Abstract
Azotobacter vinelandii is a soil bacterium that forms desiccation-resistant cysts, and the exopolysaccharide alginate is essential for this process. A. vinelandii also produces alginate under vegetative growth conditions, and this production has biotechnological significance. Poly-β-hydroxybutyrate (PHB) is another polymer synthetized by A. vinelandii that is of biotechnological interest. The GacS/A two-component signal transduction system plays an important role in regulating alginate production, PHB synthesis, and encystment. GacS/A in turn controls other important regulators such as RpoS and the ncRNAs that belong to the Rsm family. In A. vinelandii, RpoS is necessary for resisting oxidative stress as a result of its control over the expression of the catalase Kat1. In this work, we characterized a new ncRNA in A. vinelandii that is homologous to the P16/RsgA reported in Pseudomonas. We found that the expression of rgsA is regulated by GacA and RpoS and that it was essential for oxidative stress resistance. However, the activity of the catalase Kat1 is unaffected in rgsA mutants. Unlike those reported in Pseudomonas, RgsA in A. vinelandii regulates biofilm formation but not polymer synthesis or the encystment process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the last decade, small non-coding RNAs (ncRNAs) have gained great relevance in bacteriology. The ncRNAs are involved in many and diverse processes such as the stress response, motility, biofilm formation, control of metabolic fluxes, virulence, and many others [37]. In recent years, massive-scale RNA sequencing (RNAseq) and in silico genomic analysis have revealed hundreds of bacterial ncRNAs, but at present, the function of only a few of them is fully understood [27].
Genomic studies in Pseudomonas species revealed that one of the most ubiquitous ncRNAs is P16, also named RgsA [13]. In Pseudomonas fluorescens and Pseudomonas syringae, RgsA is related to the oxidative stress response, and in both cases, the ncRNA acts as a positive regulator [14, 28]. In P. syringae, rgsA mutants showed enhanced resistance to heat stress [28]. In P. aeruginosa, P. fluorescens, and P. syringae, the P16/rgsA ncRNA gene is transcriptionally controlled by GacA and the RpoS sigma factor [14, 28]. The two-component signal transduction system GacS/GacA is conserved in γ proteobacteria, and in many species, Gacs/A directly controls the expression of ncRNAs belonging to the Rsm family. The Rsm ncRNAs counteract the repressive activity of RsmA, an RNA-binding protein that functions as a translational repressor of its target genes [20]. Rsm ncRNAs show several stem-loop structures with a GGA-binding motif at the loops, which are the binding sites of RsmA [20]. The RgsA/P16 predicted secondary structure shows a single stem-loop structure with a GGA motif and resembles the Rsm ncRNA, but there are reports that ruled out a connection between RsgA and RsmA [14]. Contrary to observations for the rsm ncRNA genes, the regulatory region of rsgA does not contain a gacA-dependent activating sequence.
Azotobacter vinelandii is a soil bacterium that, under adverse environmental conditions, forms desiccation-resistant cysts [34]. Alginate is an exopolysaccharide that is essential for cyst formation [34]. Also vegetative cells produce alginate with other purposes. In this condition, alginate is proposed to act as a protective barrier against heavy metal toxicity or as a hydrophilic barrier that provides protection against adverse environmental conditions. Alginate is used for a variety of industrial purposes as a stabilizing or gelling agent [9]. Poly-β-hydroxybutyrate (PHB) is also synthetized by A. vinelandii and functions as an intracellular carbon and energy storage material [33]; it is a polyester of biotechnological interest because it can be used as a substrate to manufacture biodegradable plastics [12]. In A. vinelandii, GacS/GacA is involved in the control of alginate and PHB production [6]. A mutation in gacS abrogates both alginate and PHB synthesis and also significantly reduces the encystment capacity [6]. GacS/A in A. vinelandii regulates the transcription of the ncRNA genes rsmZ1 and rsmZ2. RsmZ1 and RsmZ2 partially relieve the repression carried out by RsmA over alginate synthesis [24]. On the other hand, GacA positively controls the transcription of the gene rpoS [7], encoding the stationary phase sigma factor essential for oxidative stress resistance in A. vinelandii [10, 32].
