Abstract
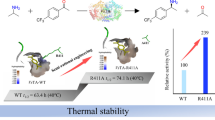
(R)-Selective ω-transaminase (ω-TA) is a key enzyme for the asymmetric reductive amination of carbonyl compounds to produce chiral amines which are essential parts of many therapeutic compounds. However, its practical industrial applications are hindered by the low catalytic efficiency and poor thermostability of naturally occurring enzymes. In this work, we report the molecular modification of (R)-selective ω-TA from Aspergillus terreus (AtTA) to allow asymmetric reductive amination of 4-hydroxy-2-butanone, producing (R)-3-amino-1-butanol. Based on substrate docking analysis, 4 residues in the substrate tunnel and binding pocket of AtTA were selected as mutation hotspots. The screening procedure was facilitated by the construction of a “small-intelligent” library and the use of thin-layer chromatography for preliminary screening. The resulting mutant AtTA-M5 exhibited a 9.6-fold higher kcat/Km value and 9.4 °C higher \( {\mathrm{T}}_{1/2}^{10\min } \) than that of wild-type AtTA. Furthermore, the conversion of 20 and 50 g L−1 4-hydroxy-2-butanone by AtTA-M5 reached 90.8% and 79.1%, suggesting significant potential for production of (R)-3-amino-1-butanol. Under the same conditions, wild-type AtTA achieved less than 5% conversion. Moreover, the key mutation (S215P in AtTA) was validated in 7 other (R)-selective ω-TAs, indicating its general applicability in improving the catalytic efficiency of homologous (R)-selective ω-TAs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, approximately 40% of the new chemical entities (NCEs) approved by the US Food and Drug Administration (FDA) have contained one or more chiral amine moieties (Aleku et al. 2017). As an important chiral building block, the synthesis of chiral amines has attracted much attention, and asymmetric reductive amination has emerged as one of the most powerful synthetic methods (Chen et al. 2018; Huang et al. 2016). Compared with traditional chemical synthesis, ω-transaminase (ω-TA)-catalyzed asymmetric reductive amination of prochiral ketones has the advantages of mild reaction conditions, high enantioselectivity, and environmental friendliness. The atom utilization can reach 100%, in line with good practices of “atomic economy” and “green chemistry.” Therefore, it has risen in popularity as an efficient method to synthesize chiral amines (Gomm and O’Reilly 2018; Guo and Berglund 2017; Sheldon and Pereira 2017). ω-TAs belong to the family of pyridoxal-5-phosphate (PLP)-dependent transferases, which consist of (S)-selective and (R)-selective types that can be used to synthesize chiral amines with different configurations (Mathew and Yun 2012; Planchestainer et al. 2019) .
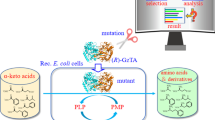
(R)-3-Amino-1-butanol is a crucial chiral intermediate for the synthesis of the new anti-HIV drug dolutegravir, which is a second-generation integrase strand transfer inhibitor developed by GSK (Johns et al. 2013). The sales of dolutegravir products reached £4.42 billion in 2018 and are expected to continue to increase. Due to the huge market demand, there is an urgent need for the development of an efficient and green method for synthesis of the chiral intermediate (R)-3-amino-1-butanol. (R)-Selective ω-TAs can catalyze the asymmetric reductive amination of 4-hydroxy-2-butanone to deliver (R)-3-amino-1-butanol, which has a significant potential to address this market need.
Before 2010, only the amino acid sequence of (R)-selective ω-TA from Arthrobacter sp. was publicly available, as described in a patent (Yamada et al. 2001). In 2010, 17 (R)-selective ω-TAs from different microbes were identified based on information about catalytic mechanism, protein structure, and sequence alignment (Höhne et al. 2010). To date, more than 25 (R)-selective ω-TAs from different microbes have been reported (Jiang et al. 2015; Kelly et al. 2019; Kim et al. 2018; Telzerow et al. 2018). Nevertheless, enzymes used for large-scale manufacturing of chiral amines will always suffer harsh conditions, such as non-natural substrates, high substrate or product concentrations, and pH or temperatures beyond physiological conditions (Sheldon and Pereira 2017). Accordingly, very few naturally occurring (R)-selective ω-TAs have been reported for industrial-scale production of chiral amines. In 2018, the process for preparation of (R)-3-amino-1-butanol (with a maximum substrate concentration of 12 g L−1) using (R)-selective ω-TAs was described in a patent. The enzymes used in that invention were obtained commercially, and the amino acid sequences have not yet been disclosed (Khatik and Husain 2018). Recently, a novel (R)-selective ω-TA from Actinobacteria sp. was used for the biosynthesis of (R)-3-amino-1-butanol, and a substrate fed-batch strategy was applied to solve the bottleneck of poor substrate tolerance for the naturally occurring enzyme. The maximum yield reached 29.6 g L−1 after the optimization of reaction conditions (Tang et al. 2019). However, in order to meet the needs of industrial applications, a strategy other than optimization of reaction conditions will need to further improve the catalytic efficiency of naturally occurring enzymes.
Along with the advances in protein engineering technologies, modified (R)-selective ω-TAs have emerged as a promising and environmentally friendly catalyst used for practical applications. The most outstanding example is the efficient chiral amine asymmetric synthesis by a modified (R)-selective ω-TA from Arthrobacter sp. to manufacture the anti-diabetic compound sitagliptin, which won the Presidential Green Chemistry Award Challenge in 2010 (Desai 2011; Savile et al. 2010). In addition, efforts aiming to improve the thermostability of (R)-selective ω-TA have been advanced through in silico design (Huang et al. 2017; Xie et al. 2019). Herein, we report the molecular modification of (R)-selective ω-TA from Aspergillus terreus (AtTA) for asymmetric reductive amination of 4-hydroxy-2-butanone to deliver (R)-3-amino-1-butanol. Through testing the minimum mutants with mutations in the substrate tunnel and binding pocket, we obtained evolved AtTAs with high catalytic efficiency, good thermostability, and excellent enantioselectivity toward 4-hydroxy-2-butanone. In addition, we also identified the key mutation (S215P) that could be crucial for improving the catalytic efficiency of other homologous (R)-selective ω-TAs.
