Abstract
Methicillin-resistant Staphylococcus aureus (MRSA) has become a worrisome superbug, due to its wide distribution and multidrug resistance. To characterize effects of a newly identified plantaricin GZ1-27 on MRSA, transcriptomic and proteomic profiling of MRSA strain ATCC43300 was performed in response to sub-MIC (16 μg/mL) plantaricin GZ1-27 stress. In total, 1090 differentially expressed genes (padj < 0.05) and 418 differentially expressed proteins (fold change > 1.2, p < 0.05) were identified. Centralized protein expression clusters were predicted in biological functions (biofilm formation, DNA replication and repair, and heat-shock) and metabolic pathways (purine metabolism, amino acid metabolism, and biosynthesis of secondary metabolites). Moreover, a capacity of inhibition MRSA biofilm formation and killing biofilm cells were verified using crystal violet staining, scanning electron microscopy, and confocal laser-scanning microscopy. These findings yielded comprehensive new data regarding responses induced by plantaricin and could inform evidence-based methods to mitigate MRSA biofilm formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Methicillin-resistant Staphylococcus aureus (MRSA) is considered a global pandemic threat (Rossolini et al. 2014). As reported, the MRSA incidence exceeds 20% in all 85 investigated World Health Organization (WHO) member states (Álvarez et al. 2019). Since it was first reported in 1961 (Jevons 1961), MRSA has thrived in hospitals, communities, and livestock worldwide (Wendlandt et al. 2013). In the past decade, various MRSA genotypes were identified in raw meat (pork, poultry, and beef) and milk, indicating a high risk of MRSA in food-borne transmission (Titouche et al. 2019). As one of the most common antibiotic-resistant bacteria, MRSA were resistant to most β-lactam antibiotics, including methicillin, penicillin, carbapenems, cephalosporins, and their derivatives (Doyle et al. 2012).
A biofilm is a community of organisms encased in a protective and adhesive matrix. Biofilms allow MRSA strains to adhere to various biotic and abiotic surfaces, including clinical devices and materials used in the food industry (Vergara et al. 2017). Biofilm production is recognized as an important virulence factor of bacteria. The capability of producing biofilm contributes to persistence and dissemination of MRSA in the variety of environments. Biofilms colonized on the clinical equipment surfaces can promote MRSA associated infections in hospitals and thereby infect patients with surgical implants. S. aureus biofilm on food-contact surfaces poses a serious risk of contaminating food, responsible for outbreaks related to consumption of fresh and processed foods (Doulgeraki et al. 2017a). In addition, due to restricted penetration and antimicrobial-destroying enzymes in the extracellular matrix, biofilms have low susceptibility to conventional antimicrobials (del Pozo and Patel 2007). Thus, there is a pressing need for new types of antimicrobial compounds to mitigate MRSA.
Bacteriocins, natural antimicrobial proteins or peptides, are mainly produced by lactic acid bacteria. Due to relatively high specificity, low toxicity and low risk of resistance development, bacteriocins are considered a promising antibacterial agent for treating infectious bacterial disease (Cotter et al. 2013). Bacteriocins have been widely studied for extending shelf-life in a variety of food by inhibiting spoilage-inducing bacteria (e.g., Brochothrix, Aeromonas, and Pseudomonas), as well as food-borne pathogens (e.g., Listeria monocytogenes, S. aureus, Escherichia coli) (Silva et al. 2018) (Du et al. 2019). Also, nisin, the most representative bacteriocin, has been approved as an antimicrobial additive for various foods in the United States, Canada and Europe (Cotter et al. 2013). Regarding effects of bacteriocin on MRSA, most studies have focused on purification and characterization of bacteriocins (Ansari et al. 2018; Hong et al. 2018) and evaluated their activities (Thomsen et al. 2016). However, antibacterial mechanisms against MRSA remain unclear. In addition, although it is generally demonstrated that bacteriocins primarily affect cell membrane integrity (Cotter et al. 2013), detailed mechanisms and global responses are lacking.
With rapid enhancement of “omics” technologies, transcriptomic and proteomic approaches are powerful tools to study responses of bacteria to antibacterial agents. Using isobaric tags for relative and absolute quantitation (iTRAQ)-based proteome analysis, it is speculated that LI-F type peptides AMP-jsa9, a LI-Fs/fusaricidins family cyclic lipodepsipeptide antimicrobial peptide, act against Bacillus cereus through inhibition of bacterial sporulation, thiamine biosynthesis, energy metabolism, DNA transcription and translation, and cell wall biosynthesis (Han et al. 2017). By characterizing a proteomic response library of Bacillus subtilis against lantibiotics, reliable marker proteins for interacting with cytoplasmic membranes were speculated (Wenzel et al. 2012). Global transcriptional responses to the subinhibitory concentrations colicin M in E. coli altered expression of genes involved in envelope, osmotic, exopolysaccharide production, and cell motility (Ek and Gur-Bertok 2013). In B. subtilis 168, transcriptome and proteome analyses identified that eight B. subtilis genes are likely to contribute to inherent nisin resistance (Hansen et al. 2009). Furthermore, integration of transcriptome and proteome should provide more reliable and comprehensive insights in bacteriocin response.
Plantaricins are bacteriocins which are produced by Lactobacillus plantarum. They have attracted great research interests because of their structural diversity and high antimicrobial activity (Zhao et al. 2016; Zhu et al. 2014). Previously, we purified a new plantaricin GZ1-27 from L. plantarum GZ1-27 and identified its peptide sequence: VSGPAGPPGTH (Du et al. 2018). Plantaricin GZ1-27 has high thermostability and pH stability. In addition, plantaricin GZ1-27 was effective against several gram-positive bacteria, including MRSA strains, according to minimal inhibitory concentrations (MICs; Supplemental Table S1). Here our objectives were to (1) investigate an MRSA strain in response to GZ1-27 treatment at gene transcriptional and expression levels and (2) verify the effect of GZ1-27 on biofilm formation and biofilm removal.
Materials and methods
MRSA inhibition curves
The MRSA strain ATCC 43300 used in this study was purchased from Guangdong Institute of Microbiology Culture Center, China, and stored in our lab. Plantaricin GZ1-27 was purified as described (Du et al. 2018). The MIC of plantaricin GZ1-27 against ATCC 43300 was 32 μg/mL. For growth curve assay, a single colony was picked up into 5 mL of Tryptic Soy Broth (TSB, Beijing Land Bridge Technology CO. LTD., China) and shaken (225 rpm) at 37 °C overnight (14 h) (Okuda 2013). Then, 1 mL of overnight culture was added to 100 mL of TSB and shaken (225 rpm) at 37 °C until optical density at 600 nm (OD600) reached 0.4 (~ 108 colony-forming units (CFU)/mL). At that time, 1 mL of plantaricin GZ1-27 (finally concentration corresponding to 1/8×, 1/4×, 1/2×, 1×, and 2× MIC) or disodium hydrogen phosphate-citric acid buffer (solvent for plantaricin GZ1-27; 2.5 mM, pH 3.8; control) were added to log-phase culture, with continuous shaking and OD600 monitoring (UV-2600 UV–VIS spectrophotometer, SHIMADZU Company, Kyoto, Japan) every 30 min.
