Abstract
Endophytic fungi are characterized as microorganisms found within internal tissues of living plants without any immediate, overtly negative effects. The present study was carried out to isolate, taxonomically characterize and determine the spatiotemporal distribution of endophytic fungi associated with leaf, stem, trunk, and root of mandarin (Citrus reticulata cv. Siyahoo). To do so, the sampling program was done seasonally in four geographically isolated mandarin growing areas of Hormozgan province of Iran, including Siyahoo, Ahmadi, Sikhoran, and Roudan. In total, 702 fungal isolates were obtained from leaf, stem, trunk, and root of healthy mandarin trees divided into 26 distinct morphotypes based on morphological characteristics. The morphotypes were taxonomically characterized through phylogenetic analysis of the ITS1-5.8S-ITS4 rDNA region sequences. Accordingly, 10 different fungal orders from 5 fungal classes were identified, i.e., Saccharomycetes (Saccharomycetales), Eurotiomycetes (Eurotiales), Dothideomycetes (Capnodiales, Pleosporales, Dothideales), and Sordariomycetes (Diaporthales, Hypocreales, Microascales, Togniniales), all from Ascomycota, which represented 97.2% and Ustilaginomycetes (Ustilaginales) from Basidiomycota which represented 2.8% of the isolates. The Aureobasidium pullulans, Penicillium citrinum, and Dothideomycetes sp. were the most frequent isolates. The trunk and leaf showed the highest and lowest total colonization frequency and species richness of endophytic fungi, respectively, in all sampling periods. The results showed that the colonization frequency of endophytes in Hormozgan province was higher in autumn than that in spring, winter, and summer. The trunk showed the maximum diversity of endophytes over all seasons. The Shannon–Wiener (H′) and Simpson indices had significant correlation with sampling cites and tissue type and the maximum value of Shannon and Simpson indices (H′ = 3.05 and 1 − D = 0.94) was found in the specimens collected from Siyahoo. In conclusion, the three factors (season, location, and tissue type) all in together could determine fungal endophyte composition of C. reticulata.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It is estimated that each of nearly 270,000 plant species existing on the earth is in association with one or more endophytes [42, 43]. Fungal endophytes, as an important group of organisms existing within plant tissues [9, 47], are in interaction with their hosts without any visible disease symptoms [7, 10, 53]. The symbiotic relationships and ecological functions of the endophytes with the host plants can be highly variable [18, 38, 54]. Some endophytic fungi can produce metabolites similar to that produced by the hosts, and others help the hosts by producing natural compounds lacking in the host plants. Suitability of such compounds to be used in medicine, agriculture, and industry has been documented [1, 42, 44]. Furthermore, the role of endophytes in mutualistic relations, decreased herbivory, and increased abiotic stress resistance has been shown [42]. Endophytes are able to allocate ecological niches in the host plants which may be occupied by plant pathogens, so they can augment disease resistance indirectly and increase growth of their host [15, 31]. The ability of endophytes in suppressing pests and diseases damage has been shown by various researchers [see; 30, 48]. On the other hand, since some fungal endophyte species have been reported frequently as pathogens on the hosts, it is likely that they may be pathogens in a latent phase of their life cycle [4, 41], therefore, detailed characterization of fungal endophytes communities and their interactions is crucial to understand fungal diseases of host plants and is a prerequisite for best management practices [10].
Owing to the unknown role of endophytic fungi existing in healthy Citrus tissues and since these endophytes may be responsible for different functions [11], it is necessary that the type of host–endophyte association is elucidated in each Citrus species. Furthermore, it is important to determine whether the isolated endophytes have the potential to be cultured or not, because it can facilitate the subsequent studies, e.g., manipulation of endophytic fungi activities and extraction of endophyte-based natural products [50].
Siyahoo mandarin (Citrus reticulata Blanco cv. Siyahoo) is a popular cultivar of mandarin in Iran due to its special taste, texture, and flavor. Here, we aimed to firstly, isolate and taxonomically identify the fungal endophytes associating leaf, stem, trunk, and root of C. reticulata; and secondly to determine the spatiotemporal impacts on fungal endophytes diversity. Since no published documents are available dealing with the fungal endophytes diversity in mandarin, this study can provide a platform for other researchers working on efficiency of the endophytes against biotic and abiotic stresses.
Materials and Methods
Sampling Sites and Mandarin Host Species
During May 2016 to February of 2017, a total of 86 trees of Citrus reticulata from four geographically isolated mandarin growing areas of Hormozgan province (Iran) were explored individually to isolate and taxonomically identify endophytic fungi (Table 1). Regarding probable seasonal dynamism of endophytes and to reach the maximum diversity, the sampling was repeated seasonally from May 2016 to February 2017. To do so, the fresh tissues of leaf, stem, trunk, and root of mature (5–8 years old) and healthy mandarin trees were collected and processed for endophyte isolation separately.
