Abstract
River-bay system is a transitional zone connecting land and ocean and an important natural source for methane emission. Methanogens play important roles in the global greenhouse gas budget and carbon cycle since they produce methane. The abundance and community assemblage of methanogens in such a dynamic system are not well understood. Here, we used quantitative PCR and high-throughput sequencing of the mcrA gene to investigate the abundance and community composition of methanogens in the Shenzhen River-Bay system, a typical subtropical river-bay system in Southern of China, during the wet and dry seasons. Results showed that mcrA gene abundance was significantly higher in the sediments of river than those of estuary, and was higher in wet season than dry season. Sequences of mcrA gene were mostly assigned to three orders, including Methanosarcinales, Methanomicrobiales, and Methanobacteriales. Specifically, Methanosarcina, Methanosaeta, and Methanobacterium were the most abundant and ubiquitous genera. Methanogenic communities generally clustered according to habitat (river vs. estuary), and salinity was the major factor driving the methanogenic community assemblage. Furthermore, the indicator groups for two habitats were identified. For example, Methanococcoides, Methanoculleus, and Methanogenium preferentially existed in estuarine sediments, whereas Methanomethylovorans, Methanolinea, Methanoregula, and Methanomassiliicoccales were more abundant in riverine sediments, indicating distinct ecological niches. Overall, these findings reveal the distribution patterns of methanogens and expand our understanding of methanogenic community assemblage in the river-bay system.
Key Points
• Abundance of methanogens was relatively higher in riverine sediments.
• Methanogenic community in estuarine habitat separated from that in riverine habitat.
• Salinity played a vital role in regulating methanogenic community assemblage.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Methane, as a greenhouse gas, contributes 20% to global warming. Its emission from natural sources such as wetlands, hydrothermal vents, and oceans account for more than 40% of the global total methane emission, among which wetland is the largest natural source (Bridgham et al. 2013; Dalal and Allen 2008; Lyu et al. 2018). Coastal wetlands are important source of methane emission and the methane fluxes from coastal wetlands range from 100 to 231 × 102 g a−1 (Tong et al. 2010; Welti et al. 2016). The area of coastal wetlands in China are estimated at more than 1.2 × 104 km2, making up a considerable proportion of the natural wetlands (Huang et al. 2006). Methane emission rates are higher from polluted wetlands than those from unpolluted ones (Purvaja and Ramesh 2000; Zheng et al. 2018).
Methane is produced by methanogens through the process of methanogenesis under anaerobic conditions (Liu and Whitman 2008). Traditionally, methanogens from the phylum Euryarchaeota can be classified into seven orders (Methanococcales, Methanopyrales, Methanobacteriales, Methanomicrobiales, Methanocellales, Methanosarcinales, and Methanomassiliicoccales). Recent genome binning has revealed that the new class of Methanofastidiosa, the new phyla of Verstraetearchaeota, Korarchaeota, and Nezhaarchaeota also have potentials for methanogenesis (McKay et al. 2019; Nobu et al. 2016; Vanwonterghem et al. 2016; Wang et al. 2019b). Methanogens harbor three major pathways of methanogenesis, i.e., hydrogenotrophic, acetoclastic, and methylotrophic pathways (Conrad 2009). All the pathways have a common enzyme, methyl-coenzyme M reductase (MCR), for the final step of methane synthesis (Liu and Whitman 2008). The mcrA gene, which encodes alpha subunit of MCR, is a commonly used gene marker for surveying diversity of methanogens (Yang et al. 2014).
Previous studies based on mcrA gene have observed a vast diversity of methanogens in various environments including animal digestive tract, soil, sediments, and anaerobic digesters (Lu et al. 2015; Mihajlovski et al. 2008; Wilkins et al. 2015). The distribution and environmental preferences of methanogenic groups are also diverse. For example, Methanofastidiosa genomes are obtained from anaerobic sludge digester (Nobu et al. 2016); Methanocellales has been isolated from paddy field soils (Lyu and Lu 2012; Sakai et al. 2008); Methanococcales is preferentially identified in marine habitats (Liu and Whitman 2008); Methanopyrales is the only methanogens so far that could produce methane at temperature higher than 100 °C (Liu 2010); Methanomassiliicoccales has been isolated from human feces (Borrel et al. 2012). These studies indicate that methanogenic community composition could be influenced by environmental factors, such as salinity and temperature.
The river-bay system is located in coastal wetlands connecting land and ocean. Sediments in such environments are under tidal influences and are characterized by an absence of oxygen some or all of the time. In addition, coastal areas are normally densely populated and remarkably disturbed by anthropogenic activities. Due to lack of oxygen and relatively high carbon inputs, coastal sediments are optimal environments for methanogens (Taketani et al. 2010). Methanogenic communities are susceptible to environmental changes across the river-bay continuum where continental freshwater meets oceanic water. Understanding the spatial and seasonal variability of methanogens in the river-bay system is important for recognizing major players involved in methane production. However, few studies have reported the transition of methanogenic communities in the river-bay system.
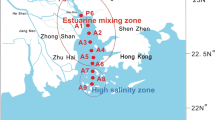
The Shenzhen River-Bay system is located in subtropical coastal areas in south China. It is an economically developed and densely populated area in China (Wang et al. 2019a). It extends the freshwater-saltwater interface from the Dasha River to Shenzhen Bay and is subjected to tidal influence (Fig. S1). The underlying mechanism of methanogenic community assemblage in such a dynamic and fluctuated system is still not well understood. Since riverine and estuarine sediments in the river-bay system are two types of habitats, some groups of methanogens might be found in distinct niches. In the current study, we analyzed samples collected during two seasons (wet and dry seasons) from the Shenzhen River-Bay system to explore the spatial and seasonal variation of methanogenic abundance and community composition. Our objectives are (i) to explore the transition of methanogen community structure in the Shenzhen River-Bay system, and (ii) to reveal the driving factors for the methanogenic community assemblage in the Shenzhen River-Bay system.
