Abstract
Purpose
The purpose of this study is to evaluate diagnostic performance of a commercially available radiomics research prototype vs. an in-house radiomics software in the binary classification of CT images from patients with pancreatic ductal adenocarcinoma (PDAC) vs. healthy controls.
Materials and methods
In this retrospective case–control study, 190 patients with PDAC (97 men, 93 women; 66 ± 9 years) from 2012 to 2017 and 190 healthy potential renal donors (96 men, 94 women; 52 ± 8 years) without known pancreatic disease from 2005 to 2009 were identified from radiology and pathology databases. 3D volume of the pancreas was manually segmented from preoperative CT scans. Four hundred and seventy-eight radiomics features were extracted using in-house radiomics software. Eight hundred and fifty-four radiomics features were extracted using a commercially available research prototype. Random forest classifier was used for binary classification of PDAC vs. normal pancreas. Accuracy, sensitivity, and specificity of commercially available radiomics software were compared to in-house software.
Results
When 40 radiomics features were used in the random forest classification, in-house software achieved superior sensitivity (1.00) and accuracy (0.992) compared to the commercially available research prototype (sensitivity = 0.950, accuracy = 0.968). When the number of features was reduced to five features, diagnostic performance of the in-house software decreased to sensitivity (0.950), specificity (0.923), and accuracy (0.936). Diagnostic performance of the commercially available research prototype was unchanged.
Conclusion
Commercially available and in-house radiomics software achieve similar diagnostic performance, which may lower the barrier of entry for radiomics research and allow more clinician-scientists to perform radiomics research.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Radiomics converts imaging data into high-dimensional mineable features, which have the potential to yield imaging biomarkers for tumor classification and prognostication [1]. This is currently a “hot topic” and exciting frontier in radiology research. Radiomics and other artificial intelligence approaches dominate discussions at our radiology meetings and our publications [2, 3]. These publications usually report impressive results with high level of statistical significance [4,5,6], but most if not all of these publications end with the same disclaimers related to reproducibility [5, 7, 8]. This is because there currently are no standardized image acquisition and post-processing protocols in radiomics. Technical parameters related to image acquisition (dose, phase) [9, 10], reconstruction (slice thickness, reconstruction kernel) [9, 11], segmentation technique [10], and radiomics feature extraction [12]. A number of studies have explored using compensation techniques or deep-learning based algorithms to mitigate the effect of these technical parameters on radiomics features [13,14,15,16]. In addition, there is high barrier to entry in radiomics research due to amount of data required for analysis and many of the published reports have used in-house radiomics software that required expertise in computer science. To our knowledge, only one publication has investigated the effect of in-house vs. freely available radiomics software in the calculated radiomics features [12]. Foy et al. evaluated radiomics features extracted from regions of interests from 40 mammograms and 39 head and neck CTs using two in-house radiomics software programs and two freely available radiomics packages, and found that there were significant variations in the calculated values across software platforms [12]. In this study, the authors evaluated the effect of software on the calculated radiomics features but they did not evaluate whether these variations affected the overall performance in a classification task. The purpose of this study is to evaluate the diagnostic performance of a commercially available radiomics research prototype vs. an in-house radiomics software in binary classification of CT images from patients with pancreatic ductal adenocarcinoma (PDAC) vs. healthy control subjects.
Materials and methods
Patients
This study was an Institutional Review Board-approved HIPAA-compliant retrospective study. The same dataset of patients with PDAC and healthy controls [6] was used for both in-house radiomics software and commercially available software. Results for the in-house software were previously published [6]. Briefly, 190 patients with surgically resected PDAC were identified from the radiology and pathology databases at our institution from 2012 to 2017. One hundred and ninety healthy renal donors without known pancreatic disease were identified from the radiology database from 2005 to 2009. Patients with suspected PDAC based on imaging features without surgical proof were excluded. Medical records of potential renal donors were reviewed to exclude participants with pancreatic disease (e.g., pancreatitis, pancreatic mass) and diabetes mellitus. Preoperative CT scans of patients with PDAC and healthy control subjects were analyzed. The dataset was divided into 255 training cases (125 healthy control cases and 130 PDAC cases) and 125 validation cases (65 healthy control cases and 60 PDAC cases). The training and testing cases were randomly selected from total 380 cases (190 PDAC + 190 normal) as the 2/3 for training (255 cases, 67%) and the remaining 1/3 (125 cases, 33%) for testing. The number of training cases was twice of the testing cases so that the samples can statistically cover the distribution of testing cases.
