Abstract
Fatty acid desaturase catalyzes the desaturation reactions by inserting double bonds into the fatty acyl chain, producing unsaturated fatty acids, which play a vital part in the synthesis of polyunsaturated fatty acids. Though soluble fatty acid desaturases have been described extensively in advanced organisms, there are very limited studies of membrane fatty acid desaturases due to their difficulties in producing a sufficient amount of recombinant desaturases. However, the advancement of technology has shown substantial progress towards the development of elucidating crystal structures of membrane fatty acid desaturase, thus, allowing modification of structure to be manipulated. Understanding the structure, mechanism, and biosynthesis of fatty acid desaturase lay a foundation for the potential production of various strategies associated with alteration and modifications of polyunsaturated fatty acids. This manuscript presents the current state of knowledge and understanding about the structure, mechanisms, and biosynthesis of fatty acid desaturase. In addition, the role of unsaturated fatty acid desaturases in health and diseases is also encompassed. This will be useful in understanding the molecular basis and structural protein of fatty acid desaturase that are significant for the advancement of therapeutic strategies associated with the improvement of health status.
Key points
• Current state of knowledge and understanding about the biosynthesis, mechanisms, and structure of fatty acid desaturase.
• The role of unsaturated fatty acid desaturase.
• The molecular basis and structural protein elucidated the crystal structure of fatty acid desaturase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fatty acid desaturases (FADs) are a group of enzymes that catalyze desaturation reaction by introducing double bonds in a wide range of different positions and desaturating saturated fatty acids into unsaturated fatty acids. This reaction takes place in the presence of molecular oxygen and reducing equivalents (which are delivered by an electron transport system) (Los and Murata 1998). The FADs have an Enzyme Commission (EC) number of 1.14.19.X and have been classified as oxidoreductases that act on single donors by incorporating with molecular oxygen (oxygenase), usually by utilizing NADP or NAD + as co-factors. The distribution of FADs is nearly universal with a few exceptions, such as Escherichia coli (Los and Murata 1998). The FADs can aid in the desaturation of fatty acids, which results in monounsaturated fatty acids (MUFAs) and polyunsaturated fatty acids (PUFAs). In the production of PUFAs, FADs are essential; PUFAs serve as precursors of eicosanoids, pheromones, growth regulators, and hormones, among other biologically active compounds (Los and Murata 1998). The biosynthesis of PUFAs can occur via different pathways, and they benefit in the development of medicines and nutritional supplements (Lindqvist et al. 1996). Fatty acids, such as eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA) (found primarily in seafood sources, fish-oil supplements, seeds, and nuts), are the end products of desaturation and elongation processes, and are essential for human health and the physiology of the human body as they are a nutritional necessity. PUFAs are produced in large quantities by fungi, algae, and protozoa, and other microbes. These microorganisms are expected to have all the enzymes required to produce EPA and DHA (Pereira et al. 2003).
Recent findings on the challenges and strategies for protein production enabled the determination of membrane protein structural biology, making it possible for the membrane protein to be solved. The advancement of technology has made the crystallization and structural elucidation of membrane-bound desaturase as one of the most significant achievements of the recent years. The number of solved crystal structures of FADs has increased tremendously. X-ray structure of a Δ9 mouse stearoyl-CoA desaturase has been crystallized and solved at 2.6 Å resolution by Bai et al. (2015). Another crystal structure of Δ9 in human stearoyl-coenzyme A desaturase in complex with substrate has been elucidated (Wang et al. 2015a, b). Both structures exhibit a tertiary fold that resembles a mushroom, with the catalytic cap domain and terminal ends on the cytosolic side of the membrane (Nachtschatt et al. 2020). Mouse and human Δ9 FAD crystal structure with the di-iron center were studied; however, in the catalytic site of both proteins, zinc ions were found instead of iron ions, which thereby triggered the enzymes to become inactive (Bai et al. 2015; Wang et al. 2015a, b). In this manuscript, the biosynthesis pathway, catalytic mechanism, architecture, and roles of unsaturated FADS are discussed.
Classification of FADs
Two groups of FADs have been found in nature: soluble and membrane-bound desaturases. These groups have been broadly classified into two evolutionary unrelated groups of soluble acyl-acyl carrier protein (ACP) and membrane-bound desaturases (Lou et al. 2014). Soluble desaturase is present in the plastids of higher plants that require NADPH and oxygen. It is part of an electron transport chain that includes ferredoxin-NADPH reductase. Each acyl-ACP desaturase forms a reactive complex with oxygen by binding two iron atoms (Shanklin and Cahoon 1998). The conserved motif for soluble FADs is 2xD/EX2H (Shanklin and Cahoon 1998). On the other hand, membrane FADs is found in most prokaryotes and eukaryotic organisms. The membrane-bound group is further subdivided into acyl-lipid FADs e and acyl-coenzyme A (CoA) FADs. Membrane FADs use ferredoxin or cytochrome b5. In monogalactosyl diacylglycerol, cyanobacterial membrane FADs could desaturate stearic (18:0) and oleic (18:1 n-9) fatty acyl chain (Tocher et al. 1998). Most of this type of FADs has 300–350 amino acid residues, and it is a hydrophobic protein that spans membranes four times (Murata and Wada 1995). The endoplasmic reticulum membranes of animals, yeasts, and fungi contain acyl-CoA FADs, and they obtain electrons from cytochrome b5 and NADH-dependent cytochrome b5 reductase. All mammalian FADs that have been identified are grouped under acyl-CoA desaturases (Tocher et al. 1998).
The amino acid sequences of membrane-bound FADs suggest that these enzymes contain two long hydrophobic domains that can traverse the membrane bi-layer twice (Alberts 2002). Three conserved His-box motifs were discovered through sequence comparison: HX3-4H, HX2-3HH, and H/QX2-3HH (H-Histidine; X-variable amino acid; Q-glutamine), all of which include eight histidine residues. These histidine residues operate in the catalytic center of desaturases as possible ligands for iron atoms (Nakamura and Nara 2004). These iron atoms are defined as nonheme and di-iron containing; thus, they can bind the iron ions with conserved histidine residues. All known desaturases have been characterized by the presence of three histidine clusters, which are strongly conserved in the amino acid sequence (Wang et al. 2013). Table 1 shows the comparison of two types of fatty acid desaturases. According to Nachtschatt et al. (2020), linear saturated fatty acids have been created by the fatty acid synthase enzyme system known as FAS I or FAS II, which has the potential to elongate acetyl CoA by condensing it with malonyl-CoA molecules under the presence of carbon dioxide and water in cyclical manner. Even-numbered carbons of fatty acids are the most prevalent, and palmitic acids along with stearic acid are the most prevalent products of fatty acid synthesis (Li et al. 2016a, b, c). Membrane-bound desaturases can be later further distinguished by the position of fatty acid where the double bond has been introduced (Nachtschattt et al. 2020).
Desaturases can be further classified as first desaturases, methyl-end desaturases, and front-end desaturases, based on the position of the introduced double bond (Li et al. 2016a, b, c). First desaturase introduces first double bond into the saturated acyl chain, and the predominant member is stearoyl-coenzyme A desaturase which generally introduces a double bond at the 9th position of palmitic acid (C16:0) or stearic acid (C18:0). Front-end desaturases such as Δ4, Δ5, and Δ6 can introduce double bonds between existing double bonds and the carboxyl group (-COOH) to generate PUFAs (Lee et al. 2016). According to International Union of Pure and Applied Chemistry (IUPAC) in the Compendium of Chemical Terminology (Castro et al. 2016), an enzyme from this group is referred to as Δx (delta X desaturase), where X is the number corresponding to the position from the carboxyl end at which the double bond is inserted. Desaturases with a new double bond between an existing unsaturation and the PUFA’s methyl terminus (–CH3) are known as methyl-end desaturases and referred to as ωx (omega X desaturases) (Monroig and Kabeya 2018). Therefore, a double bond at 12th position of fatty acyl chain counted from the carboxyl-end is known as Δ12 desaturase, and ω6 when counted from the methyl-end.
Few membrane-bound desaturases have been reported from organisms, such as Δ9, Δ12, Δ15, Δ6, Δ5, and Δ4. Δ9 FAD genes catalyze the insertion of double bond at the position of the 9th carbon in fatty acids, synthesizing long-chain monounsaturated fatty acids from long-chain saturated fatty acids, which convert stearic acid (C18:0) and palmitic acid (C16:0) to oleic acid (OA, C18:1Δ9) and palmitoleic acid (POA, C16:1Δ9) (Zhuang et al. 2022). It is commonly known as Stearoyl CoA, and has been isolated from numerous species, such as Saccharomyces cerevisiae, Mortierella alpina, Histoplasma capsulatum, Tetrahymena thermophile (Tocher et al. 1998; Sakuradani et al. 1999; Laoteng et al. 1999), and Pseudomonas oleovorans (Li et al. 2009). Two hydrophobic domains of this desaturase can penetrate the phospholipid bilayer (Zhang 2009). Formed from red algae (known as Cyanidioschyzon merolae) and yeast, Δ9 FADs are distinguished by the existence of a cytochrome b5 domain located at the C-terminus, which is obligatory for the protein’s activity (Mitchell and Martin 1995; Itoh et al. 1998). Δ9 FADs has been reported in humans (Wang et al. 2015a, b) and mice (Bai et al. 2015). Extensive studies of this protein have been conducted in these two organisms up until structure elucidation (Wang et al. 2015a, b and Bai et al. 2015).
Δ12 FADs catalyze the insertion of a double bond at the position of the 12th carbon in fatty acids, indicating the double bonds are placed at the position of the 6th carbon position counted from methyl end groups. It is a vital enzyme catalyzing oleic acid (OA, C18:1Δ9) to linoleic acid (LA, C18:2Δ9,12) (Lee et al., 2016) (Saini et al. 2021). Δ12 FADs have been isolated from Thraustochytrium aureum (yeasts) (Matsuda et al. 2012) and characterized from Mortierella alpina (Huang et al. 1999) and Mucor rouxii (Passorn et al. 1999). A study suggested that Δ12 FADs found in Mortierella alpina are extremely active and cause a conversion of 71.4% of substrate oleic acid to linoleic acid. In Lipomyces starkeyi yeast, the activity of Δ12 FADs has been measured (Lomascolo et al. 1996); in microorganisms, Δ12 FADs are quite similar to such genes in animals in terms of sequence identity. According to Zhuang et al. (2022), identification and characterization of Δ12 FADs genes from Isochrysis galbana (Han et al. 2019), Acanthamoeba castellani (Sayanova et al. 2016), Chlamydomonas sp. (Zhang et al. 2011), Calendula officinalis (Qiu et al. 2001), Helianthus annuus (Martinez-Rivas et al. 2001), Chlorella vulgaris (Lu et al. 2009), Phaedactylum tricornutum (Domergue et al. 2003), and Haematococcus pluvialis (Zhang et al. 2020) showed they shared three conserved histidine box motifs, which known as catalytic center and are critical for desaturase activity.
