Abstract
Gut microbiota play a central role in the health of animals. The bacteria that individuals acquire as they age may therefore have a profound effect on their future fitness. Since most birds are capable of flight, they can be widely distributed in and adapted to various ecosystems. Moreover, birds are also challenged by the need to digest a wide range of food resources in their guts. However, little is known regarding how the microbial community structure in birds, especially wild birds, changes with host age. Here, we used high-throughput sequencing of the 16S rRNA V3–V4 region to depict the microbial composition and structure in the adults and nestlings of Jankowski’s bunting (Emberiza jankowskii), an endangered species of bird, during the breeding season. The results showed that the phyla Proteobacteria (52.45%), Firmicutes (13.87%), Bacteroidetes (5.76%), Actinobacteria (4.95%), Planctomycetes (4.36%), Euryarchaeota (3.20%), Acidobacteria (2.59%), Fusobacteria (2.24%), and Chloroflexi (1.8%) dominated the gut microbial communities in Jankowski’s bunting. There was no significant difference in the alpha diversity and richness among different age groups. There was also no significant difference in species richness and diversity between the nestlings and adults. However, we observed different bacterial compositions at the genus level. The genera Photobacterium and Brochothrix were detected only in the nestling groups (at days 3, 6, and 9), while Diplorickettsia was detected only in the adult group. In summary, this study can provide additional information regarding the intestinal microorganisms of wild passerine and grassland birds and provide theoretical evidence for methods to protect Jankowski’s bunting.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The class Aves occupies an important ecological niche in the ecosystem and exhibits the most diverse range of ecological functions among vertebrates [24]. Similar to other vertebrates, birds harbor diverse communities of microbiomes within their guts that collectively fulfill vital roles in providing nutrients to the host and protecting the host from pathogens [31]. Since most birds are capable of flight, they can be widely distributed in and adapted to different ecosystems [24]. In addition, birds are challenged by the need to digest a wide range of food resources and thus harbor highly complex microbiome compositions in their guts. Across the field of avian microbiology, knowledge is unevenly distributed, with several species accounting for an overwhelming majority of all microbiological investigations [32]. For example, research has mainly focused on birds that are important in agricultural production, such as chickens [2, 9], turkeys [18], and ring-necked pheasant [20], ornamental and laboratory birds such as Zebra Finches [1], and birds of evolutionary significance [7, 22, 29, 31]. However, wild birds have a wide range of lifestyles (such as migration), feeding habits, physiological characteristics and developmental processes that lead to complex changes in intestinal microbial flora. Therefore, the study of intestinal microbial research in wild birds is of great significance for the comprehensive understanding of the interaction between birds and microbial flora and the coevolution of birds and microorganisms under the pressure of natural selection.
Diet is considered an important factor that affects the gut flora composition of animals. Intestinal microbial studies of birds have been performed in some birds that have unique diets. For example, recent studies on the gut microbiome of hummingbird (Trochilidae) [21], which suggested that diet had a major influence on the microbes in birds’ guts, and research on hoatzin (Opisthocomus hoazin) [7, 8] and kakapo (Strigops habroptilus) [13] showed that diet-related species and differences in dietary habits could lead to differences in the gut microbiota. All of these studies were based on adult birds, and there have been few studies on the intestinal microbial differences between chicks and adults in wild birds. Only in the study of kakapo and hoatzin did the results show that there was no significant difference in the intestinal microbial community structure between chicks and adults due to the extremely low diversity of the intestinal microbial community [7, 30].
Studies on the gut microbiome of small passerine birds in the wild are relatively rarer. Francoise studied the differences between and influential factors of the intestinal microbes in the great tit (Parus major) and blue tit (Cyanistes caeruleus) in the nestling stage. The results showed that environmental factors were the main factors affecting the intestinal microbial changes in chicks [19]. Teyssier used the same method to study the influential factors of the intestinal microbes of great tit nestlings, and similar results were obtained. The most dominant phyla were Actinobacteria and Firmicutes, followed by Proteobacteria, in the intestine of great tit nestlings [27]. However, the intestinal microbes of the sparrow (Passer montanus) were significantly different from those of the great tit and were mainly composed of Proteobacteria and Firmicutes [16]. Major species in urban and forest ecosystems have been studied, but little is known about the composition of the intestinal microbes of birds in the most threatened grassland ecosystems, which cover 30–40% of the earth's land area [35] and are most severely disturbed by human activities such as grazing [12].
