \(\delta (v_l)\) copies of node \(v_l\) and sets \(X^l\) of “half-edges” with \(\zeta (v_l)\) copies of node \(v_l\). While \(X_l\) is not empty, randomly and uniformly draw two elements \(v(l), w(l)\in X_l\) and create an edge (v(l), w(l)) (an internal connection in community l). While both \(X^1\) and \(X^2\) are non-empty, randomly and uniformly draw two elements v(1) and v(2) from \(X^1\) and \(X^2\) respectively and create an edge (v(1), v(2)) (an external connection between the two communities).
Note that this procedure does not allow multiple edges to the same nodes, and loops from a node to itself.
3 SIR models in random networks
This section introduces a low-dimensional system that models the percolation of an SIR-epidemic in a two-community network with arbitrary joint degrees. It also obtains the disease thresholds.
3.1 Our model
When a disease spreads through a network, the nodes can be in any of three exclusive states: susceptible (S), infectious (I), and recovered (R). The dynamics are as follows. An infectious node will independently infect each of its neighbors at a constant rate \(\gamma \). Infectious nodes recover at a constant rate \(\mu \), whereupon they no longer infect any neighbor. These processes will be formulated in the next section. Following the notations in the literature (Volz 2008), we let s, i and r represent the fractions of susceptible, infectious and recovered nodes, respectively.
More generally, we represent the state of a node as A or B respectively, where A and B may be S, I or R. For ease of reference, we summarize this notation in Table 1.
The model is formulated as follows:
where \(n\ne l\) and \(l,n=1,2\). This model extends the network SIR model of Volz (2008) to disease spreading in a complex two-community network with an arbitrary joint degree distribution. The model is derived in detail in “Appendix A”.
The fraction of nodes that remain susceptible at time t in community l (\(l=1, 2\)) is given by
and the fraction of nodes that remain susceptible in the whole network at time t is
To consider the number of infected individuals, we need the dynamics of the infectious nodes. The infectious class increases at rate \(-\dot{s}\) and decreases at rate \(\mu i\). Therefore, we have
3.2 Initial conditions
We randomly and uniformly select small but strictly positive fractions \(\epsilon _1\) and \(\epsilon _2\) of the nodes in Community 1 and Community 2, respectively, as initially infected nodes. Then we obtain the following condition.
where \(l, n=1,2\). To summarize,
where \(n\ne l\) and \(l,n=1,2\).
3.3 Epidemic thresholds
As noted in the literature (Volz 2008), the number of new infectious in a small time interval is proportional to \(p_S^I\), which defines the probability that a susceptible node is connected to an infectious node. If \(\dot{p}_S^I(t=0)<0\), an epidemic will necessarily die out without reaching a fraction of the population.
According to the total probability formula, we obtain
So
Assuming independent transmission from infectious nodes to their common susceptible neighbors, we have
According to Eq. (11) and the initial conditions (16), we obtain
Noting that \(\epsilon _1,\epsilon _2\ll 1\) we can ignore the higher-order terms of \(\epsilon _1\) and \(\epsilon _2\) to obtain
Similarly, we can obtain
Applying Eqs. (21) and (22) to (19), we have
where \(l\ne n\) and \(l,n=1,2\). Ignoring the higher-order terms of \(\epsilon _1\) and \(\epsilon _2\), we have
where \(l\ne n\) and \(l,n=1,2\).
Now, applying Eqs. (2), (11), (12) and (24)–(18) and noticing that \(\epsilon _1,\varepsilon _2\ll 1\), we have
After rearranging this expression, we get the critical ratio \(\gamma /\mu \) in terms of the PGF. They are two positive numbers:
If \((\gamma /\mu )^*_1>0\) and \((\gamma /\mu )^*_2>0\), we obtain the following theorem. This theorem is the main result of this section.
Theorem 1
For the SIR model described in (11), the following are true.
-
(I)
If \( \frac{\gamma }{\mu }> \max \{(\frac{\gamma }{\mu })^*_1,(\frac{\gamma }{\mu })^*_2\}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \min \{(\frac{\gamma }{\mu })^*_1,(\frac{\gamma }{\mu })^*_2\}\), the epidemic dies out in both communities;
-
(III)
If \(\min \{(\frac{\gamma }{\mu })^*_1,(\frac{\gamma }{\mu })^*_2\}<\frac{\gamma }{\mu } \le \max \{(\frac{\gamma }{\mu })^*_1,(\frac{\gamma }{\mu })^*_2\}\), the epidemic may occur in both communities, die out in both communities or occur in one community and die out in the other community.
From (6) and (7), we know that \(\frac{G_{ll}}{G_l}\) and \(\frac{G^{nn}}{G^n} (n\ne l)\) are the average numbers of transmissible neighbors in Community l of a node in communities l and n, respectively, which was newly infected by a node in Community l. Meanwhile, \(\frac{G_l^l}{G^l} \) and \(\frac{G_n^n}{G_n}\) \((n\ne l)\) are the average numbers of transmissible neighbors in Community l of a node in communities n and l, respectively, which was newly infected by a node in Community n. Therefore, the average number of transmissible neighbors in Community l is \(\frac{N_l}{N_l+N_n}(\frac{G_{ll}}{G_l}+\frac{G^{nn}}{G^n})+\frac{N_n}{N_l+N_n}(\frac{G_n^n}{G_n}+\frac{G_l^l}{G^l})\). The form of the thresholds \((\gamma /\mu )^*_1\) and \((\gamma /\mu )^*_2\) is then consistent with the critical transmissibility of the SIR epidemic model in a network with no community structure obtained by Newman (2002), i.e.