Sample preparation for transcriptomic and proteomic assays
Aliquots (50 mL) of log-phase culture (OD600 = 0.4) were prepared as described above and incubated with 1/2× MIC plantaricin GZ1-27 at 37 °C (225 rpm) for 3 h, with MRSA log-phase culture cells without GZ1-27 used as a control. After 3 h incubation, plantaricin-treated and non-treated culture were centrifuged at 5000×g for 10 min (4 °C) and pellets washed twice with phosphate buffered saline (PBS, 10 mM, pH 7.4). Three treated and three non-treated pellets were used for RNA isolation. Similarly, using the same preparation method, three treated and three non-treated pellets were used for protein extraction.
RNA library construction and sequencing
Total RNA was extracted using TransZol reagent (TransGen Biotech, Beijing, China) according to manufacturer’s instructions. Concentration and purity of total RNA were measured using NanoDrop 2000 (Thermo Scientific, Waltham, MA). The RNA Integrity Number (RIN) was assessed using Agilent 2100 (Agilent Technologies, Santa Clara, CA). Only high-quality RNA (RIN > 0.8) was used for further library construction. First, rRNA was removed using Ribo-Zero rRNA Removal kit (Illumina, San Diego, CA) and mRNA fragmented using Fragmentation Buffer (Illumina) at 94 °C for 5 min. Then, random hexamers were used for reverse transcription. After bluntness, adding A-tail and adapter, AMPure XP beads were used to select double strand cDNA (average length, 250 bp). Finally, PCR amplification and purification were performed to obtain the RNA library. The RNA library was then submitted and sequenced using an Illumina HiseqTM Xten platform at Novogene Co. Ltd. (Beijing, China), with raw RNA sequence data deposited in the National Center for Biotechnology Information (NCBI; accession SRP173085). A schematic diagram of transcriptomic assay procedures is shown in Supplemental Fig. S1a.
Transcriptomic bioinformation analysis
Raw data were filtered to obtain clean reads by depleting adapter-related sequences, selecting sequences with N > 10% (N indicates that base information cannot be determined) and selecting low-quality sequences with Qpred ≤ 20. Clean reads were mapped against S. aureus subsp. aureus NCTC8325 (https://www.ncbi.nlm.nih.gov/genome/154?genome_assembly_id=299272) genome using Bowtie2 (Langmead and Salzberg 2012). HTSeq software was used to assess gene expression level of each sample with the method of FPKM (Anders et al. 2015). The padj < 0.05 (padj is false discovery rate (FDR)-adjusted p value) was set as a threshold to determine differentially expressed genes (DEGs) between treatment and not-treatment S. aureus cells using DESeq. Cluster analysis of DEGs was performed to search for genes with similar functions. These DEGs were subjected to Gene Ontology (GO) enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis to determine their involvement in functional classification and metabolic pathways.
Protein extraction and digestion
Proteins were extracted as described (Wiśniewski et al. 2009), with some modifications. After mixing with SDT (4% (w/v) sodium dodecyl sulfate (SDS), 100 mM dithiothreitol (DTT), 100 mM Tris/HCl pH 7.6), the suspension was sonicated at 100 W (repeated 10 times) and then centrifuged at 14,000g for 40 min (4 °C). Supernatant was filtered and quantified using BCA protein assay kit (Beyotime Biotechnology, Shanghai, China), with 300 μg of protein from each sample used to perform filter-aided sample preparation (FASP). Peptides were desalted using an Empore™ SPE C18 Cartridge (3 mL, Sigma Chemical Co., St. Louis, MO).
iTRAQ labeling and SCX fraction
Digested peptides (100 μg) were labeled with iTRAQ reagents (Applied Biosystems, Waltham, MA) according to the manufacturer’s protocol. The three control and three treatment samples were labeled with 113, 114, 115 and 116, 117, 118 iTRAQ reagents, respectively (Wiese et al. 2007). After labeling, peptides were mixed and fractionated by strong cation exchange (SCX) chromatography using a AKTA purifier system (GE Healthcare, New York, NY) equipped with a PolySULFOETHYL column (4.6 × 100 mm, 5 μm; PolyLC Inc., Columbia, MD). Firstly, the chromatography column was balanced with buffer A (10 mM KH2PO4 in 25% of acetonitrile, pH 3.0). Then, peptides were gradient-eluted with following process at a flow rate of 1 ml/min. We used 0–8% buffer B (500 mM KCl, 10 mM KH2PO4 in 25% of acetonitrile, pH 3.0) for 22 min, 8–52% buffer B during 22–47 min, 52–100% buffer B during 47–50 min, 100% buffer B during 50–58 min. Collected fractions were pooled into 10 fractions, desalted using a C18 (5 μm, 250 mm × 4.6 mm i.d.) column (Agilent Technologies, Palo Alto, CA) and dried under vacuum.
LC-MS/MS analysis
Each fraction was separated by an Easy nLC (Thermo Scientific) and analyzed with a Q Exactive mass spectrometer (Thermo Scientific) for 60 min. The peptide mixture was re-suspended in buffer A (0.1% formic acid) and loaded onto a reverse-phase trap column (Acclaim PepMap100, 100 μm × 2 cm, nanoViper C18; Thermo Scientific) connected to the C18-reversed phase analytical column (Easy Column, 10 cm long, 75 μm inner diameter, 3 μm resin; Thermo Scientific). Fractions were separated with a linear gradient of buffer B (84% acetonitrile and 0.1% formic acid) at a flow rate of 300 nL/min. Then, we used 0–35% buffer for 50 min, 35–100% buffer B for 5 min and held the sample in 100% buffer for 10 min. The MS/MS analysis was operated in positive ion mode and MS data acquired using a data-dependent top10 method, dynamically choosing the most abundant precursor ions from the survey scan (300–1800 m/z) for HCD fragmentation. All raw mass spectrometry data were deposited in Proteome Xchange (Accession No. PXD012012). A schematic diagram of proteomic assay procedures was shown in Supplemental Fig. S1b.
Proteomic bioinformation analysis
The MS/MS data were searched against transcriptome results using MASCOT engine (Matrix Science, London, UK; Version 2.2) embedded into Proteome Discoverer 1.4 (Casey et al. 2017). Differentially expressed proteins (DEPs) with fold change > 1.2 and FDR-adjusted p < 0.05 were subjected to bioinformation analysis. The GO enrichment on three ontologies (biological process, molecular function and cellular component) and KEGG pathway enrichment analysis were applied, based on Fisher’s Exact test. Visualized protein-protein interaction (PPI) networks were done with STRING software (http://string-db.org/cgi/input.pl) (Szklarczyk et al. 2017) and Cytoscape software (https://cytoscape.org/,version 3.6.1) (Shannon et al. 2003). Comparative analysis of proteomic and transcriptomic data was performed in Venn diagram. Fold change of selected DEGs and DEPs were used to perform hierarchical clustering using Heml (http://hemi.biocuckoo.org/) (Deng et al. 2014).