Δ15 FADs, also known as ω3 (counted from methyl end), add a double bond at the 15th position in fatty acids, indicating the double bonds at the 3rd position of fatty acids from methyl end groups. Δ15 FADs catalyze linoleic acid (LA, C18: 2Δ9,12) and γ-linolenic acid (GLA, C18: 3Δ6,9,12) to produce α-linolenic acid (ALA, C18: 3Δ9,12,15) and stearidonic acid (SDA, C18: 4Δ6,9,12,15), respectively (Saini et al. 2021). Δ15 FADs have been discovered in various species of molds, such as Saprolegnia diclina (Pereira et al. 2004), Phytophthora infestans (Fu et al. 2013), Pythium aphanidermatum (Xue et al. 2013), and blue-green algae, Synechocystis sp. PCC6803 (Drew et al. 2001). A previous study showed that Δ15 desaturase of Phytophthora infestans converted 30.94% of arachidonic acid (AA) into eicosapentaenoic acid (EPA) (Fu et al. 2013). Δ12 and Δ15 FADs are important enzymes in the synthesis of omega-3 and omega-6 in mammals, as mammals cannot produce these fatty acids. According to phylogenetic studie, Δ12 FADs is an ancestor gene of Δ15 FADs (Wang et al. 2013).
Δ6 FADs add a double bond at the 6th carbon–carbon bond position from the carboxylic acid end in fatty acids. The desaturases are a rate-limiting enzyme in generating γ-linolenic acid (GLA, C18:3Δ6,9,12) and producing stearidonic acid (SDA, C18:4Δ6,9,12,15) from linoleic acid (LA, C18:2Δ9,12) and α-linolenic acid (ALA, C18:3Δ9,12,15) in the synthesis of PUFAs in mammals and humans (Zhuang et al. 2022). These membrane-bound FADs have been identified in Synechocystis sp. PCC6803, Mortierella alpina (Chen et al. 2014), and Mucor rouxii (Laoten et al. 2000). However, Δ6 FADs of Mucor rouxii differ from those of Mortierella alpina as it has around 500 amino acid sequences and the identity of the sequence is closely related to plant Δ6 FADs compared to animal or fungal Δ6 FADs. It also has unique histidine-conserved motifs of “HKHHSH,” which cannot be found in any other desaturase. An early study stated that the enzymatic activity is thought to be dependent on the histidine-conserved region (Laoten et al. 2000).
Δ5 FADs catalyze the addition of a double bond at the 5th carbon–carbon bond from the carboxylic acid end in fatty acids. They catalyze the conversion of dihomo-γ-linolenic acid (DGLA, C20:3Δ8,11,14) and eicosatetraenoic acid (ETA, C20:4Δ8,11,14,17) to arachidonic acid (AA, C20:4Δ5,8,11,14) and eicosapentaenoic acid (EPA, C20:4Δ5,8,11,14,17) (Lee et al. 2016). These genes have been determined first in a fungus, Mortierella alpina, and the heterokont algae, Thraustochytrium sp. (Michaelson et al. 1998). According to Hastings et al. (2001) and Wang et al. (2014), Δ5 FADs in marine invertebrates showed bifunctional activity with the desaturation reaction of Δ5 and Δ6 FADs. Δ5 FADs have also been functionally expressed in yeast. A study showed that two types of Δ5 FADs have been identified in slime mold, Dictyostelium discoideum, and they shared 66% similarities, and their protein predictions had a sequence similarity of 38.6–42% with Mortierella alpina’s Δ5 FADs (Saito et al. 2000). The amino acid sequence from these FADs contained three conserved histidine box motifs and was significantly homologous (Zhuang et al. 2022).
Biosynthesis of FADs
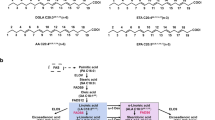
The biosynthesis of unsaturated fatty acids is synthesized de novo from acetyl-coenzyme (CoA) as a source of carbon, especially the C16:0 and C18:0 in animals (Garba et al. 2017). The pathway is catalyzed by the multienzyme fatty acid synthase (FAS) complex and acetyl-CoA carboxylase. The FAS complex is a key enzymatic complex in fatty acid synthesis from acetyl-CoA and malonyl-CoA in the presence of NADPH (Beld et al. 2015). The complex can be broadly divided into two types, Type 1 system (mainly found in animals and yeast, uses a single large multifunctional polypeptide) and the Type II system (mostly found in prokaryotes and plants, uses a series of monofunctional enzymes) (Garba et al. 2017). A protein called the acyl carrier protein (ACP) is required by both type I and type II FAS to transport the fatty acid from enzyme to enzyme (Beld et al. 2015). ACPs move and present the expanding acyl chain to the proper reaction partners for elongation and the synthesis of fatty acids.
The pathway was first started with two acetyl-CoA, one converted to malonyl-CoA by the enzyme acetyl-CoA carboxylase (Ahern and Rajagopal 2021). This enzyme is known to be phosphorylated by both AMP Kinase and Protein Kinase A. Both molecules have their CoA portions replaced by acyl-carrier protein (ACP) to form acetyl-ACP and malonyl-ACP (Ahern and Rajagopal 2021). Then, the chemical reactions resemble those of beta oxidation reversed. The ketone is reduced to a hydroxyl using NADPH, and water is removed from carbons 2 and 3 of the hydroxyl intermediate to produce a trans-doubled bonded molecule after every round of elongation. Last, the double bond is hydrogenated to yield a saturated intermediate via a chronological action by ketoactyl reductase, dehydrase, and enoyl reductase activities (Garba et al. 2017). Elongation of fatty acids to make fatty acids longer than 16 carbons is catalyzed by elongase enzymes and occurs in the endoplasmic reticulum. Fatty acids can also be extended by mitochondria, but their starting materials are typically less than 16 carbons long. Both environments use similar methods to the cytoplasm. Desaturation happens after fatty acids are produced in their saturated state (Ahern and Rajagopal 2021). Desaturases are enzymes that facilitate the cis-double bond synthesis in mature fatty acids.
Mechanism of FADS
The desaturation reaction catalyzed by fatty acid desaturase is an aerobic process (Czumaj and Sledzinski 2020). PUFAs are synthesized through chloroplasts (prokaryotic) and endoplasmic reticulum through ER (eukaryotic) pathways (He et al. 2020). The reaction requires oxygen, cytochrome b5, and NADH-dependent cytochrome b5 reductase, whereby oxygen acts as a hydrogen acceptor, and cytochrome b5 as an electron carrier (Shanklin et al. 2009). NADPH is considered essential for hydrogen donors for the synthesis of < 18C fatty acids (Chen et al. 2013). As in eukaryotic cells, acyl chains require to be the first ones to be exported to the ER (LaBrant et al. 2018; Li et al. 2016a, b, c). At the initial stage of desaturation reaction, reduced cytochrome b5 interacts with the non-heme iron active site of the desaturase, which enables the enzyme to react with oxygen (O2) and substrate, and results in the activation of hydrogen at the carbon of incipient double bond in substrate, producing a very short-lived carbon-centered radical as the product (Martin-Montalvo et al. 2016; Zhang et al. 2016). The next stage of this reaction shows that the molecule loses the second hydrogen via rapid disproportionation, which leads to the production of the final product with the formation of a new double bond, along with two molecules of water (Behrouzian and Buist 2003). Saturated fatty acids (SFAs), such as stearic acid (C18:0) and palmitic acid (C16:0), can be unsaturated by all eukaryotes and most aerobic bacteria, whereas the majority of prokaryotic organisms only contain of the so-called anaerobic fatty acid synthesis pathway that does not include any desaturase at all (Keweloh and Heipieper 1996).
Role of unsaturated FADs
Unsaturated FADs play a crucial role in the synthesis of PUFAs, inserting additional unsaturated bonds into the acyl chain producing products of these enzymes. The organism in which desaturases are expressed determines their exact function. FADs seem to exert a variable effect on microorganisms, plants, humans, and other mammals.
Role of unsaturated FADs in microorganisms
In bacteria, changes in the PUFAs composition in the plasma membrane could be vital in the adjustment towards harsh environmental conditions; thus, microorganisms can change the activity of desaturase to adapt PUFAs content towards temperature, pH, and atmospheric pressure to maintain appropriate fluidity of plasma membranes as the melting temperature of PUFAs is much lower than saturated and monounsaturated fatty acids (Yoshida et al. 2016). According to Hazel and Williams (1990), conditions of low temperature and high pressure reveal the integrity of membranes, which can be irreversible, from a fluid to a rigid state. PUFAs in microorganisms, usually with a long chain of fatty acids (around 16–18 carbons), have been found in great amounts in cold-adapted bacteria within the polar regions and deep-sea areas. Therefore, PUFAs especially EPA and DHA have been assumed as effective modulators for modifying membrane fluidity (Yoshida et al. 2016). The introduction of these bacteria into a warmer environment resulted in changes in expression of desaturases as well as changes in the PUFAs content in the plasma membrane (Diomandé et al. 2015; Yoshida et al. 2016; Siliakus et al. 2017; Bale et al. 2019).
Role of unsaturated FADs in plants
Plant composition of fatty acids is crucial for the growth and vegetation of plants as desaturation of fatty acids influences the tolerance of plants towards environmental stressors (Shanklin and Cahoon 1998; Iba 2002). According to Venegas-Caleron et al. (2009), PUFAs with more than 20 carbons cannot often be produced by higher plants, except for a few species that produce GLA and stearidonic acid. However, the production of transgenic plant oil has been developed and the first transgenic plant reported was tobacco, which used cyanobacterial delta-6 desaturase to accumulate GLA (Reddy and Thomas 1996). In plants, changes in fatty acid desaturase play roles in the facilitation of cold adaptation, maintenance of normal fluidity, and integrity of the plasma membrane (Iba 2002). A study revealed that the presence of fatty acid desaturases, FAD2 and FAD6 in seedlings, showed a significant impact on the activation by salt and osmotic stress (Zhang et al. 2009), while the deficiency of FAD2 resulted in the accumulation of sodium ions in the cytoplasm of root cells and a rise in sensitivity towards salt stress during germination and early seedling growth (Zhang et al. 2012).
