Abstract
Amylases, glycoside hydrolases widely used in several industrial processes, can be produced by many animals, plants, bacteria, and fungi. Fungal amylases from Aspergillus sp. hold remarkable importance in biotechnological applications for presenting a great catalysis efficiency in a wide range of pH and temperature. The production of amylases is mainly dependent on the genetic background of the species, i.e., Aspergillus strains, and abiotic factors. Among the major producers of amylases are the species of Aspergillus section Nigri, including Aspergillus welwitschiae. In this study, Aspergillus welwitschiae strains were evaluated for their ability to produce extracellular amylases. Among the 24 strains, wild Aspergillus welwitschiae UELAs 15.262 and mutant A. welwitschiae UELAs 15.262/35 strains showed greater potential for amylases production. The A. welwitschiae UELAs 15.262 produced more amylases (8645 U/mg) when compared to A. welwitschiae UELAs 15.262/35 (6666 U/mg). The amylases activity from partially purified crude enzymatic extract of A. welwitschiae UELAs 15.262 strain obtained at pH 5.5, 60 °C, resulted in 1.98-fold (3837 U/mg) increase in enzymatic activity. Likewise, the amylases activity from partially purified crude extract of A. welwitschiae UELAs 15.262/35 obtained at pH 5.0, 60 °C resulted in 2.2-fold (9077 U/mg) increase in amylases activity. The presence of metallic ions (Cu2+ and Fe3+) also provided an increase of amylases activity for both strains. To our knowledge, this is the first study reporting the ability of Aspergillus welwitschiae strains in order to produce amylases.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Amylases are of great biotechnological importance and can be applied in pharmaceutical, food, and even environmental industries [1, 2]. The amylases (EC 3.2.1.1) catalyze starch hydrolysis by breaking the α-1,4-O-glycosidic bonds of polysaccharides and are distributed into three major groups, the alpha-amylases (EC 3.2.2.1.1), the beta-amylases (EC 3.2.1.2), and the glucoamylases (EC 3.2.1.3) [3, 4].
This class of hydrolytic enzymes can be produced by several microorganisms, highlighting fungi such as Aspergillus niger widely used in industrial processes [2, 4]. However, the search for new enzyme-producing species like A. welwitschiae is an assiduous exercise and may also present potential for industrial and/or biotechnological applications.
The Aspergillus welwitschiae is inserted in the Aspergillus section Nigri. In this section, there are a group of species with very similar morphological characteristics that are difficult to identify, denominated as “niger aggregate,” including A. welwitschiae [5]. This species has already been isolated from sources, such as onion bulbs, garlic, and other products [6,7,8,9]. Although the production of Fumonisin B2 and Ochratoxin A has been reported in A. niger and A. welwitschiae, part of these strains do not produce these mycotoxins [7, 9, 10] and can be used in biotechnological applications.
Approximately 60% of the industrial production of enzymes is made by filamentous fungi, as they have advantageous production characteristics, such as development in wide temperature and pH ranges, and mainly produce of large amounts of enzyme [2, 11, 12]. Traditionally, amylases are produced industrially by Submerged Fermentation (SmF) allowing the monitoring of the fermentation process, followed by easy extraction and separation of the mycelium.
Regarding to amylases production by filamentous fungi, there are still some limiting factors to their availability. A greater production of these enzymes can be achieved by the association between abiotic factors and genetic background of the strains and the maximum catalytic activity, which can be obtained by characterizing the amylases.
In this sense, strategies such as the use of mutant strains for the amylases production have been developed in order to obtaining large amounts of theses enzymes. The association between abiotic and biotic factors, such as mutant strains, has provided an increase in amylases production and activity [13, 14].
The characterization of pH, temperature, and metallic ions are among the main factors involved to obtain the maximum catalytic activity of amylases. Such associated parameters adequately provide the amylases with maximum and stable catalytic activity. Variations in temperature and pH can result in protonation or deprotonation of a specific side groups at the enzyme active site, changing its chemical features, altering the enzyme conformation, and causing decrease in substrate affinity or loss activity [15]. Most amylases are holoenzymes and therefore require metallic ions as cofactors for correct active site orientation and protein stabilization [16,17,18].
Therefore, in this study it demonstrated the production and partial characterization of amylases by wild selected and mutant Aspergillus welwitschiae strains. The results showed an increase of enzyme production after optimization of abiotic parameters and partial characterization of the crude enzyme extract obtaining the maximum amylases activity.
Materials and Methods
Biological Material
This study used 24 Aspergillus strains, isolated from garlic marketed in different Brazilian states. The strains were identified as Aspergillus welwitschiae and characterized as non-producing potential Ochratoxin A and Fumonisin B2 [9]. In addition, Aspergillus welwitschiae UELAs 15.262/35 strain was introduced in this study to evaluate the production and characterization of amylases. Aspergillus welwitschiae UELAs 15.262/35 is a mutant strain for citric acid production obtained from the A. welwitschiae UELAs 15.262 wild strain, by random mutation induced by ultraviolet light [19].
