Abstract
We used high-throughput sequencing analysis, which targeted the hypervariable V3–V4 region of the bacterial 16S rRNA gene, to investigate the microbiota in fecal material from ten wild painted turtles (Chrysemys picta) captured in southeastern Wisconsin. The most predominant bacterial phylum detected in all samples was the Firmicutes (relative abundance for all samples 96.4% to 68.3%). The next most predominant phylum was Bacteroidetes (relative abundance for all samples 23.9% to 7.8%) in eight samples. Fusobacteria (relative abundance for all samples 22.2% to 0%) was the second most predominant in the other two samples.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Painted turtles (Chrysemys picta) are habitat and food generalists that are abundant throughout much of North America [1]. Although they are a relatively common species and the focus of much research [reviewed by 1], few studies have investigated the microbiota in the digestive systems of wild individuals. In fact, few studies have looked at the bacteria present in the fecal material of wild turtles, in general. Despite this, Mitchell and McAvoy [2] used a culture dependent approach to examine the enteric bacteria present in 174 wild freshwater turtles from across seven different species, including painted turtles, in Central and Southeastern Virginia. In addition, other studies centered on the enteric bacteria of wild turtles exist, but these frequently focus on the presence or prevalence of Salmonella spp. in captured individuals [e.g. 3, 4].
The aim of this preliminary investigation was to characterize the bacterial diversity in the fecal material of painted turtles living in the wild. There are numerous reasons why investigating the enteric bacterial diversity within wild turtles can be important to understanding their basic physiology and ecology. Further, assessment of internal microbes can provide information on factors that could influence turtle population dynamics (e.g., mortality, population stability, and regulation), particularly if those microbes are pathogens or contribute to other disease. The gut microbiomes of mammals, for example, are very important to the health, and perhaps the fitness, of the animal [5,6,7]. Emerging infectious diseases can pose important threats to the stability of wildlife populations and biodiversity [8]. As such, inventories of fecal microbiota can be used to screen for potential threats to population health, or act as a baseline for comparison by future researchers.
In early August 2017, we deployed four baited hoop traps at a wetland in southeastern Wisconsin (Jefferson County). Hoop traps were checked at ca 15 h post-deployment and all captured adult turtles were removed, sexed using external morphological characteristics, measured (straight-line carapace length), and weighed using drop scales (n = 10; Table S1). We then transported captured turtles to an indoor facility and placed them individually into large plastic tubs partially filled with well water. We visually inspected all turtles at regular intervals over the next 48 h to assess their general health and also to check for the presence of fecal pellets in the water. We immediately removed any fecal pellets observed, placed them in plastic vials, and stored them in a regular kitchen freezer (− 17.8 °C) for ca 14–48 h. By 48 h post-capture, all turtles were released at their original capture location. All fecal samples were then transported to a − 40 °C freezer and stored until required for laboratory investigations. In this study, samples were collected under the auspices of Institutional Animal Care and Use Committee (IACUC) approved permit K145011020Q.
The inner portion of the ten painted turtle fecal samples used in this study were analyzed using ZymoBIOMICS® service performed by Zymo Research (Irvine, CA). The ZymoBIOMICS®-96 MagBead DNA Kit (Zymo Research, Irvine, CA) was used to extract DNA using an automated platform. Bacterial 16S rRNA gene sequencing was performed using the Quick-16S™ NGS Library Prep Kit (Zymo Research, Irvine, CA). The bacterial 16S rRNA gene sequencing primers amplified the V3–V4 region of the 16S rRNA gene. The sequencing library was prepared using an innovative library preparation process in which PCR reactions were performed in real-time PCR machines to control cycles and therefore prevent limit PCR chimera formation. The final PCR products were quantified with qPCR fluorescence readings and pooled together based on equal molarity. The final pooled library was cleaned up with the Select-a-Size DNA Clean & Concentrator™ (Zymo Research, Irvine, CA), then quantified with TapeStation® and Qubit®. The final library was sequenced on Illumina® MiSeq™ with a v3 reagent kit (600 cycles). The sequencing was performed with > 10% PhiX spike-in. Unique amplicon sequences were inferred from raw reads using the Dada2 pipeline [9]. Chimeric sequences were also removed with the Dada2 pipeline. Taxonomy assignment was performed using Uclust from Qiime v.1.9.1. Composition visualization and alpha-diversity analyses were performed with Qiime v.1.9.1 [10].
