Abstract
Bioconversion using microorganisms and their enzymes is an important tool in many industrial fields. The discovery of useful new microbial enzymes contributes to the development of industries utilizing bioprocesses. Streptomyces sp. EAS-AB2608, isolated from a soil sample collected in Japan, can convert the tetrahydrobenzotriazole CPD-1 (a selective positive allosteric modulator of metabotropic glutamate receptor 5) to its hydroxylated form at the C4-(R) position. The current study was performed to identify the genes encoding the enzymes involved in CPD-1 bioconversion and to verify their function. To identify gene products responsible for the conversion of CPD-1, we used RNA sequencing to analyze EAS-AB2608; from its 8333 coding sequences, we selected two genes, one encoding cytochrome P450 (easab2608_00800) and the other encoding ferredoxin (easab2608_00799), as encoding desirable gene products involved in the bioconversion of CPD-1. The validity of this selection was tested by using a heterologous expression approach. A bioconversion assay using genetically engineered Streptomyces avermitilis SUKA24 ∆saverm3882 ∆saverm7246 co-expressing the two selected genes (strain ES_SUKA_63) confirmed that these gene products had hydroxylation activity with respect to CPD-1, indicating that they are responsible for the conversion of CPD-1. Strain ES_SUKA_63 also showed oxidative activity toward other compounds and therefore might be useful not only for bioconversion of CPD-1 but also as a tool for synthesis of drug metabolites and in optimization studies of various pharmaceutical lead compounds. We expect that this approach will be useful for bridging the gap between the latest enzyme optimization technologies and conventional enzyme screening using microorganisms.
Key points
• Genes easab2608_00800 (cyp) and easab2608_00799 (fdx) were selected by RNA-Seq.
• Selection validity was evaluated by an engineered S. avermitilis expression system.
• Strain ES_SUKA_63 showed oxidative activity toward CPD-1 and other compounds.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bioconversion using microorganisms and their enzymes is an important tool in many fields such as the food industry (Raveendran et al. 2018; Zhang et al. 2017), the chemical industry (Douka et al. 2018; Norjannah et al. 2016), and the pharmaceutical industry (Adams et al. 2019; Shaw et al. 2003). One of the benefits of enzymatic bioconversion is that it allows various types of site-selective functionalization at non-activated sites of the target compound to be achieved under mild conditions.
The opportunities for industrial applications of this technology are increasing (Chapman et al. 2018; Choi et al. 2015). In the pharmaceutical industry, for example, enzymatic bioconversion is used in a wide range of applications at all stages of the drug discovery process, such as in the preparation of drug metabolites, the optimization of lead compounds, and the production of intermediates used in the synthesis of the active pharmaceutical ingredient (Lam 2009; Rosenthal and Lütz 2018).
Bioconversion is a representative example of so-called green chemistry and is important from the viewpoint of sustainable development (Sheldon and Woodley 2018). The development of associated technologies, such as next-generation sequencers and gene synthesis technologies, is contributing significantly to innovations in the technology of bioconversion (Park and Kim 2016). Therefore, we believe that the traditional chemical tool of bioconversion will evolve further with the development of various associated technologies.
Genetic information on enzymes is accumulating day by day, and many enzymes can now be generated by conventional heterologous expression systems and be used immediately as needed. However, the discovery of new and useful enzymes requires many resources, and the development of effective approaches to finding and optimizing new enzymes is urgently needed. Attempts to discover new functional enzymes directly from metagenomic samples such as from environmental microbiomes have been reported (Colin et al. 2015; Leis et al. 2015). However, many of the industrial enzymes used today were derived from microorganisms isolated from the environment. Those microorganisms were selected by whole-cell screening, a method that is still very effective for finding useful enzymes (Liu and Kokare 2017). To identify the target enzyme from the microorganism, the genome walking method is often used (Yang et al. 2015). However, if the gene is not identified, it is necessary to use shotgun cloning or a similar technique to comprehensively evaluate the genome for the gene(s) that encode the enzyme catalyzing the desirable function. This approach requires many resources, depending on the types of enzymes to be identified. An approach that could more efficiently identify genes encoding useful enzymes possessed by these microorganisms would help accelerate the industrial use of microbial enzymes.
Actinobacteria, a group of gram-positive bacteria, produce a variety of important secondary metabolites that are useful in drug discovery (Barka et al. 2016), and many drugs and lead compounds have been extracted from actinobacterial cultures, including streptomycin (Waksman et al. 1946), avermectin (Burg et al. 1979), and FK506 (Kino et al. 1987). Actinobacteria also produce enzymes that are useful for bioconversion (Spasic et al. 2018). For example, pravastatin was converted from compactin produced by Penicillium citrinum SANK 11480 by using Streptomyces carbophilus SANK 62585 (Arai et al. 1990). The enzyme involved in the bioconversion of compactin is cytochrome P450 (CYP) P-450sca-2, which belongs to the oxidoreductase group of enzymes. Actinobacteria possess more than 20 CYP genes, which is high among eubacteria (Kelly et al. 2005). The oxidative reaction activities of bacterial CYP in general are supported by the redox partner enzymes ferredoxin (Fdx) and ferredoxin reductase (Fpr), which mediate the oxidative reaction by transferring electrons to CYP (Kelly and Kelly 2013).