When \(\gamma /\mu >(\gamma /\mu )^*_l\), the disease expands in Community l, and when \(\gamma /\mu <(\gamma /\mu )^*_l\), the disease shrinks in Community l. Hence, \(\gamma /\mu >\max \{(\gamma /\mu )^*_1, \gamma /\mu )^*_2\}\) means that the disease expands in both communities, and the epidemic spreads through the whole net. Conversely, \(\gamma /\mu <\min \{(\gamma /\mu )^*_1, \gamma /\mu )^*_2\}\) means that the disease shrinks in both communities, and the epidemic dies out throughout the whole net. However, when \(\min \{(\gamma /\mu )^*_1, \gamma /\mu )^*_2\}<\gamma /\mu <\max \{(\gamma /\mu )^*_1, \gamma /\mu )^*_2\}\), the disease expands in one community and shrinks in the other. Diseases can shift between the two communities, sometimes expanding and sometimes shrinking, so the final trend of the disease is difficult to predict.
The disease thresholds involve both the first-order and second-order moments of the network. Therefore, changing the strength Q of the community structure in a complex network changes the first order moments \(G_l\) and \(G^l\) and also the second-order moments \(G_{ll}\), \(G^{ll}\) and \(G_l^l\), \((l=1,2)\). When G is nonspecific, the impact of Q on the spread of disease cannot be easily known. In the following section, we exemplify the impact of community structure on disease spread using a specific distribution function.
3.4 An example
In this subsection, we demonstrate Theorem 1 by an example. The joint degree distribution of the community l is assumed as the following Poisson distribution, i.e.
where \(\lambda _{ln}\) (\(l,n=1,2\)) are positive numbers, and the expected number of external degrees \(\lambda _{12}\) and \(\lambda _{21}\) meet the balance condition
The corresponding PGF is then given by
From Eqs. (3) and (30), we have
and
3.4.1 Measurement of the community structure
According to Eqs. (31), (32) and the definition Eq. (10), the strength of the community structure in the above complex network is
We next study the relationship between the external degree and community structure. To this end, we first denote the average total degrees of Community 1 and 2 as \(\lambda _1\) and \(\lambda _2\), respectively. These quantities are defined as follows
Given the scales (\(N_1\) and \(N_2\)) and the average total degrees (\(\lambda _1\) and \(\lambda _2\)) of the communities, Q and \(\lambda _{12}\) are related as follows:
According to this formula, increasing the number of edges between the communities weakens the community structure, consistent with our understanding.
3.4.2 Disease thresholds
We now derive the disease thresholds in the presented example.
From Eqs. (31) and (32), we have
Assuming that
we have \((\frac{\gamma }{\mu })_1^*>0\) and \((\frac{\gamma }{\mu })_2^*>0\). Theorem 1 is then modified as follows.
Theorem 2
For the SIR model in (11), the following are true.
-
(I)
If \(\frac{\gamma }{\mu }> \max \{\frac{1}{\lambda _{11}+\lambda _{21}-1},\frac{1}{\lambda _{22}+\lambda _{12}-1}\}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \min \{\frac{1}{\lambda _{11}+\lambda _{21}-1},\frac{1}{\lambda _{22}+\lambda _{12}-1}\}\), the epidemic dies out in both communities;
-
(III)
If \(\min \{\frac{1}{\lambda _{11}+\lambda _{21}-1},\frac{1}{\lambda _{22}+\lambda _{12}-1}\}<\frac{\gamma }{\mu } \le \max \{\frac{1}{\lambda _{11}+\lambda _{21}-1},\frac{1}{\lambda _{22}+\lambda _{12}-1}\}\), the epidemic may occur in both communities, die out in both communities or occur in one community and die out in the other.
To study the effect of community structure on the disease threshold, we first study how the associations between communities affect the disease threshold in the network. Because the external degrees \(\lambda _{12}\) and \(\lambda _{21}\) meet the balance condition \(N_1\lambda _{12}=N_2\lambda _{21}\), we need only study the effect of parameter \(\lambda _{12}\). Applying Eqs. (29), (34) and (36), we have
Given the scales and average total degrees of the communities, the following investigates the effect of the external degree \(\lambda _{12}\) on the disease threshold in three cases: \(N_1=N_2\), \(N_1<N_2\) and \(N_1>N_2\).
Case 1: \(N_1=N_2\). Theorem 1 is equivalent to the following theorem.
Theorem 3
In the SIR model described in (11), the following are true.
-
(I)
If \(\frac{\gamma }{\mu }> \max \{\frac{1}{\lambda _1-1},\frac{1}{\lambda _2-1}\}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \min \{\frac{1}{\lambda _1-1},\frac{1}{\lambda _2-1}\}\), the epidemic dies out in both communities;
-
(III)
If \(\min \{\frac{1}{\lambda _1-1},\frac{1}{\lambda _2-1}\}<\frac{\gamma }{\mu } \le \max \{\frac{1}{\lambda _1-1},\frac{1}{\lambda _2-1}\}\), the epidemic may occur in both communities, die out in both communities or occur in one community and die out in the other.
At this time, the external degree \(\lambda _{12}\) and the community strength Q do not influence the disease threshold.
Case 2: \(N_1>N_2\). Let
Obviously, Eq. (37) means that
From the practical meaning of \(\lambda _{12}\) and Eq. (37), we can deduce that \(0\le \lambda _{12}<\bar{\lambda }_{12}^*\). Otherwise, we have \(\lambda _{12}\ge \bar{\lambda }_{12}^*\), meaning that \(\lambda _{22}\le 1\) or \(\lambda _{11}\le 1\), which contradicts \(\lambda _{22}>1\), \(\lambda _{11}>1\).
Phase diagram of the disease threshold \((\frac{\gamma }{\mu })^*\) versus external degree in Community 1. The blue and purple lines represent \((\frac{\gamma }{\mu })^*_2\) and \((\frac{\gamma }{\mu })^*_1\), respectively. If the parameter values fall in the red or green areas, the epidemic spreads or dies out in both communities, respectively. If the parameter values fall in the yellow area, the trend of the disease is uncertain. The parameters are \(N_1>N_2\), \(\lambda _1>\lambda _2\) in (a) and \(N_1>N_2\), \(\lambda _1<\lambda _2\) in (b) (color figure online)
When \(\lambda _2<\lambda _1\), we have \({\lambda }_{12}^*<0\). As \(\lambda _{12}\in (0,\bar{\lambda }_{12}^*)\), we can easily obtain that \((\frac{\gamma }{\mu })_2^*>(\frac{\gamma }{\mu })_1^*\). Then, from Theorem 1 and Eq. (38), we have the following theorem.
