Abstract
Pasteurella multocida is an important bacterial pathogen that can cause diseases in both animals and humans. Its elevated morbidity and mortality rates in animals result in substantial economic repercussions within the livestock industry. The prevention of diseases caused by P. multocida through immunization is impeded by the absence of a safe and effective vaccine. Outer membrane vesicles (OMVs) secreted from the outer membrane of Gram-negative bacteria are spherical vesicular structures that encompass an array of periplasmic components in conjunction with a diverse assortment of lipids and proteins. These vesicles can induce antibacterial immune responses within the host. P. multocida has been shown to produce OMVs. Nonetheless, the precise characteristics and immunomodulatory functions of P. multocida OMVs have not been fully elucidated. In this study, OMVs were isolated from P. multocida using an ultrafiltration concentration technique, and their morphology, protein constitution, and immunomodulatory properties in RAW264.7 cells were studied. Transmission electron microscopy (TEM) and nanoparticle tracking analysis (NTA) revealed that the OMVs exhibited typical spherical and bilayered lipid vesicular architecture, exhibiting an average diameter of approximately 147.5 nm. The yield of OMVs was 2.6 × 1011 particles/mL. Proteomic analysis revealed a high abundance of membrane-associated proteins within P. multocida OMVs, with the capability to instigate the host’s immune response. Furthermore, OMVs stimulated the proliferation and cellular uptake of macrophages and triggered the secretion of cytokines, such as TNF-ɑ, IL-1β, IL-6, IL-10, and TGF-β1. Consequently, our results indicated that OMVs from P. multocida could directly interact with macrophages and regulate their immune function in vitro. These results supported the prospective applicability of P. multocida OMVs as a platform in the context of vaccine development.
Key points
• Preparation and characterization of P. multocida OMVs.
• P. multocida OMVs possess a range of antigens and lipoproteins associated with the activation of the immune system.
• P. multocida OMVs can activate the proliferation, internalization, and cytokine secretion of macrophages in vitro.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the 1880s, Louis Pasteur was the first to identify the etiological role of Pasteurella multocida (Harper et al. 2006). P. multocida is a highly infectious, zoonotic pathogen that can cause numerous diseases of economic importance, including avian cholera, bovine hemorrhagic sepsis, endemic animal pneumonia, and porcine atrophic rhinitis (Boyce and Adler 2006; Roier et al. 2013; Wilkie et al. 2012). Contact with the saliva of animals colonized by P. multocida can cause soft tissue infection in humans (Wilson and Ho 2013). Other severe complications, such as pneumonia, meningitis, and sepsis from infection, may also occur in elderly individuals, immunocompromised individuals, and neonates (Piorunek et al. 2023). The importance of P. multocida as an agent in human disease has been gradually recognized. The capsule, lipopolysaccharide (LPS), and adhesins are the main virulence determinants and immunogenic structures of P. multocida that play a major role in the evasion of the host innate immune barrier and the infection of the host (Boyce and Adler 2000; Harper and Boyce 2017). Vaccine was reported to be an economical and effective antibacterial infection product which could avoid the drawbacks of conventional antibiotic therapy (Wu et al. 2022). Current vaccines against P. multocida are mainly inactivated bacteria and live attenuated bacteria. However, these vaccines have been reported to have limited immunogenicity, reactogenicity, and potential reversion to virulence (Mostaan et al. 2021). Therefore, it would be desirable to develop a novel subunit vaccine that is both safe and protective.
Bacterial membrane vesicles (MVs) were first recognized as being generated through the outer membrane budding process in Gram-negative bacteria, thus generating the designation of outer membrane vesicles (OMVs) (Gan et al. 2021). OMVs can enhance bacterial interactions with the environment, promote bacterial pathogenesis, increase bacterial viability, and modulate interactions within the microbiota (Schwechheimer and Kuehn 2015). OMVs exhibit a spherical morphology and consist of bilayered lipid membrane nanostructures, typically measuring between 20 and 250 nm in size (Kulp and Kuehn 2010). OMVs contain numerous bacterial components, including lipids, proteins, pathogen-associated molecular patterns (PAMPs), adhesins, and peptidoglycan (Kaparakis-Liaskos and Ferrero 2015). The coexistence of bacterial antigens and multiple PAMPs with immunostimulatory properties and the nanoscale structure of protein-lipid complexes make OMVs promising vaccine candidates for the prevention and treatment of bacterial infection (Lieberman 2022; van der Pol et al. 2015).
OMVs have been reported to induce antigen presentation to elicit an antigen-specific immune response by delivering both antigens and adjuvants from PAMPs to antigen-presenting cells (APCs) (Li et al. 2020). Macrophages are one of the important APCs in the innate immune system and perform key functions including phagocytosis, migration, the initiation of inflammatory responses, cytokine production, and presentation of antigens to T cells (Di Benedetto et al. 2019; Mosser and Edwards 2008). These cells express a range of innate immune receptors, including scavenger receptors, mannose-binding lectin (MBL), Fcγ receptors, toll-like receptors (TLRs), and C-type lectin-like receptors (CLRs), which are distributed throughout the cell membrane, cytoplasm, and inner membrane chamber (Mosser and Edwards 2008). During bacterial infection, macrophages recognize PAMPs via surface-exposed, vesicular, or cytoplasmic pattern recognition receptors (PRRs). Then, they phagocytose invading pathogens and release inflammatory mediators to facilitate both innate and adaptive immune responses (Weiss and Schaible 2015).
Owing to their immunogenicity and adjuvanticity, bacterial OMVs have garnered increasing attention as potential vaccines. Research has indicated that OMVs have the ability to elicit a defensive immune response in animals (Roier et al. 2013). To investigate the efficacy of OMVs as vaccinal components for preventing P. multocida infection, we isolated OMVs from P. multocida cultures and explored their morphological traits. Then, we studied the protein composition of the sample using proteomics techniques, which revealed additional information about their contents and potential functions. Finally, we investigated their role in the immune activation of macrophages by analyzing the proliferation, uptake, and cytokine secretion by RAW264.7 macrophages in vitro.
