Abstract
Three semi-selective media, DTarTA, SPbc, and SPamt, were developed and tested to isolate Pseudomonas syringae pv. maculicola (Psm) and P. cannabina pv. alisalensis (Pca) from Raphanus sativus seeds. DTarTA contained D-tartaric acid as a carbon source and potassium tellurite, ampicillin sodium, and methyl violet as antibiotics. DTarTA suppressed growth in 19 of the 24 pathovars from the P. syringae complex, whereas Psm and Pca grew and formed gray to black colonies. SPamt contained sucrose and peptone as nutrient sources and was supplemented with bromothymol blue and the same antibiotics present in DTarTA and Psm and Pca formed yellowish to dark brown colonies on the SPamt medium. SPbc contained sucrose and peptone and was supplemented with cephalexin and boric acid as antibiotics and Psm and Pca formed semi-translucent to white colonies on the SPbc medium. SPamt and SPbc suppressed the growth of several plant-associated bacteria (except the P. syringae complex). The growth of saprophytic bacteria in seeds on the different media was compared with that on King’s B medium, using five types of commercially available Raphanus sativus seeds. The suppression rate of DTarTA was 85–99% and was lower for seeds with more saprophytic bacteria. The suppression rates of SPamt and SPbc were 90–99%. In detection tests using 10,000 seed samples mixed with Pca or Psm-contaminated seeds, it was possible to selectively isolate Psm and Pca using SPamt and SPbc, even when the colony numbers of the target bacterium constituted less than 10% of the total colonies.
Key points
• Bacterial leaf spot and blight pathogens were selectively isolated from seeds.
• DTarTA medium distinguishes these pathogens from P. syringae complex pathovars.
• SPamp and SPbc media have different selectivity for plant-associated bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial leaf spot and leaf blight are global threats to the cultivation of cruciferous vegetables. Pseudomonas syringae pv. maculicola (Psm) causes bacterial leaf spots in plants such as cabbage, cauliflower, broccoli, Chinese cabbage, turnip, radish, and Japanese radish (Peters et al. 2004; Takikawa and Takahashi 2014; Zhao et al. 2000) and was first reported by McCulloch (1911). Psm is closely resembles pathovar tomato in bacteriological characteristics and genetic similarity (Peters et al. 2004; Wiebe and Campbell 1993). Both pathogens can infect each other’s hosts when artificially inoculated (Hendson et al. 1992; Takikawa et al. 1994; Wiebe and Campbell 1993). It was reported that there are at least eight genetic lineages among Psm and pathovar tomato isolates and there is a relationship between the genetic lineages and isolation sources (Gironde and Manceau 2012). Psm also closely resembles pathovar spinaciae (Bazzi et al. 1988; Inoue and Takikawa 2021; Koike et al. 2002; Ozaki et al. 1998). Psm are classified into four groups (I–IV) according to their bacteriological characteristics and the host from which they are isolated (Takikawa and Takahashi 2014; Takikawa et al. 1994). Group I consists of the majority of worldwide isolates, including pathovar tomato. Group II consists of various Japanese radish isolates, which cause small leaf spots and internal root discoloration. Group III mainly consists of Brassica rapa isolates from Japan. Group IV consists of various Japanese radish isolates which cause large necrotic lesions (now reclassified as a different pathotype). Pseudomonas cannabina pv. alisalensis (Pca) causes bacterial leaf blight in plants such as arugula, cabbage, Chinese cabbage, radish, and bristle oat (Bull et al. 2004; Ishiyama et al. 2013; Mauzey et al. 2011; Rubio et al. 2012; Sarris et al. 2010; Takahashi et al. 2013; Wechter et al. 2010) and was first reported by Cintas et al. (2002). Some strains that were isolated and identified as Psm (group IV) have now been reclassified as Pca (Rubio et al. 2012; Takahashi et al. 2013; Takikawa and Takahashi 2014). Pca is classified into two types (A and B) based on differences in host virulence and genetic diversity (Sarris et al. 2013; Takikawa and Takahashi 2014). They were divided based on utilization tests of sorbitol, malonate, caprate, and acetate. Psm and Pca invade the stomata of leaves and plant wounds, creating small chlorotic and necrotic spots with water-soaked regions on the leaves, and cause severe blight symptoms (Peters et al. 2004; Takikawa and Takahashi 2014; Zhao et al. 2000). Infections by Psm and Pca cause black brown discoloration in the core of the stem root and bacterial spots on the surface of the stem root of Japanese radish, which is a major problem in Japan (Horinouchi et al. 2009; Inoue and Takikawa 2021; Otani 2016; Takeuchi et al. 1989). Similar to other bacteria in the P. syringae group, both Psm and Pca are considered seed-borne (Schofield et al. 2012; Takimoto 1931). Therefore, the distribution of seeds contaminated with these pathogens is thought to play an important role in the occurrence of disease.