Selection of Aspergillus welwitschiae Strains for amylases Production
A. welwitschiae strains were inoculated punctually in Petri dishes, containing Czapeck-Dox agar medium (soluble starch 20 g/L; NaNO3 1 g/L; K2HPO4 1 g/L; MgSO4.7H2O 1 g/L, FeSO4 0.01 g/L, Agar 15 g/L and pH 4.5) for selection of the potential amylases production. Petri dishes were then incubated at 28 ± 2 °C for 5 days and stained with iodine (KI 1 g/100 mL; I2 0.5 g/100 mL) and then the Petri dishes were incubated again for 10 days under the same conditions. The potential amylases production was evaluated by the Enzymatic Index (EI) expressed by the relationship between the diameter of the halo of enzymatic degradation + colony growth and the diameter of the growth of the colony [20].
The 24 A. welwitschiae strains were also evaluated for the potential for amylases production in liquid Czapeck-Dox medium. Suspensions of 105 conidia/mL from each A. welwitschiae strain were prepared and they were inoculated in 5 mL of Czapeck-Dox medium, at 28 ± 2 °C for 4 days. After, the Crude Enzymatic Extract (CEE) was collected and the potential amylases production was indirectly evaluated by quantifying reducing sugars by 3,5-dinitrosalicylic acid (DNS), according to Miller [21]. All the A. welwitschiae strains were performed in experimental triplicate, the standard deviation was performed with the R software.
The A. welwitschiae strain that showed higher EI and production of reducing sugars were selected for further evaluations.
Evaluation of Abiotic Parameters for the Production of amylases by Aspergillus welwitschiae Strains
The A. welwitschiae strain selected were for evaluation of abiotic parameters of temperature and pH for amylases production according to the Rotational Central Compound Design (RCCD). The evaluated temperatures and pHs were 28 °C, 30 °C, 35 °C, 40 °C, and 42 °C and 4.2, 5.0, 7.0, 9.0, and 9.8, respectively, in liquid Czapeck-Dox medium for 4 days. All analyses were performed in experimental triplicate and the standard deviation, RCCD, Analysis of Variance (ANOVA), and Tukey test (P < 0.01) were performed with the R software.
Amylases Production by A. welwitschiae Under Submerged Fermentation
A. welwitschiae strain selected under abiotic parameters for amylases production were subjected to amylases production kinetics, under SmF, in Czapeck-Dox medium. The A. welwitschiae UELAs 15.262/35 mutant strain was also submitted to the amylases production kinetics assays using the same abiotic conditions of A. welwitschiae. Each day, a portion (a flask with 5-mL Czapeck-Dox medium and the repetition) was removed to determine amylases activity, until the eighth day. A. welwitschiae strain was performed in experimental triplicate, and the standard deviation was performed with the R software.
Determination of amylases Activity of A. welwitschiae Strains
After each fermentation period (kinetics of amylases production), the cultivation was interrupted by filtration (Whatman paper no 1) to obtain the Crude Enzymatic Extract (CEE). The amylases activity was determined according to Sperotto [22].
The determination of amylases activity was carried out by enzymatic reaction, consisting of 200 µL of citrate–phosphate buffer 0.05 M, pH 6.0 (citric acid 1.05 g/100 mL; NaH2PO4 1.38 g/100 mL), 300 µL of soluble starch (1%), and 100 µL of the CEE of each A. welwitschiae strain. The reactional mixture was incubated for 30 min at 40 °C. To interrupt the enzymatic reaction, 1.5 mL of DNS was added to each sample, followed by incubation at 100 °C for 5 min and subsequent addition of 17.9 mL of distilled water. The amount of reducing sugars was evaluated at A550nm (Biochrom Libra S22). Under such conditions, one unit (U) of amylases activity is defined as the amount of enzyme that releases 1 μmoL of reducing sugars per mL of sample per minute. Protein content was determined by the method of Lowry et al. [23].
Partial Purification of the Crude Enzymatic Extract of A. welwitschiae Strains
In a flask, a total of 680 mL of each CEE obtained by the selected A. welwitschiae strain and A. welwitschiae UELAs 15.262/35 mutant strain were added, which were precipitated with 80% (1:1 v/v) ammonium sulfate saturated solution, under shaking for 2 h at 4 °C. The resulting precipitate was collected at 9050 g (Hettich, Universal 320 R), by 15 min at 4 °C. The precipitate was solubilized with 12-mL citrate–phosphate buffer (0.05 M, pH 6.0). The solubilized precipitate was dialyzed for 24 h, with three changes of distilled water for obtaining Partially Purified Crude Enzymatic Extract (PPCEE).