Theorem 4
If \(N_1>N_2\) and \(\lambda _1>\lambda _2\), the following are true.
-
(I)
If \(\frac{\gamma }{\mu }> \frac{1}{\lambda _2-1-(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \frac{1}{\lambda _1-1+(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic dies out in both communities;
-
(III)
If \( \frac{1}{\lambda _1-1+(\frac{N_1}{N_2}-1)\lambda _{12}}<\frac{\gamma }{\mu } \le \frac{1}{\lambda _2-1-(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic may occur in both communities, die out in both communities or occur in one community and die out in the other.
The results of Theorem 4 are demonstrated in Fig. 2a.
When \(\lambda _2>\lambda _1\), we have \({\lambda }_{12}^*>0\).
For \(\lambda _{12}\in [0, \bar{\lambda }_{12}^*)\) and \(\lambda _{12}^*< \bar{\lambda }_{12}^*\) we can easily obtain that \((\frac{\gamma }{\mu })_1^*>(\frac{\gamma }{\mu })_2^*\) (as \(\lambda _{12}\in [0,\lambda _{12}^*)\)), \((\frac{\gamma }{\mu })_1^*=(\frac{\gamma }{\mu })_2^*\) (as \(\lambda _{12}=\lambda _{12}^*\)), and \((\frac{\gamma }{\mu })_1^*<(\frac{\gamma }{\mu })_2^*\) (as \(\lambda _{12}\in (\lambda _{12}^*,\bar{\lambda }_{12}^*)\)).
Applying Theorem 1, Eq. (38) and the above facts, we obtain the following theorems.
Theorem 5
If \(N_1>N_2\), \(\lambda _2>\lambda _1\) and \(\lambda _{12}\in [0,\min \{\lambda _{12}^*, \bar{\lambda }_{12}^*\})\), the following are true.
-
(I)
If \(\frac{\gamma }{\mu }> \frac{1}{\lambda _1-1+(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \frac{1}{\lambda _2-1-(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic dies out in both communities;
-
(III)
If \(\frac{1}{\lambda _2-1-(\frac{N_1}{N_2}-1)\lambda _{12}}<\frac{\gamma }{\mu } \le \frac{1}{\lambda _1-1+(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic may occur in both communities, die out in both communities or occur in one community and die out in the other.
Theorem 6
If \(N_1>N_2\), \(\lambda _2>\lambda _1\), \(\lambda _{12}^*< \bar{\lambda }_{12}^*\) and \(\lambda _{12}=\lambda _{12}^*\), the following are true.
-
(I)
If \(\frac{\gamma }{\mu }> \frac{2}{\lambda _1+\lambda _2-2}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \frac{2}{\lambda _1+\lambda _2-2}\), the epidemic dies out in both communities.
Theorem 7
If \(N_1>N_2\), \(\lambda _2>\lambda _1\), \(\lambda _{12}^*< \bar{\lambda }_{12}^*\) and \(\lambda _{12}\in (\lambda _{12}^*,\bar{\lambda }_{12}^*)\), the following are true.
-
(I)
If \(\frac{\gamma }{\mu }> \frac{1}{\lambda _2-1-(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic occurs in both communities;
-
(II)
If \( \frac{\gamma }{\mu }\le \frac{1}{\lambda _1-1+(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic dies out in both communities;
-
(III)
If \(\frac{1}{\lambda _1-1+(\frac{N_1}{N_2}-1)\lambda _{12}}<\frac{\gamma }{\mu } \le \frac{1}{\lambda _2-1-(\frac{N_1}{N_2}-1)\lambda _{12}}\), the epidemic may occur in both communities, die out in both communities or occur in one community and die out in the other.
The results of Theorems 5, 6, and 7 are demonstrated in Fig. 2b.
Case 3: \(N_1<N_2\). This case is analyzed similarly to the above cases, so the analysis is omitted.
3.4.3 Simulations
This section confirms the accuracy of our model (11) and the theorems given in Sect. 3.4.2.
To describe the scale of the disease, we adopt the cumulative epidemic incidence, defined as the fraction of infectious or recovered nodes (Volz 2008). The cumulative incidences in the whole network, Community 1, and Community 2 are denoted as J, \(J_1\), and \(J_2\), respectively. From Eqs. (12) and (13) and noting that the hazard is identical for all nodes, we have
and
First, the accuracy of model (11) is verified in numerical simulations. Figure 3 plots the temporal evolution of the cumulative epidemic incidence under model (11). The random simulations are based on the network given in Sect. 2.2, with joint degree distributions \(P_l(k,j)=\frac{\lambda _{ll}^k\lambda _{ln}^je^{-(\lambda _{ll}+\lambda _{ln})}}{k!j!}\), \(l,n=1,2,l\ne n\). The random simulation dynamics are described on page 306 of Volz (2008). The numerical results well correlate with the random simulation results, confirming the accuracy of model (11).
Random and numerical simulations of the cumulative incidence in Community 1 (\(J_1\)) and Community 2 (\(J_2\)). The dotted lines correspond to random simulations of the SIR model based on the Monte Carlo method. The curve is the average of 100 stochastic simulation runs on the same contact network with the same disease parameters. If the final epidemic size was below 20, the trial was discarded. The solid lines correspond to numerical simulations of the cumulative incidences under system (11) with the following parameter settings: a \(N_1=1000\), \(N_2=3000\), \(\lambda _{11}=10\), \(\lambda _{12}=3\), \(\lambda _{21}=1\), \(\lambda _{22}=10\), \(\gamma =0.03\), \(\mu =0.1\); b \(N_1=1000\), \(N_2=2000\), \(\lambda _{11}=5\), \(\lambda _{12}=2\), \(\lambda _{21}=1\), \(\lambda _{22}=7\), \(\gamma =0.006\), \(\mu =0.1\); (c) \(N_1=100\), \(N_2=1000\), \(\lambda _{11}=3\), \(\lambda _{12}=0.1\), \(\lambda _{21}=0.01\), \(\lambda _{22}=10\), \(\gamma =0.03\), \(\mu =0.1\)
Second, we test the correctness of Theorem 2 in examples.