Biofilm formation
Inhibition of biofilm formation under low level exposure of plantaricin was examined in 96-well polystyrene plate (Bazargani and Rohloff 2016). First, the MRSA cultures were adjusted to OD600 of 0.4, then diluted 100-fold in TSB containing 1.0% glucose. Aliquots of 100 μl culture were added to each well of the 96-well plate. Plantaricin GZ1-27 (100 μl) was added to the wells (final concentrations of 1/2× MIC and 1/4× MIC, respectively). Each concentration was replicated in eight wells and repeated in six plates for various intervals (12, 24, 36, 48, 60, and 72 h). The MRSA culture mixed with disodium hydrogen phosphate-citric acid buffer (100 μl) was used as a control. Subsequently, the plate was incubated at 37 °C without shaking. At each time point described above, biofilm mass was measured with crystal violet, as described (Du et al. 2016). Briefly, culture supernatant was discarded, plates were washed three times with PBS to remove unattached bacteria and surface-attached biofilms were fixed with methanol for 20 min. After drying at room temperature (25 °C), each well was stained with 200 μl of 1% (w/v) crystal violet solution for 15 min. Plates were then washed with PBS five times to remove unbound dye. Bound dye was re-dissolved in 200 μl of 95% (v/v) ethyl alcohol and absorbance measured at 595 nm using a microplate reader (BioTek Instruments, Winooski, VT).
SEM
Bacterial attachment of plantaricin GZ1-27-treated MRSA biofilm was assessed with acanning electron microscopy (SEM), as described (Loo et al. 2015). A log culture of S. aureus ATCC43300 (OD600 of 0.4) was diluted 100-fold in Nutrient Broth (NB) containing 0.8% glucose (Ju et al. 2018). Biofilm was formed on sterile glass coverslips (18 × 18 mm, Sail Brand, Yancheng, China) in a six-well plate (BOUANG, Haimen, China), with the coverslip placed in the bottom of each well. Then, NB diluted log culture (3.5 mL) was added to each well. In addition, 500 μl of plantaricin GZ1-27 were added to each well (final concentrations of 1/2× MIC and 1/4× MIC, respectively), with 500 μl of disodium hydrogen phosphate-citric acid buffer as control. After static incubation at 37 °C for 48 h, coverslips were washed three times with PBS and fixed overnight with 2.5% glutaraldehyde at 4 °C. After drying in ALPHA 1–4 LD plus vacuum freeze drier (Martin Christ, Osterode, Germany) for 24 h, coated samples were visualized using an EVO-LS10 SEM system (CARL ZEISS, Oberkochen, Germany).
Time-kill curve
Fresh MRSA culture reached to OD600 of 0.4 was diluted 100-fold in TSB containing 1.0% glucose. Aliquots of 200 μl culture were added per 96-well plate. The plates were incubated at 37 °C for 48 h to harvest mature biofilm. Biofilms were formed in four 96-well plates. Each plate was treated with disodium hydrogen phosphate-citric acid buffer, 2× MIC, 4× MIC and 8× MIC plantaricin GZ1-27 (200 μl), respectively. For each concentration, quintuple replication was included. At each time point (0, 1, 2, 4, 8, 12, 16, 20 and 24 h), wells were vigorously pipetted to detach the biofilm from the plate surface. Then, all biofilm cell suspension (200 μl) was collected, diluted with 1.8 mL of PBS, then serial 10-fold dilution and plated on LB plates. After incubation at 37 °C for 18 h, plates with ~ 30–300 CFU were enumerated. Alive biofilm cells were calculate using this equation: N*10*10n (n: dilution ratio. N: CFU in counting plate).
Confocal laser-scanning microscopy (CLSM)
We used CLSM to determine survival and distribution of plantaricin GZ1-27-treated biofilm cells on coverslip surfaces. The MRSA biofilm was allowed to be formed on coverslips in 6-well plates for 48 h, as described above. Then, the 48 h-biofilm was washed with PBS three times and air-dried for 10 min. The biofilm coverslip was then treated with plantaricin GZ1-27 in disodium hydrogen phosphate-citric acid buffer, at 2× MIC, 4× MIC, and 8× MIC at 37 °C for 1 h. The LIVE/DEAD BacLight Bacterial Viability Kit vital-staining probe mixture (Invitrogen, Carlsbad, CA) was used to monitor survival of plantaricin-treated biofilm. Briefly, after washing three times with PBS, the biofilm was dyed with a thoroughly mixed combination of nucleic acid binding stains SYTO®9 and propidium iodide (PI; 20 μL each). The SYTO®9 and propidium iodide (PI)-treated biofilm was incubated at room temperature (25 °C) in the dark for 15 min. After incubation, 3D images of biofilm were collected using a CLSM LEICA SP8 system (Leica, Solms, Germany) with excitation/emission wavelengths of 480/500 nm (SYTO®9) and 490/635 nm (PI). Living cells with intact membranes were stained green by SYTO®9, whereas dead cells appeared red, because PI enters into damaged-membranes cells and bind to DNA.
Statistical analysis
Microbiological data were expressed as mean ± standard deviation (SD) and figures prepared with Prism7 (GraphPad, San Diego, CA). Microbiological data were compared with one-way ANOVA, with Bonferroni multiple comparison tests used to locate differences. All statistical analyses were conducted using IBM SPSS Statistics 20.0 software (IBM, Armonk, NY).
Results
Growth of MRSA inhibited by plantaricin GZ1-27
To determine an appropriate concentration of GZ1-27 for alternating gene transcription and expression, log-phase cultures of MRSA were treated with plantaricin GZ1-27 at 5 levels of MICs (Fig. 1). After exposure to 1× MIC and 2× MIC of plantaricin GZ1-27, the OD600 of MRSA decreased to 0.315 ± 0.015 and 0.268 ± 0.019 within 30 min, respectively. However, compared to the control, there was no obvious inhibition (p > 0.05) of bacterial growth at 1/4× MIC, and 1/8× MIC of GZ1-27 at 30 min. When incubated with 1/2× MIC of GZ1-27, although growth of MRSA was inhibited within 3 h, MRSA still continually grew under GZ1-27 stress. This concentration reflected growth characteristics of cells stressed by plantaricin GZ1-27 (Yang et al. 2016). Thus, 1/2× MIC (16 μg/mL) of plantaricin GZ1-27 was selected for transcriptomic and proteomic analyses. After 3 h incubation, the optical density of treated bacteria became relatively stable (Fig. 1). Also, MRSA cells had sufficient time to produce proteins response to plantaricin GZ1-27, while minimizing the secondary unspecific effects. Thus, bacteria were harvested at 3 h after treatment with plantaricin GZ1-27.
Transcriptomic analysis on MRSA in response to plantaricin GZ1-27
After filtering raw data, high-quality clean data (error rate < 0.01%) were obtained. A total of 2766 genes were identified in plantaricin-treated and non-treated conditions, which corresponds to 81.5% of the genes of S. aureus subsp. aureus NCTC8325 (3393 genes in GenBank). Among identified genes, 1090 genes had a differential expression (padj < 0.05; Supplemental Table S2), with 552 genes up-regulated and 538 genes down-regulated in comparison to untreated MRSA cells (Fig. 2a).
