Abstract
Purpose
Microvascular invasion (MVI) is a common complication of hepatocellular carcinoma (HCC) surgery, which is an important predictor of reduced surgical prognosis. This study aimed to develop a fully automated diagnostic model to predict pre-surgical MVI based on four-phase dynamic CT images.
Methods
A total of 140 patients with HCC from two centers were retrospectively included (training set, n = 98; testing set, n = 42). All CT phases were aligned to the portal venous phase, and were then used to train a deep-learning model for liver tumor segmentation. Radiomics features were extracted from the tumor areas of original CT phases and pairwise subtraction images, as well as peritumoral features. Lastly, linear discriminant analysis (LDA) models were trained based on clinical features, radiomics features, and hybrid features, respectively. Models were evaluated by area under curve (AUC), accuracy, sensitivity, specificity, positive and negative predictive values (PPV and NPV).
Results
Overall, 86 and 54 patients with MVI− (age, 55.92 ± 9.62 years; 68 men) and MVI+ (age, 53.59 ± 11.47 years; 43 men) were included. Average dice coefficients of liver tumor segmentation were 0.89 and 0.82 in training and testing sets, respectively. The model based on radiomics (AUC = 0.865, 95% CI: 0.725–0.951) showed slightly better performance than that based on clinical features (AUC = 0.841, 95% CI: 0.696–0.936). The classification model based on hybrid features achieved better performance in both training (AUC = 0.955, 95% CI: 0.893–0.987) and testing sets (AUC = 0.913, 95% CI: 0.785–0.978), compared with models based on clinical and radiomics features (p-value < 0.05). Moreover, the hybrid model also provided the best accuracy (0.857), sensitivity (0.875), and NPV (0.917).
Conclusion
The classification model based on multimodal intra- and peri-tumoral radiomics features can well predict HCC patients with MVI.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC) is the third leading cause of cancer death [1], accounting for 90% of primary liver cancers in the world. Prognosis of HCC surgery is usually very poor, and about 50% - 70% of HCC patients are recurrent within 5 years after hepatectomy [2, 3]. Microvascular invasion (MVI) is found in 30–50% of HCC pathologies, which is regarded as a major risk factor for postsurgical recurrence [4, 5], even in patients with isolated small HCC [6]. Therefore, clinical guidelines suggest a large surgical extent for MVI-positive patients [7]. Diagnosis of MVI is thus critical in presurgical evaluation of HCC patients [8].
MVI is a nest of cancer cells within the endothelium of blood vessels, which is visible only in microscopy images [9]. Noninvasive methods have been used to predict MVI, including alpha-fetoprotein (AFP), tumor size , and two-trait predictor of venous invasion bicharacteristic predictors of venous invasion (TTPVI) [10]. However, these predictors are based on low-order features which cannot capture a full view of tumors. Radiomics technologies have been developed to extract high-throughput quantitative image features, and improve accuracy of diagnosis and prognosis in cancer studies [11, 12]. However, radiomics for live tumors usually depends on manual segmentation due to poor output of automatic segmentation [13]. The manual segmentation is time-consuming and usually inconsistent across studies. Moreover, current radiomics studies of liver cancer depend on enhanced images of certain phase, and thus ignoring the information of tumor changes in the temporal dimension [14].
To address the abovementioned issues, this study is aimed to develop an automated MVI prediction model based on 4-phase CT of patients with HCC [15, 16]. In particular, dynamic features were extracted between CT phases. The model is then built based on a fusion of clinical features, dynamic features, peritumoral and intratumoral radiomics features to improve prediction of MVI.
Materials and methods
This multi-center cross-sectional study was reviewed and approved by the Biomedical Research Ethics Committee of our hospital without patient consent. Patient data has been anonymized in this report.
Subjects
We retrospectively reviewed two datasets of HCC patients who underwent CT examination from May 2012 to July 2017 at Center 1 (***), and from January 2014 to April 2019 at Center 2 (***) (Fig. 1). The inclusion criteria were as follows: (1) pathological diagnosis as HCC, (2) available, and (3) non-contrast and contrast-enhanced abdomen CT examination within 8 weeks before hepatectomy. The exclusion criteria were as follows: (1) preoperative treatment for HCC, (2) macroscopic vascular invasion or extrahepatic, (3) image artifacts, and (4) poor registration quality. At last, a total of 81 patients from Center 1, and 59 patients from Center 2 were included for the following analysis.