The samples were rinsed gently in running water to remove dust and debris and then cut into 1 × 0.5 cm pieces with/without midribs. The stem, trunk, and root samples were cut into 0.5-1.0 cm pieces. Each sample was disinfected with 75% ethanol for 1 min followed by immersion in Sodium hypochlorite (NaOCl 3% for 3–5 min, depending on the type of samples, i.e., 3 min for leaves and 5 min for stems, trunk, and roots) and then once again in 75% ethanol for 30 s.
About 3–4 segments of each vegetative organ were placed onto Potato Dextrose Agar (PDA) containing Petri plates. Accordingly, a total of 2187 plant segments from 86 mandarin plants were investigated. The petri plates were sealed with parafilm and incubated at 27 ± 2 °C for 4–6 weeks. Most fungal growth was initiated within 10 days after inoculation. The fungi that grew out from the segments were periodically isolated and identified by transferring the hyphal tips to fresh PDA plates.
Endophyte Identification
The endophytes were identified either morphologically based on characteristics of the fungal culture, or genetically through analysis of the internal transcribed spacer regions of nuclear ribosomal DNA (ITS1-5.8S-ITS4 rDNA sequence). Fungal growth was obtained by placing the fungal endophytes onto the PDA culture medium. The plates were checked continuously for spore formation. For DNA extraction, the colony of each fungal isolate was grown in 150 ml Erlenmeyer flasks containing 20 ml Potato Dextrose Broth (PDB; Merck, Germany) at 28 °C, at 90 rpm. After 15 days, genomic DNA of each isolate was extracted using SDS-CTAB method [57] and subjected to Polymerase Chain Reaction (PCR) to amplify ITS1-5.8S-ITS4 rDNA region using following universal primers, ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) [51]. Each 50 µl reaction mixture included 4 µl of DNA, 25 µl of Taq DNA Polymerase (amplicon), 2 µl of primers (Pishgam, Iran), and 17 µl ddH2O. The PCR assay was performed in a Techne TC-572 thermocycler (Eppendorf, Hamburg, Germany) programmed for 94 °C for 5 min, 35 cycles of 94 °C for 45 s, 53 °C for 30 s, and 72 °C for 1 min, and 72 °C for 5 min. PCR products were subjected to electrophoresis on 1% agarose gel, stained with SYBR Green (SYBR safe CinnaGen, Tehran, Iran). All positive PCR products were sequenced directly by Macrogen Sequencing Service (Seoul, South Korea). The aligned and edited sequences were deposited in the GenBank database under accession numbers MH260421-53 (Table 2). The identifications were also confirmed by microscopic studies on morphological characteristics of the fungal colonies. The fungal isolates were archived as living vouchers at 4 °C, and are available upon request.
Statistical Data Analysis
The colonization frequency (CF) was calculated as the total number of segments colonized by endophytic fungi divided by the total number of incubated segments. The relative species frequency (RF) was calculated as the number of isolates of one species divided by the total number of isolates [52]. The Shannon–Weaver diversity index (H′) [39] was used to show diversity of the endophytic fungal species and was calculated as: H′ = − Ʃpi lnpi. Shannon evenness (E) was calculated as H′/Hmax, where Hmax = ln(S), S is the total number of taxa in the subsample. Simpson’s diversity (D = 1 − Ʃpi2, where pi is the proportion of isolates assigned to the ith taxa, that is Pi = ni/N) [40] was calculated to compare the species richness. A Simpson diversity index close to 1 means that the sample is highly diverse. Diversity parameters was calculated for each location, season, and tissue type using PAST software (http://folk.uio.no/ohammer/past/).
The base substitution model was implemented using MrModeltest2 [29]. To estimate invariant sites, a general time reversible model based on Akaike criterion was included among-site rate heterogeneity (GTR + G + I), in phylogenetic analyses. Phylogenetic relationships and the related tree were constructed using MrBayes v3.1.2 [35]. After discarding burn-in (25% of the samples) samples and evaluating convergence, the remaining samples, were kept for further analysis. To determine the equilibrium distribution and estimation of the Bayesian posterior probabilities of clades, the Markov chain Monte Carlo (MCMC) method within a Bayesian framework was run for 10 million generations [22] using the 50% majority rule. The Bayesian posterior probability values higher than 0.50 are presented on appropriate clades. The phylogenetic was inferred and re-drawn using Dendroscope V.3.2.8 (https://www-ab.informatik.uni-tuebingen.de/software/dendroscope) and CorelDRAW version X7, respectively.
