Abstract
Emerging evidence suggest that macrophage and osteoclast are two competing differentiation outcomes from myeloid progenitors. In this review, we summarize recent advances in the understanding of the molecular mechanisms controlling the polarization of macrophage and osteoclast. These include nuclear receptors/transcription factors such as peroxisome proliferator-activated receptor γ (PPARγ) and estrogen-related receptor α (ERRα), their transcription cofactor PPARγ coactivator 1-β (PGC-1β), metabolic factors such as mitochondrial complex I (CI) component NADH:ubiquinone oxidoreductase iron-sulfur protein 4 (Ndufs4), as well as transmembrane receptors such as very-low-density-lipoprotein receptor (VLDLR). These molecular rheostats promote osteoclast differentiation but suppress proinflammatory macrophage activation and inflammation, by acting lineage-intrinsically, systemically or cross generation. These findings provide new insights to the understanding of the interactions between innate immunity and bone remodeling, advancing the field of osteoimmunology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Two major bone cell types: osteoblast and osteoclast
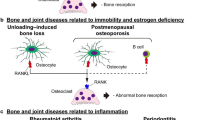
Bone is a relatively hard and dense connective tissue that is a major component of the skeleton, which not only provides structural integrity for the body but also performs many other vital functions such as movement, internal organ protection, and mineral metabolism [1]. In addition, bone provides a microenvironment that is critical for the development and maintenance of hematopoietic stem cells (HSCs) and mesenchymal stem cells (MSCs). Osteoblast and osteoclast are two major types of bone cell that are coordinated to maintain bone homeostasis [2,3,4,5,6]. Osteoblasts develop from stem cells of mesenchymal origin and are responsible for bone formation through producing an osteoid matrix that is calcified extracellularly. Osteocytes (i.e., the structural cells in the bone) are terminally differentiated osteoblasts embedded in bone matrix during the process of bone deposition. MSCs can also give rise to other cell types related to bone such as chondrocytes, marrow stromal cells, and bone marrow adipocytes. Mature osteoclasts are multinucleated giant cells that are derived from myeloid precursor cells of hematopoietic origin. Osteoclasts are formed by the fusion of myeloid precursor cells and specialized to remove mineralized bone matrix (i.e., bone resorption) through the production of lysosomal enzymes, such as tartrate-resistant acid phosphatase (TRAP) and cathepsin k (CTSK). HSCs are capable of differentiation into other immune cells such as macrophages, dendritic cells, as well as lymphoid cells. Bone formation by osteoblasts and bone resorption by osteoclasts occur mainly at the bone surface and are tightly coupled to maintain bone homeostasis. Dysregulated bone homeostasis generally leads to bone diseases, such as osteopetrosis and osteoporosis [7]. Osteopetrosis is characterized by increased bone density and a defect in bone marrow formation, while osteoporosis represents a condition with a significant increase in the risk of bone fractures due to decreased bone mass and density. Aberrantly activated osteoclastogenesis is one of the major reasons that directly lead to osteoporosis, which occurs frequently in postmenopausal women and aging population and can be exacerbated by pathological conditions such as inflammation.
Osteoclastogenesis and the RANK–RANKL–OPG signaling pathway
Bone cell communications play a crucial role in bone homeostasis. Many coupling factors that mediate these interactions between osteoblast and osteoclast have been established [8, 9]. Genetically modified mice and naturally occurring mutant mice have contributed greatly to the identification of the receptor activator of nuclear factor-κB ligand (RANKL)–RANK–osteoprotegerin (OPG) signaling pathway, as well as the associated molecular mechanisms regulating osteoclast differentiation and activation [8, 9]. Osteoblasts and their precursors can promote osteoclastogenesis by producing at least two known essential factors, macrophage colony-stimulating factor (M-CSF) and RANKL [10,11,12]. M-CSF acts as a factor for promoting the proliferation and survival of the osteoclast progenitors [13]. M-CSF functions through binding to its specific receptor c-FMS, which is a member of the receptor tyrosine kinase superfamily [14, 15]. RANKL was originally found as a T cell-derived cytokine mediating T cell proliferation and dendritic cell functions [16]. It is a transmembrane protein that belongs to the tumor necrosis factor (TNF) superfamily. In the presence of M-CSF, RANKL binding to its receptor RANK on osteoclast precursors promotes osteoclast differentiation, survival, and activation of bone resorption [17,18,19,20]. Synergistically, M-CSF stimulates the expression of RANK in osteoclast precursor cells [14], therefore rendering them being able to efficiently respond to RANKL. In addition to RANKL, osteoblasts and their precursors produce OPG, which also belongs to the TNF receptor superfamily and acts as a physiological decoy receptor of RANKL, and therefore inhibits osteoclastogenesis and bone resorption [21, 22]. The ratio between RANKL and OPG is considered to be a key determinant for osteoclast differentiation and bone resorption. Studies have indicated that osteocytes are essential source of RANKL during bone remodeling after birth [23,24,25], suggesting the interaction between osteocytes and osteoclasts during postnatal period. Binding of RANKL to RANK recruits the adapter protein TNF receptor-associated factor (TRAF) 6 (and other TRAFs) and therefore triggers a variety of intracellular signaling cascades [8], including a series of transcription factors such as mitogen-activated protein kinases (MAPKs), nuclear factor-κB (NF-κB), activator protein 1 (AP-1), and nuclear factor of activated T-cells 1/2 (NFATc1/2) [8]. These transcription factors work synergistically to induce the expression of specific genes of osteoclast including calcitonin receptor, TRAP and CTSK, leading to osteoclast differentiation, proliferation, and activation. Simultaneously, the activation of RANK signaling is also dependent on immunoglobulin-like receptors such as osteoclast-associated receptor (OSCAR), immunoglobulin-like receptor-A (PIR-A), signal regulatory protein-β1 (SIRPβ1), and triggering receptor expressed on myeloid cells 2 (TREM2), as well as immunoreceptor tyrosine-based activation motif (ITAM)-bearing molecules such as Fc-receptor common γ-subunit (FcRγ) and DAP12 [8], and all these molecules are necessary for costimulation and activation of calcium/calmodulin signaling. Sustained activation of calcium/calmodulin signaling is required for the induction and activation of NFATc1, which is a master regulator of osteoclast differentiation [26].
Osteoimmunology
The discovery of the RANK–RANKL–OPG signaling pathway has contributed enormously to the emergence and development of osteoimmunology, which represents a concept to study the interplay between immune and bone system (originally in understanding of immune regulation of osteoclasts) under both physiological and pathological conditions [27, 28]. The most direct evidence for the existence of the interplay between immune and bone system is that activated T cells express RANKL [16], and the blocked osteoclastogenesis in RANKL-deficient mice is restored by transgenic overexpression of RANKL only in T cells [29]. These studies indicate that osteoclastic bone resorption is influenced by immune system. In addition, studies have shown that osteoclasts share a number of regulatory molecules such as cytokines, receptors, signaling molecules and transcription factors with most of bone resident immune cells, including macrophages, dendritic cells, T cells, B cells, and natural killer (NK) cells [8, 30]. For example, inflammatory cytokines including interleukin-1 (IL-1), IL-6, IL-11, and TNF-α secreted from activated immune cells such as activated T lymphocytes may directly induce osteoclastogenesis and bone resorption through regulating the ratio of RANKL to OPG [31, 32], which frequently occurs in inflammatory bone diseases such as rheumatoid arthritis (RA). Conversely, some cytokines including IL-4, IL-10, and Interferon β (IFNβ) secreted from immune cells may exert the opposite effect [32]. A likely scenario is that an even more complex interaction exists between bone and immune system that exerts both positive and negative regulation of bone remodeling. Overall, the interaction between T cells and osteoclast/bone destruction was once a central subject of osteoimmunology and is quite well understood [8, 9]. Accumulating evidence suggest that the immune regulation of bone homeostasis also extends to osteoblastic bone formation which will not be discussed here [33], though the physiological and pathological significance as well as underlying molecular mechanisms are less well understood than in osteoclasts. Here, we mainly focus on the interaction between macrophage and osteoclast.
Macrophage and osteoclast
Macrophage and osteoclast are the two competing differentiation outcomes from myeloid progenitors. Macrophages are mononuclear myeloid immune cells present in nearly all tissues and known for their roles in eliminating invaded pathogens or infection, dead cells and debris, as well as orchestrating inflammatory responses [34]. Macrophages typically comprise 15 to 20% of the cells harvested from murine bone marrow [35, 36]. Bone resident tissue macrophages are termed as osteal macrophages which are predominantly located adjacent to osteoblast and may support osteoblastogenesis and bone formation [35, 37]. Thus, osteal macrophages represent a potential therapeutic target for improving fracture repair outcomes when a bone injury occurs. Considering their differential roles of macrophage and osteoclast in bone remodeling, the balance between these two types of cells regarding their differentiation and function may be especially important. Although osteoclasts share many common antigens with macrophages, there are some clear surface antigens that separate these two cell types [38, 39]. Broadly, activated macrophages can be divided into two major types—M1 and M2 [34]. M1 macrophages are pro-inflammatory ones which are classically activated by lipopolysaccharide (LPS) or T-helper 1 (Th1) cell-related cytokines such as IFNγ. M2 macrophages are identified as anti-inflammatory macrophages activated by Th2 cell-related cytokines such as interleukin IL-4 and IL-13. M1 macrophage-related cytokines such as TNF-α, IL-6, and IL-1β can induce osteoclastogenesis, while the M2 macrophage-related cytokines such as IL-4 and IL-10 can inhibit osteoclastogenesis through the inhibition of NFATc1 [8, 9]. Thus, the polarization of macrophage (M1/M2) itself is important for the determination of osteoclastogenesis, which makes the interaction between macrophage and osteoclast even more complex. Though the interaction between macrophage and osteoclast has been realized for a long while, the underlying molecular mechanisms are not well understood. In this review, we summarize recent advances in understanding of molecular mechanisms mediating the polarization of macrophage and osteoclast, which is an important aspect of osteoimmunology. Specifically, we will discuss the recent discoveries of novel modulators of the differentiation/activation of macrophage and osteoclast. These include nuclear receptors/transcription factors such as peroxisome proliferator-activated receptor γ (PPARγ) and estrogen-related receptor α (ERRα), their transcription cofactor PPARγ coactivator 1-β (PGC-1β), transmembrane receptors such as very-low-density-lipoprotein receptor (VLDLR), as well as metabolic factors such as mitochondrial complex I (CI) component NADH:ubiquinone oxidoreductase iron-sulfur protein 4 (Ndufs4). Interestingly, these molecular modulators act as rheostats to promote osteoclast differentiation but suppress proinflammatory macrophage activation and inflammation, by acting lineage-intrinsically, systemically or cross generation.