Correlation analysis of proteome and transcriptome. a Transcriptomic results for differentially expressed genes (DEGs) with padj < 0.05. Red: up-regulated genes; blue: down-regulated genes. b Proteomic results for differentially expressed proteins (DEPs) with fold change >1 .2 and p < 0.05. Red: up-regulated proteins; blue: down-regulated proteins. c Venn diagram showing all identified, as well as all significantly enriched, mRNAs and proteins and their overlap. d Correlation between significantly enriched mRNAs and proteins. Red: both up-regulated items; blue: both down-regulated items; black: opposite changes
Functional analysis classified these DEGs into 84 significantly enriched GO terms (p < 0.05), with 55 terms corresponding to biological processes (BP), 24 corresponding to molecular functions (MF), and 10 corresponding to cellular components (CC). According to GO annotation (Fig. 3a), DEGs were scattered among various biological processes including single-organism process, small molecule metabolic process and oxidation-reduction. Predominant cellular components of DEGs were intracellular non-membrane-bounded organelles, whereas terms with high number of DGEs in cellular components were amino acid binding, oxidoreductase activity and O-acyltransferase activity, respectively. In addition, based on analysis of KEGG pathways, DEGs were involved in several metabolic pathways, mainly in biosynthesis of secondary metabolites, amino acids and amino acid metabolism, carbon metabolism, ABC transporters and purine metabolism (Fig. 3b).
Proteomic analyses on MRSA in response to plantaricin GZ1-27
To obtain more accurate information regarding biological systems, iTRAQ was used for measuring protein expression. All identified peptides were mainly distributed within 10 ppm, indicating that identified results were accurate and reliable. Compared to the control group, 418 DEPs (fold change > 1.2, p < 0.05) were identified (Supplemental Table S3), including 208 up-regulated proteins and 210 down-regulated proteins (Fig. 2b). According to GO annotation, DEPs were enriched (p < 0.05) in 78 GO terms. The top 20 GO terms for BP, MF, and CC are shown (Fig. 4a). The GO terms with a high rich factor of DEPs in BP, MF, and CC were related to inosine monophosphate (IMP) biosynthetic process, cell wall and protein complex binding, respectively. The KEGG pathway analysis predicted that regulated proteins were mainly involved in purine metabolism, ABC transporters, and arginine biosynthesis (Fig. 4b).
Comparative analysis of proteomic and transcriptomic data
Data sets of DEGs and DEPs were combined to assess MRSA responses to plantaricin GZ1-27. In the Venn diagram, there was a high consistency between the identified, as well as all significantly enriched genes and proteins (Fig. 2c). Of the 1499 proteins identified, only one was not observed at transcriptional level (2766). Among the 1090 DEGs and 418 DEPs, 228 gene items were significantly regulated at both transcriptional and translation levels under plantaricin GZ1-27 stress (Fig. 2c and Supplemental Table S4). Correlation analysis of the 228 items between mRNAs and proteins changes is shown (Fig. 2d). The trend of the change of 173 proteins (red dots in Fig. 2d) was the same as of mRNAs, with ~ 75.9% (173/228) concordance. Based on these analyses, protein translation was modestly coupled to gene transcription in MRSA responses to plantaricin GZ1-27.
Functional analyses of items regulated at both transcriptional and translation levels (hypothetical proteins were removed), suggested that plantaricin GZ1-27 not only has an important role in biological functions (biofilm formation, DNA replication and repair, and stress response); but also in metabolic pathways (purine metabolism, amino acid metabolism, and biosynthesis of secondary metabolites).
Fold changes of these genes and proteins were plotted on a heatmap (Fig. 5). Eight proteins (FnBPB, SdrC, SpA, IsdB, IcaR, SarA, SigB, and ArlR) were identified to be related to biofilm formation. In the presence of GZ1-27, expression of surface proteins (FnBPB, SpA, and IsdB), the well-known global transcriptional regulatory factors (SarA and SigB), surface protein (SdrC) and negative regulator (IcaR) was up-regulated, whereas expression of the two-component regulatory factor (ArlR) was down-regulated. Besides, plantaricin GZ1-27 downregulated expression of DNA repair-related proteins RecA, XerD, XseB, and AddA and also up-regulated replication-related proteins DnaA and DnaG. Heat-shock protein GrpE, McsB, HrcA, and Asp23 were up-regulated, whereas ClpC was down-regulated. Also, proteins of operon purABCD, including PurA, PurE, PurK, PurC, PurQ, PurF, PurM, PurN, PurH, PurD, and PurB, were strongly repressed by GZ1-27.
To investigate interactive relationships between the six groups, an enriched protein-based network was constructed using the STRING database (Fig. 6). Proteins with the same function were grouped in a circle and noted with same color. Purine metabolism was closely connected with amino acid metabolism, and linked with DNA repair and replication by RecA and DnaG. Heat shock was associated with biofilm formation, purine metabolism and amino acid metabolism via HrcA.
Plantaricin GZ1-27 inhibition of biofilm formation on polystyrene surface
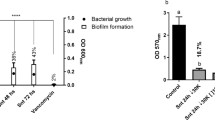
To verify if plantaricin GZ1-27-induced alternation of biofilm associated genes was capable of preventing MRSA from forming biofilm, a biofilm formation assay was conducted on a polystyrene surface. Without GZ1-27, biofilm mass continuously increased and peaked (OD595 = 2.258 ± 0.071) at 48 h (Fig. 7a). By contrast, less biofilm was formed in the presence of 1/2× MIC and 1/4× MIC GZ1-27, with biofilm mass decreased by 55.3% (1.249/2.258) and 40.2% (0.908/2.258) in the 1/2× MIC and 1/4× MIC group at the each time point at 48 h, respectively. Although after 48 h, the plantaricin-treated biofilm tended to be stable (p < 0.05), biofilm formation from both plantaricin-treated groups were still lower (p > 0.001) than in the control.
Effects of plantaricin GZ1-27 on biofilm formation. a Effect of plantaricin GZ1-27 on biofilm biomass within 72 h. Compared to Control, mean OD595 values at 1/2× MIC and 1/4× MIC in each time point decreased (p < 0.001), except at 1/4× MIC in 12 h (p < 0.01). Data were expressed as mean ± standard deviation (SD). The experiment was performed in quintuple replication. b–g SEM images of MRSA ATCC43300 biofilm cells at 48 h: b, c, d magnification × 1000; e, f, g magnification × 10,000; b, e control; c f incubated with 1/4× MIC plantaricin GZ1-27; d g incubated with 1/2× MIC plantaricin GZ1-27
Effects of plantaricin GZ1-27 on MRSA attachment on a glass surface was visualized under SEM. Without GZ1-27, MRSA formed a highly organized biofilm structure on a glass surface, comprised of numerous layers of densely concentrated cells (Fig. 7b). However, with GZ1-27, few MRSA biofilm structures were observed (Fig. 7c, d) and layered cell-to-cell connections were decreased, particularly with 1/2× MIC plantaricin GZ1-27, although few bacteria were laid as a grape-like structure, and most of the glass surface was devoid of cocci. Further, biofilm and cells surface ultrastructure from the control group were tightly connected by an extracellular polymeric substances (EPS) matrix (Fig. 7e), whereas, an EPS matrix was not clearly evident in GZ1-27-treated biofilm (Fig. 7f, g).