Materials and methods
Bacterial culture and isolation of OMVs
P. multocida (China Veterinary Drug Supervision Institute no. CVCC500) was cultured aerobically in Martin broth (MB) at 37 °C for 18 h, shaken at 200 rpm, replenished with a 1:100 dilution of MB, and allowed to grow for an additional 14 h until the stationary phase was reached. The cultured mixture was centrifuged at 5000 × g for 10 min at 4 °C, and the supernatant was subjected to a second round of centrifugation under the same conditions. The supernatant containing the OMVs was consecutively filtered through 0.45-μm and 0.22-μm filter membranes to remove live bacteria and cell debris. The filtrate was subsequently concentrated using concentrators with a filter membrane of 100 kDa (Millipore, USA). The resulting concentrated filtrate was ultracentrifuged at 100,000 × g for 2 h at 4 °C to pellet the OMVs. Finally, the OMV deposit pellet was resuspended in 1×phosphate-buffered saline (PBS), and the protein concentration was assessed using a bicinchoninic acid (BCA) assay (Solarbio, China). The prepared OMVs were stored at −80 °C until use.
Particle size, concentration, and zeta potential
The particle size–concentration and potential–concentration of the OMVs were measured via NTA (Zetaview, Particle Metrix, Germany). OMVs were diluted with a sterile PBS buffer at a ratio of 1:5000, and the average number of counted particles per frame was approximately 100–120. Then, we selected the NTA measurements from 11 distinct positions for recording and analysis.
The quantity of OMVs per unit (CFU) released by P. multocida was assessed by integrating the quantified number of OMVs using NTA with the count of colonies in the original culture obtained through colony counting.
Transmission electron microscopy
For TEM analysis, OMVs were isolated as previously described, deposited onto a 200-mesh copper grid, and stained with uranyl acetate before they were dried at room temperature. Imaging procedures were carried out using a TecnaiTM G2 Spirit BioTWIN electron microscope operating at 80 kV.
SDS–PAGE
Protein (50 μg) from the entire-cell lysates and protein (50 μg) from OMVs prepared as previously described underwent analysis with sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS–PAGE, 12% resolving gel). The resulting gel was stained with Coomassie Brilliant Blue R-250 dye.
Proteomic analysis of OMVs
Library construction and qualitative proteome analysis were carried out by Novogene Biotech (Beijing, China). The OMV sample containing 50 μg of protein was used for qualitative proteome analysis. OMV samples were collected and mixed with DB protein solution (8 M urea, 100 mM triethylammonium bicarbonate [TEAB]) and subjected to trypsin digestion at 37 °C for 4 h, followed by overnight digestion with trypsin and CaCl2. Subsequently, the digested OMV samples were mixed with formic acid, loaded onto a C18 desalting column, and washed with 0.1% formic acid and 3% acetonitrile washing buffer. This was succeeded by elution using a buffer consisting of 0.1% formic acid and 70% acetonitrile. The eluents were collected and freeze-dried for subsequent LC-MS/MS analysis.
Tryptic peptides were separated using an EASY-nLCTM 1200 UHPLC system (Thermo Fisher, Germany). Peptides eluted from OMV samples were analyzed using a Q ExactiveTM HF-X mass spectrometer (Thermo Fisher, Germany) operating in data-dependent mode with a Nanospray Flex™ (ESI) ion source. The overall scanning range spanned m/z 350 to 1500 with a resolution of 60,000 (at m/z 200). The full scan precursors were selected based on their high-to-low abundance and then fragmented via higher energy collisional dissociation (HCD). The process was carried out with a resolution of 15000 (at m/z 200), and the normalized collision energy was set to 27%. The resulting MS/MS spectra were recorded in the linear ion trap.
The resulting spectra were searched separately against the UniProt Pasteurella_multocida database (1632170-Uniprot-Pasteurella_multocida.fasta (23309 sequences)) using Proteome Discoverer (PD 2.5, Thermo Fisher). The search was performed with a maximum of two allowed missed cleavages, applying carbamidomethyl as a fixed modification, oxidation of methionine (M) as a dynamic modification, and methionine at the N-terminus as a loss. The precursor ion’s mass tolerance was set at 10 ppm, and for production, it was 0.02 Da. PD2.5 was used to analyze the retrieval results, ensuring that the identified peptide spectrum matches (PSMs) exhibited over 99% credibility. The identified proteins were required to contain at least one unique peptide, and all identified PSMs and proteins were subjected to analysis with a false discovery rate (FDR) of no more than 1.0%. Cell-mPLOC 2.0 was used to predict subcellular localization. To perform Gene Ontology (GO) functional analysis, InterproScan (version 5.22-61.0) was employed and compared with the Pfam database (http://pfam.xfam.org/). The identified proteins were subjected to BLAST searches against the online Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.genome.jp/kegg/) and Clusters of Orthologous Groups (COG) database (http://www.ncbi.nlm.nih.gov/COG/) (BLASTP, evalue ≤ 1e-4), after which the results were compared with the highest score to annotate the protein families and pathways.
Cell cultures
The RAW264.7 murine macrophage line was obtained from the Chinese Academy of Science Cell Bank and cultured in DMEM (Gibco, USA) supplemented with 10% FBS, 50 U/mL penicillin (Gibco, USA), and 50 μg/mL streptomycin (Gibco, USA). The cells were maintained at 37 °C and 5% CO2, and the medium was changed every 48 h.
Cell proliferation assay
The impact of OMVs on cell growth was evaluated using CCK-8 assay (APExBIO, USA). RAW264.7 macrophages were initially seeded into 96-well plates at a density of 5.0 × 103 per well and then incubated in a humid environment at 37 °C with 5% CO2 until adherence to the well surface occurred. After this initial phase, the cells were stimulated with 100 μL of fresh medium containing various concentrations of OMVs (62.5 μg/mL, 12.5 μg/mL, 2.5 μg/mL, 0.5 μg/mL, and 0.1 μg/mL), in addition to a positive control of LPS at 10 μg/mL. The negative control consisted of medium alone (referred to as BC). The cell cultures were then incubated for 48 h at 37 °C under 5% CO2. Each concentration was tested in 6 wells. Following incubation, the infected plates were maintained at 37 °C with a 5% CO2 environment for an additional 0.5–2 hours, after which 10 μL of CCK-8 reagent was added to each well. Finally, the growth and proliferation of the cells were determined by measuring the optical density (OD) at 450 nm.
Cellular uptake and colocalization
The pelleted OMVs were fluorescently labeled with DiD (DiIC18(5), Yeasen, China) for 30 min at 37 °C for internalization by macrophage RAW264.7 cells. RAW264.7 cells were plated at a density of 5 × 105 cells per well on 8-well slides (Ibidi, USA) and subsequently incubated at 37 °C in a 5% CO2 environment for 24 h. Then, 100 μL of labeled OMVs was added at a concentration of 5 μg/mL of protein into cells in 200 μL of medium, and the slides were incubated for 1 h, 2 h, and 4 h under 5% CO2 at 37 °C. The supernatant was discarded, and the cells were subjected to three washes with PBS and stained with LysoTracker Green DND-26 (Yeasen, China) to label the lysosomes. After 1 h of incubation, Hoechst 33342 (APExBIO, USA) was added for nuclear staining. After rinsing with PBS, the cells were fixed using 4% paraformaldehyde. Subsequently, the images were analyzed utilizing a confocal laser scanning microscope (Leica TCS-SP5).