CT acquisition
Patients with PDAC were scanned on a 64-slice MDCT scanner (Sensation 64, Siemens Healthineers) or dual-source MDCT scanner (FLASH, Siemens Healthineers), and healthy control subjects were scanned on a 64-slice MDCT scanner (Sensation 64 Siemens Healthineers). Patients with PDAC and healthy control subjects were injected with 100–120 mL of iohexol (Omnipaque, GE Healthcare) at an injection rate of 4–5 mL/s. Scanning protocols were customized for each patient to minimize dose but were in the order of 120 kVp, 300 mAs, and 0.6–0.8 pitch. Both arterial and venous phases were acquired per institution protocol, for both patients with PDAC and healthy renal donors.
Image segmentation
Venous phase 0.75 mm slices were chosen for image segmentation and radiomics analysis. The whole 3D volume of pancreas for healthy control cases and the whole 3D volume of the tumor, background pancreas, and whole pancreas (including tumor region and background normal pancreas) for PDAC cases were manually segmented by four researchers (a radiation oncologist with 30 years of experience, a CT technologist with 20 years of experience, and two post-doctoral fellows with 1 year experience) using commercial segmentation software (Velocity 3.2.0, Varian Medical Systems). The contours were verified by three abdominal radiologists with 5–30 years of experience. Features extracted from whole pancreas contours were used in radiomics analysis.
Image analysis
Image analysis using in-house software has been described in detail in [6], and 478 features were extracted. The features were based on tumor intensity, shape, texture, and wavelet features as described in [17], and the process was implemented using C++ language by our computer scientist. Because the number of features was larger than the number of training cases, it was necessary to reduce the redundancy of computed features. Minimum-redundancy maximum-relevancy feature selection was applied to the computed feature set and 40 features (and subset of 5 most relevant features) were selected for random forest classification.
Image analysis using commercial software was performed on syngo.via Frontier Radiomics prototype (syngo.via Frontier, Siemens Healthineers). Eight hundred and fifty-four radiomics features including first-order statistics, shape, and texture were extracted from the original images. Additional first-order statistics and texture features were computed from filtered images, such as wavelet filters. Feature reduction was performed and 40 most relevant features were selected for random forest classification. Decision trees for random forest were developed based on the training dataset and tested on the test set by majority voting. Performance of the algorithm was evaluated by overall sensitivity, specificity, and accuracy in binary classification of cases from patients with PDAC and healthy control subjects.
Results
Demographic information of the 190 patients with surgically resected PDAC and 190 healthy control subjects is shown in Table 1. The mean and SD of the maximal 2D diameter of the tumor was 4.1 ± 1.7 cm for the 190 patients with PDAC. The unsupervised clustering results of all 854 radiomics features extracted using commercially available research prototype for both the PDAC cases and healthy control cases are shown in Fig. 1. This heat map represents a color-coded array of all feature values (y-axis) in all cases (x-axis). For visualization, each individual radiomics feature is normalized on the basis of all 380 cases.
Forty features were selected from the commercially available research prototype using minimum-redundancy maximum-relevancy feature selection and the 10 most relevant features are shown in Table 2. The number of features was further reduced to five features to allow more direct comparison with results from the previous publication [6]. Diagnostic performance of the commercially available research prototype and in-house software in classifying CT cases from patients with PDAC and healthy control subjects is shown in Table 3. When 40 radiomics features were used, the in-house software achieved superior sensitivity (1.00) and accuracy (0.992) compared to the commercially available research prototype (sensitivity = 0.950, accuracy = 0.968). Both software achieved the same specificity (0.985). When the number of features was reduced to five features, diagnostic performance of the in-house software decreased to sensitivity (0.950), specificity (0.923), and accuracy (0.936), whereas the diagnostic performance of the commercially available research prototype was unchanged (sensitivity 0.950, specificity 0.985, and accuracy 0.968).
Although both radiomics software programs generated three false negatives when only five radiomics features were used (sensitivity = 0.950), they only shared one of the false negatives in common. The case that was misclassified as false negative by both programs was a predominantly exophytic mass arising from the head of pancreas with contiguous porta hepatic lymphadenopathy (Fig. 2). The other two false negatives that were misclassified by one radiomics software program were correctly classified by the other radiomics software (Figs. 3, 4). The discrepancy was likely due to differences in computation and selection of relevant features (Table 4).
