Abstract
ISP-SW5 is an intracellular alkaline serine protease gene from Bacillus velezensis SW5 that was heterologously expressed in Escherichia coli BL21 (DE3). Sequence analysis indicated that the ISP-SW5 gene has 960 bp open reading frame and encodes a protein of 319 amino acid residues. Three-dimensional structure of ISP-SW5 with the fibrinolytic activity from Bacillus velezensis was predicted by in silico analysis. Gly219 was the most likely active site for the fibrinolytic activity of ISP-SW5. The recombinant enzyme ISP-SW5 was purified by Ni–NTA Superflow Column. SDS-PAGE showed that this enzyme had a molecular mass of 34 kDa. The result of native-PAGE and N-terminal sequencing showed that the N-terminal propeptide of ISP-SW5 was cleaved during the maturation of protease. The optimum pH and temperature were 8.0 and 40 °C, respectively. Enzyme activity was markedly inhibited by PMSF and EDTA but enhanced by 5 mM Ca2+ and 2 mM Zn2+ by up to 143% and 115%, respectively. Additionally, ISP-SW5 retained 93%, 78%, and 49% relative enzyme activity after incubation with 0.5 M, 1 M and 2 M NaCl, respectively, at 4 °C for 12 h. The enzyme activity determined by casein as substrate was 1261 U/mg. ISP-SW5 could degrade fibrin at an activity of 3428 U/mg, and its properties reflect its potential application in developing a novel biological catalyst for efficient fibrin hydrolysis in medical treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiovascular diseases (CVDs), such as high blood pressure, acute myocardial infarction, ischemic heart disease, valvular heart disease, peripheral vascular disease, arrhythmias, and stroke, are the primary causes of death worldwide [1]. According to the World Health Organization, CVDs caused 15.6 million deaths in 2010 worldwide and are expected to cause more than 20 million deaths per year by 2020 [2]. CVDs are alleviated by removing or treating the thrombus with a thrombolytic agent. The thrombolytic agents that are currently available in the clinic are classified into two types: one is plasminogen activators, such as streptokinase, tissue-type(t)-PA and urokinase [3,4,5], which can activate plasminogen into plasmin. The other is the plasmin-like proteases, such as nattokinase [6], which can directly dissolve fibrin clots. However, these agents are commonly expensive and show undesired side effects, such as immunoreaction, inflammation, and hemorrhage [7]. Therefore, exploring low-cost, effective and safe thrombolytic agents with novel mechanisms that can dissolve a thrombus reliably is necessary. Fibrinolytic enzymes with potential thrombolytic applications have been recently purified from various sources, such as fermented food, earthworms, mushrooms, snake venom and microbial sources [8,9,10]. Amongst these sources, microbial source has attracted researchers’ attention due to its enzyme production superiority.
It is reported that intracellular alkaline serine protease from Bacillus velezensis possesses fibrinolytic activity. In this work, an intracellular alkaline serine protease gene from B. velezensis SW5 was isolated from fish sauce, successfully cloned into the pET-22b expression vector and then transferred into E. coli BL21 (DE3) as a factory for fibrinolytic protease production. The 3D structure, conserved amino acid residues associated with the fibrinolytic activity of ISP-SW5 and N-terminal sequencing of ISP-SW5 were also illustrated. Additionally, the fibrinolytic and fibrinogenolytic activities of ISP-SW5 were investigated to exploit its possibility as a thrombolytic therapy for the commercial production of pharmaceutical products.
Materials and Methods
Bacterial Strains, Plasmids and Culture Media
Bacillus velezensis SW5 isolated from fish sauce was used for genomic DNA extraction. The E. coli strains DH5α and BL21(DE3) purchased from TransGen Biotech (China) were used as recombinant plasmid host strains. These strains were grown in Luria–Bertani (LB) or LB broth agar (LB broth added with 1.8% agar) containing 100 μg/mL ampicillin. pET-22b (Biofeng Company, Shanghai, China), containing the inducible T7 and the pelB signal peptide was used as the expression vector.