To quantitatively and dynamically evaluate cellular uptake, RAW264.7 macrophages were cultured at a density of 5 × 106 cells per well in 6-well plates at 37 °C in 5% CO2 for 24 h. The cells were then exposed to varying concentrations of DiD-labeled OMVs (0.2 μg/mL, 1 μg/mL, and 5 μg/mL) and subsequently incubated at 37 °C in 5% CO2 for 1 h, 2 h, and 4 h. Following this, the cells were treated with trypsin and assessed using flow cytometry to determine the uptake of OMVs.
Cell cytokine secretion
Macrophages can stimulate immune reactions by secreting a diverse array of immune-enhancing cytokines. To investigate whether P. multocida OMVs can induce inflammatory responses in macrophages in an extracellular environment, the concentrations of cytokines released from macrophage RAW264.7 cells were quantified. RAW264.7 cells were plated in 24-well plates at a density of 4 × 105 cells per well and cocultured with varying concentrations of OMVs (62.5 μg/mL, 12.5 μg/mL, 2.5 μg/mL, and 0.5 μg/mL), as well as LPS (10 μg/mL) and medium alone as positive and negative controls, respectively. Six replicates were conducted for each concentration. The cells were then incubated at 37 °C in a 5% CO2 environment for 24 h. Following this, TNF-ɑ, IL-1β, IL-10, TGF-β1, and IL-6 were quantitatively assessed using a mouse ELISA Kit (Lianke Bio, China) after the culture supernatant was collected. The detection process strictly followed the manufacturer’s instructions.
Statistical analysis
The data were analyzed and visualized using GraphPad Prism software (version 9.5) and are presented as the mean ± standard error (SE) deviation. The “n” value represents the number of independent experiments. Each measurement was performed three times and carried out independently in separate experiments (n). Statistically significant results were analyzed using a t-test, and a one-way analysis of variance (ANOVA) was performed with Tukey’s multiple comparison test to assess the differences.
Results
Characterization of OMVs from P. multocida
OMVs were isolated from late logarithmic-phase P. multocida cultures cultivated in MB medium by the ultrafiltration concentration technique, and then the morphology and yield of the derived OMVs were studied.
The size, shape, and structure of the OMVs were visualized using TEM, which is a technique that has been frequently used for the determination of the size, shape, and cellular activities of various biological samples in previous studies (Agrawal et al. 2020). TEM result showed that the purified OMVs exhibited an irregular spherical, bilayer lipid vesicle architecture of variable size, with diameters spanning from 20 to 300 nm (Fig. 1a, b). A study has suggested that OMVs formed through blistering of the outer membrane should consist of a single bilayer membrane. However, our TEM images revealed a bilayer membrane structure for the OMVs. This observation raises the possibility that the OMVs prepared using our current method may indeed represent outer-inner membrane vesicles (OIMVs) generated by explosive cell lysis triggered by phage-derived intracellular lysins (Toyofuku et al. 2019).
Physical characteristics of OMVs derived from P. multocida. Transmission electron micrographs of OMVs (a, b). NTA results of average OMV hydrodynamic diameter and OMV size distribution (c). The zeta potential of the OMVs was measured by NTA (d). The number of OMV particles ascertained by NTA was compared to the number of CFU in bacterial cultures to establish the OMVs number of released per CFU (e). SDS–PAGE analysis was conducted on whole-cell lysates and OMVs, with lane 1 showing a 190-kDa protein ladder, lane 2 representing P. multocida whole-cell lysates, and lane 3 displaying P. multocida OMVs (f)
NTA was chosen for the monitoring and analysis of the size distribution, concentration, and zeta potential of OMVs in this study due to its superior detection limit at low concentrations compared to that of dynamic light scattering (DLS) (Mourdikoudis et al. 2018). Analysis via NTA showed an average diameter of 147.5 nm for OMVs (Fig. 1c). OMVs measuring approximately 10–50 nm in diameter constitute a small fraction of the population, while larger vesicles with diameters of 100–200 nm make up the majority of the population (Fig. 1c). These results support our previous research on the morphological characterization of OMVs via TEM. The concentration of OMVs was 2.6×1011 particles/mL (Fig. 1c), and the zeta potential was −28.86 mV (Fig. 1d). According to the NTA analysis and colony counting from the original liquid culture, an estimated 2.7 OMVs were released per bacterium (Fig. 1e). The data are the average of three independent experiments.
Proteomic analysis of P. multocida OMVs
P. multocida OMVs and whole-cell lysates were analyzed using SDS–PAGE. The results revealed the abundance of P. multocida proteins in the OMVs. Notably, the number of bands observed in the OMVs was lower compared to whole-cell lysates. However, all protein bands identified in the OMVs corresponded to those in the whole-cell lysate (Fig. 1f). These results demonstrated that the produced OMVs were free from bacterial contamination and contained a diverse array of immunogenic proteins.
Additional information for determining the contents and potential functions of P. multocida OMVs was obtained via proteomic analysis. The sequencing of 1500 peptide segments that mapped to 429 proteins of P. multocida OMVs was performed via LC–MS/MS. The major proteins identified according to the PSM data are listed in Table 1. Most of the proteins, including filamentous hemagglutinin (FHA), neuraminidase, pilus assembly proteins, pertactin (PRN), structural proteins, binding proteins, and transport proteins, were associated with virulence. In addition, prion proteins, which transport sugars, amino acids, ions, and transmembrane channel proteins, were also found.
Based on the analysis of subcellular localization, proteins within OMVs originated from diverse subcellular compartments within the bacterium. Notably, the majority of proteins (46.54%) were classified as cytoplasmic proteins, followed by periplasmic proteins (20.74%) and outer membrane proteins (17.51%) (Fig. 2).