Materials and methods
Strains, plasmids, and chemical reagents
The genes encoding 8 wild-type (WT) ω-TAs were synthesized by GenScript Biotech Co., Ltd. (Nanjing, China), enzymes that have been previously overexpressed in Escherichia coli BL21(DE3) cells using plasmid pET-28a as an expression vector. Oligonucleotide primers designed for mutagenesis were synthesized by GenScript Biotech Co., Ltd. (Nanjing, China). 4-Hydroxy-2-butanone, (R)-3-amino-1-butanol, and 3-amino-1-butanol were purchased from Makclin Biochemical Co., Ltd. (Shanghai, China). Other chemical reagents were obtained commercially.
Computational methods
Protein modeling was carried out with EasyModeller 4.0 to build a homology model of the mutant AtTA using the crystal structure of AtTA (PDB ID: 4CE5) as a template. All docking calculations were performed using AutoDock 4.2 with the docking algorithm of ligand flexibility and protein rigidity. The structure of the quinonoid intermediate was constructed with ChemBioDraw 12.0 and used as the ligand. The structures of AtTA and its mutant were used as the receptor, respectively. Analysis of the protein-ligand complex and stereo views was accomplished using Pymol.
Library construction
The residues His55, Gly126, Phe115, and Ser215 were selected to create single-site saturation mutant libraries. With the use of whole plasmid PCR, saturation mutagenesis at these positions was carried out using the primers listed in Table S1 with NNK degeneracy. The plasmid pET-28a, carrying the gene encoding AtTA, was used as the template. The reaction mixture (50 μL) contained 1 μL of DNA template (20–40 ng·μL−1), 1 μL each of forward and reverse primers (100 μM), 10 μL of 5 × PCR buffer, 4 μL of dNTPs (2.5 mM), 0.5 μL of PrimeSTAR DNA polymerase (2.5 U μL−1), and 32.5 μL ddH2O. The PCR program was as follows: 18 cycles of 20 s at 98 °C and 7 min at 68 °C. The PCR products were digested for 5 h at 37 °C with Dpn I to remove the parent plasmid. Then, the reaction mixtures were transformed into E. coli BL21 (DE3) to create the variant library for screening.
Overlap extension PCR was used to construct the gene containing mutations in 3 different positions. According to the results of single-site saturation mutagenesis, the primers were designed with degenerate codons using MDC-Analyzer (Table S1). The plasmid pET-28a, carrying the gene encoding AtTA, was used as a template. The reaction mixture (50 μL) contained 1 μL of DNA template (20–40 ng μL−1), 1 μL each of forward and reverse primers (100 μM), 10 μL of 5 × PCR buffer, 4 μL of dNTPs (2.5 mM), 0.5 μL of PrimeSTAR DNA polymerase (2.5 U μL−1), and 32.5 μL of ddH2O. The forward and reverse primers used here were AtTA-F and 55H-R, 55H-F and 126G-R, 126G-F and 215S-R, and 215S-F and AtTA-R, respectively. The PCR program was as follows: 30 cycles; pre-denaturation at 95 °C for 5 min, denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s, extension at 72 °C for 30 s. Four different PCR products were purified and used as the templates to amplify the target gene encoding AtTA with different mutations. The reaction mixture (50 μL) contained 1 μL each of 4 different PCR products (20–40 ng μL−1), 1 μL forward primer (AtTA-F, 100 μM), 1 μL reverse primers (AtTA-R, 100 μM), 10 μL 5 × PCR buffer, 4 μL dNTPs (2.5 mM), 0.5 μL PrimeSTAR DNA polymerase (2.5 U μL−1), and 32.5 μL ddH2O. The PCR program was as follows: pre-denaturation at 95 °C for 5 min, 10 cycles of 30 s at 95 °C, 30 s at 55 °C, and 1 min at 72 °C (without primers); 20 cycles of 30 s at 95 °C, 30 s at 55 °C, and 1 min at 72 °C (with primers). After purification, the resulting product was digested with NdeI and HindIII, ligated with pET-28a (linearized with NdeI and HindIII), and then transformed into E. coli BL21 (DE3) to create the variant library for screening.
Screening procedures
Mono-colonies were picked and inoculated into 96 deep well plates containing 500 μL lysogeny broth (LB) medium with kanamycin (50 μg mL−1) per well and cultivated for 12 h at 37 °C on a shaker set to 200 rpm. The colony expressing WT AtTA was used as the control. Then, 80 μL of each culture was transferred into a new 96 deep well plates containing 720 μL LB medium with kanamycin (50 mg mL−1) and incubated for 2 h at 37 °C and 200 rpm. About 0.5 mM IPTG (final concentration) was added and induced at 26 °C and 200 rpm for another 12 h. The cultures were centrifuged at 3000×g for 5 min to pellet the library of variants at the bottom of plates. The plates were frozen at − 80 °C for at least 1 h. After melting at room temperature, 300 μL disruption buffer (pH 7.5, 50 mM potassium phosphate buffer; 750 mg L−1 lysozyme; and 10 mg L−1 DNA nuclease) was added into each well and mixed for 5 min, then incubated at 37 °C for 1 h. After centrifugation at 3000×g for 20 min, the resulting supernatants (crude extracts) were transferred into new deep well plates for reaction with 4-hydroxy-2-butanone (60 mM and 120 mM), PLP (1 mM), and isopropylamine hydrochloride (300 mM and 600 mM) at 30 °C and 500 rpm for 24 h. The final volume in each well was increased to 500 μL with the addition of pH 7.5, 50 mM potassium phosphate buffer.