Cloning and Plasmid Construction
The genomic DNA of strain SW5 was extracted by using the Bacteria DNA Kit (Shanghai Sangon Biotech Company, Shanghai, China). PCR was performed to amplify the intracellular serine protease gene from the genomic DNA of strain SW5 by using specific primers containing sites for the restriction enzymes BamHI and XhoI (forward primer F1: 5′-CGACGGGATCCAATGAATGGTGAAATGCATTTGA-3′ and reverse primer R1: 5′-GCCGCTCGAGGAAAGACAGCAGCTGTGCCT-3′). The following reaction conditions were used: predenaturation for 5 min at 95 °C followed by 30 cycles of 15 s at 95 °C, 15 s at 55 °C and 1 min at 72 °C. The PCR products were purified and ligated into pET-22b (Novagen). Subsequently, the recombinant plasmid was transformed into E. coli DH5α and subjected to DNA sequencing by Shanghai Sangon Biotech Company, China.
Sequence Analysis and Molecular Modeling
Nucleotide and amino acid sequences were analyzed using the BLAST(N) program of NCBI database (https://www.ncbi.nlm.nih.gov). Molecular mass and pI were determined at the ExPASy ProtParam site (https://web.expasy.org/tools/). The signal peptide was predicted using signal 3.0 server (https://www.cbs.dtu.dk/services/SignalP-3.0/).
Conserved domain was analyzed using the CDD tool (https://www.ncbi.nlm.nih.gov/cdd). A phylogenetic tree was constructed by MEGA 7, and the protein sequences were retrieved from MERPS (https://www.ebi.ac.uk/merops/index.shtml) and UniProt databases (https://www.uniprot.org/). The multiple amino acid sequence alignment of the intracellular alkaline serine protease was determined using CLUSTAL X [11]. The conserved amino acid residues associated with the fibrinolytic activity of serine protease were analyzed on the basis of MUSCLE alignment [12]. The structural modeling of the different sites of the protein was performed using SWISS-MODEL (https://www.swissmodel.expasy.org/interactive).
Expression, Purification, And Identification
Escherichia coli BL21(DE3)carrying the recombinant plasmid was grown in LB medium supplemented with ampicillin (100 μg/mL) at 37 °C. When the absorbance at 600 nm (OD600) of the culture reached 0.6, 0.2 mM isopropyl-β-d-thiogalactopyranoside (IPTG) was added to the culture medium. After further incubation at 37 °C for 8 h under shaking (180 rpm), the cells were harvested by centrifugation at 8000×g at 4 °C for 10 min. The cell pellet was washed and resuspended in 50 mM phosphate buffer solution (PBS) buffer (pH 7.4). The resuspended cells were lysed by ultrasonication (SCIENTZ Company, Ningbo, China) (300 W, 3 s bursts and 7 s pulses for 10 min) in PBS buffer. After centrifugation at 8000×g for 20 min, cell debris was removed, and the crude extract was purified under nondenaturing conditions with Ni2+–NTA Sefinose™ Resin in accordance with the manufacturer’s instructions (Shanghai Sangon Biotech Company, Shanghai, China). To remove imidazole for downstream applications, we dialyzed the eluent with a dialysis membrane. It was then stored at − 80 °C until use. Protein concentration was determined through Bradford protein assay [13]. The purified protein was detected via sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and native-polyacrylamide gel electrophoresis (native-PAGE) as described by Rekik et al. [14]. The intracellular alkaline serine protease protein band in the SDS-PAGE gel was cut out and then subjected to N-terminal sequencing analysis using mass spectrometry methods as described by Ellis et al. [15]. The activity of the protease was determined by the Folin-phenol method using casein as the substrate, as described by Yu et al. [16].
Effect of pH and Temperature on ISP-SW5 Activity
The optimum pH of ISP-SW5 was determined under different pH values. Sodium acetate buffer (0.02 mol/L) with pH 4.0–6.0, Na2HPO4–NaH2PO4 (0.02 mol/L) with pH 6.0–8.0, Tris-HCI (0.02 mol/L) with pH 8.0–9.0 and glycine–NaOH (0.02 mol/L) with pH 9.0–11.0 were used as buffers. To investigate the pH stability, the enzyme was dissolved in different pH buffers, and the mixture was incubated for 60 min at room temperature. After readjusting the pH to 8.0, the residual activity was measured under the assay condition. The activity at the beginning of the experiment was considered as the control (100%).
The optimum temperature was investigated between 30 and 90 °C, and thermostability was examined by incubating ISP-SW5 in the absence of the substrate at various temperatures (30 °C–90 °C) for 30 and 60 min. Then, the residual activity was determined as previously described. The nonheated enzyme was considered the control (100%).
