Abstract
Anaerobic, acetogenic bacteria are promising biocatalysts for a sustainable bioeconomy since they capture and convert carbon dioxide to acetic acid. Hydrogen is an intermediate in acetate formation from organic as well as C1 substrates. Here, we analyzed mutants of the model acetogen Acetobacterium woodii in which either one of the two hydrogenases or both together were genetically deleted. In resting cells of the double mutant, hydrogen formation from fructose was completely abolished and carbon was redirected largely to lactate. The lactate/fructose and lactate/acetate ratios were 1.24 and 2.76, respectively. We then tested for lactate formation from methyl groups (derived from glycine betaine) and carbon monoxide. Indeed, also under these conditions lactate and acetate were formed in equimolar amounts with a lactate/acetate ratio of 1.13. When the electron-bifurcating lactate dehydrogenase/ETF complex was genetically deleted, lactate formation was completely abolished. These experiments demonstrate the capability of A. woodii to produce lactate from fructose but also from promising C1 substrates, methyl groups and carbon monoxide. This adds an important milestone towards generation of a value chain leading from CO2 to value-added compounds.
Key points
• Resting cells of the ΔhydBA/hdcr mutant of Acetobacterium woodii produced lactate from fructose or methyl groups + CO
• Lactate formation from methyl groups + CO was completely abolished after deletion of lctBCD
• Metabolic engineering of a homoacetogen to lactate formation gives a potential for industrial applications
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
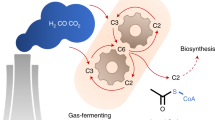
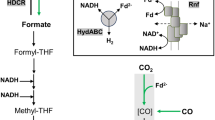
Acetogenic bacteria are a group of strictly anaerobic bacteria that oxidize one mol of hexoses such as fructose to three mol of acetate, a metabolic trait known as homoacetogenesis (Fontaine et al. 1942). Fructose is oxidized via the Embden-Meyerhof-Parnas pathway to four electrons and two mol of pyruvate which are further oxidized to two mol of acetyl-CoA, CO2 and four more electrons (Ragsdale 2003). Acetate formation yields 4 mol of ATP per hexose, the highest amount of ATP that can be obtained by fermentation (Müller 2008; Müller and Frerichs 2013). This is only possible by disposing the electrons in a special pathway for CO2 reduction to acetate, the Wood-Ljungdahl pathway (WLP) in which two CO2 are reduced by eight electrons to acetate (Müller 2003; Wood and Ljungdahl 1991). The WLP is not only an electron sink for fructose oxidation, but also allows acetogens to grow on H2 + CO2 (Schuchmann and Müller 2014; Wood et al. 1986) or other C1 compounds such as formate (Moon et al. 2021) or methanol (Balk et al. 2003; Kremp and Müller 2021; Kremp et al. 2018; van der Meijden et al. 1984). CO2 is reduced in two branches. In the methyl branch, one CO2 is first reduced to formate by a formate dehydrogenase, or more specific, by a hydrogen-dependent CO2 reductase in the model acetogen Acetobacterium woodii (Schuchmann and Müller 2013). Formate is then bound in an ATP-dependent reaction to the C1 carrier tetrahydrofolate (THF) (Himes and Harmony 1973; Lovell et al. 1988), yielding formyl-THF from which water is eliminated and the resulting methenyl-THF is reduced via methylene- to methyl-THF (Bertsch et al. 2015; Ragsdale and Ljungdahl 1984). In the second branch, CO2 is reduced to CO which is then bound to the key enzyme of the pathway, CO dehydrogenase/acetyl-CoA synthase (CODH/ACS) and combined with the methyl group and CoA to acetyl-CoA (Ragsdale 2008). The substrates formate (Moon et al. 2021) and carbon monoxide (Diekert and Thauer 1978; Diender et al. 2015; Genthner and Bryant 1982; Weghoff and Müller 2016) are intermediates of the pathway and methyl groups from, for example, methanol or glycine betaine, enter the pathway by a methyltransferase system yielding methyl-THF (Kremp and Müller 2021; Kremp et al. 2018; Lechtenfeld et al. 2018).
Acetogenic bacteria have gained much interest in recent years since they capture the greenhouse gas CO2 and reduce it to acetate. This small chain fatty acid has limited application per se, but acetate may substitute glucose in the long run to a sustainable bioeconomy as feedstock for the production of not only biofuels but also all the other products that are currently produced from sugars by, for example, Escherichia coli, Corynebacterium glutamicum or yeasts (Förster and Gescher 2014; Ingram et al. 1987; Inui et al. 2004a, b; Jojima et al. 2015a, b; Lim et al. 2018; Mohd Azhar et al. 2017). In addition to acetate, some acetogens can produce ethanol from C1 compounds such as CO2 and CO and this process is already used on an industrial scale (Liew et al. 2017, 2022; Mock et al. 2015). Higher carbon compounds are rarely produced and generally not from C1 compounds. A C1 substrate of interest is methanol which is also used by acetogens as carbon and energy source (Kremp and Müller 2021; Kremp et al. 2018; van der Meijden et al. 1984). Methanol is already produced from CO2 chemically on an industrial level and the use of methanol as a feedstock circumvents all the problems inherent to gas fermentation.
Recently, we discovered a novel metabolic trait in A. woodii, mixed acid fermentation of fructose (Moon et al. 2023a). A mutant in which the central enzyme of the WLP, the methylene-tetrahydrofolate reductase was genetically deleted, was able to grow on fructose. But acetate was not the only product; in addition molecular hydrogen, formate, ethanol and lactate were produced as end products (Moon et al. 2023a). This finding offered the possibility to engineer strains that convert fructose or even C1 compounds to reduced end products such as ethanol or lactate. Production of lactate is of great interest since it is widely used in food, pharma- and cosmetic industries as well as serves as the precursor of a biologically degradable plastic, poly lactic acid (PLA) (Ahmad et al. 2022). Here, we have chosen lactate as a target and generated a strain of A. woodii that performs heterolactate fermentation from fructose or from methyl groups plus carbon monoxide.
Materials and methods
Strains and cultivation
A. woodii wild type (DSM1030) was obtained from the Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSMZ; Braunschweig, Germany). The ∆pyrE strain was described before (Wiechmann et al. 2020). The hdcr deletion mutant ∆hdcr and the double mutant ∆hydBA/hdcr were described recently (Moon et al. 2023b). The triple mutant ∆hydBA/hdcr/lctBCD in which the genes encoding the lactate dehydrogenase were deleted in addition was generated in this study. All strains were routinely cultivated under anoxic conditions at 30 °C in bicarbonate-buffered complex medium as described before (Heise et al. 1989). As substrates for growth, 60 mM fructose + 100 mM formate, or 50 mM glycine betaine + 10% CO were used. Growth was monitored by determining the optical density at 600 nm (OD600).