PPARγ in macrophage and osteoclast
PPARγ is a ligand-activated transcription factor that belongs to the nuclear hormone receptor superfamily [40, 41]. It forms heterodimer with retinoid X receptor (RXR) to associate with PPAR responsive element (PPRE) within the promoters of the target genes, and triggers transcription upon binding to agonistic ligands [40, 41]. Their ligands include long-chain fatty acids, peroxisome proliferators, the prostaglandin D2 metabolite 15-deoxy-(Delta12, 14)-prostaglandin J2 (15d-PGJ2), and the thiazolidinedione (TZD) class of antidiabetic agents [40,41,42,43,44]. PPARγ was originally shown to play an important role in adipocyte differentiation [45] and later was shown to be involved in modulating various physiological processes such as glucose and lipid metabolism as well as inflammatory responses [46, 47].
Since we know that macrophage and osteoclast are both differentiated from myeloid progenitors, thus, an interesting question is, what is the role of PPARγ in macrophages and osteoclast, respectively? PPARγ is expressed in myeloid-lineage cells such as macrophages and is markedly upregulated in activated macrophages [46]. Its expression is induced during macrophage differentiation [48, 49]. However, the role of PPARγ in macrophage differentiation is controversial due to the differences in experimental models. Some studies have indicated that PPARγ is required for the differentiation of macrophage [49, 50]. Others using embryonic stem cells suggest that PPARγ is not required for macrophage differentiation [48, 51]. Interestingly, a study demonstrates that PPARγ only determines the differentiation of fetal monocytes into alveolar macrophages but not for macrophage differentiation in other organs [52]. It has been shown that PPARγ is a negative regulator of macrophage activation and pro-inflammatory responses [46, 50]. Consistently, a variety of PPARγ agonists inhibit the production of inflammatory cytokines or inflammatory signaling in monocytes/macrophages [46, 50, 53, 54]. Conversely, inhibition of PPARγ in myeloid-lineage cells through overexpression of dominant-negative PPARγ leads to upregulation of proinflammatory cytokines [55]. In addition, both in vitro and in vivo experiments show that PPARγ or its activation is required for maturation of alternatively activated M2 macrophages with anti-inflammatory properties in a manner dependent on IL-4 and the signal transducer and activator of transcription 6 (STAT6) [56, 57]. All these findings suggest that PPARγ in myeloid-lineage cells plays a key role in preventing proinflammatory macrophage activation. Some studies show that activation of PPARγ may induce apoptosis of macrophages [58, 59]. Thus, selective PPARγ ligands may provide a strategy for treatment of inflammatory diseases such as atherosclerosis and RA in which macrophages are activated. Actually, TZDs including troglitazone, rosiglitazone, and pioglitazone were once widely used in the treatment of insulin resistance and type 2 diabetes by activating PPARγ [60, 61]. However, a variety of harmful effects of TZDs limit their widespread use in human patients [62, 63], and one of these is that TZDs decrease bone formation and accelerate bone loss in healthy and insulin-resistant individuals, and increase the risk of bone fractures in women with type 2 diabetes [62, 64]. Similarly, TZDs have been shown to cause bone loss in rodents [65, 66]. On possibility is that activation of PPARγ by TZDs may affect osteoclast function, which is confirmed by using PPARγ genetic mouse model [67].
PPARγ deletion in the osteoclast lineage impairs the differentiation of osteoclasts through compromising RANKL signaling, leading to osteopetrosis due to decreased osteoclast number and bone resorption [67]. Importantly, ligand activation of PPARγ by rosiglitazone exacerbates osteoclast differentiation in a receptor-dependent manner [67]. Mechanistically, PPARγ acts as a direct regulator of c-fos expression, which is an essential transcription factor for osteoclastogenesis [67, 68]. These results suggest that PPARγ and its ligands promote osteoclast differentiation and bone resorption, which partly explains why use of TZDs in human patients and rodents accelerate bone loss [62, 65, 66]. In this study, mature macrophage-related inflammatory genes such as monocyte chemoattractant protein-1 (MCP-1) and TNFα were upregulated in the mutant cells and suppressed by rosiglitazone in wild-type (WT) cells [67], which is consistent with the previously reported anti-inflammatory role of PPARγ or its activation [46, 53]. Interestingly, it has been revealed that osteoclast progenitors reside in the PPARγ expressing bone marrow cells [69]. In addition to its role in osteoclast, PPARγ has also been suggested to inhibit osteoblast differentiation and bone formation [70, 71]. Collectively, PPARγ and its ligands exert effects on bone homeostasis by promoting osteoclast differentiation and bone resorption as well as inhibiting osteoblast differentiation and bone formation [72]. These findings suggest that bone loss caused by long-term use of TZDs may owe to a combination of increased bone resorption and decreased bone formation.
In summary, PPARγ acts as an important molecular rheostat to promote osteoclast differentiation/activation but suppress proinflammatory macrophage activation and inflammation. Aberrant activation and disruption of PPARγ will lead to bone diseases (e.g., osteoporosis) and inflammatory diseases, respectively. PPARγ is still an important target for improving insulin sensitivity and eliminating the side effects of TZDs including those caused to bone is a prerequisite for their widespread use for patients; thus, novel strategies are necessary to modulate PPARγ activity to enhance the beneficial effects and reduce unwanted adverse effects [73, 74]. In addition to PPARγ, studies have shown that the pro-osteoclastogenic effect of rosiglitazone is also mediated coordinately by ERRα and PGC-1β [75, 76]. Interestingly, these two factors are also involved in modulating the activation of macrophage, which will be discussed below in more detail.
ERRα in macrophage and osteoclast
ERRα is an orphan nuclear receptor that belongs to the ERR subfamily (ERRα, ERRβ, and ERRγ). It was identified initially based on their high degree of sequence homology with two estrogen receptors (ERα and ERβ) [77,78,79,80]; however, it does not bind estrogen and is not activated by estrogen directly [81]. ERRα is widely expressed in tissues of developing and adult animals, with enriched expression in tissues with high oxidative metabolic capacity [81, 82]. ERRα acts as a transcription factor that regulates metabolic homeostasis such as fatty acid oxidation (FAO) and the adaptive bioenergetic response [83,84,85,86]. For example, mice with genetic deletion of ERRα show a lean phenotype with decreased white adipose tissue, and they are resistant to high-fat diet-induced obesity and have reduced lipogenesis in adipose tissues [85]. Importantly, genes targeted by ERRα or genes mediating ERRα activity also control metabolic regulation [87]. Notably, ERRα acts as a transcriptional regulator of medium-chain acyl CoA dehydrogenase (MCAD), an enzyme involved in mitochondrial FAO [88, 89]. ERRα can be activated by its coactivators, including PGC-1α and PGC-1β, both of which play an important role in mitochondria biogenesis and respiration [90,91,92,93]. Recent studies have shown that ERRα also acts as a key mediator of the functions of macrophage and osteoclast [75, 76, 94,95,96,97,98].
ERRα expression is upregulated in activated macrophages such as those induced by IFNγ or LPS [95, 99]. Mice with genetic deletion of ERRα show excessive systemic inflammation and proinflammatory responses in macrophage upon immune challenge [95]. Further analysis shows that ERRα acts as a negative regulator of Toll-like receptor (TLR)-induced inflammatory responses through inducing TNFα-induced protein 3 (Tnfaip3) transcription and controlling the metabolic reprogramming in macrophages [95]. Consistent with these findings, Wei et al. report that proinflammatory cytokines produced by activated macrophages are suppressed by cholesterol activation of ERRα [76]. Meanwhile, other studies suggest that ERRα in macrophage together with its coactivator PGC1β play a key role in host defense such as antibacterial activity [94, 97]. ERRα or PGC-1β deficiency results in decreased mitochondrial gene expression, intracellular reactive oxygen species (ROS) level, and bacterial clearance in IFN-γ-activated macrophages [94], suggesting a link between mitochondrial oxidative metabolism and macrophage-driven antibacterial immunity. Collectively, these results indicate that ERRα is required for maintaining the homeostasis/function of macrophage such as controlling overwhelming proinflammatory responses under normal physiological conditions and promoting antibacterial activity when necessary.