Effect of Metal Ions, Inhibitors, and Surfactants on ISP-SW5 Activity
The influences of various factors, including K+, Ca2+, Mg2+, Fe2+, Cu2+, Ba2+, Zn2+, Co2+, Mn2+, A13+, PMSF, EDTA, CMC and SDS, Tween 60, Tween 80 and Trixon X-100 on enzyme activity were investigated. Besides, enzyme activity was measured to investigate salinity tolerances by incubation with NaCl (0.5, 1 and 2 M) at 4 °C for 12 h. The relative activity was calculated using enzyme activity in the absence of these chemical reagents as the control (100%).
Substrate Specificity
The substrates casein, gelatine, glycoproteins, soy protein isolate, keratin, BSA and fish peptone at a final concentration of 1.0% were added to the enzyme solution. After the samples were incubated at 50 °C for 10 min, the relative activity of the protease was determined as described above.
Fibrinolytic and Fibrinogenolytic Activity of ISP-SW5
The fibrinolytic activity of ISP-SW5 was determined using the fibrin plate method as described by Wu et al. [17]. To observe the fibrinolytic activity of ISP-SW5, 20 µL of the purified protease was added and incubated in a fibrin plate at 37 °C for 12 h. The fibrinolytic activity of ISP-SW5 was estimated by measuring the dimension of the clear zones on the fibrin plate and using the standard curve of urokinase (2–12 U) (Sigma, Shanghai, China) as the control.
The fibrinogenolytic activity was also measured by incubating 500 μg of bovine fibrinogen with 1.4 μg of ISP-SW5 in 500 μL of 50 mM PBS buffer at 37 °C. At various time intervals (0, 5, 10, 30, 60 and 120 min), the hydrolyzed products were withdrawn and analyzed against 12% SDS-PAGE.
Statistical Analysis
All determinations were performed in three independent biological replicates, and the results were expressed as the mean ± standard deviation. Statistical analysis was performed using SAS 8.1 software.
Nucleotide Sequence Accession Number
The nucleotide sequence data of ISP-SW5 (960 bp) were deposited in the GenBank database under the accession number of MN119493. The strain B. velezensis SW5 that provided the target gene was deposited in China General Microbiological Collection Centre (CGMCC) (CGMCC 1.16723).
Results and Discussion
Sequence, Structure Analysis, and Homology Model
The intracellular alkaline serine protease gene (960 bp) was amplified from the genomic DNA of B. velezensis SW5. The open reading frame (ORF) encoded 319 amino acid residues with a calculated molecular mass of 33.9 kDa and a predicted theoretical pI of 4.66. ISP-SW5 showed no signal peptide basis on SignalP analysis. The nucleic acid sequence of the ORF shared the highest homology with that from the complete genome of B. velezensis strain BIM B-439D. However, the function of this gene had not been experimentally verified. Conserved domain analysis using the CDD tool from NCBI showed that the ISP-SW5 protease sequence belonged to the peptidases S8 subtilisin subset of the peptidases_S8_S53 superfamily (Fig. 1a). The amino acid sequence of ISP-SW5 was aligned and compared with those of proteases in the serine (S) peptidases superfamily retrieved from MEROPS and UniProt databases. The phylogenetic analysis of ISP-SW5 showed close relatedness and clustering with the intracellular serine proteases belonging to the peptidase S8 family (Fig. 1b). The deduced amino acid sequence of ISP-SW5 sharing a sequence identity of 97.18% with ISP-B from Bacillus sp. [18], 86.21% with ISP-I from Bacillus sp. [19], 46.15% with ispK from B. megaterium [20] and 45.72% with NKS-21 from Bacillus sp. (Fig. 2a) [21]. Although the amino acid sequence of ISP-SW5 was highly identical to those of ISP-B and ISP-I, the properties and structures of these proteases have not been reported in detail. The homology model of ISP-SW5 was built by the I-TASSER server with the template of intracellular subtilisin protease from B. clausii (PDB ID: 2 × 8j.1.A) (Fig. 2b) [22], which exhibited 54.64% identity with ISP-SW5. According to the 3D structure constructed by I-TASSER, the three amino acid residues (Asp50, His87, and Ser246) illustrated in sticks formed a catalytic activity center similar to ISP-I and ISP-NKS21, and the residue Asn179 was an oxyanion-hole residue (Fig. 2c).
