Abstract
In the present study, three strains (ChDC F213T, ChDC F251, and ChDC F267) were classified as novel species of genus Fusobacterium based on average nucleotide identity (ANI) and genome-to-genome distance (GGD) analysis and chemotaxonomic characterization. 16S rDNA sequences of strains ChDC F213T, ChDC F251, and ChDC F267 were highly similar to that of F. periodonticum ATCC 33693T (99.6, 99.4, and 99.4%, respectively). ANI and GGD values of the three isolates with F. periodonticum ATCC 33693T ranged from 92.5 to 92.6% and 47.7 to 48.2%, respectively. Considering that threshold of ANI and GGD values for bacterial species discrimination are 95–96% and 70%, respectively, these results indicate that the three isolates represent a novel Fusobacterium species. DNA G + C contents of the three isolates were 28.0 mol% each. Cellular fatty acid analysis of these strains revealed that C14:0, C16:0, and C16:1 ω6c/C16:1 ω7c were major fatty acids. Therefore, these three strains are novel species belonging to genus Fusobacterium. Strain ChDC F213T (= KCOM 1259T = KCTC 5677T = JCM 33009T) is the type strain of a novel species of genus Fusobacterium, for which a name of Fusobacterium pseudoperiodonticum sp. nov. is proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fusobacterium periodonticum is an obligate, anaerobic, non-spore-forming, nonmotile, and Gram-negative rod bacterium isolated from periodontitis lesion [13]. F. periodonticum inhabits the oral cavity and gastrointestinal tract of human [14]. Recently, average nucleotide identity (ANI) and genome-to-genome distance (GGD) analyses instead of DNA–DNA hybridization have become new gold standards for classification of bacteria at species level [1, 11]. Three strains—ChDC F213T, ChDC F251, and ChDC F267—were isolated from human oral cavity and tentatively identified as F. periodonticum by 16S rDNA sequence analysis. Strains ChDC F213T and ChDC F251 were isolated from the tongues of two male subjects (33- and 41-year old, respectively) who had gingivitis. Strain ChDC F267 was isolated from subgingival plaque of gingivitis lesion of left lower first molar of a male (45-year old). Herein, we proposed these three strains as a novel species of the genus Fusobacterium based on the polyphasic taxonomic characterization including whole-genome analysis.
Materials and Methods
Bacterial Strains and Culture Conditions
Three strains—ChDC F213T (= KCOM 1259T = KCTC 5677T = JCM 33009T), ChDC F251 (= KCOM 1261 = KCTC 5169), and ChDC F267 (= KCOM 1263 = KCTC 5171)—and F. periodonticum ATCC 33693T were obtained from the Korean Collection for Oral Microbiology (KCOM; Gwangju, Korea) or American Type Culture Collection (ATCC; Manassas, VA, USA), respectively. All strains were cultured and maintained in tryptic soy agar (TSA; BD Difco Laboratories, Franklin Lakes, NJ, USA) supplemented with 0.5% yeast extract, 0.5 mg/ml of hemin, 0.05% cysteine HCl-H2O, and 2 μg/ml of vitamin K1 (TSA-YCHVk) [3] at 37 °C in an anaerobic chamber (Model BACTRONEZ, Sheldon Manufacturing Inc., Cornelius, OR, USA) in an atmosphere of 10% H2, 5% CO2, and 85% N2.
Phylogenetic Analysis
16S rDNA sequences of these three strains were sequenced in the course of genome sequencing. 16S rDNA sequences of type strains of Fusobacterium spp. were obtained from GenBank. 16S rRNA accession numbers of all these strains are listed in Fig. 1. Multiple sequences were aligned using CLUSTAL W algorithm. Sequence similarities were calculated using MegAlign program (DNAStar LasergeneTM 8.0, DNAStar Inc., Madison, WI, USA) [4]. Evolutionary distance was calculated in accordance with the Kimura two-parameter model [7]. Phylogenetic trees were constructed with the neighbor-joining method [12] using MEGA 6.06 software [15]. The stability of the phylogenetic tree was assessed by bootstrap analysis [5] with 1000 replicates.
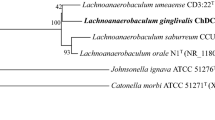
Neighbor-joining phylogenetic tree based on 16S ribosomal RNA genes. GenBank accession numbers of 16S rDNA of each strain are written in parenthesis. Stability of phylogenetic trees was assessed by bootstrap analysis of 1000 replicates using MEGA version 6.06 [15]. Bar indicates 0.002 changes per nucleotide position
Genome Sequence
Genomic DNAs of these three strains were prepared as described previously [3]. Concentration and quality of these bacterial genomic DNAs were determined using an Epoch™ Microplate Spectrophotometer (BioTek Instruments Inc., Winooski, VT, USA) at wavelengths of 260 and 280 nm [3].