In this paper, we demonstrate our approach to identifying the CYP that catalyzes a target reaction by focusing on the bioconversion of CPD-1. CPD-1, a tetrahydrobenzotriazole, is a selective positive allosteric modulator of metabotropic glutamate receptor 5 (mGluR5) with good in vitro potency (Ellard et al. 2015; Kumar et al. 2015). mGluR5 is a target for neuroprotective drugs used to treat Alzheimer’s disease and support cognitive function. Streptomyces sp. EAS-AB2608 (NBRC 114648), which was isolated from a soil sample in Japan, converts CPD-1 to CPD-2, its C4-(R)-hydroxylated form (Fig. 1). CPD-2 is a useful compound for understanding the structure–activity relationship of CPD-1 (Okubo et al. 2018). In bacterial cells, the expression levels of metabolic enzymes such as CYP involved in xenobiotic degradation are expected to change in the presence of the substrate. This hypothesis is based on the reported mechanism of induction of human CYP (Tompkins and Wallace 2007). RNA sequencing (RNA-Seq) analysis of this strain was performed to confirm the expression level of each gene in the presence of CPD-1. From the results of RNA-Seq analysis, a gene encoding CYP (EASAB2608_00800) and a gene encoding ferredoxin (EASAB2608_00799) were selected from among the 8333 coding sequences (CDS) of Streptomyces sp. EAS-AB2608 as desired gene products are involved in the bioconversion of CPD-1 to CPD-2. We then constructed Streptomyces host strains in which these enzyme genes were overexpressed. The strain co-expressing the two genes that were induced in the presence of the substrate had hydroxylation activity with respect to CPD-1. This strain also showed oxidative activity toward several compounds other than CPD-1.
Materials and methods
Whole-genome sequencing
A glycerol stock solution of Streptomyces sp. EAS-AB2608 (NBRC 114648) was prepared, and 100 μL of the stock solution was inoculated into a test tube containing 10 mL of tryptic soy broth medium (TSB; BD Biosciences, Sparks, MD, USA). The culture was then shaken for 18 h at 28 °C on a reciprocating shaker. After full growth, mycelia were harvested by centrifugation, sedimented mycelia were digested with lysozyme, and genomic DNA (gDNA) was released by adding sodium dodecyl sulfate and proteinase K (Fujifilm Wako, Osaka, Japan). This preparation scheme was based on that of Komatsu et al. (2013). The gDNA of Streptomyces sp. EAS-AB2608 was sequenced on a Miseq (Illumina, San Diego, CA, USA) and PacBio RS II sequencer (Pacific Biosciences, Menlo Park, CA, USA).
RNA-Seq analysis
One hundred microliters of Streptomyces sp. EAS-AB2608 stock solution was inoculated into a test tube containing 10 mL of TSB. The culture was shaken at 28 °C for 3 days on a reciprocating shaker. After full growth, 100 μL of culture was inoculated into test tubes containing 10 mL of fresh TSB. These second cultures were conducted at 28 °C on a reciprocating shaker. After 18 h, CPD-1 dissolved in DMSO (dimethyl sulfoxide) was added to the culture to 100 mg/L; as a negative control, the same volume of DMSO alone was added (each was prepared in duplicate). These cultures were shaken at 28 °C for 3 h on a reciprocating shaker. Total RNA (tRNA) was extracted from each culture by using an RNeasy Protect Bacteria Mini Kit (Qiagen, Hilden, Germany). After the tRNA fraction had been obtained, messenger RNA (mRNA) was enriched from the tRNA by using a MICROBExpress Bacterial mRNA Enrichment Kit (Thermo Fisher Scientific, Vilnius, Lithuania). This preparation method was based on that of Oliver et al. (2009). Complementary DNA (cDNA) libraries were constructed by using a TruSeq RNA Library Prep Kit v2 (Illumina, San Diego, CA, USA). The quality of the cDNA libraries was checked by using D1000 ScreenTape (4200 TapeStation; Agilent Technologies, Santa Clara, CA, USA). Each cDNA library concentration was confirmed by using a Kapa Library Quantification Kit (Kapa Biosystems, Cape Town, South Africa). Each cDNA library was diluted to 4 nM and mixed in equal volumes. Finally, the mixed sample was diluted to 1.5 pM and used for NextSeq500 analysis (Illumina). After sequencing, the data of each sequence read was stored in FASTQ format. These FASTQ files were used for RNA-Seq analysis. To quantify gene expression levels of Streptomyces sp. EAS-AB2608 under each experimental condition, the sequence reads were aligned to a reference genome sequence generated from a GenBank file (AP024135). Transcripts per kilobase million (TPM) was used to normalize the read count mapped to each gene. This analysis was conducted by Strand NGS v3.1 (Strand Life Sciences, Bangalore, India).
Heterologous expression
The SUKA (Special Use of Kitasato Actinobacteria) series is a series of large-deletion mutants of Streptomyces avermitilis MA-4680 (ATCC 31267, NRRL 8165, NCBIM 12804, and JCM 5070). S. avermitilis SUKA24 was isolated from S. avermitilis SUKA22 (Ikeda et al. 2014) by elimination of linear plasmid SAP1 (Ikeda et al. 2003). Furthermore, two genes encoding relatively active CYPs (SAVERM_3882 for CYP154C2; SAVERM_7426 for CYP102B2) were deleted from strain SUKA24 by site-specific recombination using Cre-loxP, and its deletion variant was designated strain SUKA24 ∆saverm3882 ∆saverm7426 (Kim et al. 2018b; Komatsu et al. 2010). S. avermitilis SUKA24 ∆saverm3882 ∆saverm7426 was used as the heterologous expression host for the selected enzymes, EASAB2608_00800 (CYP) and EASAB2608_00799 (Fdx). In this study, we used pKU565bla-tsr::Psav2794-fld-fpr-ter (Fig. 2; Kim et al. 2018b) into which codon-optimized genes encoding flavodoxin (Fld, WP_001018618) and ferredoxin reductase (Fpr, WP_000796332) were arranged as an operon (Quaderer et al. 2006). The function of Fld is similar to that of Fdx (Jenkins and Waterman 1994). The selected CYP and Fdx genes were amplified by PCR (polymerase chain reaction) using the gDNA of strain EAS-AB2608 as a template and inserted between fld and fpr (XbaI/MfeI site) on pKU565bla-tsr::Psav2794-fld-fpr-ter. The PCR primer pairs with XbaI/MfeI site were AB2608_800_1F (5′-GCtctagaTAGGTGCCTGGGGCATCTAATGAAGATCGG-3′) and AB2608_800_1R (5′-CTCGAGcaattgTTACCAGGTCACAGGGAGTTCCAGC-3′) for the CYP gene only and AB2608_800_1F and AB2608_799_1R (5′-CTCGAGcaattgTCAGCCGGTGCTGTCCCGCA-3′) for both CYP and Fdx genes. These recombinant plasmids were introduced into the heterologous expression host S. avermitilis SUKA24 ∆saverm3882 ∆saverm7426 by Escherichia coli/Streptomyces conjugation described previously (Table 1, Kim et al. 2018b).