Prediction of the conserved domain of ISP-SW5 protein (a). Phylogenetic analysis of ISP-SW5 based on amino acid sequence similarity in catalytic domains by using the neighbor-joining method (b). Bootstrap values are expressed as percentages of 1000 replications and are shown at the nodes. GenBank accession numbers of proteases are also provided in parentheses
Multiple sequence alignment of ISP-SW5 and some other typical peptidases from the S8 family of intracellular alkaline serine proteases (a). The sequence identifier was ISP-SW5 from Bacillus velezensis SW5; ISP-B from Bacillus sp. WRD-2 [18]; ISP-I from Bacillus subtilis IFO3013 [19]; ispK from Bacillus megaterium ATCC 14,945 [20] and ISP-NKS21 from Bacillus sp. NKS-21 [21], the active site residues D50, H87, S246, and oxyanion-hole residue N179 are indicated in black boxes. Three-dimensional structure modeling (b) and close view of the catalytical sites (c) of the ISP-SW5 proteins. Catalytic amino acids (Asp50, His87 and, Ser246) are marked by a stick
The Similarity of the Amino Acid Sequence of ISP-SW5 with that of Other Fibrinolytic Proteases
ISP-SW5 and other reported fibrinolytic proteases were subjected to multiple sequence analysis using MUSCLE, and the results were shown in Fig. 3. Sequence analysis and alignment revealed that ISP-SW5 was 58.44% identical to Bvsp from B. vallismortis [23], 32.43% identical to BSF1 from B. subtilis [24], 11.05% identical to Bt-vsp from Bumblebees [25], 11.80% identical to F-III-2 from Lumbricus rubellus [26], 10.03% identical to AFE from Arenicola cristata [27] and 14.24% identical to UFEII from Urechis unicinctus [28]. The five residues of Gly66, Gly93, Ala111, Gly181 and Gly219 might be the conserved amino acid residues related to the fibrinolytic activity of ISP-SW5. The serine protease from Arenicola cristata contained a highly conserved amino acid residue sequence (Gly–Asp–Ser–Gly–Gly–Pro) [27]. As shown in Fig. 3, the serine proteases contained the conserved amino acid sequence Gly–Asp–Ser–Gly–Gly–Pro and had a highly conserved amino acid residue (Gly219) at the same position. This feature suggested that Gly219 was the most likely active site for the fibrinolytic activity of ISP-SW5, followed by Gly93 and Ala111 near His50, and finally Gly181 and Gly66.
Amino acid sequence alignment of ISP-SW5 with fibrinolytic serine proteases. Identical residues are shown in black boxes. Dashes represent gaps that were introduced to preserve the alignment. The amino acid sequences of Bvsp from Bacillus vallismortis [23], BSF1 from Bacillus subtilis [24], Bt-vsp from Bumblebees [25], F-III-2 from Lumbricus rubellus [26], AFE from Arenicola cristata [27] and UFEII from Urechis unicinctus [28]
Expression and Purification of ISP-SW5
ISP-SW5 was cloned from B. velezensis SW5 and expressed in E. coli BL21(DE3) successfully with a C-terminal His-tag. The recombinant protein was purified from cell lysates using Ni2+–NTA Sefinose™, and the purified protein was then analyzed using SDS-PAGE and native-PAGE (Fig. 4). One band with a weight of approximately 34 kDa was observed (Fig. 4a, lane 3), and the result of native-PAGE indicated that the molecular weight polypeptide possessed proteolytic activity (Fig. 4b, lane 4). The molecular weight of ISP-SW5 was close to that of a cloned intracellular alkaline serine protease from Bacillus sp. WRD-2 (34 kDa) [18], Bacillus megaterium ATCC 14945 (35 kDa) [20] and Bacillus sp. LCB10 (35 kDa) [29] and differed from that of serine proteases from Virgibacillus sp. SK37 (46 kDa) [30] and Virgibacillus sp. DA1-1 (28 kDa) [31].