Generation of A. woodii ΔhydBA/hdcr/lctBCD mutant
To generate the ΔhydBA/hdcr/lctBCD triple mutant, the plasmid pMTL84151_LW_dlct was constructed in E. coli HB101 (Promega, Madison, WI, USA) and transformed into the A. woodii ΔhydBA/hdcr strain (Moon et al. 2023b), as described previously (Westphal et al. 2018). The plasmid pMTL84151_LW_dlct originated from pMTL84151 (Heap et al. 2009) but lacks a Gram-positive replicon. In pMTL84151_LW_dlct, 1000 bp of upstream flanking regions (UFR) of lctB (Awo_c08710) and 1000 bp of downstream flanking regions (DFR) of lctD (Awo_c08730) were cloned into the multiple cloning sites to delete the lctBCD genes by homologous recombination. The plasmid also has a catP marker from Clostridium perfringens coding for chloramphenicol/thiamphenicol resistance (Werner et al. 1977) and a heterologous pyrE gene from Eubacterium limosum (Wiechmann et al. 2020) as a counter selectable marker. The first selection was carried out on an agar plate with complex medium containing 20 mM fructose + 50 mM formate and 30 ng/µl thiamphenicol after transformation of pMTL84151_LW_dlct into the A. woodii ΔhydBA/hdcr strain by electroporation (625 V, 25 µF, 600 Ω, in 1 mm cuvettes). The second selection for disintegration was performed on an agar plate with minimal medium (Westphal et al. 2018) containing 20 mM fructose + 50 mM formate, 1 µg/ml uracil and 1 mg/ml 5-fluoroortate (5-FOA). The deleted region was analyzed by PCR with primers binding upstream of UFR and downstream of DFR: aus_lct_for (5′-CAGGCAATGTTTTTTAATGTCAGGA-3′) and aus_lct_rev (5′-ATAACTTTTGCCAAAGCCACAAT-3′). Consequently, PCR experiments were performed to verify the purity of the mutant, with primers binding in the lctD gene: in_lct_for (5′-GGTAATATCAGTACGAATGCCGG-3′) and in_lct_rev (5′- GAATCGCCTTGGATTTAATAATCTTCG-3′). Subsequently, the sequence of the deleted region of the mutant was verified by DNA sequencing (Sanger et al. 1977).
Preparation of resting cells
Cells were cultivated either on 60 mM fructose + 100 mM formate or 50 mM glycine betaine + 10% CO in 1 l bicarbonate-buffered complex medium to the late exponential growth phase (on 60 mM fructose + 100 mM formate, OD600 of 1.5; on 50 mM glycine betaine + 10% CO, OD600 of 0.7). Cells were harvested by centrifugation (Avanti J-25 and JA-10 Fixed-Angle Rotor; Beckman Coulter, Brea, CA, United States) at 8,000 rpm and 4 °C for 10 min, washed with 30 ml of buffer containing 50 mM imidazole (pH 7.0), 20 mM KCl, 20 mM MgSO4, 4 mM DTE and 4 µM resazurin and pelleted by centrifugation at 8,500 rpm and 4 °C for 10 min (Avanti J-25 and JA-25.50 Fixed-Angle Rotor; Beckman Coulter, Brea, CA, United States). Subsequently, the pellets were resuspended in 5 ml imidazole buffer and transferred to 16-ml Hungate tubes. All steps were performed under strictly anoxic conditions in an anoxic chamber (Coy Laboratory Products, Grass Lake, MI, United States) filled with N2/H2 (96–98%/2–4%; v/v). To get rid of residual H2 from the anoxic chamber, the gas phase of the cell suspensions was changed to 100% N2. The total protein concentration of the cell suspensions was measured as described before (Schmidt et al. 1963).
Cell suspension experiments
For fructose fermentation, the cells were resuspended in 20 ml of bicarbonate-containing imidazole buffer (50 mM imidazole, 20 mM KCl, 20 mM NaCl, 20 mM MgSO4, 60 mM KHCO3, 4 mM DTE, 4 µM resazurin, pH 7.0) in 120-ml serum flasks under a N2/CO2 atmosphere (80:20, v/v) to a final protein concentration of 2 mg/ml. As substrate, 60 mM fructose was added. For glycine betaine + CO fermentation, resting cells were prepared in 10 ml of bicarbonate-containing imidazole buffer under a N2/CO2/CO atmosphere (2 bar, 72:18:10, v/v/v) to a final protein concentration of 1 mg/ml. For the experiment under bicarbonate-depleted conditions, bicarbonate-depleted buffer (50 mM imidazole, 20 mM KCl, 20 mM NaCl, 20 mM MgSO4, 4 mM DTE, 4 µM resazurin, pH 7.0) was used and the gas phase was replaced to a N2/CO atmosphere (2 bar, 90:10, v/v). For the experiments under Na+-depleted conditions, Na+-depleted buffer (50 mM imidazole, 20 mM KCl, 20 mM MgSO4, 60 mM KHCO3, 4 mM DTE, 4 µM resazurin, pH 7.0) was used and the contaminating Na+ concentration in the buffer was determined with an Orion 84–111 ROSS sodium electrode (Thermo Electron, Witchford, UK) according to the supplier's instructions. As substrate, 50 mM glycine betaine was added to the resting cells. The resting cells were pre-incubated at 30 °C in a water bath with shaking (150 rpm) and the experiments were started by adding the substrate(s). During the experiments, 1-ml samples were routinely taken for metabolite analyses.
Metabolite analyses
The concentrations of fructose, formate, acetate, and lactate were determined by high-performance liquid chromatography as described previously (Moon et al. 2019). H2 or ethanol were analyzed by gas chromatography (Trifunović et al. 2016; Weghoff and Müller 2016).
Gene expression analyses
The ∆pyrE, ∆hdcr, ∆hydBA/hdcr mutants grown on 50 mM glycine betaine under a N2/CO2/CO atmosphere (72:18:10, v/v/v) in bicarbonate-buffered complex media were harvested in the exponential growth phase. Preparation of RNA and cDNA was performed as described before (Dönig and Müller 2018). Transcript levels of the lctB, lctC, and lctD genes were analyzed with real-time qPCR in a Rotor Gene RG-3000 qPCR cycler (Corbett Research, Cambridge, UK) using Maxima SYBR Green qPCR Master Mix (Thermo Fisher Scientific, Waltham, MA, USA) following the supplier's instructions. The housekeeping gene gyrA (Awo_c00060) was used as reference and the relative gene expression levels were calculated using the 2−ΔΔCt method (Livak and Schmittgen 2001). For the amplification, following primers were used: qlctB_for (5′-GCGCTGATGAGGGTTGTTTA-3′) and qlctB_rev (5′-TCACCCAATCGTTTGGTG-3′) for lctB, qlctC_for (5′-GTCGATCATATTGAAGGCCAGAT-3′) and qlctC_rev (5′-ACAAGGCATAAACCGGATGT-3′) for lctC, and qlctD_for (5′-GATTCCAACGGCGATTGAAT-3′) and qlctD_rev (5′-TATAAGCGTTGCTACTGGAGTC-3′) for lctD.