Bioconversion
One hundred microliters of Streptomyces sp. EAS-AB2608 stock solution was inoculated into Erlenmeyer flasks containing 10 mL of SY-32 medium (1% [w/v] d-glucose, 1% [w/v] soluble starch, 0.5% [w/v] soy peptone [Bacto Soytone, BD Biosciences], 0.5% [w/v] yeast extract [Oriental Yeast Co., Ltd, Tokyo, Japan], 0.2% [w/v] ammonium sulfate, 0.2% [w/v] NaCl, and 2.3% [w/v] N-Tris(hydroxymethyl)methyl-2-aminoethanesulfonic acid [Dojindo Laboratories, Kumamoto, Japan], pH 8.0). This culture was shaken at 28 °C for 3 days on a rotary shaker. After cultivation, this pre-culture was inoculated into Erlenmeyer flasks containing fresh SY-32 medium (inoculation volume was 1% [v/v] based on medium volume). This culture was shaken at 28 °C for 2 days on a rotary shaker. After cultivation, the mycelia were harvested by centrifugation, and sedimented mycelia were washed with 22 mM phosphate buffer (pH 7.0). Washed mycelia were placed in a reaction buffer (5 mM d-glucose-6-phosphate, 0.5 mM β-NADPH, 5 mM MgCl2, and 1 unit/mL d-glucose-6-phosphate dehydrogenase [Oriental Yeast Co., Ltd] dissolved in 66 mM phosphate buffer [pH 7.0]). The mycelial concentration of each strain was made uniform by using the turbidity method. Each of the six substrates tested was dissolved in DMSO and added to the reaction buffer to a final concentration of 0.1 mM. The bioconversion was conducted at 28 °C for 24 h on a rotary shaker.
Screening
Bioconversion screening using the above procedure was performed on six substrates. In addition to CPD-1, screening was performed on indomethacin (nonsteroidal anti-inflammatory drug), rosiglitazone (antidiabetic drug), lapachol (natural quinone compound derived from, e.g., Bignoniaceae), carbamazepine (anticonvulsant or anti-epileptic drug), and genistein (isoflavone derived from, e.g., Leguminosae) (Fig. 1), which have each been reported as substrates of human CYPs or microbial enzymes (Baldwin et al. 1999; Roh et al. 2009; Silva et al. 2016; Venkatakrishnan et al. 2001). Therefore, we considered these to be suitable compounds for confirming the activity and substrate specificity of the two selected gene products. After bioconversion, each reaction solution was mixed with an equal volume of acetonitrile, and the supernatant was collected by centrifugation (over 14,000×g for 2 min, at room temperature). The supernatant was evaporated with a centrifugal thickener (DD-4X; Genevac, Warminster, PA, USA), and the residues were dissolved in a quantity of DMSO that was 1/10 the volume of the supernatant. Each sample (10 μL) was analyzed by liquid chromatography photodiode array molecular spectrometry (LC-PDA-MS; detailed information is given below). The estimated molecular weights of substrate-derived peaks were determined by ESI-MS (electrospray ionization mass spectrometry). The conversion ratio of each peak was calculated by using the area of the UV (ultraviolet) absorption peak (at 254 nm).
Isolation of CPD-2
After completion of the bioconversion reaction, 100 mL of reaction solution was extracted twice with an equal volume of ethyl acetate, and the combined organic layer was concentrated under reduced pressure to give a residue. CPD-2 was purified from the residue by reverse-phase HPLC (high-performance liquid chromatography) with gradient elution (Unison US-C18 20 mm ⌀ × 250 mm [Imtakt, Kyoto, Japan], 20 to 80% [v/v] acetonitrile in water at a flow rate of 5 mL/min over 130 min). After fractionation, the fractions containing CPD-2 were combined, acetonitrile was removed by evaporation, and the sample was diluted with water to 25 mL. Thereafter, CPD-2 was extracted twice with an equal volume of ethyl acetate. The combined organic layer was dried over anhydrous Na2SO4, followed by concentrating under reduced pressure to give CPD-2. Elucidation of the CPD-2 structure was conducted by LC-PDA-MS and 1H and 13C NMR (nuclear magnetic resonance).
LC-PDA-MS analysis
LC-PDA-MS analyses were performed by using an HPLC system (1200 Series, Agilent Technologies) equipped with a photodiode array detector (G1315D, Agilent Technologies) coupled with a quadrupole LC/MS system (6130, Agilent Technologies). Nitrogen was used as the nebulizer gas for MS. The column oven temperature was maintained at 28 °C. Each sample was analyzed by reverse-phase LC with gradient elution (Unison UK-C18 4.6 mm ⌀ × 50 mm [Imtakt], 1 to 99% [v/v] acetonitrile in aqueous 0.1% [v/v] formic acid added at a flow rate of 2 mL/min over 6 min). The API-ESI-MS (atmospheric pressure ionization/electrospray ionization mass spectrometry) spray chamber was set at positive and negative capillary voltages of 4 kV.
NMR analysis
NMR spectra were acquired at 30 °C by using a Bruker Advance spectrometer (Bruker, Billerica, MA, USA) equipped with a DCH (dual carbon/proton) cryoprobe operating at 600.13 MHz for 1H and at 150.90 MHz for 13C. Samples were dissolved in CD3OD. Chemical shifts were referenced to internal standard peaks at δH 3.30 for CHD2OD and δC 49.0 for CD3OD, respectively. The following NMR spectra were recorded: 1H NMR, chemical shift-correlated spectroscopy (COSY); nuclear Overhauser effect spectroscopy (NOESY); 13C NMR, heteronuclear single quantum coherence (HSQC); and heteronuclear multiple-bond correlation (HMBC) spectrum. The NOESY spectrum was obtained with a mixing time of 700 ms. The HMBC experiment was performed to measure the coupling signals at 8 Hz.