To identify the involved metabolic and signaling pathways, KEGG pathway analysis was conducted on the OMV proteins. Among the proteins identified in our study, 3.4% are involved in cellular processes, 9.8% are involved in environmental information processing, and 30.5% are involved in genetic information processing and contribute to metabolism. The largest group of proteins is involved in metabolic pathways, which could be attributed to the biogenetic mechanism of OMVs (Fig. 3). The data acquired indicate associations between the majority of vesicle proteins identified in COG analysis and diverse functions of P. multocida OMVs. Specifically, 66 proteins are linked to translation, ribosomal structure, and biogenesis, while an additional 55 proteins are associated with the cell wall, membrane, and envelope biogenesis. Moreover, other proteins predominantly play roles in carbohydrate and amino acid transport and metabolism, as well as inorganic ion transport and metabolism. Additionally, these proteins are involved in intracellular transport, secretion, vesicle transport, and defense mechanisms (Fig. 4). Subsequently, we conducted GO enrichment analysis of these proteins. The molecular function terms of the identified proteins encompassed various categories, including biological processes linked to OMV proteins enriched in amino acid, lipid, and carbohydrate transport, as well as oxidation–reduction, metabolism, and translation processes. In terms of cellular components, OMV proteins were enriched in ribosomes, cell membranes, outer membranes, the cytoplasm, and the periplasm. For molecular functions, OMV proteins were enriched in nucleotide and ribonucleotide binding, as was ribosome binding (Fig. 5).
OMVs stimulate the proliferation of RAW264.7 cells
In this study, mouse RAW264.7 cells were utilized as the macrophage model due to their established suitability for mimicking macrophage behavior including pinocytosis and phagocytosis (Kong et al. 2019). The proliferation of RAW264.7 macrophages was assessed through CCK-8 assay following exposure to varying concentrations of OMVs. As shown in Fig. 6, RAW264.7 macrophages were stimulated with OMVs ranging from 0.1 to 62.5 μg/mL (n = 6). LPS served as the positive control, while BC was utilized as the negative control. The results indicated that OMVs ranging from 0.1 to 2.5 μg/mL could significantly increase the proliferation of RAW264.7 cells compared with BC (negative control). OMVs ranging from 12.5 to 62.5 μg/mL also showed a promotion effect, but was not significant compared with BC.
OMVs were successfully taken up by RAW264.7 cells
To investigate whether OMVs derived from P. multocida can be efficiently phagocytosed by APCs, confocal laser scanning microscopy and flow cytometry were employed to evaluate the interaction between macrophages and DiD-labeled OMVs.
Confocal laser scanning microscopy images clearly demonstrated the clearly illustrating the successful uptake of P. multocida OMVs by macrophages. After 1 h of incubation, a limited number of DiD-labeled OMVs were observed within the cells, with minimal intracellular localization. However, over the subsequent 2- and 4-h periods, there was a gradual increase in vesicular internalization within the macrophages (Fig. 7). Notably, macrophages exhibited notably high phagocytosis efficiency, with nearly all the cells demonstrating internalization of the vesicles within 4 h of incubation.
Confocal laser scanning microscopy (CLSM) of DiD (red) labeled OMVs from P. multocida uptake by RAW264.7 macrophages at 1 h (up row: 20 μm; down row: 10 μm), 2 h (up row: 20 μm; down row: 10 μm), and 4 h (up row: 20 μm; down row: 10 μm). LysoTracker Green DND-26 (green) was used for lysosome labeling, and Hoechst 33342 (blue) was utilized for nuclear staining. Representative images from three independent experiments are presented
A flow cytometry assay was applied to quantitatively analyze the uptake of DiD-labeled OMVs by RAW264.7 macrophages. Figure 8a shows representative data from three repetitions of different treatments, and Fig. 8b shows the quantitative data. The results revealed rapid interactions after a 1-h incubation period. The phagocytosis rates of the low-, medium-, and high-concentration groups were 5.18%, 20.7%, and 69.3%, respectively. By the 2nd hour, the phagocytosis rates modestly increased to 6.31%, 23.7%, and 74.4%, respectively. After 4 h of incubation, the percentage of cells increased to 8.7%, 37.4%, and 95.5% respectively. The nearly complete uptake of OMVs within the high-concentration group was consistent with previous observations highlighting the substantial phagocytosis efficiency of P. multocida OMVs by RAW264.7 macrophages.
OMVs induced inflammatory cytokine release from RAW264.7 cells
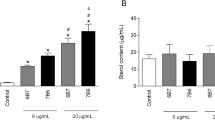
The production of cytokines by RAW264.7 cells stimulated with OMVs was assessed using ELISA to investigate the immune activation effects of OMVs on macrophages in vitro (Fig. 9). Our results demonstrated that significant increases in the production of TNF-ɑ, IL-1β, IL-10, and IL-6 in cells were stimulated with 2.5, 12.5, and 62.5 μg/mL OMV protein, compared to those in the negative control (DMEM) (Fig. 9a–d). Notably, TNF-ɑ and IL-6 by P. multocida OMVs suggest the potential concurrent induction of significant Th1 and Th2 immune responses. Moreover, higher levels of the cytokine TGF-β1 were observed at a higher OMV protein concentration (62.5 μg/mL) (Fig. 9e). These findings highlighted a dose-dependent correlation between the secretion of these five cytokines by RAW264.7 cells in response to OMVs and the capacity of P. multocida OMVs to stimulate cytokine secretion. Importantly, OMVs triggered markedly higher levels of TNF-ɑ secretion compared to those of the other four cytokines.
Discussion
OMVs are primarily enriched in PAMPs, including LPS, phospholipids, nucleic materials, outer membrane–associated proteins, and periplasmic molecules. These substances have the potential to potentially interact with immune cells and initiate a protective immune response (Gan et al. 2021; Tiku and Tan 2021). OMVs have recently shown promising prospects in biomedical applications, for example, as vaccines, adjuvants, drug delivery vehicles, and antibacterial adhesion agents (Acevedo et al. 2014). By investigating the biophysical characteristics, protein composition, and immunomodulatory effect of P. multocida OMVs on immune cells, we aimed to establish a foundation for P. multocida OMV-based vaccine development.
We initially extracted P. multocida OMVs using the ultrafiltration enrichment method. Subsequent analysis via TEM and NTA identified the characteristic double-membrane irregular spherical nanostructure of the P. multocida OMVs. The average diameter was 147.5 nm, which was marginally smaller than that previously reported, suggesting potential variations arising from distinct preparatory conditions or characterization methodologies (Fernández-Rojas et al. 2014).
Proteomic analysis of the full spectrum proteins of P. multocida OMVs aimed to elucidate their specific protein composition and functionality, with the aim of exploring their potential applicability.