Analysis of Partially Purified Crude Enzymatic Extract in Sodium Dodecyl Sulfate–Polyacrylamide gel electrophoresis (SDS-PAGE)
The PPCEE of A. welwitschiae strains were mixed with an equal volume of 2 × protein loading buffer and boiled for 95 °C for 10 min. The samples were subjected to SDS-PAGE using 4% stacking gels and 12% resolving gels in an electrophoresis mini-system. The gels were visualized using Coomassie-Brilliant Blue R-250 0.1% (w/v), methanol 50% (v/v), and glacial acetic acid 7% (v/v). The standard molecular weight protein marker (AMRESCO® Mid/Low-Range Protein Molecular Weight Marker) in a range of 14.4 – 97.4 KDa were used.
Characterization of amylases of the Partially Purified Crude Enzymatic Extract of A. welwitschiae Strains
The influence of pH on the amylases production of (PPCEE) was evaluated by incubating 100 µL of A. welwitschiae strain PPCEE for 30 min, at 40 °C in 0.05 M citrate–phosphate buffer at pHs 5.5, 6.0, 6.5, and 7.0 and Tris/HCl buffer at pHs 7.5 and 8.0. The amylases activity was determined [22]. The pH that provided the increased amylases activity for each A. welwitschiae strain was used for further evaluations.
Likewise, the influence of temperature on amylases activity was performed using 100 µL of PPCEE. The temperature range was 30 °C, 35 °C, 40 °C, 45 °C, 50 °C, 55 °C, 60 °C, 65 °C, and 70 °C for 30 min. The amylases activity was determined [22].
Under conditions of pH and temperature favorable to increased amylases activity, the thermal stability of PPCEE was analyzed. The amylases reaction containing 700 µL of PPCEE was incubated for different time periods (0, 20, 40, 60, and 100 min). For each period of reaction, the amylases activity was analyzed [22].
The influence of metal ions on the amylase activity of A. welwitschiae strains was also evaluated. So, 500 µL of PPCEE were incubated with different solutions of metallic salts (1:1 v/v) of cations: Ca2+, K+, Mg2+, Mn2+, Cu2+, Fe3+, and Hg2+ at 5 mM at 30 °C for 1 h. Aliquots of 100 µL were collected and amylases activity was evaluated [22]. The characterization of amylases of the PPCEE of A. welwitschiae strains was performed in experimental triplicate, and the standard deviation was performed with the R software.
Partially Purified Crude Enzymatic Extract zymogram of amylases
The zymographic analysis of the PPCEE of A. welwitschiae strains were loaded in 10% denaturing gel run. Previously, the PPCEE were mixed with an equal volume of 2 × protein loading buffer (without β-mercaptoethanol) and boiled at 95 °C for 10 min. Gel was incubated for 1 h in 50 mM sodium acetate buffer pH 5.5. So, the gel was incubated again for 12 h at 4 °C in a solution containing 50 mM sodium acetate pH 5.5 and 0.5% starch. The gel was rinsed with water and incubated for 2 h at 37 °C in 50 mM sodium acetate buffer and treated with a solution of 1.3% I2/3% KI to stop the reaction and stain the unreacted starch background.
Results
Selection and evaluation of abiotic parameters of Aspergillus welwitschiae strains for amylases production
The 24 A. welwitschiae strains showed potential for amylases production (Table 1). The A. welwitschiae UELAs 15.262 strains were able to produce amylases, both in solid and liquid Czapeck-Dox medium (Table 1). Due to the higher production in both conditions A. welwitschiae UELAs 15.262 strain was selected for further evaluations.
The significant abiotic parameters for amylases production were obtained at 40 °C, pH 5.0–8.5 mg/mL and 35 °C, pH 7.0–7.8 mg/mL (Table 2). ANOVA showed a significant influence of pH (Supplemental Fig. S1) and considering future applications, the abiotic parameter of 35 °C, pH 7.0 for the production of amylases by A. welwitschiae UELAs 15.262 for kinetic of amylases production was selected.
Kinetics of amylases Production by A. welwitschiae in Submerged Fermentation
The kinetics of amylases production was evaluated by Smf using the previously selected abiotic parameters. At this stage under the same conditions, we introduced the A. welwitschiae UELAs 15.262/35 mutant strain, because this strain was previously selected for citric acid production.
There was an increase in the amylases production by A. welwitschiae UELAs 15.262 throughout the fermentation process, with higher production (951 U/mL, corresponding to 8645 U/mg and 0.11 mg/mL of total protein) of these enzymes on the seventh day. For A. welwitschiae UELAs 15.262/35 strain, the increase of amylases production was observed, only until the sixth day of fermentation, (580 U/mL, corresponding to 6666 U/mg and 0.087 mg/mL), according to Fig. 1.
Characterization of amylases from Partially Purified Crude Enzymatic Extract of A. welwitschiae
The PPCEE visualized on SDS-PAGE revealed up-regulated distinct proteins for both A. welwitschiae strains. Two protein bands with molecular weight of approximately 66 kDa and 100 kDa were observed in PPCEE of A. welwitschiae UELAs 15.262 and two bands of approximately 40 kDa and 50 kDa in A. welwitschiae UELAs 15.262/35 (Fig. 2A).