SDS-PAGE (a) and casein zymography (b) of recombinant ISP-SW5. The samples were electrophoresed on a 12% SDS-PAGE (lanes 1–3) and 12% native-PAGE (lanes 4). After electrophoresis, the gel was stained with Coomassie brilliant blue (lanes 1–3) or detected for protease activity (lanes 4). M: protein markers; Lane 1: Noninduced whole cell lysate; Lane 2: IPTG-induced cell lysate; Lane 3: Purified recombinant ISP-SW5; Lane 4: Purified recombinant ISP-SW5 in casein zymogram
The estimated molecular weight of ISP-SW5 containing the pelB signal peptide from the expression vector was 39 kDa. However, the results of SDS-PAGE showed that the molecular weight of the purified protein was 34 kDa. Therefore, this protein was subjected to N-terminal sequencing, through which six amino acid residues at the N-terminal were identified as Ile–Ala–Val–Leu–Asp–Thr. The obtained sequence was in accordance with the fragment of the 46th to 51th amino acid residues of ISP-SW5. On the basis of this finding, 45 amino acid residues of the primary translation product were removed, and a mature protein was then produced. Literature survey showed that the N-terminal propeptide of the protein was cleaved during protease maturation [32, 33], for example, Jeong et al. reported that the propeptide at the N-terminal of ispK was cleaved during protease maturation [20]. For verification of the propeptide role, the first 45 amino acids of ISP-SW5 were removed and re-subcloned to pET-22b, and the recombinant plasmid was transformed into E. coli BL21(DE3). The crude enzyme solution showed no proteolytic activity after induction and ultrasonication mainly because the propeptide gene-defective ISP-SW5 could not form an active 3D structure. These results suggested that the N-terminal sequence of the protein was crucial for protease maturation.
Effect of pH and Temperature on the Activity and Stability of ISP-SW5
The optimal condition and stability for ISP-SW5 were determined in various buffers with different pH values (pH 4.0–11.0) and different temperature ranges (20 °C–80 °C). Enzyme activity increased progressively with increasing pH over the pH range of 4.0–8.0 and peaked at pH 8.0. This result was in good agreement with reports for alkaline proteases from B. cereus VITSN04 [34]. Slight and marked decreases in activity were observed in the pH ranges of 8.0–9.0 and 9.0–11.0, respectively. (Fig. 5a). For the pH stability, more than 87% of the remaining enzyme activity was observed at pH 6.0–9.0 after 30 min of incubation (Fig. 5b).
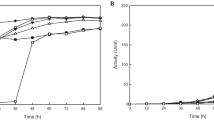
The optimal temperature of ISP-SW5 was 40 °C. Enzyme activity decreased when the temperature was increased to more than 40 °C. A similar optimal temperature (40 °C) was also reported for an alkaline serine protease from Bacillus sp. ZJ1502 [16]. ISP-SW5 possessed good thermal stability and retained 81% and 28% of its relative enzyme activity when incubated at 40 °C and 50 °C, respectively for 30 min. (Fig. 5c). Moreover, ISP-SW5 showed excellent thermostability at low temperatures. Its enzyme stability was constant after incubation at 20 °C–40 °C for 30 and 60 min, but its thermostability sharply decreased to 25% and 4% relative enzyme activity after incubation at 50 °C and 60 °C, respectively, for 60 min (Fig. 5d).
Effect of Metal Ions, Inhibitors, and Surfactants on ISP-SW5 Activity
The effects of various metals, inhibitors, and surfactants on ISP-SW5 activity were summarized in Table 1. Enzyme activity slightly increased by Triton-X100 at the concentration of 0.5% and 1% and increased to 143% and 115% in the presence of 5 mM Ca2+ and 2 mM Zn2+, respectively. PMSF is a sulfonating agent that completely inhibits the enzymatic activity of the protease by sulfonating the essential serine residue at the active site [35]. EDTA, a chelating agent, can inhibit metalloproteases indirectly by removing divalent cations [36]. EDTA and PMSF almost completely inhibited enzyme activity. This effect suggested that ISP-SW5 was a typical serine metalloprotease of the subtilase superfamily, which had calcium dependence as their well-known characteristic that was essential for their enzyme activity [30, 37]. This finding explains why calcium increases its enzyme activity. ISP-SW5 retained 93%, 78% and 49% relative enzyme activity after incubation with 0.5, 1 and 2 M NaCl, respectively, mainly because ISP-SW5 originated from a moderately halophilic bacterium B. velezensis SW5.
Substrate Specificity
Substrate specificity analysis was performed to determine the optimum substrate for the ISP-SW5-catalyzed reaction (Table 2). These experiments showed that casein was the ideal suitable substrate in these test substrates, the protease activity of casein was considered 100%. The relative enzyme activities of 51%, 45%, and 33% were detected on soy protein isolate, fish protein, and glycoproteins, respectively. By contrast, ISP-SW5 showed only 21% and 14% relative enzyme activity against keratin and bovine serum albumin (BSA) substrates, respectively. ISP-SW5 exhibited the lowest activity of 3% against gelatine. ISP-SW5 could barely hydrolyze gelatine likely because gelatine was mainly composed of left-handed α-helix and some amino acids with low molecular weights [16]. Besides, the specific activity of the purified ISP-SW5 with casein as the substrate was 1261 U/mg under optimal conditions. This value was estimated to be 26.8-fold higher than that of the cell lysate of ISP-SW5, which had a specific activity of 47 U/mg.