Numerical simulations of cumulative incidence in Community 1 (\(J_1\)) and Community 2 (\(J_2\)) under model (11) with the following parameter settings: a \(N_1=100\), \(N_2=1000\), \(\lambda _{11}=3\), \(\lambda _{12}=0.1\), \(\lambda _{21}=0.01\), \(\lambda _{22}=10\), \(\gamma =0.08\), \(\mu =0.1\); b \(N_1=100\), \(N_2=1000\), \(\lambda _{11}=3\), \(\lambda _{12}=0.1\), \(\lambda _{21}=0.01\), \(\lambda _{22}=10\), \(\gamma =0.008\), \(\mu =0.1\)
In panels (a) and (b) of Fig. 4, the epidemic spreaded through both communities and died out in both communities, respectively.
To show the uncertainty of the epidemic dynamics under the condition \(\frac{\gamma }{\mu } \in (\min \{(\frac{\gamma }{\mu })^*_1, (\frac{\gamma }{\mu })^*_2\}, \max \{(\frac{\gamma }{\mu })^*_1,(\frac{\gamma }{\mu })^*_2\})\), we varied the parameter settings in model (11). The simulation results are shown in Fig. 5. In this case, the epidemic may occur in community 2 and die out in community 1 (Fig. 5a), occur in both communities (Fig. 5b), or die out in both communities (Fig. 5c).
Numerical simulations of cumulative incidence in Community 1 (\(J_1\)) and Community 2 (\(J_2\)) under model (11) with the following parameter settings: a \(N_1=100\), \(N_2=1000\), \(\lambda _{11}=3\), \(\lambda _{12}=0.1\), \(\lambda _{21}=0.01\), \(\lambda _{22}=10\), \(\gamma =0.03\), \(\mu =0.1\); b \(N_1=1000\), \(N_2=2000\), \(\lambda _{11}=5\), \(\lambda _{12}=2\), \(\lambda _{21}=1\), \(\lambda _{22}=7\), \(\gamma =0.1\), \(\mu =0.6\); c \(N_1=1000\), \(N_2=2000\), \(\lambda _{11}=5\), \(\lambda _{12}=2\), \(\lambda _{21}=1\), \(\lambda _{22}=7\), \(\gamma =0.1\), \(\mu =0.7\)
4 Effect of community structure on disease spread
As mentioned above, the impact of Q on the spread of disease in a network is difficult to investigate when G is nonspecific. Using the example in Sect. 3.4, we instead show how the community structure affects the dynamics of model (11). We first study how the associations between the two communities affect the transmission of disease through the network, given \(N_1\), \(N_2\), \(\lambda _1\) and \(\lambda _2\).
Considering the symmetry of the conclusions, the influence of community structure on disease transmission is investigated in five cases, namely, (a) \(N_1=N_2\) and \(\lambda _1=\lambda _2\), (b) \(N_1>N_2\) and \(\lambda _1=\lambda _2\), (c) \(N_1=N_2\) and \(\lambda _1>\lambda _2\), (d) \(N_1>N_2\) and \(\lambda _1>\lambda _2\) and (e) \(N_1>N_2\) and \(\lambda _1<\lambda _2\).
We first study epidemic spreading under the influence of \(\lambda _{12}\), which is inversely proportional to Q. The temporal evolutions of the cumulative epidemic incidence for various values of \(\lambda _{12}\) in the five cases are shown in Figs. 6, 7, 8 and 9. From these figures, we can observe two interesting points: the first point is, when the values of the human-to-human transmissibility of the virus (i.e. \(\frac{\gamma }{\mu }\)) are larger, the cumulative incidence plots are more ‘wobbly’ than for standard epidemics; secondly, the influence of community structure on the cumulative epidemic incidence is dependent on the values of \(\frac{\gamma }{\mu }\). In order to dig deeper into the biological phenomena here, we will show both prevalence (frequency of population infected) plots and the connection between the variation in final epidemic cumulative incidence and the human-to-human transmissibility of the virus.
a Numerical simulations of cumulative incidence in the whole network (J) in Case (a) of model (11) with the following parameter settings: \(N_1=1000\), \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=10\), \(\gamma =0.02\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.1, 1, 5, 8; b numerical simulations of cumulative incidence in the whole network (J) in Case (b) of model (11) with the following parameter settings: \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=10\), \(\gamma =0.013\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.05, 0.1, 1, 3
Numerical simulations of cumulative incidence in the whole network (J) in Case (c) of model (11) with the following parameter settings: \(N_1=1000\), \(N_2=1000\), \(\lambda _{1}=20\), \(\lambda _{2}=10\), a \(\gamma =0.02\), \(\mu =0.1\) and \(\lambda _{12}=0.01\), 0.1, 1, 5, 8; b \(\gamma =0.0075\), \(\mu =0.1\) and \(\lambda _{12}=0.01\), 0.1, 1, 2, 3, 4, 8; c \(\gamma =0.006\), \(\mu =0.1\) and \(\lambda _{12}=0.01\), 0.1, 1, 3, 5, 8
Numerical simulations of cumulative incidence in the whole network (J) in Case (d) of model (11) with the following parameter settings: \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=20\), \(\lambda _{2}=10\), a \(\gamma =0.015\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.05, 0.1, 0.5, 1, 2, 3; b \(\gamma =0.008\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.1, 1, 2, 2.5, 3; c \(\gamma =0.006\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.05, 0.1, 0.3, 0.5, 0.7, 1, 2, 3
Numerical simulations of cumulative incidence in the whole network (J) in Case (e) of model (11) with the following parameter settings: \(N_1\)=3000, \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=20\), a \(\gamma =0.013\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.05, 0.1, 0.5, 1, 3, 5, 6, 6.3; b \(\gamma =0.009\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.1, 0.3, 0.5, 0.7, 1, 3, 5, 6, 6.3; c \(\gamma =0.0065\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.1, 0.5, 0.8, 1, 3, 5, 6.3
Numerical simulations of the prevalence in the whole network of model (11) with the following parameter settings: a \(N_1=1000\), \(N_2=1000\), \(\lambda _{1}=20\), \(\lambda _{2}=10\), \(\gamma =0.02\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.04, 0.1, 0.5, 1; b \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=20\), \(\lambda _{2}=10\), \(\gamma =0.015\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.04, 0.1, 0.5, 1; c \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=20\), \(\gamma =0.013\), \(\mu =0.1\), and \(\lambda _{12}=0.01\), 0.04, 0.1, 0.5, 1
For the first point, we now show the prevalence plots in Fig. 10. Since the ‘wobbly’ phenomenon is obvious in Figs. 7a, 8a and 9a, we give the prevalence plots under these three sets of parameters. Figure 10 shows that two peaks of prevalence appear when \(\lambda \) is small. With the increase in \(\lambda \), the second peak moves towards the first peak until it merges with the first one. At the same time, the first peak is raised, the arriving time of the second peak is advanced and the duration of the epidemic is shortened.