Results
Species Diversity
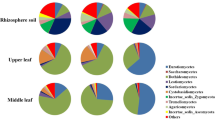
During 2016–2017, a total of 702 endophytic fungal isolates were recovered from 2187 plant tissue segments of C. reticulata in Hormozgan province of Iran (Fig. 1). Of these, 115, 155, 231, and 201 isolates were recovered from leaves, stems, trunks, and roots, respectively (Table 2). Overall, endophyte colonization frequency in C. reticulata tissues was 32.05%. The maximum colonization rate occurred in trunk (47.43%), followed by root (33.78), stems (26.4%), and leaves (22.2%). Sampling Season influenced fungal colonization rate; the maximum colonization rate was observed in autumn followed by spring, summer, and winter. In all seasons, the CF index in leaves was significantly lower than that of the other plant tissues.
The relationship between the number of isolated strains (Table 3) and sampling sites was also investigated. Results show that Siyahoo and Ahmadi yielded the highest numbers of isolates per sampling site, regarding number of strains (276 and 252, respectively) (Table 3). Also, when considering the number of sampling campaigns in each location, Siyahoo was more diverse than the other sites, because from 736 investigated tissues 276 isolates were obtained from samples of Siyahoo. The maximum colonization rate occurred in Siyahoo (37.5%) followed by Ahmadi (32. 8), Sikhoran (25.05%), and Roudan (26.25%).
Phylogeny
A total of 702 endophytic fungal isolates were recovered seasonally from the asymptomatic leaf, stem, trunk, and root tissues of spatially separated mandarin orchards. Prior to molecular identification, the isolates were morphologically grouped and divided into 26 distinct taxa. Then a single isolate of each unidentified morphotype was subjected to molecular identification and phylogenetic analysis. The accession numbers of all 26 sequenced isolates besides other supplementary information have been depicted in Table 4.
The phylogenetic tree inferred from ITS1-5.8S-ITS2 gene sequences is shown in Fig. 2. All endophytic taxa were bunched in four Ascomycetous (Dothideomycetes, Sordariomycetes, Saccharomycetes, Eurotiomycetes) and one Basidiomycetous (Ustilaginomycetes) classes. A total of 97.2% endophytic fungi belonged to the phylum Ascomycota and 2.28% to Basidiomycota (Table 4; Fig. 2). According to the percentage of species composition (Table 4), the richest and most abundant class was Dothideomycetes represented by the orders Pleosporales (6 species), Dothideales (4 species), and Capnodiales (2 species). Class Sordariomycetes represented by the orders Hypocreales (3 species), Togniniales (1 species), Microascales (1 species), and Diaporthales (1 species) was ranked second in terms of species composition and third for fungal frequency. Eurotiales (5 species) corresponded to Class Eurotiomycetes in which the identified species had a high frequency. Saccharomycetes (Ascomycota) and Ustilaginomycetes (Basidiomycota) represented by the orders Saccharomycetales and Ustilaginales (each with 1 species), respectively, had the least richness. The dendrogram created using ITS sequences of endophytic taxa and reference taxa retrieved from NCBI database shows that the endophyte assemblages of C. reticulata included representative taxa of the Ascomycota and Basidiomycota (Fig. 2). All The members of Ascomycota and Basidiomycota formed 18 different clades within the dendrogram.
Endophyte Communities in Leaf, Stem, Trunk, and Root Tissues
Seven fungal species were found repeatedly in all four tissue types. The maximum endophyte diversity was observed in the trunk (22 taxa) in which the most frequent isolates (from high to low) were Dothideomycetes sp. (26 isolates), Aureobasidium pullulans (19 isolates), and Aspergillus pallidofulvus (15 isolates). The leaves harbored 115 fungal isolates belonged to 15 taxonomically different taxa (Table 5). The most frequent isolates were Penicillium citrinum (16 isolate), Cladosporium cladosporioides (13 isolate), and Dothideomycetes sp. (13 isolate). A total of 155 fungal isolates belonged to 18 taxonomically different taxa were found in the stem (Table 5). The genus Aureobasidium was the most prevalent endophyte in the stem. The root endophyte community consisted of 202 fungal isolates which belonged to 16 taxonomically different taxa (Table 5). The genus Fusarium (82 isolates) and Alternaria (74 isolates) were the most repeated endophytes of the root.
Among the identified genera isolated from different tissues, genus Aureobasidium, Fusarium, and Alternaria had the maximum number of colonies. In contrast, Neosetophoma sp. and Pseudozyma flocculosa with the minimum number of colonies (4 and 8 colonies, respectively) were observed just in the stem samples.