Results
Selection of gene products by RNA-Seq analysis
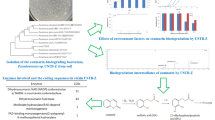
The genome data indicated that Streptomyces sp. EAS-AB2608 possesses 45 putative CYP genes. These CYP genes and their expression levels are shown in Fig. 3a. The expression level of easab2608_00800 (cyp) increased approximately 8-fold when CPD-1 (substrate) was present relative to that with the DMSO control (control TPM = 199.5 ± 4.6; substrate TPM = 1589.3 ± 461.5). The expression level of easab2608_00799 (fdx) was also increased approximately 8-fold under the same conditions (control TPM = 196.7 ± 1.2; substrate TPM = 1665.9 ± 439.3; Fig. 3b). We found that these genes were each induced to a similar degree by adding CPD-1. In contrast, no substrate-induced increase in fpr expression level was found in this analysis (Fig. 3c). Some of the other genes also showed slight positive fold changes under conditions of substrate addition. Considering the relative expression level of each gene, we advanced genes whose expression level showed a positive fold change and TPM value exceeding 500 to the verification step by using the heterologous expression approach. Specifically, we selected EASAB2608_00800 and EASAB2608_00799 as candidates for enzymes involved in the bioconversion of CPD-1. The intergenic distance and expression levels indicated that these two enzyme genes are co-regulated by the same operon. Information on each of the genes shown in Fig. 3 and their expression levels are listed in the supplementary material (Supplemental Table S1).
Expression levels of enzyme genes with and without added CPD-1 (substrate). (a) Expression levels of cytochrome P450 genes. (b) Expression levels of ferredoxin genes. (c) Expression levels of ferredoxin reductase genes. Bars denote the mean value obtained from duplicate samples. Error bars indicate standard deviation. Control TPM (white bars), expression level in sample group treated with DMSO (dimethyl sulfoxide), normalized by transcripts per million. Substrate TPM (red bars), expression level in sample group treated with CPD-1, normalized by transcripts per million. CYP, cytochrome P450. Fdx, ferredoxin. Fpr, ferredoxin reductase. TPM, transcripts per kilobase million
Identification of responsible gene products
Strains EAS-AB2608, ES_SUKA_62, ES_SUKA_63, and ES_SUKA_70 (Table 1) were used for bioconversion screening. The results of bioconversion of CPD-1 after 24-h culture with each strain are shown in Fig. 4. CPD-2 was not detected in the sample from the negative control strain (ES_SUKA_70 without cyp gene), and therefore we consider that the host (SUKA24 ∆saverm3882 ∆saverm7426) does not have hydroxylation activity with respect to CPD-1. Although strain ES_SUKA_62, which expressed only the CYP, did convert CPD-1 to CPD-2, its hydroxylation activity was very weak (the conversion ratio was less than 1% and was detectable only by the ESI-MS spectrum). Conversely, the strain co-expressing the CYP gene together with a gene encoding its redox partner, ES_SUKA_63, had far higher conversion efficiency (40.0%) than strain ES_SUKA_62. This conversion efficiency was almost equivalent to that of Streptomyces sp. EAS-AB2608 (42.2%). Therefore, we considered that EASAB2608_00799 is involved in enhancing the hydroxylation activity of EASAB2608_00800. In addition, a byproduct (−177 Da) was detected in the bioconversion by strains EAS-AB2608 and ES_SUKA_63. Therefore, we consider that EASAB2608_00800 and EASAB2608_00799 were also involved in its generation.
Application of ES_SUKA_63
To more fully understand the conversion activity and substrate specificity of each of the strains used in this study, we investigated the conversion of six compounds (Table 2). Strains ES_SUKA_62 and ES_SUKA_70 did not show conversion activity of greater than 10% with any of the compounds. Strain ES_SUKA_63 converted indomethacin and rosiglitazone to peaks estimated to be demethylated forms (−14 Da). Similarly, lapachol was converted to a putative oxidized form (+16 Da) by strain ES_SUKA_63. These peaks were not detected in the case of negative control strain ES_SUKA_70. Therefore, we suggest that EASAB2608_00800 (CYP) also has oxidative activity toward these substrates other than CPD-1. However, strain ES_SUKA_63 showed no conversion activity with respect to carbamazepine or genistein. Streptomyces sp. EAS-AB2608 showed conversion results different from those of strain ES_SUKA_63. Strain EAS-AB2608 converted indomethacin and genistein to the peaks estimated to be their glycosylated forms (+178 Da and +176 Da). In addition, strain EAS-AB2608 converted rosiglitazone and lapachol to various analogs (the conversion rate was less than 10% in each case). Therefore, strain EAS-AB2608 was considered to possess other functional enzymes involved in xenobiotic metabolism, such as glycosyltransferase, in addition to the CYP identified in this study.
Properties of CPD-1 and CPD-2
CPD-2 was recovered from the reaction solution of strains EAS-AB2608 and ES_SUKA_63 in >99% enantiomeric excess, approximately 40% isolated yield, and >99% purity (LC-PDA-MS) after 24-h reaction. The chemical structures of CPD-1 and CPD-2 were confirmed by the MS and NMR data. The positive ESI-MS analysis of CPD-1 showed [M+H]+ and [2M+Na]+ at m/z 341 and 703, whereas these ions were at m/z 357 and 735 for CPD-2, indicating an addition of 16 Da relative to CPD-1. NMR data confirmed the molecular structures of CPD-1 and CPD-2 to be as in Fig. 1. In the HSQC spectra, the C4 methylene signal of CPD-1 (δH 3.31–3.38 ppm/δC 29.36 ppm) changed to a C4 methine signal of CPD-2 (δH 5.19 ppm/δC 67.88 ppm). This signal change from a methylene group to a methine group with a downfield chemical shift indicated that hydroxylation of CPD-2 occurred on the C4 position of the tetrahydrobenzotriazole ring. In the 1H spectrum of CPD-2, the coupling constant of H4 (δH 5.19 ppm) and H5 (δH 4.53 ppm) was 7.9 Hz, which suggested that H4/H5 was an axial–axial configuration. Moreover, this conformation of the tetrahydrobenzotriazole ring was supported by NOESY correlations. From these data, the structure of CPD-2 was determined to be the C4-(R)-hydroxylated form of CPD-1.