For the second point, we firstly plotted the epidemic final cumulative incidences as functions of modularity Q. The plots are presented in Figs. 11, 12, 13 and 14.
As shown in Fig. 11, the community structure does not affect the final epidemic cumulative incidence when \(\lambda _1=\lambda _2\). The increase of Q will reduce the final epidemic cumulative incidence for larger \(\frac{\gamma }{\mu }\) (Figs. 12, 13, 14). Furthermore, increasing Q accelerates the decrease in the final cumulative incidence. When \(\frac{\gamma }{\mu }\) is near the critical point, with the increase of Q, the final epidemic cumulative incidence increases first and the increment becomes less and less until it reaches zero. Then the final size decreases with increasing Q and the decrement becomes bigger and bigger. However, when \(\frac{\gamma }{\mu }\) is markedly below the critical point, the final epidemic cumulative incidence increases with increasing Q.
a Final cumulative incidences as functions of modularity Q in Cases (a) and (b) of model (11) with the following parameter settings: a \(N_1=1000\), \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=10\), \(\gamma =0.02\), \(\mu =0.1\); b \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=10\), \(\gamma =0.013\), \(\mu =0.1\)
Final cumulative incidences as functions of modularity Q in Case (c) of model (11) with the following parameter settings: \(N_1=1000\), \(N_2=1000\), \(\lambda _{1}=20\), \(\lambda _{2}=10\), a \(\gamma =0.02\), \(\mu =0.1\); b \(\gamma =0.0075\), \(\mu =0.1\); c \(\gamma =0.006\), \(\mu =0.1\)
Final cumulative incidences as functions of modularity Q in Case (d) of model (11) with the following parameter settings: \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=20\), \(\lambda _{2}=10\), a \(\gamma =0.015\), \(\mu =0.1\); b \(\gamma =0.008\), \(\mu =0.1\); c \(\gamma =0.006\), \(\mu =0.1\)
Final cumulative incidences as functions of modularity Q in Case (e) of model (11) with the following parameter settings: \(N_1=3000\), \(N_2=1000\), \(\lambda _{1}=10\), \(\lambda _{2}=20\), a \(\gamma =0.013\), \(\mu =0.1\); b \(\gamma =0.009\), \(\mu =0.1\); c \(\gamma =0.0065\), \(\mu =0.1\)
According to the above results, the impact of community structure on disease spread (especially on the final epidemic cumulative incidence) depends on the value of \(\frac{\gamma }{\mu }\). To better capture the information in Figs. 12, 13 and 14, we denote by \(\varDelta J^+\) and \(\varDelta J^-\) the increment and decrement of the final epidemic cumulative incidence, respectively, as Q increases from its lower limit \(Q(\bar{\lambda }_{12}^*)\) to Q(0.01). [The relationship between Q and \(\lambda _{12}\) is given by Eq. (35)].
Figure 15 shows the variation in final epidemic cumulative incidence with the human-to-human transmissibility of the virus (\(\frac{\gamma }{\mu }\)). These results collaborate Figs. 12, 13 and 14. As \(\frac{\gamma }{\mu }\) is larger, we obtain \(\varDelta J^+=0\) and \(\varDelta J^->0\), which reflects that strengthening the community structure to reduce the size of the disease is an effective method. At the point of \(\frac{\gamma }{\mu }\) where \(\varDelta J^-\) reaches its maximum value, the effectiveness of this method is the most significant. However, when \(\frac{\gamma }{\mu }\) is near the critical point, we have \(\varDelta J^+>0\) and \(\varDelta J^->0\), indicating that strengthening the community structure either enhances or reduces the size of the disease, which is closely related to the community strength at this moment. Comparing panels (b) and (c) in Fig. 15, we find that the community structure more strongly affects the final cumulative incidence when \(N_1>N_2\) and \(\lambda _1<\lambda _2\) than when \(N_1>N_2\) and \(\lambda _1>\lambda _2\).
Final cumulative incidences as functions of \(\gamma /\mu \) in model (11) with the following parameter settings: a \(\lambda _{1}=20\), \(\lambda _{2}=10\), \(N_1=1000\), \(N_2=1000\), and \(\lambda _{12}\) decreasing from 8 to 0.01; b \(\lambda _{1}=20\), \(\lambda _{2}=10\), \(N_1=3000\), \(N_2=1000\) and \(\lambda _{12}\) decreasing from 3 to 0.01; c \(\lambda _{1}=10\), \(\lambda _{2}=20\), \(N_1=3000\), \(N_2=1000\) and \(\lambda _{12}\) decreasing from 6.3 to 0.01
5 Conclusions
In this study, we modeled the spread of a SIR-epidemic in a complex network with two communities. By representing the network as a probability generating function, we could easily reduce the dimensionality of the model system. We first generated the two-community complex network and introduced the quality function Q, which measures the strength of the community structure in complex networks. We then introduced the states of the nodes into the generating function of the degree distribution of the network and obtained the generating function of the excessive degree distribution, thereby establishing the SIR epidemic model. Third, we obtained sufficient conditions for the outbreak and extinction of the disease. Finally, we exemplified the SIR epidemic model in a network with a Poisson joint degree distribution.