Regional Endophytes Communities
Four sampling regions were different in terms of the endophytes communities. The maximum number of isolates were recovered from samples of Siyahoo (276 isolates, 24 taxa) followed by Ahmadi (253 isolates, 23 taxa), Sikhoran (111 isolates, 21 taxa), and Roudan (62 isolates, 14 taxa) (Table 5). Also, Siyahoo showed the highest percent of colonization frequency (37.5%), followed by Ahmadi, Sikhoran, and Roudan with 32.8%, 26.25.5% ,and 25.05%, respectively (Table 6). At both Siyahoo and Sikhoran, Aureobasidium pullulans was the most prevalent isolate (24 and 15 isolates, respectively) (Table 5). The endophyte Pseudozyma flocculosa was observed only in Roudan.
Seasonal Diversity of the Endophytes
Season effect was prominent with the maximum number of isolates occurring in autumn (253 isolates, 25 taxa), followed by winter (184 isolates, 24 taxa), and the minimum in summer (129 isolates, 19 taxa) (Table 5). Colonization frequency (%) value varied with season with the highest in autumn (41.13%), followed by spring (33.41%), winter (27.6%), and summer (25.8%) (Table 5). At autumn, the majority of the isolates were Dothideomycetes sp. (31 isolates), followed by Aureobasidium pullulans (26 isolates), and Penicillium citrinum (22 isolates). In summer, however, Cladosporium, Fusarium, and Alternaria exhibited the highest frequencies, as 30 isolates of Cladosporium, 30 isolates of Fusarium, and 24 isolates of Alternaria were isolated. Neosetophoma sp. was observed only in autumn season.
Aureobasidium iranianum and Aureobasidium melanogenum were observed only in autumn and winter (Table 5).
The diversity of the endophytic community isolated from tissues and sampling sites was compared using indices of α-diversity (Shannon–Wiener index and Simpson’s diversity index) and their components, i.e., species richness and evenness (Table 6). The concentration of dominance or Simpson’s dominance of endophytic fungi was almost identical in the leaves (0.09143) and stem tissues (0.09169) (Table 6). Simpson’s diversity indices of 0.942, in trunk tissues, indicate a high diversity of endophytes harbored by the host plant in this organ. Shannon–Wiener diversity index were higher in fungal endophytes of the trunk tissues (2.96) than in the other tissues (Table 6). Higher species richness of the endophytic fungi colonization was observed in the trunk (22) compared with the other organs. However, species evenness was higher in tissues of the roots. Dominance showed an inverse relationship with diversity indices, i.e., maximum to leaf (0.085) followed by stem (0.083) and least to trunk (0.057) (Table 6).
By comparing four sampling sites, we found the highest value of Shannon and Simpson indices in SI and AH followed by SK (2.81 and 0.928) and RD (2.749 and 0.909) (Table 6). However, the Dominance index was higher in RD (Table 6).
In comparison, among the four different seasons, maximum Shannon and Simpson indices were found in samples from autumn (3.029 and 0.9445), followed by winter (2.99 and 0.9396). In addition, the Dominance index was higher at summer (Table 6).
Discussion
It is estimated that 70,000–80,000 species of fungi exist on the planet [46]. These species may be advantageous for plant in coping with physiological disturbances, or tolerating environmental changes [43, 44]. Due to the possible existence of different morpho/biotypes of fungi within a single fungal species, traditional morphological and biochemical methods are unable to differentiate various morpho/biotypes of fungi [28]. By contrast, DNA analyzing method is shown to be objective, reproducible, and a rapid approach for identification, especially in non-sporulating endophytes [23, 55]. The ITS1-5.8S-ITS4 is an extremely conserved region in fungi and able to differentiate higher taxonomic levels, whereas ITS regions are highly variable and can be used for analysis of lower taxonomic ones [45]. Thus, herein, we used ITS1-5.8S-ITS4 rDNA region sequences for identification of endophytic fungi in mandarin trees.