Fibrinolytic and Fibrinogenolytic Activity
ISP-SW5 formed clear hydrolyzed zones on the fibrin plate, whereas PBS buffer, which was used as a negative control, did not show any clear zones. This phenomenon indicated that ISP-SW5 possessed fibrinolytic activity (Fig. 6). The fibrinolytic activity of ISP-SW5 was calculated to be 3428 U/mg basis on the standard curve of urokinase activity (Fig. 6b). The measured activity of ISP-SW5 was higher than that of the fibrinolytic enzyme from Rhizopus chinensis 12 (2143.4 U/mg) [38], but lower than that of serine protease from Bacillus sp. 4-L-16 (3863 U/mg) [23].
Fibrinolytic and fibrinogenolytic activities of purified ISP-SW5. The fibrinolytic activity of ISP-SW5 on the fibrin plate, 1: Negative control, 2: Purified ISP-SW5 (2.8 μg) (a); The fibrinolytic activity of urokinase, 1: Urokinase (2 U), 2: Urokinase (4 U), 3: Urokinase (6 U), 4: Urokinase (8 U), 5: Urokinase (10 U), 6: Urokinase (12 U) (b); The hydrolytic products from fibrinogen were analyzed in 12% SDS-PAGE. Lanes1-6, degradation products after 0, 5, 10, 30, 60 and 120 min of incubation, respectively (c)
The SDS-PAGE analysis of the fibrinogenolytic activity of ISP-SW5 showed that the Aα-chain of fibrin was completely hydrolyzed within the first 5 min. Then, the Bβ-chain was completely hydrolyzed within 60 min. Finally, the γ-chain was completely hydrolyzed within 120 min (Fig. 6c). This hydrolysis pattern was similar to the hydrolysis pattern of fibrinogen proteases [10, 23]. Based on the results, it could be seen that ISP-SW5 exhibited a broad pH range (7–9) and a moderate thermophilic character. The surroundings of temperature and pH were close to that of human blood [39]. Therefore, this enzyme can be further developed as a potential candidate for thrombolytic therapy. However, the thrombolytic effect of ISP-SW5 in vivo needs further experimental research.
Conclusions
In this study, the gene encoding intracellular alkaline serine protease, ISP-SW5, from Bacillus velezensis SW5 isolated from fish sauce was cloned and expressed as a soluble protein in E.coli and purified with a molecular mass was 34 kDa. Three-dimensional structure of intracellular serine protease with the fibrinolytic activity from Bacillus velezensis was first described by in silico analysis. Gly219 was the most likely active site for the fibrinolytic activity of ISP-SW5. The result of N-terminal of sequencing and Native-PAGE showed that the N-terminal propeptide of ISP-SW5 was cleaved during the maturation of protease. Protease activity analyses showed that ISP-SW5 had an optimum temperature at 40 °C, optimum pH at 8.0. Besides, Zn2+, Ca2+ enhanced the enzyme activity, but EDTA and PMSF almost completely inhibited the enzyme activity. Additionally, ISP-SW5 had ideal relative enzyme activity under high osmotic pressure (0.5, 1, 2 M NaCl). Finally, it was reported that ISP-SW5 from Bacillus velezensis SW5 had fibrinolytic activity. Analysis of its fibrinogen hydrolyzing activity revealed that Aα-chain was firstly hydrolyzed, followed by the β-chain, and the γ-chain was the last to be hydrolyzed. These results indicated that ISP-SW5 was a prospective candidate for antithrombotic drug development.