Role of unsaturated FADs in human and mammals
FADs appear to have a varied effect on human health, which is the outcome of an association among polymorphisms of desaturase towards certain metabotypes (Wang et al. 2014). It is well-recognized that among these fatty acids, ω3 and ω6 are the most important, as the biosynthesis of these two fatty acids creates health benefits throughout life. r ω3 and ω6 fatty acids are crucial in humans, as they cannot be produced de novo. Research suggested that the body could convert ALA to DHA and EPA throughout the desaturation and elongation processes; however, the conversion process produces a small number of fatty acids (Chiu et al. 2008), around 0.3% for EPA and < 0.01% for DHA (Hussein et al. 2005). Low levels of EPA and DHA are correlated with various inflammatory processes, including cardiovascular diseases, poor fetal development (Swanson et al. 2012), Alzheimer’s diseases (Swanson et al. 2012), bipolar affective disorders, schizophrenia (Liu et al. 2011; Li et al. 2009; Yao et al. 2000), and cardiovascular disease (Swanson et al. 2012). In a study that used human blood samples conducted by Bouwens et al. (2009), they mentioned the intake of EPA and DHA changed the expression of 1040 genes and resulted in a decrease in the expression of genes associated with inflammatory and atherogenesis-related pathways, such as adipogenesis and hypoxia signaling. Schiano et al. (2008) discovered that EPA and DHA supplementation improved endothelial function in patients with atherosclerosis by lowering plasma levels of soluble thrombomodulin from a median value of 33.0 to 17.0 mg/L (P = 0.04) and increasing brachial artery flow–mediated dilation from 6.7 to 10.0% (P = 0.02). Therefore, it is important to take diets with a higher content of EPA and DHA, which include seafood sources, such as fish and dietary supplements derived from fish and fish oil (Leaf and Hatcher 2009; Mann et al. 2010; Saito et al. 2008).
The beneficial effect of DHA has also been recognized in various types of chronic inflammatory conditions, as DHA can be metabolized into anti-inflammatory bioactive lipid mediators (Nagy and Tiuca 2017). These characteristics have been conservatively defined as their ability to interact with primary inflammatory signaling pathways and their inhibition of inflammatory cytokine production (Lorente-Cebrian et al. 2015). A report described that higher concentration of DHA is associated with fewer morning stiffness, swelling joints, and pain. Collagen-induced arthritis is less common and severe when these fatty acids are consumed (Nagy and Tiuca 2017). DHA may also inhibit the production of reactive oxygen species and the stimulation of TNFRI (tumor necrosis factor receptor type I) by AA, both of which have immune-modulatory qualities and can affect T and B cell activity (Brouwers et al. 2015). Through the nuclear peroxisome proliferator–activated receptor (PPAR) system, DHA and EPA may suppress the expression of genes linked to inflammation (Endo and Arita 2016). This receptor can activate and stimulate β-oxidation, lowering the levels of triglycerides and fatty acids in the bloodstream, which are responsible for the prevention of hyperplasia and adipocyte hypertrophy (Echeverria et al. 2016). A study reported that patients with rheumatoid arthritis have a low level of DHA, EPA, and some fatty acids (such as oleic acid and palmitic acid) in comparison to normal healthy controls (Brouwers et al. 2015). In addition, some MUFAs produced by the desaturation reaction have a beneficial effect on anti-inflammatory properties, such as oleic acid (Zhou et al. 2016). According to a recent study, taking a higher amount of DHA lowers the amount of AA in inflammatory cell membranes, which lowers the amount of pro-inflammatory eicosanoids (Nagy and Tiuca 2017).
As DHA is abundant in brain cells and retina for normal cell membrane function, FADs play a crucial role in proper fetal development and healthy aging (Dunstan et al. 2007; Krauss-Etschmann et al. 2007). AA, EPA, and DHA play a significant role in the growth of infants, and the development of their neurons and immune systems (Mychaleckyi et al. 2018; Powell et al. 2016; Xie et al. 2008; Muc et al. 2015; Ding et al. 2016). The key ingredients of the neuronal membranes are derived from AA and DHA (Czumaj and Sledzinski 2020). The presence of EPA and DHA in membrane phospholipids may affect a variety of biological mechanisms and pathways in the brain, including the integrity and survival of neurons, glial cells, endothelial cells, neurotransmission (dopaminergic, serotonergic, glutamatergic, and cholinergic), and neuroinflammation (Nishida and Murata 1996; Healy-Stoffel and Levant 2018; Cao et al. 2018). A study suggested that EPA and DHA are the key components associated with multiple benefits in an early stage of life (Swanson et al. 2012). As the placenta provides nutrients to the fetus, a certain amount of DHA is also transferred (Helland et al. 2008). Supplementing with EPA and DHA during pregnancy has been linked to longer gestation and higher fatty acid concentrations in fetal tissues, which may reduce the risk of preterm delivery by lowering prostaglandin E2 and prostaglandin F2a synthesis (Olsen et al. 2008; Roman et al. 2006). There is also evidence that EPA and DHA supplementation during pregnancy and lactation may protect infants against allergies, as fish oil supplementation generates a lower level of body cells linked to inflammation and immunological response (Krauss-Etschmann et al. 2008). Fatty acids are delivered to newborns through nursing, which contributes to the development of cognitive function and intellectual development (Capsi et al. 2007; Gould and Smithers 2019).
Though both ω3 and ω6 fatty acids are considered to have beneficial health effects, overconsumption of ω6 fatty acids with low intake of ω3 in modern westernized diet styles has been shown to be linked to antagonistic metabolic functions and the emergence of many chronic diseases, including inflammatory diseases, cardiovascular diseases, and some cancers related to inflammation (Mariamenatu and Abdu 2021). Large amounts of LA and AA derivatives cause cell proliferation, the endocannabinoid system to be extremely hyperactive, thrombus and atheroma formation, allergy, and inflammatory problems, especially in vulnerable individuals (Mariamenatu and Abdu 2012), even though these two PUFAs are essential as the human body is genetically incapable of synthesising them (Ruiz-Lopez et al. 2012). It was once thought that our ancestors consumed ω6 and ω3 in a ratio of 1–2/1, which is believed to be a perfect and balanced ratio during the Paleolithic period (a time of evolution) (Simopoulos 2008). To date, the current estimate for the typical ratio in western diet types is 20/1 (Sokola-Wysoczanska et al. 2018; Simopoulos 2016), whereas in South Asia, the ratio can reach 50/1 (Chaves et al. 2019). Increase in the ω3 and ω6 fatty acids ratio increases the risk of overweight and obesity (Simopoulos 2016). According to Simopoulos (2010), keeping the ratio of ω3 and ω6 fatty acids in balance helps lower the chance of developing autoimmune and possibly neurodegenerative disorders, as well as coronary heart disease, high blood pressure, cancer, type 2 diabetes, and arthritis.
Architecture of FADs
To date, there are around 59 crystal structures of FADs, and most of them are soluble desaturase, including the X-ray structure of Mycobacterium tuberculosis H37Rv DesA2 (Dyer et al. 2005), a soluble acyl-ACP desaturase (Buist 2004) and crystal structure of multifunctional Δ4 acyl-ACP desaturase isolated from English ivy (Hedera helix) plant at 1.95 Å resolution (Guy et al. 2007). Few membrane-bound FAD structures have been solved, which include X-ray structure of mouse stearoyl-CoA desaturase (Δ9 mSCD1) at 2.6 Å resolution (Bai et al. 2015), crystal structure of human stearoyl–coenzyme A (Δ9 hSCD1) desaturase at resolution 3.25 Å (Wang et al. 2015a, b), and Δ9 acyl-CoA desaturase from Mus musculus at 3.51 Å resolution (Shen et al. 2020). As of today, none of the crystal structures of membrane-bound FADs from bacteria have been established and solved; however, the structure prediction of Δ9 acyl-CoA FADs from Pseudomonas sp. AMS8 has been predicted (Garba et al. 2018) (shown in Table 2). It remains a major problem in determining crystal structures due to factors such as difficulties to express and purify (Birch et al. 2018), and identification of suitable conditions that would produce crystals for structure determination. Affinity chromatography is by far the most useful and widely used technology for purifying integral membrane proteins as done by these studies (Chen et al. 2013; Wang et al. 2017; Halim et al. 2021). Recombinant FADs of Δ9, Δ12, and Δ15 from Mortierella alpina were successfully affinity purified on His Mag Sepharose Ni beads with one-step purification in methylotrophic yeast, Pischia pastoris (Chen et al. 2013). Based on a previous study, purified Δ12 FADs was discovered at the expected size of 49 kDa, with a concentration of only 0.728 mg/mL of culture (Halim et al. 2021). However, a study reported that the concentration of 4.6 mg/L of culture for Δ15 FADs 22.5 mg/L of culture for Δ12 FADs and 37.5 mg/L of culture Δ9 FADs could be obtained through protein solubilization and affinity purifications (Chen et al. 2013). Therefore, there is a need to optimize a few parameters to acquire the desired concentration of crystallization because some membrane proteins may produce crystals at 1 to 2 mg/mL, while others may need a concentration of 20 to 30 mg/mL to crystallize (Li et al. 2014).
Joseph E. Stukey (1990), suggested a topological model for membrane-bound FADs which had four trans-membrane spanning domains with N and C termini in the cytosolic side of the membrane, as well as a catalytic site based on Δ9 stearoyl-CoA FADs in yeast and rat. The amino acid sequences of Δ9 bacterial acyl-lipid FADs contained conserved histidine boxes predicted as “HXXXH,” “HXXHH,” and “HXXHH,” which were identical to those found in mammal, yeast, and fungal cells (Fukuchi-Mizutani et al. 1995, 1998). Alignments of Δ9 acyl-CoA FADs in Pseudomonas sp. AMS8 (Garba et al. 2018) and sequences from several other Δ9 FADs (Alberts et al. 2002) revealed three conserved-histidine boxes at positions 34–39, 71–75, and 206–210, which are common to all membrane-bound FADs in bacteria (Alberts et al. 2002; Pereira et al. 2004), fungi (Wang et al. 2015a, b), and animals (Bai et al. 2015). Membrane-bound FADs are thought to have different active sites than soluble desaturases, with a cleft into which substrates enter laterally rather than deep binding cavity; thus, substrates enter in extended conformation as soluble FADs do (Shanklin et al. 2009). The putative active site for Δ9 acyl-CoA FADs in Pseudomonas sp. AMS8 was discovered within the conserved histidine-box region, which had previously been identified as a probable binding site for this enzyme (Garba et al. 2018). Certain residues, consisting of Ile, Tyr, Val, Gly, Pro, Glu, His, Arg, Thr, and Lys were found to bind the palmitic acid, which acts as a substrate for this Δ9 acyl-CoA FADs (Garba et al. 2018). In the crystal structures of Δ9 mSCD1and Δ9 hSCD1, amino acid residues, Ile, Val, Gly, and Arg, are comparable to binding residues (Wang et al. 2015a, b).