Our results indicated that the majority of the recovered endophytic fungi (97.2%) in C. reticulata belonged to the Ascomycota, which seems to be a general characteristic of the endophytic mycobiota of other woody plants [2, 27, 32], as well as the Citrus trees [10]. A total of 2.8% of the isolates belonged to Basidiomycota in our study. Indeed, the introduced endophytic fungi belonged to 10 different fungal orders from 5 fungal classes, i.e., Saccharomycetes (Saccharomycetales), Eurotiomycetes (Eurotiales), Dothideomycetes (Capnodiales, Pleosporales, Dothideales), and Sordariomycetes (Diaporthales, Hypocreales, Microascales, Togniniales), all from Ascomycota, and Ustilaginomycetes (Ustilaginales) from Basidiomycota. Endophytic dominance of Dothideomycetes and Sordariomycetes in Cupressaceae, Sordariomycetes in Fagaceae, and Leotiomycetes in Pinaceae appears to be coevolutionary phenomena [3, 42]. Here, the endophytic community was dominated by Dothideomycetes followed by Sordariomycetes, which is similar to a pattern seen in Citrus limon as well [10]. Pleosporales and Eurotiales both had the most endophytic species in association with mandarin. Members of these orders in association with Citrus plants have been mentioned by Douanla-Meli et al. [10] and Duran et al. [11]. It seems that these fungi form an important, intimate, and long-lasting relation with C. reticulata. These data may indicate coevolution of those fungi and Citrus species.
Considering the fungal species, the endophytic community of C. reticulata comprised taxa belonging to the genera Alternaria, Penicillium, Phomopsis, and Cladosporium which are previously reported as endophytes of Citrus trees [5, 10, 11, 14]. Other genera that were found in the present study and have not been commonly isolated included Pseudozyma, Talaromyces, Meyerozyma, Phaeoacremonium, Aureobasidium, Myrothecium, Neosetophoma, Scedosporium, Dothideomycetes, Didymella, and Ascochyta. However, the taxa Aureobasidium pullulans, Penicillium citrinum, Dothideomycetes sp. were the most frequent fungi in the present study. P. citrinum has been shown to be a common fungus isolated from different environmental conditions, ranging from permafrost sediments to agricultural fields and forest soils [13, 44]. P. citrinum is a well-known species due to producing mycotoxin citrinin, cellulose-digesting enzymes like cellulase, endoglucanase, and xylulase [12, 49] and gibberellins [17, 21].
Aureobasidium pullulans has been used as a microbial antagonist against a diverse array of grapevine pathogens, including postharvest fungi [30, 36, 37] due its competition for nutrients and space, production of pectolytic enzymes, polysaccharides, or antimicrobial metabolites [13, 26]. Dothideomycetes sp. is also capable of producing secondary metabolites and a large amount of 2-hydroxymethyl-3-methyl-cyclopent-2-enone, a useful scaffold for organic synthesis [6].
The genus Fusarium represented the most abundant endophytic fungi recovered from roots of C. reticulata nearly in all sampling sites. The species of Fusarium are among the most frequently isolated endophytes in tropical plants [48] and able to descend subsequent invasions made by aggressive fungi through niche partitioning [25].
In this study, it is found that some endophytic species were restricted only to one vegetative part. For instance, Scedosporium apiospermum and Pseudozyma flocculosa were isolated only from trunk and branches, respectively. Also, Phaeoacremonium parasiticum and Fusarium solani were limited only to plant roots. The tissue-dependent specialization in host–endophyte relations is not unusual and has been evidenced in other plants. It may be stemming from high affinity of endophytes to establish within a specific chemistry or texture of different host tissues [33, 34].
According to our findings, the maximum diversity of endophytes occurred in the trunk tissue. The root, stem, and leaf were ranked the next (from high to low). Low colonization frequency (%) in mandarin leaves is also observed in leaves of many other tropical plants [9, 24], as well as the other Citrus species [10]. Possibly, long-term exposure of the trunk can provide an appropriate opportunity for association of endophytic fungi. In our study, the total colonization frequency and species richness of endophytic fungi in stems were slightly higher than the leaves which is in line with former studies [16, 46]. Shorter life time of leaves compared with the other vegetative organs may explain the low colonization frequency and species richness of the endophytes in this organ. Furthermore, the structure and substrates of the stem can influence infection of the endophytic fungi and increase their colonization frequency and species richness [15, 34, 46].
We also found difference in endophyte colonization among various sampling sites. For instance, the endophyte colonization in Roudan was lower than Siyahoo and Sikhoran. The ecological and environmental conditions such as lower annual rainfall and comparatively low annual temperature may affect colonization of host tissues by endophytes. In particular, sampling from plants in their biogeographic areas of origin would reveal the ways in which introduction to novel environments changes the fungal associations with which economically important plants species associate. It is reported that cupressaceous trees cultivated in non-native areas, maintained a lower diversity of fungal endophyte than the native species [20]. Siyahoo region is attributed with tropical climate accompanied with heavy and comparatively long spanned annual rain fall. Thus, maximal diversity index and species richness at Siyahoo is in concordance with the favorable conditions found there for fungal growth and dispersion. The high dominance and low richness in the fungal endophyte community of C. reticulata in Roudan can be attributed to the extremely hard environment conditions, such as drought. Other reason could be the fact that mandarin trees in this sampling site are subjected to excessive fungicide sprays annually, that may decrease endophyte colonization. We also observed location-specific distribution of certain endophytes at least at species level. Neosetophoma sp. was specific to Siyahoo, likewise Pseudozyma flocculosa was exclusively isolated from Roudan whereas Aureobasidium iranianum and Myrothecium sp. were restricted to Siyahoo and Ahmadi. Space limited distribution of these taxa indicates spatial structuring of endophytic communities.