Materials and methods
Study sites, sediment sampling, and environmental parameter analysis
The Shenzhen River-Bay system includes Dasha River and Shenzhen Bay. The Dasha River originates in the north of Shenzhen and flows south towards Shenzhen Bay. Shenzhen Bay is a semi-enclosed bay located in the east coast of the Pearl River Estuary. We selected 6 sampling sites from Dasha River (R1, R3–R7) and 10 sampling sites from Shenzhen Bay (E1–E5, E7–E11) along salinity gradients (Fig. S1). The sampling sites were not named consecutively because the sediments of two sites (R2 and E6) were not enough for the following physical and molecular analysis and these two sites were removed. At each sampling site, sediments were collected using a stainless steel sampler in July 2016 and February 2017, representing the wet (W) and dry (D) seasons, respectively. Altogether, 32 sediment samples were collected. All samples were transferred to the laboratory on ice and stored at − 40 °C before analyses.
Water temperature, salinity, and pH were measured in situ by using a multiparameter water quality sonde (EXO2, USA). Sediments were air-dried in the laboratory for a few days until the weight kept unchanged. Sediment ammonium, nitrite, and nitrate were extracted from air-dried sediments with 1 M KCl and determined by a Continuous Segmented Flow Analyzer (SEAL AutoAnalyzer 3 HR, USA). Total organic carbon (TOC) and total nitrogen (TN) contents were measured using TOC Analyzer (Shimadzu, Japan), as previously described (Zhang et al. 2019).
DNA extraction and qPCR
Total DNA was extracted from 0.3 g of sediments using the DNeasy PowerSoil Kit (Qiagen, Germany) according to the manufacturer’s instructions. The quantity and quality of the extracted DNA were examined using NanoDrop® ND-2000c UV-Vis spectrophotometer (NanoDrop Technologies, USA). The DNA samples were stored at − 20 °C and used for later molecular analysis. Abundance of the mcrA genes was determined by quantitative PCR (qPCR) on an iCycler iQ 5 thermocycler (Bio-Rad, USA) using the primer pair mlas-mod-F (5′-GGY GGT GTM GGD TTC ACM CAR TA-3′) and mcrA-rev-R (5′-CGT TCA TBG CGT AGT TVG GRT AGT-3′) (Steinberg and Regan 2008). Each reaction with 25 μl contained 12.5 μl of 2 × SYBR® Premix Ex Taq™ (Takara Biotechnology, Japan), 0.5 μl of each primer (10 μM), and 2 μl of diluted DNA template (1–10 ng). The reaction protocol was as follows: 95 °C for 3 min, 30 cycles of 30 s at 95 °C, 45 s at 55 °C, and 30 s at 72 °C. Standard curves were generated using 10-fold serial dilutions of a plasmid containing the mcrA gene fragments. The PCR efficiency for different assays ranged between 90 and 100%, with R2 value of 0.99.
DNA sequencing and data processing
For amplicon sequencing, mcrA gene fragments were amplified using the primer pair mlas-mod-F and mcrA-rev-R (Steinberg and Regan 2008). The purified amplification products were sequenced on the Illumina MiSeq platform at Novogene Bioinformatics Technology Co., Ltd. (Tianjin, China). Across the 32 samples, the high-throughput sequencing yielded 2,965,757 mcrA gene sequences in total, and the minimum sequence number for individual sample was 10,983.
The paired-end raw reads were merged using the join_paired_ends.py script of Quantitative Insights into Microbial Ecology (QIIME1) (Caporaso et al. 2010). The sequence data set was then analyzed using a modified QIIME2 pipeline (Bolyen et al. 2019). Briefly, chimeras were removed and sequences were trimmed by sequence length (478 bp) using DADA2. After denoising, feature table and feature sequences were generated. Each feature in the feature table will be represented by exactly one sequence. Feature sequences were BLAST against the NCBI-nr database to assign taxonomy. All the samples were randomly rarefied to the minimum number (10,983) of sequences per sample to ensure equal sampling depth.
Statistical analysis
The mcrA gene copies were log-transformed prior to statistical analysis to satisfy the normality assumptions. Shannon index, which was commonly used to represent the alpha diversity, was calculated by diversity function in “vegan” R package. Species abundance distribution (SAD) patterns of methanogenic genera were estimated according to their relative abundance and occurrence. According to the sampling habitats (R = river: Dasha River; E = estuary: Shenzhen Bay) and sampling seasons (W = wet season; D = dry season), the samples were separated into four groups: WR, DR, WE, and DE. The significant differences of the physicochemical properties, abundance, Shannon index, and relative abundances of major orders/families across groups were evaluated by single-factor analysis of variance (ANOVA) followed with Duncan’s test (P < 0.05) by “agricolae” R package. Principal coordinate analysis (PCoA) and permutational multivariate analysis of variance (PERMANOVA) were performed to investigate shifts of the methanogenic community based on the genus level across groups using “vegan” R package. The indicator lineages, which were preferentially found in samples grouped by habitat (river vs. estuary), were calculated by IndVal index in “labdsv” R package (De Cáceres and Legendre 2009). The indicator lineages analysis combined relative abundance and frequency of occurrence of methanogenic genera.