A study revealed that three transmembrane domains traverse the membrane bi-layer twice, with both protein termini towards the cytoplasm, using CCTOP to predict the structure of Pseudomonas sp. AMS8 Δ9 acyl-CoA FADs, whereby each domain of TM1, TM2, and TM3 has 20 (Leu13-Leu33), 25 (Leu135-Ile159), and 20 (Met162-Tyr181) amino acid residues, respectively (Garba et al. 2018). This is contrary to the predicted structures of Δ9 mSCD1 (Bai et al. 2015) and Δ9 hSCD1 that had a stem of four transmembrane proteins in each structure (Wang et al. 2015a, b). The topology of Δ9 hSCD1 crystal structure matches the prior predictions of the topology of Δ9 mSCD1. For Δ9 mSCD1, the trans-membrane α-helices (TM) are built in a cone-like shape, where TM1 and TM2 sandwich TM4. TM2 and TM4 protrude, and the membrane makes three helical twists on the cytoplasmic side, which offers a few amino acids for the di-metal active site (Bai et al. 2015). Δ9 hSCD1 exposes four α-helices of trans-membrane and a cytoplasmic domain, which has mushroom-like architecture (Wang et al. 2015a, b). TM1–TM4 of the TM stem apparently span the ER membrane (Fig. 1). The TM stem joins through two short ER-lumen loops (LL1 and LL2) and a long polypeptide linkage, which extends into the cytoplasmic side of the ER (Wang et al. 2015a, b).
For acyl-lipid FADs, according to the projected positioning of Δ12 acyl-lipid FADs in relation to the membrane, the enzyme traverses the membrane four times and exposes three histidine clusters that comprise the desaturase catalytic center (Los and Murata 1998). A study suggested that the structure of acyl-lipid FADs is like acyl-CoA FADs, as they are mainly built up of transmembrane proteins (Murata and Wada 1995). Synechocystis sp. PCC 6803’s Δ12 acyl-lipid FADs revealed that a conserved histidine residue substituted by different amino acid residues might lead to the loss of active enzyme as the residues are unable to bind to ferric iron where it is needed (Avelange-Macherel et al. 1995; Schneider et al. 1992). The residues Lys, Glu, and His are also present in probable acyl-ACP FAD genes desA2, with the metal-binding ligands of a plant Δ9 FADs (Alberts et al. 2002). Δ12 acyl-lipid FADs from Synechocystis sp. PCC 6803 had four transmembrane domains, exposing three histidine-box motifs to the cytoplasmic side (Avelange-Macherel et al. 1995) like the crystal structure. However, Δ5 acyl-lipid FADs from Bacillus subtilis was reported with six transmembrane domains (two additional from the proposed topology of membrane desaturase in bacteria) and one membrane-associated domain, which likely represent a substrate-binding motif with three conserved histidine motifs found on the membrane of cytoplasm (Diaz et al. 2002). The presence of two additional trans-membrane domains might be essential for the desaturation of acyl chain in the activity of acyl-lipid desaturase (Yeagle 2016).
Catalytic mechanisms
The catalytic sites of membrane-bound FADs are often situated on the cytosolic side of the membrane, where they have access to substrates and co-factors such as NADH, NADPH, AMP, and ATP (Wang et al. 2013). The predicted structure of Δ9 FADs in Pseudomonas sp. AMS8 revealed that three conserved-histidine boxes found in all classes of these enzymes play a role in binding to two irons at the catalytic core (Garba et al. 2018). Site-directed mutagenesis of rat stearoyl-CoA Δ9 FADs established the importance of these histidine residues in the conserved histidine-box motifs, while residues bordering the conserved area exhibit essential catalytic features in plant FAD2 desaturases and similar enzymes (Meesapyodsuk et al. 2007). Two metal-binding sites in the cytoplasmic cap domain of Δ9 hSCD1 have been discovered. Zn1+ coordinated with five histidine residues: His120, His125, His157, His161, and His301, whereas Zn2+ coordinated with four histidine residues: His160, His269, His298, and His302, as well as a water molecule (Wang et al. 2015a, b) as shown in Fig. 2. The two zinc ions were within 4.5–5.5 Å of stearoyl-CoA carbons 9 and 10, identifying the Δ9 hSCD1 dehydrogenation catalytic center. Two zinc metal ions have also been found in Δ9 mSCD1, with the first zinc surrounded by four histidine residues and the second zinc surrounded by five histidine residues (Wang et al. 2015a, b). The positions of the first and second zinc are identified as 5.2 Å from carbon 9 and 4.7 Å from carbon 10, respectively, while the positions of nine histidine residues are highly conserved (Bai et al. 2015). According to a study, eight histidine residues correspond to three conserved histidine residue motifs, which are one of the characteristics of membrane-bound desaturase (shown in Table 1), alkane hydroxylase, and xylene monooxygenases (Sperling et al. 2003; Shanklin and Cahoon 1998). In addition, with one missing ligand, the coordination of both zinc ions was consistent with an octahedral shape. Nagao et al. (2019) stated that nine conserved histidine residues in humans (His120, His125, His157, His160, His161, His298, His301, and His 302), in mouse (His116, His121, His153, His156, His157, His265, His297, His298), and one conserved Asparagine residue (Asn265 in human; Asn261 in mouse), respectively, in TM2, TM4, the cytosolic loop between TM2 and TM3, and the C-terminal domain. However, substitution of a single histidine residue among the conserved histidine residues eliminates the enzyme’s ability to complement the growth defects (Shanklin et al. 1994).
The sequence identity of Δ9 hSCD1 and Δ9 mSCD1 is 84% similar, and the structures of both are almost identical to each other with an RMSD of 0.35 Å (Shen et al. 2020). However, the structural fold of SCD1 is different compared to soluble acyl-ACP desaturase that catalyzes a comparable reaction (Behrouzian et al. 2002; Whittle et al. 2008). A study suggested that the existence of two zinc ions, instead of two iron ions, in the catalytic center might cause the enzymes to be inactive. According to a study published in Δ9 mSCD1, the distance between the two metal ions is 6.4 Å, which is the largest distance between two metal ions ever observed in any soluble di-iron enzymes’ solved structures (Hogbom et al. 2002; Lindqvist et al. 1996; Sazinsky and Lippard 2006). Therefore, a recent work was undertaken to insert iron into Δ9 mSCD1 and the structure was resolved by molecular replacement, by utilizing the preceding Δ9 mSCD1; the search model with the presence of both metal ions and histidine residues was at 3.5 Å (6WF2) (Shen et al. 2020). A study revealed that the structures of Δ9 mSCD1 and Δ9 hSCD1 were almost similar, whereby the structure of mSCD1 with insertion of iron was nearly identical to Δ9 mSCD1’s prior structure, with RMSD of 0.3 Å for all backbone atoms (Shen et al. 2020). The solved structure also exhibited the di-metal center, coordinated by the conserved histidine residues, which also portrayed similar coordination geometry (Bai et al. 2015). Furthermore, as Zn2+ has the ionic radius and charge property like Fe2+, it can be used as a suitable surrogate to maintain structural integrity; misincorporation of Zn2+ fails to accelerate the enzyme reaction mechanism. However, it is not unique to detect Zn2+ in the sites that bind iron; few reports suggested the presence of Fe2+ in the structure of yeast fatty acid α-hydroxylase (Zhu et al. 2015) and quinol-dependent nitric oxide reductase (PDB 3AYF) (Hino et al. 2010). A study revealed that Zn2+ has an ionic radius of 0.88 Å, whereby the size and charge are similar to Fe2+ (0.92 Å), so it might have the potential to be a substitute (Shen et al. 2020). Through comparisons of the di-iron clusters of several di-iron-containing enzymes, relevant information about the tuning of di-iron centers about various chemical reactivities has been made available (Shanklin and Cahoon 1998; Shanklin et al. 2009; Soon and Lippard 2004). This indicates that the iron-bound oxidant is present in the experiment (Shanklin and Cahoon, 1998).
Stearoyl-CoA binding site
Desaturation begins with the energy-intensive abstraction of hydrogen from a methylene group using stearoyl-CoA as the substrate, which results in the insertion of double bonds within the fatty acyl chain (Shanklin et al., 2009) (shown in Fig. 3). Multiple proteins share an active-site di-iron cluster in this process, including methane monooxygenase, ribonucleotide reductase, rubrerythrins, and a variety of oxidase enzymes to recruit and activate molecular oxygen (Shanklin et al. 2009; Soon and Lippard 2004). A study reported the interaction between SCD1 with the stearoyl-CoA, in which the polar residues of the surface of SCD1 guide the fatty acyl chain towards the tunnel located between TM2 and TM4 (Wang et al. 2015a, b). The substrate must induce a cis conformation between C9 and C10 before it aligns proximally at the di-metal center for the fatty acyl chain to fit the tunnel (Wang et al. 2015a, b). The di-metal center’s relative orientation to C9 and C10 is necessary for the reaction of pro-R dehydrogenation, which demands additional elements (Enoch et al. 1976). The CoA groups (adenosine, diphosphate, and pantothenate) interact with the polar remains on SCD1’s surface via hydrogen-bonding and electrostatic interactions in this CoA-SCD1 interface (shown in Fig. 4). The adenosine group of CoA mainly forms hydrogen-bonding connections with the side chains of residues Arg155 and Asp156 from CH2 and with the main chain carbonyl group of Gly197 (Wang et al. 2015a, b).
Current study and future perspectives of FADs
Cellular biology and biochemistry make membrane protein an interesting subject of study. It represents between 20 and 30% of the proteins of most organisms and more than 60% of drug targets (Attwood and Schioth, 2021) important for pharmaceuticals, and yet very few structures of these proteins have been solved. Over the last 15 years, membrane protein structural biology has made great leaps despite their instability (Birch et al. 2020). To date, the database of the “Membrane Proteins of Known 3D Structure” has shown an additional 1472 unique membrane protein structures, with 6252 coordinate files of membrane proteins deposited in the Protein Data Bank (PDB) in August 2022. This contrasts with only 2588 coordinated files of membrane proteins deposited in the PDB with a list of 1141 unique membrane protein structures in September 2020 (Birch et al. 2020). As of today, none of the crystal structures of membrane-bound FADs from bacteria has been established and solved. It remains a major problem in determining crystal structures of FADs for some reasons, such as difficulties in expressing and purifying them (Birch et al. 2020), as well as the identifications of suitable conditions for production of crystals for structure determination. However, different strategies, such as using mutant host strains E. coli C41(DE3) and C43(DE3) known as “Walker strain” and shifting towards eukaryotic targets and expression systems might be promising to improve the protein production yield of FADs (Karyolaimos and de Gier 2021). An effective recombinant membrane protein production plan must consider a broad range of factors, including construct design, expression system, extraction, and purification methods (Birch et al. 2020). As more information becomes available, the easier it may become to develop predictors and other computer-aided tools that can assist in designing strategies to enhance protein production yield (Cetnar and Salis 2021). Furthermore, having purified FADs paves the way towards the characterization biochemically and biophysically (Halim et al. 2021) and may be beneficial for the specification of function, structure, and interactions of membrane proteins.