Considering the similar origin for osteoclast and macrophage, it is not surprising that ERRα is also found to be expressed in osteoclasts [82, 100]. ERRα has been demonstrated to be involved in the regulation of osteoclast adhesion and transmigration [101]. Later its role in osteoclasts has been systematically analyzed in ERRα knockout (KO) mice [75]. ERRα KO mice exhibit osteopetrosis due to osteoclast defects and decreased bone resorption, suggesting that ERRα is an important regulator of osteoclasteogenesis. Interestingly, rosiglitazone activation of PPARγ induces the expression of ERRα, while ERRα deletion compromises the expression of several key genes of osteoclast and formation of mature osteoclasts stimulated by RANKL and rosiglitazone. Concomitantly, ERRα deletion completely abolishes the rosiglitazone induction of genes related to mitochondrial biogenesis/activation in osteoclasts. These results suggest that ERRα is a direct PPARγ target gene in the specific context of osteoclastogenesis, and it acts as a key PGC-1β (as an ERRα coactivator) target to promote osteoclast function and mitochondrial biogenesis/activation. This study reveals a previously unrecognized role of ERRα in promoting osteoclastogenesis by inducing the expression of mitochondrial genes via a PGC1β-dependent mechanism. No natural ligand has been found until a study by Wei et al. identifies cholesterol as an endogenous ERRα agonist [76]. In this study, the rheostat role of ERRα in macrophage and osteoclast is further confirmed in another manner, as cholesterol activation of ERRα inhibits the proinflammatory cytokines produced by macrophage and promotes osteoclast differentiation, while all these effects caused by cholesterol are ablated by ERRα deletion. Concomitantly, pharmacological effects of statin and bisphosphonate (for depletion of cholesterol) on bone resorption and skeletal remodeling are mediated by ERRα [76]. In line with this report, ERRα also acts as a mediator of the driving effect of MYC-dependent oxidative metabolism on osteoclastogenesis [98]. In addition, ERRα has also been reported to inhibit osteoblast development and bone formation [102, 103].
Together, these findings identify ERRα as a critical regulator of bone homeostasis through its functions in osteoclast and osteoblast, as well as a regulator of macrophage function, suggesting that proper ERRα activity is important to maintain the balance between osteoclast and macrophage.
PGC-1β in macrophage and osteoclast
PGC-1-related transcription coactivators are usually expressed in tissues with a high oxidative capacity, such as muscle, brown fat, and liver [104]. Similar with roles of the other two members of PGC-1 family, PGC-1α and PGC-1-related coactivator (PRC), PGC-1β is a strong activator of mitochondrial biogenesis and respiration and therefore regulates multiple aspects of energy metabolism, including adaptive thermogenesis and FAO [91, 105,106,107,108,109,110,111]. Their regulating functions are achieved by activating specific target transcription factors including PPARγ and ERRα. Genetically manipulated mouse models greatly promote the understanding of the in vivo physiological functions of PGC-1β. For example, transgenic mice with overexpression of PGC-1β are resistant to obesity due to the elevated energy expenditure [91]. Muscle fibers with transgenic PGC-1β overexpression are rich in mitochondria and highly oxidative [112], while hepatic-specific activation of PGC-1β protects against steatohepatitis [113]. PGC-1β-deficient mice show less resistance to acute cold exposure due to impaired adaptive thermogenesis [107, 110], insulin resistance [109] as well as hepatic steatosis on a high-fat diet [107, 110]. More or less, mitochondrial dysfunction is observed in different tissues with PGC-1β ablation [107, 109, 110, 114, 115].
As a coactivator of PPARγ and ERRα, one intriguing question is, does PGC-1β have a similar role as PPARγ and ERRα in the regulation of macrophage and osteoclast? Evidence show that PGC-1β expression and oxidative metabolism are induced during alternative macrophage activation (M2 macrophage activation) [116]. Transgenic expression of PGC-1β promotes the differentiation and activation of M2 macrophages possibly through programming macrophage for FAO and mitochondrial biogenesis and attenuates the expression of proinflammatory cytokines, whereas inhibition of mitochondria respiration or blocking PGC-1β impairs alternative macrophage activation [116]. This is consistent with the results from a recent study showing that repolarization of inflammatory macrophages to an M2 phenotype is prevented by the inhibition of mitochondrial oxidative respiration [117]. Mechanistic analyses show that PGC-1β co-activates the transcriptional functions of STAT6 to metabolically shift macrophages to the alternative state. Another study shows that PGC-β and FAO is downregulated in bone marrow derived macrophages (BMDM) with tuberous sclerosis 1 (TSC1) deletion, and these macrophages fail to upregulate the M2 program [118], further suggesting that PGC1-β-mediated oxidative metabolism is crucial for alternative macrophage functions and prevention of proinflammatory responses. Consistently, several other reports also show the anti-inflammatory property of PGC-1β under different settings [119, 120]. All these studies suggest that PGC-1β may be a key player linking mitochondrial oxidative metabolism to the anti-inflammatory program of alternative macrophage activation.
Interestingly, PGC-1β is induced during osteoclast differentiation by cyclic adenosine monophosphate (cAMP) response element–binding protein (CREB) as a result of ROS [121]. Knockdown of PGC-1β in vitro inhibits osteoclast differentiation and mitochondria biogenesis, whereas mice with global PGC-1β deletion exhibits impaired osteoclast function leading to increased bone mass [121]. Notably, PGC-1β-deficient osteoblasts also show defects [121]. A study by Wei et al. specifically dissects the mechanisms for how PGC-1β mediates the induction of osteoclast differentiation and bone loss by rosiglitazone [75]. Wei et al. found that PGC-1β is highly induced by rosiglitazone during osteoclast differentiation in a PPARγ-dependent manner [75]. Rosiglitazone-activated PPARγ indirectly induces PGC-1β expression by downregulating β-catenin and derepressing c-jun. In vitro experiments show that PGC-1β is required for the pro-osteoclastogenic effect of rosiglitazone. Importantly, it has been demonstrated that PGC-1β deletion in the osteoclast lineage confers complete resistance to rosiglitazone-induced bone loss. These findings identify PGC-1β as an essential co-activator of PPARγ to promote osteoclastogenesis in vivo.
In summary, as a coactivator of PPARγ and ERRα, PGC-1β is an important molecular rheostat in promoting osteoclast differentiation and mitochondrial function but suppressing pro-inflammatory responses through programming alternative macrophage activation. Thus, a transcriptional network consisting at least PPARγ, ERRα, and PGC-1β is required for maintaining the proper polarization of macrophage and osteoclast.
Ndufs4 in macrophage and osteoclast
Ndufs4 is an 18 kDa subunit of mitochondrial complex I (CI), which is located in the mitochondrial inner membrane [122, 123]. CI is comprised of at least 45 different proteins and is a critical component for electron transport and generation of a proton gradient across the mitochondrial inner membrane to drive adenosine triphosphate (ATP) production [122, 123]. CI-associated defects are the most common mitochondrial disorders which lead to metabolic diseases such as impaired oxidative metabolism, as well as neurological diseases such as Alzheimer, Parkinson, and Leigh syndrome [124,125,126,127,128]. Accumulating evidence have shown that Ndufs4 is essential for proper assembly of CI [129, 130], and its mutation causes Leigh syndrome in humans and is associated with various defects including retarded growth, developmental delay, visual defects, muscular hypotonia, encephalomyopathy, cardiomyopathy, lethargy, and failure to thrive [129,130,131,132,133,134]. Global Ndufs4 KO mice or mice with loss of Ndufs4 specifically in the brain shows a similar phenotype of encephalopathy resembling Leigh syndrome including retarded growth, loss of motor ability, breathing abnormalities, and death by ~ 7 weeks old [128, 135].
Interestingly, global Ndufs4 KO pups exhibit a transient alopecia phenotype [128], and this phenotype has been shown to be caused by a systemic inflammatory response [136]. Detailed ex vivo experiments show that Ndufs4 deletion in macrophages leads to the upregulation of pro-inflammatory genes, indicating that the systemic inflammation in Ndufs4 KO pups is likely attributed to, at least in part, the activation of macrophages [136]. Experiments using mice with hematopoietic-specific deletion of Ndufs4 further confirm the cell-autonomous role of Ndufs4 in the prevention of proinflammatory macrophage activation and inflammation. Similarly, treatment with a CI inhibitor rotenone also elevates the expression of proinflammatory genes in WT macrophages. These results indicate that Ndufs4 (or CI) in macrophage indeed plays a cell-autonomous role in prevention of proinflammatory macrophage activation and inflammation.
Both global deletion of Ndufs4 and hematopoietic deletion of Ndufs4 decrease bone resorption and increase bone mass due to the compromised osteoclastogenesis [136]. These results indicate that Ndufs4 (or CI) plays a cell intrinsic role in the osteoclast lineage to promote osteoclastogenesis and decrease bone mass. Ndufs4 deletion specifically in liver causes a metabolic shift from FAO to glycolysis, leading to accumulation of fatty acids and lactate in the circulation. The accumulated fatty acids and lactate further activate Ndufs4 KO macrophages via ROS induction and diminish osteoclast lineage commitment in Ndufs4 KO myeloid progenitors; both inflammation and osteopetrosis in Ndufs4 KO mice are attenuated by TLR4 deletion. These findings uncover TLR4 as an essential mediator of the systemic inflammation in the Ndufs4 KO pups. In summary, using deletion of the essential CI subunit Ndufs4 as a model for mitochondrial dysfunction, this study reports that normal mitochondrial oxidative metabolism suppresses proinflammatory macrophage activation and inflammation while promotes osteoclast differentiation and bone resorption via both cell-autonomous and systemic regulation. These findings reveal the essential CI subunit Ndufs4 as a critical rheostat in macrophage and osteoclast polarization.