Results
Strain design
There are two hydrogenases encoded in the genome of A. woodii, the HydA2 subunit of the HDCR and the electron-bifurcating HydABC hydrogenase (Poehlein et al. 2012); both have been deleted solely or in tandem (Moon et al. 2023b; Wiechmann et al. 2020). There is only one known lactate dehydrogenase in A. woodii, the electron bifurcating LDH/ETH complex, encoded by lctBCD (Awo_c08710 – Awo_c08730) (Poehlein et al. 2012). This enzyme complex is known to be responsible for lactate oxidation during growth of A. woodii on lactate (Weghoff et al. 2015). Recently, it has been reported that the lctBCD genes were highly expressed in the ΔmetVF mutant grown on fructose where lactate was formed as a side product (Moon et al. 2023a). Therefore, to verify that a possible lactate formation was indeed catalyzed by LctBCD we genetically deleted the LDH/ETF complex. For the generation of the ΔhydBA/hdcr/lctBCD mutant, the suicide plasmid pMTL_84151_LW_dlct was constructed, which contains each 1000 bp of upstream flanking region (UFR) of lctB and downstream flanking region (DFR) of lctD leaving only the start codon of lctB and the stop codon of lctD (Fig. 1a). For selection, this plasmid carries the pyrE gene from Eubacterium limosum (Wiechmann et al. 2020) and the chloramphenicol/thiamphenicol resistance cassette (catP) from Clostridium perfringens (Werner et al. 1977). The plasmid was integrated into the chromosome of the ΔhydBA/hdcr mutant by homologous recombination at one flanking region in the presence of thiamphenicol and subsequently, disintegration was carried out by counter-selection with 5-fluoroorotate. Single colonies were picked on agar plates with fructose + formate as carbon and energy source. PCR experiments with primers binding outside the deleted region revealed that the lctBCD genes were successfully deleted (Fig. 1b), and the lctD gene could not be amplified with primers binding inside of lctD (Fig. 1c). Subsequently, the absence of the lctBCD genes in the chromosome was confirmed by DNA sequencing (Sanger et al. 1977).
Deletion of the lctBCD genes in the chromosome of the ΔhydBA/hdcr mutant. (a) Genetic organization after deletion of the lctBCD genes using plasmid pMTL_LW_dlct. Only 3 bp of the lctB gene and 3 bp of the lctD gene remained in the ΔhydB/hdcr/lctBCD mutant. Genotypic analyses of the ΔhydBA/hdcr/lctBCD mutant were carried out by colony PCR with primers binding outside the deleted region (b) (aus_lct_for and aus_lct_rev) or inside (c) (in_lct_for and in_lct_rev)
Heterolactate fermentation with fructose in the ∆hydBA/hdcr double mutant
In a previous study we have found conversion of fructose to molecular hydrogen, formate, ethanol and lactate as end products in a ΔmetVF mutant of A. woodii (Moon et al. 2023a). Here, we aimed to redirect fructose metabolism to lactate. Since ethanol was only produced in very minor amounts, and since A. woodii has eleven different alcohol dehydrogenases, it was not attempted to genetically delete ethanol production. Hydrogen was produced in huge amounts (Moon et al. 2023a) and therefore we analyzed whether H2 production would be abolished in the ∆hdcr, and the ΔhydBA/hdcr double mutant. The growth phenotype of these mutants has been described before; in brief, they do not grow on fructose, H2 + CO2, methanol, or formate (Moon et al. 2023b). Therefore, the mutants were grown on fructose + formate, harvested in the exponential growth phase and we then analyzed the fermentation balance from fructose in resting cells. Since we have seen that high concentrations of sugars stimulated production of a reduced end product, ethanol, under certain conditions (Moon and Müller 2021), we performed the experiments with 60 mM instead of 20 mM fructose.
Upon addition of fructose to resting cells of the ∆hdcr mutant, 21.4 ± 1.4 mM fructose was consumed, and 21.0 ± 0.4 mM acetate was produced, giving a fructose:acetate ratio of 1:1 (Fig. 2a). Formate was not produced, as expected. As seen before with the ΔmetVF mutant (Moon et al. 2023a), hydrogen was still formed in huge amounts (45.4 ± 2.2 mM) with a fructose:H2 ratio of 1:2.1. Ethanol (1.5 ± 0.0 mM) and lactate (2.1 ± 0.9 mM) were only formed in very minor amounts. Since electrons were apparently released as hydrogen, we checked the effect of deletion of the hydrogenase HydABC in the ∆hdcr background. In resting cells of the ∆hydBA/hdcr mutant, hydrogen formation was completely abolished, and less acetate (14.0 ± 2.5 mM) was produced from 31.1 ± 1.1 mM fructose with a fructose:acetate ratio of only 1:0.45 (Fig. 2b). Ethanol formation increased a bit (4.5 ± 0.6 mM) with a fructose:ethanol ratio of 1:0.14 and an acetate:ethanol ratio of 1:0.32. In contrast, lactate production increased dramatically from almost zero to 38.6 ± 2.1 mM, giving a fructose:lactate ratio of 1:1.24 and an acetate:lactate ratio of 1:2.76.
Conversion of fructose in resting cells of A. woodii. Cells of the Δhdcr (a), ΔhydBA/hdcr (b) and ΔhydBA/hdcr/lctBCD mutants (c) were grown in bicarbonate-buffered complex media under a N2/CO2 atmosphere (80:20, v/v) with 60 mM fructose + 100 mM formate and harvested in the early stationary growth phase. The cell suspensions were prepared in 10 ml of cell suspension buffer (50 mM imidazole, 20 mM MgSO4, 20 mM KCl, 20 mM NaCl, 60 mM KHCO3, pH 7.0) in 120 ml serum flasks under a N2/CO2 atmosphere at a final protein concentration of 2 mg/ml. 60 mM fructose was given to the cell suspensions as carbon and energy source. Fructose (●), acetate (■), ethanol (▲), formate (▼), H2 (♦) and lactate ( ×) were determined. Each data point presents a mean with standard deviation (SD); n = 2 independent experiments
Since the lctBCD genes are the only genes annotated to encode a lactate dehydrogenase (Poehlein et al. 2012), we expected a complete loss of lactate formation and increase in ethanol production in the triple mutant ∆hydBA/hdcr/lctBCD. However, this was not observed. Lactate production had a longer lag phase of around 8 h, compared to the double mutant, but lactate was then produced with rates and yields similar to the double mutant (Fig. 2c).