Time killing of plantaricin GZ1-27 on biofilm cells formed on polystyrene surfaces
To assess effectiveness of GZ1-27 in permeation and killing MRSA biofilm cells, mature biofilms (48 h) were incubated with plantaricin GZ1-27 at concentrations of 8× MIC, 4× MIC and 2× MIC (Fig. 8a). Without GZ1-27, MRSA biofilm cells remained at 8.73 log10 CFU over 24 h incubation. Overall, bactericidal activities against biofilm cells were observed for all three concentrations (p < 0.05). However, 8× MIC plantaricin GZ1-27 had the strongest killing effect, initiating reduction as soon as at 1 h (1.51 log10 CFU). With increasing duration of incubation, the killing effect continued and at 24 h, viable biofilm cells were not detectable. In the presence of 4× MIC plantaricin GZ1-27, the maximum reduction reached 5.91 log10 CFU at 24 h. Furthermore, 2× MIC plantaricin GZ1-27 was less effective in removing MRSA biofilm, enabling a maximum 2.1 log10 CFU reduction at 20 h.
Effects of plantaricin GZ1-27 on biofilm cells. a Time killing of plantaricin GZ1-27 on biofilm cells. Data were expressed as mean ± standard deviation (SD). The experiment was performed in quintuple replication. CLSM images of MRSA ATCC43300 biofilm treated with 2× MIC (c), 4× MIC (d), and 8× MIC (e) plantaricin GZ1-27
Survival of plantaricin GZ1-27-treated MRSA biofilms on glass surface
Biofilms were visualized using CLSM to examine survival and localization of MRSA biofilm with post-treatment of GZ1-27. In the absence of GZ1-27, a green biofilm completely covered the glass surface (Fig. 8a). After incubation with plantaricin GZ1-27, biofilms exhibited extensive bacterial death (red). And 8× MIC plantaricin GZ1-27 killed almost all of the biofilm cells at 1 h. Furthermore, with increased concentrations of GZ1-27, the proportion of dead cells was increased (Fig. 8c, d, e). These results were consistent with CFU counts of viable cells after incubation with plantaricin GZ1-27 (Fig. 8a). In addition, based on Z-stack images, dead cells were distributed in biofilms from top to base, demonstrating that plantaricin GZ1-27 could penetrate the S. aureus biofilm matrix and kill bottom cells.
Discussion
Based on transcriptome or proteome technology, several studies on MRSA responses to various antibiotics (e.g., cefotaxime (Brochmann et al. 2018), erythromycin derivative and oxacillin combination (Liu et al. 2016)) and natural extracts (lupeol and stigmasterol (Ibrahim and Yaacob 2017), rocket (Doulgeraki et al. 2017b), gambogic acid and neogambogic acid (Hua et al. 2019)) have been reported. This study was apparently the first to profile the MRSA transcriptome and proteome in response to a bacteriocin. The most prominent functional groups affected by plantaricin GZ1-27 were those with functions associated with biofilm formation, DNA replication and repair, heat-shock proteins, purine metabolism, amino acid metabolism, and biosynthesis of secondary metabolites.
Biofilm is initiated by bacteria attaching to abiotic and biotic surfaces, followed by cell adhesion and multiplication, leading to a mature biofilm (Lister and Horswill 2014). Biofilm formation is recognized as an important virulence factor of MRSA strains. Based on analyses of the transcriptome and proteome, plantaricin GZ1-27 suppressed the formation of surface proteins (FnBPB, SpA, IsdB, SdrC, ArlR) and polysaccharide intercellular adhesin PIA (IcaR, SarA, and SigB). Surface-associated proteins such as fibrinogen-binding proteins FnBPB, protein A (SpA), the serine-aspartate repeat protein SdrC, and iron-responsive surface determinants IsdB, play a role both in attachment and accumulation (Feuillie et al. 2017; Mazmanian et al. 2002). Specific homophilic interactions between these surface proteins represent an important mechanism of cell accumulation during biofilm formation. The two-component system ArlS–ArlR modifies the activity of extracellular serine protease which plays a role in biofilm maturation (Fournier et al. 2001). PIA is another major constituent of the biofilm matrix which is synthesized by enzymes encoded by the icaADBC operon. IcaR is a negative regulator of icaADBC, and global transcriptional regulatory factors accessory regulator A (SarA) and sigma B (SigB) are implicated in control of icaADBC (David et al. 2012; Handke et al. 2007). It is noteworthy that fnBPB, sdrC, spa, and isdB contribute to the formation of the extracellular matrix, whereas icaR, sarA, sigB, and arlR are regulatory factors of biofilm formation. Thus, we speculate that plantaricin GZ1-27 regulated formation of MRSA biofilm by inhibiting production of the extracellular matrix and supressing the activity of several regulatory factors. This assumption was supported by SEM images, with surface matrix being dramatically reduced in the treatment group. Similar results were reported for norgestimate, which inhibits MRSA biofilm formation by inhibiting production of polysaccharide intercellular adhesin and proteins in the extracellular matrix (Yoshii et al. 2017).
Meanwhile, based on crystal violet and SEM, biofilm formation of MRSA ATCC43300 was efficiently prevented by plantaricin GZ1-27 at 1/4 × and 1/2× MIC. However, 1/4 × MIC plantaricin GZ1-27 showed limited effect on the growth of MRSA. These results indicated that the inhibition of biofilm formation caused by plantaricin GZ1-27 was not related to the MRSA growth. This corresponds with the results of curcumin which suggested that the mechanism of biofilm inhibition was due to the prevention of biofilm formation process rather than the bactericidal effect (Loo et al. 2015). Furthermore, concentrated plantaricin GZ1-27 (4× and 8× MIC) penetrated the biofilm matrix and had sustained concentration-dependent bactericidal activity against MRSA biofilm cells (Fig. 8). Therefore, we inferred that the new bacteriocin plantaricin GZ1-27 is a potential candidate for S. aureus anti-biofilm therapy.
Repair of DNA is indispensable in maintaining genomic integrity and cell growth; consequently, bacterial enzymes responsible for DNA repair are targets for some antimicrobial agents. For example, lactobionic acid inhibited DNA repair in MRSA, acting via common DNA repair proteins, RuvB, UvrA, and MutS (Kang et al. 2019). In the present study, plantaricin GZ1-27 decreased expression of DNA repair, and homologous recombination-related proteins RecA, AddA, XerD, and XseB. DNA recombination protein RecA and helicase-nuclease AddA are important repair proteins in the repair of DNA double-strand break, where RecA catalyzes an ATP-dependent DNA strand-exchange reaction and AddA initiates the bi-directional degradation and resection of DNA ends (Badrinarayanan et al. 2017). In addition, efficient site-specific chromosomally encoded tyrosine recombinase XerD resolves the chromosome dimers back to monomers in homologous recombination process (Castillo et al. 2017). XseB is an important component of ssDNA-specific nucleases exonuclease VII (ExoVII) (Poleszak et al. 2012). The down-regulation of RecA, AddA, XerD, and XseB observed with plantaricin GZ1-27 treatment indicate that the DNA repair pathways were blocked by plantaricin GZ1-27, resulting in instability of the MRSA genome that leads to cell damage or death.