The accuracy of our main results was confirmed in simulations on this network. Fixing the total degrees and scales of the two communities, we studied the epidemic dynamics in networks with various community structures, and obtained the following results:
-
(1)
Increasing the external degree of the communities reduced the strength of the communities (expressed in terms of the modularity Q).
-
(2)
When \(N_1>N_2\), the community structure exerted a stronger effect on the final cumulative incidence when \(\lambda _1<\lambda _2\) than when \(\lambda _1>\lambda _2\).
-
(3)
In a network with strong community structure, the disease tends to be kept within the community and it may die out before spreading to the other community. However, when the human-to-human transmissibility of the virus is large enough to cause the disease to break out in all two communities, the strengthening of the community structure will lead to appearance of the second peak in prevalence. Since the stronger the community structure, the easier it is to spread the disease within the community, and the more difficult to spread the disease to the other community. So the strengthening of the community structure will bring the drop in the second peak, the delay of the arrival of the second peak and the prolongation of the epidemic duration.
-
(4)
In two communities of different sizes, enhancing the community structure reduced and enhanced the final cumulative incidence of the epidemic under strong and weak transmissibility of the virus between humans (i.e. \(\frac{\gamma }{\mu }\)), respectively. The reduction (increase) in cumulative incidence was attenuated (amplified) as \(\frac{\gamma }{\mu }\) increased. When the transmissibility was near the critical point of the outbreak, strengthening the community structure either increased or decreased the final cumulative epidemic incidence (the actual effect was uncertain). Furthermore, the increment (decrement) was attenuated (amplified) with increasing \(\frac{\gamma }{\mu }\).
How can we explain the above results? We suggest that when the transmissibility of the virus between humans is strong, a disease outbreak in the community is likely. However, whereas a strong community structure will probably retain the disease within the community, the dense network connections in a strongly connected community contain many redundant edges. Consequently, the stronger the community structure, the smaller the final cumulative epidemic incidence. Conversely, when the transmissibility of the virus between humans is weak, a disease outbreak is unlikely. In this case, strengthening the community structure (i.e. increasing the number of network connections in the community) increases the chance of disease breakout within that community. Therefore, the stronger the community structure, the larger the final cumulative epidemic incidence. When the transmissibility is near the critical point of the outbreak, the situation becomes more complicated. Initially, strengthening the community structure encourages the disease outbreak by adding new links within the community, thus increasing the final cumulative epidemic incidence. Once the disease outbreak has peaked, the disease-transmission efficiency of the newly added links steadily reduces. This occurs because the community structure gains redundant internal links while losing effective external links. Consequently, the final cumulative epidemic incidence decreases.
In summary, the size of a menacing disease can be effectively reduced toby strengthening the community structure. Furthermore, the effect becomes more obvious in stronger community structures.
In this paper, we demonstrated the SIR model in a network with a Poisson joint degree distribution as an example, and studied the detailed effects of community structure on disease spread. In future works, we will extend our SIR model to networks with power-law or other joint degree distributions.
References
Ball F (1983) The threshold behavior of epidemic models. J Appl Probab 20(2):227–241
Ball F, Neal P (2008) Network epidemic models with two levels of mixing. Math Biosci 212:69–87
Ball F, Mollison D, Scalia-Tomba G (1997) Epidemics in populations with two levels of mixing. Ann Appl Probab 7:46–89
Ball FG, Sirl DJ, Trapman P (2009) Threshold behaviour and final outcome of an epidemic on a random network with household structure. Adv Appl Probab 41:765–796
Ball FG, Sirl DJ, Trapman P (2010) Analysis of a stochastic SIR epidemic on a random network incorporating household structure. Math Biosci 224:53–73
Ball F, Britton T, Sirl D (2013) A network with tunable clustering, degree correlation and degree distribution and an epidemic thereon. J Math Biol 66:979–1019
Chris TB (2005) The spread of infectious diseases in spatially structured populations: an invasory pair approximation. Math Biosci 198:217–237
Eames KTD (2008) Modelling disease spread through random and regular contacts in clustered populations. Theor Popul Biol 73:104–111
Gleeson JP (2009) Bond percolation on a class of clustered random networks. Phys Rev E 80:036107
Gleeson JP, Melnik S, Adam H (2010) How clustering affects the bond percolation threshold in complex networks. Phys Rev E 81:066114
Gross T, DLima C, Blasius B (2006) Epidemic dynamics on an adaptive network. Phys Rev Lett 96(20):208701
House T, Keeling MJ (2010) Epidemic prediction and control in clustered populations. J Theor Biol 272(1):1–7
House T, Keeling MJ (2011) Insights from unifying modern approximations to infections on networks. J R Soc Interface 8:67–73
Huang W, Li C (2007) Epidemic spreading in scale-free networks with community structure. J Stat Mech 1(1):01014
Jin EM, Girvan M, Newman MEJ (2001) Structure of growing social networks. Phys Rev E 64(2):322–333
Kiss IZ, Miller JC, Simon PL (2017) Mathematics of epidemics on networks: from exact to approximate models. Springer, New York