Lactate formation from glycine betaine and carbon monoxide
Next, we analyzed whether cells would produce lactate from C1 compounds. The wild type of A. woodii was shown to grow on methanol + CO which are converted to acetate; the methyl-group and CO are intermediates of the WLP which are condensed by CODH/ACS to acetyl-CoA (Litty et al. 2022). The HDCR is not involved in that metabolism. Since the ∆hdcr and the ∆hydBA/hdcr mutants do not grow on methanol (Moon et al. 2023b) regardless of the presence or absence of CO, we tested for growth on another methyl group-containing substrate, glycine betaine, that A. woodii can use as carbon and energy source (Lechtenfeld et al. 2018). We recently showed that the ∆hdcr and the ∆hydBA/hdcr mutants grow on glycine betaine and produce formate as final product alongside acetate (Moon et al. 2023b). Glycine betaine serves as methyl group donor and dimethylglycine is excreted by the cells (Lechtenfeld et al. 2018). The ∆pyrE as well as the ∆hdcr, ∆hydBA/hdcr, ∆hydBA/hdcr/lctBCD mutants grew well on 50 mM glycine betaine + CO and produced only acetate (∆pyrE, 47.5 ± 1.3 mM; ∆hdcr, 46.6 ± 2.1 mM; ∆hydBA/hdcr, 44.9 ± 1.8 mM; ∆hydBA/hdcr/lctBCD, 45.5 ± 0.6 mM) via the WLP similar to growth on methanol + CO (Litty et al. 2022) (Fig. 3). We then checked for product formation in resting cells. Resting cells of the ∆pyrE strain produced 50.9 ± 1.6 mM acetate from 50 mM glycine betaine and CO (Fig. 4a) and the same was true for the HDCR mutant (48.2 ± 1.0 mM) (Fig. 4b), as expected. Cells produced hydrogen (0.5 mM in both strains), most likely from CO oxidation. CO oxidation is coupled to reduction of ferredoxin followed by the production of molecular hydrogen in two steps: first, reduced ferredoxin is reoxidized by the Rnf complex (with reduction of NAD) (Hess et al. 2013) and the electron-bifurcating hydrogenase then forms hydrogen from reduced ferredoxin and NADH (Schuchmann and Müller 2012). Therefore, we reasoned that deletion of the electron bifurcating hydrogenase should redirect electrons to another acceptor. Indeed, resting cells of the ∆hydBA/hdcr double mutant no longer produced H2 but lactate instead, alongside with acetate (Fig. 4c). Acetate production was a bit faster, but final acetate and lactate concentrations were similar. From 50 mM glycine betaine + CO, 18.1 ± 1.1 mM acetate and 20.4 ± 0.5 mM lactate were formed with an acetate:lactate ratio of 1:1.1. As a minor product, we also detected 2.5 mM ethanol. In agreement with the lactate production, we found that the lctBCD genes were highly upregulated in the ∆hydBA/hdcr mutant during glycine betaine + CO fermentation (Fig. 5). Compared to the ∆pyrE strain, the lctB gene in the ∆hydBA/hdcr mutant was upregulated with a log2 fold change of 9.9 ± 0.6. The same was true for the lctC gene with a log2 fold change of 10.0 ± 0.2 and the lctD gene with a log2 fold change of 11.0 ± 0.3. Lactate must have been formed from acetyl-CoA via carboxylation to pyruvate by pyruvate:ferredoxin oxidoreductase (PFOR), and indeed, a reduced lactate formation was observed under CO2/bicarbonate-depleted conditions (Fig. 6a) compared to CO2/bicarbonate-rich conditions (cf. Figure 4c). Since NADH is required for lactate production by the LDH/ETF complex, the Rnf complex must be involved i.e., the lactate production must be Na+ dependent. Indeed, lactate production (cf. Figure 4c) was completely abolished in the absence of NaCl and the ∆hydBA/hdcr mutant produced only acetate (44.0 ± 1.8 mM) (Fig. 6b).
Growth of the A. woodii strains on glycine betaine + CO. Growth experiments were performed in 20 ml bicarbonate-buffered complex medium in 120-ml serum flasks with 50 mM glycine betaine under a N2/CO2/CO atmosphere (72:18:10, v/v/v) at 30 °C. Depicted are the optical densities of the ∆pyrE (●), ∆hdcr (■), ∆hydBA/hdcr (▲), and the ∆hydBA/hdcr/lctBCD mutant (▼). Additionally, acetate (open symbols) was determined during growth. Each data point presents a mean ± SD; n = 2 independent experiments
Conversion of glycine betaine + CO in resting cells of A. woodii. Cells of the ΔpyrE (a), Δhdcr (b) and ΔhydBA/hdcr mutants (c) were grown in bicarbonate-buffered complex media under a N2/CO2/CO atmosphere (72:18:10, v/v/v) with 50 mM glycine betaine and harvested in the early stationary growth phase. The cell suspensions were prepared in 10 ml of cell suspension buffer (50 mM imidazole, 20 mM MgSO4, 20 mM KCl, 20 mM NaCl, 60 mM KHCO3, pH 7.0) in 120-ml serum flasks with 50 mM glycine betaine under 2 bar of a N2/CO2/CO (72:18:10, v/v/v) atmosphere at a final protein concentration of 1 mg/ml. Acetate (●) and lactate (▲) were determined. Each data point presents a mean ± SD; n = 2 independent experiments
Quantification of transcript levels of the lctB, lctC, and lctD genes in the ΔhydBA/hdcr mutant during growth on glycine betaine + CO. cDNA was synthesized from the ∆pyrE, Δhdcr and ΔhydBA/hdcr mutants grown on 50 mM glycine betaine in bicarbonate-buffered complex media under a N2/CO2/CO atmosphere (72:18:10, v/v/v). The transcript levels of the lctB, lctC, and lctD genes in the Δhdcr (grey bars) and ΔhydBA/hdcr mutants (black bars) were analyzed with quantitative real-time PCR and the relative expression was normalized to a house keeping gene gyrA. As control, cDNA of the ∆pyrE strain was used (white bars). Each data bar presents a mean ± SD; n = 3 independent experiments
Conversion of glycine betaine + CO in resting cells of the ΔhydBA/hdcr mutant under (a) CO2/HCO3−- or (b) Na+-depleted conditions. Cells of the ΔhydBA/hdcr mutants were grown in bicarbonate-buffered complex media under a N2/CO2/CO atmosphere (72:18:10, v/v/v) with 50 mM glycine betaine and harvested in the early stationary growth phase. The cell suspensions were prepared in 10 ml of (a) bicarbonate-depleted (50 mM imidazole, 20 mM MgSO4, 20 mM KCl, 20 mM NaCl, pH 7.0) or (b) Na+-depleted cell suspension buffer (50 mM imidazole, 20 mM MgSO4, 20 mM KCl, 60 mM KHCO3, pH 7.0) in 120-ml serum flasks with 50 mM glycine betaine under 2 bar of a (A) N2/CO (90:10, v/v) or (B) N2/CO2/CO (72:18:10, v/v/v) atmosphere at a final protein concentration of 1 mg/ml. The contaminating Na+ concentration was 0.1 mM. Acetate (●) and lactate (▲) were determined. Each data point presents a mean ± SD; n = 2 independent experiments
In the ∆hydBA/hdcr/lctBCD triple mutant, lactate formation was nearly completely abolished (Fig. 7), demonstrating that lactate is produced by the electron bifurcating LDH/ETF complex. Interestingly, the ΔhydBA/hdcr/lctBCD mutant produced double the amount of ethanol (6.0 ± 0.2 mM) compared to the ΔhydBA/hdcr mutant, indicating electrons are partially shifted towards ethanol production in the absence of the LDH/ETF complex.