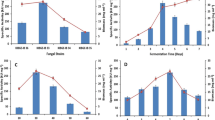
Characterization of amylases activity from Partially Purified Crude Enzymatic Extract of A. welwitschiae UELAs 15.262 and UELAs 15.262/35 strains. A SDS-PAGE of partially purified crude enzymatic extract of amylases (1- UELAs 15.262, 2- UELAs 15.262/35 strains); B influence of pH on amylases activity; C influence of temperature on amylases activity; D influence of thermal stability on amylases activity; E influence of metallic ions on amylases activity; F zymographic analysis of amylases (1- UELAs 15.262/35; 2- UELAs 15.262 strains). In the B, C, D, and E, the data were expressed using Mean ± standard deviation
The highest amylases activity of A. welwitschiae UELAs 15.262 was obtained with pH 5.5 (798.7 U/mL), while for UELAs 15.262/35 the highest enzymatic activity was observed in pH 5.0 (908.7 U/mL), as shown in Fig. 2B.
The highest amylases activity was obtained at 55 °C (1477 U/mL), 60 °C (1587 U/mL), and 65 °C (1510 U/mL) by A. welwitschiae UELAs 15.262. Furthermore, for A. welwitschiae UELAs 15.262/35 strain, the highest amylases activity was obtained at 60 °C (1982 U/mL) and 65 °C (1973 U/mL). At 70 °C there was 66% reduction in amylases activity from A. welwitschiae UELAs 15.262. In contrast, only 10% reduction in amylases activity by A. welwitschiae UELAs 15.262/35 (Fig. 2C).
Under the conditions previously established, there was decrease in amylases activity over the time. The amylases activity of A. welwitschiae UELAs 15.262 strain was more stable in the first 40 min, when compared to amylases of A. welwitschiae UELAs 15.262/35 strain. However, amylases activity from the A. welwitschiae UELAs 15.262/35 strain showed more thermal stability between 60 and 100 min, as shown in Fig. 2D.
The Cu2+ and Fe3+ ions provided greater increase on the amylases activity for both A. welwitschiae UELAs 15.262 (2410 U/mL and 2324 U/mL, respectively) and A. welwitschiae UELAs 15.262/35 (2976 U/mL and 3150 U/mL, respectively) strains (Fig. 2E). The other metallic ions (Ca2+, Mg2+, Mn2+, Hg2+) were not efficient as cofactors. So, the amylases activity from PPCEE of A. welwitschiae UELAs 15.262 strain obtained at pH 5.5, 60 °C in the presence of Cu+2 ion was 2410 U/mL or 3837 U/mg and 0.628 mg/mL of the total protein, while A. welwitschiae UELAs 15.262/35 strain at pH 5.0, 60 °C in the presence of Fe+3 ion was 3150 U/mL or 9077 U/mg and 0.347 mg/mL of the total protein.
The zymogram was performed to confirm the activity of amylases produced by A. welwitschiae strains. The starch degradation activity of the separated protein was visualized by the presence of a light band on the iodine-stained gel (Fig. 2F) indicating amylases activity of PPCEE from A. welwitschiae strains.
Discussion
The increase of amylases production by microorganisms is dependent on the genetic background of strains and/or species and abiotic factors during the fermentation process, such as temperature, pH, and type of substrate among others [2, 11, 24]. The selection of strains with genetic characteristics that favor the increase of amylases production has been of great importance. Aspergillus strains are described as good amylases producers, among which Aspergillus section Nigri [25, 26]. A. niger has been the most studied species of Aspergillus section Nigri regarding the production of amylases [27, 28]. However, in this study we have detected that A. welwitschiae is a potential amylases producer. To our knowledge, this is the first study reporting the production of amylases by A. welwitschiae strains.
In addition, A. niger and A. welwitschiae harbor potentially producing strains of Ochratoxin A and Fumonisin B2 [29, 30]. This fact highlights the importance of using strains capable of producing amylases, which are safe in terms of the inability to produce Ochratoxin A and Fumonisin B2. In this sense, all A. welwitschiae strains in this study were evaluated as genotype no-producing both mycotoxins [9] and all A. welwitschiae strains amylases produced.
A. welwitschiae UELAs 15.262 strain stood out in terms of production of these enzymes. The screening of Aspergillus strains regarding the potential for amylases production has also been reported [31].
Besides the genetic background of the strains, the abiotic factors interfere in the increase of amylases production [32]. Temperature and pH are crucial factors that have provided the amylases production. The amylases production by A. welwitschiae UELAs 15.262 strain was favored at slightly acidic pH (pH 5.0) and temperature between 35 °C and 40 °C. These data are according to reports of amylases production by species of A. niger, pH 4.0, 30 ± 2 °C [33] and Aspergillus sp., pH 5.5, 30 °C [34].