Koch D, Illner R, Ma JL (2013) Edge removal in random contact networks and the basic reproduction number. J Math Biol 67:217–238
Liu Z, Hu B (2005) Epidemic spreading in community networks. EPL 72(2):315–321
Liu JZ, Wu JS, Yang ZR (2004) The spread of infectious disease with household-structure on the complex networks. Physica A 141:273–280
Luo XF, Zhang X, Sun GQ, Jin Z (2014) Epidemical dynamics of SIS pair approximation models on regular and random networks. Physica A 410:144–153
Ma J, Van DP, Willeboordse FH (2013) Effective degree household network disease model. J Math Biol 66:75–94
Miller J (2009a) Percolation and epidemics in random clustered networks. Phys Rev E 80:020901R
Miller J (2009b) Spread of infectious disease through clustered populations. J R Soc Interface 6:1121–1134
Miller JC (2011) A note on a paper by Erik Volz: SIR dynamics in random networks. J Math Biol 62:349–358
Miller JC, Volz EM (2013) Incorporating disease and population structure into models of SIR disease in contact networks. PLoS One 8:1–14
Molloy M, Reed R (1995) A critical point for random graphs with a given degree sequence. Random Struct Algorithms 6(2–3):161–180
Neal P (2007) Coupling of two SIR epidemic models with variable susceptibility and infectivity. J Appl Probab 44:41–57
Newman MEJ (2002) Spread of epidemic disease on networks. Phys Rev E 66:016128
Newman MEJ (2003a) Mixing patterns in networks. Phys Rev E 67(2):241–251
Newman MEJ (2003b) Properties of highly clustered networks. Phys Rev E 68:026121
Newman MEJ, Girvan M (2004) Finding and evaluating community struture in networks. Phys Rev E 69(2):026113
Orman K, Labatut V, Cherifi H (2013) An empirical study of the relation between community structure and transitivity. Complex Netw SCI 424:99–110
Peng XL, Small M, Xu XJ, Fu X (2013) Temporal prediction of epidemic patterns in community networks. New J Phys 15(11):17161–17175
Rowthorn RE, Laxminarayan R, Gilligan CA (2009) Optimal control of epidemics in metapopulations. J R Soc Interface 6:1135–1144
Salathé M, Jones JH (2010) Dynamics and control of diseases in networks with community structure. PLOS Comput Biol 6(4):e1000736
Sun HJ, Gao ZY (2007) Dynamical behaviors of epidemics on scale-free networks with community structure. Physica A 381:491–496
Traud A, Kelsic E, Mucha P, Porter M (2009) Community structure in online collegiate social networks. APS March Meet 53(3):526–543
Tunc I, Shaw LB (2014) Effects of community structure on epidemic spread in an adaptive network. Phys Rev E 90(2):022801
Volz E (2008) SIR dynamics in random networks with heterogeneous connectivity. J Math Biol 56:293–310
Wang B, Cao L, Suzuki H, Aihara K (2010) Epidemic spread in adaptive networks with multitype agents. J Phys A Math Theor 44:035101
Wang B, Cao L, Suzuki H, Aihara K (2012) Impacts of clustering on interacting epidemic. J Theor Biol 304:121–130
Wu XY, Liu ZH (2008) How community structure influences epidemic spread in social networks. Physica A 387:623–630
Yan G, Fu ZQ, Ren J, Wang WX (2007) Collective synchronization induced by epidemic dynamics on complex networks with communities. Phys Rev E 75(1 Pt 2):112–118
Zachary W (1977) An information flow model for conflict and fission in small groups. J Anthropol Res 33(4):452–473
Zhou J, Liu ZH (2009) Epidemic spreading in communities with mobile agents. Physica A 388:1228–1236
Acknowledgements
This work is supported by the National Natural Science Foundations of China under Grant (Nos. 11571210, 11331009, 11501339, 11101251, 11001157, 11471197, 11701348) and the Youth Science Foundation of Shanxi Province (No. 2010021001-1).The authors wish to thank the anonymous referees for their helpful feedback and suggestions.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix A: the dynamics of SIR epidemic model in a network with two communities
In this section, we develop an SIR-epidemic model involving the variables \(\theta _{mn}\), \(p^{I_m}_{S_n}\) and \(p^{S_m}_{S_n}\), \(m,n=1,2\). The method used is similar to that (Volz 2008).
Firstly, we introduce the dynamics of \(\theta _{mn}\), \(m,n=1,2.\) Consider a susceptible node ego in community l at time t with a joint degree (k, j), there are a set of k internal arcs \((\text{ ego }^l,\text{ alter }^{in}_{1})\), \((\text{ ego }^l,\text{ alter }^{in}_{2})\), \(\cdots \), \((\text{ ego }^l, \text{ alter }^{in}_{k})\) and a set of j external arcs \((\text{ ego }^l,\text{ alter }^{out}_{1})\), \((\text{ ego }^l,\text{ alter }^{out}_{2})\), \(\cdots \), \((\text{ ego }^l,\text{ alter }^{out}_{j})\) corresponding to the ego. We assume that for each arc \((\text{ ego }^l,\text{ alter }^{in}_{m})\) for \(m=1,2,\cdots , k\) and \((\text{ ego }^l,\text{ alter }^{out}_{n})\) for \(n=1,2,\cdots , j\), there will be uniform probabilities \(p^{I_l}_{S_l}=M^{I_l}_{S_l}/M^{in}_{S_l}\) and \(p^{I_n}_{S_l}=M^{I_n}_{S_l}/M^{out}_{S_l}\) for \(n\ne l\) that \(alter^{in}_{m}\) and \(alter^{out}_{n}\) are infectious, respectively. In a time dt, an expected number \(\gamma (k p^{I_l}_{S_l}+j p^{I_n}_{S_l})dt\) for \(n\ne l\) of these will be such that the infectious alter transmits to ego. Consequently, the hazard for ego becoming infected at time t is
Now let \(u^l_{(k,j)}(t)\) represent the fraction of nodes with joint degree (k, j) in community l which remain susceptible at time t. Using Eq. (42), we have
Subsequently, let
Then
Thus, the fraction of nodes which remain susceptible at time t in community l for \(l=1, 2\) are
respectively and the fraction of nodes which remain susceptible in the whole network at time t is
Similarly, for \(l,n=1,2\) and \(n\ne l\), we have
In terms of \(A^{in}_l=NG_l{(1,1)}\) and Eq. (46), we have
According to Eq. (44), the dynamics of \(\theta _{ln}\), \(l, n=1, 2\) is
which depends on the variable \(p^{I_n}_{S_l}\). To close the above system, we have to calculate the dynamics of \(p^{I_n}_{S_l}\).