Variation partitioning approach (VPA) was used to evaluate the relative importance of habitats, seasons, and environmental factors on methanogenic community using “vegan” R package. Furthermore, redundancy analysis (RDA) was used to explore relationships between methanogenic community and environmental factors using “vegan” R package. Here, environmental factors included water temperature, pH, salinity, and nutrients (ammonia, nitrite, nitrate, TOC, and TN) contents. Multivariate regression tree (MRT) was performed to predict relationships between methanogenic genera and environmental characteristics using “mvpart” R package (De’Ath 2002). The size of tree to be generated was set as 5. Pie chart under every leaf showed the community composition and how the relative abundances of the methanogenic genera contributed to the separation. Relationships between the relative abundances of methanogenic genera and environmental factors were calculated using Spearman’s rank order correlations. All statistical analyses were performed using R software version 3.5.1 (Team 2018).
Results
Environmental parameters
Environmental properties in the Shenzhen River-Bay system varied according to habitat (river vs. estuary) and season (wet vs. dry) (Fig. S2). In general, the concentrations of nutrients (ammonia, nitrate, TOC, and TN) in riverine sediments were significantly higher than those in estuarine sediments (P < 0.001). Nitrite was higher in the wet season than those in the dry season (P < 0.05). Notably, pH and salinity were comparatively higher in estuarine sediments than those in riverine sediments (P < 0.001). In addition, salinity in the dry season was significantly higher than those in the wet season (P < 0.001). Water temperature in the wet season was significantly higher than in the dry season.
Abundance and alpha diversity of methanogens
Quantitative PCR analysis showed that the mcrA gene copies varied significantly according to habitat and season (Fig. 1a). It was significantly higher in the sediments of river than those of estuary (P < 0.05), and significantly higher in the wet season than in the dry season (P < 0.05). Alpha diversity, as measured by Shannon index, showed that there were no significant differences according to habitats and seasons (Fig. 1b).
Abundance and alpha diversity of methanogens. The samples were grouped by different habitats (river and estuary), and seasons (wet and dry seasons). a Log-transformed abundance of the mcrA genes. b Alpha diversity of methanogens, estimated with the Shannon index. Box plots were constructed to show median, interquartile range, and 1.5 × the interquartile range (n = 6 for riverine samples; n = 10 for estuarine samples). The lowercase letters indicate significant differences among groups (one-way ANOVA; Duncan test; P < 0.05). Sample names: W = wet season; D = dry season; R = river, Dasha River; E = estuary, Shenzhen Bay
Community composition of methanogens
The community composition of methanogens at each sampling site in the Shenzhen River-Bay system is presented in Fig. 2. The mcrA gene sequences were mostly assigned into one class and five orders from the phyla Euryarchaeota. Methanosarcinales, Methanomicrobiales, and Methanobacteriales were the top three abundant orders, contributing 44.4%, 30.6%, and 20.2% of the average relative abundance of methanogenic community, respectively. Methanomassiliicoccales, Methanocellales, and Methanofastidiosa were detected but at relatively low abundance, the average relative abundance of which were 2.4%, 1.9%, and 0.4%, respectively. Furthermore, the relative abundances of different genera were plotted against their occurrence to show the SAD patterns (Fig. S3). Methanosarcina, Methanosaeta, and Methanobacterium were the most abundant and ubiquitous genera in the Shenzhen River-Bay system.
There were significant differences of the relative abundance of methanogenic taxon according to habitat (river vs. estuary). The relative abundance of Methanomicrobiaceae was significantly higher in estuarine sediments than in the riverine zones, while those of Methanoregulaceae and Methanomassiliicoccales showed the opposite trends (Fig. S4). Specifically, Methanomicrobiaceae contributed 5.0% of the average relative abundance of the riverine community and 18.3% of the estuarine community. Conversely, Methanoregulaceae comprised 20.1% of the average relative abundance of the riverine community and 6.8% of the estuarine community. Methanomassiliicoccales contributed 3.5% of the average relative abundance of the riverine community and 1.7% of the estuarine community.
Beta diversity of methanogenic community and indicator groups for habitats
PCoA revealed that habitat (river and estuary) was a strong structuring factor (Fig. 3). Methanogenic community in estuarine habitat (WE and DE) clearly separated from that in riverine habitat (WR and DR), regardless of the season. PERMANOVA based on Bray-Curtis dissimilarity corroborated that habitat showed a significant influence on methanogenic community composition (P = 0.001, R2 = 0.137).
To further identify the methanogenic lineages responsible for the observed community variations according to habitat (river vs. estuary), indicator taxa analysis was conducted. Overall, 7 methanogen genera showed a significant IndVal (P < 0.01) for habitat (Fig. 4). These genera were affiliated to Methanomicrobiales, Methanosarcinales, and Methanomassiliicoccales. Among them, Methanolinea, Methanoregula affiliated to Methanomicrobiales, and Methanomethylovorans affiliated to Methanosarcinales were more abundant in riverine sediments. By contrast, Methanoculleus and Methanogenium affiliated to Methanomicrobiales were more abundant in estuarine sediments. Methanococcoides affiliated to Methanosarcinales only occurred in estuarine sediments and were absent from riverine sediments.