Fungal endophytes colonization was also influenced by season in our study. We found high colonization frequency of most endophytic species in autumn followed by winter and spring. The season of sampling affected either colonization frequency and species richness or type of taxa throughout the sampling season. In summer, Cladosporium, Fusarium, and Alternaria exhibited the highest frequencies. Aureobasidium iranianum and Aureobasidium melanogenum were observed only in autumn and winter. Further, increase in frequencies of Penicillium citrinum and Dothideomycetes sp, in autumn indicates seasonal effect on fungal endophytic communities. Environmental factors, such as temperature, humidity, and ecological niches may affect endophyte variation and have a determinant role in spread and germination success of endophytic fungal spores [39]. This seasonal variation in the fungal endophytic communities is in accordance with former reports [16, 27]. Greater species richness of mandarin trees in autumn compared to spring could be due to higher rainfall. Rain splashes help in release of inoculum materials, and high humidity and low temperature help fungal spore germination and reproduction, causing high infection rate and fungal establishment in autumn and winter seasons. There is a discrepancy regarding the effect of seasons on colonization frequency. For instance, it is reported the highest endophyte colonization frequency in needle of Pinus tabulaeformis occurred in spring [16]. Findings of Collado et al. [8] in Quercus ilex also confirmed the highest species richness of endophytes in spring. In contrast, Helander et al. [19] showed that season of sampling has no effect on colonization frequency of endophytes in old needles of Scots pine, whereas in young needles the colonization frequency was increased in summer.
Overall, our findings indicate that the fungal endophytes in the mandarin tree can be affected by the tissue type, host plant location, and season. Although this is the first work on fungal endophytes of the mandarin tree and on factors that may structure their communities, more research is required to identify the functional and ecological significance of these fungal endophytes. Some of the species identified have been described as having antagonistic characteristics and potentials to promote plant growth. Therefore, a better understanding of this complex network of interactions between the mandarin tree and fungal endophytes and/or the consequence of these interactions would help to enhance mandarin’s productivity and sustainability.
References
Aly AH, Debbab A, Kjer J, Proksch P (2010) Fungal endophytes from higher plants: a prolific source of phytochemicals and other bioactive natural products. Fungal Divers 4(1):1–16
Angelini P, Rubini A, Gigante D, Reale L, Pagiotti R, Venanzoni R (2012) The endophytic fungi communities associated with the leaves and roots of the common reed (Phragmites australis) in lake Trasimeno (Perugia, Italy) in declining and healthy stands. Fungal Ecol 5:683–693
Arnold AE, Henk DA, Eells RL, Lutzoni F, Vilgalys R (2007) Diversity and phylogenetic affinities of foliar fungal endophytes in loblolly pine inferred by culturing and environmental PCR. Mycologia 99:185–206
Cheplick GP, Faeth S (2009) Ecology and evolution of the grassendophyte symbiosis. Oxford University Press, Oxford
Childs J, Kopp L, Johnson R (1965) A species of Physoderma present in Citrus and related species. Phytopathol 55(6):681–687
Chomcheon P, Wiyakrutta S, Sriubolmas N, Ngamrojanavanich N, Mahidol C, Ruchirawat S, Kittakoop P (2009) Metabolites from the endophytic mitosporic Dothideomycete sp. LRUB20. Phytochem 70(1):121–127
Clay K, Shearin ZR, Bourke KA, Bickford WA, Kowalski KP (2016) Diversity of fungal endophytes in non-native Phragmites australis in the Great Lakes. Biol invasions 18(9):2703–2716
Collado J, Platas G, Gonzalez I, Pelaez F (1999) Geographical and seasonal influences on the distribution of fungal endophytes in Quercus ilex. New Phytol 144:525–532
Douanla-Meli C, Langer E (2012) Diversity and molecular phylogeny of fungal endophytes associated with Diospyros crassiflora. Mycology 3:175–187