Jankowski’s bunting is a small grassland bird [14] that was listed as ‘Endangered’ on the IUCN Red List of Threatened Species in 2018 (https://dx.doi.org/10.2305/IUCN.UK.2018-2.RLTS.T22720905A132004685.en.). It has disappeared from most of its former breeding distributions, with the exceptions of Dagang, Xiergen, and Tumiji. Recently, 13 new breeding sites were discovered in Inner Mongolia. The population size of Jankowski’s bunting is approximately 10,000 individuals, which are distributed within discrete patches [10]. The study of its intestinal microbes could enable us to understand the community structure and composition of gut microbes in grassland birds.
We had two goals in this study. Our primary aim for the current study was to describe the gut microbial community structure and composition of wild Jankowski’s buntings using 16S rRNA gene high-throughput sequencing. Second, we aimed to compare the difference in the gut microbiota between nestling and adult Jankowski’s buntings and to understand the succession of the gut microbiota community structure in Jankowski’s bunting. Our study not only enriched the research available on Jankowski’s bunting and provided a theoretical basis for its protection but also enriched related research on wild birds.
Materials and Methods
Sample Collection and Preservation
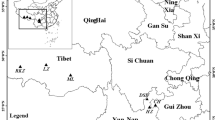
We searched for Jankowski’s bunting nests in the breeding season (May–July, 2018) in the breeding areas in Zhalute Qi, Tongliao City, Inner Mongolia, China (43 50′ 13″–45 35′ 31″ N, 119 34′ 48″–121 56′ 50″ E). The nestlings′ feces were collected during the nestling period (3 days, 6 days, and 9 days) by holding the nestlings in gloved hands for one minute. Because the amount of feces collected from the nestlings was particularly low, we pooled the feces of three nestlings into one sample. To reduce the likelihood of nest abandonment as a result of early interference, adults were captured by mist nets during the nestling period, 6–8 days after hatching, and paper was placed in the cage under the captive bird to collect feces samples. Because Jankowski’s bunting is an endangered bird, our sample size was relatively small, but this could be appropriate for a preliminary study to answer the proposed research questions. We collected 24 chick feces samples from 10 breeding nests and 6 adult bird feces samples from 6 breeding nests. Information regarding the feces samples is shown in Table 1. All samples were frozen immediately in liquid nitrogen in the wild and stored at − 80 °C after being transported to the laboratory.
DNA Extraction, PCR Amplification, and High-Throughput Sequencing
DNA was extracted from the feces samples with the EZAN Mag-Bind Soil DNA Kit (OMEGA, USA). A Qubit2.0 DNA detection Kit was used to precisely quantify genomic DNA to determine the amount of DNA to be added to the PCR. We amplified the hypervariable region V3–V4 of the 16S rRNA gene using the primer pair 341F: CCTACGGGNGGCWGCAG and 805R: GACTACHVGGGTATCTAATCC [11]. Amplification was performed in a total volume of 30 μl, with 20 ng DNA template, 15 µl 2 × master mix, 1 µl Bar-PCR primer F (10 µM), 1 µl Primer R (10 µM), and H2O to a final volume of 30 µl. The PCR program was as follows: 3 min of denaturation at 94 °C; 5 cycles of 30 s at 94 °C, 20 s at 45 °C, and 30 s at 65 °C; 20 cycles of 20 s at 94 °C, 20 s at 55 °C, and 30 s at 72 °C; and a final elongation for 5 min at 72 °C. In the second PCR, with the same reaction mix as that described above, the conditions were 3 min of denaturation at 95 °C; 5 cycles of 20 s at 94 °C, 20 s at 55 °C, and 30 s at 72 °C; and a final elongation for 5 min at 72 °C. High-throughput sequencing was performed using the Illumina MiSeq platform following the manufacturer’s instructions at Sangon, Shanghai, China.
Sequence Analysis and Statistical Analysis
Cutadapt (1.2.1; to cut joint sequences), PEAR (0.9.6; to merge sequences) [38], and Prinseq (0.20.4; for quality control) [23] software was used to preprocess the obtained sequences. Usearch (5.2.236) [5] was used to remove the nonamplified sequence region after the pretreatment of data, sequence correction, and identification of chimera by Uchime (4.2.40) [4]. Then, we performed blastn comparisons between discarded chimera sequences and representative database sequences. Results below the threshold were considered to be the outer sequence of the target region and thus eliminated. All sequences were classified into operational taxonomic units (OTUs) at a threshold of 97% similarity by Usearch, and representative sequences of each OTU were generated simultaneously. Taxonomic assignment of representative sequences was performed with the Ribosomal Database Project (RDP) naïve Bayesian rRNA classifier at an 80% confidence level [34]. The OTUs were used to analyze the alpha diversity (Shannon index and Simpson index) and richness (ACE index and Chao1 index) of the gut microbiota community structure of Jankowski’s bunting in different stages, and ANOVA was used to determine the differences among the different nestlings’ groups and between the nestling and adult groups. All statistical analyses were carried out in R.