Thin-layer chromatography was chosen for preliminary screening with ethyl acetate-methanol-ammonia (45:54:1, v/v/v) as the developing solvent of and a mixture of KMnO4–Na2CO3 as the chromogenic reagent. Compared with WT AtTA, those mutants that exhibited similar or higher conversion were selected for further study (Fig. S1). The product and remaining substrate in the reaction mixture were extracted using an equal volume of ethyl acetate for further determination via GC analysis (Fig. S2).
Protein expression and purification
E. coli BL21 (DE3) cells containing the recombinant plasmid were cultivated in LB medium containing kanamycin (50 μg mL−1) at 37 °C. When the optical density reached 0.8, IPTG was added to a final concentration of 0.1 mM, and then was incubated at 26 °C for 12 h to induce the expression of AtTA. After centrifugation at 6000×g for 10 min at 4 °C, the cells were resuspended in buffer A (pH 7.5, 50 mM potassium phosphate buffer, 0.3 M NaCl, 20 mM imidazole, 0.1 mM PLP) and lysed by sonication at 4 °C. The supernatant was collected by centrifugation at 12,000×g for 30 min at 4 °C. Recombinant enzymes with His-tags at the N terminus were purified using affinity chromatography with a HisTrap™ FF crude column (GE Healthcare) according to the manufacturer’s instructions. Purified enzymes were eluted in buffer B (pH 7.5, 50 mM potassium phosphate buffer, 0.3 M NaCl, 300 mM imidazole, 0.1 mM PLP), desalted with a HiPrep 26/10 Desalting column (GE Healthcare) with 50 mM potassium phosphate buffer (pH 7.5), and analyzed by SDS-PAGE (Fig. S3). The protein concentration was measured by the Bradford method.
Determination of enzyme activity, conversion, and enantioselectivity
Enzyme activity was determined by measuring the amount of (R)-3-amino-1-butanol generated from 4-hydroxy-2-butanone. The reaction mixture (pH 8.0, 0.5 mL) contained 5 mM 4-hydroxy-2-butanone, 1 mM PLP, 10 mM isopropylamine hydrochloride, and an appropriate amount of purified enzyme. The reaction was performed at 30 °C for 5 min and then terminated by the addition of 0.5 mL acetonitrile. The amount of (R)-3-amino-1-butanol formed was determined by GC and extrapolated from a standard curve. One unit of enzyme activity was defined as the amount of enzyme that released 1 μmol of (R)-3-amino-1-butanol per minute. Enantioselectivity was determined by HPLC with precolumn derivatization using 1-fluro-2,4-dinitrobenzene (DNFB). The chromatographic conditions are listed in Table S2. The conversion of acetophenone was determined by GC as previously described (Mutti et al. 2011).
Characterization of AtTA and its mutants
The kinetic parameters were obtained by measuring the initial velocities of the enzymic reactions and curve-fitting according to the Michaelis–Menten equation. In experiments determining the Km of 4-hydroxy-2-butanone, the substrate concentrations were ranged from 0.1 to 5 mM with a fixed concentration of isopropylamine (10 mM). When determining the Km for isopropylamine, the substrate concentrations ranged from 0.1 to 10 mM with a fixed concentration of 4-hydroxy-2-butanone (5 mM).
To detect thermal inactivation, purified AtTA and its mutants (0.1 mg·mL−1) were incubated in the range of 20–60 °C for 10 min; after which, the enzymes were immediately placed on ice for 10 min to decrease the temperature. The residual activity was recorded as a percentage of the original activity. The \( {\mathrm{T}}_{1/2}^{10\mathit{\min}} \) value was defined as the temperature at which the enzyme activity was reduced to 50% for 10 min and was calculated by fitting the residual activity at different temperatures to a Boltzmann sigmoidal function using the Origin software. The temperature dependence of AtTA and its mutants was determined by a standard assay from 20 to 60 °C.
Preparative synthesis of (R)-3-amino-1-butanol
For synthesis of (R)-3-amino-1-butanol, 1 g and 2.5 g 4-hydroxy-2-butanone were added into 50 mL potassium phosphate buffer (pH 7.5, 100 mM) containing 0.5 g lyophilized cells expressing AtTA-M5, 15 mg PLP, and 4.8 g isopropylamine hydrochloride. The bioconversion was performed by magnetic agitation at 35 °C for 48 h, and the pH was maintained at 7.5 using 2 M isopropylamine. WT AtTA was used as the control. The product was extracted with ethyl acetate at pH 10 (NaOH addition) and obtained in the form of oil after drying and distillation, which was then characterized by chiral HPLC and NMR (Fig. S4 and S5).
Validation of the S215P mutation in other (R)-selective ω-TAs
In order to confirm our findings from AtTA, we made mutations to residues of (R)-selective ω-TAs from other organisms that corresponded to Ser215 of AtTA (Fig. S6); whole plasmid PCR was used to introduce S215P mutation into (R)-selective ω-TAs from Aspergillus fumigatus (AfTA), Aspergillus oryzae (AoTA), Neosartorya fisheri (NfTA), Gibberella zeae (GzTA), and Arthrobacter sp. (ArTA), the P230S mutation into (R)-selective ω-TA from Mycobacterium vanbaalenii (MvTA), and the A213P mutation into (R)-selective ω-TA from Hyphomonas neptunium (HnTA). The recombinant plasmids carrying the genes encoding different (R)-selective ω-TAs were used as templates. The primers used in this experiment are listed in Table S1. The reaction mixture and PCR program were the same as those used for single-site saturation mutagenesis.
The catalytic efficiency of different WT enzymes and mutants was confirmed by the measurement of conversion. The reaction conditions were as follows: 50 mM acetophenone, 250 mM d-alanine, 1 mM PLP, 10 U mL−1d-amino acid dehydrogenase, 10 g L−1 lyophilized cells expressing (R)-selective ω-TA, 10 U mL−1 formate dehydrogenase, and 100 mM ammonium formate, 1 mM NAD+, 5% (v/v) DMSO, 100 mM potassium phosphate buffer (pH 7.5) at 35 °C for 24 h. The reaction mixture was extracted with ethyl acetate, and the conversion was determined by GC.