In general, tests to infer the pathogen-free status of seeds require the isolation of live pathogenic bacteria. Thus, a selective medium with high selectivity for the target pathogenic bacterium is required. KBC medium (Mohan and Schaad 1987) is widely used for the isolation of bacteria in the P. syringae group (Asaad et al. 2017; Bull and Koike 2017; Randhawa et al. 2017; Shepherd and Block 2017). However, KBC has been developed to isolate P. syringae pv. syringae from beans, and it is difficult to distinguish between the saprophytic bacteria and target bacteria, Psm and Pca based on their colony morphology in this medium; furthermore, the suppression of saprophytic bacteria by this medium is imperfect. Thus, the development of selective media that suppress the growth of more saprophytic bacteria and allow Psm and Pca to form colonies with distinctive morphological characteristics would help in the isolation of these pathogens. KBC contains cephalexin and boric acid which act as antibacterial substances in King’s B medium (KB; King et al. 1954). By changing the composition of the antibacterial component, it may be possible to prepare a new medium that differs from KBC in terms of its selectivity to saprophytic bacteria. MSP (Mohan and Schaad 1987) and SPTPsjA (Mori et al. 2018) media have also been developed as semi-selective media to isolate bacteria in the P. syringae group. MSP uses sucrose as its main carbon source and is suitable for separating the P. syringae complex from other Pseudomonas spp. SPTPsjA enhances the selection of P. syringae pv. syringae using L-serine as the sole carbon source and contains multiple antibiotics. By combining the characteristics of these media, it may be possible to develop new selective media. In this study, three semi-selective media for Psm and Pca were developed based on the aforementioned media.
Materials and methods
Bacterial strains and culture conditions
Sixty-five strains of plant-associated bacteria were used in these experiments (Table 1). The Pseudomonas strains, including five strains of Pca, eight strains of Psm, and 23 other strains belonging to the P. syringae group, as well as an Acidovorax strain, were cultured on PPGA medium (Nishiyama 1978) at 25 °C. The Xanthomonas and Ralstonia strains were cultured on potato semisynthetic medium (PSA) (Wakimoto 1960) at 27 °C, and those in the Erwinia group and Burkholderia strains were cultured on yeast-peptone agar medium (YPA: 5 g yeast extract, 10 g peptone, 15 g agar, and 1000 mL of distilled water; pH 6.8) at 27 °C. These cultures were suspended in sterile distilled water at an optical density of 0.3 at 600 nm (~ 108 CFU/mL in each case) and appropriately diluted for subsequent examinations. Rifampicin-resistant strains KMrR-R03 (derived from MAFF (Ministry of Agriculture, Forestry and Fisheries, Japan) 106,179) and NMH-R1 (derived from MAFF 106,156) (Inoue and Takikawa 2021) were used to create pathogen-infected seeds.
D-tartaric acid, tellurite, and ampicillin medium
To the base of the SPTPsjA (KH2PO4: 1.3 g; Na2HPO4\(\cdot\) 12H2O: 1.2g; MgSO4\(\cdot\) 7H2O : 0.25 g; (NH4)2SO4 : 5 g)
medium, 5 g of D-tartaric acid was added. This was dissolved in 1 L of distilled water, and the pH was adjusted to 6.8. Fifteen grams of agar was added, and the medium was autoclaved at 115 °C for 15 min and then cooled to approximately 50 °C. To this, 100 µL each of ampicillin sodium (100 mg/mL in sterile water), potassium tellurite (250 mg/mL in sterile water), methyl violet (10 mg/mL in 70% ethanol), and cycloheximide (100 mg/mL in 70% ethanol) was added. The medium was mixed well. Approximately 15 mL of the medium was added to a 9-cm sterile Petri dish to prepare a plate medium.
Sucrose-peptone medium with ampicillin, methyl violet, and potassium tellurite
The determined composition (KH2PO4: 1 g; MgSO4 \(\cdot\) 7H2O: 0.2 g; Bacto-peptone: 5 g; sucrose: 20 g) was dissolved in 1 L of distilled water; 1 mL of bromothymol blue (BTB, dissolved in ethanol at 50 mg/mL) was added; and the pH was corrected to 6.2, 6.4, 6.6, 6.8, and 7.0. Fifteen grams of agar was added to this, and the mixture was autoclaved at 121 °C for 20 min. The autoclaved medium was cooled to approximately 50 °C. To this, 100 µL each of ampicillin sodium (100 mg/mL in sterile water), potassium tellurite (250 mg/mL in sterile water), and methyl violet (10 mg/mL in 70% ethanol), as well as 3.5 mL of nystatin (10 mg/mL in 70% ethanol), were added. The medium was mixed well, and approximately 15 mL of it was added to a 9-cm sterile Petri dish to prepare a plate medium.
Sucrose-peptone medium with boric acid and cephalexin
The basic composition of the medium (KH2PO4: 1 g; MgSO4 \(\cdot\) 7H2O: 0.2 g; Bacto-peptone: 5 g; sucrose: 20 g) was the same as that used for the sucrose-peptone medium with ampicillin, methyl violet and potassium tellurite (SPamt). The components were dissolved in 900 mL of distilled water. The pH was adjusted to 7.0 because the addition of boric acid after sterilization lowered the pH. Fifteen grams of agar was added, and the medium was autoclaved at 121 °C for 20 min and then cooled to approximately 50 °C. To this, 100 mL of boric acid (1.5 g/mL in sterile water), 8 mL of cephalexin (10 mg/mL in sterile water), and 3.5 mL of nystatin (10 mg/mL in 70% ethanol) were added. The medium was mixed well, and approximately 15 mL of it was added to a 9 cm sterile petri dish to prepare a plate medium.