Analysis of CPD-1
The molecular formula is C18H21FN6. 1H NMR (600 MHz, methanol-d4): δ ppm 1.70 (s, 9H), 2.41–2.51 (m, 2H), 3.10 (ddd, J = 16.4, 8.5, 8.0 Hz, 1H), 3.17 (ddd, J = 16.6, 5.0, 4.7 Hz, 1H), 3.31–3.38 (m, 2H), 4.82–4.88 (m, 1H), 7.17 (br t, J = 8.8 Hz, 2H), 8.04 (br dd, J = 8.8, 5.4 Hz, 2H), 8.53 (s, 1H). 13C NMR (151 MHz, methanol-d4): δ ppm 22.36, 29.36, 29.70, 30.39, 56.73, 62.23, 116.56 (d, 2JC-F = 22.4 Hz), 128.50 (d, 4JC-F = 2.8 Hz), 129.41 (d, 3JC-F = 8.1 Hz), 132.13, 142.80, 144.94, 162.43, 165.02 (d, 1JC-F = 247.8 Hz).
Analysis of CPD-2
The molecular formula is C18H21FN6O. 1H NMR (600 MHz, methanol-d4): δ ppm 1.71 (s, 9H), 2.40 (dddd, J = 13.7, 5.4, 3.2, 2.8 Hz, 1H), 2.55–2.63 (m, 1H), 3.13 (dddd, J = 16.6, 10.5, 5.6, 1.0 Hz, 1H), 3.20 (ddd, J = 16.6, 5.7, 3.2 Hz, 1H), 4.53 (ddd, J = 11.3, 8.0, 3.1 Hz, 1H), 5.19 (d, J = 7.9 Hz, 1H), 7.17 (br t, J = 8.8 Hz, 2H), 8.05 (br dd, J = 8.8, 5.4 Hz, 2H), 8.53 (s, 1H). 13C NMR (151 MHz, methanol-d4): δ ppm 22.55, 28.83, 29.73, 62.32, 64.48, 67.88, 116.55 (d, 2JC-F = 22.1 Hz), 128.57 (d, 4JC-F = 3.2 Hz), 129.43 (d, 3JC-F = 8.5 Hz), 133.20, 146.33, 146.38, 162.68, 165.00 (d, 1JC-F = 247.3 Hz).
Discussion
Transcriptional analysis by RNA-Seq has a wide dynamic range and enables comprehensive comparison of levels of gene expression between samples (Wang et al. 2009). RNA-Seq has been used to gain a deep understanding of the metabolic and degradation pathways of specific compounds (Kim et al. 2018a; Zhang et al. 2019). Furthermore, attempts to explore for genes encoding functional CYP from genomes of eukaryotes by RNA-Seq have already been reported (Chen et al. 2018; Hori et al. 2018). In this study, we applied RNA-Seq analysis to a whole-cell bioconversion system using Actinobacteria.
Streptomyces sp. EAS-AB2608 has the ability to convert CPD-1 to its hydroxylated form (CPD-2). By using RNA-Seq analysis, we were able to efficiently select the CYP (EASAB2608_00800) involved in the hydroxylation of CPD-1 from among the 45 putative CYPs possessed by Streptomyces sp. EAS-AB2608. In parallel, Fdx EASAB2608_00799 was identified as the redox partner of CYP EASAB2608_00800. In the case of bioconversion of CPD-1, strain ES_SUKA_62, which expressed only the CYP, showed far weaker hydroxylation activity than did ES_SUKA_63 (the strain co-expressing the CYP and the Fdx) or EAS-AB2608. This suggests that Fdx EASAB2608_00799 is selectively recognized by CYP EASAB2608_00800 and that neither the other endogenous ferredoxins possessed by Streptomyces sp. EAS-AB2608 nor flavodoxins act as redox partners. Pandey et al. (2014) analyzed gene expressions of Fdx and Fpr by using quantitative reverse transcription polymerase chain reaction in order to identify the optimal redox partner genes of CYP105D7. They clarified the expression levels of sav7470 (fdx) and sav5675 (fpr) that are induced by daidzein (an isoflavone derived from, e.g., Leguminosae) and found that CYP105D7 shows hydroxylation activity with respect to daidzein only by the combination of SAV7470 (Fdx) and SAV5675 (Fpr) among the endogenous redox partner enzymes possessed by S. avermitilis MA-4680. Therefore, in the exploration of functional CYPs, the selection of their optimal redox partners by transcriptional analysis is important. From this perspective, RNA-Seq analysis could help us understand the relationship and function of each enzyme, especially CYPs and their redox partners, involved in whole-cell bioconversion systems.
In contrast, the ferredoxin reductase corresponding to EASAB2608_00799 was not identified, and the enzyme activities and functions of the other CYPs not selected in this study were not investigated. The Streptomyces have highly regulated networks of gene expression. Streptomyces possess many transcriptional factors including DNA-binding response regulators and sigma factors, and approximately 20 gene families have been reported as transcriptional regulators involved in primary metabolism, biosynthesis of natural products, and antibiotics resistance (Romero-Rodríguez et al. 2015). However, no transcriptional regulators of metabolic enzymes such as CYPs involved in the metabolism of xenobiotics have been clarified. The regulatory mechanism for the transcription of an operon encoding EASAB2608_00800 and EASAB2608_00799 identified in this study is also still unclear.