Results
Initial screening for appropriate enzyme scaffolds
To identify a suitable scaffold for the present work, a group of naturally occurring (R)-selective ω-TAs was screened using isopropylamine as the amine donor and 4-hydroxy-2-butanone as the amine acceptor. Among different enzymes, only AtTA and (R)-selective ω-TA from Arthrobacter sp. showed detectable activity, with conversion rates of 24.3% and 10.8% (Table S3), respectively. These two enzymes also exhibited excellent enantioselectivity (ee value > 99.9%). Considering the relatively high conversion, AtTA was selected as the initial scaffold for further protein engineering.
Exploring key residues of AtTA by substrate docking analysis
To more accurately identify the residues affecting the binding of 4-hydroxy-2-butanone in AtTA, a docking simulation was carried out using the crystal structure of AtTA as accepter and the quinonoid intermediate composed of prochiral ketone and PLP as ligand. The docking result indicated that the methyl group of 4-hydroxy-2-butanone was located at the small binding pocket, and the hydroxyl group on the other side was located at the large binding pocket. As shown in Fig. 1, two residues (His55 and Gly126) from chain B were located at the substrate tunnel, which may affect substrate accessibility. The residue Phe115 from chain A was located at the large binding pocket, which may result in disfavored interaction with the substrate. Another residue, Ser215 from chain A, was located at a β-turn close to the substrate, which may also affect the substrate binding orientation. These 4 residues were determined to likely participate in the interaction of the substrate with AtTA, and so were selected as the mutation hotspots of interest.
The docking structure of WT AtTA (PDB ID: 4CE5) and quinonoid intermediate. Monomer A and monomer B are colored in cyan and orange, respectively. The substrate tunnel and substrate binding pocket lie at the dimer interface. The hotspot residues are shown as yellow (Phe115 and Ser215 from chain A) and magenta (His55 and Gly126 from chain B) sticks. The quinonoid intermediate is shown as a green stick
Asymmetric reductive amination of 4-hydroxy-2-butanone by WT AtTA and its mutants. a The conversion of wild-type (WT) AtTA and positive single-site mutants at the substrate concentration of 60 mM. b The conversion of WT AtTA, AtTA-M1 (S215P), AtTA-M2 (H55S/S215P), AtTA-M3 (H55A/S215P), AtTA-M4 (S215P/G126F), and AtTA-M5 (H55A/S215P/G126F) at a substrate concentration of 120 mM. The optical purity of the (R)-3-amino-1-butanol produced by the various mutants was the same as that produced by WT AtTA
Reshaping the substrate binding region of AtTA
A saturation mutagenesis library comprising all 20 natural amino acids at each of these 4 positions was constructed. Subsequently, 96 colonies were picked up in the mutant library of each position in order to achieve 95% library coverage (Kille et al. 2013). The mutants showing higher conversion than that of WT AtTA were selected for DNA sequencing to identify the beneficial mutations. As shown in Fig. 2a, the conversion was improved significantly (1.9- to 3.0-fold higher than that of WT AtTA) when 55His was changed to residues with small side chain, such as Ser (3.0-fold), Ala (2.8-fold), Cys (2.6-fold), and Thr (1.9-fold). At position 126, the conversion of mutants G126P, G126Y, G126F, and G126S was 2.0- to 3.0-fold higher than WT AtTA. In addition, S215P and S215A improved the conversion 3.0- and 2.2-fold relative to WT AtTA, respectively. Nevertheless, none of the mutants at 115 had improved conversion. All of the above single-site mutants exhibited the same enantioselectivity as the WT AtTA (ee value > 99.9%).
To further improve the catalytic efficiency of AtTA for 4-hydroxy-2-butanone, a combination of positive mutations at the 3 positions was tested. On the basis of the mount of codons in the primers used for triple-site mutagenesis (Table S1), approximately 450 colonies (3-fold oversampling, 7 × 7 × 3 × 3 = 441) were screened in order to achieve 95% library coverage, and the most active single-site mutant AtTA-M1 (S215P) was used as the control. Several different mutants that exhibited significant improvement in the catalytic efficiency without any loss of enantioselectivity were identified by DNA sequencing. Corresponding mutation sites and conversion of the mutants are shown in Fig. 2b. The triple-site mutant AtTA-M5 (H55A/G126F/S215P) exhibited the best catalytic efficiency, with an impressive 9.4-fold and 2.1-fold increase in conversion compared with that of WT AtTA and AtTA-M1. In addition, 3 double-point mutants AtTA-M2 (H55S/S215P), AtTA-M3 (H55A/S215P), and AtTA-M4 (S215P/G126F) displayed 6.6-fold, 7.7-fold, and 6.2-fold increases in conversion relative to that of WT AtTA, respectively.
Characterization of WT AtTA and its mutants
The most active single-site (AtTA-M1), double-site (AtTA-M3), and triple-site mutants (AtTA-M5), along with WT AtTA, were purified and characterized in kinetic and thermostability experiments. As shown in Table 1, all 3 mutants showed higher catalytic efficiency (kcat/Km) toward 4-hydroxy-2-butanone and isopropylamine relative to WT AtTA. For 4-hydroxy-2-butanone, AtTA-M1 displayed an increase in kcat value to 0.53 S−1 compared with 0.17 S−1 of WT AtTA, and AtTA-M3 had a decrease in Km value to 0.28 mM compared with 0.46 mM of AtTA-M1. In addition, the AtTA-M5 exhibited a further decrease in Km value to 0.17 mM, and the final kcat/Km value was 9.8-fold higher than that of WT AtTA. In addition, a similar tendency was observed for isopropylamine, and the kcat/Km value of AtTA-M5 was 2.5-fold higher than that of WT AtTA.