Bubble plot showing the average relative abundance of methanogenic genera in the samples grouped following Fig. 1. The genera represent significant indicator lineages according to the IndVal index (P < 0.05). Sample names: W = wet season; D = dry season; R = river, Dasha River; E = estuary, Shenzhen Bay
Driving factors of methanogenic community assemblage
VPA revealed that habitat, season, and environmental factors contributed to the variations of methanogenic communities together. Environmental factors explained the largest proportion of variation (29.6%), followed by habitat (11.1%), and season (2.0%). RDA revealed that water temperature, pH, salinity, TN, TOC, ammonia, nitrite, and nitrate were significantly correlated with the variation of methanogenic communities. Among all the measured environmental factors, salinity and ammonia were more strongly associated with methanogenic community variation than other environmental factors (Fig. S5). MRT analysis was carried out to link the methanogenic community composition and environmental effects (Fig. 5). The analysis showed that salinity could divide the MRT into three major branches, one contained samples with freshwater condition (salinity < 1.11‰), one contained samples with saline condition (1.11‰ < salinity < 27.07‰), and the other one contained samples with hypersaline condition (salinity > 27.07‰). The most discriminated groups accounting for the separation were Methanoregula, Methanoculleus, and Methanogenium. Methanoregula was more abundant in freshwater environment, while Methanoculleus and Methanogenium were more abundant in saline environment. The branch with salinity less than 1.11‰ was then spilt by water temperature. The branch with salinity between 1.11‰ and 27.07‰ could be further divided by pH value.
Multivariate regression tree (MRT) showing the relationships between the methanogenic community and environmental factors. Five-split tree is visualized. Statistics information is listed under tree, including the residual error, cross-validated error, and standard error. Pie charts under each leaf represent the methanogenic community composition at genus level. The corresponding residual error and sample numbers under each leaf are also listed
Next, correlation analyses were performed to examine the relationships between the environmental factors and relative abundances of methanogenic genera (Fig. 6). Spearman’s rank analysis found that pH and salinity were significantly positively correlated with relative abundances of Methanogenium, Methanoculleus, and Methanococcoides (P < 0.05). Moreover, significant positive correlations were found between sediment nutrients concentration (TOC, TN, ammonia, and nitrate) and relative abundances of Methanoregula, Methanomethylovorans, and Candidatus Methanomethylophilus (P < 0.05).
Discussion
In the current study, we systematically investigated the transition of methanogenic abundance and community composition in the Shenzhen River-Bay system. The presented findings provided novel evidence that salinity was a major factor driving the methanogen community assemblage in the river-bay system. Salinity and nutrients’ contents regulated the relative abundance of a few particular methanogenic groups. For example, Methanoregula were positively influenced by nutrients’ contents, whereas Methanogenium and Methanoculleus were positively influenced by salinity. These results implied that particular methanogenic groups had environmental preference.
Quantitative PCR analysis revealed that mcrA gene copies ranged from 104 to 108 per g sediment and differed according to habitats and seasons. The abundance of methanogens was significantly higher in the riverine sediments than that in the estuarine sediments. One of the possible reasons is that methanogenic abundance is associated with nutrients availability (Zhou et al. 2014). Human activities, such as industrial and domestic wastewater, discharge relatively high contents of nutrients in riverine sediments, which might provide substrates for methanogenesis (Liu et al. 2018). Higher concentration of nutrients in the riverine sediments we measured in the current study supported the above opinion. We also found that the methanogenic abundance was higher in the wet season than in the dry season. It might be because of a higher temperature in the wet season than in the dry season, which promoted the growth of methanogens (Chafee et al. 2017; Lu et al. 2015).
We identified diverse methanogens, including one class and five orders from the phylum Euryarchaeota, in the Shenzhen River-Bay system. Methanosarcinales, Methanomicrobiales, and Methanobacteriales were the three dominant methanogen orders identified in the current study. This was consistent with previous reports that Methanomicrobiales and Methanosarcinales were abundant in coastal sediments (Chen and Yin 2013). Methanomassiliicoccales, Methanocellales, and Methanofastidiosa were also detected, but their relative abundance were low, indicating that these methanogens contribute less to methane production in the river-bay system. Methanomassiliicoccales has been isolated from human feces and considered to be the seventh order of methanogens according to phylogenetic analysis (Borrel et al. 2013; Dridi et al. 2012). It has been reported that Methanomassiliicoccales contain two main clades: environmental clade and gastro-intestinal tract (GIT) clade (Söllinger and Urich 2019). Methanomassiliicoccales in river-bay system has to face higher variations in environmental parameters such as salinity, oxygen exposure, and nutrient availability than Methanomassiliicoccales in the GIT clade, which might explain the low abundance in the river-bay system. Methanocellales has been isolated from paddy field soils (Lyu and Lu 2012; Sakai et al. 2008). Methanocellales are key methanogen groups in rice soils, and they are at a small fraction in other environments including riverine sediments (Angel et al. 2012; Li et al. 2012). Methanofastidiosa is a newly described class of methanogens. There is no pure culture of this class for now. The first draft genomes retained from the anaerobic digester were reported in 2016 (Nobu et al. 2016). Methanofastidiosa is suspected to produce methane using methylated thiol as substrates strictly. In addition, genome annotations revealed that Methanofastidiosa lack abilities to fix carbon/nitrogen and synthesize many amino acids (Nobu et al. 2016). Since they require exogenous organic carbon as a carbon source for growth, there is no competitive advantage over other methanogens in the river-bay system.
In the current study, methanogenic communities were clustered according to habitat (river vs. estuary). Salinity was the major environmental factor regulating the methanogenic community assemblage, as indicated by RDA and MRT analysis. It is in agreement with previous studies disclosing that salinity is a primary factor shaping microbial communities (Wang et al. 2019a; Wen et al. 2017). Effects of salinity on growth and metabolisms of methanogens depend on the level of salinity and the pathways of methanogenesis. Low salinity (0.1–0.2‰) is essential to methanogens, probably because of its role in the oxidation of NADH and in the formation of ATP (Lu et al. 2019). In contrast, high salinity (larger than 8‰) inhibits hydrogenotrophic and acetotrophic methanogens but not methylotrophic methanogens. It is because high levels of salinity favor growth of sulfate reducing bacteria (SRB). Due to higher affinity for H2 and acetate, SRB have a thermodynamic advantage over hydrogenotrophic and acetotrophic methanogens. Methylotrophic methanogens could coexist with SRB because they use methyl compounds as substrates (Xiao et al. 2017).