Expression systems using Actinobacteria have already been reported as tools for heterologous expression of CYPs. Imoto et al. (2011) performed the conversion of vitamin D3 to 25-hydroxyvitamin D3 by using a Rhodococcus erythropolis expression system. R. erythropolis host cells can be treated with nisin (lantibiotic) to improve their membrane permeability. In general, heterologous expression of membrane-bound CYPs derived from eukaryotes is difficult in bacterial hosts due to differences in prokaryotic and eukaryotic membrane composition, but Felpeto-Santero et al. (2019) evaluated 11β-hydroxylation activity with respect to cortexolone (anti-androgenic drug) of membrane-bound CYP from Cochliobolus lunatus by using Corynebacterium glutamicum. In the current study, strain ES_SUKA_63 showed oxidative activity with respect to a variety of substrates, including CPD-1, indomethacin, and rosiglitazone. In the case of bioconversion using strain ES_SUKA_63, no side reaction due to glycosyltransferase or the like was detected. Therefore, the combination of S. avermitilis SUKA24 ∆saverm3882 ∆saverm7426 and expression vector pKU565bla-tsr::Psav2794-fld-fpr-ter (or other type expression vector) will be a useful tool for performing selective oxidative reactions and will contribute to the drug discovery process, such as in the preparation of drug metabolites and in lead optimization phase studies as well. Our expression system complements other expression systems used to understand the function of CYPs and their redox partners.
We have applied this approach to six bacterial strains so far, including this study (data not shown). Although the criteria for gene selection by RNA-Seq were changed for each strain, we succeeded in identifying the genes encoding functional CYPs and their redox partners in five out of six strains. The one unsuccessful strain converted the substrate to multiple oxidative metabolites. When whole-cell bioconversion shows low selectivity with respect to the substrate, it can be difficult to identify the responsible gene products under the experimental conditions of this study. Therefore, sample preparation and analysis methods need to be optimized for each particular compound and microorganism. Currently, we are confirming whether this approach is applicable to the identification of gene products of metabolic enzymes other than CYP. This approach is expected to be a useful for bridging the gap between the latest enzyme optimization technologies and conventional enzyme screening using microorganisms.
References
Adams JP, Brown MJ, Diaz-Rodriguez A, Lloyd RC, Roiban GD (2019) Biocatalysis: a pharma perspective. Adv Synth Catal 361:2421–2432. https://doi.org/10.1002/adsc.201900424
Arai M, Naito A, Okazaki T, Serizawa N, Iwado S (1990) Application of actinomycetes in the production of pravastatin, a novel cholesterol-lowering agent (in Japanese with English abstract). Actinomycetologica 4:95–102. https://doi.org/10.3209/saj.4_95
Baldwin SJ, Clarke SE, Chenery RJ (1999) Characterization of the cytochrome P450 enzymes involved in the in vitro metabolism of rosiglitazone. Br J Clin Pharmacol 48:424–432. https://doi.org/10.1046/j.1365-2125.1999.00030.x
Barka EA, Vatsa P, Sanchez L, Gaveau-Vaillant N, Jacquard C, Klenk HP, Clément C, Ouhdouch Y, van Wezel GP (2016) Taxonomy, physiology, and natural products of Actinobacteria. Microbiol Mol Biol Rev 80:1–43. https://doi.org/10.1128/MMBR.00019-15
Burg RW, Miller BM, Baker EE, Birnbaum J, Currie SA, Hartman R, Kong Y, Monaghan RL, Olson G, Putter I, Tunac JB, Wallick H, Stapley EO, Oiwa R, Ōmura S (1979) Avermectins, new family of potent anthelmintic agents: producing organism and fermentation. Antimicrob Agents Chemother 15:361–367. https://doi.org/10.1128/aac.15.3.361
Chapman J, Ismail AE, Dinu CZ (2018) Industrial applications of enzymes: recent advances, techniques, and outlooks. Catalysts 8:238. https://doi.org/10.3390/catal8060238
Chen C, Wang C, Liu Y, Shi X, Gao X (2018) Transcriptome analysis and identification of P450 genes relevant to imidacloprid detoxification in Bradysia odoriphaga. Sci Rep 8:1–9. https://doi.org/10.1038/s41598-018-20981-2
Choi JM, Han SS, Kim HS (2015) Industrial applications of enzyme biocatalysis: current status and future aspects. Biotechnol Adv 33:1443–1454. https://doi.org/10.1016/j.biotechadv.2015.02.014
Colin PY, Kintses B, Gielen F, Miton CM, Fischer G, Mohamed MF, Hyvönen M, Morgavi DP, Janssen DB, Hollfelder F (2015) Ultrahigh-throughput discovery of promiscuous enzymes by picodroplet functional metagenomics. Nat Commun 6:1–12. https://doi.org/10.1038/ncomms10008
Douka A, Vouyiouka S, Papaspyridi LM, Papaspyrides CD (2018) A review on enzymatic polymerization to produce polycondensation polymers: the case of aliphatic polyesters, polyamides and polyesteramides. Prog Polym Sci 79:1–25. https://doi.org/10.1016/j.progpolymsci.2017.10.001
Ellard JM, Madin A, Philps O, Hopkin M, Henderson S, Birch L, O’Connor D, Arai T, Takase K, Morgan L, Reynolds D, Talma S, Howley E, Powney B, Payne AH, Hall A, Gartlon JE, Dawson LA, Castro L, Atkinson PJ (2015) Identification and optimisation of a series of tetrahydrobenzotriazoles as metabotropic glutamate receptor 5-selective positive allosteric modulators that improve performance in a preclinical model of cognition. Bioorg Med Chem Lett 25:5792–5796. https://doi.org/10.1016/j.bmcl.2015.10.050