Maternal VLDLR in offspring macrophage and osteoclast
VLDLR is a transmembrane lipoprotein receptor that belongs to the low-density-lipoprotein (LDL) receptor superfamily [137, 138]. It is abundantly expressed in fatty-acid-utilizing tissues (heart, skeletal muscle, and adipose tissue), brain and macrophages, but essentially absent in liver and intestine. VLDLR regulates lipid metabolism via modulating the uptake of its ligands such as VLDL [139, 140]. It is also involved in the regulation of brain development by forming a heterodimer with apolipoprotein E receptor 2 (ApoER2) [141]. Activation of VLDLR by its ligand Reelin (RELN) triggers downstream signaling events that modulate neuronal functions [141, 142]. In addition, VLDLR has been implicated in metabolic homeostasis [140, 143,144,145]. Disruption of VLDLR in mice leads to a reduction in adipose tissue [143, 146]. On an LDL receptor-deficient background, VLDLR deletion increases serum triglyceride in mice on a high-fat diet, while VLDLR overexpression decreases serum triglyceride [147]. VLDLR in macrophage has shown to be a pro-atherogenic factor [145], as reconstitution of macrophage VLDLR expression in VLDLR KO mice largely increases atherosclerotic lesion development, probably by mediating the accumulation of atherogenic lipoproteins.
The unique role of VLDLR in macrophage and osteoclast has been revealed at the maternal-offspring interface [148,149,150]. Maternal deletion of VLDLR in mice leads to the production of defective milk with diminished levels of platelet-activating factor acetylhydrolase (PAFAH) due to the impaired expression of phospholipase A2 group 7 (PLA2G7) in macrophages [148]. Platelet-activating factors (PAFs) are a class of potent proinflammatory lipids that are present in neonates, and are elevated in infants suffering inflammatory disorders such as necrotizing enterocolitis [151, 152]. PAFs can be removed by the degradation enzyme PAFAH. Studies have shown that secreted form of PAFAH exists in milk, which may function to suppress the proinflammatory activity of PAF in the nursing neonates [151, 153, 154]. PAFs accumulate in the nursing pups with ingestion of the PAFAH-deficient milk from VLDLR KO mother, resulting in symptoms related to systemic inflammation such as alopecia, anemia and growth retardation [148]. Interestingly, these neonatal defects can be rescued by oral supplementation of PAFAH to the pups. Therefore, macrophage VLDLR promotes PAFAH secretion in mother’s milk and suppresses systemic inflammation in nursing neonates. This study reveals a novel anti-inflammatory role of VLDLR in macrophage at the maternal-offspring interface and demonstrates the physiological significance of maternal VLDLR in ensuring milk quality.
Provocatively, a later study shows that ingestion of defective milk can exert a long-term impact on the offspring to adulthood [149]. In this study, the role of maternal and offspring VLDLR in osteoclasts is investigated. The deficient milk from VLDLR KO mothers blunts the differentiation of osteoclast and bone resorption and consequently causes osteopetrosis in the offspring [149]. The detailed pharmacological and genetic rescue experiments reveal that the milk defects are largely due to an excessive activity of mammalian target of rapamycin (mTOR) signaling in the adipocytes, which can be reversed by maternal rapamycin treatment during lactation. Meanwhile, excessive activity of mTOR in the adipocytes also results in an elevated expression of 3-hydroxy-3-methylglutaryl coenzyme A reductase (HMGCR), a rate-limiting enzyme in the cholesterol biosynthetic pathway, consequently leading to increased levels of cholesterol precursors in the milk. Together, these findings uncover the cellular, molecular and biochemical mechanisms for how maternal VLDLR ensures milk quality, protects the neonatal offspring from systemic inflammation, as well as enhances normal differentiation of osteoclast in adult offspring. Thus, maternal VLDLR control offspring macrophage osteoclast polarization by increasing osteoclastogenesis but decreasing proinflammatory macrophage activation. In contrast, in the offspring, VLDLR suppresses osteoclast differentiation by promoting RANKL signaling, leading to osteoporosis, although maternal VLDLR plays a dominant role over offspring VLDLR in the regulation of bone resorption [149]. Moreover, VLDLR in macrophages of adipose tissue aggravates adipose inflammation and insulin resistance in obesity [155], suggesting that in the offspring, VLDLR also modulates macrophage osteoclast polarization by inhibiting osteoclastogenesis but activating proinflammatory macrophages. Together, these interesting findings identify maternal and offspring VLDLR as critical yet opposite rheostats that controls myeloid lineage allocation into osteoclast or macrophage.
Conclusions and perspectives
In this review, we have summarized and discussed the recently discovered modulators of the polarization between macrophage and osteoclast. These include nuclear receptors and transcription factors such as PPARγ and ERRα, their transcription cofactor PGC-1β, metabolic factors such as mitochondrial CI component Ndufs4, as well as transmembrane receptors such as VLDLR. Intriguingly, these molecular rheostats are all related to energy metabolism, and at the same time all function to promote osteoclast differentiation but suppress proinflammatory macrophage activation, by acting lineage-intrinsically, systemically or cross generation (Fig. 1). Particularly, the convergence of the functions of ERRα, PGC-1β and Ndufs4 at mitochondrial oxidative metabolism implies an important role of metabolic control of the lineage specification for macrophage and osteoclast. Several studies have already highlighted mitochondrial oxidative metabolism/metabolic (re)programming as a key player in inflammatory signaling in macrophage [117, 156, 157]. Thus, mitochondrial metabolism may represent a novel therapeutic target for both inflammatory diseases and bone resorption disorders. These findings provide new insights into the understanding of the interaction between innate immunity and bone remodeling. These significant findings open an exciting path to future research to identify additional factors and mechanisms that control osteoclast and macrophage polarization. We believe that more progress will be made to deepen our understanding of the interaction between innate immunity and bone remodeling.
Macrophage and osteoclast are two competing differentiation outcomes from myeloid progenitors. The polarization of macrophage and osteoclast is regulated by molecular rheostats. Peroxisome proliferator-activated receptor γ (PPARγ) and its ligand rosiglitazone (Rosi), estrogen-related receptor α (ERRα) and its endogenous agonist cholesterol (Chol), PPARγ coactivator 1-β (PGC-1β), mitochondrial complex I (CI) component NADH:ubiquinone oxidoreductase iron-sulfur protein 4 (Ndufs4), as well as maternal very-low-density-lipoprotein receptor (VLDLR) act as rheostats to promote osteoclast differentiation but suppress proinflammatory macrophage activation and inflammation, by acting lineage-intrinsically, systemically or cross generation. M-CSF macrophage colony-stimulating factor, RANKL receptor activator of nuclear factor-κB ligand
References
Clarke B (2008) Normal bone anatomy and physiology, Clinical. J Am Soc Nephrol 3(Supplement 3):S131–S139
Manolagas SC (2000) Birth and death of bone cells: basic regulatory mechanisms and implications for the pathogenesis and treatment of osteoporosis*. Endocr Rev 21(2):115–137
Harada S-i, Rodan GA (2003) Control of osteoblast function and regulation of bone mass. Nature 423:349
Seeman E, Delmas PD (2006) Bone quality — the material and structural basis of bone strength and fragility. N Engl J Med 354(21):2250–2261
Aubin JE (2001) Regulation of osteoblast formation and function. Rev Endocr Metab Disord 2(1):81–94
Teitelbaum SL (2000) Bone resorption by osteoclasts. Science 289(5484):1504–1508
Zelzer E, Olsen BR (2003) The genetic basis for skeletal diseases. Nature 423:343