References
Simkhada JR, Mander P, Cho SS, Yoo JC (2010) A novel fibrinolytic protease from Streptomyces sp. CS684. Process Biochem 45(1):88–93
Hu YL, Yu D, Wang ZT, Hou JJ, Tyagi R, Liang YX, Hu YM (2019) Purification and characterization of a novel, highly potent fibrinolytic enzyme from Bacillus subtilis DC27 screened from Douchi, a traditional Chinese fermented soybean food. Sci Rep 9:1–10
Arnesen H, Heilo A, Jakobsen E, Ly B, Skaga E (1978) A prospective study of streptokinase and heparin in the treatment of deep vein thrombosis. Acta Med Scand 203(6):457–463
Broderick JP, Tomsick TA (2013) Reimbursement for thrombectomy devices in patients who are ineligible for intravenous tissue-type plasminogen activator. Stroke 44(5):1215–1216
Carriero MV, Stoppelli MP (2011) The urokinase-type plasminogen activator and the generation of inhibitors of urokinase activity and signaling. Curr Pharm Des 17(19):1944–1961
Kwon EY, Kim KM, Kim MK, Lee IY, Kim BS (2011) Production of nattokinase by high cell density fed-batch culture of Bacillus subtilis. Bioproc Biosyst Eng 34(7):789–793
Kool KB, Suh HJ, Ra KS, Kim YH, Joo HS, Choi JW (2010) Fibrinolytic activity of a novel serine protease from the hemolymph of a polychaeta, Periserrula leucophryna. J Korean Soc Appl Bio 53(2):149–157
Deng ZH, Wang SH, Li Q, Ji X, Zhang LZ, Hong M (2010) Purification and characterization of a novel fibrinolytic enzyme from the polychaete, Neanthes japonica (Iznka). Bioresour Technol 101(6):1954–1960
Patel GK, Kawale AA, Sharma AK (2012) Purification and physicochemical characterization of a serine protease with fibrinolytic activity from latex of a medicinal herb Euphorbia hirta. Plant Physiol Biochem 52(3):104–111
Yao Z, Kim JA, Kim JH (2019) Characterization of a fibrinolytic enzyme secreted by Bacillus velezensis BS2 isolated from sea squirt jeotgal. J Microbiol Biotechn 29(3):347–356
Jeanmougin F, Thompson JD, Gouy M, Higgins DG, Gibson TJ (1998) Multiple sequence alignment with Clustal X. Trends Biochem Sci 23(10):403–405
Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter SC, Finn RD et al (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 47(W1):W636–W641
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254
Rekik H, Zarai Jaouadi N, Gargouri F, Bejar W, Frikha F, Jmal N, Bejar S, Jaouadi B (2019) Production, purification and biochemical characterization of a novel detergent-stable serine alkaline protease from Bacillus safensis strain RH12. Int J Biol Macromol 121(1):1227–1239
Ellis S, Fairwell T, Lovins RE (1972) Quantitative protein sequencing using mass spectrometry: N-terminal sequence analysis of small quantities of peptides from the mass spectral analysis of the N-methylthiourea derivatives. Biochem Biophys Res Commun 49(6):1407–1413
Yu P, Huang XX, Ren Q, Wang XX (2019) Purification and characterization of a H2O2-tolerant alkaline protease from Bacillus sp. ZJ1502, a newly isolated strain from fermented bean curd. Food Chem 274(2):510–517
Wu B, Wu LC, Chen DJ, Yang ZJ, Luo MY (2009) Purification and characterization of a novel fibrinolytic protease from Fusarium sp. CPCC 480097. J Ind Microbiol Biot 36(3):451–459
An SY, Ok M, Kim JY, Jang MS, Cho YS, Choi YL, Kim CH, Lee YC (2004) Cloning, high-level expression and enzymatic properties of an intracellular serine protease from Bacillus sp. WRD-2. Indian J Biochem Biol 41(4):141–147
Koide Y, Nakamura A, Uozumi T, Beppu T (1986) Cloning and sequencing of the major intracellular serine protease gene of Bacillus subtilis. J Bacteriol 167(1):110–116
Jeong YJ, Baek SC, Kim H (2018) Cloning and characterization of a novel intracellular serine protease (IspK) from Bacillus megaterium with a potential additive for detergents. Int J Biol Macromol 108(3):808–816
Yamagata Y, Ichishima E (1995) A new alkaline serine-protease from alkalophilic Bacillus sp. cloning, sequencing, and characterization of an intracellular protease. Curr Microbiol 30(6):357–366