Felpeto-Santero C, Galán B, Luengo JM, Fernández-Cañon JM, Del Cerro C, Medrano FJ, García JL (2019) Identification and expression of the 11β-steroid hydroxylase from Cochliobolus lunatus in Corynebacterium glutamicum. Microb Biotechnol 12:856–868. https://doi.org/10.1111/1751-7915.13428
Hori K, Yamada Y, Purwanto R, Minakuchi Y, Toyoda A, Hirakawa H, Sato F (2018) Mining of the uncharacterized cytochrome P450 genes involved in alkaloid biosynthesis in California poppy using a draft genome sequence. Plant Cell Physiol 59:222–233. https://doi.org/10.1093/pcp/pcx210
Ikeda H, Ishikawa J, Hanamoto A, Shinose M, Kikuchi H, Shiba T, Sakaki Y, Hattori M, Omura S (2003) Complete genome sequence and comparative analysis of the industrial microorganism Streptomyces avermitilis. Nat Biotechnol 21:526–531. https://doi.org/10.1038/nbt820
Ikeda H, Shi-ya K, Omura S (2014) Genome mining of the Streptomyces avermitilis genome and development of genome-minimized hosts for heterologous expression of biosynthetic gene clusters. J Ind Microbiol Biotechnol 41:233–250. https://doi.org/10.1007/s10295-013-1327-x
Imoto N, Nishioka T, Tamura T (2011) Permeabilization induced by lipid II-targeting lantibiotic nisin and its effect on the bioconversion of vitamin D3 to 25-hydroxyvitamin D3 by Rhodococcus erythropolis. Biochem Biophys Res Commun 405:393–398. https://doi.org/10.1016/j.bbrc.2011.01.038
Jenkins CM, Waterman MR (1994) Flavodoxin and NADPH-flavodoxin reductase from Escherichia coli support bovine cytochrome P450c17 hydroxylase activities. J Biol Chem 269:27401–27408
Kelly SL, Kelly DE (2013) Microbial cytochromes P450: biodiversity and biotechnology. Where do cytochromes P450 come from, what do they do and what can they do for us? Philos Trans R Soc Lond Ser B Biol Sci 368:20120476. https://doi.org/10.1098/rstb.2012.0476
Kelly SL, Kelly DE, Jackson CJ, Warrilow AG, Lamb DC (2005) The diversity and importance of microbial cytochromes P450. In: Paul R (ed) Ortiz de Montellano (ed) Cytochrome P450. Springer, Boston, MA, pp 585–617. https://doi.org/10.1007/0-387-27447-2_13
Kim D, Park HJ, Sul WJ, Park H (2018a) Transcriptome analysis of Pseudomonas sp. from subarctic tundra soil: pathway description and gene discovery for humic acids degradation. Folia Microbiol (Praha) 63:315–323. https://doi.org/10.1007/s12223-017-0573-0
Kim JH, Komatsu M, Shin-Ya K, Omura S, Ikeda H (2018b) Distribution and functional analysis of the phosphopantetheinyl transferase superfamily in Actinomycetales microorganisms. Proc Natl Acad Sci U S A 115:6828–6833. https://doi.org/10.1073/pnas.1800715115
Kino T, Hatanaka H, Miyata S, Inamura N, Nishiyama M, Yajima T, Goto T, Okuhara M, Kohsaka M, Aoki H, Ochiai T (1987) FK-506, a novel immunosuppressant isolated from a Streptomyces. J Antibiot 40:1256–1265. https://doi.org/10.7164/antibiotics.40.1249
Komatsu M, Uchiyama T, Ōmura S, Cane DE, Ikeda H (2010) Genome-minimized Streptomyces host for the heterologous expression of secondary metabolism. Proc Natl Acad Sci U S A 107:2646–2651. https://doi.org/10.1073/pnas.0914833107
Komatsu M, Komatsu K, Koiwai H, Yamada Y, Kozone I, Izumikawa M, Hashimoto J, Takagi M, Omura S, Shin-ya K, Cane DE, Ikeda H (2013) Engineered Streptomyces avermitilis host for heterologous expression of biosynthetic gene cluster for secondary metabolites. ACS Synth Biol 2:384–396. https://doi.org/10.1021/sb3001003
Kumar A, Dhull DK, Mishra PS (2015) Therapeutic potential of mGluR5 targeting in Alzheimer’s disease. Front Neurosci 9:215. https://doi.org/10.3389/fnins.2015.00215
Lam KS (2009) Application of whole-cell biotransformation in the pharmaceutical industry. In: Tao J (ed) Biocatalysis for the pharmaceutical industry: discovery, development, and manufacturing. John Wiley & Sons (Asia), Singapore, pp 213–227
Leis B, Angelov A, Mientus M, Li H, Pham VT, Lauinger B, Bongen P, Pietruszka J, Goncalves LG, Santos H, Liebl W (2015) Identification of novel esterase-active enzymes from hot environments by use of the host bacterium Thermus thermophilus. Front Microbiol 6:275. https://doi.org/10.3389/fmicb.2015.00275
Liu X, Kokare C (2017) Microbial enzymes of use in industry. In: Brahmachari G (ed) Biotechnology of microbial enzymes: production, biocatalysis and industrial applications. Academic Press, Cambridge, MA, pp 267–298. https://doi.org/10.1016/B978-0-12-803725-6.00011-X
Norjannah B, Ong HC, Masjuki HH, Juan JC, Chong WT (2016) Enzymatic transesterification for biodiesel production: a comprehensive review. RSC Adv 6:60034–60055. https://doi.org/10.1039/C6RA08062F
Okubo S, Shibuguchi T, Ishihara Y, Shin K, Ena E, Okuda A (2018) Application of microbial C-H bond activation to chemical synthesis for drug discovery [Poster session]. The 36th Medicinal Chemistry Symposium, Kyoto, Japan (November 28-30)
Oliver HF, Orsi RH, Ponnala L, Keich U, Wang W, Sun Q, Cartinhour SW, Filiatrault MJ, Wiedmann M, Boor KJ (2009) Deep RNA sequencing of L. monocytogenes reveals overlapping and extensive stationary phase and sigma B-dependent transcriptomes, including multiple highly transcribed noncoding RNAs. BMC Genomics 10:641. https://doi.org/10.1186/1471-2164-10-641
Pandey BP, Lee N, Choi KY, Kim JN, Kim EJ, Kim BG (2014) Identification of the specific electron transfer proteins, ferredoxin, and ferredoxin reductase, for CYP105D7 in Streptomyces avermitilis MA4680. Appl Microbiol Biotechnol 98:5009–5017. https://doi.org/10.1007/s00253-014-5525-x