Growth test of plant-associated bacteria on DTarTA, SPamt, and SPbc
Suspensions of all the bacterial strains shown in Table 1, except the Psm strain SUPP (Shizuoka University Plant Pathology) 1331, were prepared. A suspension of eight bacterial strains was added to each well in the horizontal rows starting in the left top corner of a 96-well plate. The second, fourth, and sixth rows were not used. A suspension of the bacterial strains was added to each of the eight wells in the third, fifth, and seventh row. A total of 32 strains were used per plate. The volume of each bacterial suspension was 150 μL. The pins of a copy plate (TK-CP96-1/2; Tokken, Chiba, Japan) were inserted into the wells to attach the bacterial suspensions, and the copy plate was then placed on the medium. The copy plate was removed after 10 s, and the surface of the medium was allowed to dry (with the cover open) for 5 min on a clean bench. The medium was then incubated at 25 °C. Bacterial growth was evaluated after 3, 5, and 7 days for D-tartaric acid, tellurite, and ampicillin (DTarTA) and after 3 and 5 days for SPamt and sucrose-peptone medium with boric acid and cephalexin (SPbc). The tests were performed twice.
To compare the plating efficiencies of diluted media, six strains of Psm and four strains of Pca were used. The bacterial suspensions were diluted with 1 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) buffer (pH 7.0) to approximately 103 CFU/mL. Using three plates per strain, 75 µL of each diluted bacterial suspension was spread on the plate. After incubating at 25 °C in the dark for 3 and 5 days, the number of colonies formed on the medium was counted and the average for the three plates was calculated in terms of strain, dilution, and incubation period. The bacterial growth on these media was compared to those on KB and KBC. This test was repeated three times, and differences in the number of colonies between media for each strain were evaluated using Tukey’s HSD test.
Evaluation of the media for the inhibition of saprophytic bacteria associated with Raphanus sativus seeds
For this experiment, commercially available Raphanus sativus seeds that had not been seed-treated were used. The five seed types were categorized as A (Japanese radish of unknown variety, provided by the Center for Seeds and Seedlings (NARO)), B (Hataro, Takii Seeds, Kyoto, Japan), C (Cyoukouaonaga, Takii Seeds), D (Sakuranbo, Sakata Seeds, Kanagawa, Japan), and E (Yukikomachi, Sakata Seeds). Bacteria were extracted from the seeds using the same method as that used for black rot (Export and International Affairs Bureau 2016; International Seed Testing Association 2017). The weight of 100 seeds was measured 10 times, the average weight was calculated, and 100 times the average weight of the 100 seeds was recorded as the weight of 10,000 seeds. Seed sets were placed in 500 mL or 1000 mL conical flasks. When the weight of 10,000 seeds was 100 g or less, a 2.5 × volume of wash buffer (0.85% NaCl, 0.02% Tween20) of the seed weight was added. When the weight was 100 g or more, a 2.25 × volume of buffer of the seed weight was added. The sample was shaken at 120 rpm for 2.5 h, and then 10 mL of the supernatant was collected. The seed extract was diluted using tenfold dilution steps, and 100 µL of the diluted solution from each step was spread on two plates each of DTarTA, SPamt, SPbc, KBC, and KB media. The number of colonies formed in the medium was counted seven days after inoculation. The suppression rate was determined from the ratio of the number of colonies in the experimental medium to the number of colonies in the KB medium (Mohan and Schaad 1987).
Recovery of Psm and Pca from seed samples containing inoculated seeds
The infected seeds were inoculated with KMrR-R03 and NMH-R1 (Inoue and Takikawa 2021). The seeds were infected with approximately 0–500 CFUs of the inoculated strain per seed. Seed sets were mixed with either 50 Pca-inoculated seeds or 50 Psm-inoculated seeds to a total of 10,000 seeds per set, and seed extracts were prepared from these seed sets. The seed extract was diluted in 1 mM HEPES buffer (pH 7.0). Finally, 100 μL each of the undiluted extract, a tenfold diluted solution, and a 100-fold diluted solution were applied to two plates of each medium. To measure the density of target bacteria in the liquid, 100 μL of the undiluted extract was applied to two plates of YPA medium containing 20 mg/L of rifampicin and 35 mg/L of nystatin, and the number of colonies was counted after 3 days. As a control (to observe the morphological characteristics of colonies for colony identification), 100 µL of a suspension of KMrR-R03 and NMH-R1 (diluted to approximately 103 CFU/mL) was also added to the medium. After 5–7 days for DTarTA and 3–5 days for SPamt, SPbc, and KBC, colonies were transferred from the medium surface with a sterile toothpick to YPA containing 20 mg/L rifampicin. At that time, the colonies judged to be similar to those of the reference strain based on morphological characteristics like size, color, surface appearance, edge, and elevation were noted. The colonies that grew 2–3 days after subculturing were considered to be the target strain, and the ratio of target strain colonies to all colonies was calculated. The proportion of target bacteria that could or could not be predicted as target bacteria at the time of colony transfer was also calculated. Using some of the isolated colonies, a specific PCR test was performed on Psm and Pca (Inoue and Takikawa 2021) to confirm that the bacteria were indeed Psm or Pca.