Lactate formation from glycine betaine + CO was abolished in resting cells of the ΔhydBA/hdcr/lctBCD mutant. Cells of the ΔhydBA/hdcr/lctBCD mutant were grown in bicarbonate-buffered complex media under a N2/CO2/CO atmosphere (72:18:10, v/v/v) with 50 mM glycine betaine and harvested in the early stationary growth phase. The cell suspensions were prepared in 10 ml of cell suspension buffer (50 mM imidazole, 20 mM MgSO4, 20 mM KCl, 20 mM NaCl, 60 mM KHCO3, pH 7.0) in 120-ml serum flasks with 50 mM glycine betaine under 2 bar of a N2/CO2/CO (72:18:10, v/v/v) atmosphere at a final protein concentration of 1 mg/ml. Acetate (●) and lactate (▲) were determined. Each data point presents a mean ± SD; n = 2 independent experiments
Discussion
Acetogenic bacteria are prime candidates as biocatalysts required to transform our bioeconomy to a sustainable, sugar-free bioeconomy. This group of bacteria does not require oxygen, is easy to handle under strict anoxic conditions, grows robust even in industrial size fermenters, and can use carbon monoxide (Diekert and Thauer 1978; Diender et al. 2015; Genthner and Bryant 1982; Savage et al. 1987; Weghoff and Müller 2016), or more reduced C1 compounds such as formate (Moon et al. 2021) or methyl groups derived from various methyl group donors such as methanol or glycine betaine as building blocks for acetyl-CoA (Kremp and Müller 2021; Kremp et al. 2018; Lechtenfeld et al. 2018; Litty et al. 2022; van der Meijden et al. 1984). Electrons for the reduction can be derived from the oxidation of molecular hydrogen, carbon monoxide or organic substrates such as sugars. Moreover, many acetogens can grow mixotrophically on sugars and molecular hydrogen thus increasing the potential for a zero carbon-emission technology (Schuchmann and Müller 2016).
While acetate is the main product for all acetogens, some can naturally produce reduced end products such as ethanol from C1 compounds (Abrini et al. 1994; Köpke et al. 2010; Wilkins and Atiyeh 2011). Production of lactate has rarely been observed from C1 compounds. Lactate is a compound of significant industrial value due to its role as the precursor of PLA (Ahmad et al. 2022). A. woodii is one of the best studied acetogens and not only the biochemistry and bioenergetics of the WLP has been studied to a great detail, but also the metabolic pathways that feed C1 substrates into the WLP such as methanol, glycine betaine or CO (Kremp and Müller 2021; Schuchmann and Müller 2014). Recently, we have shown that a methylene-THF reductase deletion mutant performed mixed acid fermentation and produced lactate along with other fermentation products (Moon et al. 2023a). Here, we further investigated lactate production using genetically engineered strains, the ∆hdcr and ∆hydBA/hdcr mutants. When the electron bifurcating hydrogenase was deleted, lactate was the main product of fructose fermentation, implying that the electrons generated during glycolysis were used for lactate production. Unexpectedly, the ∆hydBA/hdcr/lctBCD mutant still produced lactate, although no other ldh genes could be identified in the genome. Interestingly, in some microbes NAD+-dependent LDH requires fructose-1,6-bisphosphate, an intermediate of the glycolysis, for catalytic activity (Arai et al. 2002; Brown and Wittenberger 1972; Freier and Gottschalk 1987; Machida et al. 1985a, b). In the triple mutant, fructose-1,6-biphosphate could have been accumulated due to slow fructose conversion and triggered the formation/activation of an alternative unknown LDH. But there is also an alternative way to produce lactate during fructose fermentation. An intermediate of glycolysis, dihydroxyacetone phosphate (DHAP) can be converted to methylglyoxal and further reduced to lactaldehyde. Then, lactaldehyde can be reoxidized to lactate (Bhowal et al. 2020; Stewart et al. 2013). The genome of A. woodii encodes enzymes that may catalyze these reactions (Poehlein et al. 2012). However, this way does not reoxidize reducing equivalents formed by glycolysis. How exactly lactate is produced from fructose by the double mutant must be investigated by further mutant analyses. Noteworthy, deletion of the LDH/ETF complex abolished lactate formation from C1 compounds (see below), indicating the need for (partial) glycolysis to trigger the alternative LDH way.
Production of lactate from C1 compounds is most attractive for biotechnological applications. Recently, a lctBCD deletion mutant of A. woodii harboring a lactate dehydrogenase gene from Leuconostoc mesenteroides fused to fluorescence-activating and absorption-shifting tag protein (FAST) was shown to produce lactate from H2 + CO2 (Mook et al. 2022). This strain produced 18.8 mM lactate from H2 + CO2 in batch experiments, but lactate was a side product with a lactate/acetate ratio of 0.33 (Mook et al. 2022). For exploring lactate production from more reduced C1 compounds, we chose glycine betaine as a methyl group donor plus CO as substrate. As described before for methanol plus CO (Litty et al. 2022), resting cells of ∆pyrE strain produced only acetate from glycine betaine + CO according to:
A likely scenario for lactate formation from glycine betaine + CO in the ∆hydBA/hdcr mutant is depicted in Fig. 8. The methyl group of glycine betaine is first transferred to THF by the methyltransferase system, yielding methyl-THF which is then condensed with CO and CoA on the CODH/ACS complex for acetyl-CoA production; 0.5 mol acetyl-CoA are then converted to acetate yielding 0.5 mol acetate. The other 0.5 mol of acetyl-CoA have to be reduced to 0.5 mol pyruvate via PFOR and the required reduced ferredoxin and CO2 were generated from oxidation of CO by the CODH. To produce 0.5 mol lactate, one mol NADH should be required which is produced by the Rnf complex. In sum, 0.5 mol acetate and 0.5 mol lactate are produced from one mol glycine betaine and 2 mol CO according to Eq. 2:
Biochemistry and bioenergetics of lactate production from glycine betaine + CO in the ΔhydBA/hdcr mutant of A. woodii. Fd, ferredoxin; PFOR, pyruvate:ferredoxin oxidoreductase; LDH/ETF, electron-bifurcating lactate dehydrogenase, GB, glycine betaine; DMG, dimethylglycine; THF, tetrahydrofolate; CODH/ACS, CO dehydrogenase/acetyl-coenzyme A synthase; CoFeSP, corrinoid iron-sulfur protein; MTI, methyltransferase I; MTII, methyltransferase II; CoP, corrinoid protein. The stoichiometry of the ATP synthase is 3.3 Na+/ATP (Matthies et al. 2014) and for the Rnf complex a stoichiometry of 2 Na+/2 e− is assumed
During growth on glycine betaine + CO, the ∆hydBA/hdcr mutant produced only acetate, similar to the ∆pyrE and ∆hdcr mutants; the ATP gain of this fermentation is 0.5 mol per mol of carbon of products or educts. On the other hand, during heterolactate fermentation, the ATP gain decreased to 0.37 mol per carbon of products or educts. Therefore, acetogenesis appears to be more favorable over heterolactate fermentation during growth but in resting cells, where a maximum ATP gain is not required, lactate fermentation is obviously preferred for unknown reasons. Moreover, pyruvate produced during growth is probably not accumulated, instead, utilized to build up biomass.