Under such specific abiotic conditions for amylases production, A. welwitschiae UELAs 15.262 and A. welwitschiae UELAs 15.262/35 strains produced the greatest amounts of amylases in the period of seven and six days of fermentation, respectively. In fact, studies have reported the highest amylases production by Aspergillus strains in periods of time longer than four days of Smf (502 U/mL—6 days – 30 °C – 180 rpm [35]; 1780 U/mL—4 days – 40 °C – pH 4,5 – under shaking [36]), under specific conditions. Aspergillus strains under Smf up to four days produced less amylases (33.52 U/mL—35 °C—4 days [37], 70.29 U/mL—30 °C—88 h—200 rpm [38], 230 U/mL—35 °C—3 days—200 rpm [39], 0.48 U/mL—28 °C—2 days—200 rpm [4]).
On the other hand, A. welwitschiae UELAs 15.262/35 strain showed ability to increase amylases production in a shorter time, under favorable conditions to future applications. It well known that the use of mutant strains has contributed to increase the production of metabolites, such as amylases [14]. The A. welwitschiae UELAs 15.262/35 mutant strain was selected for the highest accumulation of citric acid, and according to Hu et al. [39], strains that accumulate citric acid from starch materials consequently up-regulate the production of amylases.
The fact that A. welwitschiae UELAs 15.262 and UELAs 15.262/35 presented considerable amylases production rates, and the characterization of PPCEE was relevant. The proteins detected in A. welwitschiae UELAs 15.262/35 may correspond to alpha-amylases, whose molecular weight in Aspergillus sp. has been reported to be between 45 and 65 kDa [40, 41], whereas proteins detected in A. welwitschiae UELAs 15.262 may correspond to fungal glucose oxidase [42]. According to Pel et al. [43], A. niger (a species highly related to A. welwitschiae) contains four putative extracellular amylases—AmyA, AmyB, AmyC, and AamA. All these amylases are also found in the A. welwitschiae genome with high similarity. In this study, we possibly have different amylases being expressed in A. welwitschiae UELAs 15.262 and UELAs 15.262/35.
Enzyme activity is generally higher post-purification, even if partially purified, due to the larger contact surface. And in fact, in this study, the amylases activity after semi-purification reached 2410 U/mL and 3150 U/mL, corresponding to 3837 U/mg and 9077 U/mg in A. welwitschiae UELAs 15.262 and UELAs 15.262/35, respectively.
The amylases activity from PPCEE of A. welwitschiae UELAs 15.262 strain obtained at pH 5.5, 60 °C provided 1.98-fold increase in amylases activity. Similarly, the amylases activity from PPCEE of A. welwitschiae UELAs 15.262/35 strain obtained at pH 5.0, 60 °C increased 2.2-fold the activity of these enzymes. Most of the amylases produced by Aspergillus species have good enzymatic activity with pH around 4.5 to 6.0, between 50 °C and 65 °C [44, 45].
Another factor that influences enzymatic activity is the thermal stability of the enzyme, one of the main requirements for its application. The reduction of 35% and 30% in the amylases activity of A. welwitschiae UELAs 15.262 and A. welwitschiae UELAs 15.262/35 strains, respectively, at the end of 100 min allows the use of amylases produced by A. welwitschiae in future applications. Studies that evaluated the amylases activity produced by Aspergillus sp. obtained between 40 and 70% of reduction in the activity of these enzymes, in a period between 50 and 180 min and between 50 °C and 60 °C [4, 46].
Still regarding the PPCEE amylases activity of Aspergillus strains, there was increase of 25% and 20% for amylases activity of A. welwitschiae UELAs 15.262 strain and 21% and 29% for A. welwitschiae UELAs 15.262/35 strain, in the presence of Cu+2 and Fe+3 ions, respectively. Selim [47], evaluating amylases activity from A. niger, obtained increase in amylases activity of 53% using the Cu2+ ion and 38% with Fe3+ ion. Xian and Feng [48] using the ions of Cu2+ and Fe3+ (5 mM) obtained increase of 14% and 23%, respectively, in the enzymatic activity of amylases produced by the species A. tritici WZ99.
The association between pH, temperature, and metallic ions allows to obtain maximum catalytic activity, a factor of great relevance in the different sectors that use amylases, since one of the limiting obstacles to the applications of these enzymes is the poor catalytic activity. It is also noted that maximum catalytic activity provides better use of the substrate and consequent cost reduction. Several studies have shown that variations in catalysis parameters contribute to achieving the maximum catalysis activity of amylases [49, 50].
Thus, the set of abiotic factors together with the strains genetic background provide greater amylases production and similarly, specific parameters provide maximum amylases activity. This is the first report of the production and characterization of amylases from A. welwitschiae UELAs 15.262 and A. welwitschiae UELAs 15.262/35, which demonstrated that both strains have greater potential for amylases production, under the established conditions. The results showed the greater amylases production of A. welwitschiae UELAs 15.262 in relation to A. welwitschiae UELAs 15.262/35. On the other hand, under such conditions, the greater amylases activity by A. welwitschiae UELAs 15.262/35 is slightly higher compared to A. welwitschiae UELAs 15.262, suggesting that both strains could be used in future applications.