False negative case of pancreatic ductal adenocarcinoma that is misclassified with both commercially available and in-house radiomics software, using five radiomics features for classification. Coronal IV contrast-enhanced CT image of a 85-year-old man shows a predominantly exophytic mass arising from head of pancreas that is contiguous with locoregional lymphadenopathy (arrows)
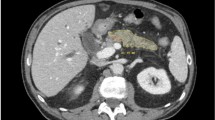
False negative cases of pancreatic ductal adenocarcinoma that are misclassified by the commercially available software but were correctly classified by the in-house software, using five radiomics features for classification. a Axial IV contrast-enhanced CT image of a 62-year-old woman shows diffusely infiltrative tumor throughout the pancreatic body and tail (white arrow) associated with dilatation of the pancreatic duct (black arrow). b Axial IV contrast-enhanced CT image of a 65-year-old woman shows an exophytic hypoenhancing mass arising from the uncinate process of the pancreas (arrow)
False negative cases of pancreatic ductal adenocarcinoma that are misclassified by the in-house software but were correctly classified by the commercially available software, using five radiomics features for classification. a Coronal IV contrast-enhanced CT image of a 59-year-old man shows dilated common bile duct and pancreatic duct with a subtle isoenhancing mass at the transition point in the ampullary region (arrow). b Axial IV contrast-enhanced CT image of a 69-year-old man shows a hypoenhancing mass in the pancreatic body (arrow)
Discussion
Radiomics has the potential to generate imaging biomarkers for classification and prognostication. Technical parameters from image acquisition to feature extraction and analysis have the potential to affect radiomics features [9,10,11,12]. The current study used the same CT images with manual segmentation on both a commercially available research prototype and in-house radiomics software to control for any variability at the image acquisition step and compared the diagnostic performance of the two programs. Both programs achieved similar diagnostic performance in the binary classification of CT images from patients with PDAC and healthy control subjects, despite differences in the radiomics features they employed (854 features in commercial program vs. 478 features in in-house program). This is reassuring that even though there may be variations in the computed values for radiomics features, the differences do not seem to significantly impact the overall diagnostic performance of the constellation of radiomics features. This is important for the broader implementation of radiomics research. Currently, many radiomics studies have been performed using proprietary in-house software, which requires in-house expertise in computer science, a luxury that only a few academic centers can afford. The results of this study show that commercially available radiomics software may be a viable alternative to in-house computer science expertise, which can lower the barrier of entry for radiomics research and allow clinicians to validate findings of the published studies with their own local datasets.
In the previously published study [6], we observed a decrease in diagnostic performance when the number of features was reduced from 40 features to 5 features. In the current study, there is no change in performance when the number of features was reduced. Interesting, although both programs achieve the same sensitivity (0.950) using five features, the false negative cases are not the same across both programs, likely due to differences in computation and selection of relevant features. The cases that were misclassified as false negatives by one or both software show diverse imaging appearance ranging from small subtle isoenhancing mass to diffuse tumor infiltration of the pancreas. These algorithms appear to be focusing on different imaging features as the basis for the classification. It may be possible to combine different algorithms to achieve superior performance. Due to “blackbox” nature of radiomics, it is not easy pinpoint the exact cause of the difference in performance of these programs. Future research is needed to understand these differences.
This study has a number of limitations. First, it was a retrospective study with a relatively small sample size. The study population was selected based on the previously published study to allow for direct comparison of diagnostic performance between the commercially available and the in-house software. Second, this study compared the performance of two software on one specific application. Future research is needed to determine if other commercially available radiomics software will achieve similar results and if these software will achieve similar results for other clinical applications and imaging modality. Third, there is currently no standardization of imaging protocol for radiomics studies. In the future, these radiomics software will require validation across different institutions, vendors, and scanning protocols.
Conclusion
This study showed that a commercially available radiomics software may be able to achieve similar diagnostic performance as an in-house radiomics software. The results obtained from one radiomics software may be transferrable to another system. Availability of commercial radiomics software may lower the barrier of entry for radiomics research and allow more researchers to engage in this exciting area of research.