In conclusion, this study shows that a directed genetic engineering of a homoacetogen leads to lactate formation not only from sugar fermentation but also from C1 compounds, which gives a new perspective for industrial applications.
Data availability
All datasets and material generated or analyzed in this study are available from the corresponding author upon reasonable request.
References
Abrini J, Naveau H, Nyns EJ (1994) Clostridium autoethanogenum, sp. nov., an anaerobic bacterium that produces ethanol from carbon monoxide. Arch Microbiol 161:345–351. https://doi.org/10.1007/BF00303591
Ahmad A, Banat F, Alsafar H, Hasan SW (2022) An overview of biodegradable poly (lactic acid) production from fermentative lactic acid for biomedical and bioplastic applications. Biomass Conv Bioref. https://doi.org/10.1007/s13399-022-02581-3
Arai K, Hishida A, Ishiyama M, Kamata T, Uchikoba H, Fushinobu S, Matsuzawa H, Taguchi H (2002) An absolute requirement of fructose 1,6-bisphosphate for the Lactobacillus casei L-lactate dehydrogenase activity induced by a single amino acid substitution. Protein Eng 15:35–41. https://doi.org/10.1093/protein/15.1.35
Balk M, Weijma J, Friedrich MW, Stams AJ (2003) Methanol utilization by a novel thermophilic homoacetogenic bacterium, Moorella mulderi sp. nov., isolated from a bioreactor. Arch Microbiol 179:315–320. https://doi.org/10.1007/s00203-003-0523-x
Bertsch J, Öppinger C, Hess V, Langer JD, Müller V (2015) Heterotrimeric NADH-oxidizing methylenetetrahydrofolate reductase from the acetogenic bacterium Acetobacterium woodii. J Bacteriol 197:1681–1689. https://doi.org/10.1128/JB.00048-15
Bhowal B, Singla-Pareek SL, Sopory SK, Kaur C (2020) From methylglyoxal to pyruvate: a genome-wide study for the identification of glyoxalases and D-lactate dehydrogenases in Sorghum bicolor. BMC Genomics 21:145. https://doi.org/10.1186/s12864-020-6547-7
Brown AT, Wittenberger CL (1972) Fructose-1,6-diphosphate-dependent lactate dehydrogenase from a cariogenic Streptococcus: purification and regulatory properties. J Bacteriol 110:604–615. https://doi.org/10.1128/jb.110.2.604-615.1972
Diekert GB, Thauer RK (1978) Carbon monoxide oxidation by Clostridium thermoaceticum and Clostridium formicoaceticum. J Bacteriol 136:597–606. https://doi.org/10.1128/jb.136.2.597-606.1978
Diender M, Stams AJ, Sousa DZ (2015) Pathways and bioenergetics of anaerobic carbon monoxide fermentation. Front Microbiol 6:1275. https://doi.org/10.3389/fmicb.2015.01275
Dönig J, Müller V (2018) Alanine, a novel growth substrate for the acetogenic bacterium Acetobacterium woodii. Appl Environ Microbiol 84:e02023-e2118. https://doi.org/10.1128/AEM.02023-18
Fontaine FE, Peterson WH, McCoy E, Johnson MJ, Ritter GJ (1942) A new type of glucose fermentation by Clostridium thermoaceticum. J Bacteriol 43:701–715. https://doi.org/10.1128/jb.43.6.701-715.1942
Förster AH, Gescher J (2014) Metabolic engineering of Escherichia coli for production of mixed-acid fermentation end products. Front Bioeng Biotechnol 2:16. https://doi.org/10.3389/fbioe.2014.00016
Freier D, Gottschalk G (1987) L(+)-lactate dehydrogenase of Clostridium acetobutylicum is activated by fructose-1,6-bisphosphate. FEMS Microbiol Lett 43:229–233. https://doi.org/10.1111/j.1574-6968.1987.tb02128.x
Genthner BR, Bryant MP (1982) Growth of Eubacterium limosum with carbon monoxide as the energy source. Appl Environ Microbiol 43:70–74. https://doi.org/10.1128/aem.43.1.70-74.1982
Heap JT, Pennington OJ, Cartman ST, Minton NP (2009) A modular system for Clostridium shuttle plasmids. J Microbiol Methods 78:79–85. https://doi.org/10.1016/j.mimet.2009.05.004
Heise R, Müller V, Gottschalk G (1989) Sodium dependence of acetate formation by the acetogenic bacterium Acetobacterium woodii. J Bacteriol 171:5473–5478. https://doi.org/10.1128/jb.171.10.5473-5478.1989
Hess V, Schuchmann K, Müller V (2013) The ferredoxin:NAD+ oxidoreductase (Rnf) from the acetogen Acetobacterium woodii requires Na+ and is reversibly coupled to the membrane potential. J Biol Chem 288:31496–31502. https://doi.org/10.1074/jbc.M113.510255
Himes RH, Harmony JA (1973) Formyltetrahydrofolate synthetase. CRC Crit Rev Biochem 1:501–535. https://doi.org/10.3109/10409237309105441
Ingram LO, Conway T, Clark DP, Sewell GW, Preston JF (1987) Genetic engineering of ethanol production in Escherichia coli. Appl Environ Microbiol 53:2420–2425. https://doi.org/10.1128/aem.53.10.2420-2425.1987
Inui M, Kawaguchi H, Murakami S, Vertes AA, Yukawa H (2004a) Metabolic engineering of Corynebacterium glutamicum for fuel ethanol production under oxygen-deprivation conditions. J Mol Microbiol Biotechnol 8:243–254. https://doi.org/10.1159/000086705
Inui M, Murakami S, Okino S, Kawaguchi H, Vertes AA, Yukawa H (2004b) Metabolic analysis of Corynebacterium glutamicum during lactate and succinate productions under oxygen deprivation conditions. J Mol Microbiol Biotechnol 7:182–196. https://doi.org/10.1159/000079827
Jojima T, Igari T, Moteki Y, Suda M, Yukawa H, Inui M (2015a) Promiscuous activity of (S, S)-butanediol dehydrogenase is responsible for glycerol production from 1,3-dihydroxyacetone in Corynebacterium glutamicum under oxygen-deprived conditions. Appl Microbiol Biotechnol 99:1427–1433. https://doi.org/10.1007/s00253-014-6170-0
Jojima T, Noburyu R, Sasaki M, Tajima T, Suda M, Yukawa H, Inui M (2015b) Metabolic engineering for improved production of ethanol by Corynebacterium glutamicum. Appl Microbiol Biotechnol 99:1165–1172. https://doi.org/10.1007/s00253-014-6223-4