CT acquisition
All patients underwent unenhanced and contrast-enhanced multiphasic abdominal CT examinations using one of the following systems: GE Revolution CT, TOSHIBA Aquilion ONE, SIEMENS SOMATOM Definition, SIEMENS SOMATOM Definition Flash, UIH uCT 510 or UIH uCT 760. Scanning parameters were as follows: tube voltage of 100 or 120 kVp; tube current of 60-380 mAs; the field of view = 350 × 350 mm; matrix = 512 × 512; slice thickness = 0.625 mm. A 1.3–1.5 mL/kg body weight bolus of contrast material iodixanol (Ultravist 300 or Ultravist 370, Bayer, Germany) and ioversol (Optiray 320 or Optiray 350, Guerbet, France) was injected intravenously at a flow rate of 3.0–4.0 mL/sec. Arterial phase scanning was initiated with about 20 seconds delay after enhancement of the descending aorta to 100 HU, as measured using a bolus-tracking technique; portal venous and delayed phase images were obtained at 60–75 and 150–180 seconds, respectively, after injection of contrast.
Image analysis
Clinical features and semantic imaging assessment
The data on clinical characteristics collected through electronic health records included age, sex, hepatitis B virus (HBV), Barcelona clinic liver cancer stage, cirrhosis, Child-Pugh (CHILD), AFP [26], alanine aminotransferase (ALT), and aspartate aminotransferase (AST).
Two readers (***, *** with 15, and 7 years of experience in abdomen imaging, respectively) were blinded and independently reviewed all images to evaluate the semantic imaging features. Any controversies were settled by a majority vote. The following four semantic imaging features were evaluated: (1) pseudo–capsule, (2) TTPVI, (3) peritumoral enhancement, (4) margin, and (5) tumor size. The definition of TTPVI is the presence of tumors with internal arteries and without hypodense halos.
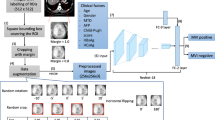
Overview of image processing and radiomics analysis. Four phases enhanced CT images were registered using Elastix and multimodal segmentation of tumor areas using nnU-Net. Three broad categories of images were designed: Original images, Subtraction images and Peritumoral images. Finally, feature extraction and selecting were performed and machine learning models were established
Image pre-processing
The raw data (Fig. 3A_I–A_IV) of all 4 phases of CT images were resampled to 0.695 × 0.695 × 5 mm3 to eliminate differences in rotation and slice thickness (Appendix E1). The first three phases (arterial, delay and nonenhanced) were transformed to corresponding portal venous phase image by rigid transformation. The same organs were then placed in the same positions, and all images share the same origins and spacings. Then Elastix software was used to register the four phases of CT images based on liver mask (Appendix E2). At last, pairwise subtraction was performed between the 4-phase images to generate 6 new images (Fig. 3B). The phase with lower average intensity is subtracted by those with larger average intensities.
Images detail display (both were male patients from ***). In A, I–IV and VI–IX are the original and four phase CT images after registration, respectively; B contains six subtraction images, from left to right, a–v, d–v, n–v, a–d, n–a, n-d (Appendix E5), respectively; C shows the peritumoral expansion area. From left to right, they are expanded by 2–20mm (increment = 2mm). V and X are spliced images, which are composed of different quarters of the focal parts of the original phase 4 CT and the post-registration phase 4 CT, respectively. The spliced images after registration is are closer to the mask predicted by the network than that before registration
Tumor segmentation
Each original transverse plane was reviewed and the tumor area was delineated on portal venous phase images of all patients by two radiologists using the ITK-SNAP software (http://www.itksnap.org) with 15 and 20 years of experience (*** and ***, respectively). Furthermore, all evaluators were blinded to the patients’ pathological and clinical data, and the results were determined by consensus.
Subsequently, aA tTotal of randomly -selected images from 98 patients were used as the training set, and those from the left 42 patients were used as the testing dataset. The nnU-Net was used for semantic segmentation of tumors. The preprocessed four CT phase images (Fig. 3A_VI–A_IX) were concatenated as input of the nnU-Net. The specific details and parameters of the network implementation can be described in Appendix E3.