References
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: Images Are More than Pictures, They Are Data. Radiology 278 (2):563–577. https://doi.org/10.1148/radiol.2015151169
Bluemke DA (2020) Top Publications in Radiology, 2019. Radiology 294 (1):2–3. https://doi.org/10.1148/radiol.2019194017
Carlos RC, Kahn CE, Halabi S (2018) Data Science: Big Data, Machine Learning, and Artificial Intelligence. J Am Coll Radiol 15 (3 Pt B):497–498. https://doi.org/10.1016/j.jacr.2018.01.029
Lubner MG, Smith AD, Sandrasegaran K, Sahani DV, Pickhardt PJ (2017) CT Texture Analysis: Definitions, Applications, Biologic Correlates, and Challenges. Radiographics : a review publication of the Radiological Society of North America, Inc 37 (5):1483–1503. https://doi.org/10.1148/rg.2017170056
Bodalal Z, Trebeschi S, Nguyen-Kim TDL, Schats W, Beets-Tan R (2019) Radiogenomics: bridging imaging and genomics. Abdominal radiology 44 (6):1960–1984. https://doi.org/10.1007/s00261-019-02028-w
Chu LC, Park S, Kawamoto S, Fouladi DF, Shayesteh S, Zinreich ES, Graves JS, Horton KM, Hruban RH, Yuille AL, Kinzler KW, Vogelstein B, Fishman EK (2019) Utility of CT Radiomics Features in Differentiation of Pancreatic Ductal Adenocarcinoma From Normal Pancreatic Tissue. AJR American journal of roentgenology:1–9. https://doi.org/10.2214/ajr.18.20901
Kumar V, Gu Y, Basu S, Berglund A, Eschrich SA, Schabath MB, Forster K, Aerts HJ, Dekker A, Fenstermacher D, Goldgof DB, Hall LO, Lambin P, Balagurunathan Y, Gatenby RA, Gillies RJ (2012) Radiomics: the process and the challenges. Magnetic resonance imaging 30 (9):1234–1248. https://doi.org/10.1016/j.mri.2012.06.010
Kocak B, Durmaz ES, Erdim C, Ates E, Kaya OK, Kilickesmez O (2019) Radiomics of Renal Masses: Systematic Review of Reproducibility and Validation Strategies. AJR American journal of roentgenology:1–8. https://doi.org/10.2214/ajr.19.21709
Meyer M, Ronald J, Vernuccio F, Nelson RC, Ramirez-Giraldo JC, Solomon J, Patel BN, Samei E, Marin D (2019) Reproducibility of CT Radiomic Features within the Same Patient: Influence of Radiation Dose and CT Reconstruction Settings. Radiology 293 (3):583–591. https://doi.org/10.1148/radiol.2019190928
Yamashita R, Perrin T, Chakraborty J, Chou JF, Horvat N, Koszalka MA, Midya A, Gonen M, Allen P, Jarnagin WR, Simpson AL, Do RKG (2020) Radiomic feature reproducibility in contrast-enhanced CT of the pancreas is affected by variabilities in scan parameters and manual segmentation. European radiology 30 (1):195–205. https://doi.org/10.1007/s00330-019-06381-8
Kim H, Park CM, Gwak J, Hwang EJ, Lee SY, Jung J, Hong H, Goo JM (2019) Effect of CT Reconstruction Algorithm on the Diagnostic Performance of Radiomics Models: A Task-Based Approach for Pulmonary Subsolid Nodules. AJR American journal of roentgenology 212 (3):505–512. https://doi.org/10.2214/ajr.18.20018
Foy JJ, Robinson KR, Li H, Giger ML, Al-Hallaq H, Armato SG (2018) Variation in algorithm implementation across radiomics software. J Med Imaging (Bellingham) 5 (4):044505. https://doi.org/10.1117/1.jmi.5.4.044505
Orlhac F, Frouin F, Nioche C, Ayache N, Buvat I (2019) Validation of A Method to Compensate Multicenter Effects Affecting CT Radiomics. Radiology 291 (1):53–59. https://doi.org/10.1148/radiol.2019182023
Park S, Lee SM, Do KH, Lee JG, Bae W, Park H, Jung KH, Seo JB (2019) Deep Learning Algorithm for Reducing CT Slice Thickness: Effect on Reproducibility of Radiomic Features in Lung Cancer. Korean journal of radiology : official journal of the Korean Radiological Society 20 (10):1431–1440. https://doi.org/10.3348/kjr.2019.0212
Zhovannik I, Bussink J, Traverso A, Shi Z, Kalendralis P, Wee L, Dekker A, Fijten R, Monshouwer R (2019) Learning from scanners: Bias reduction and feature correction in radiomics. Clin Transl Radiat Oncol 19:33–38. https://doi.org/10.1016/j.ctro.2019.07.003
Choe J, Lee SM, Do KH, Lee G, Lee JG, Seo JB (2019) Deep Learning-based Image Conversion of CT Reconstruction Kernels Improves Radiomics Reproducibility for Pulmonary Nodules or Masses. Radiology 292 (2):365–373. https://doi.org/10.1148/radiol.2019181960
Aerts HJ, Velazquez ER, Leijenaar RT, Parmar C, Grossmann P, Carvalho S, Bussink J, Monshouwer R, Haibe-Kains B, Rietveld D, Hoebers F, Rietbergen MM, Leemans CR, Dekker A, Quackenbush J, Gillies RJ, Lambin P (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nature communications 5:4006. https://doi.org/10.1038/ncomms5006
Funding
Linda C. Chu, Seyoun Park, Satomi Kawamoto, Alan L. Yuille, and Elliot K. Fishman received research support from the Lustgarten Foundation. Linda C. Chu, Seyoun Park, and Elliot K. Fishman received additional research support from the Emerson Collective. Other authors have no disclosures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethics approval
This was an IRB-approved retrospective study.
Informed consent
Patient consent was waived given retrospective nature of the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chu, L.C., Solmaz, B., Park, S. et al. Diagnostic performance of commercially available vs. in-house radiomics software in classification of CT images from patients with pancreatic ductal adenocarcinoma vs. healthy controls. Abdom Radiol 45, 2469–2475 (2020). https://doi.org/10.1007/s00261-020-02556-w
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00261-020-02556-w