The success of the purification and crystallization of membrane proteins has greatly depended on the detergents’ ability to maintain a target membrane protein in a functional, folded state in the absence of membrane (Kermani 2020). It is crucial to identify the ideal combination of constructs and detergents as it is important to work with stable, correctly folded protein (Birch et al. 2020). The primary method for membrane protein crystallization has relied on detergents' ability and successful crystallization screening kits (Healey et al. 2021) such as MemGold, MemAdvantage, and MemMeso, which are credited to advances in tools and reagents used to manipulate membrane proteins. This improvement has been accompanied by the development of cryogenic electron microscopy (cryo-EM) and X-ray crystallography as 80% of membrane protein structures have been solved by this leading technique (Kermani 2020). Every new discovery in membrane protein structural biology shifts the bottlenecks in a new direction. The understanding of the correlative techniques combining super resolution light microscopy with cryo-correlative light and electron microscopy (cryo-CLEM), cryo-ET, or cryo-X-ray tomography will promote understanding of the structural information of membrane proteins (Birch et al. 2020). These emerging innovations are promising for future research efforts to overcome the intrinsic difficulties and should have a major impact on understanding of membrane proteins as the ability of this electron microscope with advances in cryo-technology may provide high-resolution images of detailed structural and chemical information about proteins.
Conclusion
In summary, this manuscript shows that FADs have been isolated from all groups of organisms. Class of soluble desaturases has been extensively studied, whereas the membrane-bound class of desaturases lacks sufficient information on its structure–function relationships due to its difficult nature to purify. Biosynthesis of FADs revealed that this enzyme is responsible for the production of polyunsaturated fatty acids through the mechanism of desaturation reaction. Fatty acid desaturation involves an enzymatic reaction in which a double bond is introduced into an acyl chain, and a molecule of oxygen is completely reduced to water. Role of unsaturated FADs tend to exert a variety effect in microorganisms, plants, humans, and other mammals. The 3D structures of FADs demonstrate almost similar structural architecture, specifically in humans and mammals, with four transmembrane helices arranged in a cone shape. FADs possess a unique di-iron catalytic site, which occurs due to different di-iron metal ions; however, they can maintain the integrity of the structure.
References
Alberts B, Johnson A, Lewis J (2002) Molecular biology of the cell. 4th edition. New York: Garland Science; Membrane Proteins. ISBN-10: 0-8153-4072-9
Ahern K, Rajagopal I (2021) Chapter 6.12 Fatty acid synthesis, Libretexts, Oregon State University. https://bio.libretexts.org/@go/page/2999
Attwood MM, Schiöth HB (2021) Characterization of five transmembrane proteins: with focus on the Tweety, Sideroflexin, and YIP1 Domain Families. Front Cell Dev Biol 9:708754. https://doi.org/10.3389/fcell.2021.708754
Avelange-Macherel MH, Macherel D, Wada H, Murata N (1995) Site-directed mutagenesis of histidine in the Δ12 acyl-lipid desaturase of Synechocystis. FEBS Letter 361:111–114. https://doi.org/10.1016/0014-5793(95)00163-4
Bai Y, McCoy JG, Levin EJ, Sobrado P, Rajashankar KR, Fox BG, Zhou M (2015) X-ray structure of a mammalian stearoyl-CoA desaturase. Nature 524:252–256. https://doi.org/10.1038/nature14549
Bale NJ, Rijpstra WIC, Sahonero-Canavesi DX, Oshkin IY, Belova SE, Dedysh SN, Sinninghe Damsté JS (2019) Fatty Acid and hopanoid adaption to cold in the methanotroph Methylovulum psychrotolerans. Front Microbiol 10:589. https://doi.org/10.3389/fmicb.2019.00589
Behrouzian B, Savile CK, Dawson B, Buist PH, Shanklin J (2002) Exploring the hydroxylation−dehydrogenation connection: novel catalytic activity of castor stearoyl-ACP Δ9 desaturase. J Am Chem Soc 124:3277–3283. https://doi.org/10.1021/ja012252l
Behrouzian B, Buist PH (2003) Mechanism of fatty acid desaturation: a bioorganic perspective. Prostaglandins Leukot Essent Fatty Acids 68:107–112. https://doi.org/10.1016/s0952-3278(02)00260-0
Beld J, Lee DJ, Burkart MD (2015) Fatty acid biosynthesis revisited: structure elucidation and metabolic engineering. Mol BioSyst 11(1):38–59. https://doi.org/10.1039/c4mb00443d
Birch J, Axford D, Foadi J, Meyer A, Eckhardt A, Thielmann Y, Moraes I (2018) The fine art of integral membrane protein crystallisation. Methods 147:150–162. https://doi.org/10.1016/j.ymeth.2018.05.014
Birch J, Cheruvara H, Gamage N, Harrison PJ, Lithgo R, Quigley A (2020) Changes in membrane protein structural biology. Biology 9(11):401. https://doi.org/10.3390/biology9110401
Bouwens M, van de Rest O, Dellschaft N, Bromhaar MG, de Groot LC, Geleijnse JM, Afman LA (2009) Fish-oil supplementation induces antiinflammatory gene expression profiles in human blood mononuclear cells. Am J Clin Nutr 90(2):415–424. https://doi.org/10.3945/ajcn.2009.27680
Brouwers H, von Hegedus J, Toes R, Kloppenburg M, Ioan-Facsinay A (2015) Lipid mediators of inflammation in rheumatoid arthritis and osteoarthritis. Best Pract Res Clin Rheumatol 29:741–755. https://doi.org/10.1016/j.berh.2016.02.003
Buist PH (2004) Fatty acid desaturases: selecting the dehydrogenation channel. Nat Prod Rep 21:249. https://doi.org/10.1039/b302094k
Cao B, Wang D, Sun X, Yan J, Ren B, Lu Q, Liu Y, Zeng J, Huang N, Xie Q, Gu H, Wang J (2018) Alterations of eicosanoids and related mediators in patients with schizophrenia meta-analysis View project Alterations of eicosanoids and related mediators in patients with schizophrenia. J Psychiatr Res 102:168–178. https://doi.org/10.1016/j.jpsychires.2018.04.002
Caspi A, Williams B, Kim-Cohen J, Craig IW, Milne BJ, Poulton R, Schalkwyk LC, Taylor A, Werts H, Moffitt TE (2007) Moderation of breastfeeding effects on the IQ by genetic variation in fatty acid metabolism. Proc Natl Acad Sci USA 104:18860–18865. https://doi.org/10.1073/pnas.0704292104
Castro LFC, Tocher DR, Monroig O (2016) Long-chain polyunsaturated fatty acid biosynthesis in chordates: insights into the evolution of Fads and Elovl gene repertoire. Progr Lipid Res 62:25–40. https://doi.org/10.1016/j.plipres.2016.01.001
Cetnar DP, Salis HM (2021) Systematic quantification of sequence and structural determinants controlling mRNA stability in bacterial operons. ACS Synth Biol 10(2):318–332. https://doi.org/10.1021/acssynbio.0c00471
Chaves H, Singh RB, Khan S, Wilczynska A, Takahashi T (2019) Chapter 14 - High omega-6/omega-3 fatty acid ratio diets and risk of noncommunicable diseases. The role of functional food security in global health. 217–259. Academic Press. https://doi.org/10.1016/b978-0-12-813148-0.00014-1
Chen G, Shujie Q, Wang Q, Bian F, Peng Z, Zhang Y, Ge H, Yu J, Xuan N, Bi Y, He Q (2014) Transgenic expression of delta-6 and delta-15 fatty acid desaturases enhances omega-3 polyunsaturated fatty acid accumulation in Synechocystis sp. PCC6803. Biotechnol Biofuels 7:32. https://doi.org/10.1186/1754-6834-7-32
Chen H, Gu Z, Zhang HM, Chen W, Lowther WT, Chen YQ (2013) Expression and purification of integral membrane fatty acid desaturases. PLoS ONE 8:e58139. https://doi.org/10.1371/journal.pone.0058139
Chiu CC, Su KP, Cheng T-C, Liu H-C, Chang C-J, Dewey ME, Stewart R, Huang S-Y (2008) The effects of omega-3 fatty acids monotherapy in Alzheimer’s disease and mild cognitive impairment: a preliminary randomized double-blind placebo-controlled study. Prog Neuropsychopharmacol Biol Psychiatry 32:1538–1544. https://doi.org/10.1016/j.pnpbp.2008.05.015
Czumaj A, Sledzinski T (2020) Biological role of unsaturated fatty acid desaturase in health and disease. Nutrients 12:356. https://doi.org/10.3390/nu12020356
Diaz AR, Mansilla MC, Vila AJ, de Mendoza D (2002) Membrane topology of the acyl-lipid desaturase from Bacillus subtilis. J Biol Chem 277:48099–48106. https://doi.org/10.1074/jbc.M208960200
Ding Z, Liu G-L, Li X, Chen X-Y, Wu Y-X, Cui C-C, Zhang X, Yang G, Xie L (2016) Association of polyunsaturated fatty acids in breast milk with fatty acid desaturase gene polymorphisms among Chinese lactating mothers. Prostaglandins Leukot Essent Fat Acid 109:66–71. https://doi.org/10.1016/j.plefa.2016.03.009
Diomandé SE, Guinebretière M-H, De Sarrau B, Nguyen-the C, Broussolle V, Brillard J (2015) Fatty acid profiles and desaturase-encoding genes are different in thermo- and psychrotolerant strains of the Bacillus cereus Group. BMC Res Notes 8:329. https://doi.org/10.1186/s13104-015-1288-4
Domergue F, Spiekermann P, Lerchl J, Beckmann C, Kilian O, Kroth PG (2003) New insight into Phaeodactylum tricornutum fatty acid metabolism cloning and functional characterization of plastidial and microsomal delta12-fatty acid desaturases. Plant Physiol 131:1648–1660. https://doi.org/10.1104/pp.102.018317
Drew DE, von Heijne G, Nordlund P, de Gier JW (2001) Green fluorescent protein as an indicator to monitor membrane protein overexpression in Escherichia coli. FEBS Lett 507:220–224. https://doi.org/10.1016/s0014-5793(01)02980-5
Dunstan JA, Mitoulas LR, Dixon G, Doherty DA, Hartmann PE, Simmer K, Prescott SL (2007) The effects of fish oil supplementation in pregnancy on breast milk fatty acid composition over the course of lactation: a randomized controlled trial. Pediatr Res 62:689–694. https://doi.org/10.1203/PDR.0b013e318159a93a
Dyer DH, Lyle KS, Rayment I, Fox BG (2005) X-ray structure of putative acyl-ACP desaturase DesA2 from Mycobacterium tuberculosis H37Rv. Protein Sci 14:1508–1517. https://doi.org/10.1110/ps.041288005
Echeverria F, Ortiz M, Valenzuela R, Videla LA (2016) Long-chain polyunsaturated fatty acids regulation of PPARs, signaling: relationship to tissue development and aging. Prostag Leukot Essent Fat Acids 114:28–34. https://doi.org/10.1016/j.plefa.2016.10.001
Endo J, Arita M (2016) Cardio protective mechanism of omega-3 polyunsaturated fatty acids. J Cardiol 67:22–27. https://doi.org/10.1016/j.jjcc.2015.08.002
Enoch HG, Catala A, Strittmatter P (1976) Mechanism of rat liver microsomal stearyl- CoA desaturase: studies of the substrate specificity, enzyme-substrate interactions, and the function of lipid. J Biol Chem 251:5095–5103.