Data Availability
Not applicable.
Material Availability
Not applicable.
Code Availability
Not applicable.
References
Gopinath SCB, Anbu P, Arshad MKM, Lakshmipriya T, Voon CH, Hashim U, Chinni SV (2017) Biotechnological processes in microbial amylase production. BioMed Res Int. https://doi.org/10.1155/2017/1272193
Sundarram A, Murthy TPK (2014) α-amylase production and applications a review. J Appl Environ Microbiol 2(4):166–175. https://doi.org/10.12691/jaem-2-4-10
Gupta R, Gigras P, Mohapatra H, Goswami VK, Chauhan B (2003) Microbial α-amylases: a biotechnological perspective. Proc Biochem 38:1599–1616. https://doi.org/10.1016/S0032-9592(03)00053-0
Sethi S, Gupta S (2015) Isolation, characterization and optimization of cultural conditions for amylase production from fungi. J Biosci 4(9):3356–3363
Samson RA, Visagie CM, Houbraken J, Hong SB, Hubka V, Klaassen CH, Frisvad JC (2014) Phylogeny, identification and nomenclature of the genus Aspergillus. Stud Mycol 78:141–173. https://doi.org/10.1016/j.simyco.2014.07.004
Massi FP, Sartori D, Ferranti LS, Iamanaka BT, Taniwaki MH, Vieira MLC, Fungaro MHP (2016) Prospecting for the incidence of genes involved in ochratoxin and fumonisin biosynthesis in Brazilian strains of Aspergillus niger and Aspergillus welwitschiae. Int J Food Microbiol 221:19–28. https://doi.org/10.1016/j.ijfoodmicro.2016.01.010
Massi FP, Iamanaka BT, Barbosa RL, Sartori D, Ferrranti L, Taniwaki MH, Fungaro MHP (2020) Molecular analysis of Aspergillus section Nigri isolated from onion samples reveals the prevalence of A. welwitschiae. Braz J Microbiol. https://doi.org/10.1007/s42770-020-00390-2
Gherbawy Y, Elhariry H, Kocsubé S, Bahobial A, Deeb BE, Altalhi A, Varga J, Vágvölgyi C (2015) Molecular characterization of black Aspergillus species from onion and their potential for Ochratoxin A and Fumonisin B2 production. Foodborne Pathog Dis 12(5):414–423. https://doi.org/10.1089/fpd.2014.1870
Vanzela DOA, Massi FP, Oliveira ALM, Fungaro MHP, Sartori D (2020) Isolation and Identification of Aspergillus Section Nigri, and Genotype Associated with Ochratoxin A and Fumonisin B2 Production in Garlic Marketed in Brazil. Curr Microbiol 77:1150–1158. https://doi.org/10.1007/s00284-020-01915-6
Palumbo JD, O’keeffe TL, (2014) Detection and discrimination of four Aspergillus section Nigri species by PCR. Lett Appl Microbiol 60(2):188–195. https://doi.org/10.1111/lam.12358
Saini R, Saini HS, Dahiya A (2017) Amylases: Characteristics and industrial applications. J Pharmacogn Phytochem 6(4):1865–1871
Melnichuk N, Braia MJ, Anselmi PA, Meini MR, Romanini D (2020) Valorization of two agroindustrial wastes to produce alpha-amylase enzyme from Aspergillus oryzae by solid-state fermentation. Waste Manag 106:155–161. https://doi.org/10.1016/j.wasman.2020.03.025
Abdullah R, Ikram-Ul-HAQ, (2015) Purification and characterization of α-amylase produced by mutant strain of Aspergillus oryzae EMS-18. Nat Prod Res 29(8):710–716. https://doi.org/10.1080/14786419.2014.982648
Abdullah R, Ikram-Ul-HAQ MJ (2011) Optimization of cultural conditions for the production of alpha amylase by wild and mutant strain of Aspergillus oryzae in stirred fermenter. Pak J Bot 43(1):715–723
Devasena T (2010) Enzymology, 1st edn. Oxford University Press, New Delhi, pp 9–12
Kellis J, Todd R, Arnold F (1991) Protein stabilization by engineered metal chelation. Biotechnol 9:994–995. https://doi.org/10.1038/nbt1091-994
Saboury AA, Karbassi F (2000) Thermodynamic studies on the interaction of calcium ions with alpha-amylase. Thermochim Acta 362:121–129. https://doi.org/10.1016/S0040-6031(00)00579-7
Sarojini R, Deepika K, Rangabhashiyam S (2012) Production, characterization, kinetic studies of glucoamylase through solid-state fermentation by Aspergillus niger using agricultural residues as substrate. Int J Curr Rsrch 4:198–201
Sartori D, Ribeiro MM, Castilho POS, Bossa LF, Amador IR (2021) Produção de Ácido Cítrico a partir de linhagem mutante de A. welwitschiae UELAs 15.262/35. 2021, Brasil. Patente: Privilégio de Inovação. Número do registro: BR10202100913, título: "Produção de Ácido Cítrico a partir de linhagem mutante de A. welwitschiae UELAs 15.262/35", Instituição de registro: INPI - Instituto Nacional da Propriedade Industrial. Depósito: 11/05/2021