Results
Growth of Psm and Pca on DTarTA
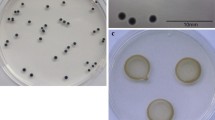
The growth of plant-associated bacteria on DTarTA was evaluated. In the P. syringae complex, Psm, Pca, and P. syringae pvs. mellea, spinaciae, and tomato grew on DTarTA (Table 1). Psm grew in all seven tested strains, and Pca grew in four of the five tested strains. Fluorescent Pseudomonas such as P. cichorii, P. viridiflava, and the P. fluorescens complex also grew vigorously (Table 1, Fig. S1a). Psm and Pca formed initially translucent and eventually black colonies on DTarTA in approximately 3–5 days (Fig. 1). In relation to the Psm strains, the ratio of the number of colonies growing on DTarTA to that on KB was 0.64 to 0.80. Regarding the Pca strains, the ratio of the number of colonies growing on DTarTA to that on KB was as low as 0.45–0.71 (Fig. 2). MAFF 730087 did not form colonies (Fig. 2).
Selectivity of DTarTA
Using the aforementioned five sets of seed samples, the growth-suppressing effects of DTarTA on seed bacteria were investigated. The suppression efficiency varied depending on the seed sample, and the number of colonies was reduced by 85–100% compared to that on the KB medium (Table 2). The colonies of several saprophytic bacteria were black in color and appeared similar to the colonies of Psm or Pca (Fig. S2). Seeds inoculated with Psm or Pca were mixed into these seed lots, and an attempt was made to isolate the inoculated bacteria. In seed samples B, C, and D (approximately 104 CFU/mL of saprophytic bacteria in the extract), 2.2–100% of the colonies grown on the medium were Psm or Pca, and it was possible to distinguish between saprophytic and inoculated bacteria based on morphological characteristics (Table 3). In seed samples A and E (approximately 105 CFU/mL of saprophytic bacteria in the extract), Psm or Pca comprised approximately 0–10% of the colonies grown on the medium. However, it was difficult to distinguish these colonies from those of the saprophytic bacteria based on morphological characteristics because there were too many colonies on the medium, and many of these colonies had similar morphologies to the Psm and Pca.
Growth of Psm and Pca on SPamt
The lower pH of SPamt suppressed not only the growth of seed-associated bacteria but also the growth of Psm and Pca (Fig. S3). Therefore, the pH of SPamt was set at 6.8–6.6. The pH of the medium changed slightly (± 0.05 or less) after sterilization at 121 °C for 20 min. Colonies were visible two days after the application of Psm and Pca to SPamt. After 3–5 days, yellowish to coffee brown colonies were formed by Psm, and brown to dark brown colonies were formed by Pca (Fig. 1). Pseudomonas cichorii, P. fuscovaginae, P. viridiflava, and bacteria in the P. syringae complex grew on SPamt, while other bacteria did not grow or grew only slightly on the media (Table 1, Fig. S1b). The ratio of the number of bacterial colonies growing on SPamt to that on KB was 0.8–1.2 (Fig. 3). No significant differences were observed in any of the strains.
Ratio of the number of colonies of Pseudomonas syringae pv. maculicola and P. cannabina pv. alisalensis on KBC, SPbc, and SPamt to that on KB. Vertical bars indicate the standard error for three independent experiments. Asterisks indicate a significant difference following Tukey’s HSD test (* P < 0.05, ** P < 0.01)
Growth of Psm and Pca on SPbc
Psm and Pca formed colonies 2–3 days after plating on SPbc, and formed translucent to milky white shiny colonies at 3–5 days (Fig. 1). Bacteria in the P. syringae group, P. fuscovaginae, P. viridiflava, and some bacteria in the P. fluorescens group also grew on this medium (Table 1, Fig. S1c). The Psm strains MAFF 302783, 302723, and 730088 formed significantly fewer colonies on SPbc than that on KB medium (Fig. 3). Also, the plating efficiencies of MAFF 301175 on SPbc were also 45.5%, 107.0%, and 37.6% of those on KB, and the number of colonies formed was 50% or less, except in one test. For other strains, the ratio of the number of bacterial colonies growing on SPbc to the number of bacterial colonies growing on KB was 0.8–1.3.