Douanla-Meli C, Langer E, Mouafo FT (2013) Fungal endophyte diversity and community patterns in healthy and yellowing leaves of Citrus limon. Fungal Ecol 6:212–222
Duran EL, Ploper LD, Ramallo JC, Piccolo Grandi RA, Hupper Giancoli AC, Azevedo JL (2005) The foliar fungal endophytes of Citrus limon in Argentina. Can J Bot 83:350–355
Dutta T, Sahoo R, Sengupta R, Ray SS, Bhattacharjee A, Ghosh S (2007) Novel cellulases from an extremophilic filamentous fungi Penicillium citrinum: production and characterization. J Ind Microbiol Biotechnol 35:275–282
El-Tarabily KA, Sivasithamparam K (2006) Potencial of yeast as biocontrol agents of soil-borne fungal plant pathogens and as plant growth promoters. Mycoscience 47:25–35
Glienke-Blanco C, Aguilar-Vildoso CI, Vieira MLC, Barroso PAV, Azevedo JL (2002) Genetic variability in the endophytic fungus Guignardia citricarpa isolated from citrus plants. Genet Mol Biol 25(2):251–255
Golparyan F, Azizi A, Soltani J (2018) Endophytes of Lippia citriodora (Syn. Aloysia triphylla) enhance its growth and antioxidant activity. Eur J Plant Pathol 152:759–768
Guo LD, Huang GR, Wang Y (2008) Seasonal and tissue age influences on endophytic fungi of Pinus tabulaeformis (pinaceae) in the dongling mountains, Beijing. J Int Plant Biol 50(8):997–1003
Hamayun M, Khan SA, Iqbal I, Ahmad B, Lee IJ (2010) Isolation of a gibberellin-producing fungus (Penicillium sp. MH7) and growth promotion of crown caisy (Chrysanthemum coronarium). J Microbiol Biotechnol 20:202–207
Hamilton C, Gundel PE, Helander M, Saikkonen K (2012) Endophytic mediation of reactive oxygen species and antioxidant activity in plants: a review. Fungal Divers 54:1–10
Helander ML, Sieber TN, Petrini O, Neuvonen S (1994) Endophytic fungi in Scots pine needles: spatial variation and consequences of simulated acid rain. Can J Bot 72(8):1108–1113
Hoffman M, Arnold AE (2007) Geographic locality and host identity shape fungal endophyte communities in cupressaceous trees. Mycol Res 112:331–344
Khan SA, Hamayun M, Yoon H, Kim HY, Suh SJ, Hwang SK, Kim JM, Lee IJ, Choo YS, Yoon UH, Kong WS (2008) Plant growth promotion and Penicillium citrinum. BMC Microbiol 8(1):231
Larget B, Simon DL (1999) Markov chain Monte Carlo algorithms for the Bayesian analysis of phylogenetic trees. Mol Biol Evol 16:750–759
Lee JS, Lee HK, Hur JS, Andreev M, Hong SG (2008) Diversity of the lichenized fungi in King George Island, Antarctica, revealed by phylogenetic analysis of partial large subunit rDNA sequences. J Microbiol Biotechnol 18:1016–1023
Linnakoski R, Puhakka-Tarvainen H, Pappinen A (2012) Endophytic fungi isolated from Khaya anthotheca in Ghana. Fungal Ecol 5:298–308
Lockwood JL (1981) Exploitation competion. In: Wicklow DT, Carrol GC (eds) The fungal community: its organization and role in ecosystem. Marcel Dekker, New York, pp 319–349
Martini M, Musetti R, Grisan S, Polizzotto R, Borselli S, Pavan F, Osler R (2009) DNA-dependent detection of the grapevine fungal endophytes Aureobasidium pullulans and Epicoccum nigrum. Plant Dis 93:993–998
Martins F, Pereira JA, Bota P, Bento A, Baptista P (2016) Fungal endophyte communities in above-and belowground olive tree organs and the effect of season and geographic location on their structures. Fungal Ecol 20:193–201
Mitchell AM, Strobel GA, Hess WM, Vargas PN, Ezra D (2008) Muscodor crispans, a novel endophyte from Ananas ananassoides. in the Bolivian Amazon. Fungal Divers 31:37–43
Nylander JAA (2004) MrModeltest V2. Evolutionary Biology Centre, Uppsala University, Uppsala
Ownley HB, Gwinn DK, Vega EF (2010) Endophytic fungal entomopathogens with activity against plant pathogens: ecology and evolution. Biocontrol 55:113–128
Pakvaz S, Soltani J (2016) Endohyphal bacteria from fungal endophytes of the Mediterranean cypress (Cupressus sempervirens) exhibit in vitro bioactivity. For Path 46:569–581
Pancher M, Ceol M, Corneo EC, Longa CMO, Yousaf S, Pertot I, Campisano A (2012) Fungal endophytic communities in Grapevines (Vitis vinifera L.) respond to crop management. Appl Environ Microbiol 78:4308–4317
Photita W, Lumyong S, Lumyong P, Hyde KD (2001) Endophytic fungi of wild banana (Musa acuminata) at Doi Suthep Pui National Park, Thailand. Mycol Res 105:1508–1513