A. vinelandii possesses two catalases called Kat1 (also named CCC and encoded by the cccA gene) and Kat2 (encoded by katG) [31]. It was previously shown that Kat1 activity is necessary for survival under oxidative stress; its enzymatic activity is RpoS dependent and is predominantly observed in the stationary growth phase [32]. To date, it is not clear how RpoS controls Kat1 activity, and it is unknown whether σS directly regulates cccA transcription or if other types of regulation exist. Although Kat2 is expressed in the exponential growth phase, Kat2 activity is also detected in the stationary phase of cultures with high aeration [32]. A third type of catalase activity (called Kat3) can be detected; Kat3 is proposed as an isoform of Kat2, and its activity is detected under similar conditions to Kat2 [31, 32].
Recently, physiological studies in A. vinelandii revealed that the presence of endogenous or exogenous reactive oxygen species (ROS) is an important condition to promote biofilm formation [36]. On the other hand, general catalase activity is enhanced in biofilm-embedded cells compared with planktonic bacteria [36]. However, currently, there are no reports of genetic regulatory elements related to biofilm formation.
In this work, we report the identification and characterization of the RsgA/P16 homologue in A. vinelandii. As in Pseudomonas spp, rgsA is regulated by GacA and RpoS, and it is involved in oxidative stress resistance. However, in contrast to that reported in Pseudomonas, we show here that RgsA regulates biofilm formation but that this effect is not related to the synthesis of alginate.
Materials and Methods
Microbiological Procedures
The bacterial strains and plasmids used in this study are listed in Table 1. The medium and growth conditions were as follows: A. vinelandii was grown at 30 °C in Burk’s nitrogen-free salts medium [18] supplemented with 2 % sucrose (BS) or peptone yeast extract with 2 % sucrose (PY). E. coli strain DH5α was grown on Luria–Bertani medium (LB) at 37 °C. Antibiotic concentrations used (in micrograms per milliliter) for A. vinelandii and E. coli, respectively, were as follows: tetracycline, 20 and 20; ampicillin (Ap), not used and 100; nalidixic acid (Nal) and gentamycin (Gm), 1.5 and 10. A. vinelandii transformation was carried out as previously described [2]. The encystment assay was carried out as described by Moreno et al. [25].
Nucleic Acid Procedures
DNA isolation, cloning, and Northern blot procedures were carried out as described previously [30]. The A. vinelandii DJ genome sequence was used for designing the oligonucleotides used for PCR amplifications. The DreamTaq polymerase (Thermo Fisher Scientific) was used for PCR amplifications.
Cloning the A. vinelandii rgsA Locus
A 1.4-kb DNA fragment containing rgsA was amplified by PCR from A. vinelandii AEIV and DJ chromosomal DNA with the primers DcomrgsA (5′-GTTGCAGCGCCAGCGACTTATG-3′) and RcomrgsA (5′-CGAGACCGGGCTGGACTATCACTA-3′). This fragment was cloned into pGEM-T Easy (Promega). The plasmids generated were named pGErgsA1.4 and pGDJrsgA1.4 and were used to determine the nucleotide sequences of the rgsA locus in the AEIV and DJ strains, respectively.
Generation of A. vinelandii rgsA Mutants
To generate A. vinelandii rsgA mutants, an rsgA deletion was generated by inverse PCR using the primers FinvrsgA (5′-GGAAGGTACCAGCGGGCGAC-3′) and RinvrgsA (5′-AGAGGAGGTACCGTCGTGGAT-3′), which contain KpnI sites. Inverse PCR was carried out with pGErgsA1.4 as the template. The resulting fragment was digested with KpnI and ligated to generate the plasmid pGrsgA1.2, which has a 160 bp deletion of the rsgA locus. This plasmid was digested with KpnI, and a Gm cassette from pBSL41 [1] was ligated to generate pGrsgA1.2Gm. The rsgA deletion and the cassette insertions in the recombinant plasmid were verified by PCR and DNA sequencing. Plasmid pGrsgA1.2Gm, which is unable to replicate in A. vinelandii, was introduced into strains AEIV and DJ. Transformants resistant to Gm were isolated and confirmed by PCR analysis to carry the rgsA::Gm mutation (data not shown). The resultant mutants were named AEIVrgsA and DJrsgA.