Selectivity of SPamt and SPbc
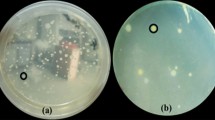
Using the five sets of seed samples, the growth-suppressing effects of the media were investigated. The number of colonies on SPamt was 0.01–0.1 of that on KB (Table 4). The colonies of several saprophytic bacteria on SPamt were dark blue with an area of blue discoloration around them (Fig. S2). They were distinguished from those of Psm and Pca (Fig. 4) based on morphological characteristics. The colonies were confirmed to be Psm or Pca by using a growth test on YPA containing 20 mg/L of rifampicin and a specific PCR. Some bacteria formed fluid colonies and spread over the surrounding colonies. The number of colonies on SPbc was also 0.01–0.1 of that o KB (Table 4). The growth inhibitory effect of this medium was the same as that of KBC, and as the culturing period increased, the number of colonies increased significantly in some samples. The colonies of several saprophytic bacteria on SPbc were flat and pale yellow (Fig. S2) and could be distinguished from those of Psm and Pca (Fig. 4) based on morphological characteristics. The colonies were confirmed to be Psm or Pca by using a growth test on YPA containing 20 mg/L of rifampicin and a specific PCR.
Effect of Raphanus sativus seed-associated bacteria on the recovery of Pseudomonas syringae pv. maculicola and P. cannabina pv. alisalensis on SPamt and SPbc media. The sample contained seeds in group B (Hataro, Takii Seeds, Kyoto, Japan) mixed with MAFF 106156- or MAFF 106179-inoculated seeds. Arrows point to a typical MAFF 106156 or 106179 colony
Seeds inoculated with Psm or Pca were mixed into these seed sets, and an attempt was made to isolate the inoculated bacteria. In seed samples B, C, and D (approximately 104 CFU/mL of saprophytic bacteria in the extract), 76.5–100% of the colonies growing on the medium were Psm or Pca, and the bacteria could be identified (Table 5). In seed samples A and E (approximately 105 CFU/mL of saprophytic bacteria in the extract), Pca accounted for approximately 10–50% of the colonies growing on the media and could be distinguished from saprophytic bacteria. Among the bacteria isolated from sample A growing on SPbc, Psm accounted for 28.6–43.8% of the colonies growing on the medium; however, the target strains in the colonies growing in the other combinations were as low as 0–15% (Table 5). Nevertheless, it was possible to identify and isolate Psm.
Discussion
To date, KBC medium has been often used to isolate bacteria in the P. syringae group on seeds (Asaad et al. 2017; Bull and Koike 2017; Randhawa et al. 2017; Shepherd and Block 2017). However, my preliminary studies suggested that the growths of some strains of Psm were significantly suppressed on KBC (Fig. 2) and that it was difficult to distinguish between target and saprophytic bacteria (Fig. S2). Therefore, it was necessary to develop new selective media that suppress the growth of saprophytic bacteria and enables the formation of colonies in which Psm and Pca are morphologically different from the saprophytic bacteria.
The availability of multiple carbon sources was investigated during the development of SPTPsjA (Mori et al. 2019), and several carbon sources were evaluated in more detail in this study. Among the carbon sources evaluated, D-tartaric acid was highly selective for Psm and Pca (Table S1). Billing (1970) reported that D-tartaric acid is rarely used by any bacteria other than pv. tomato in the P. syringae group. Psm and pv. tomato are closely related (Peters et al. 2004; Wiebe and Campbell 1993), and both used D-tartaric acid and grew on DTarTA in this study. In contrast, pvs. actinidiae and theae (belonging to the same genetic group as Psm) and pvs. lachrymans and sesami (belonging to the same genetic group as Pca) did not grow in a medium containing D-tartaric acid. D-tartaric acid is rarely used in P. cichorii (Billing 1970), and the observed differences in the use of D-tartaric acid may be related to the host range. DTarTA medium was considered to help distinguish Psm and Pca from other strains of P. syringae complex.
In the DTarTA medium, it was possible to identify Psm or Pca in a sample with a bacterial density of approximately 104 CFU/mL in the seed extract. However, identification was difficult in a sample with a bacterial density of 105 CFU/mL or more, as several colonies on DTarTA were similar to those of Psm and Pca. The 16S rDNA sequences of some of the colonies on DTarTA confirmed that these were Pseudomonas spp. (Supplementary Information). Pseudomonas spp. have also been reported to survive in rapeseed and wild cabbage seeds (Granér et al. 2003; Rybakova et al. 2017; Tyc et al. 2020). In DTarTA, it was difficult to isolate Psm and Pca from seeds containing a large amount of Pseudomonas spp.
In this study, Spamt and SPbc media were developed. Although bacteria in the P. syringae complex use sucrose for energy and produce acid, several fluorescent Pseudomonas spp. do not exhibit such metabolism (Lelliott et al. 1966). By using sucrose as the carbon source, Psm and Pca grew as rapidly as saprophytic bacteria. Peptone can be used as a nitrogen source, but the production of biosurfactants increases with the amount of peptone added (Arino et al. 1996). In SPamt, many saprophytic bacteria used peptone to raise the pH, which turned the medium and colonies blue in color. However, Psm and Pca used sucrose to produce acid, which suppressed the blue coloring. This difference in color contributed to the distinctiveness of the colonies. In SPamt, 0.5% peptone promoted blue coloring by saprophytic bacteria. In contrast, in SPbc, Psm and Pca formed milky white glossy colonies, which appeared different from the colonies of saprophytic bacteria. Therefore, BTB was not added, and the peptone content was reduced to 0.2% to suppress the growth of saprophytic bacteria.