Furthermore, recombinase RecA is implicated as a universal drug target in pathogenic bacteria. RecA has a critical role in activation of the DNA damage response (SOS response), which associated with emergence of antimicrobial resistance (Žgur-Bertok 2013). In S. aureus, deletion of recA significantly reduced antibiotic-induced resistance (Singh et al. 2010). RecA provided new insights of mitigating drug resistance for treatment of MRSA infections (Pavlopoulou 2018). In our work, expression of RecA was decreased by plantaricin GZ1-27. We speculate that plantaricin GZ1-27 is a possible therapeutic strategy to suppress development of multi-resistance. Similar effect happened to a small-molecule inhibitor IMP-1700. It inhibited bacterial DNA repair mediated by AddAB/RecBCD protein complexes as a means to sensitize MRSA to DNA damage to the fluoroquinolone antibiotic ciprofloxacin (Lim et al. 2019). Further experiments are undergone to verify the suppression effect of plantaricin GZ1-27 on RecA expression, and whether GZ1-27 is able to impair drug resistance, survival and pathogenicity of MRSA.
Expression of certain heat-shock proteins (GrpE, McsB, HrcA, and Asp23) of MRSA ATCC43300 was induced by plantaricin GZ1-27. In parallel with our observation, enhanced expression of heat-shock proteins was observed when alpha-mangostin was used against Staphylococcus epidermidis RP62A (Clpb, Clpc, Groel, and Asp23) (Sivaranjani et al. 2019) and oxacillin against S. aureus RN450 (GroES) (Singh et al. 2001). We inferred that upregulation of heat-shock proteins may be an integral part of stress responses against sub-MIC antibacterial substances.
In addition to perturbation of biological functions, plantaricin GZ1-27 also targeted fundamental metabolic processes of MRSA, in particular, the purine pathway. In that regard, inosine monophosphate (IMP), a precursor of purine metabolism, is essential for biosynthesis of RNA and DNA. Almost all genes of operon purABCD, including purA, purE, purK, purC, purQ, purF, purM, purN, purH, purD, and purB were strongly suppressed by GZ1-27 (Fig. 6). Similarly, in response to mupirocin exposure, strong repression of the pur and pyr pathways occurred in S. aureus COL (Reiß et al. 2012). In addition, purine involves in enhancement resistance to stress. Mutants purB and purM had defective persistence compared to the parental S. aureus strain USA300 under multiple stress conditions, including various antibiotics (Yee et al. 2015), with potential as a new combination therapy. The inhibition of purine metabolism by plantaricin GZ1-2 may hinder the synthesis of genetic material and reduce MRSA’s resistance to environmental stress.
Based on transcriptomic and proteomic data, multiple amino acid metabolisms and the biosynthesis of secondary metabolites were affected by plantaricin GZ1-27, respectively. However, expressions of some genes (e.g., argH, argG, ilvC, mtnN, and dapH) in transcriptome analysis were not consistent with the proteome. This apparent discrepancy may reflect limitations of transcriptome and proteomic technology, e.g., lower sensitivity of the proteomic analysis. Importantly, divergent efficiencies of transcriptional and translational levels in MRSA ATCC43300 may have caused post-translational turnover and modifications (Darby et al. 2014; Petersen et al. 2015).
In conclusion, plantaricin GZ1-27 may prevent biofilm formation of MRSA by inhibiting production of surface matrix-associated proteins and restraining functions of regulatory factors. In addition, plantaricin GZ1-27 were capable of disturbing multiple targets, including DNA repair, purine pathways, biosynthesis pathways for amino acid and secondary metabolites, which are essential for cell synthesis. This study advanced current knowledge regarding molecular mechanisms of anti-MRSA of plantaricin and provided mechanistic principles for development of plantaricin-based antimicrobials.
References
Álvarez A, Fernández L, Gutiérrez D, Iglesias B, Rodríguez A, García P (2019) Methicillin-resistant Staphylococcus aureus (MRSA) in hospitals: latest trends and treatments based on bacteriophages. J Clin Microbiol 57(2):e01006–e01019. https://doi.org/10.1128/JCM.01006-19
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31(2):166–169. https://doi.org/10.1093/bioinformatics/btu638
Ansari A, Zohra RR, Tarar OM, Qader SAU, Aman A (2018) Screening, purification and characterization of thermostable, protease resistant Bacteriocin active against methicillin resistant Staphylococcus aureus (MRSA). BMC Microbiol 18(1):1–10. https://doi.org/10.1186/s12866-018-1337-y
Badrinarayanan A, Le TB, Spille J-H, Cisse II, Laub MT (2017) Global analysis of double-strand break processing reveals in vivo properties of the helicase-nuclease complex AddAB. PLoS Genet 13(5):e1006783. https://doi.org/10.1371/journal.pgen.1006783
Bazargani MM, Rohloff J (2016) Antibiofilm activity of essential oils and plant extracts against Staphylococcus aureus and Escherichia coli biofilms. Food Control 61:156–164. https://doi.org/10.1016/j.foodcont.2015.09.036
Brochmann P, Hesketh A, Jana B, Brodersen G, Guardabassi L (2018) Transcriptome analysis of extended-spectrum β-lactamase-producing Escherichia coli and methicillin-resistant Staphylococcus aureus exposed to cefotaxime. Sci Rep 8(1):1–11. https://doi.org/10.1038/s41598-018-34191-3
Casey TM, Khan JM, Bringans SD, Koudelka T, Takle PS, Downs RA, Livk A, Syme RA, Tan K-C, Lipscombe RJ (2017) Analysis of reproducibility of proteome coverage and quantitation using isobaric mass tags (iTRAQ and TMT). J Proteome Res 16(2):384–392. https://doi.org/10.1021/acs.jproteome.5b01154
Castillo F, Benmohamed A, Szatmari G (2017) Xer site specific recombination: double and single recombinase systems. Front Microbiol 8:1–18. https://doi.org/10.3389/fmicb.2017.00453
Cotter PD, Ross RP, Hill C (2013) Bacteriocins—a viable alternative to antibiotics? Nat Rev Microbiol 11(2):95–105. https://doi.org/10.1038/nrmicro2937
Darby AC, Gill AC, Armstrong SD, Hartley CS, Xia D, Wastling JM, Makepeace BL (2014) Integrated transcriptomic and proteomic analysis of the global response of Wolbachia to doxycycline-induced stress. ISME J 8(4):925–937. https://doi.org/10.1038/ismej.2013.192
David C, Lei MG, Lee CY (2012) Genetic regulation of the intercellular adhesion locus in staphylococci. Front Cell Infect Microbiol 2:1–13. https://doi.org/10.3389/fcimb.2012.00038