Takayanagi H (2007) Osteoimmunology: shared mechanisms and crosstalk between the immune and bone systems. Nat Rev Immunol 7:292
Nakashima T, Hayashi M, Takayanagi H (2012) New insights into osteoclastogenic signaling mechanisms. Trends Endocrinol Metab 23(11):582–590
Tacken PJ, Teusink B, Jong MC, Harats D, Havekes LM, van Dijk KW, Hofker MH (2000) LDL receptor deficiency unmasks altered VLDL triglyceride metabolism in VLDL receptor transgenic and knockout mice. J Lipid Res 41(12):2055–2062
Teitelbaum SL, Ross FP (2003) Genetic regulation of osteoclast development and function. Nat Rev Genet 4:638
Theill LE, Boyle WJ, Penninger JM (2002) RANK-L and RANK: T cells, bone loss, and mammalian evolution. Annu Rev Immunol 20(1):795–823
Tanaka S, Takahashi N, Udagawa N, Tamura T, Akatsu T, Stanley ER, Kurokawa T, Suda T (1993) Macrophage colony-stimulating factor is indispensable for both proliferation and differentiation of osteoclast progenitors. J Clin Invest 91(1):257–263
Arai F, Miyamoto T, Ohneda O, Inada T, Sudo T, Brasel K, Miyata T, Anderson DM, Suda T (1999) Commitment and differentiation of osteoclast precursor cells by the sequential expression of C-Fms and receptor activator of nuclear factor κb (Rank) receptors. J Exp Med 190(12):1741–1754
Ross FP, Teitelbaum SL (2005) αvβ3 and macrophage colony-stimulating factor: partners in osteoclast biology. Immunol Rev 208(1):88–105
Anderson DM, Maraskovsky E, Billingsley WL, Dougall WC, Tometsko ME, Roux ER, Teepe MC, DuBose RF, Cosman D, Galibert L (1997) A homologue of the TNF receptor and its ligand enhance T-cell growth and dendritic-cell function. Nature 390:175
Fuller K, Wong B, Fox S, Choi Y, Chambers TJ (1998) TRANCE is necessary and sufficient for osteoblast-mediated activation of bone resorption in osteoclasts. J Exp Med 188(5):997–1001
Lacey DL, Timms E, Tan HL, Kelley MJ, Dunstan CR, Burgess T, Elliott R, Colombero A, Elliott G, Scully S, Hsu H, Sullivan J, Hawkins N, Davy E, Capparelli C, Eli A, Qian YX, Kaufman S, Sarosi I, Shalhoub V, Senaldi G, Guo J, Delaney J, Boyle WJ (1998) Osteoprotegerin ligand is a cytokine that regulates osteoclast differentiation and activation. Cell 93(2):165–176
Yasuda H, Shima N, Nakagawa N, Yamaguchi K, Kinosaki M, Mochizuki S-i, Tomoyasu A, Yano K, Goto M, Murakami A, Tsuda E, Morinaga T, Higashio K, Udagawa N, Takahashi N, Suda T (1998) Osteoclast differentiation factor is a ligand for osteoprotegerin/osteoclastogenesis-inhibitory factor and is identical to TRANCE/RANKL. Proc Natl Acad Sci 95(7):3597–3602
Wong BR, Josien R, Choi Y (1999) TRANCE is a TNF family member that regulates dendritic cell and osteoclast function. J Leukoc Biol 65(6):715–724
Simonet WS, Lacey DL, Dunstan CR, Kelley M, Chang MS, Lüthy R, Nguyen HQ, Wooden S, Bennett L, Boone T, Shimamoto G, DeRose M, Elliott R, Colombero A, Tan HL, Trail G, Sullivan J, Davy E, Bucay N, Renshaw-Gegg L, Hughes TM, Hill D, Pattison W, Campbell P, Sander S, Van G, Tarpley J, Derby P, Lee R, Boyle WJ (1997) Osteoprotegerin: a novel secreted protein involved in the regulation of bone density. Cell 89(2):309–319
Tsuda E, Goto M, Mochizuki S-i, Yano K, Kobayashi F, Morinaga T, Higashio K (1997) Isolation of a novel cytokine from human fibroblasts that specifically inhibits osteoclastogenesis. Biochem Biophys Res Commun 234(1):137–142
Xiong J, Onal M, Jilka RL, Weinstein RS, Manolagas SC, O'Brien CA (2011) Matrix-embedded cells control osteoclast formation. Nat Med 17:1235
Nakashima T, Hayashi M, Fukunaga T, Kurata K, Oh-hora M, Feng JQ, Bonewald LF, Kodama T, Wutz A, Wagner EF, Penninger JM, Takayanagi H (2011) Evidence for osteocyte regulation of bone homeostasis through RANKL expression. Nat Med 17:1231
Xiong J, Piemontese M, Thostenson JD, Weinstein RS, Manolagas SC, O'Brien CA (2014) Osteocyte-derived RANKL is a critical mediator of the increased bone resorption caused by dietary calcium deficiency. Bone 66:146–154
Takayanagi H, Kim S, Koga T, Nishina H, Isshiki M, Yoshida H, Saiura A, Isobe M, Yokochi T, Inoue J-i, Wagner EF, Mak TW, Kodama T, Taniguchi T (2002) Induction and activation of the transcription factor NFATc1 (NFAT2) integrate RANKL signaling in terminal differentiation of osteoclasts. Dev Cell 3(6):889–901
Arron JR, Choi Y (2000) Bone versus immune system. Nature 408:535
Takayanagi H, Ogasawara K, Hida S, Chiba T, Murata S, Sato K, Takaoka A, Yokochi T, Oda H, Tanaka K, Nakamura K, Taniguchi T (2000) T-cell-mediated regulation of osteoclastogenesis by signalling cross-talk between RANKL and IFN-γ. Nature 408:600
Kim N, Odgren PR, Kim DK, Marks SC Jr, Choi Y (2000) Diverse roles of the tumor necrosis factor family member TRANCE in skeletal physiology revealed by TRANCE deficiency and partial rescue by a lymphocyte-expressed TRANCE transgene. Proc Natl Acad Sci U S A 97(20):10905–10910
Walsh MC, Kim N, Kadono Y, Rho J, Lee SY, Lorenzo J, Choi Y (2006) Osteoimmunology: interplay between the immune system and bone metabolism. Annu Rev Immunol 24(1):33–63
S.R. Goldring, Pathogenesis of bone and cartilage destruction in rheumatoid arthritis, Rheumatology 42(suppl_2) (2003) ii11-ii16
Walsh NC, Gravallese EM (2004) Bone loss in inflammatory arthritis: mechanisms and treatment strategies. Curr Opin Rheumatol 16(4):419–427
Lorenzo J, Horowitz M, Choi Y (2008) Osteoimmunology: interactions of the bone and immune system. Endocr Rev 29(4):403–440
Varol C, Mildner A, Jung S (2015) Macrophages: Development and tissue specialization. Annu Rev Immunol 33(1):643–675
Chang MK, Raggatt L-J, Alexander KA, Kuliwaba JS, Fazzalari NL, Schroder K, Maylin ER, Ripoll VM, Hume DA, Pettit AR (2008) Osteal tissue macrophages are intercalated throughout human and mouse bone lining tissues and regulate osteoblast function in vitro and in vivo. J Immunol 181(2):1232–1244
Sinder BP, Pettit AR, McCauley LK (2015) Macrophages: their emerging roles in bone. J Bone Miner Res 30(12):2140–2149
Nicolaidou V, Wong MM, Redpath AN, Ersek A, Baban DF, Williams LM, Cope AP, Horwood NJ (2012) Monocytes induce STAT3 activation in human mesenchymal stem cells to promote osteoblast formation. PLoS One 7(7):e39871
Athanasou NA, Heryet A, Quinn J, Gatter KC, Mason DY, McGe JOD (1986) Osteoclasts contain macrophage and megakaryocyte antigens. J Pathol 150(4):239–246
Athanasou NA, Quinn J (1990) Immunophenotypic differences between osteoclasts and macrophage polykaryons: immunohistological distinction and implications for osteoclast ontogeny and function. J Clin Pathol 43(12):997–1003
Kersten S, Desvergne B, Wahli W (2000) Roles of PPARs in health and disease. Nature 405:421
Mangelsdorf DJ, Thummel C, Beato M, Herrlich P, Schütz G, Umesono K, Blumberg B, Kastner P, Mark M, Chambon P, Evans RM (1995) The nuclear receptor superfamily: the second decade. Cell 83(6):835–839
T. Lemberger, BD and, W. Wahli, Peroxisome proliferator-activated receptors: a nuclear receptor signaling pathway in lipid physiology, Annu Rev Cell Dev Biol 12(1) (1996) 335–363
Lehmann JM, Moore LB, Smith-Oliver TA, Wilkison WO, Willson TM, Kliewer SA (1995) An antidiabetic thiazolidinedione is a high affinity ligand for peroxisome proliferator-activated receptor γ (PPARγ). J Biol Chem 270(22):12953–12956
Kliewer SA, Lenhard JM, Willson TM, Patel I, Morris DC, Lehmann JM (1995) A prostaglandin J2 metabolite binds peroxisome proliferator-activated receptor γ and promotes adipocyte differentiation. Cell 83(5):813–819
Rosen ED, Sarraf P, Troy AE, Bradwin G, Moore K, Milstone DS, Spiegelman BM, Mortensen RM (1999) PPARγ is required for the differentiation of adipose tissue in vivo and in vitro. Mol Cell 4(4):611–617
Ricote M, Li AC, Willson TM, Kelly CJ, Glass CK (1998) The peroxisome proliferator-activated receptor-γ is a negative regulator of macrophage activation. Nature 391:79
Rangwala SM, Lazar MA (2004) Peroxisome proliferator-activated receptor γ in diabetes and metabolism. Trends Pharmacol Sci 25(6):331–336
Moore KJ, Rosen ED, Fitzgerald ML, Randow F, Andersson LP, Altshuler D, Milstone DS, Mortensen RM, Spiegelman BM, Freeman MW (2001) The role of PPAR-γ in macrophage differentiation and cholesterol uptake. Nat Med 7:41