For industrial applications, enzyme robustness is a vital factor. As shown in Fig. 3a, the \( {\mathrm{T}}_{1/2}^{10\mathit{\min}} \) value of WT AtTA was 37.8 °C. AtTA-M5 exhibited the best thermostability with a 9.4 °C higher \( {\mathrm{T}}_{1/2}^{10\mathit{\min}} \) compared with that of WT AtTA. AtTA-M1 and AtTA-M3 showed similar \( {\mathrm{T}}_{1/2}^{10\mathit{\min}} \) values, being 5.6 and 5.1 °C higher than that of WT AtTA, respectively. The determination of temperature-dependent activity (Fig. 3b) indicated that AtTA-M5 showed high activity between 40 and 55 °C and reached maximum activity at 50 °C which was 5 °C higher than WT AtTA (45 °C). The AtTA-M1 and AtTA-M3 exhibited the same optimal temperatures as WT AtTA but had better catalytic performance than WT AtTA at higher temperatures.
Effect of temperature on the stability and activity of WT AtTA and its mutants. a Determination of \( {\mathrm{T}}_{1/2}^{10\mathit{\min}} \) of WT, AtTA-M1 (S215P), AtTA-M3 (H55A/S215P), and AtTA-M5 (H55A/S215P/G126F) at different temperatures. The data points were fitted with a Boltzmann sigmoidal function. b Temperature-dependent activity of WT, AtTA-M1 (S215P), AtTA-M3 (H55A/S215P), and AtTA-M5 (H55A/S215P/G126F). The residual activity was recorded as a percentage of the maximum activity for each enzyme
Preparative synthesis of (R)-3-amino-1-butanol
To validate the practical potential of the mutant obtained in these studies, the most active mutant AtTA-M5, was used for asymmetric synthesis of (R)-3-amino-1-butanol on a preparative scale with WT AtTA as a control. As shown in Fig. 4, AtTA-M5 exhibited high conversion when the reaction was carried out for 48 h. The highest conversions reached 90.8% at a substrate concentration of 20 g L−1 and 79.1% at substrate concentration of 50 g L−1 while displaying excellent enantioselectivity (ee value > 99.9%). Under the same conditions, WT AtTA achieved less than 5% conversion. In the end, the purified product was confirmed to be (R)-3-amino-1-butanol by chiral HPLC and NMR analysis (Fig. S4 and S5).
Validation of the identified mutation in other (R)-selective ω-TAs
In the present work, the S215P mutation was present in all of the most active single-site, double-site, and triple-site mutants. In addition, the amino acid residue at position 215 was highly conserved among (R)-selective ω-TAs from different microbes. Herein, we introduced the mutation into 7 other (R)-selective ω-TAs from other species. As shown in Fig. 5, the corresponding mutants of ArTA, AfTA, AoTA, NfTA, GzTA, and HnTA exhibited higher conversion toward acetophenone relative to the WT enzymes, in good agreement with the results from AtTA. Similarly, in MvTA, the corresponding wild-type residue was originally Pro230, but the introduction of the P230S mutation reduced the conversion of acetophenone from 42.4 to 15.2%. These results indicated that proline in this position was a key residue for improving the catalytic efficiency of homologous (R)-selective ω-TAs.
Asymmetric reductive amination of acetophenone by different WT (R)-selective ω-TAs and single-site mutants. (R)-selective ω-TAs from Aspergillus fumigatus, Aspergillus oryzae, Neosartorya fisheri, Gibberella zeae, Mycobacterium vanbaalenii, Hyphomonas neptunium, and Arthrobacter sp. were abbreviated to AfTA, AoTA, NfTA, GzTA, MvTA, HnTA, and ArTA, respectively
Discussion
Methods for asymmetric reductive amination using ω-TAs have become a topic of significant interest in the field of chiral amine synthesis, and (R)-selective ω-TAs play an irreplaceable role in the efficient manufacturing of chiral amines, especially in the synthesis of (R)-amines. Asymmetric synthesis is the most convenient and economically favored route to produce a target chiral amine. Nevertheless, the downside of this strategy is the common requirement for an unfavorable reaction equilibrium (Seo et al. 2011). Several equilibrium displacement techniques have been established, such as enzymatic cascades in order to remove co-products, use of a large excess of the amine donor, and others. In practical applications, d-alanine and isopropylamine are the most common amine donors used for (R)-selective ω-TAs. When d-alanine is used, the d-amino acid dehydrogenase/formate dehydrogenase or lactate dehydrogenase/glucose dehydrogenase enzymatic cascades will always be introduced to recycle or remove pyruvate, a process which also requires the co-factor NADH (Pavlidis et al. 2016; Schätzle et al. 2011). In comparison with d-alanine, isopropylamine is a better amine donor for industrial asymmetric synthesis because it is cheap and achiral, and the by-product (acetone) can be easily removed from the reaction solution without additional enzymatic cascades (Dawood et al. 2018). Therefore, isopropylamine was selected as the amine donor for (R)-3-amino-1-butanol synthesis in this work.
Among different WT ω-TAs, AtTA was selected for molecular modification due to its relatively high base conversion of 4-hydroxy-2-butanone. We found that the crystal structure of AtTA was a homodimer, and the active center was located at the interface between the two subunits. The substrate had to enter the substrate binding pocket through a substrate tunnel (Fig. 1). The substrate binding pocket consisted of two binding pockets, a general feature of ω-TA. The large binding pocket could accommodate bulky or charged groups, such as aromatic rings, aliphatic chains, and carboxyl groups. On the contrary, the small binding pocket can only accommodate small groups such as methyl groups due to strict steric constraints. The substrate scope is restricted by this special architecture to a preference toward methyl ketone (Calvelage et al. 2017; Lyskowski et al. 2014; Nobili et al. 2015). Based on this structural analysis, docking simulations were carried out in this work. The quinonoid compound was the most energetically demanding intermediate in terms of its catalytic mechanism, and its planar character creates a significant steric hurdle, making the orientation of quinonoid intermediate binding in AtTA less variable. Therefore, the quinonoid intermediate was selected as the ligand for docking simulation. Additionally, considering that the small binding pocket of AtTA can accommodate the methyl group well, the determination of mutation hotspots in this work was limited to the substrate tunnel and the large binding pocket.