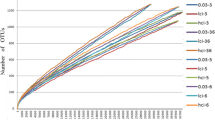
In the present study, a total of 1,084,590 bacterial sequences were analyzed from animal samples that possessed 392,328 sequences of sufficient quality. The predominant bacterial phylum detected was Firmicutes, with a relative abundance between 96.37 and 57.38% of the sequences (Table 1). The second most predominant phylum was Bacteroidetes in eight samples with a relative abundance between 23.9 and 2.4% for all samples. Fusobacteria was the second most predominant group bacteria detected in the other two samples, with a relative abundance between 22.0 and 0.0% for all samples (Table 1). The relative abundance of sequences for eight different phyla is shown in Table 1. Most bacteria detected were members of the order Clostridiales, ranging from 96.4 to 56.0% of samples analyzed (Table S2). A rarefaction curve was generated to show the completeness of the sampling, by plotting the number of observed species from the painted turtle fecal samples against the number of sequences (Supplementary Fig. S1).
The calculated Chao1 index values from our samples ranged from 25 to 83, which indicates that all of the samples have similar species richness. The Shannon–Weaver index value, calculated as a way to quantify the diversity of bacterial species in our samples, ranged from 2.922 to 4.891.
Past studies of aquatic animals have generally reported similar, but not identical, results to ours. For example, varying abundances of bacteria from the phyla Firmicutes and Bacteroidetes have been detected in the gut in both terrestrial and marine mammals [11]. In a study using cloacal swab samples from captured and rehabilitated green sea turtles (Chelonia mydas) previously living in the wild, Firmicutes were the most predominant. This was followed by Bacteroidetes and Proteobacteria [12]. In a recent study by Ahasan et al. [13], eight cloacal samples from four different green sea turtles which were undergoing rehabilitation were collected. In the samples from pre-hospitalization turtles, Proteobacteria was the most dominant followed by Firmicutes and then Bacteroidetes. Proteobacteria was again the most dominant phyla of bacteria found in samples from post-rehabilitated turtles, followed by Bacteroidetes and then Firmicutes. Interestingly, in a juvenile green sea turtle sample WP940 both a cloacal and a fecal sample were collected and the microbiota was investigated [14]. In the cloacal sample, Firmicutes was the most dominant followed by Bacterioides with a relative abundance of 61.1% and 28.9%, respectively. In the fecal sample, Proteobacteria was the most dominant followed by Bacterioides with a relative abundance of 42.5% and 34.3%, respectively [J. T. Price pers. comm.].
In all vertebrates, Firmicutes are the most widespread and common phylum detected. This is likely due to their ability to absorb nutrients and to harvest energy from material which was ingested by the host [15]. In ruminants, for example, Firmicutes are important in the breaking down of fiber and cellulose [16]. In wild-captured green turtles the high number of Firmicutes could be a reflection of healthy turtles living in a natural state [12]. Also, in green turtles, bacteria in the genera Cellulosilyticum, Peptoclostridium, and Clostridium could aid turtle digestion, facilitating the metabolism of different plant-derived polysaccharides [17, 18]. Even though painted turtles are not ruminants, our current study detected bacteria of the genera Clostridium (15.1–53.6%), Peptoclostridium (0–0.8%), and Cellulosilyticum (0–11.6%) in fecal samples, which could also aid in the digestion of complex polysaccharides.
Members of Bacteroidetes are common microbes associated with the gut. They are reported in several aquatic and terrestrial vertebrate animals including herbivorous mammals [11, 19]. The main function of Bacteroidetes is to break down carbohydrates and proteins, while also facilitating in the development of the gastrointestinal immune system [20, 21]. The members of the phylum Fusobacteria have been identified from both aquatic and terrestrial vertebrates including humans. However, the phylum is poorly studied and consists of approximately 32 species with an overall undefined phylogenetic position [11, 22].
Our study represents an important first step in elucidating the fecal bacterial communities’ in adult painted turtles. Unfortunately, other groups which are also an important part of the microbiota, such as archaea and fungi, remain poorly characterized and need to be investigated in future.