Furthermore, some indicator genera of methanogens were observed to show preferences for freshwater or saline environments, explaining the observed community structure variation. The relative abundances of Methanomethylovorans, Methanolinea, and Methanoregula were significantly higher in freshwater riverine sediments (Fig. 4). Methanomethylovorans, a methanogen belonging to Methanosarcinales, have been isolated from freshwater sediment (Lomans et al. 1999). Methanolinea and Methanoregula, affiliated to the order Methanomicrobiales, are detected from diverse habitats including digester sludge, wastewater treatment systems, rice field soils, and riverine sediments (Chen and Yin 2013; Kuroda et al. 2015; Sakai et al. 2012; Yashiro et al. 2011). The relative abundance of Methanoregula was positively and significantly correlated to nutrient contents, which might explain its preference for riverine sediments. The current results also showed that the relative abundance of Methanomassiliicoccales was significantly higher in riverine sediments during wet season. Methanomassiliicoccales could utilize methyl-compounds such as methanol, methylamine, and methyl thiol for methane production (Borrel et al. 2014). The upstream riverine zone (sites R3) had a larger input of nutrients due to discharges of wastewater from the nearby Xili wastewater treatment plants, thus Methanomassiliicoccales had relatively higher abundance in the riverine sediments.
In contrast, the relative abundances of Methanococcoides, Methanoculleus, and Methanogenium were significantly higher in saline estuarine sediments (Fig. 4). Methanococcoides, a genus in Methanosarcinales, was absent from riverine sediments, suggesting that Methanococcoides is indicator group of saline environments. It has been reported that Methanococcoides is a predominant lineage in marine sediments and barely exist in riverine sediments (Wen et al. 2017; Xiao et al. 2017). It might be explained by that Methanococcoides are methylotrophic methanogens and could compete with sulfate reducer in saline environments (L’Haridon et al. 2014). Methanoculleus and Methanogenium, belonging to the order Methanomicrobiales, were significantly more abundant in estuary than in riverine sediments. It is in agreement with the previous analyses that Methanoculleus and Methanogenium are commonly detected in marine and estuarine environments, but rarely found in river habitats (Romesser et al. 1979; Wen et al. 2017; Weng et al. 2015). They are hydrogenotrophic methanogens and well adapt to low H2 concentration. Therefore, the above genera might have an advantage over other methanogens in saline environments.
It is observed that some methanogenic genera such as Methanosarcina, Methanosaeta, and Methanobacterium were abundant and ubiquitous in all the samples, indicating they had high adaptation to various salinity conditions including freshwater to saline environments. Methanosarcina could utilize multiple substrates including H2 + CO2, acetate, and methyl compounds for methanogenesis, which gave them an advantage to exist in different environments (Thauer et al. 2008). Methanosaeta was acetoclastic methanogens and was frequently detected in wetlands (Narrowe et al. 2017; Welte and Deppenmeier 2014). It has been reported that Methanobacterium were detected in diverse environments, such as freshwater sediments and anaerobic digester (Kern et al. 2015; Schirmack et al. 2014) Methanobacterium was proposed to be involved in syntrophic methane production with syntrophs such as Geobacter, Desulfovibrio, Pelobacter, Pelotomaculum, and Syntrophomonas (Losey et al. 2017; Yin et al. 2019), which might favor the widespread of Methanobacterium. In addition, we used traditional mcrA gene primer pair that targeted Euryarchaeotal methanogens in this study. It has low specificity to ANaerobic MEthanotrophic (ANME) archaea and the novel methanogens in other phyla such as Verstraetearchaeota, Korarchaeota, and Nezhaarchaeota, potentially not reaching the highest diversity of methanotrophs and methanogens (McKay et al. 2017). Therefore, the response obtained could slightly vary if the groups not considered here were taken into account. Further studies should be conducted to design new primers and investigate the novel methanogens in future.
In conclusion, we found that abundance of methanogens was significantly higher in the riverine than estuarine sediments. Community of methanogens was separated by habitat (river vs. estuary). According to RDA and MRT analysis, salinity was the major factor regulating methanogenic community assemblage. Interestingly, the indicator groups for riverine and estuarine environments were identified, suggesting some methanogenic groups were environment-specific. Methanomethylovorans, Methanolinea, Methanoregula, and Methanomassiliicoccales preferentially dwelled in riverine sediments, while Methanococcoides, Methanoculleus, and Methanogenium preferred existing in estuarine sediments. Collectively, the current study contributes to the understanding of the spatial and seasonal patterns of methanogens and addresses the underlying mechanisms explaining methanogenic community assemblage in the Shenzhen River-Bay system.
Data availability
All the raw reads in the current study have been deposited in the sequencing read archive (SRA) of NCBI under BioProject accession number “PRJNA601619”.