To reshape the substrate binding region, a mutant library containing mutations at different positions was constructed. The CASTing method has been shown to be a valuable means of constructing small focused saturation mutagenesis libraries. Herein, we constructed a “small-intelligent” library using a modified CASTing method based on single-site saturation mutagenesis to further reduce the screening library size. The beneficial mutations in each position were combined using degenerate primers designed by an MDC-Analyzer (Tang et al. 2014). Consequently, no more than 850 colonies were required for this work. The mutant AtTA-M5 displayed the greatest increase in catalytic efficiency (kcat/Km), which was 9.6-fold (4-hydroxy-2-butanone) and 2.5-fold (isopropylamine) higher than that of WT AtTA. Among the different mutations, H55A and G126F led to a lower Km, contributing to an increase in catalytic efficiency, and S215P resulted in a higher kcat and consequently higher catalytic efficiency. Meanwhile, G126F and S215P also made significant contributions to the enhanced thermostability of AtTA-M5.
To further investigate the possible mechanisms of improved catalytic efficiency and thermostability, we constructed a homology model of AtTA-M5. As shown in Fig. 6a and b, His55 was located at the inner end of the substrate tunnel near the bound substrate. The H55A mutation resulted in an obvious enlargement of the substrate tunnel by reducing the volume of the amino acid side chain, and consequently, the substrate was able to easily enter the substrate binding site. This also provided a good explanation for why the H55S/C/T mutants displayed higher conversion of 4-hydroxy-2-butanone. Another residue, Gly126, was located at the loop 121-136 from chain B, which formed the outer end of the substrate tunnel. A structural comparison (Fig. 6c) indicated that the loop 121-136 of AtTA-M5 displayed significantly different conformation from that of WT AtTA. Indeed, the backbone of loop 121-136 was moved outward due to the bulky side chain of Phe126, accompanied by the enlargement of the substrate tunnel. Meanwhile, the side chain of the nearby residue Arg128 was predicted to move significantly in the presence of the G126F mutation. The distance between the guanidine group of Arg128 and the carboxyl group of Glu151 was shortened from 5.8 to 2.8 Å. This alteration would be predicted to strengthen the electrostatic interactions between the side chains of Arg128 and Glu151 that can stabilize the loop 121-136. This result was consistent with the enhanced thermostability of AtTA-M5. It has been reported that the 121-136 loop was not conserved among different (R)-selective ω-TAs; however, the conformation alteration was already observed in (R)-selective ω-TA from Aspergillus fumigatus. In the “open” conformation, the loop moved outward and was stabilized by the hydrogen bond between Arg126 and Glu132 (corresponding to Arg128 and Glu134 of AtTA), accompanied by the broadened entrance to the active site (Skalden et al. 2015). Although the conformational change of loop 121-136 has not been reported in WT AtTA, the G126F mutation in this work could give rise to a similar “open” conformation of the loop. In (R)-selective ω-TA from Arthrobacter sp., the similar mutation G136F significantly reduced the catalytic efficiency toward pyruvate due to the bulky side chain of phenylalanine, which could cause the key residue Arg138 (important for pyruvate recognition) to move away from the substrate binding site; the loop 121-136 could be a hotspot region for altering the substrate specificity of (R)-selective ω-TAs (Guan et al. 2015). As for position 215, it was located at a β-turn in the active center. When the mutation S215P was introduced, an additional van der Waals interaction among the side chains of Leu187, Pro202, Pro215, and Phe217 was observed that probably contributed to improved thermostability (Fig. 6d). Meanwhile, proline was the most rigid residue that could be introduced to rigidify flexible regions (Yu et al. 2015). The structural stability of the β-turn could also be strengthened by this mutation. In addition, the residue Phe217 next to the β-turn was reported to participate in the PLP binding (Lyskowski et al. 2014). The strengthened structural stability of the β-turn will probably enhance the binding of PLP in AtTA, resulting in an improvement of the catalytic efficiency of the mutants with S215P. In the biocatalytic synthesis of sitagliptin intermediate, the corresponding mutation (S223P) has been shown to be able to increase the activity of ATA117 toward a methyl ketone analog (Savile et al. 2010). We tested the generalizability of this finding by mutating the corresponding proline residue in 8 homologous (R)-selective ω-TAs, and found significant improvements in the conversion of acetophenone.
Local environments of WT AtTA and AtTA-M5. a, b Enlargement of the substrate tunnel caused by the mutations of H55A and G126F. Stereo view of the surface is shown for WT AtTA (a) and AtTA-M5 (b). The locations of the 55th, 126th, and 128th residues are labeled in red, green, and blue, respectively. c Structural comparison of the loop 121-136 between WT AtTA and AtTA-M5. The loop121-136 of WT AtTA and AtTA-M5 are shown in cyan and orange, respectively. The side chain of Glu151 is shown in yellow, and the distances between Glu151 and Arg128 in WT AtTA (5.8 Å) and AtTA-M5 (2.8 Å) are indicated by the yellow dashed line. d Close-up of Pro215 in AtTA-M5. The van der Waals radii of the side chains of Leu187, Pro202, Pro215, and Phe217 are represented as yellow dots. The β-turn is shown in orange, and the quinonoid intermediate is indicated by the green stick
In conclusion, this work improved the catalytic efficiency and thermostability of AtTA toward 4-hydroxy-2-butanone without any loss of enantioselectivity. The screening procedure was facilitated by the construction of a “small-intelligent” library and the use of thin-layer chromatography for preliminary screening. Three key residues in the substrate tunnel and binding pocket of AtTA that impacted the interaction between 4-hydroxy-2-butanone and AtTA were identified. The best mutant, AtTA-M5, displayed significantly enhanced catalytic efficiency and thermostability toward 4-hydroxy-2-butanone, suggesting the potential for efficient production of (R)-3-amino-1-butanol. Moreover, the key mutation (S215P in AtTA) was further validated in 7 other (R)-selective ω-TAs, indicating its general applicability in improving the catalytic efficiency of homologous (R)-selective ω-TAs.