Subcellular localization analysis indicated that, in addition to outer membrane proteins, P. multocida OMVs prepared using the ultrafiltration concentration method also contain cytoplasmic, periplasmic, and intracellular proteins. Previously, such observations were attributed to sample contamination with inner membrane proteins and cytoplasmic proteins. However, recent studies have revealed that distinct pathways for MV production result in diverse vesicle subtypes. The explosive cell lysis pathway involves the formation of vesicles through fragment coiling and self-fracturing; these vesicles include proteins from the outer membrane, periplasm, inner membrane, cytoplasm DNA, and peptidoglycan (Toyofuku et al. 2019; Turnbull et al. 2016).
Among the proteins involved in our comprehensive functional analysis, a significant proportion was associated with translation, ribosomal structure, and biogenesis processes, which may have a correlation with the pathway of OMV formation. The subsequent prominent category consisted of proteins involved in cell wall/membrane/envelope biogenesis processes. Studies have shown that the bacterial outer membrane contains transmembrane proteins and lipoproteins, which can act as virulence factors and enzymes, activating the innate immune system and playing a crucial role in the immune response following host infection (Smithers et al. 2021; Verma et al. 2013).
We discovered several immunogenic proteins within the functional category of cell wall/membrane/coating biosynthesis (Supplementary Table S1). Among these proteins, OmpA (A0A2J9QJ69, I2BGA6) exhibited the highest number of PSMs, OmpA originates from the outer membrane, and its molecular weight is 38 kDa. According to the KEGG database, these proteins are classified as opaque proteins or related surface antigens. Moreover, OmpA is a versatile protein with adhesin activity that mediates the formation of bacterial biofilms, stimulates the production of proinflammatory cytokines, and participates in the adhesion and invasion of host cells by bacteria (Dabo et al. 2003; E-Kobon et al. 2017; Katoch et al. 2014; McClean 2012; Yang et al. 2023). Additionally, OmpH (A0MCG2) is a 37.2-kDa molecule originating from the outer membrane. KEGG describes it as a porin outer membrane protein. However, studies have shown that OmpH is a virulence protein that facilitates the diffusion of diverse molecules and possesses specific and cross-reactive epitopes that are abundantly expressed on the bacterial surface. In animal models, OmpH can induce protective immunity against P. multocida (Tan et al. 2010). Furthermore, BamA (A0A191VYV1) is also an immunogenic protein associated with P. multocida antigens, with a molecular weight of 87.7 kDa and originating from the outer membrane. BamA belongs to the outer membrane protein insertase family and is functionally classified as a bacterial surface antigen according to its COG database.
In other COG functional categories, we also identified filamentous hemagglutinin protein (A0A379BCW7) and sialidase (V4PX95). Filamentous hemagglutinin, as classified in the GO system, is a protein associated with pathogenic mechanisms. It serves as a potential virulence factor, facilitating bacterial adhesion to host cells (Tatum et al. 2005). Sialidases, on the other hand, can be generated by various respiratory mucosal pathogens and promote the adhesion and colonization of bacteria (Xu et al. 2009). Notably, both proteins are extracellular, possibly because explosive cell lysis may occur during bacterial culture. After ultracentrifugation, extracellular proteins appear in the OMVs. Furthermore, lipoprotein E (A5H9S0) has also been identified. Lipoprotein E plays a critical role as an immunogenic membrane protein with variable serotype coverage, immunogenicity, antigenicity, and accessible antibodies and is responsible for facilitating complement-mediated killing (Li et al. 2022). Our findings show that P. multocida OMVs are abundant in immunogenic proteins. We also identified the peptide ABC transporter substrate-binding protein (V4NAK3), which is derived from a periplasmic protein and involved in the quorum-sensing process of P. multocida by KEGG. TonB-dependent hemoglobin (Q9CNT8) plays a key role in iron uptake as a virulence factor for bacteria that ensures survival within the host (Krewulak and Vogel 2008). TolC acts as an outer membrane transporter, participating in the type I secretion of macromolecular proteins and the removal of smaller, toxic compounds (Buchanan 2001).
Taken together with previously reported results, KEGG pathway analysis, COG analysis, and GO enrichment analysis indicated that the proteins contained within P. multocida OMVs likely participate in various processes. In addition to activating and regulating the host immune response, these proteins may also modulate the biogenesis of OMVs. A study proposed a new mechanism of OMVs formation in which the peptidoglycan layer of a bacterial cell is weakened by autolysin, allowing the inner membrane to protrude into the periplasm, and the vesicles are eventually pinched away from the cell surface along with the surrounding outer membrane (Toyofuku et al. 2019). Proteins involved in cell wall/membrane biogenesis, including OmpA, OmpW, and MipA (Supplementary Table S1), may play a role in this process (Moon et al. 2012; Lee et al. 2007). Furthermore, the protein cargo of P. multocida OMVs is likely to play a critical role in promoting bacterial survival and facilitating communication with other bacteria. However, additional experiments are necessary to validate these functions.
In summary, our data revealed abundant antigenic proteins associated with virulence and infection mechanisms in isolated P. multocida OMVs. These proteins are crucial for bacterial invasion into the host, suggesting that OMVs could be promising candidates for vaccine development against P. multocida. However, the presence of proteins with unknown functions in P. multocida OMVs highlights the need for further exploration and investigation.
Macrophages play a crucial role in the body’s initial response to pathogen invasion and perform a range of functions such as phagocytosis, migration, cytokine secretion, antigen presentation, and initiation of inflammatory responses (Pidwill et al. 2020; Weiss and Schaible 2015). These functions collectively contribute to the body’s defense against pathogens. We cultured P. multocida OMVs and RAW264.7 macrophages to investigate the potential of P. multocida OMVs to modulate cellular responses in an in vitro setting.
The internalization and eradication of pathogens by macrophages are key parts of the innate immune response and contribute to antigen presentation and the development of acquired immunity. Phagocytosis is an essential process in macrophages (Chen et al. 2023). Through the insights gained from confocal microscopy and flow cytometry, it was observed that P. multocida OMVs were swiftly internalized by RAW264.7 macrophages and were distributed randomly within the lysosomes of the macrophages. This finding underscores the potential for the relevant antigens present on the surface of P. multocida OMVs to be presented, facilitating their recognition and uptake by macrophages, thereby delivering antigen information.