Hankin L, Anagnostakis SL (1975) The use of solid media for detection of enzyme production by fungi. Mycologia 67:597–607. https://doi.org/10.2307/3758395
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428. https://doi.org/10.1021/ac60147a030
Sperotto RA (2014) Protocolos e métodos de análise em laboratórios de biotecnologia agroalimentar e de saúde humana. Editora Univates, Lajeado
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with folin phenol reagent. J Biol Chem 193:265–275
Ahmad MA, Isah U, Raubilu IA, Muhammad SI, Ibrahim D (2019) An overview of the enzyme: Amylase and its industrial potentials. Bayero J Pure Appl Sci 12(1):352–358. https://doi.org/10.4314/bajopas.v12i1.53S
Saranraj P, Stella D (2013) Fungal amylase—a review. Int J Microbiol Res 4(2):203–211. https://doi.org/10.5829/idosi.ijmr.2013.4.2.75170
Uguru GC, Akinayanju JÁ, SanI A (1997) The use of Yam peel for growth of locally isolated Aspergillus niger and amylase production. Enzyme Microb Technol 21:48–51. https://doi.org/10.1016/S0141-0229(96)00225-6
Hernández MS, Rodríguez MR, Guerra NP, Rosés RP (2006) Amylase production by Aspergillus niger in submerged cultivation on two wastes from food industries. J Food Eng 73(1):93–100. https://doi.org/10.1016/j.jfoodeng.2005.01.009
Weerasooriya MKB, Piyarathne SAPM (2019) Production of extracellular amylase by Aspergillus niger under submerged fermentation using jack fruit rag as the carbon source. Indian J Tradit Knowl 19(1):158–163
Abarca ML, Bragulat MR, Castella G, Cabanes FJ (1994) Ochratoxin A production by strains of Aspergillus niger var. niger. Appl Environ Microbiol 60(7):2650–2652. https://doi.org/10.1128/aem.60.7.2650-2652.1994
Frisvad JC, Smedsgaard J, Samson RA, Larsen TO, Thrane U (2007) Fumonisin B2 production by Aspergillus niger. J Agric Food Chem 55:9727–9732. https://doi.org/10.1021/jf0718906
Saleem A, Ebrahim MKH (2014) Production of amylase by fungi isolated from legume seeds collected in Almadinah Almunawwarah. Saudi Arabia J Taibah Univ Sci 8(2):90–97. https://doi.org/10.1016/j.jtusci.2013.09.002
Sudharhsan S, Senthilkumar S, Ranjith K (2007) Physical and nutritional factors affecting the production of amylase from species of Bacillus isolated from spoiled food waste. Afr J Biotechnol 6:430–435
Spier MR, Woiciechowski AL, Vandenberghe LPS, Soccol CR (2006) Production and characterization of amylases by Aspergillus niger under solid state fermentation using agro industrials products. Int J Food Eng. https://doi.org/10.2202/1556-3758.1116
Chimata MK, Sasidhar P, Challa S (2010) Production of extracellular amylase from agricultural residues by a newly isolated Aspergillus species in solid state fermentation. Afr J Biotechnol 9(32):5162–5169
Suleimenova ZB, Saduyeva ZK, Rakhmetova ZK (2016) Alpha-amylase production from Aspergillus oryzae in submerged fermentation. Biotechnol Acta 9(4):77–82. https://doi.org/10.15407/biotech9.04.077
Sreelakshmi SN, Paul A, Vasanthi NS, Saravanan D (2014) Low-temperature acidic amylases from Aspergillus for desizing of cotton fabrics. J Text Inst 105(1):59–66. https://doi.org/10.1080/00405000.2013.810019
Ali E, El-Nagdy M, Al-Garni S, Ahmed M, Rawaa A (2017) Enhancement of alpha amylase production by Aspergillus flavus AUMC 11685 on mandarin (Citrus reticulata) peel using submerged fermentation. Eur J Biol Res 7(3):154–164. https://doi.org/10.5281/zenodo.818271
Ratnasri PV, Lakshmi BKM, Devi KA, Hemalatha KPJ (2014) Isolation, characterization of Aspergillus fumigatus and optimization of cultural conditions for amylase production. Int J Res Eng Technol 3(2):457–463
Hu W, Li W, Chen H, Liu J, Wang S, Chen J (2017) Changes in transcript levels of starch hydrolysis genes and raising citric acid production via carbono ion irradiation mutagenesis of Aspergillus niger. PLoS ONE 12(6):e0180120. https://doi.org/10.1371/journal.pone.0180120