Radiomic analysis
Feature extraction
Image transformation was performed on aligned four phase CT images and their subtraction images, including logarithm, square root, exponential, gradient, wavelet and Laplacian of Gaussian transforms. Then, PyRadiomics (https://pyradiomics.readthedocs.io) was used to extract radiomics features from the original and transformed images. Radiomics features include first-order statistics, shape, Gray level co-occurrence matrix (GLCM), Gray level size zone matrix (GLSZM), Gray level run length matrix (GLRLM), Gray level difference matrix (GLDM) and Neighborhood gray-tone difference matrix (NGTDM). Additionally, Combinations of image transformations were implemented to find the optimal subset of radiomics features for different image types of radiomics models.
Peritumoral analysis
Peritumoral features were extracted by dilating tumor regions within the liver mask. The tumor regions were dilated by 2 to 20 mm with an incremental of 2 mm. Dilated images were subtracted by original tumor regions to generate shell-like mask. These shells were then intersected with the liver mask to generate 10 peritumoral regions for each patient. A total of 6752 radiomic features were extracted from each peritumoral region.
Model construction
Z-score normalization was performed for all extracted radiomics features and clinical features. Wilcoxon rank sum test was carried out to select MVI-related features. Then maximum correlation optimum redundancy algorithm (mRMR) was performed to reduce redundancy across features. A combination of image transforms and Wilcoxon rank sum test thresholds (p-value < 0.01 and 0.05) (Appendix E4) was used to select the best parameters.
Three prediction models were built sequentially based on radiomic features: (1) The original modal (Ori model) was established using linear discriminant analysis (LDA) based on radiomic features from original 4 CT phase images. (2) Then, a radiomics model (Ori_sub model) was based on features from both original and subtraction images. (3) Peritumoral features were added to LDA classification model to construct the Ori_sub_edge model. In addition, we also built a LDA model based on clinical features. Finally, all of the above features were integrated to build a hybrid Ecos Model (Fig. 2) and We ranked importance of radiomics features through mRMR.
Model evaluation
The discriminability of predication models was evaluated by receiver operating characteristic curves (ROC). sensitivity, specificity, accuracy, PPV, NPV, and AUC were calculated. The clinical utility of the model was assessed through decision curve analysis (DCA), which involved measuring the net benefit at various threshold probabilities. This helped to determine the threshold probabilities at which the model could provide the greatest clinical benefit.
Statistical analysis
Statistical analysis was performed using IBM SPSS Statistics (Version 25) and MedCalc (Version 20.123) software(https://www.medcalc.org/). Patients enrolled in the MVI study were randomly assigned to the training and testing groups in a ratio of 7:3.
All continuous variables were tested for normal distribution and homogeneity of variance. The independent sample t test was used to compare the differences between the two groups. Delong test was used to compare the performance of the models. The discriminant performance of the model to predict MVI state was investigated using AUC as an indicator, and tested in the testing cohort. The difference was statistically significant (p-value < 0.05). Automatic segmentation and artificial mapping of interobserver and intraobserver dice similarity coefficients (DSCs) between tumors have been reported.
Results
Demographic and clinical information
The patient characteristics are shown in Table 1. In this study, 86 MVI− patients (mean age 55.92±9.624 years, 68 men) and 54 MVI+ patients (mean age 53.59±11.469 years, 43 men) were included. The diagnostic model was developed based on a training set involving 60 patients with MVI− (mean age: 56.3±9.962, 45 males) and 38 patients with MVI+ (mean age: 53.92±10.066, 30 males) and an independent testing set involving 26 patients with MVI− (mean age: 54.96±8.906, 23 men) and 16 MVI+ patients (mean age 52.81±14.625, 13 men), the two radiologists had very good inter-observer agreement on images features, with k statistics ranging from 0.93 to 0.99. The interobserver ICC was also good for tumor size, ranging from 0.84 to 0.95. The independent sample T test shows that there is no significant difference between the MVI+ group and the MVI− group in gender, margin, cirrhosis, and HBV between the training set and the verification set. Significant differences were observed in tumor size, pseudo–capsule and TTPVI (p-value < 0.05).