Macrophages, as professional APCs, recognize bacteria through PRRs and subsequently bind bacteria to initiate phagocytosis (Weiss and Schaible 2015). This process allows them to phagocytose and digest pathogens, presenting antigens to T cells through major histocompatibility complex II (MHC-II), establishing a connection between the innate and adaptive immune systems. This sequential process plays a pivotal role in the initiation of adaptive immune responses (Dale et al. 2008). Furthermore, this process triggers intracellular reactions that lead to macrophage proliferation and enhanced immune function (Ren et al. 2017).
Our quantitative analysis of macrophage proliferation induced by P. multocida OMVs using the CCK-8 method yielded evidence indicating that macrophage RAW264.7 cells underwent proliferation upon acquiring antigen information from P. multocida OMVs. This discovery supports the notion that P. multocida OMVs contribute to macrophage proliferation and activation in an in vitro setting. Additionally, upon sensing pathogenic signals, macrophages release cytokines to facilitate both innate and adaptive immune responses (Brown 2006; Weiss and Schaible 2015). Macrophages can be categorized into M1 and M2 types, each of which is involved in distinct immune responses. M1 macrophages are associated with type 1 immune responses and play a role in killing intracellular pathogens, characterized by heightened synthesis of proinflammatory cytokines such as IL-6, TNF-ɑ, and IL-1β. In contrast, M2-type macrophages are involved in immune regulation and promote the type 2 immune response by producing inflammatory chemokines such as TGF-β1 and IL-10 (Zhang and Wang 2014; Shapouri-Moghaddam et al. 2018). The findings revealed that OMVs derived from P. multocida were capable of promoting cytokine secretion in RAW264.7 cells. Notably, compared with the other four cytokines, OMVs significantly increased the secretion of TNF-ɑ in RAW264.7 cells. TNF-ɑ, produced by effector T cells or innate immune cells, has been shown to activate both CD4+ and CD8+ T cells, enhance T-cell proliferation, and increase cytokine production (Mehta et al. 2018). Additionally, the levels of IL-6 were higher than those of the other three cytokines except TNF-ɑ, potentially due to the regulatory effect of TNF-ɑ on stimulating IL-6 secretion. IL-6 serves as a link between the immune responses of T cells and B cells, contributes to the specific differentiation of naive CD4 T cells, and prompts the differentiation of CD8 T cells into cytotoxic T cells. This linkage effectively bridges the innate and adaptive immune responses in the host following P. multocida infection, thereby playing a central role in anti-infection defense. Furthermore, IL-6 stimulates lymphocytes and facilitates the production of antibodies, which are essential for eliminating P. multocida within the host (Tanaka et al. 2014; Tanaka et al. 2016). Furthermore, the results showed that P. multocida OMVs also stimulated the secretion of Th1-type polarized cytokines (IL-1β) and Th2-type polarized cytokines (IL-10 and TGF-β1) in macrophages. IL-1β promotes B-cell proliferation and, in conjunction with antigens, serves as a costimulator of T-cell function (Dinarello 2009). IL-10 activates mast cells and enhances the function of CD8+ T cells, NK cells, and B cells (Moore et al. 2001; Saraiva and O'Garra 2010). TGF-β promotes the production of peripheral (p)Treg, Th17, Th9, and Tfh cells and plays a pivotal role in the development and maturation of immune cells, as well as in regulating the immune response to pathogens (Lodyga and Hinz 2020; Sanjabi et al. 2017). These results provide evidence that the interaction between P. multocida OMVs and macrophages may lead to the presentation of associated antigens on the cell surface, activating the protective immune response of immune cells and mediating and promoting inflammation.
In summary, P. multocida OMVs can modulate cellular responses in vitro, augment phagocytic activity, promote proliferation and internalization, and induce cytokine secretion, thereby promoting the expedited clearance of pathogens. However, further investigations are needed to comprehensively elucidate the intricate regulatory mechanism of immune cells mediated by OMVs. This study provides the molecular and cellular basis for considering P. multocida OMVs as promising subunit vaccine antigen candidates, thereby bolstering the development of novel vaccines against P. multocida infections.
Data availability
The MS proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository with the dataset identifier PXD046592.
References
Acevedo R, Fernández S, Zayas C, Acosta A, Sarmiento ME, Ferro VA, Rosenqvist E, Campa C, Cardoso D, Garcia L, Perez JL (2014) Bacterial outer membrane vesicles and vaccine applications. Front Immunol 5:121. https://doi.org/10.3389/fimmu.2014.00121
Agrawal A, Varshney R, Gattani A, Kirthika P, Khan MH, Singh R, Kodape S, Patel SK, Singh P (2020) Gold nanoparticle based immunochromatographic biosensor for rapid diagnosis of Mycobacterium avium subspecies paratuberculosis infection using recombinant protein. J Microbiol Methods 177:106024. https://doi.org/10.1016/j.mimet.2020.106024
Boyce JD, Adler B (2000) The capsule is a virulence determinant in the pathogenesis of Pasteurella multocida M1404 (B:2). Infect Immun 68(6):3463–3468
Boyce JD, Adler B (2006) How does Pasteurella multocida respond to the host environment? Curr Opin Microbiol 9(1):117–122
Brown GD (2006) Macrophage Receptors and Innate Immunity: Insights from Dectin‐1. In: Innate Immunity to Pulmonary Infection: Novartis Foundation Symposium. Chichester, UK, John Wiley & Sons, Ltd., p 279
Buchanan SK (2001) Type I secretion and multidrug efflux: transport through the TolC channel-tunnel. Trends Biochem Sci 26(1):3–6