The SPbc medium suppressed the growth of some Psm strains (Fig. 2), which was similar to that observed in the KBC medium. This suppression was presumed to be due to the antibiotic composition. The strains were isolated from Brassica rapa (Chinese cabbage and turnip) and Raphanus (Japanese radish). Psm are classified into groups I, II, and III. Group III contained many bacteria isolated from B. rapa and group II contained several bacteria isolated from Raphanus (Matsuda and Takikawa 2003; Takikawa and Takahashi 2014). Although there was no suppression of Psm and Pca growth in SPamt compared to that in KB, fluid colonies were formed in some seed extracts. The fluid colonies grew fast, making it difficult to identify and isolate Psm and Pca. No fluid colonies were formed on SPbc. Thus, both media have advantages and disadvantages in isolating these target bacteria from seeds. The combination of these media can be expected to compensate for the shortcomings of both and allow efficient isolation of these bacterial strains from seeds.
It is difficult to obtain a medium with complete selectivity for the target bacterium. In such cases, bacterial isolation would be performed using multiple semi-selective media. The seed test for black rot in cruciferous vegetables uses FS and mCS20ABN media (International Seed Testing Association 2017). Neither medium is completely selective for black rot pathogens; however, the growth characteristics of seed-borne saprophytic bacteria are different because of the various growth-inhibiting activities of these media against other bacteria. Thus, these two media are sufficient to selectively isolate black rot pathogens and allow their easy identification. For the isolation of Psm and Pca from Raphanus seeds, the KBTA medium was prepared using a KB medium containing the same antibiotics as those found in DTarTA (Inoue 2022). The KBTA medium was used in combination with KBC (National Agriculture and Food Research Organization 2021). KBTA promoted the growth of the Japanese radish isolates of Psm (Fig. S4); however, it showed poor growth of Pca and allowed the formation of fluid colonies. The SPamt and SPbc media developed in the current study have selectivity equal to or higher than that of KBC. Both media enhance the visibility of Psm and Pca. Based on these results, it is believed that these media can be used for the efficient identification and isolation of these bacterial strains.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
Arino S, Marchal R, Vandecasteele J-P (1996) Identification and production of a rhamnolipidic biosurfactant by a Pseudomonas species. Appl Microbiol Biotechnol 45:162–168
Asaad S, Sands DC, Mohan SK (2017) Detection of Pseudomonas syringae pv. syringae in wheat seeds. In: Fatmi M, Walcott RR, Schaad NW (eds) Detection of plant-pathogenic bacteria in seed and other planting material. APS Press, St. Paul, pp 21–26
Bazzi C, Gozzi R, Stead D, Sellwood J (1988) A bacterial leaf spot of spinach (Spinacia oleracea L.) caused by a non-fluorescent Pseudomonas syringae van Hall (pathovar). Phytopathol Mediterr 27:103–107
Billing E (1970) Pseudomonas viridiflava (Burkholder, 1930; Clara 1934). J Appl Bacteriol 33:492–500
Bull CT, Koike ST (2017) Detection of Pseudomonas pathogens from crucifer seeds. In: Fatmi M, Walcott RR, Schaad NW (eds) Detection of plant-pathogenic bacteria in seed and other planting material. APS Press, St. Paul, pp 165–172
Bull CT, Goldman P, Koike ST (2004) Bacterial blight on arugula, a new disease caused by Pseudomonas syringae pv alisalensis in California. Plant Dis 88:1384. https://doi.org/10.1094/pdis.2004.88.12.1384a
Bull CT, Manceau C, Lydon J, Kong H, Vinatzer BA, Fischer-Le Saux M (2010) Pseudomonas cannabina pv. cannabina pv. nov., and Pseudomonas cannabina pv. alisalensis (Cintas Koike and Bull, 2000) comb. nov., are members of the emended species Pseudomonas cannabina (ex Šutič & Dowson 1959) Gardan, Shafik, Belouin, Brosch, Grimont & Grimont 1999. Syst Appl Microbiol 33:105–115. https://doi.org/10.1016/j.syapm.2010.02.001
Cintas NA, Koike ST, Bull CT (2002) A new pathovar, Pseudomonas syringae pv. alisalensis pv. nov., proposed for the causal agent of bacterial blight of broccoli and broccoli raab. Plant Dis 86:992–998. https://doi.org/10.1094/pdis.2002.86.9.992
Export and International Affairs Bureau (2016) Manual of detection method of black rot pathogen from Japanese radish seeds (in Japanese). https://www.maff.go.jp/j/kanbo/tizai/brand/b_syokubut/attach/pdf/index-43.pdf. Accessed 10 February 2022