Complementation of rsgA Mutants
To complement the rgsA mutants, a 0.9-kb DNA fragment that contains the rsgA gene plus its 0.5 kb upstream sequence containing the promoter and URS region was amplified by PCR using AEIV chromosomal DNA and the primers DcomrgsA and RsonrgsA (5′-TCGCCCGCTGATACTTTC-3′). This fragment was cloned into pGEM-T Easy (Promega) to generate pG0.9rgsA. To delimit the rgsA gene with its regulatory region, a fragment was excised with StuI (insertion site) and PstI (vector site) cuts. The 0.35 kb fragment (StuI-PstI) was cloned into the pUMATc vector [10]. The resultant plasmid was called pUMATcrsgA. pUMATc is an integrative suicidal vector that promotes the integration of the cloned DNA in the melA locus of the A. vinelandii chromosome. Plasmid pUMATcrsgA was introduced by transformation into strains AEIVrgsA and DJrgsA. Transformants resistant to Tc were isolated and confirmed by PCR analysis to carry the rgsA wild-type gene recombined in the melA locus using the primers DsonrgsA (5′-ACTGACGTGGCATTTCCC-3′) and RsonrgsA (data not shown). The resultant recombinants were named AEIVrgsA −/+ and DJrgsA −/+.
Oxidative Stress Assay and Heat Survival Assay
The assay was carried out as reported by Cocotl-Yañez et al. [11]. To carry out the heat survival assay, once cells were resuspended to standardize the cellular concentration, the cultures were incubated at 48 °C, taking samples prior to incubation and after 30, 60, and 120 min. The colony-forming units (CFU) in each sample were determined by serial dilutions.
Biofilm Assay
Biofilm formation in microtiter plates was quantified by crystal violet staining as reported by O’Toole and Kolter [26], with some modifications; we normalized biofilm formation to the concentration of protein instead, using optical density (OD) as growth parameter. Biofilm formation was reported as OD/mg of protein.
Analytical Methods
The protein concentration was determined by the Lowry method [23]. Alginate production was determined as previously described [3]. The PHB content of the bacteria was assayed by the method of Law and Slepecky [22]. The zymographic catalase analysis was conducted as described by Woodbury et al. [38].
Results
Identification of the A. vinelandii rgsA Gene
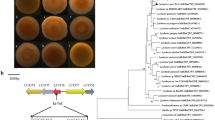
To identify ncRNAs in the A. vinelandii DJ genome, we analyzed the intergenic regions with the INFERNAL algorithm using the Rfam database as a comparison pattern (http://rfam.xfam.org/). One region located within nucleotides 1479172–1479292 of the DJ chromosome sequence presented 56 % identity with the rsgA homologue of P. fluorescens. The predicted secondary structure of the A. vinelandii RsgA ncRNA was similar to the Pseudomonas RgsA, showing only one stem-loop structure with a predicted GGA motif in the loop. rgsA was located between a gene encoding a putative TatD family hydrolase (Avin_15000) and a gene encoding a putative IS4 transposase (Avin_15010). The genome of the DJ strain has been sequenced and published [35] and is considered the type strain of A. vinelandii. This strain does not produce alginate due to an IS in the algU gene, which is essential for alginate production [25]. Hence, in this study, we worked with the AEIV strain, a wild-type mucoid strain of A. vinelandii [21].