References
Angel R, Claus P, Conrad R (2012) Methanogenic archaea are globally ubiquitous in aerated soils and become active under wet anoxic conditions. ISME J 6(4):847–862. https://doi.org/10.1038/ismej.2011.141
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet C, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope E, Da Silva R, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley G, Janssen S, Jarmusch AK, Jiang L, Kaehler B, Kang KB, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu Y-X, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton J, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson IIMS, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852-857. https://doi.org/10.1038/s41587-019-0209-9
Borrel G, Harris HMB, Tottey W, Mihajlovski A, Parisot N, Peyretaillade E, Peyret P, Gribaldo S, O’Toole PW, Brugèrea J-F (2012) Genome sequence of “Candidatus Methanomethylophilus alvus” Mx1201, a methanogenic archaeon from the human gut belonging to a seventh order of methanogens. J Bacteriol 194(24):6944–6945. https://doi.org/10.1128/JB.01867-1210.1371/
Borrel G, O'Toole PW, Harris HM, Peyret P, Brugere JF, Gribaldo S (2013) Phylogenomic data support a seventh order of methylotrophic methanogens and provide insights into the evolution of Methanogenesis. Genome Biol Evol 5(10):1769–1780. https://doi.org/10.1093/gbe/evt128
Borrel G, Parisot N, Harris HM, Peyretaillade E, Gaci N, Tottey W, Bardot O, Raymann K, Gribaldo S, Peyret P, O’Toole PW, Brugère J-F (2014) Comparative genomics highlights the unique biology of Methanomassiliicoccales, a Thermoplasmatales-related seventh order of methanogenic archaea that encodes pyrrolysine. BMC Genomics 15:679. https://doi.org/10.1186/1471-2164-15-679
Bridgham SD, Cadillo-Quiroz H, Keller JK, Zhuang Q (2013) Methane emissions from wetlands: biogeochemical, microbial, and modeling perspectives from local to global scales. Glob Chang Biol 19(5):1325–1346. https://doi.org/10.1111/gcb.12131
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. https://doi.org/10.1038/nmeth.f.303
Chafee M, Fernàndez-Guerra A, Buttigieg PL, Gerdts G, Eren AM, Teeling H, Amann RI (2017) Recurrent patterns of microdiversity in a temperate coastal marine environment. ISME J 12(1):237–252. https://doi.org/10.1038/ismej.2017.165
Chen J, Yin X (2013) Stratified communities of methanogens in the Jiulong River estuarine sediments, southern China. Indian J Microbiol 53(4):432–437. https://doi.org/10.1007/s12088-013-0397-9
Conrad R (2009) The global methane cycle: recent advances in understanding the microbial processes involved. Environ Microbiol Rep 1(5):285–292. https://doi.org/10.1111/j.1758-2229.2009.00038.x
Dalal RC, Allen DE (2008) Greenhouse gas fluxes from natural ecosystems. Aust J Bot 56:369–407. https://doi.org/10.1071/BT07128
De Cáceres M, Legendre P (2009) Associations between species and groups of sites: indices and statistical inference. Ecology 90(12):3566–3574. https://doi.org/10.1890/08-1823.1
De’Ath G (2002) Multivariate regression trees: a new technique for modeling species - environment relationships. Ecology 83(4):1105–1117. https://doi.org/10.1890/0012-9658(2002)083[1105:MRTANT]2.0.CO;2
Dridi B, Fardeau ML, Ollivier B, Raoult D, Drancourt M (2012) Methanomassiliicoccus luminyensis gen. nov., sp. nov., a methanogenic archaeon isolated from human faeces. Int J Syst Evol Microbiol 62:1902–1907. https://doi.org/10.1099/ijs.0.033712-0
Huang GL, He P, Hou M (2006) Present status and prospects of estuarine wetland research in China. Chin J Appl Ecol 17(9):1751–1756 (in Chinese)
Kern T, Linge M, Rother M (2015) Methanobacterium aggregans sp. nov., a hydrogenotrophic methanogenic archaeon isolated from an anaerobic digester. Int J Syst Evol Microbiol 65(6):1975–1980. https://doi.org/10.1099/ijs.0.000210
Kuroda K, Hatamoto M, Nakahara N, Abe K, Takahashi M, Araki N, Yamaguchi T (2015) Community composition of known and uncultured archaeal lineages in anaerobic or anoxic wastewater treatment sludge. Microb Ecol 69(3):586–596. https://doi.org/10.1007/s00248-014-0525-z
L’Haridon S, Chalopin M, Colombo D, Toffin L (2014) Methanococcoides vulcani sp. nov., a marine methylotrophic methanogen that uses betaine, choline and N,N-dimethylethanolamine for methanogenesis, isolated from a mud volcano, and emended description of the genus Methanococcoides. Int J Syst Evol Microbiol 64:1978–1983. https://doi.org/10.1099/ijs.0.058289-0
Li Q, Wang F, Chen Z, Yin X, Xiao X (2012) Stratified active archaeal communities in the sediments of Jiulong River estuary, China. Front Microbiol 3:311. https://doi.org/10.3389/fmicb.2012.00311
Liu Y (2010) Methanopyrales. In: Timmis KN (ed) Handbook of hydrocarbon and lipid microbiology. Springer, Berlin, pp 605–607
Liu Y, Whitman WB (2008) Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann N Y Acad Sci 1125:171–189. https://doi.org/10.1196/annals.1419.019
Liu X, Pan J, Liu Y, Li M, Gu JD (2018) Diversity and distribution of Archaea in global estuarine ecosystems. Sci Total Environ 637-638:349–358. https://doi.org/10.1016/j.scitotenv.2018.05.016