Results
Sequencing Data of Jankowski’s Bunting
After quality filtering, we obtained a total of 522,321 high-quality reads from 14 fecal samples, averaging 37,309 reads per sample, with a median sequence length of 416 bp. The Good's coverage percentage ranged from 99.54 to 99.85% for each sample, suggesting that more than 99% of the bacteria present in samples were identified in this study (Supporting Information Table S1).
Gut Microbiota Communities in Jankowski’s Bunting
A total of 6733 OTUs were identified at the 97% sequence similarity level, and each sample contained 388-1640 OTUs (Table 2). Based on phylogenetic classification, 6733 OTUs could be assigned to a total of 42 phyla, 74 classes, 109 orders, 245 families, and 852 genera from the 14 fecal samples (Table 2). The unclassified mean rates of OTUs at the phylum and genus levels were 2.03% (0.48–3.52%) and 17% (9.78–27.25%), respectively.
At the phylum level, Proteobacteria held the overwhelming majority, with an average relative abundance of 52.45%, followed by Firmicutes (13.87%), Bacteroidetes (5.76%), Actinobacteria (4.95%), Planctomycetes (4.36%), Euryarchaeota (3.20%), Acidobacteria (2.59%), Fusobacteria (2.24%), and Chloroflexi (1.8%) (Fig. 1a). The cumulative abundance of these 9 dominant phyla was above 91.22% across all samples. At the genus level, the 17 most abundant genera were Serratia (4.27%), Acinetobacter (3.87%), Escherichia/Shigella (3.84%), Phenylobacterium (3.81%), Aeromonas (3.76%), Lactobacillus (3.59%), Sphingomonas (2.24%), Cetobacterium (2.14%), Mesorhizobium (2.07%), Pseudomonas (1.75%), Methylobacterium (1.62%), Enterococcus (1.55%), Plesiomonas (1.52%), Lactococcus (1.41%), Delftia (1.24%), Methanothrix (1.13%), and Staphylococcus (1.04%) (Fig. 1b). Among these top 17 genera, 11 genera belonged to the phylum Proteobacteria, 4 genera belonged to the phylum Firmicutes, and the other 2 genera belonged to the phyla Fusobacteria and Euryarchaeota. The compositions of the microbial community at the levels of class, order, and family are shown in Supporting Information Figs. S1–S3.
Comparison of the Gut Microbiota Compositions in Nestlings and Adults
Our samples included three 3-day-old nestlings, two 6-day-old nestlings, three 9-day-old nestlings, and six adult birds. There were no significant differences in the gut microbiota community richness (measured by the Chao1 index and ACE index) and alpha diversity (Shannon index and Simpson index) among the different stages of the nestling period, nestlings, and adults (Table 3). However, we observed different bacterial compositions at the genus level. The genera Photobacterium and Brochothrix were detected only in the nestling groups (at days 3, 6, and 9), while Diplorickettsia was detected only in the adult group. With increasing age, the genera Serratia, Delftia, and Streptococcus tended to decrease in abundance, while the genera Phenylobacterium and Methylobacterium increased with age (Supporting Information Table S2).
The results of ANOVA showed that at the phylum level, there was a significant difference between nestlings and adults in the abundance of Woesearchaeota, Euryarchaeota, and Crenarchaeota (P < 0.05). At the genus level, there was a significant difference in Woesearchaeota Incertae Sedis AR16, Methanothrix, Brachybacterium, Roseburia, Delftia, Methanomethylovorans, Rudaea, and Phenylobacterium.
Discussion
In this study, we compared the gut microbiota community structure of nestlings and adults of the critically endangered species Jankowski’s bunting and discussed the compositions and changes with age of the gut microbiota community structure of this typical grassland bird species. We found that the microbial community compositions and structures changed from nestlings to adults.