The DNA fragments from the A. vinelandii AEIV and DJ strains containing the rgsA gene, including its promoter and terminator sequences, were amplified by PCR and cloned as described in Materials and Methods. Sequence analysis revealed 100 % identity between the two A. vinelandii strains. To detect rsgA transcripts, Northern blot analysis was conducted and confirmed the presence of the ncRNA in the AEIV and DJ strains (Fig. 1b).
a Alignments between P. aeruginosa rgsA, P. fluorescens rgsA, and P. syringae rgsA with A. vinelandii rsgA homologue. +1 putative transcription start and URS boxes are indicated. The −10 and −35 σS putative promoter sequences are boxed. A predicted terminator sequence is indicated by divergent arrows. b Northern blot analysis of rgsA in AEIV, AEIVrgsA, and AEIVrgsA −/+ (left panel) and DJ, DJrgsA, and DJ rgsA −/+ (right panel). The RNAs were isolated from stationary phase cultures (48 h of incubation in Burk’s sucrose medium). All blots were re-probed with a 16S rRNA probe to confirm that similar amounts of total RNA were analyzed
GacA and RpoS are Required for Efficient rgsA Expression
In P. fluorescens, P. aeruginosa, and P. syringae, RpoS controls rgsA transcription [14, 28]. To investigate whether RpoS controls the expression of the rgsA gene in A. vinelandii, Northern blot analysis was conducted using RNA extracted from the rpoS mutants derived from the AEIV and DJ strains. As shown in Fig. 1b, the expression of rgsA was not detected in the rpoS mutants.
GacA regulation of rgsA is a common phenomenon in Pseudomonas spp.; however, the rsgA regulatory regions do not show the typical gacA-dependent activating sequences. Interestingly, these regulatory regions show a conserved sequence called the URS (Upstream Regulatory Sequence). This box is proposed to serve as the binding site of an unknown regulator [14]. Likewise, in A. vinelandii, the rsgA regulatory region also contains an URS box (Fig. 1a). As shown in Fig. 1b, rgsA transcripts were detected in the AEIVgacA and DJ gacA mutants. In both cases, rgsA expression was diminished but not completely abrogated.
The ncRNA RsgA is an Important Factor in Oxidative Stress Resistance
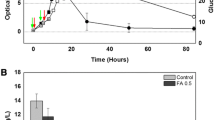
It was reported in P. fluorescens and P. syringae that RgsA is important for resisting oxidative stress. To investigate the role of RsgA in oxidative stress, an rsgA mutant derived from strain AEIV and carrying an rgsA deletion was constructed as described in Materials and Methods. The absence of the rgsA gene was confirmed by Northern blot analysis (Fig. 1b). We tested the survival of the rsgA mutant of A. vinelandii upon hydrogen peroxide exposure. As shown in Fig. 2a, the AEIVrgsA mutant exhibited enhanced sensitivity to hydrogen peroxide. The wild-type strain showed a survival of approximately 20 % after 300 mM H2O2 exposure, while the rsgA mutant showed a survival rate of approximately 4 %. Similar data were obtained for the DJrgsA mutant (data not shown). Genetic complementation of the rgsA mutant with a wild-type copy of the rsgA gene integrated into the chromosome restored the survival of the mutant, confirming the positive effect of rsgA on oxidative stress resistance.
Oxidative stress response and catalases activities in A. vinelandii strains. a Resistance to 300 mM of hydrogen peroxide of the strains AEIV, AEIVrgsA, and AEIVrgsA −/+, and the number of colony former units (CFU) after exposure to peroxide are shown. The bars represent the statistical media of six measures and its standard deviation. b Zymographic analyses of catalases activity. In situ catalase zymographs carried out with 50 μg of cell lysate from cultures of the strains AEIV, AEIVrgsA, and AEIVrgsA −/+, separated by non-denaturing PAGE. The three bands corresponding to different catalases activities are indicated
In A. vinelandii, the role of RpoS and Kat1 in the survival of oxidative stress has been well documented. We hypothesized that Kat1 catalase activity could be influenced by RgsA. To test this hypothesis, we carried out zymographic analysis to detect catalase activity in the A. vinelandii AEIVrgsA mutant. As shown in Fig. 2b, no differences in Kat1 activity were observed, indicating that RsgA is necessary for oxidative stress resistance in a Kat1-independent manner.