Chen S, Saeed AFUH, Liu Q, Jiang Q, Xu H, Xiao GG, Rao L, Duo Y (2023) Macrophages in immunoregulation and therapeutics. Signal Transduct Target Ther 8(1):207. https://doi.org/10.1038/s41392-023-01452-1
Dabo SM, Confer AW, Quijano-Blas RA (2003) Molecular and immunological characterization of Pasteurella multocida serotype A:3 OmpA: evidence of its role in P. multocida interaction with extracellular matrix molecules. Microb Pathog 35(4):147–157
Dale DC, Boxer L, Liles WC (2008) The phagocytes: neutrophils and monocytes. Blood 112(4):935–945. https://doi.org/10.1182/blood-2007-12-077917
Di Benedetto P, Ruscitti P, Vadasz Z, Toubi E, Giacomelli R (2019) Macrophages with regulatory functions, a possible new therapeutic perspective in autoimmune diseases. Autoimmun Rev 18(10):102369. https://doi.org/10.1016/j.autrev.2019.102369
Dinarello CA (2009) Immunological and inflammatory functions of the interleukin-1 family. Annu Rev Immunol 27:519–550. https://doi.org/10.1146/annurev.immunol.021908.132612
E-Kobon T, Leeanan R, Pannoi S, Anuntasomboon P, Thongkamkoon P, Thamchaipenet A (2017) OmpA protein sequence-based typing and virulence-associated gene profiles of Pasteurella multocida isolates associated with bovine haemorrhagic septicaemia and porcine pneumonic pasteurellosis in Thailand. BMC Vet Res 13(1):243. https://doi.org/10.1186/s12917-017-1157-6
Fernández-Rojas MA, Vaca S, Reyes-López M, de la Garza M, Aguilar-Romero F, Zenteno E, Soriano-Vargas E, Negrete-Abascal E (2014) Outer membrane vesicles of Pasteurella multocida contain virulence factors. MicrobiologyOpen 3(5):711–717. https://doi.org/10.1002/mbo3.201
Gan Y, Li C, Peng X, Wu S, Li Y, Tan JPK, Yang YY, Yuan P, Ding X (2021) Fight bacteria with bacteria: bacterial membrane vesicles as vaccines and delivery nanocarriers against bacterial infections. Nanomedicine 35:102398. https://doi.org/10.1016/j.nano.2021.102398
Harper M, Boyce JD (2017) The myriad properties of Pasteurella multocida lipopolysaccharide. Toxins 9(8). https://doi.org/10.3390/toxins9080254
Harper M, Boyce JD, Adler B (2006) Pasteurella multocida pathogenesis: 125 years after Pasteur. FEMS Microbiol Lett 265(1)
Kaparakis-Liaskos M, Ferrero RL (2015) Immune modulation by bacterial outer membrane vesicles. Nat Rev Immunol 15(6):375–387. https://doi.org/10.1038/nri3837
Katoch S, Sharma M, Patil RD, Kumar S, Verma S (2014) In vitro and in vivo pathogenicity studies of Pasteurella multocida strains harbouring different ompA. Vet Res Commun 38(3):183–191. https://doi.org/10.1007/s11259-014-9601-6
Kong L, Smith W, Hao D (2019) Overview of RAW264.7 for osteoclastogensis study: phenotype and stimuli. J Cell Mol Med 23(5):3077–3087. https://doi.org/10.1111/jcmm.14277
Krewulak KD, Vogel HJ (2008) Structural biology of bacterial iron uptake. Biochim Biophys Acta Bioenerg 1778(9):1781–1804
Kulp A, Kuehn MJ (2010) Biological functions and biogenesis of secreted bacterial outer membrane vesicles. Annu Rev Microbiol 64:163–184. https://doi.org/10.1146/annurev.micro.091208.073413
Lee EY, Bang JY, Park GW, Choi DS, Kang JS, Kim HJ, Park KS, Lee JO, Kim YK, Kwon KH, Kim KP, Gho YS (2007) Global proteomic profiling of native outer membrane vesicles derived from Escherichia coli. Proteomics 7(17):3143–3153. https://doi.org/10.1002/pmic.200700196
Li M, Zhou H, Yang C, Wu Y, Zhou X, Liu H, Wang Y (2020) Bacterial outer membrane vesicles as a platform for biomedical applications: an update. J Control Release 323:253–268. https://doi.org/10.1016/j.jconrel.2020.04.031
Li Y, Xiao J, Chang Y-F, Zhang H, Teng Y, Lin W, Li H, Chen W, Zhang X, Xie Q (2022) Immunogenicity and protective efficacy of the recombinant Pasteurella multocida lipoproteins VacJ and PlpE, and outer membrane protein H from P. multocida A:1 in ducks. Front Immunol 13:985993. https://doi.org/10.3389/fimmu.2022.985993
Lieberman LA (2022) Outer membrane vesicles: a bacterial-derived vaccination system. Front Microbiol 13:1029146. https://doi.org/10.3389/fmicb.2022.1029146
Lodyga M, Hinz B (2020) TGF-β1 - a truly transforming growth factor in fibrosis and immunity. Semin Cell Dev Biol 101:123–139. https://doi.org/10.1016/j.semcdb.2019.12.010
McClean S (2012) Eight stranded β -barrel and related outer membrane proteins: role in bacterial pathogenesis. Protein Pept Lett 19(10):1013–1025
Mehta AK, Gracias DT, Croft M (2018) TNF activity and T cells. Cytokine 101:14–18. https://doi.org/10.1016/j.cyto.2016.08.003
Moon DC, Choi CH, Lee JH, Choi C-W, Kim H-Y, Park JS, Kim SI, Lee JC (2012) Acinetobacter baumannii outer membrane protein A modulates the biogenesis of outer membrane vesicles. J Microbiol 50(1):155–160. https://doi.org/10.1007/s12275-012-1589-4
Moore KW, de Waal MR, Coffman RL, O'Garra A (2001) Interleukin-10 and the interleukin-10 receptor. Annu Rev Immunol 19:683–765
Mosser DM, Edwards JP (2008) Exploring the full spectrum of macrophage activation. Nat Rev Immunol 8(12):958–969. https://doi.org/10.1038/nri2448
Mostaan S, Ghasemzadeh A, Asadi Karam MR, Ehsani P, Sardari S, Shokrgozar MA, Abolhassani M, Nikbakht Brujeni G (2021) Pasteurella multocida plpE protein polytope as a potential subunit vaccine candidate. Vector Borne Zoonotic Dis 21(11):870–874. https://doi.org/10.1089/vbz.2020.2758
Mourdikoudis S, Pallares RM, Thanh NTK (2018) Characterization techniques for nanoparticles: comparison and complementarity upon studying nanoparticle properties. Nanoscale 10(27):12871–12934. https://doi.org/10.1039/c8nr02278j
Pidwill GR, Gibson JF, Cole J, Renshaw SA, Foster SJ (2020) The role of macrophages in staphylococcus aureus infection. Front Immunol 11:620339. https://doi.org/10.3389/fimmu.2020.620339
Piorunek M, Brajer-Luftmann B, Trafas T, Schneider A, Walkowiak J (2023) Lower respiratory infection in humans caused by pasteurella multocida. Respir Physiol Neurobiol 315:104091. https://doi.org/10.1016/j.resp.2023.104091