Fu Y, Fan X, Li X, Wang H, Chen H (2013) The desaturase OPIN17 from Phytophthora infestans converts arachidonic acid to eicosapentaenoic acid in CHO cells. Biotechnol Appl Biochem 171:975–988. https://doi.org/10.1007/s12010-013-0332-x
Fukuchi-Mizutani M, Savin K, Cornish E, Tanaka Y, Ashikari T, Kusimi T, Murata N (1995) Senescence-induced expression of a homologue of Δ9 desaturase in rose petals. Plant Mol Biol 29:627635. https://doi.org/10.1007/BF00041154
Fukuchi-Mizutani M, Tasaka Y, Tanaka Y, Ashikari T, Kusimi T, Murata N (1998) Characterization of Δ9 acyl-lipid desaturase homologues from Arabidopsis thaliana. Plant Cell Physiol 39:247–253. https://doi.org/10.1093/oxfordjournals.pcp.a029364
Garba L, Ali MSM, Oslan SN, Rahman RNZRA (2017) Review on fatty acid desaturases and their roles in temperature acclimatisation. J Appl Sci 17:282–295. https://doi.org/10.3923/jas.2017.282.295
Garba L, Yusoff MAM, Halim KBA, Ishak SNH, Ali MSM, Oslan SN, Rahman RNZRA (2018) Homology modeling and docking studies of a Δ9-fatty acid desaturase from a Cold-tolerant Pseudomonas sp. AMS8. Peer J 6:e4347(1–21). https://doi.org/10.7717/peerj.4347
Gould JF, Smithers LG (2019) Prenatal n-3 long-chain polyunsaturated fatty acids and children’s executive functions. Omega Fat Acids Brain Neurol Heal :83–105. Academic Press. https://doi.org/10.1016/B978-0-12-815238-6.00006-7
Guy JE, Whittle E, Kumaran D, Lindqvist Y, Shanklin J (2007) The crystal structure of the Ivy Δ4-16:0-ACP desaturase reveals structural details of the oxidized active site and potential determinants of regioselectivity. J Biol Chem 282:19863–19871. https://doi.org/10.1074/jbc.M702520200
Halim NFAA, Ali MSM, Leow ATC, Rahman RNZRA (2021) Membrane-bound Δ12 fatty acid desaturase (FAD12); from Brassica napus to E. coli expression system. Int J Biol Macromol 180:242–251. https://doi.org/10.1016/j.ijbiomac.2021.03.072
Han XT, Wang S, Zheng L, Liu WS (2019) Identification and characterization of a delta-12 fatty acid desaturase gene from marine microalgae Isochrysis galbana. Acta Oceanol Sin 38:107–113. https://doi.org/10.1007/s13131-019-1354-1
Hastings N, Agaba M, Tocher DR, Leaver MJ, Dick JR, Sargent JR, Teale AJ (2001) A vertebrate fatty acid desaturase with Δ5 and Δ6 activities. Proc Natl Acad Sci USA 98:14304–14309. https://doi.org/10.1073/pnas.251516598
Hazel JR, Williams EE (1990) The role of alterations in membrane lipid composition in enabling physiological adaptation of organisms to their physical environment. Prog Lipid Res 29:167–227. https://doi.org/10.1016/0163-7827(90)90002-3
He M, Qin C-X, Wang X, Ding N-Z (2020) Plant unsaturated fatty acids: biosynthesis and regulation. Frontiers in Plant Science 11:390. https://doi.org/10.3389/fpls.2020.00390
Healey RD, Basu S, Humm A-S, Leyrat C, Cong X, Golebiowski DF, Pica A, Granier S, Márquez JA (2021) An automated platform for structural analysis of membrane proteins through serial crystallography. Cell Reports Methods 1:6. https://doi.org/10.1016/j.crmeth.2021.100102
Healy-Stoffel M, Levant B (2018) N-3 (Omega-3) Fatty acids: effects on brain dopamine systems and potential role in the etiology and treatment of neuropsychiatric disorders. CNS Neurol Disord Drug Targets 17:216–232. https://doi.org/10.2174/1871527317666180412153612
Helland IB, Smith L, Blomen B, Saarem K, Saugstad OD, Drevon CA (2008) Effect of supplementing pregnant and lactating mothers with n-3 very- long-chain fatty acids on children’s IQ and body mass index at 7 years of age. Pediatrics 122:472–479. https://doi.org/10.1542/peds.2007-2762
Hino T, Matsumoto Y, Nagano S, Sugimoto H, Fukumori Y, Murata T, Iwata S, Shiro Y (2010) Structural basis of biological N2O generation by bacterial nitric oxide reductase. Science 330:1666–1670. https://doi.org/10.1126/science.1195591
Hogbom M, Huque Y, Sjoberg BM, Nordlund P (2002) Crystal structure of the di-iron/radical protein of ribonucleotide reductase from Corynebacterium ammoniagenes. Biochemistry 41:1381–1389. https://doi.org/10.1021/bi011429l
Huang YS, Chaudhary SJ, Thurmond M Jr, Bobik EG, Yuan L, Chan GM, Kirchner SJ, Mukerji P, Knutzon DS (1999) Cloning of delta12- and delta6-desaturases from Mortierella alpina and recombinant production of gamma-linolenic acid in Saccharomyces cerevisiae. Lipids 34:649–659. https://doi.org/10.1007/s11745-999-0410-8
Hussein N, Ah-Sing E, Wilkinson P, Leach C, Griffin BA, Millward DJ (2005) Long-chain conversion of [13C] linoleic acid and alpha-linolenic acid in response to marked changes in their dietary intake in men. J Lipid Res 46:269–280. https://doi.org/10.1194/jlr.M400225-JLR200
Iba K (2002) Acclimative response to temperature stress in higher plants: approaches of gene engineering for temperature tolerance. Annu Rev Plant Biol 53:225–245. https://doi.org/10.1146/annurev.arplant.53.100201.160729
Itoh R, Toda K, Takahashi H, Takan H, Kuroiwa T (1998) Δ-9 desaturase gene containing a carboxyl-terminal cytochrome b5 domain from the red alga Cyanidioschyzon merolae. Curr Genet 33:165–170. https://doi.org/10.1007/s002940050323
Karyolaimos A, de Gier J-W (2021) Strategies to enhance periplasmic recombinant protein production yields in Escherichia coli. Front Bioeng Biotechnol 9:797334. https://doi.org/10.3389/fbioe.2021.797334
Kermani AA (2020) A guide to membrane protein X-ray crystallography. FEBS J 288:5788–5804. https://doi.org/10.1111/febs.15676
Keweloh H, Heipieper HJ (1996) Trans unsaturated fatty acids in bacteria. Lipids 31(2):129–137. https://doi.org/10.1007/bf02522611
Krauss-Etschmann S, Hartl D, Rzehak P, Heinrich J, Shadid R, del Carmen Ramirez-Tortosa M, Campoy C, Pardillo S, Schendel DJ, Decsi T, Demmelmair H, Koletzko BV, Nutraceuticals for Healthier Life Study Group (2008) Decreased cord blood IL-4, IL-13, and CCR4 and increased TGF- beta levels after fish oil supplementation of pregnant women. J Allergy Clin Immunol 121:464–470. https://doi.org/10.1016/j.jaci.2007.09.018
Krauss-Etschmann S, Shadid R, Campoy C, Hoster E, Demmelmair H, Jimenez M, Gil A, Montserrat R, Veszpremi B, Decsi T, Koletzko BV, (NUHEAL) Study Group (2007) Effects of fish- oil and folate supplementation of pregnant women on maternal and fetal plasma concentrations of docosahexaenoic acid and eicosapentaenoic acid: a European randomized multicenter trial. Am J Clin Nutr 85:1392-1400. https://doi.org/10.1093/ajcn/85.5.1392
LaBrant E, Barnes AC, Roston RL (2018) Lipid transport required to make lipids of photosynthetic membranes. Photosynth Res 138:345–360. https://doi.org/10.1007/s11120-018-0545-5
Laoteng K, Anjard C, Rachadawong S, Tanticharoen M, Maresca B, Cheevadhanarak S (1999) Mucor rouxii Δ9-desaturase gene is transcriptionally regulated during cell growth and by low temperature. Mol Biol Res Commun 1:36–43. https://doi.org/10.1006/mcbr.1999.0107
Laoten K, Mannontarat R, Tanticharoen M, Cheevadhanarak S (2000) Δ6-desaturase of Mucor rouxii with high similarity to plant Δ6-desaturse and its heterologous expression in Saccharomyces cerevisiae. Biochem Biophys Res Commun 279:17–22. https://doi.org/10.1006/bbrc.2000.3856
Leaf DA, Hatcher L (2009) The effect of lean fish consumption on triglyceride levels. Phys Sportsmed 37:37–43. https://doi.org/10.3810/psm.2009.04.1681
Lee JM, Lee H, Kang SB, Park WJ (2016) Fatty acid desaturases, polyunsaturated fatty acid regulation, and biotechnological advances. Nutrients 8(1):23. https://doi.org/10.3390/nu8010023
Li D, Howe N, Dukkipati A, Shah STA, Bax BD, Edge C, Bridges A, Hardwicke P, Singh OMP, Giblin G, Pautsch A, Pfau R, Schnapp G, Wang M, Olieric V, Caffrey M (2014) Crystallizing membrane proteins in the lipidic mesophase. Experience with human prostaglandin E2 synthase 1 and an evolving strategy 14(4): 2034–2047. https://doi.org/10.1021/cg500157x
Li N, Xu C, Li-Beisson Y, Philippar K (2016) Fatty acid and lipid transport in plant cells. Trends Plant Sci 21:145–158. https://doi.org/10.1016/j.tplants.2015.10.011
Li D, Moorman R, Vanhercke T, Petrie J, Singh S, Jackson CJ (2016) Classification and substrate head-group specificity of membrane fatty acid desaturases. Comput Struct Biotechnol J 14:341–349. https://doi.org/10.1016/j.csbj.2016.08.003
Li Y, Dietrich M, Schmid RD, He B, Ouyang P, Urlacher VB (2009) Identification and functional expression of a Δ9 fatty acid desaturase from the marine bacterium Pseudoalteromonas sp. MLY15. J Mol Catal b: Enzym 56:96–101. https://doi.org/10.1007/s11745-007-3150-5
Li Q, Shen W, Qian Z, Fowler DB, Zou J (2016) Adjustment of lipid pathways in plant adaptation to temperature stress. Plant Signal Behave 11(1):e105846. https://doi.org/10.1080/15592324.2015.1058461
Liu Y, McNamara RK (2011) Elevated Delta-6 desaturase (FADS2) gene expression in the prefrontal cortex of patients with bipolar disorder. J Psychiatr Res 45:269–272. https://doi.org/10.1016/j.jpsychires.2010.06.010