Additionally, we determined the high-temperature resistance response for the AEIVrgsA mutant. Contrary to published findings in P. syringae, differences between the rgsA mutant and its parental strain were not observed (Fig. 3).
RsgA Controls Biofilm Formation
As mentioned above, in A. vinelandii, an increase in ROS promotes biofilm formation [36]. Therefore, we investigated whether rgsA was involved in biofilm generation; the experiment was conducted as described in Materials and Methods. As expected, the AEIVrgsA mutant showed enhanced capacity for developing biofilms in both rich and minimal media (Fig. 4). The complementant strain (AEIVrgsA −/+) showed a similar behavior to the wild type.
Biofilm formation of AEIV, AEIVrgsA, and AEIVrgsA −/+ strains on polystyrene microtitrer plates. The determinations were carried out in minima medium (BS) and rich medium (PYS) as reported by O’Toole and Kolter [26]. The data represent the media of six individual experiments with its statistical standard deviation
RgsA is not Involved in Alginate Production, PHB Biosynthesis, or Encystment in A. vinelandii
In A. vinelandii, the two metabolites controlled by the GacS/A system are alginate and PHB [6, 7]. To investigate whether RgsA was involved in these GacA-regulated-pathways, we measured both polymers in rgsA. As shown in Table 2, alginate production by the AEIVrgsA mutant showed wild-type levels, indicating that RsgA is not involved in the regulation of this polymer. Likewise, PHB in the DJ genetic background was not affected by the rsgA mutation, DJrgsA, indicating that RgsA is not involved in the control that GacA exerts upon these polymers.
As both GacA and RpoS are essential for cyst formation and since these regulators control rsgA expression, the possible involvement of RgsA in this process was also evaluated. As shown in Table 2, the encystment capacity of the mutant AEIVrgsA was not affected, indicating that RsgA is not involved in this phenomenon.
Discussion
In this work, we reported the first study of rgsA outside of Pseudomonas spp. The Azotobacter and Pseudomonas groups belong to the Pseudomonadaceae family. In silico analysis revealed rsgA homologues in three A. vinelandii strains (DJ, CA and CA6). In this work, we characterized an rgsA homologue in a fourth strain (AEIV). We also localized an rsgA homologue in the recently published genome of Azotobacter chrococcum [ 29]. In all cases, the gene downstream of rgsA is a tatD homologue. The gene upstream of rgsA varies between the two Azotobacter species; in A. vinelandii, the 5′ neighbor is a transposase belonging to the IS4 family and in A. chrococcum, it is a class V aminotransferase. Our bioinformatics search did not find rgsA homologues outside of Pseudomonadaceae family, suggesting that RsgA is an ncRNA that is only related to this phylogenetic family.
The RgsA stem-loop structure with a GGA motif resembles the secondary structures of the Rsm ncRNAs. However, the current interaction model between RsmZ and RsmA involves two contiguous GGA motifs that bind an RsmA dimer, so it is unlikely that RgsA interacts with RsmA. The role of the RgsA GGA motif remains to be elucidated. Otherwise, in P. syringae, RgsA interacts with the RNA chaperone Hfq, which suggests that this chaperone promotes the interaction between the ncRNAs and their mRNA targets [28]. The predicted gene Avin 7540 of the A. vinelandii DJ genome encodes a putative homologue of Hfq, which could mediate RgsA activity; whether an RgsA–Hfq interaction occurs and its functional mechanism with its mRNA targets remain to be established.