Tontonoz P, Nagy L, Alvarez JGA, Thomazy VA, Evans RM (1998) PPARγ promotes monocyte/macrophage differentiation and uptake of oxidized LDL. Cell 93(2):241–252
Heming M, Gran S, Jauch S-L, Fischer-Riepe L, Russo A, Klotz L, Hermann S, Schäfers M, Roth J, Barczyk-Kahlert K (2018) Peroxisome proliferator-activated receptor-γ modulates the response of macrophages to lipopolysaccharide and glucocorticoids. Front Immunol 9(893)
Chawla A, Barak Y, Nagy L, Liao D, Tontonoz P, Evans RM (2001) PPAR-γ dependent and independent effects on macrophage-gene expression in lipid metabolism and inflammation. Nat Med 7:48
Schneider C, Nobs SP, Kurrer M, Rehrauer H, Thiele C, Kopf M (2014) Induction of the nuclear receptor PPAR-γ by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat Immunol 15:1026
Jiang C, Ting AT, Seed B (1998) PPAR-γ agonists inhibit production of monocyte inflammatory cytokines. Nature 391:82
Wan Y, Evans RM (2010) Rosiglitazone activation of PPARγ suppresses fractalkine signaling. J Mol Endocrinol 44(2):135–142
Wu L, Yan C, Czader M, Foreman O, Blum JS, Kapur R, Du H (2012) Inhibition of PPARγ in myeloid-lineage cells induces systemic inflammation, immunosuppression, and tumorigenesis. Blood 119(1):115–126
Odegaard JI, Ricardo-Gonzalez RR, Goforth MH, Morel CR, Subramanian V, Mukundan L, Eagle AR, Vats D, Brombacher F, Ferrante AW, Chawla A (2007) Macrophage-specific PPARγ controls alternative activation and improves insulin resistance. Nature 447:1116
Bouhlel MA, Derudas B, Rigamonti E, Dièvart R, Brozek J, Haulon S, Zawadzki C, Jude B, Torpier G, Marx N, Staels B, Chinetti-Gbaguidi G (2007) PPARγ activation primes human monocytes into alternative M2 macrophages with anti-inflammatory properties. Cell Metab 6(2):137–143
Chinetti G, Griglio S, Antonucci M, Torra IP, Delerive P, Majd Z, Fruchart J-C, Chapman J, Najib J, Staels B (1998) Activation of proliferator-activated receptors α and γ induces apoptosis of human monocyte-derived macrophages. J Biol Chem 273(40):25573–25580
Bodles AM, Varma V, Yao-Borengasser A, Phanavanh B, Peterson CA, McGehee RE, Rasouli N, Wabitsch M, Kern PA (2006) Pioglitazone induces apoptosis of macrophages in human adipose tissue. J Lipid Res 47(9):2080–2088
Saltiel AR, Olefsky JM (1996) Thiazolidinediones in the treatment of insulin resistance and type ii diabetes. Diabetes 45(12):1661–1669
Soccio RE, Chen ER, Lazar MA (2014) Thiazolidinediones and the promise of insulin sensitization in type 2 diabetes. Cell Metab 20(4):573–591
Grey A, Horne A, Wattie D, Reid IR, Bolland M, Gamble G, Davidson J (2007) The peroxisome proliferator-activated receptor-γ agonist rosiglitazone decreases bone formation and bone mineral density in healthy postmenopausal women: a randomized, controlled trial. J Clin Endocrinol Metab 92(4):1305–1310
Kahn SE, Haffner SM, Heise MA, Herman WH, Holman RR, Jones NP, Kravitz BG, Lachin JM, O'Neill MC, Zinman B, Viberti G (2006) Glycemic durability of rosiglitazone, metformin, or glyburide monotherapy. N Engl J Med 355(23):2427–2443
Kahn SE, Zinman B, Lachin JM, Haffner SM, Herman WH, Holman RR, Kravitz BG, Yu DH, Heise MA, Aftring RP, Viberti G, for the A Diabetes Outcome Progression Trial (ADOPT) study group (2008) Rosiglitazone-Associated Fractures in Type 2 Diabetes An analysis from A Diabetes Outcome Progression Trial (ADOPT). Diabetes Care 31(5):845–851
Rzonca SO, Lecka-Czernik B, Gaddy D, Montague DC, Suva LJ (2004) Bone is a target for the antidiabetic compound rosiglitazone. Endocrinology 145(1):401–406
Sottile V, Seuwen K, Kneissel M (2004) Enhanced marrow adipogenesis and bone resorption in estrogen-deprived rats treated with the PPARgamma agonist BRL49653 (rosiglitazone). Calcif Tissue Int 75(4):329–337
Wan Y, Chong L-W, Evans RM (2007) PPAR-γ regulates osteoclastogenesis in mice. Nat Med 13:1496
Grigoriadis A, Wang Z, Cecchini M, Hofstetter W, Felix R, Fleisch H, Wagner E (1994) c-Fos: a key regulator of osteoclast-macrophage lineage determination and bone remodeling. Science 266(5184):443–448
Wei W, Zeve D, Wang X, Du Y, Tang W, Dechow PC, Graff JM, Wan Y (2011) Osteoclast progenitors reside in the peroxisome proliferator-activated receptor γ-expressing bone marrow cell population. Mol Cell Biol 31(23):4692–4705
Akune T, Ohba S, Kamekura S, Yamaguchi M, Chung U-i, Kubota N, Terauchi Y, Harada Y, Azuma Y, Nakamura K, Kadowaki T, Kawaguchi H (2004) PPAR γ insufficiency enhances osteogenesis through osteoblast formation from bone marrow progenitors. J Clin Invest 113(6):846–855
Lazarenko OP, Rzonca SO, Lecka-Czernik B, Swain FL, Suva LJ, Hogue WR (2007) Rosiglitazone induces decreases in bone mass and strength that are reminiscent of aged bone. Endocrinology 148(6):2669–2680
Wan Y (2010) PPARγ in bone homeostasis. Trends Endocrinol Metab 21(12):722–728
Stechschulte LA, Czernik PJ, Rotter ZC, Tausif FN, Corzo CA, Marciano DP, Asteian A, Zheng J, Bruning JB, Kamenecka TM, Rosen CJ, Griffin PR, Lecka-Czernik B (2016) PPARG post-translational modifications regulate bone formation and bone resorption. EBioMedicine 10:174–184
Choi S, Chung S, Park K (2018) Re-highlighting the action of PPAR? in treating metabolic diseases [version 1; referees: 2 approved]. F1000Research 7(1127)
Wei W, Wang X, Yang M, Smith LC, Dechow PC, Sonoda J, Evans RM, Wan Y (2010) PGC1β mediates pparγ activation of osteoclastogenesis and rosiglitazone-induced bone loss. Cell Metab 12(2):202
Wei W, Schwaid AG, Wang X, Wang X, Chen S, Chu Q, Saghatelian A, Wan Y (2016) Ligand activation of ERRα by cholesterol mediates statin and bisphosphonate effects. Cell Metab 23(3):479–491
Giguère V, Yang N, Segui P, Evans RM (1988) Identification of a new class of steroid hormone receptors. Nature 331(6151):91–94
Hong H, Yang L, Stallcup MR (1999) Hormone-independent transcriptional activation and coactivator binding by novel orphan nuclear receptor ERR3. J Biol Chem 274(32):22618–22626
Heard DJ, Vissing H, Norby PL, Holloway J (2000) Human ERRγ, a third member of the estrogen receptor-related receptor (ERR) subfamily of orphan nuclear receptors: tissue-specific isoforms are expressed during development and in the adult. Mol Endocrinol 14(3):382–392
Giguére V (2002) To ERR in the estrogen pathway. Trends Endocrinol Metab 13(5):220–225
Kallen J, Schlaeppi J-M, Bitsch F, Filipuzzi I, Schilb A, Riou V, Graham A, Strauss A, Geiser M, Fournier B (2004) Evidence for ligand-independent transcriptional activation of the human estrogen-related receptor α (ERRα): crystal structure of ERRα ligand binding domain in complex with peroxisome proliferator-activated receptor coactivator-1α. J Biol Chem 279(47):49330–49337
Bonnelye E, Vanacker JM, Spruyt N, Alric S, Fournier B, Desbiens X, Laudet V (1997) Expression of the estrogen-related receptor 1 (ERR-1) orphan receptor during mouse development. Mech Dev 65(1):71–85
Giguère V (2008) Transcriptional control of energy homeostasis by the estrogen-related receptors. Endocr Rev 29(6):677–696
Audet-walsh É, Giguére V (2014) The multiple universes of estrogen-related receptor α and γ in metabolic control and related diseases. Acta Pharmacol Sin 36:51
Luo J, Sladek R, Carrier J, Bader J-A, Richard D, Giguère V (2003) Reduced fat mass in mice lacking orphan nuclear receptor estrogen-related receptor α. Mol Cell Biol 23(22):7947–7956
Huss JM, Imahashi K-i, Dufour CR, Weinheimer CJ, Courtois M, Kovacs A, Giguère V, Murphy E, Kelly DP (2007) The nuclear receptor ERRα is required for the bioenergetic and functional adaptation to cardiac pressure overload. Cell Metab 6(1):25–37
Vanacker JM, Pettersson K, Gustafsson JÅ, Laudet V (1999) Transcriptional targets shared by estrogen receptor-related receptors (ERRs) and estrogen receptor (ER) α, but not by ERβ. EMBO J 18(15):4270–4279
Sladek R, Bader JA, Giguère V (1997) The orphan nuclear receptor estrogen-related receptor alpha is a transcriptional regulator of the human medium-chain acyl coenzyme A dehydrogenase gene. Mol Cell Biol 17(9):5400–5409