Vevodova J, Gamble M, Kunze G, Ariza A, Dodson E, Jones DD, Wilson KS (2010) Crystal structure of an intracellular subtilisin reveals novel structural features unique to this subtilisin family. Structure 18(6):744–755
Cheng QP, Xu FY, Hu N, Liu XS, Liu ZD (2015) A novel Ca2+-dependent alkaline serine-protease (Bvsp) from Bacillus sp. with high fibrinolytic activity. J Mol Catal B 117(7):69–74
Agrebi R, Haddar A, Hmidet N, Jellouli K, Manni L, Nasri M (2009) BSF1 fibrinolytic enzyme from a marine bacterium Bacillus subtilis A26: purification, biochemical and molecular characterization. Process Biochem 44(11):1252–1259
Qiu Y, Choo YM, Yoon HJ, Jia J, Cui Z, Wang D, Kim DH, Sohn HD, Jin BR (2011) Fibrin(ogen)olytic activity of bumblebee venom serine protease. Toxicol Appl Pharmacol 255(2):207–213
Sugimoto M, Nakajima N (2001) Molecular cloning, sequencing, and expression of cDNA encoding serine protease with fibrinolytic activity from earthworm. Biosci Biotechnol Biochem 65(7):1575–1580
Zhao CL, Ju JY (2015) Cloning, expression and activity analysis of a novel fibrinolytic serine protease from Arenicola cristata. J Ocean Univ China 14(3):533–540
Bi QQ, Chu JX, Feng YL, Jiang ZQ, Han BQ, Liu WS (2013) Purification and characterization of a new serine protease with fibrinolytic activity from the marine invertebrate Urechis unicinctus. Appl Biochem Biotech 170(3):525–540
Hou Y, Lu F, Tian J, Tian Y (2019) Cloning, heterologous expression and characterization of an intracellular serine protease from Bacillus sp. LCB10. Appl Biochem Micro 55(5):482–488
Phrommao E, Yongsawatdigul J, Rodtong S, Yamabhai M (2011) A novel subtilase with NaCl-activated and oxidant-stable activity from Virgibacillus sp. SK37. BMC Biotechnol 11:65
Chen X, Zhou C, Xue Y, Shi J, Ma Y (2018) Cloning, expression, and characterization of an alkaline protease, AprV, from Vibrio sp. DA1-1. Bioprocess Biosyst Eng 41(10):1437–1447
Choi JH, Lee SY (2004) Secretory and extracellular production of recombinant proteins using Escherichia coli. Appl Microbiol Biot 64(5):625–635
Mergulhao FJ, Summers DK, Monteiro GA (2005) Recombinant protein secretion in Escherichia coli. Biotechnol Adv 23(3):177–202
Sundararajan S, Kannan CN, Chittibabu S (2011) Alkaline protease from Bacillus cereus VITSN04: potential application as a dehairing agent. J Biosci Bioeng 111(2):128–133
Ibrahim ASS, Elbadawi YB, El-Tayeb MA, Al-maary KS, Maany DAF, Ibrahim SSS, Elagib AA (2019) Alkaline serine protease from the new halotolerant alkaliphilic Salipaludibacillus agaradhaerens strain AK-R: purification and properties. 3 Biotech 9(11):391
Salihi A, Asoodeh A, Aliabadian M (2017) Production and biochemical characterization of an alkaline protease from Aspergillus oryzae CH93. Int J Biol Macromol 94(Pt B):827–835
Siezen RJ (1996) Subtilases: subtilisin-like serine proteases. Adv Exp Med Biol 379:75–93
Liu XL, Lian-xiang D, Fu-ping L, Xi-qun Z, Jing X (2005) Purification and characterization of a novel fibrinolytic enzyme from Rhizopus chinensis 12. Appl Microbiol Biotechnol 67(2):209–214
Zhao N, Ma L, Bai S, Wang C (2018) Correlation of blood glucose level and blood pH value with cardiac enzyme, amylase and other markers in patients with diabetic ketoacidosis. Int J Clin Exp Med 11(9):10208–10214
Acknowledgements
This work was financially supported by the Natural Science Fund Project of Ningbo (2018A610337) and Public Welfare Project of Zhejiang (GG19C200003).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: HY, YL, YN—Performed the experiments: HY, CW—Analyzed the data: HY, NL—Contributed reagents/materials/analysis tools: ZW, PW, XZ—Wrote the manuscript: HY—Revised and approved the final version of this work: HY ZW.
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest declared.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, H., Liu, Y., Ning, Y. et al. Characterization of an Intracellular Alkaline Serine Protease from Bacillus velezensis SW5 with Fibrinolytic Activity. Curr Microbiol 77, 1610–1621 (2020). https://doi.org/10.1007/s00284-020-01977-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-01977-6