Liver tumor registration and tumor segmentation
The liver mask-based registration improved consistency of tumor location, size, and shape across different phases, as shown in Fig. 3(A_V, A_X).
The performance of the multi-modal deep learning segmentation model is shown in Table E3. In manually segmented observers, DSCs was good, ranging from 0.76 to 0.83 in the four test sequences (arterial phase, delay phase, nonenhanced phase, and portal venous phase). In addition, the inter-observer DSCs for manual segmentation was moderate, with DSCs ranging from 0.74 to 0.79 in the four sequences. The mean DSCs values between manual description and deep learn-based automatic segmentation were 0.76 and 0.82 in the single-mode (venous phase) and multi-mode segmentation models in the testing set, respectively.
Clinical model
The clinical model was established according to five clinical characteristics, including tumor size, pseudo–capsule, TTPVI, margin, and AFP (Table 2). The AUC of the final clinical model in the training cohort and testing cohort were 0.860 (95% [CI], 0.775–0.922) and 0.841 (95% [CI], 0.696–0.936). Calibration curves show good calibration with the training queue (p-value = 0.88) and testing queue (p-value = 0.94), respectively, Fig E1d.
Radiomics model
Models based on a combination of logarithm, wavelet, and square root transforms obtained the best performance for original four phase CT images and their subtraction images. A combination of all 6 transforms (logarithm, square root, exponential, gradient, wavelet, and Laplacian of Gaussian) showed the best performance in peritumoral radiomics model.
Violin maps of the radiomics features included in the construction model (f1–f24 represents 24 radiomics features, as shown in Table 2 respectively)
In feature selection, the Ori model, Ori_sub model, and Ori_sub_edge model selected 20, 13, and 24 radiomic features, respectively. As shown in Table 3 and Fig. 5(a,b), the Ori_sub_edge model outperformed the Ori model and Ori_sub model, and achieved the best AUC, accuracy, sensitivity, specificity, PPV, and NPV in both training and testing sets. The calibration curves of the three radiomics models on the training set and the testing set all show good calibration degree, the confusion matrix represents the overall performance of the model in distinguishing between MVI− groups and MVI+ groups in the training and testing queues (Fig E1).
ROC and AUC of five MVI− and MVI+ discrimination models were presented in training set (a) and testing set (b). Training set (c) and testing set (d) show the interrelation of decision curves of the five models. The black and blue lines represent the assumption that MVI is absent and present in all patients, respectively
The features of the established radiomics model are shown in Table 2. Among the selected features, the proportion of peritumoral features and subtraction image features is significantly higher than that of intratumoral features. Fig. 4 shows the numerical distribution of radiomics features. The heat map of the radiomics features of a selected patient was also visualized in Fig. 6. In the feature importance ranking, the top five features were the radiomics features of subtraction images, and among the top ten features, eight dimensions were peritumoral radiomics features (Table E2).
MVI− patients versus MVI+ cancer patients. A, B, and C respectively represent post-registration images of arterial phase, subtraction images of arterial phase and delay phase, and 6mm images of peritumoral dilatation of arterial phase. Some characteristic heat maps of three kinds of images are shown respectively. A_I to A_IV, B_I to B_IV, and C_I to C_IV were all male 1 (57 years old, MVI−); A_V to A_VIII, B_V to B_VIII, and C_V to C_VIII were male 2 (63 years old, MVI+) (both patients were from ***). A_III, A_VII and A_IV, A_VIII are the features of log-sigma-1-0-mm-3D_glszm_LowGrayLevelZoneEmphasis and log-sigma-3-0-mm-3D_firstorder_Kurtosis in the arterial period, respectively. B_III, B_VII and B_IV, B_VIII are the characteristics of wavelet-HHH_glszm_Zone Entropy and the wavelet-HLL_firstorder_Maximum in the subtracted image (a–d) of the arterial period minus the delay period, respectively. C_III, C_VII and C_IV, C_VIII are the characteristics of 6mm wavelet-LLH_glcm_Correlation and logarithm_glcm_Idmn in the peritumoral area, respectively. It can be seen that the features of different images have a good complementary effect
Model evaluation
By integrating 5-dimensional clinical features and 20-dimensional radiomics features, a hybrid prediction Model, Ecos Model, was obtained. In the training cohort, Delong test found that there was a significant difference between Clinical Model and Ecos Model (p-value = 0.003). There are also significant differences between Ori_sub_edge Model and Ecos Model (p-value = 0.032). In the testing cohort, the difference was marginal between the Clinical Model and the Ecos Model (p-value = 0.036), as well as between the Ori_sub_edge Model and the Ecos Model (p-value = 0.047), with significant differences. Calibration curves of Ecos Model on training set and testing set show optimal calibration, the confusion matrix represents the overall performance of the model in distinguishing between MVI− groups and MVI+ in the training and testing queues.