Rodrigues KF (1994) The foliar fungal endophytes of the Amazonian palm Euterpe oleracea. Mycologia 86:376–385
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Schena L, Ippolito A, Zahavi T, Cohen L, Nigro F, Droby S (1999) Genetic diversity and biocontrol activity of Aureobasidium pullulans isolates against postharvest rots. Postharvest Biol Technol 17:189–199
Schena L, Nigro F, Pentimone I, Ligorio A, Ippolito A (2003) Control of postharvest rots of sweet cherries and table grapes with endophytic isolates of Aureobasidium pullulans. Postharvest Biol Technol 30:209–220
Schulthess FM, Faeth SH (1998) Distribution, abundances and association of the endophytic fungal community of Arizona fescue (Festuca arizonica). Mycologia 90:569–578
Shannon CE, Weaver W (1963) The mathematical theory of communication. University of Illinois Press, Urbana
Simpson GG (1964) Species density of North American recent mammals. Syst Zool 13:57–73
Slippers B, Wingfield MJ (2007) Botryosphaeriaceae as endophytes and latent pathogens of woody plants: diversity, ecology and impact. Fungal Biol Rev 21:90–106
Soltani J (2017) Endophytism in Cupressoideae (Coniferae): a model in endophyte biology and biotechnology. In: Maheshwari D (ed) Endophytes: biology and biotechnology. Sustainable development and biodiversity, vol 15. Springer, Cham, pp 127–143
Soltani J, Zaheri Shoja M, Hamzei J, Moghaddam SH, Pakvaz M S (2016) Diversity and bioactivity of endophytic bacterial community of Cupressaceae. For Path 46:353–361
Soltani J, Moghaddam MSH (2015) Fungal endophyte diversity and bioactivity in the mediterranean cypress Cupressus sempervirens. Curr microbiol 70(4):580–586
Sugita T, Nishikawa A (2003) Fungal identification method based on DNA sequence analysis. Reassessment of the methods of the pharmaceutical society of Japan and the Japanese pharmacopoeia. J Health Sci 49:531–533
Sun X, Ding Q, Hyde KD, Guo LD (2012) Community structure and preference of endophytic fungi of three woody plants in a mixed forest. Fungal Ecol 5:624–632
Tadych M, Bergen MS, Johnson-Cicalese J, Polashock JJ, Vorsa N, White JF Jr (2012) Endophytic and pathogenic fungi of developing cranberry ovaries from flower to mature fruit: diversity and succession. Fungal Divers 54:101–116
Vega FE, Simpkins A, Aime MC, Posada F, Pehner SA, Infante F, Castillo A, Arnold AE (2010) Fungal endophyte diversity in coffee plants from Colombia, Hawaii, Mexico and Puerto Rico. Fungal Ecol 3:122–138
Wakiyama M, Tanaka H, Yoshihara K, Hayashi S, Ohta K (2008) Purification and properties of family-10 endo-1,4-β-xylanase from Penicillium citrinum and structural organization of encoding gene. J Biosci Bioeng 105(4):367–374
Wearn JA, Sutton BC, Morley NJ, Gange AC (2012) Species and organ specificity of fungal endophytes in herbaceous grassland plants. J Ecol 100(5):1085–1092
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Yuan Z, Zhang C, Lin F, Kubicek CP (2010) Identity, diversity, and molecular phylogeny of the endophyticmycobiota in the roots of rare wild rice (Oryza granulate) from a nature reserve in Yunnan, China. Appl Environ Microbiol 76:1642–1652
Zabalgogeazcoa I (2008) Fungal endophytes and their interactions with plant pathogens. Spanish J Agric Res 6:138–146
Zabalgogeazcoa I, Gundel P, Helander M, Saikkonen K (2013) Nonsystemic fungal endophytes in Festuca rubra plants infected by Epichloe festucae in subarctic habitats. Fungal Divers 60:25–32
Zhang D, Yang Y, Castelbury LA, Cerniglia CE (1996) A method for large scale isolation of high transformation efficiency genomic DNA. FEMS Microbiol Lett 145:261–226
Acknowledgements
This research was supported by the research grant from University of Hormozgan, Iran, to Fatemeh Sadeghi in 2017.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sadeghi, F., Samsampour, D., Seyahooei, M.A. et al. Diversity and Spatiotemporal Distribution of Fungal Endophytes Associated with Citrus reticulata cv. Siyahoo. Curr Microbiol 76, 279–289 (2019). https://doi.org/10.1007/s00284-019-01632-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01632-9