Genomic DNAs of ChDC F213T, ChDC F251, and ChDC F267 were sequenced using a PacBio RSII platform by Macrogen Inc. (Seoul, Korea). DNA libraries (20-kb) were constructed in accordance with the manufacturer’s protocol and sequenced using single-molecule real-time sequencing with P6 DNA polymerase and C4 chemistry [6]. From genomic DNAs of ChDC F213T, ChDC F251, and ChDC F267, 1,175,912,373 bp (487.3 × coverage), 664,376,116 bp (280.0 × coverage), and 1,149,364,741 bp (433.5 × coverage) were generated, respectively. De novo assembly was performed using RS HGAP Assembly 3.0 [2] with default option. Genome annotation was conducted using the NCBI Prokaryotic Genome Annotation Pipeline through the NCBI Genome Submission Portal (https://submit.ncbi.nlm.nih.gov/subs/genome) [16]. GenBank accession numbers for genomes of the three strains—ChDC F213T, ChDC F251, and ChDC F267—are listed in Table 1.
ANI and GGD Analyses
ANI and GGD analyses were conducted using the calculator provided by ChunLab (Seoul, Korea) (http://www.ezbiocloud.net/tools/ani) [9] and the German Collection of Microorganisms and Cell Cultures (DSMZ; Braunschweig, Germany) (http://ggdc.dsmz.de) [10], respectively, to determine the genome relatedness. Whole-genome sequences of these strains and type strains of Fusobacterium spp. were downloaded from GenBank database (https://www.ncbi.nlm.nih.gov/genome/?term=fusobacterium+periodonticum). GenBank accession numbers of these strains are listed in Table 1.
Morphological and Physiological Characterization, Biochemical Analysis, and Chemotaxonomic Characteristics
Cell shape and size of the three isolates were determined by scanning electron microscopy (SEM) as described previously [4]. Growth at different temperatures (25–45 °C at intervals of 5 °C and 37 °C) was determined using TSB-YCHVk medium [4] after incubation for 3 days. Growth at various pH conditions (5–10 at intervals of 0.5) was assessed in TSB-YCHVk medium [4] after culturing at 37°C for 3 days. Growth at various NaCl concentrations was assessed in TP-YCHVk medium [4] containing 0°C, 1°C, 2°C, or 3% (w/v) NaCl (pH 7) after culturing at 37 °C for 3 days.
API ZYM and API 20A test strips (bioMerieux, Marcy-l’Etoile, France) were used to analyze enzyme activities, biochemical traits, and sugar fermentation patterns of these bacterial strains in accordance with the manufacturer’s instructions. Cellular fatty acid compositions of these bacterial strains were determined using MIDI/Hewlett Packard Microbial Identification System (MIDI, Microbial ID, Newark, DE, USA) in accordance with the manufacturer’s instructions by the Korean Culture Center of Microorganisms (Seoul, Korea). Fatty acids were analyzed using a gas chromatograph (Model 6890N and Auto-sampler 7683; Agilent, Santa Clara, CA, USA) and identified using SherlockTM Microbial Identification System (version 6.3).
Results and Discussion
Phylogenetic analysis revealed that 16S rDNAs of all these strains belonged to the same cluster (C1). 16S rDNA sequences of strains ChDC F213T, ChDC F251, and ChDC F267 were most closely related to that of F. periodonticum ATCC 33693T (identities: 99.6, 99.5, and 99.4%, respectively, Supplementary Table S1). They were clearly separated from F. periodonticum ATCC 33693T with bootstrap value of 99% by neighbor-joining, maximum likelihood, and minimum evolution methods (Fig. 1 and Supplementary Fig. S1). Average percent similarity of 16S rDNAs among strains belonging to cluster C1 was 99.7% (range 99.3% to 100%) (Supplementary Table S1). The genomic DNA sequences of 14 strains belonging to cluster C1 were deposited in GenBank as F. periodonticum (https://www.ncbi.nlm.nih.gov/genome/?term=fusobacterium periodonticum). 16S rDNA sequences of these strains closely related to that of F. periodonticum ATCC 33693T (average 99.6%; range 99.4%, and 99.4%, respectively) and F. hwasookii KCOM 1249T (average 99.6%; range 99.4–99.8%) and 99.0% (range 98.8–99.2%) (Supplementary Table S2). These data indicate that these strains in cluster C1 belong to genus Fusobacterium.