Our results showed that the microbial community of Jankowski’s bunting was mainly composed of Proteobacteria, Firmicutes, Bacteroidetes, and Actinobacteria at the phylum level, accounting for 77.03% of the total community abundance, which coincided with previously described core gut microbial communities in wild birds [31, 32]. The gut microbiota community of Jankowski’s bunting was dominated by Proteobacteria (52.45%) and Firmicutes (13.87%), accounting for 66.32% of the total community abundance, which was similar to the results of Kohl's study on sparrows [16]. The dominant phyla of intestinal flora vary among birds, for example, hoatzin [7, 8], kakapo [13], barn swallow, and bar-headed goose [15]. Different birds consume different diets, and the decomposition activities of the microbiota in the guts are different [32]. Jankowski’s bunting feeds on locusts [6], and its intestinal microflora is dominated by Proteobacteria, which include a variety of species with different metabolic activities to degrade organic substances for energy [25], specifically for the growth of nestlings and energy expenditure of adults in the nestling period. Hoatzin feed on plants [7]; therefore, the dominant phyla were Firmicutes and Bacteroidetes, which can decompose carbohydrates and plant cell wall components.
The taxonomic composition of the guts of Jankowski’s bunting nestlings was characterized by a predominance of Proteobacteria and Firmicutes and a low abundance of Actinobacteria. This was obviously different from the intestinal microbial community structure of great tit nestlings, which was dominated by Firmicutes and Actinobacteria and a low abundance of Proteobacteria [27]. Although the great tit and Jankowski’s bunting feed on insects, the species of insects vary. The great tit feeds on forest pests such as pine caterpillar and moths [36], while Jankowski’s bunting feeds on locusts. The different microbial compositions in different insects and environments may lead to the significant differences in microbial community structure between the nestlings of these two birds.
Our results are somewhat similar to those of other studies comparing microbial diversity between nestlings and adult birds. There were no significant differences in the alpha diversity of the microbiota between nestlings and adult hoatzins [7] or kakapos [30]. However, the alpha diversity metrics were higher in adult than in nestling barn swallows [17], house sparrows [16], and black-legged kittiwakes [28]. Differences in diet or other evolutionary characteristics may underlie the differences in microbial succession between these avian hosts [16].
We detected no significant differences in the gut microbiota within the nestling period. This result is similar to studies on barn swallows [17] and house swallows [16]. Although these studies revealed significant differences in the gut microbiota of adults and nestlings, it could be for other reasons. First, Kohl found that there was a significant difference between the intestinal microbial community structure of sparrow nestlings and adults. However, he collected the adult feces samples after the breeding season (in Sept–Oct). During this period, the diet of sparrows changes, and it includes more than just insects, as some of them feed on grass seeds and have a plant-based diet. Therefore, the proportion of Firmicutes in their intestines is slightly higher. We collected feces samples from adults in the nestling period, and the adults and nestlings were both feeding on locusts. However, we observed different bacterial compositions in the guts between the two groups. The intestinal microbial community structure of the nestlings was not only influenced by diet but also by the living environment of the nestlings (i.e., the microbial community structure in the nest material) [27]. In the nestling period, the environment in which the nestlings live changed, and the gut microbiota community structure of the nestlings was similar to that of the adults. Second, the sample size was insufficient, and only 14 samples were obtained in this study (8 nestling samples, 6 adult samples). Third, some birds, such as barn swallows, have nearly unchanged diets in both the adult and during the nestling period [17]; therefore, the host development of the immune system and other physiological systems could drive the changes in the microbial community structure.
The ANOVA results indicated that, at the phylum level, Woesearchaeota, Euryarchaeota, and Crenarchaeota were significantly lower in nestlings than in adults. The reason for this finding is that these three phyla are archaebacteria, the main function of which is carbon dioxide fixation, and these phyla also participate in some carbon fixation pathways in the organism [37]. The adults were stronger than the nestlings in their capacity to fix carbon. At the genus level, Photobacterium and Brochothrix were detected only in the nestling groups (at days 3, 6, and 9). Brochothrix is a genus consisting of putrefying bacteria and is harmful to nestlings [3]. Photobacterium is an indicator of organic pollutants [33]; the complete metabolic system had not yet developed in the nestlings’ bodies, and organic pollutants could not be metabolized out of the body. The adults had a complete metabolic system and could discharge some harmful substances from their body. Photobacterium and Brochothrix were eliminated in the competition between microbial species in the adult guts. Diplorickettsia was detected only in the adult group, which may be the result of adult birds being bitten by certain ticks (Ixodes ricinus) or infected with certain parasites while feeding [26]. The genera Serratia, Delftia, and Streptococcus are pathogenic bacteria, and as the immune capacity of the organism increased with age, the contents of these genera decreased. Phenylobacterium and Methylobacterium metabolize the organic waste generated in the organism. With the increase in the age of the nestlings, the organic waste generated by the organism increased, and the content of these two bacterial groups increased.