Del Pozo JL, Patel R (2007) The challenge of treating biofilm-associated bacterial infections. Clin Pharmacol Ther 82(2):204–209. https://doi.org/10.1038/sj.clpt.6100247
Deng W, Wang Y, Liu Z, Cheng H, Xue Y (2014) HemI: a toolkit for illustrating heatmaps. PLoS One 9(11):e111988. https://doi.org/10.1371/journal.pone.0111988
Doulgeraki AI, Di Ciccio P, Ianieri A, Nychas GE (2017a) Methicillin-resistant food-related Staphylococcus aureus: a review of current knowledge and biofilm formation for future studies and applications. Res Microbiol 168(1):1–15. https://doi.org/10.1016/j.resmic.2016.08.001
Doulgeraki AI, Efthimiou G, Paramithiotis S, Pappas KM, Typas MA, Nychas G-J (2017b) Effect of rocket (Eruca sativa) extract on MRSA growth and proteome: metabolic adjustments in plant-based media. Front Microbiol 8:1–9. https://doi.org/10.3389/fmicb.2017.00782
Doyle ME, Hartmann FA, Lee Wong AC (2012) Methicillin-resistant staphylococci: implications for our food supply? Anim Health Res Rev 13(2):157–180. https://doi.org/10.1017/S1466252312000187
Du H, Li X, Lu Z, Bie X, Zhao H, Zhang C, Lu F (2019) Effect of plantaricin 163 in combination with thymol and surfactin on Crucian carp (Carassius auratus). J Food Prot 82(8):1283–1291. https://doi.org/10.4315/0362-028X.JFP-18-439
Du H, Pang M, Dong Y, Wu Y, Wang N, Liu J, Awan F, Lu C, Liu Y (2016) Identification and characterization of an Aeromonas hydrophila oligopeptidase gene pepF negatively related to biofilm formation. Front Microbiol 7:1–12. https://doi.org/10.3389/fmicb.2016.01497
Du H, Yang J, Lu X, Lu Z, Bie X, Zhao H, Zhang C, Lu F (2018) Purification, characterization, and mode of action of plantaricin GZ1-27, a novel bacteriocin against Bacillus cereus. J Agric Food Chem 66(18):4716–4724. https://doi.org/10.1021/acs.jafc.8b01124
Ek SK, Gur-Bertok D (2013) Global transcriptional responses to the bacteriocin colicin M in Escherichia coli. BMC Microbiol 13(1):1–13. https://doi.org/10.1186/1471-2180-13-42
Feuillie C, Formosa-Dague C, Hays LM, Vervaeck O, Derclaye S, Brennan MP, Foster TJ, Geoghegan JA, Dufrêne YF (2017) Molecular interactions and inhibition of the staphylococcal biofilm-forming protein SdrC. Proc Natl Acad Sci U S A 114(14):3738–3743. https://doi.org/10.1073/pnas.1616805114
Fournier B, Klier A, Rapoport G (2001) The two-component system ArlS-ArlR is a regulator of virulence gene expression in Staphylococcus aureus. Mol Microbiol 41(1):247–261. https://doi.org/10.1046/j.1365-2958.2001.02515.x
Han J, Gao P, Zhao S, Bie X, Lu Z, Zhang C, Lv F (2017) iTRAQ-based proteomic analysis of LI-F type peptides produced by Paenibacillus polymyxa JSa-9 mode of action against Bacillus cereus. J Proteome 150:130–140. https://doi.org/10.1016/j.jprot.2016.08.019
Handke LD, Slater SR, Conlon KM, O'Donnell ST, Olson ME, Bryant KA, Rupp ME, O'Gara JP, Fey PD (2007) σB and SarA independently regulate polysaccharide intercellular adhesin production in Staphylococcus epidermidis. Can J Microbiol 53(1):82–91. https://doi.org/10.1139/w06-108
Hansen ME, Wangari R, Hansen EB, Mijakovic I, Jensen PR (2009) Engineering of Bacillus subtilis 168 for increased nisin resistance. Appl Environ Microbiol 75(21):6688–6695. https://doi.org/10.1128/AEM.00943-09
Hong J, Kim J, Quan LH, Heu S, Roh E (2018) Purification and characterization of pasteuricin produced by Staphylococcus pasteuri RSP-1 and active against multidrug-resistant Staphylococcus aureus. J Food Prot 81(11):1768–1775. https://doi.org/10.4315/0362-028x.Jfp-18-111
Hua X, Jia Y, Yang Q, Zhang W, Dong Z, Liu S (2019) Transcriptional analysis of the effects of gambogic acid and neogambogic acid on methicillin-resistant Staphylococcus aureus. Front Pharmacol 10:1–14. https://doi.org/10.3389/fphar.2019.00986
Ibrahim N, Yaacob WA (2017) Transcriptome analysis of methicillin-resistant Staphylococcus aureus in response to stigmasterol and lupeol. J Glob Antimicrob Resist 8:48–54. https://doi.org/10.1016/j.jgar.2016.10.006
Jevons MP (1961) “Celbenin”-resistant staphylococci. Br Med J 1(5219):124–125. https://doi.org/10.1136/jcp.14.4.385
Ju X, Li J, Zhu M, Lu Z, Lv F, Zhu X, Bie X (2018) Effect of the luxS gene on biofilm formation and antibiotic resistance by Salmonella serovar Dublin. Food Res Int 107:385–393. https://doi.org/10.1016/j.foodres.2018.02.039
Kang S, Kong F, Liang X, Li M, Yang N, Cao X, Yang M, Tao D, Yue X, Zheng Y (2019) Label-free quantitative proteomics reveals the multitargeted antibacterial mechanisms of lactobionic acid against methicillin-resistant Staphylococcus aureus (MRSA) using SWATH-MS technology. J Agric Food Chem 67(44):12322–12332. https://doi.org/10.1021/acs.jafc.9b06364
Langmead B, Salzberg S (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. https://doi.org/10.1038/nmeth.1923
Lim CSQ, Ha KP, Clarke RS, Gavin L-A, Cook DT, Hutton JA, Sutherell CL, Edwards AM, Evans LE, Tate EW, Lanyon-Hogg T (2019) Identification of a potent small-molecule inhibitor of bacterial DNA repair that potentiates quinolone antibiotic activity in methicillin-resistant Staphylococcus aureus. Biorg Med Chem 27(20):1–6. https://doi.org/10.1016/j.bmc.2019.06.025
Lister JL, Horswill AR (2014) Staphylococcus aureus biofilms: recent developments in biofilm dispersal. Front Cell Infect Microbiol 4:1–9. https://doi.org/10.3389/fcimb.2014.00178
Liu X, Pai P-J, Zhang W, Hu Y, Dong X, P-y Q, Chen D, Lam H (2016) Proteomic response of methicillin-resistant S. aureus to a synergistic antibacterial drug combination: a novel erythromycin derivative and oxacillin. Sci Rep 6:1–12. https://doi.org/10.1038/srep19841
Loo C-Y, Rohanizadeh R, Young PM, Traini D, Cavaliere R, Whitchurch CB, Lee W-H (2015) Combination of silver nanoparticles and curcumin nanoparticles for enhanced anti-biofilm activities. J Agric Food Chem 64(12):2513–2522. https://doi.org/10.1021/acs.jafc.5b04559
Mazmanian SK, Ton-That H, Su K, Schneewind O (2002) An iron-regulated sortase anchors a class of surface protein during Staphylococcus aureus pathogenesis. Proc Natl Acad Sci U S A 99(4):2293–2298. https://doi.org/10.1073/pnas.032523999
Okuda K (2013) Effects of bacteriocins on methicillin-resistant Staphylococcus aureus biofilm. Antimicrob Agents Chemother 57(11):5572–5579. https://doi.org/10.1128/AAC.00888-13