The zymographic analysis of the catalases of A. vinelandii suggests that there is no regulatory relation between Kat1 activity and RgsA. The zymogram reveals predominant activity of Kat3, which is an isoform of Kat2; a similar phenomenon has been described by Sandercock et al. [32] in the late stationary phase of cultures with high aeration. The authors suggest that the Kat2/Kat3 activities occur in response to high levels of ROS caused by the high aeration. Therefore, the presence of Kat3 activity in the zymogram gel (Fig. 2b) could be due to increased ROS in the culture conditions. Interestingly, only a faint Kat2 band was detected in the rgsA mutant and was not present in the complemented strain, probably as a compensatory response to the defective oxidative stress response of the mutant.
Mucoid strains of A. vinelandii produce an alginate-dense biofilm [15]. Recently, Villa et al. [36] have shown that A. vinelandii responds to the increase in ROS by enhancing biofilm formation. As expected, we showed that RgsA is necessary to contend with oxidative stress and that its absence promotes a major biofilm formation. Further, the enhanced catalase activity in biofilms produced by cells expressing the alkyl hydroperoxide reductase gene (ahpC) is increased in the presence of endogenous ROS [36]; it would be interesting to determine the ahpC expression expression in rgsA mutants.
In P. fluorescens [16], P. syringae [8], and A. vinelandii [7], GacA has been shown to control rpoS expression and to regulate the transcription of ncRNAs from the Rsm family [20]. Therefore, the gacA mutant phenotypes could be related to RpoS, the Rsm system or both. In A. vinelandii, GacS/A controls alginate and PHB production, and this regulation is explained partially by the control exerted over RsmZ1 and RsmZ2 [24]; so it could be that RgsA will control the synthesis of polymers, fact that was ruled out in this study. Thus, it is possible that GacA will control some targets exclusively through RpoS.
In A. vinelandii, the rpoS mutant cysts completely lacked the exine and intine layers of the cyst capsule and are unable to resist desiccation [11]. Interestingly, inactivation of rpoS does not prevent alginate synthesis. Therefore, the inability of the rpoS mutant to form the capsule layers is not caused by the absence of alginate [11]. Based on the findings of Cocotl-Yanez et al. [11] it is likely that other genes controlled by RpoS are essential for cyst formation. rgsA could be one of these genes, although the encystment assay we developed ruled out this hypothesis. Similarly, in P. syringae, RpoS controls virulence genes independent of RgsA [28]. In both bacteria, the data suggest the existence of alternative regulation pathways controlled by RpoS, and one of these (RpoS–RgsA) controls the oxidative stress response. It remains to be determined whether other functions regulated by the RpoS–RsgA pathway exist, as well as its target genes.
In order to establish possible interaction targets of RsgA, we carried out a search using the IntaRNA algorithm (http://rna.informatik.uni-freiburg.de/IntaRNA/Input.jsp) [5]. The software found two interesting targets in the genome sequence of the DJ strain: HexR1 (Avin_27270) and AlgW (Avin_12950). HexR1 is a transcriptional regulator of Entner–Doudoroff pathway (ED). In Pseudomonas putida, HexR1 has been related with the increasing of NADPH production under oxidative stress. This fact suggests that the ED pathway regulates the cell redox status resulting in improved tolerance to oxidative stress [19]. AlgW is a homologue of the protease HtrA, interestingly, in P. aeruginosa AlgW is also involved in the resistance to oxidative stress [4].
In summary, in A. vinelandii, the expression of rgsA is controlled by RpoS and GacA. Besides, we found that RgsA is an important factor to resist the oxidative stress and plays a significant role in the biofilm formation.