In addition, a series of comparative tests were conducted to verify the optimal peri-tumor expansion distance (Fig. 3C). The results of 10 peritumoral expansion comparison models are shown in Table E1. In the training set, the 6mm region around the tumor outperformed the other nine models on AUC, Sen, and NPV. In the test set, the 6mm peritumor region performed optimally on AUC, accuracy, specificity, and PPV among the 10 models. In addition, the optimal values of the remaining evaluation indicators in both training set and testing set were mainly concentrated around the 10 and 12mm peritumoral models, marginal significantly (p-value < 0.15). In the 12–20mm model, the AUC of the testing set is all lower than 0.9, while the average value of other indicators is lower than that of the model within 10mm.
Clinical utility
DCA (Fig. 5c–d) shows that in both the training set and testing set, using all five LDA-based models to distinguish MVI− from MVI+ was more beneficial than the “treat all” approach and the “treat none” approach, respectively. Additionally, the Ecos model outperforms other four models in a relatively large threshold range.
Discussion
This study developed a hybrid model which combined radiomic features and clinical features to predict preoperative MVI. Proposed hybrid model achieved good discrimination in both training (AUC = 0.96) and test datasets (AUC = 0.91), and outperformed the clinical model (AUC = 0.86 and 0.84 for training and test datasets, respectively) and radiomics model (AUC = 0.90 and 0.87). The proposed hybrid model achieved the state-of-the-art classification level, and provided an automated non-invasive tool to discriminate MVI- patients from MVI+ patients.
Co-registration between different CT phases improved both manual delineation and automated segmentation of tumors [17, 18]. Because of movement during CT examination, tumors can locate at various positions between different phases, which calls for tedious manual delineation on all CT phases [19]. Co-registration compensated effects of movement, therefore only one manual delineation is required for one patient. Information of different CT phases can be utilized at the same time, and thus improvinged the quality and intra-subject consistence of delineation [20]. In the actual situation, the problem of inconsistent delineation between phases of CT at different periods can be avoided to a large extent. Moreover, a deep learning method was implemented to segment tumors based on all 4 co-registered CT phases [21]. The proposed segmentation method can capture comprehensive tumor features from different CT phases, and thus can outperform single-phase- based methods [22]. As far as we know, this is the first automatic segmentation method based on multiple CT phases to distinguish MVI− and MVI+.
In this study, dynamic features between CT phases were investigated by subtraction images. Radiomic features of subtraction images dramatically improved discrimination of MVI− and MVI+. Our finding is in consistence with previous study studies that subtraction of CT images improved detection of HCC [23,24,25]. Furthermore, radiomic features of subtraction images played a more important role than differences between radiomic features of original phases. Dynamic information, referred by radiomic features of subtraction images, was closely related to MVI pathology. It implied that more attention should be payed paid to the dynamic changes between CT phases in clinical diagnosis of liver tumors.
Results showed that tumor size, pseudo–capsule, TTPVI, margin, and AFP were important predictors of MVI, which was consistent with previous studies [26, 27]. Furthermore, radiomic features were extracted to improve the discrimination between MVI+ and MVI−. In the pool of radiomic features, the selected radiomics features of subtraction images focus on the first-order statistical features and GLSZM features after wavelet transformation. First-order statistics can provide information about the overall brightness and contrast of the tumor, and GLSZM represents information about different gray levels and continuous pixel region distribution, respectively. First-order statistics can be used to evaluate the perfusion of tumor, in accordance with the fact that microvascular invasion is often associated with the blood supply to the tumor [28]. GLSZM features quantify distribution of different gray levels and the size of contiguous regions within tumor regions, possibly related to the diffusion and density of blood vessels. Proposed model concentrated on features from subtraction images more than those from original images (features number = 11 vs. 2). The wavelet transform features of subtraction images greatly improved the model, and the top five features of mRMR screening all belonged to features of subtraction images. It means subtraction images can enhance the visibility of vascular structures and highlight the tumor's internal composition, leading to more accurate and detailed radiomic features that can aid in better characterizing the tumor and guiding clinical decision-making.