Genome sizes of strains ChDC F213T, ChDC F251, and ChDC F267 were 2,413,021 bp, 2,372,833 bp, and 2,651,098 bp, respectively. DNA G + C contents of these three strains were 28.0 mol % each, similar to the G + C content of F. periodonticum ATCC 33693T (27.8 mol%) [3]. ANI value between F. periodonticum ATCC 33693T and strain ChDC F213T, ChDC F251, or ChDC F267 were 92.7, 92.6, or 92.5%, respectively (Table 2). GGD value between F. periodonticum ATCC 33693T and strain ChDC F213T, ChDC F251, or ChDC F267 was 48.2, 48.2, or 47.7%, respectively (Table 2). Considering that threshold values of ANI and GGD for bacterial species discrimination are 95–96% and 70%, respectively [9, 10], these results indicate that strains ChDC F213T, ChDC F251, and ChDC F267 represent a novel Fusobacterium species.
ANI and GGD values of strains in cluster C1, except strains ChDC F213T, ChDC F251, ChDC F267, and 1_1_41FAA, compared with F. peridodonticum ATCC 33693T were from 90.6% to 93.0% and 47.7% to 48.8%, respectively (Table 2). ANI and GGD values of strains in cluster C1 except for strains ChDC F213T, ChDC F251, ChDC F267, and 1_1_41FAA, compared with strain ChDC F213T were from 95.3% to 96.9% and 61.8% to 72.5%, respectively (Table 2). ANI and GGD values between strain 1_1_41FAA and ChDC F213T were 94.7% and 58.0%, respectively (Table 2). This ANI value is almost borderline ANI value to discriminate bacteria at species level. The phylogenetic tree based on 16S rDNA showed that strain 1_1_41FAA and ChDC F213T had the same cluster (Fig. 1 and Supplementary Fig. S1). Percent similarity of 16S rDNA between strain 1_1_41FAA and strain ChDC F213T (99.9%) was higher than that between strain 1_1_41FAA and F. periodonticum ATCC 33693T (99.6%) (Supplementary Tables S1, S2). Based on these data, strain 1_1_41FAA might belong to the same species as strain ChDC F213T, but not F. periodonticum ATCC 33693T. ANI and GGD values of 14 strains in cluster C1 and type strains of F. nucleatum, F. polymorphum, F. vincentii, F. animalis, and F. hwasookii that were closely related to strain ChDC F213T by 16S rDNA sequence analysis were below 93.0% and 31.8%, respectively (Supplementary Tables S3, S4). These results indicate that these 14 strains in cluster C1 are members of a novel Fusobacterium spp.
In the API ZYM panel, tests for acid phosphatase and naphthol-AS-BI-phosphohydrolase were positive for the three strains (ChDC F213T, ChDC F251, and ChDC F267), but negative for F. periodonticum ATCC 33693T (Table 3). In a previous study, tests for acid phosphatase were negative for four subspecies of Fusobacterium nucleatum (now reclassified as four novel species [8]) and Fusobacterium hwasookii [3]. Tests for alkaline phosphatase and leucine arylamidase were positive for strains ChDC F213T and ChDC F251 (Table 3). Esterase lipase (C8) was positive in strains ChDC F213T and ChDC F267 (Table 3). The remaining 13 tests in the API ZYM panel were negative for the three strains. An API 20A test for indole production was positive (Table 3). Strains ChDC F213T and ChDC F267 fermented glucose (Table 3). Strains ChDC F213T and ChDC F267 hydrolyzed gelatin and esculin, respectively (Table 3). The remaining 17 tests, including test for catalase, were negative for the three strains (Supplementary Table S5). Biochemical test results for these three strains were similar to those for F. periodonticum ATCC 33693T.
Morphological characteristics and optimal growth conditions of the three strains are summarized in Supplementary Table S6.
Based on molecular, chemical, and phenotypic evidence presented in the present study, we propose that these three strains—ChDC F213T, ChDC F251, and ChDC F267—should be assigned to a novel species of Fusobacterium, for which a name of Fusobacterium pseudoperiodonticum sp. nov. is proposed.
Description of Fusobacterium pseudoperiodonticum sp. nov.