Second, we introduce the dynamics of \(p^{I_n}_{S_l}\) for \(l,n=1,2\). The definition of \(p^{I_n}_{S_l}\) is
Hence,

From Eq. (47), we easily get
and
where \(l,n=1,2\) and \(n\ne l\).
The dynamics of \(M^{I_n}_{S_l}\) for l, \(n=1,2\) depends on the state of excessive neighbors (not counting the alter \(I_n\) in the chosen arc \((S_l, I_n)\)) of the ego node \(S_l\) for an random chosen arc \((S_l, I_n)\) in \(A^{I_n}_{S_l}\) for l, \(n=1,2\). The following notations in Table 2 are needed to clarify subsequent calculations.
The PGF for the excessive degree distribution of the ego node \(S_l\) in the selected arc \((S_l, I_n)\) are
The derivation of the Eq. (53) is presented in the “Appendix B” as an example.
Because arcs are distributed polynomially to nodes in sets S, I, R, we have \(g_{S_1I_1}{(X_{S_1},X_{I_1},X_{R_1},X_{S_2},X_{I_2},X_{R_2})}=g_{S_1S_1}{(X_{S_1},X_{I_1},X_{R_1},X_{S_2},X_{I_2},X_{R_2})}\), which is easy to verify by repeating the calculation in Eq. (53).
By Eqs. (53) and (54), we have
We next obtain the dynamics of \(M_{S_l}^{I_l}\). Volz (2008) introduces the approach to model SIR dynamics on networks. Here, we briefly present this approach.
There are three reasons for the reduction of the fraction of arcs between \(S_l\) and \(I_l\). One reason is the newly infectious nodes in community l. In a time infinitesimal dt, the fraction of newly infectious nodes is \(-\dot{s}_l\). According to Eq. (45), we can obtain
in which \(-\frac{N}{N_l}G_l(\theta _{ll},\theta _{ln})\dot{\theta }_{ll}\) describes a fraction nodes infected by infectious alters in community l and \(-\frac{N}{N_l}G^l(\theta _{ll},\theta _{ln})\dot{\theta }_{ln}\) describes a fraction nodes infected by infectious alters in community n for \(l\ne n\). Therefore, \(M^{I_l}_{S_l}\) decreases at rate
Because \(\delta _{S_lI_l}(I_l)\) does not count the arc along which a node was infected, another reason is the transmission from the infectious alter to the susceptible ego in this arc, which results in a reduction at a rate
The third reason is the recovery of the infectious alter in the arcs, which brings about a reduction at a rate
On the other hand, the increase of \(M^{I_l}_{S_l}\) is only due to the infection of the fraction of the susceptible egos which have excessive susceptible neighbors. So \({M^{I_l}_{S_l}}\) increases at the rate
From Eqs. (48), (59)–(60), we have
Now applying Eqs. (47), (48), (51) and (61) to Eq. (50), we obtain the dynamics of \( p^{I_l}_{S_l}\), i.e.
Similarly, we can obtain the dynamics of \(p^{I_n}_{S_l}\), \(p^{S_l}_{S_l}\) and \(p^{S_n}_{S_l}\) for \(l\ne n\) and \(l, n=1, 2\).
Appendix B: the PGF for the excessive degree distribution of the ego node in an arc
We give PGF for the excessive degree distribution of the ego node \(S_l\) in an arc \((S_l, I_n)\), which is a conditional PGF involving the state of the ego node.
For an arbitrary chosen arc \(\xi \), define some random variables. Easy to find, we list them in Table 3. Let A, B and C represent the state of a node and they may be S, I or R. Let \(P(i_1, j_1, l_1, i_2, j_2, l_2\mid \xi )\) be the conditional probability that \(\xi _{S_1}=i_1\), \(\xi _{I_1}=j_1\), \(\xi _{R_1}=l_1\), \(\xi _{S_2}=i_2\), \(\xi _{I_2}=j_2\) and \(\xi _{R_2}=l_2\) for an arbitrary chosen arc \(\xi \). The amount of excessive neighbors in state S, I and R and in community l, \(l=1,2\), of ego node in state A and in the chosen arc are distributed polynomially with probability \(p^{I_l}_{A}\), \(p^{S_l}_{A}\) and \(p^{R_l}_{A}=1-p^{I_l}_{A}-p^{S_l}_{A}\), \(l=1,2\), respectively.
The PGF for the excessive degree distribution of the ego node \(S_1\) in the chosen arc \((S_1, I_1)\), is
From the definition of the conditional probability, we have
Grounded on total probability formula, we have
and
where
Since the amount of excessive neighbors of ego node in state S and in the chosen arc \((S_1,I_1)\) are distributed polynomially with probability \(p_{S_1}^{S_l}\), \(p_{S_1}^{I_l}\), \(p^{R_l}_{S_1}=1-p^{S_l}_{S_1}-p_{S_1}^{I_l}\), \(l=1,2\), respectively, we have
From Eqs. (63), (64), (65), (66) and (67), we have
Furthermore, Eq. (68) leads to
where
and
Thus, from Eqs. (70), (71) and (72), we have
where \(\alpha =p^{S_1}_{S_1}X_{S_1}+p^{I_1}_{S_1}X_{I_1}+p^{R_1}_{S_1}X_{R_1}\) and \(\beta =p^{S_2}_{S_1}X_{S_2}+p^{I_2}_{S_1}X_{I_2}+p^{R_2}_{S_1}X_{R_2}\). As in the above analysis, we have
From Eqs. (69), (73) and (74), we have
Rights and permissions
About this article
Cite this article
Li, J., Wang, J. & Jin, Z. SIR dynamics in random networks with communities. J. Math. Biol. 77, 1117–1151 (2018). https://doi.org/10.1007/s00285-018-1247-5
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-018-1247-5