References
Aleku G, France S, Man H, Mangas-Sanchez J, Montgomery S, Sharma M, Leipold F, Hussain S, Grogan G, Turner N (2017) A reductive aminase from Aspergillus oryzae. Nat Chem 9(10):961–969. https://doi.org/10.1038/nchem.2782
Calvelage S, Dörr M, Höhne M, Bornscheuer UT (2017) A systematic analysis of the substrate scope of (S)- and (R)-selectiveamine transaminases. Adv Synth Catal 359(23):4235–4243. https://doi.org/10.1002/adsc.201701079
Chen F, Zheng G, Liu L, Li H, Chen Q, Li F, Li C, Xu J (2018) Reshaping the active pocket of amine dehydrogenases for asymmetric synthesis of bulky aliphatic amines. ACS Catal 8(3):2622–2628. https://doi.org/10.1021/acscatal.7b04135
Dawood AW, Weiß MS, Schulz C, Pavlidis IV, Iding H, de Souza RO, Bornscheuer UT (2018) Isopropylamine as amine donor in transaminase-catalyzed reactions: better acceptance through reaction and enzyme engineering. ChemCatChem 10(18):3943–3949. https://doi.org/10.1002/cctc.201800936
Desai AA (2011) Sitagliptin manufacture: a compelling tale of green chemistry, process intensification, and industrial asymmetric catalysis. Angew Chem Int Ed 50(9):1974–1976. https://doi.org/10.1002/anie.201007051
Gomm A, O’Reilly E (2018) Transaminases for chiral amine synthesis. Curr Opin Chem Biol 43:106–112. https://doi.org/10.1016/j.cbpa.2017.12.007
Guan L, Ohtsuka J, Okai M, Miyakawa T, Mase T, Zhi Y, Hou F, Ito N, Iwasaki A, Yasohara Y, Tanokura M (2015) A new target region for changing the substrate specificity of amine transaminases. Sci Rep 5:10753. https://doi.org/10.1038/srep10753
Guo F, Berglund P (2017) Transaminase biocatalysis: optimization and application. Green Chem 19(2):333–360. https://doi.org/10.1039/C6GC02328B
Höhne M, Schätzle S, Jochens H, Robins K, Bornscheuer UT (2010) Rational assignment of key motifs for function guides in silico enzyme identification. Nat Chem Biol 6(11):807–813. https://doi.org/10.1038/nchembio.447
Huang H, Liu X, Zhou L, Chang M, Zhang X (2016) Direct asymmetric reductive amination for the synthesis of chiral beta-arylamines. Angew Chem Int Ed 55(17):5309–5312. https://doi.org/10.1002/anie.201601025
Huang J, Xie D-F, Feng Y (2017) Engineering thermostable (R)-selective amine transaminase from Aspergillus terreus through in silico design employing B-factor and folding free energy calculations. Biochem Biophys Res Commun 483(1):397–402. https://doi.org/10.1016/j.bbrc.2016.12.131
Jiang J, Chen X, Zhang D, Wu Q, Zhu D (2015) Characterization of (R)-selective amine transaminases identified by in silico motif sequence blast. Appl Microbiol Biotechnol 99(6):2613–2621. https://doi.org/10.1007/s00253-014-6056-1
Johns BA, Kawasuji T, Weatherhead JG, Taishi T, Temelkoff DP, Yoshida H, Akiyama T, Taoda Y, Murai H, Kiyama R (2013) Carbamoyl pyridone HIV-1 integrase inhibitors 3. A diastereomeric approach to chiral nonracemic tricyclic ring systems and the discovery of dolutegravir (S/GSK1349572) and (S/GSK1265744). J Med Chem 56(14):5901–5916. https://doi.org/10.1021/jm400645w
Kelly S, Magill D, Megaw J, Skvortsov T, Allers T, McGrath J, Allen C, Moody T, Gilmore B (2019) Characterisation of a solvent-tolerant haloarchaeal (R)-selective transaminase isolated from a Triassic period salt mine. Appl Microbiol Biotechnol 103(14):5727–5737. https://doi.org/10.1007/s00253-019-09806-y
Khatik AG, Husain M (2018) Enzymatic process for the preparation of (R)-3-aminobutan-1-ol, useful in preparation of dolutegravir. WO patent, WO2018020380A1
Kille S, Acevedo-Rocha CG, Parra LP, Zhang Z-G, Opperman DJ, Reetz MT, Acevedo JP (2013) Reducing codon redundancy and screening effort of combinatorial protein libraries created by saturation mutagenesis. ACS Synth Biol 2(2):83–92. https://doi.org/10.1021/sb300037w
Kim E, Park J, Kim B, Seo J (2018) Identification of (R)-selective omega-aminotransferases by exploring evolutionary sequence space. Enzym Microb Technol 110:46–52. https://doi.org/10.1016/j.enzmictec.2017.12.002
Lyskowski A, Gruber C, Steinkellner G, Schurmann M, Schwab H, Gruber K, Steiner K (2014) Crystal structure of an (R)-selective omega-transaminase from Aspergillus terreus. PLoS One 9(1):e87350. https://doi.org/10.1371/journal.pone.0087350
Mathew S, Yun H (2012) ω-Transaminases for the production of optically pure amines and unnatural amino acids. ACS Catal 2(6):993–1001. https://doi.org/10.1021/cs300116n
Mutti FG, Fuchs CS, Pressnitz D, Sattler JH, Kroutil W (2011) Stereoselectivity of four (R)-selective transaminases for the asymmetric amination of ketones. Adv Synth Catal 353(17):3227–3233. https://doi.org/10.1002/adsc.201100558
Nobili A, Steffen-Munsberg F, Kohls H, Trentin I, Schulzke C, Höhne M, Bornscheuer UT (2015) Engineering the active site of the amine transaminase from Vibrio fluvialis for the asymmetric synthesis of aryl-alkyl amines and amino alcohols. ChemCatChem 7(5):757–760. https://doi.org/10.1002/cctc.201403010