Köpke M, Held C, Hujer S, Liesegang H, Wiezer A, Wollherr A, Ehrenreich A, Liebl W, Gottschalk G, Dürre P (2010) Clostridium ljungdahlii represents a microbial production platform based on syngas. Proc Natl Acad Sci USA 107:13087–13092. https://doi.org/10.1073/pnas.1004716107
Kremp F, Müller V (2021) Methanol and methyl group conversion in acetogenic bacteria: biochemistry, physiology and application. FEMS Microbiol Rev 45:fuaa040. https://doi.org/10.1093/femsre/fuaa040
Kremp F, Poehlein A, Daniel R, Müller V (2018) Methanol metabolism in the acetogenic bacterium Acetobacterium woodii. Environ Microbiol 20:4369–4384. https://doi.org/10.1111/1462-2920.14356
Lechtenfeld M, Heine J, Sameith J, Kremp F, Müller V (2018) Glycine betaine metabolism in the acetogenic bacterium Acetobacterium woodii. Environ Microbiol 20:4512–4525. https://doi.org/10.1111/1462-2920.14389
Liew F, Henstra AM, Köpke M, Winzer K, Simpson SD, Minton NP (2017) Metabolic engineering of Clostridium autoethanogenum for selective alcohol production. Metab Eng 40:104–114. https://doi.org/10.1016/j.ymben.2017.01.007
Liew FE, Nogle R, Abdalla T, Rasor BJ, Canter C, Jensen RO, Wang L, Strutz J, Chirania P, De Tissera S, Mueller AP, Ruan Z, Gao A, Tran L, Engle NL, Bromley JC, Daniell J, Conrado R, Tschaplinski TJ, Giannone RJ, Hettich RL, Karim AS, Simpson SD, Brown SD, Leang C, Jewett MC, Kopke M (2022) Carbon-negative production of acetone and isopropanol by gas fermentation at industrial pilot scale. Nat Biotechnol 40:335–344. https://doi.org/10.1038/s41587-021-01195-w
Lim HG, Lee JH, Noh MH, Jung GY (2018) Rediscovering acetate metabolism: Its potential sources and utilization for biobased transformation into value-added chemicals. J Agric Food Chem 66:3998–4006. https://doi.org/10.1021/acs.jafc.8b00458
Litty D, Kremp F, Muller V (2022) One substrate, many fates: different ways of methanol utilization in the acetogen Acetobacterium woodii. Environ Microbiol 24:3124–3133. https://doi.org/10.1111/1462-2920.16011
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆Ct method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lovell CR, Przybyla A, Ljungdahl LG (1988) Cloning and expression in Escherichia coli of the Clostridium thermoaceticum gene encoding thermostable formyltetrahydrofolate synthetase. Arch Microbiol 149:280–285. https://doi.org/10.1007/BF00411642
Machida M, Matsuzawa H, Ohta T (1985a) Fructose 1,6-bisphosphate-dependent L-lactate dehydrogenase from Thermus aquaticus YT-1, an extreme thermophile: activation by citrate and modification reagents and comparison with Thermus caldophilus GK24 L-lactate dehydrogenase. J Biochem 97:899–909. https://doi.org/10.1093/oxfordjournals.jbchem.a135132
Machida M, Yokoyama S, Matsuzawa H, Miyazawa T, Ohta T (1985b) Allosteric effect of fructose 1,6-bisphosphate on the conformation of NAD+ as bound to L-lactate dehydrogenase from Thermus caldophilus GK24. J Biol Chem 260:16143–16147. https://doi.org/10.1016/S0021-9258(17)36212-9
Matthies D, Zhou W, Klyszejko AL, Anselmi C, Yildiz O, Brandt K, Müller V, Faraldo-Gomez JD, Meier T (2014) High-resolution structure and mechanism of an F/V-hybrid rotor ring in a Na+-coupled ATP synthase. Nat Commun 5:5286. https://doi.org/10.1038/ncomms6286
Mock J, Zheng Y, Müller AP, Ly S, Tran L, Segovia S, Nagaraju S, Köpke M, Dürre P, Thauer RK (2015) Energy conservation associated with ethanol formation from H2 and CO2 in Clostridium autoethanogenum involving electron bifurcation. J Bacteriol 197:2965–2980. https://doi.org/10.1128/JB.00399-15
Mohd Azhar SH, Abdulla R, Jambo SA, Marbawi H, Gansau JA, Mohd Faik AA, Rodrigues KF (2017) Yeasts in sustainable bioethanol production: A review. Biochem Biophys Rep 10:52–61. https://doi.org/10.1016/j.bbrep.2017.03.003
Mook A, Beck MH, Baker JP, Minton NP, Dürre P, Bengelsdorf FR (2022) Autotrophic lactate production from H2 + CO2 using recombinant and fluorescent FAST-tagged Acetobacterium woodii strains. Appl Microbiol Biotechnol 106:1447–1458. https://doi.org/10.1007/s00253-022-11770-z
Moon J, Müller V (2021) Physiology and genetics of ethanologenesis in the acetogenic bacterium Acetobacterium woodii. Environ Microbiol 23:6953–6964. https://doi.org/10.1111/1462-2920.15739
Moon J, Henke L, Merz N, Basen M (2019) A thermostable mannitol-1-phosphate dehydrogenase is required in mannitol metabolism of the thermophilic acetogenic bacterium Thermoanaerobacter kivui. Environ Microbiol 21:3728–3736. https://doi.org/10.1111/1462-2920.14720
Moon J, Dönig J, Kramer S, Poehlein A, Daniel R, Müller V (2021) Formate metabolism in the acetogenic bacterium Acetobacterium woodii. Environ Microbiol 23:4214–4227. https://doi.org/10.1111/1462-2920.15598
Moon J, Schubert A, Poehlein A, Daniel R, Müller V (2023a) A new metabolic trait in an acetogen: Mixed acid fermentation of fructose in a methylene-tetrahydrofolate reductase mutant of Acetobacterium woodii. Environ Microbiol Rep in Press. https://doi.org/10.1111/1758-2229.13160
Moon J, Schubert A, Waschinger LM, Müller V (2023b) Reprogramming the metabolism of an acetogenic bacterium to homoformatogenesis. ISME J. https://doi.org/10.1038/s41396-023-01411-2. (in Press)
Müller V (2003) Energy conservation in acetogenic bacteria. Appl Environ Microbiol 69:6345–6353. https://doi.org/10.1128/aem.69.11.6345-6353.2003