Our study was the first to focus on an endangered bird in the grassland ecosystem and studied the composition of the gut microbiota community and the process of microbiota community development in Jankowski’s bunting. However, because Jankowski’s bunting is an endangered bird, the number of samples collected was relatively small, which may not be able to fully reveal the community structure of the intestinal microorganisms. In the future, increasing attention should be paid to the potential sources of microbes and the interaction between microbes and hosts to more comprehensively reveal the mechanism underlying the development of the gut microbiota community structure in wild birds.
References
Benskin CM, Rhodes G, Pickup RW, Wilson K (2010) Diversity and temporal stability of bacterial communities in a model passerine bird, the zebra finch. Mol Ecol 19:5531–5544. https://doi.org/10.1111/j.1365-294X.2010.04892.x
Bjerrum L, Engberg RM, Leser TD, Jenson BB (2006) Microbial community composition of the ileum and cecum of broiler chickens as revealed by molecular and culture-based techniques. Poultry Sci 85:1151–1164. https://doi.org/10.1093/ps/85.7.1151
Du HC, Li XX, Lu ZX, Bie XM (2018) Antibacterial activity and mechanism of action of Plantaricin 163 against Brochothrix thermosphacta. Microbiol CHN 45:2439–2448. https://doi.org/10.13344/j.microbiol.china.171049
Edgar RC, Haas BJ, Clemente JC, Quince C (2011) UCHIME improves sensitivity and speed of chimeradetection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Wei G (2002) Ecology in Jankowski’s bunting. Jilin Science and Technology Press, Changchun
Godoy-Vitorino F, Goldfarb KC, Brodie EL, Garcia-Amado MA (2010) Developmental microbial ecology of the crop of the folivorous hoatzin. ISME J 4:611. https://doi.org/10.1038/ismej.2009.147
Godoy-Vitorino F, Goldfarb KC, Karaoz U, Leal S (2012) Comparative analyses of foregut and hindgut bacterial communities in hoatzins and cows. ISME J 6:531. https://doi.org/10.1038/ismej.2011.131
Gong J, Si W, Forster RJ, Huang R (2006) 16S rRNA gene-based analysis of mucosa-associated bacterial community and phylogeny in the chicken gastrointestinal tracts: from crops to ceca. FEMS Microbiol Eco 59:147–157. https://doi.org/10.1111/j.1574-6941.2006.00193.x
Han Z, Zhang LS, Qin B, Wang L (2018) Updated breeding distribution and population status of Jankowski’s Bunting Emberiza jankowskii in China. Bird Conserv Int. https://doi.org/10.1017/S0959270917000491
Herlemann DPR, Labrenz M, Jürgens K, Bertilsson S (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571. https://doi.org/10.1038/ismej.2011.41
Hoekstra JM, Boucher TM, Ricketts TH, Roberts C (2005) Confronting a biome crisis: global disparities of habitat loss and protection. Ecol Lett 8:23–29. https://doi.org/10.1111/j.1461-0248.2004.00686.x
Horrocks M, Salter J, Braggins J, Nichol S (2008) Plant microfossil analysis of coprolites of the critically endangered kakapo (Strigops habroptilus) parrot from New Zealand. Rev Palaeobot Palyno 149:229–245. https://doi.org/10.1016/j.revpalbo.2007.12.009
Jiang YL, Gao W, Lei FM, Wang HT (2008) Nesting biology and population dynamics of Jankowski's Bunting Emberiza jankowskii in Western Jilin, China. Bird Conserv Int 18:153–163. https://doi.org/10.1017/S0959270908000154
Kenzaka T, Katsuji T (2017) Public health implications of intestinal microbiota in migratory birds. Metagenomics for gut microbes. IntechOpen. https://doi.org/10.5772/intechopen.72456
Kohl KD, Brun A, Caviedes-Vidal E, Karasov WH (2019) Age-related changes in the gut microbiota of wild House Sparrow nestlings. Ibis 161:184–191. https://doi.org/10.1111/ibi.12618
Kreisinger J, Kropáčková L, Petrželková A et al (2017) Temporal stability and the effect of transgenerational transfer on fecal microbiota structure in a long distance migratory bird. Front Microbiol 8:50. https://doi.org/10.3389/fmicb.2017.00050
Lu J, Santo Domingo J (2008) Turkey fecal microbial community structure and functional gene diversity revealed by 16S rRNA gene and metagenomic sequences. J Microbiol 46:469–477. https://doi.org/10.1007/s12275-008-0117-z