Lindqvist Y, Huang W, Schneider G, Shanklin J (1996) Crystal structure of delta9 stearoyl-acyl carrier protein desaturase from castor seed and its relationship to other di-iron proteins. EMBO J 15:4081–4092
Lomascolo A, Dubreucq E, Galzy P (1996) Study of the Δ12-desaturase system of Lipomyces starkeyi. Lipids 31:253–259. https://doi.org/10.1007/BF02529871
Lorente-Cebrián S, Costa AGV, Navas-Carretero S, Zabala M, Laiglesia LM, Martínez JA, Moreno-Aliaga MJ (2015) An update on the role of omega-3 fatty acids on inflammatory and degenerative diseases. J Physiol Biochem 71:341–349. https://doi.org/10.1007/s13105-015-0395-y
Los DA, Murata N (1998) Structure and expression of fatty acid desaturases. BBA 1394:3–15. https://doi.org/10.1016/S0005-2760(98)00091-5
Lou Y, Schwender J, Shanklin J (2014) FAD2 and FAD3 desaturases form heterodimers that facilitate metabolic channelingin vivo. J Biol Chem 289(26):17996–18007. https://doi.org/10.1074/jbc.m114.572883
Lu YD, Chi XY, Yang QL, Li ZX, Liu SF, Gan QH (2009) Molecular cloning and stress-dependent expression of a gene encoding Δ12-fatty acid desaturase in the Antarctic microalga Chlorella vulgaris NJ-7. Extremophiles 13:875–884. https://doi.org/10.1007/s00792-009-0275-x
Mann NJ, O’Connell SL, Baldwin KM, Singh I, Meyer BJ (2010) Effects of seal oil and tuna-fish oil on platelet parameters and plasma lipid levels in healthy subjects. Lipids 45:669–681. https://doi.org/10.1007/s11745-010-3450-z
Martin-Montalvo A, Sun Y, Diaz-Ruiz A, Ali A, Gutierrez V, Palacios HH, Curtis J, Siendones E, Ariza, J, Abulwerdi GA, Sun X, Wang AX (2016) Cytochrome b5 reductase and the control of lipid metabolism and healthspan. NPJ Aging Mech 2. https://doi.org/10.1038/npjamd.2016.6. eCollection
Martinez-Rivas JM, Sperling P, Luehs W, Heinz E (2001) Spatial and temporal regulation of three different microsomal oleate desaturase genes (FAD2) from normal-type and high-oleic varieties of sunflower (Helianthus annuus L). Mol Breed 8:159–168. https://doi.org/10.1023/A:1013324329322
Matsuda T, Sakaguchi K, Hamaguchi R, Kobayashi T, Abe E, Hama Y, Hayashi M, Honda D, Okita Y, Sugimoto S, Okino N, Ito M (2012) Analysis of Δ12-fatty acid desaturase function revealed that two distinct pathways are active for the synthesis of PUFAs in T. aureum ATCC 34304. J Lipid Res 53:1210–1222. https://doi.org/10.1194/jlr.M024935
Meesapyodsuk D, Reed DW, Covello PS, Qiu X (2007) Primary structure, regioselectivity, and evolution of the membrane-bound fatty acid desaturases of Claviceps purpurea. J Biol Chem 282:20191–20199. https://doi.org/10.1074/jbc.M702196200
Michaelson LV, Lazarus CM, Griffiths G, Napier JA, Stobart AK (1998) Isolation of a Δ5-fatty acid desaturase gene from Mortierella alpine. J Biol Chem 273:19055–19059. https://doi.org/10.1074/jbc.273.30.19055
Mitchell AG, Martin CE (1995) A novel cytochrome b5-like domain is linked to the carboxyl terminus of the Saccharomyces cerevisiae Δ-9 fatty acid desaturase. J Biol Chem 270:29766–29772. https://doi.org/10.1074/jbc.270.50.29766
Monroig Ó, Kabeya N (2018) Desaturases and elongases involved in polyunsaturated fatty acid biosynthesis in aquatic invertebrates: a comprehensive review. Fish Sci 84:911–928. https://doi.org/10.1007/s12562-018-1254-x
Muc M, Kreiner-Møller E, Larsen J, Birch S, Pedersen S, Bisgaard H, Lauritzen L (2015) Maternal fatty acid desaturase genotype correlates with infant immune responses at 6 months. Br J Nutr 114:891–898. https://doi.org/10.1017/S0007114515002561
Murata N, Wada H (1995) Acyl-lipid desaturases and their importance in the tolerance and acclimatization to cold of cyanobacteria. Biochem J 308:1–8. https://doi.org/10.1042/bj3080001
Mychaleckyj JC, Nayak U, Colgate ER, Zhang D, Carstensen T, Ahmed S, Ahmed T, Mentzer AJ, Alam M, Kirkaptrick BD, Haque R, Faruque ABS, Petri WA Jr (2018) Multiplex genomewide association analysis of breast milk fatty acid composition extends the phenotypic association and potential selection of FADS1 variants to arachidonic acid, a critical infant micronutrient. J Med Genet 55:459–468. https://doi.org/10.1093/ajcn/85.5.1392
Nachtschatt M, Okada S, Speight R (2020) Integral membrane fatty acid desaturases: a review of biochemical, structural, and biotechnological advances. Eur J Lipid Sci Technol 122(12):1–15. https://doi.org/10.1002/ejlt.202000181
Nagao K, Murakami A, Umeda M (2019) Structure and function of Δ9-fatty acid desaturase. Chem Pharm Bull 67(4):327–332. https://doi.org/10.1248/cpb.c18-01001
Nagy K, Tiuca I-D (2017) Importance of fatty acids in physiopathology of human body, Chapter 1: Fatty Acids. IntechOpen. https://doi.org/10.5772/67407
Nakamura MT, Nara TY (2004) Structure, function, and dietary regulation of Δ6, Δ5 and Δ9 desaturases. Annu Rev Nutr 24:345–376. https://doi.org/10.1146/annurev.nutr.24.121803.063211
Nishida I, Murata N (1996) Chilling sensitivity in plants and cyanobacteria: the crucial contribution of membrane lipids. Annu Rev Plant Physiol Plant Mol Biol 47:541–568. https://doi.org/10.1146/annurev.arplant.47.1.541
Olsen SF, Osterdal ML, Salvig JD, Mortensen LM, Rytter D, Secher NJ, Henriksen TB (2008) Fish oil intake compared with olive oil intake in late pregnancy and asthma in the offspring: 16 y of registry-based follow-up from a randomized controlled trial. Am J Clin Nutr 88:167–175. https://doi.org/10.1093/ajcn/88.1.167
Passorn S, Laoteng K, Rachadawong S, Tanticharoen M, Cheevadhanarak S (1999) Heterologous expression of Mucor rouxii Δ (12)-desaturase gene in Saccharomyces cerevisiae. Biochem Bioph Res Co 263:47–51. https://doi.org/10.1006/bbrc.1999.1258
Pereira SL, Leonard AE, Mukerji P (2003) Recent advances in the study of fatty acid desaturases from animals and lower eukaryotes. Prostag Leukot Ess 68:97–104. https://doi.org/10.1016/S0952-3278(02)00259-4
Pereira SL, Huang YS, Bobik EG, Kinney AJ, Stecca KL, Packer JC, Mukerji P (2004) A novel omega 3-fatty acid desaturase involved in the biosynthesis of eicosapentaenoic acid. Biochem J 378:665–671. https://doi.org/10.1042/BJ20031319
Powell DR, Gay JP, Smith M, Wilganowski N, Harris A, Holland A, Reyes M, Kirkham L, Kirpatrick LL, Zambrowicz B, Hansen G, Platt KA, van Slightenhorst I, Ding Z-M, Desai U (2016) Fatty acid desaturase 1 knockout mice are lean with improved glycemic control and decreased development of atheromatous plaque. Diabetes Metab Syndr Obes 9:185–199. https://doi.org/10.2147/DMSO.S106653
Qiu X, Reed DW, Hong H, MacKenzie SL, Covello PS (2001) Identification and analysis of a gene from calendula officinalis encoding a fatty acid conjugase. Plant Physiol 125:847–855. https://doi.org/10.1104/pp.125.2.847
Reddy AS, Thomas TL (1996) Expression of a cyanobacterial Δ6-desaturase gene results in γ-linolenic acid production in transgenic plants. Nat Biotechnol 14:639–642. https://doi.org/10.1038/nbt0596-639
Roman AS, Schreher J, Mackenzie AP, Nathanielsz PW (2006) Omega-3 fatty acids and decidual cell prostaglandin production in response to the inflammatory cytokine IL-1beta. Am J Obstet Gynecol 195:1693–1699. https://doi.org/10.1016/j.ajog.2006.04.009
Ruiz-Lopez N, Sayanova O, Napier JA, Haslam RP (2012) Metabolic engineering of the omega-3 long chain polyunsaturated fatty acid biosynthetic pathway into transgenic plants. J Exp Bot 63(7):2397–2410. https://doi.org/10.1093/jxb/err454
Saini RK, Prasad P, Sreedhar RV, Akhilender NK, Shang X, Keum Y-S (2021) Omega−3 polyunsaturated fatty acids (PUFAs): emerging plant and microbial sources, oxidative stability, bioavailability, and health benefits—a review. Antioxidants 10:1627. https://doi.org/10.3390/antiox10101627
Saito T, Morio T, Ochiai T (2000) A second functional Δ5 fatty acid desaturase the cellular slime mould, Dictyostelium discoideum. Eur J 267:1813–1818. https://doi.org/10.1046/j.1432-1327.2000.01180.x
Saito Y, Yokoyama M, Origasa H, Matsuzaki M, Matsuzawa Y, Ishikawa Y, Oikawa S, Sasaki J, Hishida H, Itakura H, Kita T, Kitabatake A, Nakaya N, Sakata T, Shimada K, Shirato K (2008) Effects of EPA on coronary artery disease in hypercholesterolemic patients with multiple risk factors: sub-analysis of primary prevention cases from the Japan EPA Lipid Intervention Study (JELIS). Atherosclerosis 200:135–140. https://doi.org/10.1016/j.atherosclerosis.2008.06.003
Sakuradani E, Kobayashi M, Shimizu S (1999) Δ9-fatty acid desaturase from arachidonic acid producing fungus. Unique gene sequence and its heterologous expression in fungus Aspergillus. Eur J 260:208–216. https://doi.org/10.1046/j.1432-1327.1999.00131.x
Sayanova O, Haslam R, Guschina I, Lloyd D, Christie WW, Harwood JL (2016) A bifunctional delta 12, delta 15-desaturase from Acanthamoeba castellanii directs the synthesis of highly unusual n-1 series unsaturated fatty acids. J Bio Chem 281:36533–36541. https://doi.org/10.1074/jbc.M605158200
Sazinsky MH, Lippard SJ (2006) Correlating structure with function in bacterial multicomponent monooxygenases and related diiron proteins. Acc Chem Res 39:558–566. https://doi.org/10.1021/ar030204v