Gironde S, Manceau C (2012) Housekeeping gene sequencing and multilocus variable-number tandem-repeat analysis to identify subpopulations within Pseudomonas syringae pv. maculicola and Pseudomonas syringae pv. tomato that correlate with host specificity. Appl Environ Microbiol 78:3266–3279. https://doi.org/10.1128/aem.06655-11
Granér G, Persson P, Meijer J, Alström S (2003) A study on microbial diversity in different cultivars of Brassica napus in relation to its wilt pathogen, Verticillium longisporum. FEMS Microbiol Lett 224:269–276
Hendson M, Hildebrand DC, Schroth MN (1992) Relatedness of Pseudomonas syringae pv. tomato, Pseudomonas syringae pv. maculicola and Pseudomonas syringae pv. antirrhini. J Appl Bacteriol 73:455–464. https://doi.org/10.1111/j.1365-2672.1992.tb05005.x
Horinouchi H, Watanabe H, Shirakawa T, Hasegawa J, Mamiya T, Kuwabara K (2009) Occurrence and control of root browning symptom of Japanese radish at Gifu highland region (in Japanese). Ann Rept Kansai Plant Prot 51:45–47
Inoue Y, Takikawa Y (2021) Primers for specific detection and identification of Pseudomonas syringae pv. maculicola and P. cannabina pv. alisalensis. Appl Microbiol Biotechnol 105:1575–1584
Inoue Y (2022) Development of medium for detecting bacterial leaf spot and bacterial leaf blight pathogen (Abstract in Japanese). Proc Kanto Tosan Plant Prot Soc 69 (in press)
International Seed Testing Association (2017) International rules for seed testing, Edition 2017, Annexe to Chapter 7: Seed Health Testing Methods, pp. 7–019a: Detection of Xanthomonas campestris pv. campestris on Brassica spp
Ishiyama Y, Yamagishi N, Ogiso H, Fujinaga M, Takahashi F, Takikawa Y (2013) Bacterial brown spot on Avena storigosa Schereb. caused by Pseudomonas syringae pv. alisalensis. J Gen Plant Pathol 79:155–157. https://doi.org/10.1094/pdis.2004.88.12.1384a
King EO, Ward MK, Raney DE (1954) Two simple media for the demonstration of pyocyanin and fluorescin. J Lab Clin Med 44:301–307
Koike ST, Azad HR, Cooksey DC (2002) First Report of Bacterial Leaf Spot of Spinach Caused by a Pseudomonas syringae pathovar in California. Plant Dis 86:921. https://doi.org/10.1094/pdis.2002.86.8.921a
Lelliott RA, Billing E, Hayward AC (1966) A determinative scheme for the fluorescent plant pathogenic pseudomonads. J Appl Bacteriol 29:470–489
Matsuda D, Takikawa Y (2003) Classification of maculicola/tomato complex in Pseudomonas syringae (abstract in Japanese). Jpn J Phytopathol 69:302–303
Mauzey SJ, Koike ST, Bull CT (2011) First report of bacterial blight of cabbage (Brassica oleracea var. capitata) caused by Pseudomonas cannabina pv alisalensis in California. Plant Dis 95:71. https://doi.org/10.1094/PDIS-09-10-0642
McCulloch L (1911) A spot disease of cauliflower. Bulletin, Bureau of Plant Industry, United States Department of Agriculture 225: 1–15
Mohan SK, Schaad NW (1987) An improved agar plating assay for detecting Pseudomonas syringae pv. syringae and P. s. pv. phaseolicola in contaminated been seed. Phytopathology 77:1390–1395
Mori M, Sogou K, Inoue Y (2019) Development of a selective medium and antisera for Pseudomonas syringae pv. syringae from seeds of barley and wheat. J Gen Plant Pathol 85:211–220
National Agriculture and Food Research Organization, “Method for detection of bacterial leaf spot and bacterial leaf blight pathogens from radish seeds”, National Agriculture and Food Research Organization. https://www.naro.go.jp/project/results/4th_laboratory/carc/2020/20_041.html. Accessed 11 November 2021
Nishiyama K (1978) Shokubutsu byogen saikin kan-i doteiho no shian (in Japanese). Plant Prot 32:283–288
Otani Y (2016) Notes on the development of root rot and blackening symptoms on Japanese radish infected with Pseudomonas syringae pv. maculicola (in Japanese with English summary). Ann Rept Kansai Plant Prot 58:23–26. https://doi.org/10.4165/kapps.58.23
Ozaki K, Kimura T, Matsumoto K (1998) Pseudomonas syringae pv. spinaciae pv. nov., the causal agent of bacterial leaf spot of spinach in Japan. Jpn J Phytopathol 64:264–269
Peters BJ, Ash GJ, Cother EJ, Hailstones DL, Noble DH, Urwin NAR (2004) Pseudomonas syringae pv. maculicola in Australia: pathogenic, phenotypic and genetic diversity. Plant Pathol 53:73–79. https://doi.org/10.1111/j.1365-3059.2004.00946.x
Randhawa P, Pradhanang P, Schaad NW (2017) Detection of Pseudomonas syringae pv. tomato in tomato seeds. In: Fatmi M, Walcott RR, Schaad NW (eds) Detection of plant-pathogenic bacteria in seed and other planting material. APS Press, St. Paul, pp 119–124