Lucas Françoise S, Heeb P (2005) Environmental factors shape cloacal bacterial assemblages in great tit Parus major and blue tit P. caeruleus nestlings. J Avian Bio 36:510–516. https://doi.org/10.1111/j.0908-8857.2005.03479.x
Pinto RM, Tortelly R, Rodrigo CM, Delir CG (2004) Trichurid nematodes in ring-necked pheasants from backyard flocks of the State of Rio de Janeiro, Brazil: frequency and pathology. Mem I Oswaldo Cruz 99:721–726. https://doi.org/10.1590/S0074-02762004000700010
Preest MR, Folk DG, Beuchat CA (2003) Decomposition of nitrogenous compounds by intestinal bacteria in hummingbirds. Auk 120:1091–1101. https://doi.org/10.1093/auk/120.4.1091
Roggenbuck M, Schnell IB, Blom N, Bælum J (2014) The microbiome of New World vultures. Nat Commun 5:5498. https://doi.org/10.1038/ncomms6498
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics 27(6):863–864. https://doi.org/10.1093/bioinformatics/btr026
Sekercioglu CH (2006) Increasing awareness of avian ecological function. Trends Ecol Evol 21:464–471. https://doi.org/10.1016/j.tree.2006.05.007
Na-Ri S, Whon TW, Jin-Woo B (2015) Proteobacteria: microbial signature of dysbiosis in gut microbiota. Trends Biotechnol 33:496–503. https://doi.org/10.1016/j.tibtech.2015.06.011
Subramanian G, Mediannikov O, Angelakis E, Socolovschi C (2012) Diplorickettsia massiliensis as a human pathogen. Eur J Clin Microbiol 31:365–369. https://doi.org/10.1007/s10096-011-1318-7
Teyssier A, Lens L, Matthysen E, White J (2018) Dynamics of gut microbiota diversity during the early development of an avian host: evidence from a cross-foster experiment. Front Microbiol 9:1524. https://doi.org/10.3389/fmicb.2018.01524
van Dongen WFD, White J, Brandl HB et al (2013) Age-related differences in the cloacal microbiota of a wild bird species. BMC Ecol 13(1):11. https://doi.org/10.1186/1472-6785-13-11
Videvall E, Elin SJ, Bensch HM, Strandh M (2018) The development of gut microbiota in ostriches and its association with juvenile growth. bioRxiv. https://doi.org/10.1101/270017
Waite DW, Eason DK, Taylor MW (2014) Influence of hand rearing and bird age on the fecal microbiota of the critically endangered kakapo. Appl Environ Microb 80:4650–4658. https://doi.org/10.1128/AEM.00975-14
Waite DW, Taylor MW (2014) Characterizing the avian gut microbiota: membership, driving influences, and potential function. Front Microbiol 5:223. https://doi.org/10.3389/fmicb.2014.00223
Waite DW, Taylor M (2015) Exploring the avian gut microbiota: current trends and future directions. Front Microbiol 6:673. https://doi.org/10.3389/fmicb.2015.00673
Wang HQ, Dong YY, Wang LW (2018) Study on the toxicity of three emerging pollutants to Photobacterium phosphoreum. Asia J Ecotoxicol 13:179–184
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
World Resources Institute (2000) World Resources: People and ecosystems: the fraying web of life. World Resources Institute, Washington
Xie QK (1985) Location observation of breeding and feeding habits of Great tits. Liaoning For Sci Technol. 05.
Zhang C, Pan Y, Gu J, Li M (2018) Archaea diversity and carbon metabolism in mangrove sediments. Acta Microbiol Sin 58:608–617. https://doi.org/10.13343/j.cnki.wsxb.20170519
Zhang J, Kobert K, Flouri T, Stamatakis A (2013) PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30:614–620. https://doi.org/10.1093/bioinformatics/btt593
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics Statement
This study conformed to the guidelines for the care and use of experimental animals established by the Ministry of Science and Technology of the People's Republic of China (Approval number: 2006-398). The research protocol was reviewed and approved by the Ethical Committee of Jilin Agriculture University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shang, W., Li, S., Zhang, L. et al. The Composition of Gut Microbiota Community Structure of Jankowski’s Bunting (Emberiza jankowskii). Curr Microbiol 77, 3731–3737 (2020). https://doi.org/10.1007/s00284-020-02048-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02048-6