In addition, the model also focused on peritumoral radiomics features extracted from the surrounding region of the tumor. These periphery features included GLCM features and GLDM features. GLCM provides statistical information about the relationships between gray levels of different pixels in the image, while GLDM describes the gray level differences between different pixels in the image. The GLCM features represent the texture characteristics in the peritumoral area, giving insights into the patterns and structures of the neighboring tissues. On the other hand, the GLDM features offer valuable information about the arrangement of microvessels and changes in blood perfusion in the peritumoral region. These radiomics features provide a more comprehensive understanding of the peritumoral environment, enhancing the model's ability to analyze and predict tumor behavior and treatment outcomes.
Our study also evaluated peritumoral regional features for MVI discrimination [29]. The quantitative evaluation showed that peritumoral 6mm model showed consistent and excellent results in both training set and testing set, which means that the radiomics features of the peritumoral 6mm ring region contributed the most to the final prediction model, and the imaging features in the peritumoral 8mm-12mm region had a good prediction effect, which was in line with clinical experience [30,31,32]. Our finding is consistent with previous study that proved portal vein phase within 12mm of the tumor core and peripheral areas as potential quantitative imaging biomarkers [32]. Moreover, peritumoral parts larger than 12mm also have vascular invasion. Due to a certain degree of overfitting, it is not enough to improve the prediction results of the model. However, theoretically, resection is required in clinical surgery, which requires further discussion and experimentation [33,34,35]. In conclusion, this study provides a good quantitative reference for the clinician to delimit the lesion area before surgery and resection during surgery [36]. At the same time, peritumoral dilation can alleviate, to a certain extent, the problems such as the loss of effective features or the extraction of invalid features caused by the inaccurate edge of the lesion caused by automatic segmentation error.
There are still several limitations in this study. First, although this study included more patients compared with previous studies (140 vs 128 and 111) [26, 32], the sample size could still be relatively small. Second, different CT scans from two centers may affect selection of radiomic features. This issue could be partially solved by image normalization in this study, however, alignment of radiomic features needs further studies. Third, this study mixed the patients from two centers and randomly assigned them to a training and a testing dataset, future studies may include more patients from multiple centers to validate the findings of this study.
In summary, we have developed an end-to-end automated diagnostic process that facilitates preoperative differentiation between MVI− and MVI+. This model quantified multimodal image features including clinical features, dynamic features, peritumoral and intratumoral radiomics features, and thus could comprehensively represent image characteristics of HCC. Results showed that proposed Ecos model can efficiently distinguish MVI− from MVI+ to facilitate clinical decision-making.
References
Sperandio RC, Pestana RC, Miyamura BV, Kaseb AO (2022) Hepatocellular carcinoma immunotherapy. Annu Rev Med 73:267–278. https://doi.org/10.1146/annurev-med-042220-021121
Romain D, Amaia L (2022) The liver cancer immune microenvironment: Therapeutic implications for hepatocellular carcinoma. HEPATOLOGY. https://doi.org/10.1002/hep.32740
Lima HA, Moazzam Z, Endo Y, et al (2023) TBS-Based preoperative score to predict non-transplantable recurrence and identify candidates for upfront resection versus transplantation for hepatocellular carcinoma. Ann Surg Oncol. https://doi.org/10.1245/s10434-023-13273-1