References
Alexeyev MF, Shokolenko IN, Croughan TP (1995) Improved antibiotic-resistance gene cassette and omega elements for Escherichia coli vector construction and in vitro deletion/insertion mutagenesis. Gene 160:63–67
Bali A, Blanco G, Hill S, Kennedy C (1992) Excretion of ammonium by a nifL mutant of Azotobacter vinelandii fixing nitrogen. Appl Environ Microbiol 58:1711–1718
Blumenkrantz N, Asboe-Hansen G (1973) New method for quantitative determination of uronic acids. Anal Biochem 54:484–489
Boucher JC, Martinez-Salazar J, Schurr MJ, Mudd MH, Yu H, Deretic V (1996) Two distinct loci affecting conversion to mucoidy in Pseudomonas aeruginosa in cystic fibrosis encode homologs of the serine protease HtrA. J Bacteriol 178:511–523
Busch A, Richter AS, Backofen R (2008) IntaRNA: efficient prediction of bacterial sRNA targets incorporating target site accessibility and seed regions. Bioinformatics 24:2849–2856
Castañeda M, Guzmán J, Moreno S, Espín G (2000) The GacS sensor kinase regulates alginate and poly-β-hydroxybutyrate production in Azotobacter vinelandii. J Bacteriol 182:2624–2628
Castañeda M, Sánchez J, Moreno S, Núñez C, Espín G (2001) The global regulators GacA and σs form part of a cascade that controls alginate production in Azotobacter vinelandii. J Bacteriol 183:6787–6793
Chatterjee A, Cui Y, Yang H, Collmer A, Alfano JR, Chatterjee AK (2003) GacA, the response regulator of a two-component system, acts as a master regulator in Pseudomonas syringae pv. tomato DC3000 by controlling regulatory RNA, transcriptional activators, and alternate sigma factors. Mol Plant Microbe Interact 12:1106–1117
Clementi F (1998) Alginate production by Azotobacter vinelandii. Crit Rev Biotechnol 17:327–361
Cocotl-Yañez M, Moreno S, Encarnación S, López-Pliego L, Castañeda M, Espín G (2014) A small heat-shock protein (Hsp20) regulated by RpoS is essential for cyst desiccation resistance in Azotobacter vinelandii. Microbiology 160:479–487
Cocotl-Yañez M, Sampieri A, Moreno S, Núñez C, Castañeda M, Segura D, Espín G (2011) Roles of RpoS and PsrA in cyst formation and alkylresorcinol synthesis in Azotobacter vinelandii. Microbiology 157:1685–1693
Galindo E, Peña C, Núñez C, Segura D, Espín G (2007) Molecular and bioengineering strategies to improve alginate and polydydroxyalkanoate production by Azotobacter vinelandii. Microb Cell Fact 6:7. doi:10.1186/1475-2859-6-7
Gómez-Lozano M, Marvig RL, Molina-Santiago C, Tribelli PM, Ramos JL, Molin S (2015) Diversity of small RNAs expressed in Pseudomonas species. Environ Microbiol Rep 2:227–236
González N, Heeb S, Valverde C, Kay E, Reimmann C, Junier T, Hass D (2008) Genome-wide search reveals a novel GacA-regulated small RNA in Pseudomonas species. BMC Genom 9:167
Hashimoto W, Miyamoto Y, Yamamoto M, Yoneyama F, Murata K (2013) A novel bleb-dependent polysaccharide export system in nitrogen-fixing Azotobacter vinelandii subjected to low nitrogen gas levels. Int Microbiol 16:35–44
Heeb S, Valverde C, Gigot-Bonnefoy C, Haas D (2005) Role of the stress sigma factor RpoS in GacA/RsmA-controlled secondary metabolism and resistance to oxidative stress in Pseudomonas fluorescens CHA0. FEMS Microbiol Lett 243:251–258
Hernandez-Eligio A, Moreno S, Castellanos M, Castañeda M, Nuñez C, Muriel-Millan LF, Espin G (2012) RsmA post-transcriptionally controls PhbR expression and polyhydroxybutyrate biosynthesis in Azotobacter vinelandii. Microbiology 158:1956–1963
Kennedy C, Gamal R, Humphrey R, Ramos J, Brigle K, Dean D (1986) The nifH, nifM, and nifN genes of Azotobacter vinelandii: characterization by Tn5 mutagenesis and isolation from pLARF1 gene banks. Mol Gen Genet 205:318–325