Park ST, Kim J (2016) Trends in next-generation sequencing and a new era for whole genome sequencing. Int Neurourol J 20:S76. https://doi.org/10.5213/inj.1632742.371
Quaderer R, Omura S, Ikeda H, Cane DE (2006) Pentalenolactone biosynthesis. Molecular cloning and assignment of biochemical function to PtlI, a cytochrome P450 of Streptomyces avermitilis. J Am Chem Soc 128:13036–13037. https://doi.org/10.1021/ja0639214
Raveendran S, Parameswaran B, Beevi Ummalyma S, Abraham A, Kuruvilla Mathew A, Madhavan A, Rebello S, Pandey A (2018) Applications of microbial enzymes in food industry. Food Technol. Biotechnol 56:16–30. https://doi.org/10.17113/ftb.56.01.18.5491
Roh C, Seo SH, Choi KY, Cha M, Pandey BP, Kim JH, Park JS, Kim DH, Chang IS, Kim BG (2009) Regioselective hydroxylation of isoflavones by Streptomyces avermitilis MA-4680. J Biosci Bioeng 108:41–46. https://doi.org/10.1016/j.jbiosc.2009.02.021
Romero-Rodríguez A, Robledo-Casados I, Sánchez S (2015) An overview on transcriptional regulators in Streptomyces. Biochim Biophys Acta Gene Regul Mech 1849:1017–1039. https://doi.org/10.1016/j.bbagrm.2015.06.007
Rosenthal K, Lütz S (2018) Recent developments and challenges of biocatalytic processes in the pharmaceutical industry. Curr Opin Green Sustain Chem 11:58–64. https://doi.org/10.1016/j.cogsc.2018.03.015
Shaw NM, Robins KT, Kiener A (2003) Lonza: 20 years of biotransformations. Adv Synth Catal 345:425–435. https://doi.org/10.1002/adsc.200390049
Sheldon RA, Woodley JM (2018) Role of biocatalysis in sustainable chemistry. Chem Rev 118:801–838. https://doi.org/10.1021/acs.chemrev.7b00203
Silva EO, Ruano-Gonzalez A, Dos Santos RA, Sanchez-Maestre R, Furtado NA, Collado IG, Aleu J (2016) Antifungal and cytotoxic assessment of lapachol derivatives produced by fungal biotransformation. Nat Prod Commun 11:95–98. https://doi.org/10.1177/1934578X1601100128
Spasic J, Mandic M, Djokic L, Nikodinovic-Runic J (2018) Streptomyces spp. in the biocatalysis toolbox. Appl Microbiol Biotechnol 102:3513–3536. https://doi.org/10.1007/s00253-018-8884-x
Tompkins LM, Wallace AD (2007) Mechanisms of cytochrome P450 induction. J Biochem Mol Toxicol 21:176–181. https://doi.org/10.1002/jbt.20180
Venkatakrishnan K, von Moltke LL, Greenblatt DJ (2001) Human drug metabolism and the cytochromes P450: application and relevance of in vitro models. J Clin Pharmacol 41:1149–1179. https://doi.org/10.1177/00912700122012724
Waksman SA, Reilly HC, Johnstone DB (1946) Isolation of streptomycin-producing strains of Streptomyces griseus. J Bacteriol 52:393
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57–63. https://doi.org/10.1038/nrg2484
Yang W, Cao H, Xu L, Zhang H, Yan Y (2015) A novel eurythermic and thermostale lipase LipM from Pseudomonas moraviensis M9 and its application in the partial hydrolysis of algal oil. BMC Biotechnol 15:94. https://doi.org/10.1186/s12896-015-0214-0
Zhang W, Zhang T, Jiang B, Mu W (2017) Enzymatic approaches to rare sugar production. Biotechnol Adv 35:267–274. https://doi.org/10.1016/j.biotechadv.2017.01.004
Zhang C, Hao Q, Zhang S, Zhang Z, Zhang X, Sun P, Pan H, Zhang H, Sun F (2019) Transcriptomic analysis of chlorimuron-ethyl degrading bacterial strain Klebsiella jilinsis 2N3. Ecotoxicol Environ Saf 183:109581. https://doi.org/10.1016/j.ecoenv.2019.109581
Acknowledgements
We thank Dr. Tadashi Kadowaki (DENSO Corporation) for instruction and support with regard to RNA-Seq analysis, which is an important aspect of this study. This work was supported in part by the Japan Agency for Medical Research and Development (AMED) under grant number JP20ae0101045 to K.S.
Availability of data and materials
Streptomyces sp. EAS-AB2608 was deposited with National Institute of Technology and Evaluation, Biological Resource Center (www.nite.go.jp/en/nbrc/) as NBRC 114648. The genome sequence information of Streptomyces sp. EAS-AB2608 is available in the DDBJ/EMBL/GenBank databases under accession number AP024135. The RNA-Seq raw data obtained in this study is also available in the DDBJ Sequence Read Archive under accession number DRA011062.
Code availability
Strand NGS v3.1 (www.strand-ngs.com/) was used for RNA-Seq analysis.
Author information
Authors and Affiliations
Contributions
SO, HI, and KS conceived the experiments. YN, MF, and NS contributed genome sequencing. HI contributed development of expression system (S. avermitilis SUKA24 ∆saverm3882 ∆saverm7426 and expression vector pKU565bla-tsr::Psav2794-fld-fpr-ter). IK and JH performed heterologous expression. SO performed RNA-Seq analysis, bioconversion, and LC-PDA-MS analyses. AO performed purification of CPD-2. EE performed NMR analyses. SO wrote the manuscript. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Shinya Okubo now belongs to the Global Health Research Section, hhc Data Creation Center, Eisai Co., Ltd.
Rights and permissions
About this article
Cite this article
Okubo, S., Ena, E., Okuda, A. et al. Identification of functional cytochrome P450 and ferredoxin from Streptomyces sp. EAS-AB2608 by transcriptional analysis and their heterologous expression. Appl Microbiol Biotechnol 105, 4177–4187 (2021). https://doi.org/10.1007/s00253-021-11304-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11304-z