Müller V, Frerichs J (2013) Acetogenic bacteria. In: Encyclopedia of life sciences. John Wiley & Sons Ltd (ed), Chichester. https://doi.org/10.1002/9780470015902.a0020086.pub2
Müller V (2008) Bacterial fermentation. In: Encyclopedia of life sciences. John Wiley & Sons Ltd (ed), Chichester. https://doi.org/10.1002/9780470015902.a0001415.pub2
Poehlein A, Schmidt S, Kaster A-K, Goenrich M, Vollmers J, Thürmer A, Bertsch J, Schuchmann K, Voigt B, Hecker M, Daniel R, Thauer RK, Gottschalk G, Müller V (2012) An ancient pathway combining carbon dioxide fixation with the generation and utilization of a sodium ion gradient for ATP synthesis. PLoS One 7:e33439. https://doi.org/10.1371/journal.pone.0033439
Ragsdale SW (2003) Pyruvate ferredoxin oxidoreductase and its radical intermediate. Chem Rev 103:2333–2346. https://doi.org/10.1021/cr020423e
Ragsdale SW (2008) Enzymology of the Wood-Ljungdahl pathway of acetogenesis. Ann N Y Acad Sci 1125:129–136. https://doi.org/10.1196/annals.1419.015
Ragsdale SW, Ljungdahl LG (1984) Purification and properties of NAD-dependent 5,10-methylenetetrahydrofolate dehydrogenase from Acetobacterium woodii. J Biol Chem 259:3499–3503. https://doi.org/10.1007/BF00411642
Sanger FS, Nickelen F, Coulson AR (1977) DNA-sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467. https://doi.org/10.1073/pnas.74.12.5463
Savage MD, Wu ZG, Daniel SL, Lundie LL Jr, Drake HL (1987) Carbon monoxide-dependent chemolithotrophic growth of Clostridium thermoautotrophicum. Appl Environ Microbiol 53:1902–1906. https://doi.org/10.1128/aem.53.8.1902-1906.1987
Schmidt K, Liaaen-Jensen S, Schlegel HG (1963) Die Carotinoide der Thiorhodaceae. Arch Mikrobiol 46:117–126. https://doi.org/10.1007/BF00408204
Schuchmann K, Müller V (2012) A bacterial electron bifurcating hydrogenase. J Biol Chem 287:31165–31171. https://doi.org/10.1074/jbc.M112.395038
Schuchmann K, Müller V (2013) Direct and reversible hydrogenation of CO2 to formate by a bacterial carbon dioxide reductase. Science 342:1382–1385. https://doi.org/10.1126/science.1244758
Schuchmann K, Müller V (2014) Autotrophy at the thermodynamic limit of life: a model for energy conservation in acetogenic bacteria. Nat Rev Microbiol 12:809–821. https://doi.org/10.1038/nrmicro3365
Schuchmann K, Müller V (2016) Energetics and application of heterotrophy in acetogenic bacteria. Appl Environ Microbiol 82:4056–4069. https://doi.org/10.1128/AEM.00882-16
Stewart BJ, Navid A, Kulp KS, Knaack JL, Bench G (2013) D-Lactate production as a function of glucose metabolism in Saccharomyces cerevisiae. Yeast 30:81–91. https://doi.org/10.1002/yea.2942
Trifunović D, Schuchmann K, Müller V (2016) Ethylene glycol metabolism in the acetogen Acetobacterium woodii. J Bacteriol 198:1058–1065. https://doi.org/10.1128/JB.00942-15
van der Meijden P, van der Drift C, Vogels GD (1984) Methanol conversion in Eubacterium limosum. Arch Microbiol 138:360–364. https://doi.org/10.1007/bf00410904
Weghoff MC, Müller V (2016) CO metabolism in the thermophilic acetogen Thermoanaerobacter kivui. Appl Environ Microbiol 82:2312–2319. https://doi.org/10.1128/AEM.00122-16
Weghoff MC, Bertsch J, Müller V (2015) A novel mode of lactate metabolism in strictly anaerobic bacteria. Environ Microbiol 17:670–677. https://doi.org/10.1111/1462-2920.12493
Werner H, Krasemann C, Gorniak W, Hermann A, Ungerechts J (1977) Die Thiamphenicol- und Chloramphenicol-Empfindlichkeit von Anaerobiern. Zentralbl Bakteriol Orig A 237:358–371
Westphal L, Wiechmann A, Baker J, Minton NP, Müller V (2018) The Rnf complex is an energy coupled transhydrogenase essential to reversibly link cellular NADH and ferredoxin pools in the acetogen Acetobacterium woodii. J Bacteriol 200:e00357-e418. https://doi.org/10.1128/JB.00357-18
Wiechmann A, Ciurus S, Oswald F, Seiler VN, Müller V (2020) It does not always take two to tango: “Syntrophy” via hydrogen cycling in one bacterial cell. ISME J 14:1561–1570. https://doi.org/10.1038/s41396-020-0627-1
Wilkins MR, Atiyeh HK (2011) Microbial production of ethanol from carbon monoxide. Curr Opin Biotechnol 22:326–330. https://doi.org/10.1016/j.copbio.2011.03.005
Wood HG, Ljungdahl LG (1991) Autotrophic character of the acetogenic bacteria. In: Shively JM, Barton LL (eds) Variations in autotrophic life. Academic press, San Diego, pp 201–250
Wood HG, Ragsdale SW, Pezacka E (1986) The acetyl-CoA pathway of autotrophic growth. FEMS Microbiol Rev 39:345–362
Funding
Open Access funding enabled and organized by Projekt DEAL. This project has received funding from the European Research Council (ERC) under the European Union's Horizon 2020 research and innovation program (grant agreement number 741791).
Author information
Authors and Affiliations
Contributions
V.M designed and supervised the research, analyzed the data, and wrote the manuscript. J.M designed the research, conducted the experiments, analyzed the data, and wrote the manuscript. L.M.W generated the deletion mutant, performed the experiments, and analyzed the data. The manuscript was approved by all authors.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Moon, J., Waschinger, L.M. & Müller, V. Lactate formation from fructose or C1 compounds in the acetogen Acetobacterium woodii by metabolic engineering. Appl Microbiol Biotechnol 107, 5491–5502 (2023). https://doi.org/10.1007/s00253-023-12637-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12637-7