Kim J, Park W (2014) Oxidative stress response in Pseudomonas putida. Appl Microbiol Biotechnol 98:6933–6946
Lapouge K, Schubert M, Allain FH, Haas D (2008) Gac/Rsm signal transduction pathway of gamma-proteobacteria: from RNA recognition to regulation of social behaviour. Mol Microbiol 67:241–253
Larsen B, Haug A (1971) Biosynthesis of alginate. 1. Composition and structure of alginate produced by Azotobacter vinelandii (Lipman). Carbohydr Res 17:287–296
Law J, Slepcky A (1961) Assay of poly-β-hydroxybutyric acid. J Bacteriol 82:33–36
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Manzo J, Cocotl-Yañez M, Tzontecomani T et al (2011) Post-transcriptional regulation of the alginate biosynthetic gene algD by the Gac/Rsm system in Azotobacter vinelandii. J Mol Microbiol Biotechnol 21:147–159
Moreno S, Guzmán J, Nájera R, Soberón-Chávez G, Espín G (1998) Role of the alternative σ factor AlgU in encystment of Azotobacter vinelandii. J Bacteriol 180:2766–2769
O’Toole GA, Kolter R (1998) Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol 30:295–304
Papenfort K, Vanderpool CK (2015) Target activation by regulatory RNAs in bacteria. FEMS Microbiol Rev 39:362–378
Park SH, Butcher BG, Anderson Z, Pellegrini N, Bao Z, D´Amico K, Filiatrault MJ (2013) Analysis of the small RNA P16/RgsA in the plant pathogen Pseudomonas syringae pv. tomato strain DC3000. Microbiology 159:296–306
Robson RL, Jones R, Robson RM, Schwartz A, Richardson TH (2015) Azotobacter genomes: The genome of Azotobacter chroococcum NCIMB 8003 (ATCC 4412). PLoS One 10(6):e0127997. doi:10.1371/journal.pone.0127997
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor
Sandercock JR, Page WJ (2008) Identification of two catalases in Azotobacter vinelandii: a KatG homologue and a novel bacterial cytochrome c catalase, CCCAv. J Bacteriol 190:954–962
Sandercock JR, Page WJ (2008) RpoS expression and the general stress response in Azotobacter vinelandii during carbon and nitrogen diauxic shifts. J Bacteriol 190:946–953
Segura D, Cruz T, Espín G (2003) Encystment and alkylresorcinol production by Azotobacter vinelandii strains impaired in poly-beta-hydroxybutyrate synthesis. Arch Microbiol 179:437–443
Segura D, Núñez C, Espín G (2014) Azotobacter Cysts. In: eLS. Wiley, Chichester. doi:10.1002/9780470015902.a0000295.pub2
Setubal JC, dos Santos P, Goldman BS et al (2009) Genome sequence of Azotobacter vinelandii, an obligate aerobe specialized to support diverse anaerobic metabolic processes. J Bacteriol 191:4534–4545
Villa F, Remelli W, Forlani F, Gambino M, Landini P, Cappitelli F (2012) Effects of chronic sub-lethal oxidative stress on biofilm formation by Azotobacter vinelandii. Biofouling. 28:823–833
Wagner EG, Romby P (2015) Small RNAs in bacteria and archaea: who they are, what they do, and how they do it. Adv Genet 90:133–208
Woodbury W, Spencer AK, Stahman MA (1971) An improved procedure using ferricyanide for detecting catalase isozymes. Anal Biochem 44:301–305
Acknowledgments
This work was supported by CONACyT Grant CB-2009-01-129525. We thank Cinthia Núñez for helpful comments regarding the manuscript. We also thank Daniel Segura for helpful discussions regarding the results. J. M. Huerta thanks CONACyT for M.Sc. scholarships.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huerta, J.M., Aguilar, I., López-Pliego, L. et al. The Role of the ncRNA RgsA in the Oxidative Stress Response and Biofilm Formation in Azotobacter vinelandii . Curr Microbiol 72, 671–679 (2016). https://doi.org/10.1007/s00284-016-1003-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-016-1003-2