Lomans BP, Maas R, Luderer R, Den Camp HJMO, Pol A, Drift CV, Vogels GD (1999) Isolation and characterization of Methanomethylovorans hollandica gen. nov., sp. nov., isolated from freshwater sediment, a methylotrophic methanogen able to grow on dimethyl sulfide and methanethiol. Appl Environ Microbiol 65(8):3641–3650
Losey NA, Mus F, Peters JW, Le HM, McInerney MJ (2017) Syntrophomonas wolfei uses an NADH-dependent, ferredoxin-independent [FeFe]-hydrogenase to reoxidize NADH. Appl Environ Microbiol 83(20). https://doi.org/10.1128/AEM.01335-17
Lu Y, Fu L, Lu Y, Hugenholtz F, Ma K (2015) Effect of temperature on the structure and activity of a methanogenic archaeal community during rice straw decomposition. Soil Biol Biochem 81:17–27. https://doi.org/10.1016/j.soilbio.2014.10.031
Lu Q, Yu Z, Yu S, Liang Z, Li H, Sun L, Wang S (2019) Organic matter rather than salinity as a predominant feature changes performance and microbiome in methanogenic sludge digesters. J Hazard Mater 377:349–356. https://doi.org/10.1016/j.jhazmat.2019.05.075
Lyu Z, Lu Y (2012) Methanocella conradii sp. nov., a thermophilic, obligate hydrogenotrophic methanogen, isolated from Chinese rice field soil. PLoS One 7(4):e35279. https://doi.org/10.1371/journal.pone.0035279
Lyu Z, Shao N, Akinyemi T, Whitman WB (2018) Methanogenesis. Curr Biol 28(13):R727–R732. https://doi.org/10.1016/j.cub.2018.05.021
McKay LJ, Hatzenpichler R, Inskeep WP, Fields MW (2017) Occurrence and expression of novel methyl-coenzyme M reductase gene (mcrA) variants in hot spring sediments. Sci Rep 7(1):7252. https://doi.org/10.1038/s41598-017-07354-x
McKay LJ, Dlakic M, Fields MW, Delmont TO, Eren AM, Jay ZJ, Klingelsmith KB, Rusch DB, Inskeep WP (2019) Co-occurring genomic capacity for anaerobic methane and dissimilatory sulfur metabolisms discovered in the Korarchaeota. Nat Microbiol 4(4):614–622. https://doi.org/10.1038/s41564-019-0362-4
Mihajlovski A, Alric M, Brugere JF (2008) A putative new order of methanogenic Archaea inhabiting the human gut, as revealed by molecular analyses of the mcrA gene. Res Microbiol 159(7–8):516–521. https://doi.org/10.1016/j.resmic.2008.06.007
Narrowe AB, Angle JC, Daly RA, Stefanik KC, Wrighton KC, Miller CS (2017) High-resolution sequencing reveals unexplored archaeal diversity in freshwater wetland soils. Environ Microbiol 19(6):2192–2209. https://doi.org/10.1111/1462-2920.13703
Nobu MK, Narihiro T, Kuroda K, Mei R, Liu WT (2016) Chasing the elusive Euryarchaeota class WSA2: genomes reveal a uniquely fastidious methyl-reducing methanogen. ISME J 10(10):2478–2487. https://doi.org/10.1038/ismej.2016.33
Purvaja R, Ramesh R (2000) Human impacts on methane emission from mangrove ecosystems in India. Reg Environ Chang 1(2):86–97. https://doi.org/10.1007/PL00011537
Romesser JA, Wolfe RS, Mayer F, Spiess E, Walther-Mauruschat A (1979) Methanogenium, a new genus of marine methanogenic bacteria, and characterization of Methanogenium cariaci sp. nov. and Methanogenium marisnigri sp. nov. Arch Microbiol 121:147–153. https://doi.org/10.1007/BF00689979
Sakai S, Imachi H, Hanada S, Ohashi A, Harada H, Kamagata Y (2008) Methanocella paludicola gen. nov., sp. nov., a methane-producing archaeon, the first isolate of the lineage 'Rice cluster I', and proposal of the new archaeal order Methanocellales ord. nov. Int J Syst Evol Microbiol 58:929–936. https://doi.org/10.1099/ijs.0.65571-0
Sakai S, Ehara M, Tseng IC, Yamaguchi T, Brauer SL, Cadillo-Quiroz H, Zinder SH, Imachi H (2012) Methanolinea mesophila sp. nov., a hydrogenotrophic methanogen isolated from rice field soil, and proposal of the archaeal family Methanoregulaceae fam. nov. within the order Methanomicrobiales. Int J Syst Evol Microbiol 62:1389–1395. https://doi.org/10.1099/ijs.0.035048-0
Schirmack J, Mangelsdorf K, Ganzert L, Sand W, Hillebrand-Voiculescu A, Wagner D (2014) Methanobacterium movilense sp. nov., a hydrogenotrophic, secondary-alcohol-utilizing methanogen from the anoxic sediment of a subsurface lake. Int J Syst Evol Microbiol 64:522–527. https://doi.org/10.1099/ijs.0.057224-0
Söllinger A, Urich T (2019) Methylotrophic methanogens everywhere—physiology and ecology of novel players in global methane cycling. Biochem Soc Trans. https://doi.org/10.1042/BST20180565/862468/bst-2018-0565c.pdf
Steinberg LM, Regan JM (2008) Phylogenetic comparison of the methanogenic communities from an acidic, oligotrophic fen and an anaerobic digester treating municipal wastewater sludge. Appl Environ Microbiol 74(21):6663–6671. https://doi.org/10.1128/AEM.00553-08
Taketani RG, Yoshiura CA, Dias AC, Andreote FD, Tsai SM (2010) Diversity and identification of methanogenic archaea and sulphate-reducing bacteria in sediments from a pristine tropical mangrove. Antonie Van Leeuwenhoek 97(4):401–411. https://doi.org/10.1007/s10482-010-9422-8
Team RC (2018) R: A language and environment for statistical computing. https://www.R-project.org/