Rubio I, Hiddink G, Asma M, Bull CT (2012) First report of crucifer pathogen Pseudomonas cannabina pv alisalensis causing bacterial blight on radish (Raphanus sativus) in Germany. Plant Dis 96:804. https://doi.org/10.1094/PDIS-01-12-0043-PDN
Rybakova D, Mancinelli R, Wikiström M, Birch-Jensen A-S, Postma J, Ehlers R-U, Goertz S, Berg G (2017) The structure of the Brassica napus seed microbiome is cultivar-dependent and affects the interactions of symbionts and pathogens. Microbiome 5:104. https://doi.org/10.1186/s40168-017-0310-6
Sarris PF, Karri IV, Goumas DE (2010) First report of Pseudomonas syringae pv. alisalensis causing bacterial blight of arugula (Eruca vesicaria subsp sativa) in Greece. New Dis Rep 22:22. https://doi.org/10.5197/j.2044-0588.2010.022.022
Sarris PF, Trantas EA, Baltrus DA, Bull CT, Wechter WP, Yan S, Ververidis F, Almeida NF, Jones CD, Dangl JL, Panopoulos NJ, Vinatzer BA, Goumas DE (2013) Comparative genomics of multiple strains of Pseudomonas cannabina pv. alisalensis a potential model pathogen of both monocots and dicots. PLOS One 8:e59366. https://doi.org/10.1371/journal.pone.0059366
Schofield DA, Bull CT, Rubio I, Wechter WP, Westwater C, Molineux IJ (2012) Development of an engineered bioluminescent reporter phage for detection of bacterial blight of crucifers. Appl Environ Microbiol 78:3592–3598. https://doi.org/10.1128/AEM.00252-12
Shepherd LM, Vidaver AK (2017) Detection of Pseudomonas savastanoi pv. glycinea in soybean seeds. In: Fatmi M, Walcott RR, Schaad NW (eds) Detection of plant-pathogenic bacteria in seed and other planting material. APS Press, St. Paul, pp 85–88
Takahashi F, Ogiso H, Fujinaga M, Ishiyama Y, Inoue Y, Shirakawa T, Takikawa Y (2013) First report of bacterial blight of crucifers caused by Pseudomonas cannabina pv. alisalensis in Japan. J Gen Plant Pathol 79:260–269. https://doi.org/10.1007/s10327-013-0458-2
Takeuchi K, Tsuchiya K, Kagawa H, Kase M (1989) Occurrence of root browning symptom on Japanease radish caused by Pseudomonas syringae pv. maculicola (in Japanese). Proc Kanto Tosan Plant Prot Soc 36:60–62
Takikawa Y, Nishiyama N, Ohba K, Tsuyumu S, Goto M (1994) Synonymy of Pseudomonas syringae pv. maculicola and Pseudomonas syringae pv. tomato. In: LeMattre M, Freigoun S, Rudolph K, Swings JG (Eds.) Plant pathogenic bacteria. Proc. 8th Int. Conf., Versailles, pp. 199–204
Takikawa Y, Takahashi F (2014) Bacterial leaf spot and blight of crucifer plants (Brassicaceae) caused by Pseudomonas syringae pv. maculicola and P. cannabina pv. alisalensis. J Gen Plant Pathol 80:466–474. https://doi.org/10.1007/s10327-014-0540-4
Takimoto S (1931) Bacterial black spot of cruciferous plants II (in Japanese with English summary). Bult Sci Fak Terkult Kyushu Imp Univ 4:545–559
Tyc O, Putra R, Gols R, Harvey JA, Garbeva P (2020) The ecological role of bacterial seed endophytes associated with wild cabbage in the United Kingdom. Microbiol Open 9:e00954. https://doi.org/10.1002/mbo3.954
Wakimoto S (1960) Classification of strains of Xanthomonas oryzae on the basis of their susceptibility against bacteriophages. Jpn J Phytopathol 25:193–198
Wechter WP, Keinath AP, Farnham MW, Smith JP (2010) First report of bacterial leaf blight on broccoli and cabbage caused by Pseudomonas syringae pv alisalensis in South Carolina. Plant Dis 94:132. https://doi.org/10.1094/pdis-94-1-0132c
Wiebe WL, Campbell RN (1993) Characterization of Pseudomonas syringae pv. maculicola and comparison with P. s. tomato. Plant Dis 77:414–419
Zhao Y, Damicone JP, Demezas DH, Rangaswamy V, Bender CL (2000) Bacterial leaf spot of leafy crucifers in Oklahoma caused by Pseudomonas syringae pv. maculicola. Plant Dis 84:1015–1020. https://doi.org/10.1094/PDIS.2000.84.9.1015
Acknowledgements
I thank Dr. Nobutaka Someya (Institute for Plant Protection, NARO) for their technical assistance. I also thank Dr. Yuichi Takikawa (Graduate School of Science and Technology, Shizuoka University, Japan) for distributing the strains and Dr. Kohei Osaki, Dr. Masatoshi Sato (Center for Seeds and Seedlings, NARO), and Mr. Mitsuru Igarashi (Sakata Seeds Corporation, Japan) for providing the radish seeds.
Author information
Authors and Affiliations
Contributions
YI conceived and designed the research, conducted the experiments, analyzed data, and wrote the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The author declares no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Inoue, Y. Three semi-selective media for Pseudomonas syringae pv. maculicola and P. cannabina pv. alisalensis. Appl Microbiol Biotechnol 106, 5741–5755 (2022). https://doi.org/10.1007/s00253-022-12092-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12092-w