Wei Y, Pei W, Qin Y, et al (2021) Preoperative MR imaging for predicting early recurrence of solitary hepatocellular carcinoma without microvascular invasion. Eur J Radiol 138:109663. https://doi.org/10.1016/j.ejrad.2021.109663
Imura S, Teraoku H, Yoshikawa M, et al (2018) Potential predictive factors for microvascular invasion in hepatocellular carcinoma classified within the Milan criteria. Int J Clin Oncol 23:98–103. https://doi.org/10.1007/s10147-017-1189-8
Yang C, Liu X, Ling W-W, et al (2020) Primary isolated hepatic tuberculosis mimicking small hepatocellular carcinoma. Medicine. https://doi.org/10.1097/MD.0000000000022580
Xie D, Ren Z, Zhou J, et al (2020) 2019 Chinese clinical guidelines for the management of hepatocellular carcinoma: updates and insights. Hepatobiliary Surg Nutr v.9(4). https://doi.org/10.21037/hbsn-20-480
Song L, Li J, Luo Y (2021) The importance of a nonsmooth tumor margin and incomplete tumor capsule in predicting HCC microvascular invasion on preoperative imaging examination: a systematic review and meta-analysis. Clin Imaging 76:77–82. https://doi.org/10.1016/j.clinimag.2020.11.057
Zhang H-M, Wen D-G, Chen J, et al (2023) A diagnostic test of three-dimensional magnetic resonance elastography imaging for preoperative prediction of microvascular invasion in patients with T1 stage clear cell renal carcinoma. Transl Androl Urol 12. https://doi.org/10.21037/tau-23-94
Peng J, Zhang J, Zhang Q-F, et al (2018) A radiomics nomogram for preoperative prediction of microvascular invasion risk in hepatitis B virus-related hepatocellular carcinoma. Diagn Interv Radiol. https://doi.org/10.5152/dir.2018.17467
Lambin P, Leijenaar RT, Deist TM, et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 14:749–762. https://doi.org/10.1038/nrclinonc.2017.141
Jha AK, Mithun S, Purandare NC, Kumar R, et al (2022) Radiomics: a quantitative imaging biomarker in precision oncology. Nucl Med Commun 43:483–493. https://doi.org/10.1097/MNM.0000000000001543
Jiang Y-Q, Cao S-E, Cao S, et al (2021) Preoperative identification of microvascular invasion in hepatocellular carcinoma by XGBoost and deep learning. J Cancer Res Clin Oncol 147:821–833. https://doi.org/10.1007/s00432-020-03366-9
Jiang C, Zhao L, Xin B, et al (2022) 18F-FDG PET/CT radiomic analysis for classifying and predicting microvascular invasion in hepatocellular carcinoma and intrahepatic cholangiocarcinoma. Quant Imaging Med Surg 12:4135–4150. https://doi.org/10.21037/qims-21-1167
Park S, Kim JH, Kim J, et al (2023) Development of a deep learning-based auto-segmentation algorithm for hepatocellular carcinoma (HCC) and application to predict microvascular invasion of HCC using CT texture analysis: preliminary results. Acta Radiol 64:907–917. https://doi.org/10.1177/02841851221100318
Yao W, Yang S, Ge Y, et al (2022) Computed tomography radiomics-based prediction of microvascular invasion in hepatocellular carcinoma. Front Med 9. https://doi.org/10.3389/fmed.2022.819670
Azam M, Khan K, Ahmad M, et al (2021) Multimodal medical image registration and fusion for quality enhancement. Comput Mater Contin 68:821–840. https://doi.org/10.32604/cmc.2021.016131
Albers J, Svetlove A, Alves J, et al (2021) Elastic transformation of histological slices allows precise co-registration with microCT data sets for a refined virtual histology approach. Sci Rep 11:10846. https://doi.org/10.1038/s41598-021-89841-w
Rietzel E, Pan T, Chen GTY (2005) Four-dimensional computed tomography: Image formation and clinical protocol. Med Phys 32:874–889. https://doi.org/10.1118/1.1869852
Decaux N, Conze P-H, Ropars J, et al (2023) Semi-automatic muscle segmentation in MR images using deep registration-based label propagation. Pattern Recognit 140:109529. https://doi.org/10.1016/j.patcog.2023.109529
Li L, Zhao X, Lu W, Tan S (2020) Deep learning for variational multimodality tumor segmentation in PET/CT. Neurocomputing 392:277–295. https://doi.org/10.1016/j.neucom.2018.10.099
Zhang Y, Peng C, Peng L, et al (2021) Multi-phase liver tumor segmentation with spatial aggregation and uncertain region inpainting, Medical Image Computing and Computer Assisted Intervention. https://doi.org/10.1007/978-3-030-87193-2_7