Thauer RK, Kaster AK, Seedorf H, Buckel W, Hedderich R (2008) Methanogenic archaea: ecologically relevant differences in energy conservation. Nat Rev Microbiol 6(8):579–591. https://doi.org/10.1038/nrmicro1931
Tong C, Wang WQ, Zeng CS, Marrs R (2010) Methane (CH4) emission from a tidal marsh in the Min River estuary, Southeast China. J Environ Sci Health A Tox Hazard Subst Environ Eng 45(4):506–516. https://doi.org/10.1080/10934520903542261
Vanwonterghem I, Evans PN, Parks DH, Jensen PD, Woodcroft BJ, Hugenholtz P, Tyson GW (2016) Methylotrophic methanogenesis discovered in the archaeal phylum Verstraetearchaeota. Nat Microbiol 1:16170. https://doi.org/10.1038/nmicrobiol.2016.170
Wang Y, Pan J, Yang J, Zhou Z, Pan Y, Li M (2019a) Patterns and processes of free-living and particle-associated bacterioplankton and archaeaplankton communities in a subtropical river-bay system in South China. Limnol Oceanogr 65:S161–S179. https://doi.org/10.1002/lno.11314
Wang Y, Wegener G, Hou J, Wang F, Xiao X (2019b) Expanding anaerobic alkane metabolism in the domain of Archaea. Nat Microbiol 4(4):595–602. https://doi.org/10.1038/s41564-019-0364-2
Welte C, Deppenmeier U (2014) Bioenergetics and anaerobic respiratory chains of aceticlastic methanogens. Biochim Biophys Acta 1837(7):1130–1147. https://doi.org/10.1016/j.bbabio.2013.12.002
Welti N, Hayes M, Lockington D (2016) Seasonal nitrous oxide and methane emissions across a subtropical estuarine salinity gradient. Biogeochemistry 132(1–2):55–69. https://doi.org/10.1007/s10533-016-0287-4
Wen X, Yang S, Horn F, Winkel M, Wagner D, Liebner S (2017) Global biogeographic analysis of methanogenic archaea identifies community-shaping environmental factors of natural environments. Front Microbiol 8:1339. https://doi.org/10.3389/fmicb.2017.01339
Weng CY, Chen SC, Lai MC, Wu SY, Lin S, Yang TF, Chen PC (2015) Methanoculleus taiwanensis sp. nov., a methanogen isolated from deep marine sediment at the deformation front area near Taiwan. Int J Syst Evol Microbiol 65:1044–1049. https://doi.org/10.1099/ijs.0.000062
Wilkins D, Lu XY, Shen Z, Chen J, Lee PK (2015) Pyrosequencing of mcrA and archaeal 16S rRNA genes reveals diversity and substrate preferences of methanogen communities in anaerobic digesters. Appl Environ Microbiol 81(2):604–613. https://doi.org/10.1128/AEM.02566-14
Xiao KQ, Beulig F, Kjeldsen KU, Jorgensen BB, Risgaard-Petersen N (2017) Concurrent methane production and oxidation in surface sediment from Aarhus Bay, Denmark. Front Microbiol 8:1198. https://doi.org/10.3389/fmicb.2017.01198
Yang S, Liebner S, Alawi M, Ebenhöh O, Wagner D (2014) Taxonomic database and cut-off value for processing mcrA gene 454 pyrosequencing data by MOTHUR. J Microbiol Methods 103:3–5. https://doi.org/10.1016/j.mimet.2014.05.00610.5880/GFZ.4.5.2014.001
Yashiro Y, Sakai S, Ehara M, Miyazaki M, Yamaguchi T, Imachi H (2011) Methanoregula formicica sp. nov., a methane-producing archaeon isolated from methanogenic sludge. Int J Syst Evol Microbiol 61:53–59. https://doi.org/10.1099/ijs.0.014811-0
Yin Q, Wang Z, Wu G (2019) Impacts of environmental factors on microbial diversity, distribution patterns and syntrophic correlation in anaerobic processes. Arch Microbiol 201(5):603–614. https://doi.org/10.1007/s00203-019-01627-x
Zhang C-J, Pan J, Duan C-H, Wang Y-M, Liu Y, Sun J, Zhou H-C, Song X, Li M (2019) Prokaryotic diversity in mangrove sediments across southeastern China fundamentally differs from that in other biomes. mSystems 4(5):e00442–e00419. https://doi.org/10.1128/mSystems.00442-19
Zheng X, Guo J, Song W, Feng J, Lin G (2018) Methane emission from mangrove wetland soils is marginal but can be stimulated sgnificantly by anthropogenic activities. Forests 9(12):738. https://doi.org/10.3390/f9120738
Zhou Z, Chen J, Cao H, Han P, Gu JD (2014) Analysis of methane-producing and metabolizing archaeal and bacterial communities in sediments of the northern South China Sea and coastal Mai Po Nature Reserve revealed by PCR amplification of mcrA and pmoA genes. Front Microbiol 5:789. https://doi.org/10.3389/fmicb.2014.00789
Funding
This work was financially supported by the Shenzhen Science and Technology Program (JCYJ20170818091727570 and KQTD20180412181334790), the National Natural Science Foundation of China (91851105, 31622002, and 31600093), the Key Project of Department of Education of Guangdong Province (2017KZDXM071) to ML, and a China Postdoctoral Science Foundation (2018M630977) to CZ.
Author information
Authors and Affiliations
Contributions
ML conceived the study. YC and YW sampled sediments in the field and conducted experiments. CZ, YC, and JP analyzed data. CZ wrote the manuscript with help from all coauthors. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 423 kb).
Rights and permissions
About this article
Cite this article
Zhang, CJ., Chen, YL., Pan, J. et al. Spatial and seasonal variation of methanogenic community in a river-bay system in South China. Appl Microbiol Biotechnol 104, 4593–4603 (2020). https://doi.org/10.1007/s00253-020-10613-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10613-z