Li S, Zuo Z, Niu D, Singh S, Permaul K, Prior BA, Shi G, Wang Z (2011) Gene cloning, heterologous expression. and characterization of a high maltose-producing α-amylase of Rhizopus oryzae. Appl Biochem Biotechnol 164:581–592. https://doi.org/10.1007/s12010-011-9159-5
Sidkey N, Abo-Shadi M, Balahmar R, Sabry R, Badrany G (2011) Purification and characterization of α-amylase from a newly isolated Aspergillus flavus F 2 Mbb. Int Res J Microbiol 2:96–103
Derakshan FK, Darvishi F, Dezfulian M, Madzak C (2017) Expression and characterization of glucose oxidase from Aspergillus niger in Yarrowia lipolytica. Mol Biotechnol 59:307–314. https://doi.org/10.1007/s12033-017-0017-8
Pel HJ, de Winde JH, Archer DB, Dyer PS, Hofmann G, Schaap PJ, Turner G, de Vries RP, Albang R, Albermann K, Andersen MR, Bendtsen JD, Benen JA, van den Berg M, Breestraat S, Caddick MX, Contreras R, Cornell M, Coutinho PM, Danchin EG, Debets AJ, Dekker P, van Dijck PW, van Dijk A, Dijkhuizen L, Driessen AJ, d’Enfert C, Geysens S, Goosen C, Groot GS, de Groot PW, Guillemette T, Henrissat B, Herweijer M, van den Hombergh JP, van den Hondel CA, van der Heijden RT, van der Kaaij RM, Klis FM, Kools HJ, Kubicek CP, van Kuyk PA, Lauber J, Lu X, van der Maarel MJ, Meulenberg R, Menke H, Mortimer MA, Nielsen J, Oliver SG, Olsthoorn M, Pal K, van Peij NN, Ram AF, Rinas U, Roubos JA, Sagt CM, Schmoll M, Sun J, Ussery D, Varga J, Vervecken W, van de Vondervoort PJ, Wedler H, Wösten HA, Zeng AP, van Ooyen AJ, Visser J, Stam H (2007) Genome sequencing and analysis of the versatile cell factory Aspergillus niger. Nat Biotechnol 25(2):221–231. https://doi.org/10.1038/nbt1282
Aisien ET, Igbinosa IH (2019) Production, purification, and characterization of α-amylase from Aspergillus niger, Aspergillus flavus and Penicillium expansum using cassava peels as substrate. Niger J Biotechnol 36(2):114–126. https://doi.org/10.4314/njb.v36i2.12
Oluwabunmi MA, Adesola AA, Grace IO (2019) Production and characterization of partially purified α-amylase from Aspergillus niger. J Phys. https://doi.org/10.1088/1742-6596/1378/4/042077
Wang J, Zhang Y, Wang X, Shang J, Li Y, Zhang H, Lu F, Liu F (2018) Biochemical characterization and molecular mechanism of acid denaturation of a novel α-amylase from Aspergillus niger. Biochem Eng J 137:222–231. https://doi.org/10.1016/j.bej.2018.06.004
Selim M (2016) Optimization of glucoamylase production by local isolate of Aspergillus niger using agro-industrial substrates under solid state fermentation. J Agric Chem Biotechnol 7(12):303–309. https://doi.org/10.21608/jacb.2016.41143
Xian L, Feng JX (2018) Purification and biochemical characterization of a novel mesophilic glucoamylase from Aspergillus tritici WZ99. Int J Biol Macromol 107:1122–1130. https://doi.org/10.1016/j.ijbiomac.2017.09.095
Ezugwu AL, Ottah VE, Eze SOO, Chilaka FC (2016) Effect of pH, various divalent metal ion and diferente substrates on glucoamylase activity obtained from Aspergillus niger using amylopectin from tiger nut starch as carbon source. Afr J Biotechnol 15(21):980–988. https://doi.org/10.5897/AJB2015.14886
Okwuenu PC, Agbo KU, Ezugwu AL, Eze SO, Chilaka FC (2017) Effect of divalent metal ions on glucoamylase activity of glucoamylase isolated from Aspergillus niger. Ferment Technol 6:141. https://doi.org/10.4172/2167-7972.1000141
Funding
This study was financed by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) (M. M. Ribeiro, Grant Code 001.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: (DS). Performed the experiments: (MMR), (CB), (MIR), and (DS). Analyzed the data: (MMR), (MIR), and (DS). Contributed reagents/materials/analysis tools: (MMR), (MIR), and (DS). Wrote the paper: (MMR), (CB), (MIR), and (DS).
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical Approval
Not applicable.
Consent to Participate
The authors had given their consent to participate.
Consent to Publication
The authors had given their consent for publication in the present journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ribeiro, M.M., Rezende, M.I., Baldo, C. et al. Aspergillus welwitschiae: A Potential amylases Producer. Curr Microbiol 79, 307 (2022). https://doi.org/10.1007/s00284-022-03005-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-03005-1