Pavlopoulou A (2018) RecA: a universal drug target in pathogenic bacteria. Front Biosci 23:36–42. https://doi.org/10.2741/4580
Petersen HO, Höger SK, Looso M, Lengfeld T, Kuhn A, Warnken U, Nishimiya-Fujisawa C, Schnölzer M, Krüger M, Özbek S (2015) A comprehensive transcriptomic and proteomic analysis of hydra head regeneration. Mol Biol Evol 32(8):1928–1947. https://doi.org/10.1093/molbev/msv079
Poleszak K, Kaminska KH, Dunin-Horkawicz S, Lupas A, Skowronek KJ, Bujnicki JM (2012) Delineation of structural domains and identification of functionally important residues in DNA repair enzyme exonuclease VII. Nucleic Acids Res 40(16):8163–8174. https://doi.org/10.1093/nar/gks547
Reiß S, Pané-Farré J, Fuchs S, François P, Liebeke M, Schrenzel J, Lindequist U, Lalk M, Wolz C, Hecker M (2012) Global analysis of the Staphylococcus aureus response to mupirocin. Antimicrob Agents Chemother 56(2):787–804. https://doi.org/10.1128/AAC.05363-11
Rossolini GM, Arena F, Pecile P, Pollini S (2014) Update on the antibiotic resistance crisis. Curr Opin Pharmacol 18:56–60. https://doi.org/10.1016/j.coph.2014.09.006
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Silva CCG, Silva SPM, Ribeiro SC (2018) Application of bacteriocins and protective cultures in dairy food preservation. Front Microbiol 9:1–15. https://doi.org/10.3389/fmicb.2018.00594
Singh R, Ledesma KR, Chang K-T, Tam VH (2010) Impact of recA on levofloxacin exposure-related resistance development. Antimicrob Agents Chemother 54(10):4262–4268. https://doi.org/10.1128/AAC.00168-10
Singh VK, Jayaswal R, Wilkinson BJ (2001) Cell wall-active antibiotic induced proteins of Staphylococcus aureus identified using a proteomic approach. FEMS Microbiol Lett 199(1):79–84. https://doi.org/10.1111/j.1574-6968.2001.tb10654.x
Sivaranjani M, Leskinen K, Aravindraja C, Saavalainen P, Pandian SK, Skurnik M, Ravi AV (2019) Deciphering the antibacterial mode of action of alpha-mangostin on Staphylococcus epidermidis RP62A through an integrated transcriptomic and proteomic approach. Front Microbiol 10:1–16. https://doi.org/10.3389/fmicb.2019.00150
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, von Mering C (2017) The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45(D1):D362–D368. https://doi.org/10.1093/nar/gkw937
Thomsen TT, Mojsoska B, Cruz JC, Donadio S, Jenssen H, Lobner-Olesen A, Rewitz K (2016) The lantibiotic NAI-107 efficiently rescues Drosophila melanogaster from infection with methicillin-resistant Staphylococcus aureus USA300. Antimicrob Agents Chemother 60(9):5427–5436. https://doi.org/10.1128/aac.02965-15
Titouche Y, Hakem A, Houali K, Meheut T, Vingadassalon N, Ruiz-Ripa L, Salmi D, Chergui A, Chenouf N, Hennekinne (2019) Emergence of methicillin-resistant Staphylococcus aureus (MRSA) ST8 in raw milk and traditional dairy products in the Tizi Ouzou area of Algeria. J Dairy Sci 102:6876–6884. https://doi.org/10.3168/jds.2018-16208
Vergara A, Normanno G, Di Ciccio P, Pedonese F, Nuvoloni R, Parisi A, Santagada G, Colagiorgi A, Zanardi E, Ghidini S (2017) Biofilm formation and its relationship with the molecular characteristics of food-related methicillin-resistant Staphylococcus aureus (MRSA). J Food Sci 82(10):2364–2370. https://doi.org/10.1111/1750-3841.13846
Wendlandt S, Schwarz S, Silley P (2013) Methicillin-resistant Staphylococcus aureus: a food-borne pathogen? Annu Rev Food Sci Technol 4:117–139. https://doi.org/10.1146/annurev-food-030212-182653
Wenzel M, Kohl B, Münch D, Raatschen N, Albada HB, Hamoen L, Metzler-Nolte N, Sahl H-G, Bandow J (2012) Proteomic response of Bacillus subtilis to lantibiotics reflects differences in interaction with the cytoplasmic membrane. Antimicrob Agents Chemother 56(11):5749–5757. https://doi.org/10.1128/AAC.01380-12
Wiese S, Reidegeld KA, Meyer HE, Warscheid B (2007) Protein labeling by iTRAQ: a new tool for quantitative mass spectrometry in proteome research. J Proteome 7(3):340–350. https://doi.org/10.1002/pmic.200600422
Wiśniewski JR, Zougman A, Nagaraj N, Mann M (2009) Universal sample preparation method for proteome analysis. Nat Methods 6(5):359–362. https://doi.org/10.1038/nmeth.1322
Yang X-Y, Shi T, Du G, Liu W, Yin X-F, Sun X, Pan Y, He Q-Y (2016) iTRAQ-based proteomics revealed the bactericidal mechanism of sodium new houttuyfonate against Streptococcus pneumoniae. J Agric Food Chem 64(32):6375–6382. https://doi.org/10.1021/acs.jafc.6b02147
Yee R, Cui P, Shi W, Feng J, Zhang Y (2015) Genetic screen reveals the role of purine metabolism in Staphylococcus aureus persistence to rifampicin. Antibiotics 4(4):627–642. https://doi.org/10.3390/antibiotics4040627
Yoshii Y, Okuda KI, Yamada S, Nagakura M, Sugimoto S, Nagano T, Okabe T, Kojima H, Iwamoto T, Kuwano K, Mizunoe Y (2017) Norgestimate inhibits staphylococcal biofilm formation and resensitizes methicillin-resistant Staphylococcus aureus to beta-lactam antibiotics. NPJ Biofilms Microbiol 3:1–9. https://doi.org/10.1038/s41522-017-0026-1
Žgur-Bertok D (2013) DNA damage repair and bacterial pathogens. PLoS Pathog 9(11):e1003711. https://doi.org/10.1371/journal.ppat.1003711
Zhao S, Han J, Bie X, Lu Z, Zhang C, Lv F (2016) Purification and characterization of plantaricin JLA-9: a novel bacteriocin against Bacillus spp. produced by Lactobacillus plantarum JLA-9 from Suan-Tsai, a traditional Chinese fermented cabbage. J Agric Food Chem 64(13):2754–2764. https://doi.org/10.1021/acs.jafc.5b05717
Zhu X, Zhao Y, Sun Y, Gu Q (2014) Purification and characterisation of plantaricin ZJ008, a novel bacteriocin against Staphylococcus spp. from Lactobacillus plantarum ZJ008. Food Chem 165(3):216–223. https://doi.org/10.1016/j.foodchem.2014.05.034
Acknowledgments
Appreciation is expressed to Dr. John Kastelic (University of Calgary) for manuscript editing.
Funding
This study is financially supported by the National Research Program of China (No. 2015BAD16B04).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
This research did not involve any human participants.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Du, H., Zhou, L., Lu, Z. et al. Transcriptomic and proteomic profiling response of methicillin-resistant Staphylococcus aureus (MRSA) to a novel bacteriocin, plantaricin GZ1-27 and its inhibition of biofilm formation. Appl Microbiol Biotechnol 104, 7957–7970 (2020). https://doi.org/10.1007/s00253-020-10589-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10589-w