Aoki T, Kamiya T, Lu H, et al (2021) CT temporal subtraction: techniques and clinical applications. Quant Imaging Med Surg 11:2214–2223. https://doi.org/10.21037/qims-20-1367
Kim DH, Choi SH, Byun JH, et al (2019) Arterial subtraction images of gadoxetate-enhanced MRI improve diagnosis of early-stage hepatocellular carcinoma. J Hepatol 71:534–542. https://doi.org/10.1016/j.jhep.2019.05.005
Lee J, Kim KW, Kim SY, et al (2015) Automatic detection method of hepatocellular carcinomas using the non-rigid registration method of multi-phase liver CT images. J X-Ray Sci Technol 23:275–288. https://doi.org/10.3233/XST-150487
Zhang T, Pandey G, Xu L, et al (2020) The value of TTPVI in prediction of microvascular invasion in hepatocellular carcinoma. Cancer Manag Res 12:4097. https://doi.org/10.2147/CMAR.S245475
Peng J, Zhang J, Zhang Q, et al (2018) A radiomics nomogram for preoperative prediction of microvascular invasion risk in hepatitis B virus-related hepatocellular carcinoma. Diagn Interv Radiol 24:121. https://doi.org/10.5152/dir.2018.17467
Jiang Y, Cao S, Cao S, et al (2021) Preoperative identifcation of microvascular invasion in hepatocellular carcinoma by XGBoost and deep learning. Journal of Cancer Research and Clinical Oncology 147:821–833. https://doi.org/10.1007/s00432-020-03366-9
Wu X, Dong D, Zhang L, et al (2021) Exploring the predictive value of additional peritumoral regions based on deep learning and radiomics: a multicenter study. Med Phys 48:2374–2385. https://doi.org/10.1002/mp.14767
Chong H-H, Yang L, Sheng R-F, et al (2021) Multi-scale and multi-parametric radiomics of gadoxetate disodium–enhanced MRI predicts microvascular invasion and outcome in patients with solitary hepatocellular carcinoma≤ 5 cm. Eur Radiol 31:4824–4838. https://doi.org/10.1007/s00330-020-07601-2
Chong H, Gong Y, Pan X, et al (2021) Peritumoral dilation radiomics of gadoxetate disodium-enhanced MRI excellently predicts early recurrence of hepatocellular carcinoma without macrovascular invasion after hepatectomy. J Hepatocell Carcinoma 545–563. https://doi.org/10.2147/JHC.S309570
Zhang W, Yang R, Liang F, et al (2021) Prediction of microvascular invasion in hepatocellular carcinoma with a multi-Disciplinary team-Like radiomics fusion model on dynamic contrast-Enhanced computed tomography. Front Oncol 11:660629. https://doi.org/10.3389/fonc.2021.660629
Shipe ME, Deppen SA, Farjah F, Grogan EL (2019) Developing prediction models for clinical use using logistic regression: an overview. J Thorac Dis 11:S574. https://doi.org/10.21037/jtd.2019.01.25
Azad TD, Ehresman J, Ahmed AK, et al (2021) Fostering reproducibility and generalizability in machine learning for clinical prediction modeling in spine surgery. Spine J 21:1610–1616. https://doi.org/10.1016/j.spinee.2020.10.006
Steyerberg EW, Steyerberg EW (2019) Overfitting and optimism in prediction models. Clin Predict Models Pract Approach. https://doi.org/10.1007/978-3-030-16399-0_5
Guo Y, Mokany K, Ong C, et al (2023) Plant species richness prediction from DESIS hyperspectral data: A comparison study on feature extraction procedures and regression models. ISPRS J Photogramm Remote Sens 196:120–133. https://doi.org/10.1016/j.isprsjprs.2022.12.028
Funding
This work was supported by National Natural Science Foundation of China (Nos. 62171230, 62101365, 92159301, 91959207, 62301263, 62301265, 62302228, 82302291, 82302352).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors of this manuscript declare no relationships with each other and any companies, whose products or services may be related to the subject matter of the article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Z., Xia, T., Zhang, T. et al. Prediction of preoperative microvascular invasion by dynamic radiomic analysis based on contrast-enhanced computed tomography. Abdom Radiol 49, 611–624 (2024). https://doi.org/10.1007/s00261-023-04102-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00261-023-04102-w