Schiano V, Laurenzano E, Brevetti G, De Maio JI, Lanero S, Scopacasa F, Chiariello M (2008) Omega-3 polyunsaturated fatty acid in peripheral arterial disease: effect on lipid pattern, disease severity, inflammation profile, and endothelial function. Clin Nutr 27(2):241–247. https://doi.org/10.1016/j.clnu.2007.11.0
Schneider G, Lindqvist Y, Shanklin J, Somerville C (1992) Preliminary crystallographic data for stearoyl-acyl carrier protein desaturase from castor seed. J Mol Biol 225:561–564. https://doi.org/10.1016/0022-2836(92)90941-c
Shanklin J, Cahoon EB (1998) Desaturation and related modifications of fatty acids. Annu Rev Plant Physiol Plant Mol Biol 49:611–641. https://doi.org/10.1146/annurev.arplant.49.1.611
Shanklin J, Guy E, Mishra G, Lindqvist Y (2009) Desaturases: emerging models for understanding functional diversification of diiron-containing enzymes. J Biol Chem 284:18559–18563. https://doi.org/10.1074/jbc.R900009200
Shanklin J, Whittle E, Fox BG (1994) Eight histidine residues are catalytically essential in a membrane-associated iron enzyme, stearoyl-CoA desaturase, and are conserved in alkane hydroxylase and xylene monooxygenase. Biochemistry 33:12787–12794. https://doi.org/10.1021/bi00209a009
Shen J, Wu G, Tsai A-L, Zhou M (2020) Structure and mechanism unique diiron center in mammalian Stearoyl-CoA desaturase. J Mol Biol 432:5152–5161. https://doi.org/10.1016/j.jmb.2020.05.017
Siliakus MF, van der Oost J, Kengen SWM (2017) Adaptations of archaeal and bacterial membranes to variations in temperature, pH and pressure. Extremophiles 21:651–670. https://doi.org/10.1007/s00792-017-0939-x
Simopoulos AP (2008) The importance of the omega-6/omega-3 fatty acid ratio in cardiovascular disease and other chronic diseases. Exp Biol Med 233(6):674–688. https://doi.org/10.3181/0711-mr-311
Simopoulos AP (2010) The omega-6/omega-3 fatty acid ratio: health implications. Oléagineux, Corps Gras, Lipides 17(5):267–275. https://doi.org/10.1051/ocl.2010.0325
Simopoulos A (2016) An increase in the omega-6/omega-3 fatty acid ratio increases the risk for obesity. Nutrients 8(3):128. https://doi.org/10.3390/nu8030128
Sokoła-Wysoczańska E, Wysoczański T, Wagner J, Czyż K, Bodkowski R, Lochyński P-S (2018) Polyunsaturated fatty acids and their potential therapeutic role in cardiovascular system disorders—a review. Nutrients 10(10):1561. https://doi.org/10.3390/nu10101561
Sperling P, Ternes P, Zank TK, Heinz E (2003) The evolution of desaturases. Prostaglandins Leukotrienes Essent Fatty Acids 68(2):73–95. https://doi.org/10.1016/s0952-3278(02)00258-2
Stukey JE, McDonough VM, Martin CE (1990) The OLE1 gene of Saccharomyces cerevisiae encodes the delta 9 fatty acid desaturase and can be functionally replaced by the rat stearoyl-CoA desaturase gene. J Biol Chem 265(33):20144–20149. https://doi.org/10.1016/S0021-9258(17)30481-7
Swanson D, Block R, Mousa SA (2012) Omega-3 fatty acids EPA and DHA: health benefits throughout life. Adv Nutr 3:1–7. https://doi.org/10.3945/an.111.000893
Tocher DR, Leaver MJ, Hodgson PA (1998) Recent advances in the biochemistry and molecular biology of fatty acyl desaturases. Prog Lipid Res 37:73–117. https://doi.org/10.1016/s0163-7827(98)00005-8
Venegas-Caleron M, Napier SO, JA, (2009) An alternative to fish oils: metabolic engineering of oil-seed crops to produce omega-3 long chain polyunsaturated fatty acids. Prog Lipid Res 49:108–119. https://doi.org/10.1016/j.plipres.2009.10.001
Wang H, Klein MG, Zou H, Lane W, Snell G, Levin I, Li K, Sang B-C (2015) Crystal structure of human Stearoyl-coenzyme A desaturase in complex with substrate. Nature 22:581–586. https://doi.org/10.1038/nsmb.3049
Wang L, Athinarayanan S, Jiang G, Chalasani N, Zhang M, Liu W (2015b) Fatty acid desaturase 1 gene polymorphisms control human hepatic lipid composition. Hepatology 61(1):119–128. https://doi.org/10.1002/hep.27373
Wang M, Chen H, Gu Z, Zhang H, Chen W, Chen YQ (2013) ω3 fatty acid desaturases from microorganisms: structure, function, evolution, and biotechnological use. Appl Microbiol Biotechnol 97:10255–10262. https://doi.org/10.1007/s00253-013-5336-5
Wang M, Chen H, Ailati A, Chen W, Chilton FH, Todd LW, Chen YQ (2017) Substrate specificity and membrane topologies of the iron-containing ω3 and ω6 desaturases from Mortierella alpina. Appl Microbiol Biotechnol 102(1):211–223. https://doi.org/10.1007/s00253-017-8585-x
Wang Y-D, Peng K-C, Wu J-L, Chen J-Y (2014) Transgenic expression of salmon delta-5 and delta-6 desaturase in zebrafish muscle inhibits the growth of Vibrio alginolyticus and affects fish immunomodulatory activity. Fish Shellfish Immunol 39:223–230. https://doi.org/10.1016/j.fsi.2014.04.021
Whittle EJ, Tremblay AE, Buist PH, Shanklin J (2008) Revealing the catalytic potential of an acyl-ACP desaturase: tandem selective oxidation of saturated fatty acids. Proc Natl Acad Sci 105:14738–14743. https://doi.org/10.1073/pnas.0805645105
Xie L, Innis SM (2008) Genetic variants of the FADS1 FADS2 gene cluster are associated with altered (n-6) and (n-3) essential fatty acids in plasma and erythrocyte phospholipids in women during pregnancy and in breast milk during lactation. J Nutr 138:2222–2228. https://doi.org/10.3945/jn.108.096156
Xue Z, He H, Hollerbach D, Macool DJ, Yadav NS, Zhang H, Szostek B, Zhu Q (2013) Identification and characterization of new Delta-17 fatty acid desaturases. Appl Microbiol Biotechnol 97:1973–1985. https://doi.org/10.1007/s00253-012-4068-2
Yao JK, Leonard S, Reddy RD (2000) Membrane phospholipid abnormalities in postmortem brains from schizophrenic patients. Schizophr Res 42:7–17. https://doi.org/10.1016/s0920-9964(99)00095-x
Yeagle PL (2016) Membrane Proteins. The Membranes of Cells 219–268. eBook
Yoshida K, Hashimoto M, Hori R, Adachi T, Okuyama H, Orikasa Y, Nagamine T, Shimizu S, Ueno A, Morita N (2016) Bacterial long-chain polyunsaturated fatty acids: their biosynthetic genes, functions, and practical use. Mar Drugs 14:94. https://doi.org/10.3390/md14050094
Zhang J, Liu H, Sun J, Li B, Zhu Q, Chen S, Zhang H (2012) Arabidopsis fatty acid desaturase FAD2 is required for salt tolerance during seed germination and early seedling growth. PLoS ONE 7(1):e30355. https://doi.org/10.1371/journal.pone.0030355
Zhang J-T, Zhu J-Q, Zhu Q, Liu H, Gao X-S, Zhang H-X (2009) Fatty acid desaturase-6 (Fad6) is required for salt tolerance in Arabidopsis thaliana. Biochem Biophys Res Commun 390:469–474. https://doi.org/10.1016/j.bbrc.2009.09.095
Zhang L, Chen WB, Yang SP, Zhang YB, Xu JL, Yang DJ (2020) Identification and functional characterization of a novel delta 12 fatty acid desaturase gene from Haematococcus pluvialis. J Ocean U China 19:1362–1370. https://doi.org/10.1007/s11802-020-4418-0
Zhang PY, Liu SH, Cong BL, Wu GT, Liu CL, Lin XZ (2011) A novel omega-3 fatty acid desaturase involved in acclimation processes of polar condition from Antarctic ice algae Chlamydomonas sp. ICE-l Mar Biotechnol 13:393–401. https://doi.org/10.1007/s10126-010-9309-8
Zhang Y (2009) Protein structure prediction: when is it useful? Curr Opin Struct Biol 19:145–155. https://doi.org/10.1016/j.sbi.2009.02.005
Zhang Y, Wang H, Zhang J, Hu Y, Zhang L, Wu X, Su X, Li T, Zou X, Liang B (2016) The cytochrome b5 reductase HPO-19 is required for biosynthesis of polyunsaturated fatty acids in Caenorhabditis elegans. BBA - Mol Cell Biol Lipids 1861:310–319. https://doi.org/10.1016/j.bbalip.2016.01.009
Zhou J, Chen J, Hu C, Xie Z, Li H, Wei S, Wang D, Wen C, Xu G (2016) Exploration of the serum metabolite signature in patients with rheumatoid arthritis using gas chromatography-mass spectrometry. J Pharm Biomed Anal 127:60–67. https://doi.org/10.1016/j.jpba.2016.02.004
Zhu G, Koszelak-Rosenblum M, Connelly SM, Dumont ME, Malkowski MG (2015) The crystal structure of an integral membrane fatty acid alpha-hydroxylase. J Biol Chem 290:29820–29833. https://doi.org/10.1074/jbc.M115.680124
Zhuang X-Y, Zhang Y-H, Xiao A-F, Zhang A-H, Fang B-S (2022) Key enzymes in fatty acid synthesis pathway for bioactive lipids biosynthesis. Front Nutr 9:851402. https://doi.org/10.3389/fnut.2022.851402
Funding
The authors received financial support from Yayasan Penyelidikan Antartika Sultan Mizan—2017 for this project and financial assistance and scholarship from Graduate Research Fellowship (GRF) for NFAAH.
Author information
Authors and Affiliations
Contributions
NFAAH drafted the content, analyzed the subjects, and wrote the manuscript. NMY, RNZ, MSMA, and ATCL gave critical insights on the subjects, reviewed, and improved the manuscript.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies involving human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Halim, N.F.A.A., Ali, M.S.M., Leow, A.T.C. et al. Membrane fatty acid desaturase: biosynthesis, mechanism, and architecture. Appl Microbiol Biotechnol 106, 5957–5972 (2022). https://doi.org/10.1007/s00253-022-12142-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12142-3