References
Ernst CH, Lovich JE (2009) Turtles of the United States and Canada, 2nd edn. Johns Hopkins University Press, Baltimore, p. 840
Mitchell JC, McAvoy BV (1990) Enteric bacteria in natural populations of freshwater turtles in Virginia. Virginia J Sci 41:233–242
Richards JM, Brown JD, Kelly TR, Fountain AL, Sleeman JM (2004) Absence of detectable Salmonella cloacal shedding in free-living reptiles on admission to the wildlife center of Virginia. J Zoo Wildl Med 35:562–563
Saelinger CA, Lawbart GA, Christian LS, Lemons CL (2006) Prevalence of Salmonella spp. in cloacal, fecal, and gastrointestinal mucosal samples from wild North American turtles. J Amer Vet Med Assoc 229:266–268
Ley RE, Lozupone CA, Hamady M, Knight R, Gordon JI (2008) Worlds within worlds: evolution of the vertebrate gut microbiota. Nat Rev Microbiol 6:776–788
McFall-Ngai M, Hadfield MG, Bosch TC, Carey HV, Domazet-Lošo T, Douglas AE, Dubilier N, Eberl G, Fukami T, Gilbert SF, Hentschel U, King N, Kjelleberg S, Knoll AH, Kremer N, Mazmanian SK, Metcalf JL, Nealson K, Pierce NE, Rawls JF, Reid A, Nishino R, Mikami K, Takahashi H, Tomonaga S, Furuse M, Hiramoto T, Aiba Y, Koga Y, Sudo N (2013) Commensal microbiota modulate murine behaviors in a strictly contamination-free environment confirmed by culture-based methods. Neurogastroenterol Motil 25:521–528
Koeth RA, Wang Z, Levison BS, Buffa JA, Org E, Sheehy BT, Britt EB, Fu X, Wu Y, Li L, Smith JD, DiDonato JA, Chen J, Li H, Wu GD, Lewis JD, Warrier M, Brown JM, Krauss RM, Tang WH, Bushman FD, Lusis AJ, Hazen SL (2013) Intestinal microbiota metabolism of l-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat Med 19:576–585
Tompkins DM, Carver S, Jones ME, Krkosek M, Skerratt LF (2015) Emerging infectious diseases of wildlife: a critical perspective. Trends Parasit 31:149–159
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJ, Holmes SP (2016) DADA2: high resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Nelson TM, Rogers TL, Brown MV (2013) The gut bacterial community of mammals from marine and terrestrial habitats. PLoS ONE 8:e83655
Ahasan MS, Waltzek TB, Huerlimann R, Ariel E (2017) Fecal bacterial communities of wild-captured and stranded green turtles (Chelonia mydas) on the great barrier reef. FEMS Microbiol Ecol 1:93
Ahasan MS, Waltzek TB, Huerlimann R, Ariel E (2018) Comparative analysis of gut bacterial communities of green turtles (Chelonia mydas) pre-hospitalization and post-rehabilitation by high-throughput sequencing of bacterial 16S rRNA gene. Microbiol Res 207:91–99
Price JT, Paladino FV, Lamont MM, Witherington BE, Bates ST, Soule T (2017) Characterization of the juvenile green turtle (Chelonia mydas) microbiome throughout an ontogenetic shift from pelagic to neritic habitats. PLoS ONE 12:e0177642
Wang W, Zheng S, Sharshov K, Sun H, Yang F, Wang X, Li L, Xiao Z (2017) Metagenomic profiling of gut microbial communities in both wild and artificially reared Bar-headed goose (Anser indicus). Microbiologyopen 6:e00429
Thoetkiattikul H, Mhuantong W, Laothanachareon T, Tangphatsornruang S, Pattarajinda V, Eurwilaichitr L, Champreda V (2013) Comparative analysis of microbial profiles in cow rumen fed with different dietary fiber by tagged 16S rRNA gene pyrosequencing. Curr Microbiol 67:130–137
Uffen RL (1997) Xylan degradation: a glimpse at microbial diversity. J Ind Microbiol Biotechnol 19:1–6
Uz I, Ogram AV (2006) Cellulolytic and fermentative guilds in eutrophic soils of the Florida Everglades. FEMS Microbiol Ecol 57:396–408
Hong PY, Wheeler E, Cann IK, Mackie RI (2011) Phylogenetic analysis of the fecal microbial community in herbivorous land and marine iguanas of the Galapagos Islands using 16S rRNA-based pyrosequencing. ISME J 5:1461–1470
Meital NO, Hadar N, Omry K (2016) Microbial changes during pregnancy, birth, and infancy. Front Microbiol 7:1031
Spence C, Wells WG, Smith CJ (2006) Characterization of the primary starch utilization operon in the obligate anaerobe Bacteroides fragilis: regulation by carbon source and oxygen. J Bacteriol 188:4663–4672
Keenan SW, Engel AS, Elsey RM (2013) The alligator gut microbiome and implications for archosaur symbioses. Sci Rep 3:2877
Acknowledgements
This research was done as part of a Provost Honors project under the leadership of Zina Haywood, Executive Vice President/Provost. We thank Karina Rebman and Jarod Lorenz for their assistance with turtle surveys. We also thank Jennifer Cumpston and Donald Zakutansky for their enthusiastic support of this research.
Funding
This project was supported by funding provided by Gateway Technical College and by the Gateway Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fugate, H.M., Kapfer, J.M. & McLaughlin, R.W. Analysis of the Microbiota in the Fecal Material of Painted Turtles (Chrysemys picta). Curr Microbiol 77, 11–14 (2020). https://doi.org/10.1007/s00284-019-01787-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01787-5