Vega R, Kelly D (1997) A role for estrogen-related receptor α in the control of mitochondrial fatty acid β-oxidation during brown adipocyte differentiation. J Biol Chem 272:31693–31699
Huss JM, Kopp RP, Kelly DP (2002) Peroxisome proliferator-activated receptor coactivator-1α (PGC-1α) coactivates the cardiac-enriched nuclear receptors estrogen-related receptor-α and -γ: identification of novel leucine-rich interaction motif within PGC-1α. J Biol Chem 277(43):40265–40274
Kamei Y, Ohizumi H, Fujitani Y, Nemoto T, Tanaka T, Takahashi N, Kawada T, Miyoshi M, Ezaki O, Kakizuka A (2003) PPARγ coactivator 1β/ERR ligand 1 is an ERR protein ligand, whose expression induces a high-energy expenditure and antagonizes obesity. Proc Natl Acad Sci 100(21):12378–12383
Schreiber SN, Knutti D, Brogli K, Uhlmann T, Kralli A (2003) The transcriptional coactivator PGC-1 regulates the expression and activity of the orphan nuclear receptor estrogen-related receptor α (ERRα). J Biol Chem 278(11):9013–9018
Wende AR, Huss JM, Schaeffer PJ, Giguère V, Kelly DP (2005) PGC-1α coactivates PDK4 gene expression via the orphan nuclear receptor ERRα: a mechanism for transcriptional control of muscle glucose metabolism. Mol Cell Biol 25(24):10684–10694
Sonoda J, Laganière J, Mehl IR, Barish GD, Chong L-W, Li X, Scheffler IE, Mock DC, Bataille AR, Robert F (2007) Nuclear receptor ERRα and coactivator PGC-1β are effectors of IFN-γ-induced host defense. Genes Dev 21(15):1909–1920
Yuk J-M, Kim TS, Kim SY, Lee H-M, Han J, Dufour CR, Kim JK, Jin HS, Yang C-S, Park K-S, Lee C-H, Kim J-M, Kweon GR, Choi H-S, Vanacker J-M, Moore DD, Giguère V, Jo E-K (2015) Orphan nuclear receptor ERRα controls macrophage metabolic signaling and A20 expression to negatively regulate TLR-induced inflammation. Immunity 43(1):80–91
He X, Ma S, Tian Y, Wei C, Zhu Y, Li F, Zhang P, Wang P, Zhang Y, Zhong H (2017) ERRα negatively regulates type I interferon induction by inhibiting TBK1-IRF3 interaction. PLoS Pathog 13(6):e1006347
Kim SY, Yang C-S, Lee H-M, Kim JK, Kim Y-S, Kim Y-R, Kim J-S, Kim TS, Yuk J-M, Dufour CR, Lee S-H, Kim J-M, Choi H-S, Giguère V, Jo E-K (2018) ESRRA (estrogen-related receptor α) is a key coordinator of transcriptional and post-translational activation of autophagy to promote innate host defense. Autophagy 14(1):152–168
Bae S, Lee MJ, Mun SH, Giannopoulou EG, Yong-Gonzalez V, Cross JR, Murata K, Giguère V, van der Meulen M, Park-Min K-H (2017) MYC-dependent oxidative metabolism regulates osteoclastogenesis via nuclear receptor ERRα. J Clin Invest 127(7):2555–2568
Ocampo CB, Downes M, Yu RT, Evans RM, Barish GD, Alaynick WA, Bookout AL, Mangelsdorf DJ (2005) A nuclear receptor atlas: macrophage activation. Mol Endocrinol 19(10):2466–2477
Bonnelye E, Kung V, Aubin JE, Laplace C, Galson DL (2002) Estrogen receptor-related receptor α impinges on the estrogen axis in bone: potential function in osteoporosis. Endocrinology 143(9):3658–3670
Bonnelye E, Saltel F, Chabadel A, Zirngibl RA, Aubin JE, Jurdic P (2010) Involvement of the orphan nuclear estrogen receptor-related receptor α in osteoclast adhesion and transmigration. J Mol Endocrinol 45(6):365–377
Studer A, Morvan F, Evans G, Delhon I, Wyder L, Kneissel M, Gutzwiller S, Fournier B, Rangwala S (2009) Absence of estrogen receptor-related-α increases osteoblastic differentiation and cancellous bone mineral density. Endocrinology 150(10):4463–4472
Teyssier C, Gallet M, Rabier B, Monfoulet L, Dine J, Macari C, Espallergues J, Horard B, Giguère V, Cohen-Solal M, Chassande O, Vanacker J-M (2009) Absence of ERRα in female mice confers resistance to bone loss induced by age or estrogen-deficiency. PLoS One 4(11):e7942
Lin J, Puigserver P, Donovan J, Tarr P, Spiegelman BM (2002) Peroxisome proliferator-activated receptor γ coactivator 1β (PGC-1β), a novel PGC-1-related transcription coactivator associated with host cell factor. J Biol Chem 277(3):1645–1648
Lin J, Tarr PT, Yang R, Rhee J, Puigserver P, Newgard CB, Spiegelman BM (2003) PGC-1β in the regulation of hepatic glucose and energy metabolism. J Biol Chem 278(33):30843–30848
St-Pierre J, Lin J, Krauss S, Tarr PT, Yang R, Newgard CB, Spiegelman BM (2003) Bioenergetic analysis of peroxisome proliferator-activated receptor γ coactivators 1α and 1β (PGC-1α and PGC-1β) in muscle cells. J Biol Chem 278(29):26597–26603
Sonoda J, Mehl IR, Chong L-W, Nofsinger RR, Evans RM (2007) PGC-1β controls mitochondrial metabolism to modulate circadian activity, adaptive thermogenesis, and hepatic steatosis. Proc Natl Acad Sci 104(12):5223–5228
Lai L, Leone TC, Zechner C, Schaeffer PJ, Kelly SM, Flanagan DP, Medeiros DM, Kovacs A, Kelly DP (2008) Transcriptional coactivators PGC-1α and PGC-lβ control overlapping programs required for perinatal maturation of the heart. Genes Dev 22(14):1948–1961
Vianna CR, Huntgeburth M, Coppari R, Choi CS, Lin J, Krauss S, Barbatelli G, Tzameli I, Kim Y-B, Cinti S, Shulman GI, Spiegelman BM, Lowell BB (2006) Hypomorphic mutation of PGC-1β causes mitochondrial dysfunction and liver insulin resistance. Cell Metab 4(6):453–464
Lelliott CJ, Medina-Gomez G, Petrovic N, Kis A, Feldmann HM, Bjursell M, Parker N, Curtis K, Campbell M, Hu P, Zhang D, Litwin SE, Zaha VG, Fountain KT, Boudina S, Jimenez-Linan M, Blount M, Lopez M, Meirhaeghe A, Bohlooly-Y M, Storlien L, Strömstedt M, Snaith M, Orešič M, Abel ED, Cannon B, Vidal-Puig A (2006) Ablation of PGC-1β results in defective mitochondrial activity, thermogenesis, hepatic function, and cardiac performance. PLoS Biol 4(11):e369
Gali Ramamoorthy T, Laverny G, Schlagowski A-I, Zoll J, Messaddeq N, Bornert J-M, Panza S, Ferry A, Geny B, Metzger D (2015) The transcriptional coregulator PGC-1β controls mitochondrial function and anti-oxidant defence in skeletal muscles. Nat Commun 6:10210
Arany Z, Lebrasseur N, Morris C, Smith E, Yang W, Ma Y, Chin S, Spiegelman BM (2007) The transcriptional coactivator PGC-1β drives the formation of oxidative type IIX fibers in skeletal muscle. Cell Metab 5(1):35–46
Bellafante E, Murzilli S, Salvatore L, Latorre D, Villani G, Moschetta A (2013) Hepatic-specific activation of peroxisome proliferator-activated receptor γ coactivator-1β protects against steatohepatitis. Hepatology 57(4):1343–1356
Enguix N, Pardo R, González A, López VM, Simó R, Kralli A, Villena JA (2013) Mice lacking PGC-1β in adipose tissues reveal a dissociation between mitochondrial dysfunction and insulin resistance. Mol Metab 2(3):215–226
Chambers KT, Chen Z, Crawford PA, Fu X, Burgess SC, Lai L, Leone TC, Kelly DP, Finck BN (2012) Liver-specific PGC-1beta deficiency leads to impaired mitochondrial function and lipogenic response to fasting-refeeding. PLoS One 7(12):e52645
Vats D, Mukundan L, Odegaard JI, Zhang L, Smith KL, Morel CR, Greaves DR, Murray PJ, Chawla A (2006) Oxidative metabolism and PGC-1β attenuate macrophage-mediated inflammation. Cell Metab 4(1):13–24
Van den Bossche J, Baardman J, Otto NA, van der Velden S, Neele AE, van den Berg SM, Luque-Martin R, Chen H-J, Boshuizen MCS, Ahmed M, Hoeksema MA, de Vos AF, de Winther MPJ (2016) Mitochondrial dysfunction prevents repolarization of inflammatory macrophages. Cell Rep 17(3):684–696
Byles V, Covarrubias AJ, Ben-Sahra I, Lamming DW, Sabatini DM, Manning BD, Horng T (2013) The TSC-mTOR pathway regulates macrophage polarization. Nat Commun 4:2834
Chen H, Liu Y, Li D, Song J, Xia M (2016) PGC-1β suppresses saturated fatty acid-induced macrophage inflammation by inhibiting TAK1 activation. IUBMB Life 68(2):145–155
Eisele PS, Furrer R, Beer M, Handschin C (2015) The PGC-1 coactivators promote an anti-inflammatory environment in skeletal muscle in vivo. Biochem Biophys Res Commun 464(3):692–697
Ishii K-a, Fumoto T, Iwai K, Takeshita S, Ito M, Shimohata N, Aburatani H, Taketani S, Lelliott CJ, Vidal-Puig A, Ikeda K (2009) Coordination of PGC-1β and iron uptake in mitochondrial biogenesis and osteoclast activation. Nat Med 15:259
Carroll J, Fearnley IM, Shannon RJ, Hirst J, Walker JE (2003) Analysis of the subunit composition of complex I from bovine heart mitochondria. Mol Cell Proteomics 2(2):117–126