Ren Y, Khan FA, Pandupuspitasari NS, Zhang S (2017) Immune evasion strategies of pathogens in macrophages: the potential for limiting pathogen transmission. Curr Issues Mol Biol 21:21–40
Roier S, Fenninger JC, Leitner DR, Rechberger GN, Reidl J, Schild S (2013) Immunogenicity of Pasteurella multocida and Mannheimia haemolytica outer membrane vesicles. Int J Med Microbiol 303(5):247–256. https://doi.org/10.1016/j.ijmm.2013.05.001
Sanjabi S, Oh SA, Li MO (2017) Regulation of the immune response by TGF-β: from conception to autoimmunity and infection. Cold Spring Harb Perspect Biol 9(6). https://doi.org/10.1101/cshperspect.a022236
Saraiva M, O'Garra A (2010) The regulation of IL-10 production by immune cells. Nat Rev Immunol 10(3):170–181. https://doi.org/10.1038/nri2711
Schwechheimer C, Kuehn MJ (2015) Outer-membrane vesicles from Gram-negative bacteria: biogenesis and functions. Nat Rev Microbiol 13(10):605–619. https://doi.org/10.1038/nrmicro3525
Shapouri-Moghaddam A, Mohammadian S, Vazini H, Taghadosi M, Esmaeili S-A, Mardani F, Seifi B, Mohammadi A, Afshari JT, Sahebkar A (2018) Macrophage plasticity, polarization, and function in health and disease. J Cell Physiol 233(9):6425–6440. https://doi.org/10.1002/jcp.26429
Smithers L, Olatunji S, Caffrey M (2021) Bacterial lipoprotein posttranslational modifications. new insights and opportunities for antibiotic and vaccine development. Front Microbiol 12:788445. https://doi.org/10.3389/fmicb.2021.788445
Tan HY, Nagoor NH, Sekaran SD (2010) Cloning, expression and protective capacity of 37 kDa outer membrane protein gene (ompH) of Pasteurella multocida serotype B:2. Trop Biomed 27(3):430–441
Tanaka T, Narazaki M, Kishimoto T (2014) IL-6 in inflammation, immunity, and disease. Cold Spring Harbor Perspect Biol 6(10):a016295. https://doi.org/10.1101/cshperspect.a016295
Tanaka T, Narazaki M, Masuda K, Kishimoto T (2016) Regulation of IL-6 in immunity and diseases. Adv Exp Med Biol 941:79–88
Tatum FM, Yersin AG, Briggs RE (2005) Construction and virulence of a Pasteurella multocida fhaB2 mutant in turkeys. Microb Pathog 39(1-2)
Tiku V, Tan M-W (2021) Host immunity and cellular responses to bacterial outer membrane vesicles. Trends Immunol 42(11):1024–1036. https://doi.org/10.1016/j.it.2021.09.006
Toyofuku M, Nomura N, Eberl L (2019) Types and origins of bacterial membrane vesicles. Nat Rev Microbiol 17(1):13–24. https://doi.org/10.1038/s41579-018-0112-2
Turnbull L, Toyofuku M, Hynen AL, Kurosawa M, Pessi G, Petty NK, Osvath SR, Cárcamo-Oyarce G, Gloag ES, Shimoni R, Omasits U, Ito S, Yap X, Monahan LG, Cavaliere R, Ahrens CH, Charles IG, Nomura N, Eberl L, Whitchurch CB (2016) Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat Commun 7:11220. https://doi.org/10.1038/ncomms11220
van der Pol L, Stork M, van der Ley P (2015) Outer membrane vesicles as platform vaccine technology. Biotechnol J 10(11):1689–1706. https://doi.org/10.1002/biot.201400395
Verma S, Sharma M, Katoch S, Verma L, Kumar S, Dogra V, Chahota R, Dhar P, Singh G (2013) Profiling of virulence associated genes of Pasteurella multocida isolated from cattle. Vet Res Commun 37(1):83–89. https://doi.org/10.1007/s11259-012-9539-5
Weiss G, Schaible UE (2015) Macrophage defense mechanisms against intracellular bacteria. Immunol Rev 264(1):182–203. https://doi.org/10.1111/imr.12266
Wilkie IW, Harper M, Boyce JD, Adler B (2012) Pasteurella multocida: diseases and pathogenesis. Curr Top Microbiol Immunol 361. https://doi.org/10.1007/82_2012_216
Wilson BA, Ho M (2013) Pasteurella multocida: from zoonosis to cellular microbiology. Clin Microbiol Rev 26(3):631–655. https://doi.org/10.1128/CMR.00024-13
Wu Y, Deng G, Song Z, Zhang K, Deng J, Jiang K, Han H (2022) Enhancing antibacterial immunotherapy for bacterial pneumonia via nanovaccines coated with outer membrane vesicles. Chem Eng J 436. https://doi.org/10.1016/j.cej.2022.135040
Xu G, Ryan C, Kiefel MJ, Wilson JC, Taylor GL (2009) Structural studies on the Pseudomonas aeruginosa sialidase-like enzyme PA2794 suggest substrate and mechanistic variations. J Mol Biol 386(3):828–840. https://doi.org/10.1016/j.jmb.2008.12.084
Yang X, Fu Q, Zhang W, An Q, Zhang Z, Li H, Chen X, Chen Z, Cheng Y, Chen S, Man C, Du L, Chen Q, Wang F (2023) Overexpression of Pasteurella multocida OmpA induces transcriptional changes and its possible implications for the macrophage polarization. Microb Pathog:106212. https://doi.org/10.1016/j.micpath.2023.106212
Zhang L, Wang C-C (2014) Inflammatory response of macrophages in infection. Hepatob Pancreatic Dis Int 13(2):138–152
Acknowledgements
This work was supported by “Pioneer” and “Leading Goose” R&D Program of Zhejiang (Grant No. 2023C02047), National Natural Science Foundation of China (Grant No. 32002323), China Agriculture Research System of MOF and MARA (Grant No. CARS-43-C-2), and the Key Research and Development Program of Zhejiang Province (No. 2021C02007 and No. 2019C02052).
Author information
Authors and Affiliations
Contributions
YH, YL, and GB conceived and designed the research. JS, YH, XL, and XX conducted the experiments. JS and FH collected the samples. JS and YH analyzed the data. JS, YH, and CC edited the article. YH, YL, and CC critically reviewed the manuscript. XC and QJ contributed reagents and materials. All the authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
Table S1 List of the functions of the identified important P. multocida OMV proteins are described in order of decreasing PSMs. (XLSX 13 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, J., Huang, Y., Li, X. et al. Characterization and immunological effect of outer membrane vesicles from Pasteurella multocida on macrophages. Appl Microbiol Biotechnol 108, 238 (2024). https://doi.org/10.1007/s00253-024-13060-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13060-2