Fusobacterium pseudoperiodonticum [Gr. adj. pseudês, false; N.L. n. periodonticum, a bacterial specific epithet; N.L. n. pseudoperiodonticum, a false (Fusobacterium) periodonticum].
Fusobacterium pseudoperiodonticum is a Gram-negative, anaerobic, and fusiform-shaped bacterium with variable size. Cell size was ranged from 0.3–0.4 × 2.2–106.5 μm. Colonies were pigmented in grayish brown and spread to a diameter of approximately 0.7–1.0 mm after growing on TSA-YCHVk agar at 37°C for 2 days. Growth occurred in the range of 30–37 °C (optimum 35–37 °C). The optimum pH for growth for these strains was 7.0–7.5. Acid phosphatase and naphthol-AS-BI-phosphohydrolase were positive. Indole production test was positive. Cellular fatty acids were mainly composed of C14:0, C16:0, and C16:1 ω6c/C16:1 ω7c (Table 4). G + C contents of all strains were 28.0 mol%.
The type strain of Fusobacterium pseudoperiodonticum is ChDC F213T (= KCOM 1259T = KCTC 5677T = JCM 33009T). It was isolated from the tongue of a Korean. It can hydrolyze gelatin. This strain produces esterase (C4). The DNA G + C content is 28.0 mol%.
References
Auch AF, von Jan M, Klenk HP, Göker M (2010) Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci 2:117–134. https://doi.org/10.4056/sigs.531120
Chin CS, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10:563–569. https://doi.org/10.1038/nmeth.2474
Cho E, Park SN, Lim YK, Shin Y, Paek J, Hwang CH, Chang YH, Kook JK (2015) Fusobacterium hwasookii sp. nov., isolated from a human periodontitis lesion. Curr Microbiol 70:169–175. https://doi.org/10.1007/s00284-014-0692-7
Cho E, Park SN, Shin Y, Lim YK, Paek J, Kim HK, Hwang CH, Jo E, Jin D, Chang YH, Kook JK (2015) Peptoniphilus mikwangii sp. nov., isolated from a clinical specimen of human origin. Curr Microbiol 70:260–266. https://doi.org/10.1007/s00284-014-0712-7
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Jo E, Park SN, Lim YK, Paek J, Shin Y, Kim H, Kim SH, Shin JH, Chang YH, Kook JK (2018) Capnocytophaga endodontalis sp. nov., isolated from a human refractory periapical abscess. Curr Microbiol 75:420–425. https://doi.org/10.1007/s00284-017-1397-5
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kook JK, Park SN, Lim YK, Cho E, Jo E, Roh H, Shin Y, Paek J, Kim HS, Kim H, Shin JH, Chang YH (2017) Genome-based reclassification of Fusobacterium nucleatum subspecies at the species level. Curr Microbiol 74:1137–1147. https://doi.org/10.1007/s00284-017-1296-9
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinf 14:60. https://doi.org/10.1186/1471-2105-14-60
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Slots J, Potts TV, Mashimo PA (1983) Fusobacterium periodonticum, a new species from the human oral cavity. J Dent Res 62:960–963. https://doi.org/10.1177/00220345830620090901
Strauss J, White A, Ambrose C, McDonald J, Allen-Vercoe E (2008) Phenotypic and genotypic analyses of clinical Fusobacterium nucleatum and Fusobacterium periodonticum isolates from the human gut. Anaerobe 14:301–309. https://doi.org/10.1016/j.anaerobe.2008.12.003
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP, Zaslavsky L, Lomsadze A, Pruitt KD, Borodovsky M, Ostell J (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Acknowledgements
This research was supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science and ICT (No. 2017M3A9B8065844) and in part by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (2017R1A2B4004894).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
284_2019_1675_MOESM2_ESM.pptx
Maximum likelihood (A) and the minimum evolution (B) phylogenetic tree based on 16S ribosomal genes of strains. GenBank accession number of 16S rDNA of each strain was written in parenthesis. Stability of phylogenetic trees was assessed by a bootstrap analysis of 1000 replicates using MEGA version 6.06 [15]. Bars indicate 0.002 (A) or 0.002 (B) changes per nucleotide position. Supplementary material 2 (PPTX 83 kb)
Rights and permissions
About this article
Cite this article
Park, SN., Lim, Y.K., Shin, J.H. et al. Fusobacterium pseudoperiodonticum sp. nov., Isolated from the Human Oral Cavity. Curr Microbiol 76, 659–665 (2019). https://doi.org/10.1007/s00284-019-01675-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01675-y