Carroll J, Fearnley IM, Skehel JM, Shannon RJ, Hirst J, Walker JE (2006) Bovine complex I is a complex of 45 different subunits. J Biol Chem 281(43):32724–32727
D.M. Kirby, M. Crawford, M.A. Cleary, H.-H.M. Dahl, X. Dennett, D.R. Thorburn, Respiratory chain complex I deficiency: an underdiagnosed energy generation disorder 52(6) (1999) 1255–1255
Mimaki M, Wang X, McKenzie M, Thorburn DR, Ryan MT (2012) Understanding mitochondrial complex I assembly in health and disease. Biochim Biophys Acta, Bioenerg 1817(6):851–862
Finsterer J (2006) Central nervous system manifestations of mitochondrial disorders. Acta Neurol Scand 114(4):217–238
Coskun P, Wyrembak J, Schriner SE, Chen H-W, Marciniack C, LaFerla F, Wallace DC (2012) A mitochondrial etiology of Alzheimer and Parkinson disease. Biochim Biophys Acta Gen Subj 1820(5):553–564
Kruse SE, Watt WC, Marcinek DJ, Kapur RP, Schenkman KA, Palmiter RD (2008) Mice with mitochondrial complex I deficiency develop a fatal encephalomyopathy. Cell Metab 7(4):312–320
Scacco S, Petruzzella V, Budde S, Vergari R, Tamborra R, Panelli D, van den Heuvel LP, Smeitink JA, Papa S (2003) Pathological mutations of the human NDUFS4 gene of the 18-kDa (AQDQ) subunit of complex I affect the expression of the protein and the assembly and function of the complex. J Biol Chem 278(45):44161–44167
Petruzzella V, Papa S (2002) Mutations in human nuclear genes encoding for subunits of mitochondrial respiratory complex I: the NDUFS4 gene. Gene 286(1):149–154
Budde SMS, van den Heuvel LPWJ, Smeets RJP, Skladal D, Mayr JA, Boelen C, Petruzzella V, Papa S, Smeitink JAM (2003) Clinical heterogeneity in patients with mutations in the NDUFS4 gene of mitochondrial complex I. J Inherit Metab Dis 26(8):813–815
Ugalde C, Smeitink JAM, van den Heuvel LP, Nijtmans LGJ, Janssen RJRJ (2004) Differences in assembly or stability of complex I and other mitochondrial OXPHOS complexes in inherited complex I deficiency. Hum Mol Genet 13(6):659–667
van den Heuvel L, Ruitenbeek W, Smeets R, Gelman-Kohan Z, Elpeleg O, Loeffen J, Trijbels F, Mariman E, de Bruijn D, Smeitink J (1998) Demonstration of a new pathogenic mutation in human complex I deficiency: a 5-bp duplication in the nuclear gene encoding the 18-kD (AQDQ) subunit. Am J Hum Genet 62(2):262–268
Iuso A, Scacco S, Piccoli C, Bellomo F, Petruzzella V, Trentadue R, Minuto M, Ripoli M, Capitanio N, Zeviani M, Papa S (2006) Dysfunctions of cellular oxidative metabolism in patients with mutations in the NDUFS1 and NDUFS4 genes of complex I. J Biol Chem 281(15):10374–10380
Quintana A, Kruse SE, Kapur RP, Sanz E, Palmiter RD (2010) Complex I deficiency due to loss of Ndufs4 in the brain results in progressive encephalopathy resembling Leigh syndrome. Proc Natl Acad Sci 107(24):10996–11001
Jin Z, Wei W, Yang M, Du Y, Wan Y (2014) Mitochondrial complex I activity suppresses inflammation and enhances bone resorption by shifting macrophage-osteoclast polarization. Cell Metab 20(3):483–498
Oka K, Ishimura-Oka K, Chu M-j, Sullivan M, Krushkal J, Li W-H, Chan L (1994) Mouse very-low-density-lipoprotein receptor (VLDLR) cDNA cloning, tissue-specific expression and evolutionary relationship with the low-density-lipoprotein receptor. Eur J Biochem 224(3):975–982
Webb JC, Patel DD, Jones MD, Knight BL, Soutar AK (1994) Characterization and tissue-specific expression of the human ‘very low density lipoprotein (VLDL) receptor’mRNA. Hum Mol Genet 3(4):531–537
Tacken PJ, Hofker MH, Havekes LM, van Dijk KW (2001) Living up to a name: the role of the VLDL receptor in lipid metabolism. Curr Opin Lipidol 12(3):275–279
Takahashi S, Sakai J, Fujino T, Hattori H, Zenimaru Y, Suzuki J, Miyamori I, Yamamoto TT (2004) The very low-density lipoprotein (VLDL) receptor: characterization and functions as a peripheral lipoprotein receptor. J Atheroscler Thromb 11(4):200–208
Trommsdorff M, Gotthardt M, Hiesberger T, Shelton J, Stockinger W, Nimpf J, Hammer RE, Richardson JA, Herz J (1999) Reeler/disabled-like disruption of neuronal migration in knockout mice lacking the VLDL receptor and ApoE receptor 2. Cell 97(6):689–701
D'Arcangelo G, Homayouni R, Keshvara L, Rice DS, Sheldon M, Curran T (1999) Reelin is a ligand for lipoprotein receptors. Neuron 24(2):471–479
Goudriaan JR, Tacken PJ, Dahlmans VE, Gijbels MJ, van Dijk KW, Havekes LM, Jong MC (2001) Protection from obesity in mice lacking the VLDL receptor. Arterioscler Thromb Vasc Biol 21(9):1488–1493
Nguyen A, Tao H, Metrione M, Hajri T (2014) Very low density lipoprotein receptor (VLDLR) expression is a determinant factor in adipose tissue inflammation and adipocyte-macrophage interaction. J Biol Chem 289(3):1688–1703
Eck MV, Oost J, Goudriaan JR, Hoekstra M, Hildebrand RB, Bos IST, van Dijk KW, Van Berkel TJC (2005) Role of the macrophage very-low-density lipoprotein receptor in atherosclerotic lesion development. Atherosclerosis 183(2):230–237
Frykman PK, Brown MS, Yamamoto T, Goldstein JL, Herz J (1995) Normal plasma lipoproteins and fertility in gene-targeted mice homozygous for a disruption in the gene encoding very low density lipoprotein receptor. Proc Natl Acad Sci 92(18):8453–8457
Miyamoto T, Arai F, Ohneda O, Takagi K, Anderson DM, Suda T (2000) An adherent condition is required for formation of multinuclear osteoclasts in the presence of macrophage colony-stimulating factor and receptor activator of nuclear factor κB ligand. Blood 96(13):4335–4343
Du Y, Yang M, Wei W, Huynh HD, Herz J, Saghatelian A, Wan Y (2012) Macrophage VLDL receptor promotes PAFAH secretion in mother’s milk and suppresses systemic inflammation in nursing neonates. Nat Commun 3:1008–1008
Huynh H, Wei W, Wan Y (2017) mTOR inhibition subdues milk disorder caused by maternal VLDLR loss. Cell Rep 19(10):2014–2025
Yang D, Huynh H, Wan Y (2018) Milk lipid regulation at the maternal-offspring interface. Semin Cell Dev Biol 81:141–148
Caplan MS, Sun X-M, Hsueh W, Hageman JR (1990) Role of platelet activating factor and tumor necrosis factor-alpha in neonatal necrotizing enterocolitis. J Pediatr 116(6):960–964
Walker A (2010) Breast milk as the gold standard for protective nutrients. J Pediatr 156(2, Supplement):S3–S7
Furukawa M, Narahara H, Yasuda K, Johnston JM (1993) Presence of platelet-activating factor-acetylhydrolase in milk. J Lipid Res 34(9):1603–1609
Moya FR, Eguchi H, Zhao B, Furukawa M, Sfeir J, Osorio M, Ogawa Y, Johnston JM (1994) Platelet-activating factor acetylhydrolase in term and preterm human milk: a preliminary report. J Pediatr Gastroenterol Nutr 19(2):236–239
Shin KC, Hwang I, Choe SS, Park J, Ji Y, Kim JI, Lee GY, Choi SH, Ching J, Kovalik J-P, Kim JB (2017) Macrophage VLDLR mediates obesity-induced insulin resistance with adipose tissue inflammation. Nat Commun 8(1):1087
Huang SC-C, Smith AM, Everts B, Colonna M, Pearce EL, Schilling JD, Pearce EJ (2016) Metabolic reprogramming mediated by the mTORC2-IRF4 signaling axis is essential for macrophage alternative activation. Immunity 45(4):817–830
Mills EL, Kelly B, Logan A, Costa ASH, Varma M, Bryant CE, Tourlomousis P, Däbritz JHM, Gottlieb E, Latorre I, Corr SC, McManus G, Ryan D, Jacobs HT, Szibor M, Xavier RJ, Braun T, Frezza C, Murphy MP, O’Neill LA (2016) Succinate dehydrogenase supports metabolic repurposing of mitochondria to drive inflammatory macrophages. Cell 167(2):457–470.e13
Acknowledgments
Y.W. is Lawrence Raisz Professor in Bone Cell Metabolism and a Virginia Murchison Linthicum Scholar in Medical Research.
Funding
This work was in part supported by NIH (R01CA229487, R01CA236802, Y.W.), DOD (W81XWH-18-1-0014, Y.W.), CPRIT (RP180047, Y.W.), The Welch Foundation (I-1751, Y.W.), and UT Southwestern Endowed Scholar Startup Fund (Y.W.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This article is a contribution to the special issue on Osteoimmunology - Guest Editor: Mary Nakamura
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yang, D., Wan, Y. Molecular determinants for the polarization of macrophage and osteoclast. Semin Immunopathol 41, 551–563 (2019). https://doi.org/10.1007/s00281-019-00754-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00281-019-00754-3