Pavlidis I, Weiss M, Genz M, Spurr P, Hanlon S, Wirz B, Iding H, Bornscheuer U (2016) Identification of (S)-selective transaminases for the asymmetric synthesis of bulky chiral amines. Nat Chem 8(11):1076–1082. https://doi.org/10.1038/nchem.2578
Planchestainer M, Hegarty E, Heckmann CM, Gourlay LJ, Paradisi F (2019) Widely applicable background depletion step enables transaminase evolution through solid-phase screening. Chem Sci 10:5952–5958. https://doi.org/10.1039/C8SC05712E
Savile C, Janey J, Mundorff E, Moore J, Tam S, Jarvis W, Colbeck J, Krebber A, Fleitz F, Brands J, Devine P, Huisman G, Hughes G (2010) Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science 329(5989):305–309. https://doi.org/10.1126/science.1188934
Schätzle S, Steffen-Munsberg F, Thontowi A, Höhne M, Robins K, Bornscheuer UT (2011) Enzymatic asymmetric synthesis of enantiomerically pure aliphatic, aromatic and arylaliphatic amines with (R)-selective amine transaminases. Adv Synth Catal 353(13):2439–2445. https://doi.org/10.1002/adsc.201100435
Seo J-H, Kyung D, Joo K, Lee J, Kim B-G (2011) Necessary and sufficient conditions for the asymmetric synthesis of chiral amines using w-aminotransferases. Biotechnol Bioeng 108(2):253–263. https://doi.org/10.1002/bit.22930
Sheldon R, Pereira P (2017) Biocatalysis engineering: the big picture. Chem Soc Rev 46(10):2678–2691. https://doi.org/10.1039/c6cs00854b
Skalden L, Thomsen M, Hohne M, Bornscheuer U, Hinrichs W (2015) Structural and biochemical characterization of the dual substrate recognition of the (R)-selective amine transaminase from Aspergillus fumigatus. FEBS J 282(2):407–415. https://doi.org/10.1111/febs.13149
Tang L, Wang X, Ru B, Sun H, Huang J, Gao H (2014) MDC-Analyzer: a novel degenerate primer design tool for the construction of intelligent mutagenesis libraries with contiguous sites. Biotechniques 56(6):301–302, 304, 306-8, 310. https://doi.org/10.2144/000114177
Tang X, Zhang N, Ye G, Zheng Y (2019) Efficient biosynthesis of (R)-3-amino-1-butanol by a novel (R)-selective transaminase from Actinobacteria sp. J Biotechnol 295:49–54. https://doi.org/10.1016/j.jbiotec.2019.02.008
Telzerow A, Paris J, Håkansson M, González-Sabín J, Ríos-Lombardía N, Schürmann M, Gröger H, Morís F, Kourist R, Schwab H (2018) Amine transaminase from Exophiala Xenobiotica-crystal structure and engineering of a fold IV transaminase that naturally converts biaryl ketones. ACS Catal 9(2):1140–1148. https://doi.org/10.1021/acscatal.8b04524
Xie DF, Yang J, Lv C, Mei J, Wang H, Hu S, Zhao W, Cao J, Tu J, Huang J, Mei L (2019) Construction of stabilized (R)-selective amine transaminase from Aspergillus terreus by consensus mutagenesis. J Biotechnol 293:8–16. https://doi.org/10.1016/j.jbiotec.2019.01.007
Yamada Y, Iwasaki A, Kizaki N, Ogura M, Hasegawa J (2001) Process for producing optically active amino compounds. US patent, US6221638
Yu H, Zhao Y, Guo C, Gan Y, Huang H (2015) The role of proline substitutions within flexible regions on thermostability of luciferase. BBA-Proteins Proteom 1854(1):65–72. https://doi.org/10.1016/j.bbapap.2014.10.017
Funding
This work was financially supported by the Natural Science Fund for Colleges and Universities in Jiangsu Province (18KJB416006 and 17KJB350011), the Natural Science Foundation of Jiangsu Province (BK20160575), the National Natural Science Foundation of China (81703363), the Scientific Research Starting Foundation of Taizhou University (TZXY2017QDJJ008), and the Social Development Program of Taizhou (TS201916).
Author information
Authors and Affiliations
Contributions
Xinxing Gao and Xin Zhang performed the experiments. Xinxing Gao, Nianqing Zhu, and Yi Mou conceived and designed the research. Hailing Zhang and Xin Liu analyzed the data and supervised the projected. Xinxing Gao wrote the initial draft of the paper, and Xin Zhang and Pinghe Wei revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key points
• A “small-intelligent” library was constructed to reduce the screening library size.
• Thin-layer chromatography was applied for high-throughput preliminary screening.
• AtTA-M5 displayed significantly enhanced catalytic efficiency and thermostability.
• The identified mutation was validated in other homologous (R)-selective ω-TAs.
Electronic supplementary material
ESM 1
(PDF 767 kb)
Rights and permissions
About this article
Cite this article
Gao, X., Zhang, X., Zhu, N. et al. Reshaping the substrate binding region of (R)-selective ω-transaminase for asymmetric synthesis of (R)-3-amino-1-butanol. Appl Microbiol Biotechnol 104, 3959–3969 (2020). https://doi.org/10.1007/s00253-020-10539-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10539-6